Chromosomal translocations in non-hodgkin lymphomas
|
|
|
- Arnold Parks
- 5 years ago
- Views:
Transcription
1 Chromosomal Translocations Involved in Non-Hodgkin Lymphomas Francisco Vega, MD, PhD; L. Jeffrey Medeiros, MD Context. The discovery that recurrent chromosomal translocations are involved in the pathogenesis of non- Hodgkin lymphomas has greatly improved our understanding of these diseases and revolutionized their diagnosis. Objective. To review the mechanisms by which chromosomal translocations occur and contribute to the pathogenesis of various types of non-hodgkin lymphomas and to review the utility of molecular genetic methods for the assessment of these translocations. Data Sources and Study Selection. Primary research studies and reviews published in the English language that focus on chromosomal translocation and non-hodgkin lymphomas. Data Extraction and Synthesis. Chromosomal translocations, which usually result in oncogene activation, occur in many types of B- and T-cell lymphoma, and their detection is helpful for establishing an accurate diagnosis and monitoring disease following therapy. However, the precise mechanisms that explain how translocations occur remain unknown, although for some types of translocations a clear relationship has been established with immunoglobulin gene rearrangement mechanisms. In recent years, a number of genes deregulated by chromosomal translocations have been identified, and the detailed molecular mechanisms by which chromosomal translocations contribute to the pathogenesis of non-hodgkin lymphoma are beginning to be elucidated. Conclusions. Molecular genetic analysis has played a major role in improving our understanding of B- and T-cell non-hodgkin lymphomas and has allowed more precise definition of lymphoma types. Molecular genetic tests to detect these translocations are important ancillary tools for the diagnosis and classification of malignant lymphomas. (Arch Pathol Lab Med. 2003;127: ) Chromosomal translocations in non-hodgkin lymphomas can be subdivided into 2 general types. In the first type, an intact oncogene (or intact coding region) is juxtaposed via the translocation with another gene, usually an antigen receptor gene. As a result, the oncogene becomes transcriptionally deregulated. Historically, this was the first type of translocation discovered in non- Hodgkin lymphomas. Examples of this type include the t(8;14) in Burkitt lymphoma/leukemia and the t(14;18) in follicular lymphoma. In the second type of translocation, 2 genes (usually non antigen receptor genes) are disrupted, and portions of each gene are juxtaposed, resulting in a fusion gene, chimeric mrna, and a novel protein. This type of chromosomal translocation occurs in some types of non-hodgkin lymphoma, such as the t(2;5) in anaplastic large cell lymphoma (ALCL), but is best known in acute and chronic myeloid leukemias. MECHANISMS OF CHROMOSOMAL TRANSLOCATIONS Overall, the precise molecular mechanisms underlying chromosomal translocation remain largely unknown, and different mechanisms have been proposed to explain recurrent translocations in various lymphoma types. In some cases, the evidence suggests that translocation is a Accepted for publication February 19, From the Division of Pathology and Laboratory Medicine, University of Texas M. D. Anderson Cancer Center, Houston. Reprints: L. Jeffrey Medeiros, MD, Department of Hematopathology, University of Texas M. D. Anderson Cancer Center, Box 72, 1515 Holcombe Blvd, Houston, TX ( jmedeiro@mdanderson.org). result of aberrant V(D)J recombinase activity, because cryptic heptamer/nonamer consensus sequences are present in the immediate vicinity of the translocation breakpoints. 1,2 The antigen receptor loci of B cells may be subjected to 4 types of modification: recombination of the variable (V), diversity (D), and joining (J) regions, somatic hypermutation of the V segments, immunoglobulin heavy-chain (IgH) gene class switching, and receptor editing. Occasional failures in the control of these processes appear to play an important role in the generation of chromosomal translocations in B-cell lymphomas. These events occur mainly at 2 stages of B-cell development: during V region recombination in B-cell precursors in the bone marrow and during B-cell differentiation in the germinal center microenvironment. Chromosome translocations in T-cell lymphomas also may arise from analogous errors of T-cell receptor (TCR) gene V(D)J recombination. 2 However, somatic hypermutation of V segments is rare, and class switching of the TCR genes does not occur in T cells. V(D)J Recombination The development of B cells in the bone marrow is initiated by a site-specific recombination reaction, V(D)J recombination, of the IgH andig light-chain genes ( and ). 3 For formation of the V region exon of the IgH chain, V, D, and J gene segments are joined, whereas the V region exons of and light chains are composed only of V and J segments. 3,4 As shown for IgH in Figure 1, first a D H segment is joined with a J H segment. This joining is fol Arch Pathol Lab Med Vol 127, September 2003 Chromosomal Translocations in Lymphomas Vega & Medeiros
2 Figure 1. IgH gene rearrangement. The IgH chain gene in its germline configuration has approximately 120 variable (V) regions, up to 30 diversity (D) regions, 6 joining (J) regions, and 9 constant (C) regions. Each of these regions are discontinuous. Three gene segments (V, D, and J) are joined to create a heavy-chain V region gene. This process occurs in 2 steps: D H joins to J H and then V H joins to D H J H. Figure 2. Coding end processing. A, The endonuclease activity of the RAG protein complexes makes 2 single-strand DNA breaks at sites just 5 of each recognition signal sequence (RSS), and a DNA hairpin is created. B, The DNA hairpin is nicked open by a single-strand break, also by the RAG proteins. The nicking can happen at various points along the hairpin, which leads to sequence variability in the eventual joint. When nicks occur on only one strand, the resulting protruding strand ends with palindromic nucleotides coming from the opposite strand. C, These nucleotide additions are termed P nucleotides and are a characteristic of variable, diversity, joining (VDJ) recombination because they are the direct result of DNA hairpin resolution. D, Once coding ends are opened up, more modifications can take place. Nucleotides may be removed by exonuclease activity, and nongermline nucleotides (N sequences) may be added by the terminal deoxynucleotidyl transferase (TdT), further increasing the diversity of V region genes. E and F, Pairing of strands, exonuclease cleavage, DNA synthesis, and ligation occur to form the coding joint. The sum of those modifications of the coding ends before religation (P and N addition and nucleotide deletion) is referred to as coding end processing and constitutes the hallmark of the VDJ recombination process. Figure 3. The npm-alk fusion protein resulting from the t(2;5)(p23;q35). The major breakpoint region to the right of the transmembrane domain is indicated by a vertical arrow. OD indicates oligomerization domain; MB, metal-binding domain; AD, acidic amino acid clusters; NLS, nuclear localization signals; LBS, putative ligand binding site for pleiotropin; TM, transmembrane domain; TK, tyrosine kinase catalytic domain. Figure 4. The t(8;14) breakpoints in sporadic and endemic Burkitt lymphoma (BL). In sporadic BL, the breakpoints occur near the c-myc gene and in the switch (S) region of IgH (yellow arrows). Exon 1, including the major promoters of c-myc, is deleted; however, the coding regions (exons 2 and 3) remain structurally intact. The coding regions are indicated in white, and the noncoding regions are gray. In endemic BL, the breakpoints are located more than 100 kb 5 of the first exon of c-myc and in the joining (J) region of IgH (blue arrows). The chromosome 8 breakpoints of the variant translocations (arrowhead) occur 3 to the gene. Restriction fragment length (Southern blot) analysis can be used to detect c-myc gene rearrangements in sporadic BL but not in endemic BL. The reverse is true for detecting IgH rearrangements using genomic J region probes. lowed by a V H to D H J H joining, which can be either in frame (correct for encoding antibody sequences) or out of frame. When in-frame VDJ joining is achieved, and the precursor cell is now a B cell that can proceed with maturation by undergoing Ig light-chain gene rearrangements. Following out-of-frame VDJ joining, a second attempt at VDJ joining occurs on the other allele. Recombination involves the introduction of doublestrand breaks in DNA by the products of the recombination-activating genes, RAG-1 and RAG-2. 5 These breaks Arch Pathol Lab Med Vol 127, September 2003 Chromosomal Translocations in Lymphomas Vega & Medeiros 1149
3 occur at specific recognition signal sequences (RSSs) adjacent to the V, D, and J elements (3 to the V regions, 5 and 3 to the D regions, and 5 to the J regions). An RSS is composed of conserved heptamer and nonamer motifs separated by a relatively nonconserved spacer region of either 12 or 13 base pairs (bp), and efficient recombination requires one RSS of each type. Several other proteins are needed in this process, including DNA ligase IV; DNAdependent protein kinase; Ku, which is a heterodimer (Ku 70:Ku 80) that associates tightly with DNA-dependent protein kinase; and the DNA-repair protein, x-ray repair complementing defective in Chinese hamster 4 (XRCC4). 6 The characteristic steps in the V(D)J recombination process are illustrated in Figure 2. Defects during V(D)J recombination, during the phases in which physiological breaks are generated or in which they are joined, can result in chromosomal translocations. 7,8 At least 2 distinct mechanisms involving V(D)J recombination can lead to translocation. First, an illegitimate V(D)J recombination may occur between a legitimate Ig or TCR locus and an illegitimate proto-oncogene locus bearing a functional RSS. Translocations that involve antigen receptor gene loci in precursor T-cell lymphoblastic lymphoma/leukemia, such as the t(10;14)(q24;q11) involving the TCR / locus, seem to be mediated by such a mechanism. 9,10 In a second mechanism, only the Ig or TCR locus breaks are mediated by V(D)J recombination. Breaks at the locus bearing the proto-oncogene are initiated by other mechanisms of cleavage. The t(11;14)(q13;q32) in mantle cell lymphoma appears to be an example of this second mechanism. 9,11 Several models have been proposed to explain the occurrence of chromosomal breaks in proto-oncogene loci. Translocations may occur through elements such as crossover hotspot instigator (chi)-like sequences, which promote recombination. In agreement with this hypothesis, chi-like sequences are found within the 2 predominant breakpoint regions of the bcl-2 locus of the t(14;18) and the major translocation cluster of the bcl-1 locus of the t(11; 14). 12 Another possible mechanism has been attributed to the transpositional activity of the RAG proteins. 7,8 Somatic Hypermutation Somatic hypermutation is a process by which mutations, mainly single-nucleotide exchanges or small insertions or deletions of DNA, are introduced at a high rate into the V segments of normal germinal center B cells after exposure to antigen. 13 These mutations require double-stranded DNA breaks prior to the mutational event and thus increase the likelihood that translocations may occur. 13 Others have suggested that translocations involving the c- myc gene in endemic Burkitt lymphoma (BL) probably arise as by-products of somatic hypermutation. 13 In addition, some translocations of the bcl-6 gene in diffuse large B-cell lymphomas (DLBCL) and of the IgG FcIIb receptor in follicular lymphoma show translocation breakpoints in somatically mutated V region genes. 14,15 IgH Switching IgH switching is a process by which B cells normally replace the IgM andigd constant regions with IgG, IgA, or IgE. This process is mediated by a recombination event that occurs in germinal center B cells after antigen exposure and results in deletion of DNA between involved switch (S) regions. These S regions are arrays of short tandem repeats located upstream of each constant-region gene (with the exception of C ). The switch results in a change in the effector functions of the antibody but leaves the V(D)J region unaltered. For several types of B-cell lymphoma, chromosomal translocations into IgH chain S regions have been described, such as translocations involving c-myc in sporadic BL, 16 bcl-3 [t(14;19)] in B-cell chronic lymphocytic leukemia (CLL), 17 bcl-6 [t(3;14)] in DLBCL, 18 muc1 [t(1;14)] in large cell lymphoma, 19 and fgfr3 [t(4;14)], c-maf [t(14;16)], and mum/irf4 [t(t6;14)] in plasma cell myeloma (PCM) Receptor Editing Receptor editing is a process by which one of the expressed antibody polypeptide light chains, or, is replaced by the other. The process of Ig light-chain loci receptor editing is mediated by secondary rearrangements of the V region gene, usually involving upstream V segments and downstream J segments. This process also involves DNA-strand breaks and may play a role in the generation of chromosomal translocations in B-cell lymphomas. MECHANISMS OF ONCOGENE FUSION The molecular mechanisms by which 2 genes are disrupted resulting in the formation of a novel hybrid gene remain largely unknown. In the cases of the t(11;18) and t(2;5), breakpoints in the api2 andalk genes, respectively, are clustered, suggesting that structural features make these breakpoints sites (introns) more vulnerable to rearrangement. Different molecular mechanisms can be implicated, such as homologous recombination mediated by Alu elements, 23 TRANSLIN activity, 24 cleavage at purine/ pyrimidine repeat regions, 25 topoisomerase II subunit exchange, 26 and repair of DNA breaks with nonhomologous chromosomes. 27 In the t(11;18), Baens and colleagues 28 found AluSx repeat sequences within or close to the breakpoint region in intron 7 of api2. However, no AluSx sequences were consistently observed close to the breakpoint sites in malt1 on chromosome 18. Thus, homologous recombination between Alu elements as a mechanism for generating the t(11;18) seems unlikely. Sato and colleagues 29 found a V(D)J heptamer consensus sequence close to the breakpoint in intron 7 of api2 and several heptamer-like sequences in the malt1 gene. However, no nonamer-like sequences were found in either the api2 ormalt1 genes. One cannot exclude the possibility that the clustering of breakpoints delineates the portions of the proteins required in the fusion for its oncogenic properties. With this explanation, one must hypothesize that illegitimate random recombinations can occur during the normal cell division process. Random cleavage of the genes can be followed by attempts to repair the breaks and joining of the loci, and those clones containing the fusion transcript survive and have a growth advantage. Thus, the breakpoints appear to be clustered. However, in the case of t(11;18) and t(2;5), this explanation seems unlikely. 29,30 COMMON TRANSLOCATIONS IN NON-HODGKIN LYMPHOMAS ALCL of T- or Null-Cell Lineage ALCL accounts for approximately 3% of adult and 10% to 30% of childhood lymphomas and includes a subset of tumors that carry the t(2;5)(p23;q35). 31 This translocation 1150 Arch Pathol Lab Med Vol 127, September 2003 Chromosomal Translocations in Lymphomas Vega & Medeiros
4 Table 1. Translocation t(2;5)(p23;q35) t(1;2)(q25;p23) t(2;3)(p23;q21) inv(2)(p23q35) t(x;2)(q11-12;p23) t(2;17)(p23;q23) Chromosomal Translocations in Anaplastic Large Cell Lymphoma Involved Genes* alk and npm tpm3 and alk alk and tfg alk and atic msn and alk alk and cltcl * alk indicates anaplastic lymphoma kinase; npm, nucleophosmin; tpm3, tropomyosin 3; tfg, TRK-fused gene; atic, 5-aminoimidazole-4- carboxamide ribonucleotide formyltransferase/imp cyclohydrolase; msn, moesin; and cltcl, clathrin heavy chain like. has been identified in 40% to 70% of ALCLs of the T-cell and null-cell types, most commonly in tumors that occur in younger patients. Structure of the Translocation. The t(2;5) fuses the nucleophosmin (NPM) gene, npm, at 5q35 with the anaplastic lymphoma kinase (ALK) gene, alk, at 2p23. The fusion gene encodes a 80-kd chimeric protein, of which 117 NH 2 - terminal residues of NPM are linked to the intracytoplasmic portion of ALK (C-terminal residues ) (Figure 3). 30,32 The reciprocal alk npm fusion gene is not transcribed at significant levels and thus probably does not contribute to the pathogenesis of ALCL. 32 Other rare translocations involving the alk locus at 2p23 that also result in alk overexpression have been characterized (Table 1). All except one of the variant fusions cloned thus far have had the same breakpoint within the alk gene,ina 1935-bp intron located between the exons encoding the transmembrane and juxtamembrane domains of alk, suggesting that this chromosomal region is very susceptible to breakage. The exception is the alk breakpoint in the moesin (msn) alk translocation, which is localized in an exonic sequence 17 bp downstream of the 5 end of the first alk exon. Pathophysiology. The fusion of npm with alk encodes a constitutively activated tyrosine kinase that is a potent oncogene. 33 The causative role of this fusion protein in lymphomagenesis was confirmed by the development of lymphomas in mice transplanted with bone marrow cells transduced with human NPM ALK fusion protein. 34 Motifs within NPM regulate dimerization of NPM ALK, inducing conformational changes in the ALK component, resulting in a constitutive activation of the kinase domain. 33 In all the variant 2p23 abnormalities, the NPM protein is replaced by another protein that contains some form of oligomerization motif, suggesting that the mechanism underlying ALK activation is similar in all and that the signal transduction pathways activated are probably identical. Although the pathogenesis of ALCL is unknown, recent in vitro studies have provided evidence that several mechanisms, including ras, signal transducer and activator of transcription (STAT), phospholipase C, and phosphatidylinositol 3 kinase pathways, may be involved in deregulation of cell proliferation and apoptosis in ALCLs. 35 Rassidakis and colleagues 36 have shown that ALK-positive ALCLs have low levels of the antiapoptotic protein Bcl-2 and a higher apoptotic rate than ALK-negative ALCLs. Thus, differential expression of Bcl-2 family proteins may play a role in the good response to chemotherapy observed in patients with ALK-positive ALCLs. ALK-positive ALCLs appear to be clinically homogeneous, regardless of their morphology or whether they have the t(2;5) or other translocations involving alk. Based on the expression of ALK, several investigators have proposed restricting the definition of ALCL to include only ALK-positive tumors, leaving ALK-negative cases in a separate category, most likely anaplastic variants of peripheral T-cell lymphoma. Methods of Detection. In most pathology laboratories, ALK immunostaining, given its ease, rapidity, specificity, and low cost, is the most useful screening technique for detecting abnormalities involving the alk locus. 35 ALK is normally expressed in nerve cells but is not expressed in normal hematopoietic tissues. However, as a result of translocations or other abnormalities involving alk, ALK is overexpressed. Thus, immunostaining with antibodies specific for the cytoplasmic portion of ALK provides indirect evidence for abnormalities involving the alk gene, most often the t(2;5). ALK expression can be detected immunohistochemically with the monoclonal antibody ALK- 1 or with polyclonal antibodies (eg, p80, ALK-11). Malignant cells carrying the t(2;5) have both a cytoplasmic and nuclear pattern of staining with anti-alk antibodies because of dimerization of NPM ALK with wild-type NPM. The latter has nuclear localization motifs. 37 In some studies, 15% to 20% of ALK-positive ALCLs show only cytoplasmic expression of ALK, suggesting the presence of a variant translocation not involving npm. The lack of tight clustering within the involved npm and alk introns precludes analysis using standard polymerase chain reaction (PCR) methods, but reverse transcription (RT) or long-range PCR methods can be used. 38 These methods can detect the t(2;5) but not other variant translocations involving alk. RT-PCR requires RNA and generates amplicons of identical size, making it difficult to exclude contamination if it occurs. Long-range PCR has the advantages that gene transcription is not required and amplicons of distinct size for each case are generated. However, high-quality DNA is necessary for this approach. Conventional cytogenetic analysis provides the most complete information because these methods allow detection of the t(2;5) and all variant translocations involving alk. However, viable cells that can be induced to divide are required, which is a major drawback. Fluorescence in situ hybridization (FISH) methods can be used to detect either the t(2;5) or alk rearrangements, the latter representing either t(2;5) or variant translocations involving alk. Restriction fragment length analysis using genomic probes derived from either alk or npm also can be used to detect gene rearrangements. However, this approach is labor intensive and is seldom used routinely. Burkitt Lymphoma/Leukemia Burkitt lymphoma/leukemia is a highly aggressive B- cell neoplasm characterized by chromosomal translocations involving the c-myc gene. 39 These translocations occur in virtually all cases. The t(8;14) is most common, found in approximately 80% to 85% of cases. The t(2;8) and t(8;22), also known as the variant translocations, occur in approximately 10% to 15% of tumors. Structure of the Translocation. The t(8;14) juxtaposes the c-myc gene, normally at 8q24, 5 to the IgH locus on the derivative chromosome 14. In the t(2;8) and t(8;22), the Ig and Ig light-chain loci are juxtaposed 3 to the c-myc gene on the derivative chromosome 8. The breakpoints on chromosome 8 are variable and widely scattered (Figure Arch Pathol Lab Med Vol 127, September 2003 Chromosomal Translocations in Lymphomas Vega & Medeiros 1151
5 4). For example, in endemic (African) BL, the breakpoints are usually located far 5 ( 100 kilobases [kb]) to the c- myc gene. In contrast, in sporadic BL the chromosome 8 breakpoints occur immediately 5 to or within the c-myc gene. Similarly, the breakpoints on chromosome 14 are variable. In endemic BL, most breakpoints occur within the J region of IgH. In sporadic BL, the breakpoints occur more commonly within the S or constant (C) regions, upstream of C, C, orc (Figure 4). 40 Pathophysiology. As a result of the t(8;14) or variant translocations, Myc protein is overexpressed. Myc influences multiple cellular processes, including cell proliferation, differentiation, apoptosis, and metabolism, that contribute positively or negatively to cell transformation. The c-myc gene is one of the immediate early growth response genes that are rapidly induced in quiescent cells upon mitogenic induction, playing an important role in mediating the transition from quiescence to proliferation. Myc induces expression of cyclins D1 and D2, allowing these molecules to form complexes with cyclin-dependent kinase inhibitors. These cyclin D cyclin-dependent kinase complexes sequester the cell cycle inhibitors p27 and p21, thereby releasing the cell cycle from inhibition. 41 The c-myc gene is also able to stimulate apoptosis (programmed cell death), and abrogation of Myc-induced apoptosis is probably necessary to allow its proliferation effect. Concomitant activation of bcl-2 or alterations in the p19(arf)-mdm2-p53 pathway can nullify the apoptotic influence of c-myc. 42 These cooperative effects have been observed in cell culture. When normal cells are placed in medium with low concentrations of growth factors, such as platelet-derived growth factor, they are arrested in the G0 or G1 stages of the cell cycle but remain viable. Recombinant cells that overexpress c-myc also arrest their growth under these conditions but soon undergo apoptosis. Apparently, the cell senses an inappropriate growth signal from Myc and, in the absence of other growth signals from surface receptors, commits suicide. However, overexpression of bcl-2 rescues c-myc overexpressing cells from death. Thus, a cell with abundant Myc and Bcl-2 proteins can proliferate in the absence of normal growth factors. Methods of Detection. The c-myc gene potently drives cell division, and metaphases are easily detectable in cell cultures. As a result, the t(8;14) or variant translocations are usually detected by conventional cytogenetic methods. However, viable cells are required. FISH methods are advantageous because cell division and thus viable cells are not required, and the large size of c-myc probes allows detection of virtually all 8q24 breakpoints, as a result of the t(8;14), t(2;8), and t(8;22). 43 If c-myc and Ig gene probes are used, the specific translocation can be identified. The number of partner chromosomes and the great variability in breakpoints makes PCR assays impractical. A large number of primer pairs would be needed, and even then all translocations cannot be detected. Restriction fragment length can be used to assess c-myc gene rearrangements in sporadic BL. However, in endemic BL the c-myc breakpoints extend beyond the length of DNA fragments that can be detected using genomic c-myc probes (Figure 4). 40 In sporadic BL, a similar approach using J region probes to assess the IgH gene often do not detect rearrangements because the breakpoints occur in the S or C regions. In contrast, IgH gene rearrangements can be detected in endemic BL because the breakpoints occur in the J region (Figure 4). Chronic Lymphocytic Leukemia/Small Lymphocytic Lymphoma Chronic lymphocytic leukemia/small lymphocytic lymphoma (CLL/SLL) represents approximately 6% to 7% of non-hodgkin lymphomas in the United States. A number of molecular abnormalities have been detected in CLL/ SLL, but their roles in pathogenesis have yet to be elucidated. Using conventional cytogenetic analysis, chromosomal abnormalities have been identified in 50% to 60% of CLL/ SLL cases, including trisomy 12, abnormalities of 13q14, deletions of 11q, and abnormalities of 14q32. FISH analysis reveals that approximately 80% of the cases have abnormal karyotypes. 44,45 Deletion 13q14 is the most common abnormality in CLL/SLL, occurring in up to 50% of cases. Trisomy 12 is found in 10% to 20% of cases, and 11q deletions occur in 20% of CLL/SLL cases. 46 Chromosomal translocations occur in only a small subset of CLL/SLL cases. In approximately 5% of cases, the bcl-2 gene at 18q21 is involved in either the t(2;18)(p12; q21) or t(18;22)(q21;q11), involving the Ig (chromosome 2p12) or Ig (chromosome 22q11) gene loci. 46 In these translocations, the chromosome 18 breakpoints occur 5 to the first exon of bcl-2. Other chromosomal translocations identified uncommonly in CLL/SLL include the t(14;19) and the t(2;14). The t(14;19)(q32,q13) is a rare cytogenetic abnormality that is found in less than 5% of cases of CLL/SLL and is associated with young age at time of initial diagnosis, atypical cytologic features, and poor prognosis. 47 The t(14; 19) juxtaposes the bcl-3 gene at 19q13 with IgH in a headto-head configuration, resulting in overexpression of bcl- 3. Restriction fragment length analysis, using genomic bcl- 3 probes, is the standard method for detection of bcl-3 gene rearrangements. 47 The t(2;14)(p13;q32) is a rarely described abnormality identified in uncommon tumors that has been designated as childhood CLL. The translocation breakpoints for 2 such cases have been cloned. 48 The t(2;14) juxtaposes the bcl-11a gene at 2p13 with IgH at 14q32. The bcl-11a gene is the human homologue of mouse evi-9, which is deregulated in murine myeloid leukemias after proviral integration. The t(2;14) may be associated with the Ig-unmutated subset of CLL/SLL. 49 Diffuse Large B-Cell Lymphoma DLBCL is a molecularly heterogeneous group of B-cell lymphomas in which conventional cytogenetic analysis has shown a wide variety of molecular abnormalities (Table 2). Two of the more common translocations include those involving bcl-6 and the t(14;18). The t(14;18) is present in 20 30% of DLBCL cases. These tumors are thought to be of follicle center origin and may represent transformation of follicular lymphoma (FL) presenting as de novo DLBCL. Other chromosomal translocations identified in DLBCL include the t(14;15)(q32;q11 13), t(1;22)(q22;q11), t(1;14)(q21;q32), and t(10;14)(q24;q32) and involve the genes bcl-8, fcgriib, muc-1, and nf- b2, respectively. Approximately 4% of DLBCL carry the t(14;15) or bcl-8 gene rearrangements; the other translocations are less frequent. These translocations do not result in fusion proteins but rather in the juxtaposition of oncogenes with Ig gene loci Arch Pathol Lab Med Vol 127, September 2003 Chromosomal Translocations in Lymphomas Vega & Medeiros
6 Table 2. Chromosomal Translocations in Diffuse Large B-Cell Lymphoma Translocation t(14;18)(q32;q21) t(3;4)(q27;p13) t(3;6)(p27;p21.3) t(3;6)(p27;p21.2) t(3;7)(p27;p12) t(3;11)(p27;q23.1) t(3;3)(p27;q29) t(3;13)(q27;q14) t(3;16)(p27;p11) t(3;12)(p27;12q ) t(3;18)(p27;p11.2) t(3;14)(p27;q32) t(3;6)(p27;p12) t(3;16)(p27;p13) t(3;14)(p27;q32) t(3;2)(p27;p12) t(3;22)(p27;q11) t(14;15)(q32;q11-13) t(1;22)(q22;q11) t(1;14)(q21;q32) t(10;14)(q24;q32) t(1;13)(p32;q14) t(1;7)(q21;q22) der(6)t(6;8)(q11;q11) t(5;16)(?;q11-q12) t(19;22)(q13;q11-q13) Involved Genes IgH and bcl-2 bcl-6 and rhoh/ttf bcl-6 and H4 histone bcl-6 and pim-1 bcl-6 and ikaros bcl-6 and bob/obf1 bcl-6 and tfrr bcl-6 and 1-plastin bcl-6 and Il-21r bcl-6 and -nac bcl-6 and eif4aii bcl-6 and hsp89 bcl-6 and hsp90 bcl-6 and ciita bcl-6 and IgH bcl-6 and Ig bcl-6 and Ig IgH and bcl-8 fcgriib and Ig muc-1 and IgH nf- b2 and IgH Recently, Nanjangud and collaborators 50 used spectral karyotyping to identify new recurrent chromosomal translocations in DLBCL, including the t(1;13)(p32;q14), t(1; 7)(q21;q22), and der(6)t(6;8)(q11;q11) and potentially the t(5;16)(?;q11-q12) and t(19;22)(q13;q11-q13). The genes at these loci are not currently known. Structure and Pathophysiology of bcl-6. Rearrangements of the bcl-6 gene are found in 30% to 40% of DLBCL cases, approximately 20% of acquired immunodeficiency syndrome associated DLBCL, and 5% to 14% of FL cases. 51 The bcl-6 gene resides at the 3q27 locus, spans 26 kb, and has 10 exons that encode the Bcl-6 protein consisting of 706 amino acids. 52,53 This gene is highly promiscuous and is translocated with a number of partner chromosome loci (Table 2). The IgH gene locus is most commonly involved, in 40% to 50% of cases. The bcl-6 breakpoints are concentrated in a 4-kb region of 3q27 that includes the promoter and the first noncoding exon. In most cases, the breakpoints are located immediately 3 to the first exon, and as a result bcl-6 exons 2 10 (spanning the entire coding domain) are juxtaposed to a partner chromosome downstream from heterologous promoters (Figure 5). The bcl-6 gene encodes a zinc-finger transcription factor implicated in germinal center B-cell differentiation, antibody-affinity maturation, and T helper cell mediated responses. 53 Translocations involving bcl-6 do not involve its coding regions and thus result in overexpression of structurally intact Bcl-6 protein. The Bcl-6 protein is normally expressed by germinal center B cells, is required for normal germinal center formation, and is downregulated in post germinal center stages of B-cell maturation. Downregulation of bcl-6 during plasma cell differentiation allows expression of B lymphocyte induced maturation protein 1, a major regulator of plasma cell differentiation. B lymphocyte induced maturation protein 1 represses transcription of c-myc, and in B cells that are poised to differentiate to plasma cells, this drop in c-myc expression is an integral part of terminal differentiation. 54 Loss of normal control mechanisms regulating bcl-6 expression is thought to contribute to malignant transformation in germinal center derived B cells. Point mutations of the regulatory portion of the bcl-6 gene have been detected frequently in germinal center and post germinal center lymphomas, including FL, DLBCL, and BL, and their significance in lymphomagenesis is not clear. One study has suggested that the prognostic impact of bcl-6 translocations might depend on whether the gene is juxtaposed with Ig loci or other genes; the cases with non- Ig partner chromosomes have a poorer outcome. 55 In another study, high levels of bcl-6 gene expression were correlated with a favorable prognosis in patients with DLBCL. 56 In one study, DLBCL cases with non Ig/bcl-6 translocations had higher levels of bcl-6 mrna than cases with Ig/bcl-6 translocations. 57 Methods of Detection. Restriction fragment length analysis is used most often because bcl-6 breakpoints are relatively clustered, usually in the first intron or immediately 5 to the first exon, and thus bcl-6 gene rearrangements can be readily assessed with a bcl-6 probe. 52 FISH probes specific for bcl-6 should produce results comparable to those obtained with restriction fragment length analysis and are less labor intensive. 58 The telomeric location of 3q27 makes translocations involving this locus difficult to recognize by conventional cytogenetic techniques. The large number of partner chromosomes precludes routine PCR analysis in the clinical laboratory. Follicular Lymphoma Follicular lymphoma is a neoplasm of follicle center B cells and is characterized by the t(14;18)(q32;q21), detected by conventional cytogenetic techniques in approximately 80% to 90% of cases. 59 In the remaining 10% to 20% of FL cases, the t(14;18) is not detectable using any available method. This observation suggests the existence of other mechanisms that dysregulate bcl-2 expression. The t(14;18) is also found in 20% to 30% of DLBCL cases. These tumors are presumably of follicle center cell origin. 60 Structure of the Translocation. In the t(14;18), the bcl- 2 gene on 18q21 is juxtaposed with the IgH gene on 14q32, resulting in overexpression of structurally intact and functional Bcl-2 protein. The breakpoints on chromosome 14 are tightly clustered, occurring immediately 5 to the IgH joining regions. The majority of breakpoints on chromosome 18 also tightly clustered. Two well-known clusters, the major breakpoint cluster region (MBR) and the minor breakpoint cluster region (MCR) (Figure 6), 61 are involved in 50% to 60% and 10% to 15% of the cases of FL, respectively. Recently, Albinger-Hegyi and collaborators 62 sequenced an approximately 25-kb region of chromosome 18 between the MBR and MCR and detected a third breakpoint cluster region, which they designated the intermediate cluster region. 62 Other small breakpoint cluster regions have been reported, including a region 5 to the first exon of bcl-2 (also involved in 5% of cases of CLL/SLL). Pathophysiology. The t(14;18) has been identified in a significant proportion of healthy persons. 63 In the absence of FL (by all measurable parameters), the significance of this finding is uncertain but may suggest that the translocation is an early initiating event, with subsequent cumulative oncogenic alterations necessary for manifestation Arch Pathol Lab Med Vol 127, September 2003 Chromosomal Translocations in Lymphomas Vega & Medeiros 1153
7 Figure 5. The (3;14)(q27;q32) translocation. Most of the breakpoints are located in the first intron or immediately 5 to the first exon of the bcl- 6 gene and cause the juxtaposition of bcl-6 exons 2 10, corresponding to the entire coding domain, downstream from the promoter derived from chromosome 14. The breakpoints in chromosome 14 occur within the switch (S) region of IgH. Thus, the functional consequence of this translocation is the deregulation of bcl-6 expression by promoter substitution. Figure 6. Germline configuration of chromosome 18q21 region involved in t(14;18)(q32;q21). Most of the breakpoints occur in the major breakpoint region (MBR) with a smaller subset in the minor breakpoint cluster region (MCR) (yellow arrowheads). The black, green, orange, and white rectangles represent the range of detection using polymerase chain reaction, Southern blot (SB), fluorescent in situ hybridization, and conventional cytogenetic methods, respectively. Figure 7. Germline configuration of chromosome 11q13 region involved in t(11;14)(q13;q32). The ccnd-1 gene is indicated as a green box. The major translocation cluster region accounts for 30% to 40% of all translocations. Minor breakpoint clusters are indicated as mtc1 and mtc2. The black, green, orange, and white rectangles represent the range of detection using polymerase chain reaction, Southern blot (SB), fluorescent in situ hybridization, and conventional cytogenetic methods, respectively. Figure 8. The api2 and malt1 genes involved in t(11;18)(q21;q21). All breakpoints occur between exons 7 and 8 of the api2 gene (arrowhead), 3 to the 3 baculovirus inhibitor of apoptosis repeat domain (BIR) motifs. Large arrows indicate the position of the different malt1 breakpoints; small arrows indicate the alternative splice sites used when a breakpoint upstream of 814 base pairs occurs. CARD indicates caspase recruitment domain. The RING finger motif is a distinct zinc-chelating domain involved in mediating protein DNA and protein protein interactions. of FL. Lower incidence rates of FL have been reported in most Asian countries. However, available published data suggest that the incidence of the t(14;18) in healthy persons in the absence of FL is similar in Western and Asian populations. 63,64 The exact mechanisms by which Bcl-2 and related family members regulate apoptosis are not clear, and results of some studies have indicated that bcl-2 expression alone is not sufficient to explain its antiapoptotic activity, as can be observed in cell culture experiments. Low-grade FL cells carrying the t(14;18) undergo apoptosis when cultured in vitro, meaning that FL cells are not capable of self-maintenance and lack proper humoral and/or cellular factors that allow their survival. Therefore, other factors or regulatory mechanisms must be involved in FL cell survival. Independent of its antiapoptotic function, Bcl-2 and other Bcl-2 family members can also promote cell-cycle arrest, retarding entry into the cell cycle and accelerating withdrawal from the cell cycle. 65,66 Thus, Bcl-2 appears to have a physiological role in influencing the transition between quiescent and cycling states. Both the antiapoptotic and 1154 Arch Pathol Lab Med Vol 127, September 2003 Chromosomal Translocations in Lymphomas Vega & Medeiros
8 the cell-cycle arrest functions of Bcl-2 may explain the indolent nature of FL. Inhibition of apoptosis and cell-cycle arrest may lead to an accumulation of malignant cells over time, without increasing the proliferation rate of the affected cells. Methods of Detection. The tight physical clustering of the MBR and MCR breakpoints facilitates effective screening for the presence of the t(14;18) translocation by standard PCR assays, which are commonly used as tests of choice, particularly in the assessment of minimal residual disease. Using 2 sets of primers specific for the MBR and MCR, PCR can be used to detect the t(14;18) in approximately 60% to 75% of cases. 67 This method is also advantageous because assays for MBR and MCR can be applied to archival paraffin-embedded tissue. Several investigators have demonstrated the utility and reliability of using real-time PCR methods to detect t(14;18) and to quantify tumor burden in FL patient samples. 68,69 However, standard or real-time PCR assays will not detect a subset of cases that can be detected using other methods. At our hospital, the prognosis is thought to be poorer in patients with FL in which the t(14;18) is not detected by the standard PCR approach assessing MBR and MCR. 70 A higher detection rate for the t(14;18) may be obtained by analysis of the intermediate cluster region, which also can be assessed by standard PCR methods. 62 Detection rates higher than those obtained by standard PCR can be achieved by using long-range PCR, which can detect additional breakpoints. 62 However, this technique requires high-quality DNA and has not yet been implemented as a routine detection method in most laboratories. FISH methods using bcl-2 and IgH probes also have been used to detect the t(14;18). These methods have the potential to detect virtually all cases with the t(14;18). 71 Restriction fragment length analysis can be used to detect PCR positive cases as well as a subset of cases of FL in which the breakpoints do not occur in the MBR and MCR (Figure 6). Disadvantages of restriction fragment length analysis include the need for chromosome 14 and 18 probes to prove the presence of t(14;18) and the time-consuming and laborious nature of this method. Mantle Cell Lymphoma Mantle cell lymphoma (MCL) is a clinically aggressive B-cell lymphoma that represents approximately 6% of all non-hodgkin lymphomas in the United States. An almost constant feature in this neoplasm is the presence of the t(11;14)(q13;q32). 72 Structure of the Translocation. The t(11;14) juxtaposes the ccnd-1 gene (also known as bcl-1, prad1, and cyclin D1) at 11q13 with an enhancer of the IgH gene at 14q32. Nearly half of these translocations cluster within an 80- to 100- bp region known as the major translocation cluster region on chromosome 11q13 (Figure 7). 73,74 However, the remainder of chromosome 11 breakpoints are widely scattered over approximately 120 kb. The breakpoints on chromosome 14 occur 5 to 1 of 6 joining regions of IgH. Pathophysiology. As a result of the t(11;14), the ccnd- 1 gene is overexpressed. 72 Overexpression of cyclin D1 protein in the G1 phase of the cell cycle causes early phosphorylation of retinoblastoma protein and acceleration through G1. 75,76 Although cyclin D1 overexpression is sufficient to confer transformed properties on fibroblasts, it is insufficient to transform primary cells or to induce lymphomagenesis in E -cyclin D1 transgenic mice. 77 However, concurrent overexpression of cyclin D1 and either the n- myc or l-myc oncogenes in transgenic mice results in lymphoma formation. 78 Thus, other cellular factors in addition to ccnd-1 overexpression appear to play a role in MCL. Various proteins, including the E2F5 transcription factor, its dimerization partner dp2, the cdc28 protein kinase 1, and the products of the cell cycle genes ckd4, mdm2, c-myc, cdc8, and skp2 have been reported to be overexpressed in MCL. 79 In addition, several pathways associated with apoptosis are altered and substantially downregulated in MCL. Methods of Detection. FISH methods using probes for chromosomes 11 and 14 are very sensitive and can detect the t(11;14) in up to 95% of MCLs. 80,81 Conventional cytogenetic analysis can detect the t(11;14) in at least 70% to 80% of cases (Figure 7). Slow growth, sampling error, or other technical limitations may explain the negative results in 20% to 30% of tumors. Translocations involving the major translocation cluster are amenable to routine PCR analysis, and thus 30% to 40% of MCL cases are positive using these methods. 82 However, the large subset of negative cases makes this approach problematic for routine diagnosis. Restriction fragment length analysis, using at least 4 genomic chromosome 11 probes, can detect rearrangements of the bcl-1 locus in approximately 65% to 75% of MCLs. 73,74 However, this approach is time consuming and labor intensive, and chromosome 14 probes are also required to prove the presence of the t(11;14). An alternative approach to obtain support for the diagnosis of MCL is to assess the expression of cyclin D1, because all t(11;14) translocations theoretically result in overexpression of cyclin D1. Because normal lymphocytes express very low levels of cyclin D1, not detectable by immunohistochemical methods, these methods are convenient and specific for distinguishing MCL from other types of non-hodgkin lymphoma. However, a negative immunoreaction does not exclude the diagnosis of MCL because the immunohistochemical approach is relatively insensitive and some MCLs have relatively low levels of cyclin D1. 72 Detection of cyclin D1 mrna by quantitative RT-PCR, in contrast, is very sensitive and distinguishes low levels of cyclin D1 derived from nonoverexpressing lymphomas or normal cells from high levels of cyclin D1 mrna characteristic of MCL. 83 Marginal Zone B-Cell Lymphoma of Extranodal Mucosa-Associated Lymphoid Tissue Type Both types of chromosomal translocations (described earlier) have been implicated in the pathogenesis of extranodal marginal zone B-cell lymphoma of mucosa-associated lymphoid tissue (MALT). 84 In the t(11;18)(q21; q21), the api2 gene on 11q21, and the malt1 gene on chromosome 18q21 are disrupted and recombine to form a novel api2 malt1 fusion gene on the derivative chromosome 11. In contrast, in the other translocations reported in MALT lymphoma, listed in Table 3, the entire coding region of a gene (malt1, bcl-10) is juxtaposed with an antigen receptor gene. t(11;18)(q21;q21). The t(11;18) is the most frequent chromosomal translocation known to occur in MALT lymphoma, identified in at least one third of cases. The t(11; 18)-positive lymphomas have been reported to involve most extranodal MALT sites, most commonly the gastrointestinal tract and lung. The t(11;18) appears to be specific for MALT lymphomas and has not been identified in Arch Pathol Lab Med Vol 127, September 2003 Chromosomal Translocations in Lymphomas Vega & Medeiros 1155
9 other lymphoma types, including nodal and splenic marginal zone B-cell lymphomas Structure of the Translocation. The api2 gene is a member of the inhibitor of apoptosis protein family and contains 3 baculovirus inhibitor of apoptosis protein repeat (BIR) domains, a caspase recruitment domain (CARD), and a RING finger domain (Figure 8). The common domain of the inhibitor of apoptosis protein family is the BIR motif, which plays an essential role in inhibiting apoptosis. All breakpoints in the api2 gene occur between exons 7 and 8, 3 to the 3 BIR motifs. Thus, the BIR motif is considered essential to the function of api2 malt1. The malt1 gene is a novel gene that has homology with the Ig-like domain of CD22, the laminin-5 3b subunit, and F22D3.6 of Caenorhabditis elegans. 87 The C-terminal part of malt1 harbors 2 Ig-like C2 domains and an Ig VDJ4-like sequence. Four different breakpoints occur throughout the malt1 gene, all of which result in the 3 end of the gene being fused in frame with api2 (Figure 8). Pathophysiology. It is hypothesized that fusion of api2 to the C-terminal region of malt1 leads to an increased inhibition of apoptosis, inhibiting the biological activity of caspases 3, 7, and 9 and thereby conferring a survival advantage. The function of the malt1 gene is largely unknown. The api2 malt1 protein recently has been shown to activate nf- b, a transcription factor for several survivalrelated genes. 88 Recent studies suggest that response of gastric MALT lymphoma to antibiotic therapy can be predicted by the presence or absence of the t(11;18). 89,90 Lymphomas that carry the t(11;18) are unlikely to respond to antibiotic therapy. By contrast, those lymphomas negative for the t(11; 18) are more often associated with immunologic stimulation mediated by Helicobacter pylori and are more likely to respond to antibiotic therapy. To date, the t(11;18) only rarely has been detected in high-grade B-cell lymphomas arising in MALT sites. Thus, t(11;18)-positive low-grade MALT lymphomas seem to have a low likelihood of transforming to large B-cell lymphoma. Because low-grade MALT lymphomas are commonly associated with a highgrade component and both components have been shown to be clonally related in some patients, 91,92 there may be 2 types of low-grade MALT lymphoma: one not associated with the t(11;18) that may transform to high-grade lymphoma and the other that carries the t(11;18) and rarely transforms to high-grade lymphoma. Alternatively, the translocation may be not present in all tumor cells, implicating the presence of subclones with or without the translocation, the latter likely to transform to a high-grade lymphoma. Although the possibility that the t(11;18) is lost from the clone during tumor progression cannot be excluded, this possibility seems unlikely because most other types of low-grade B-cell lymphoma that undergo highgrade transformation retain their initial molecular abnormalities while acquiring additional genetic alterations. Methods of Detection. Cytogenetic analysis of MALT lymphomas is hampered by low yield and poor quality of metaphase spreads. These problems are related in part to small size of biopsy specimens, the slow growth of these tumors, and bacterial contamination of biopsy specimens from some extranodal sites, such as the gastrointestinal tract. RT-PCR methods used to detect the t(11;18) have great sensitivity and may be applied for evaluation of minimal residual disease. A number of different transcripts of various sizes have been detected using these methods. Table 3. Chromosomal Translocations in Marginal Zone B-Cell Lymphoma of Extranodal Mucosa- Associated Lymphoid Tissue (MALT) Type Translocation t(11;18)(q21;q21) t(14;18)(q32;q21) t(1;14)(p22;q32) t(1;2)(p22;p12) * api indicates apoptosis inhibitor. Involved Genes api2 and malt1 malt1 and IgH bcl-10 and IgH bcl-10 and Ig Baensetal 93 also developed a reliable real-time RT-PCR assay to detect the t(11;18), based on the 5 3 exonuclease activity of Taq polymerase. A major disadvantage of this approach is the need for fresh-frozen tissue specimens to allow extraction of RNA. Interphase FISH assays using api2- and malt1-specific probes can be applied to fresh and archival tissues or nuclei. 86 FISH is easy to interpret, nearly as sensitive as RT-PCR, and highly specific and may detect translocations of malt1 involving partner genes other than api2 as well as translocations of api2 involving partner genes other than malt1. t(14;18)(q32;q21). The t(14;18)(q32;q21) has been recently identified in approximately 20% of MALT lymphomas. 94 In this translocation, the malt1 gene is juxtaposed with the IgH gene at 14q32, resulting in malt1 overexpression. To date, the t(14;18) in MALT lymphomas has been identified by conventional cytogenetics and FISH methods. The t(11;18) and t(14;18) appear to be mutually exclusive in MALT lymphomas. 94 t(1;14)(p22;q32). The t(1;14)(p22;q32) has been identified rarely in MALT lymphomas. 95 As a result of the t(1; 14), the entire bcl-10 gene, at chromosome 1p22, is juxtaposed with the IgH gene, resulting in Bcl-10 overexpression. 96 Another rare chromosomal translocation involving bcl-10 also has been identified recently in MALT lymphoma, the t(1;2)(q22;p12). 97 In this translocation, the bcl-10 gene is juxtaposed with the Ig gene on 2p12 (Table 3). Structure of the Translocation. The bcl-10 gene has 4 exons within an 11.7-kb segment of the genome. The gene encodes a protein of 233 amino acids, with residues forming a CARD similar to that of api2, whereas 132 amino acids at the C-terminus of the protein contain no known motifs. 96 Pathophysiology. The bcl-10 and IgH genes on the derivative chromosome 14 are oriented in a head-to-head fashion. As a result, bcl-10 comes under the control of the IgH gene enhancers resulting in overexpression of Bcl Wild-type bcl-10 encodes a protein containing an NH 2 -terminal CARD that weakly promotes apoptosis, activates nf- b, and behaves as a tumor suppressor gene in in vitro assays. 96,98 The t(1;14) truncates the bcl-10 gene at its C- terminus, 3 to the CARD motif. Truncated Bcl-10 protein loses its proapoptotic function but retains its ability to activate nf- b. Mutations of bcl-10 also occur infrequently, in about 5% of MALT lymphomas. 99 Recently, Maes et al 100 reported that bcl-10 genomic mutations do not play an important role in the pathogenesis or the progression of gastric MALT lymphomas. Methods of Detection. Because the t(1;14) is rare, its presence is not assessed routinely with molecular diagnostic methods. Conventional cytogenetic analysis is the main method of detection. Expression of Bcl-10 protein can be detected by immunohistochemistry, and in a study using these methods Bcl-10 was found in the cytoplasm 1156 Arch Pathol Lab Med Vol 127, September 2003 Chromosomal Translocations in Lymphomas Vega & Medeiros
GENETIC MARKERS IN LYMPHOMA a practical overview. P. Heimann Dpt of Medical Genetics Erasme Hospital - Bordet Institute
 GENETIC MARKERS IN LYMPHOMA a practical overview P. Heimann Dpt of Medical Genetics Erasme Hospital - Bordet Institute B and T cell monoclonalities Rearrangement of immunoglobin and TCR genes may help
GENETIC MARKERS IN LYMPHOMA a practical overview P. Heimann Dpt of Medical Genetics Erasme Hospital - Bordet Institute B and T cell monoclonalities Rearrangement of immunoglobin and TCR genes may help
Malignant lymphomas are neoplasms that arise from B
 Overview of the Role of Molecular Methods in the Diagnosis of Malignant Lymphomas L. Jeffrey Medeiros, MD; Jeanne Carr, PhD Objective. To review the role of molecular genetics in the diagnosis of malignant
Overview of the Role of Molecular Methods in the Diagnosis of Malignant Lymphomas L. Jeffrey Medeiros, MD; Jeanne Carr, PhD Objective. To review the role of molecular genetics in the diagnosis of malignant
Non-Hodgkin lymphomas (NHLs) Hodgkin lymphoma )HL)
 Non-Hodgkin lymphomas (NHLs) Hodgkin lymphoma )HL) Lymphoid Neoplasms: 1- non-hodgkin lymphomas (NHLs) 2- Hodgkin lymphoma 3- plasma cell neoplasms Non-Hodgkin lymphomas (NHLs) Acute Lymphoblastic Leukemia/Lymphoma
Non-Hodgkin lymphomas (NHLs) Hodgkin lymphoma )HL) Lymphoid Neoplasms: 1- non-hodgkin lymphomas (NHLs) 2- Hodgkin lymphoma 3- plasma cell neoplasms Non-Hodgkin lymphomas (NHLs) Acute Lymphoblastic Leukemia/Lymphoma
Molecular Pathology of Lymphoma (Part 1) Rex K.H. Au-Yeung Department of Pathology, HKU
 Molecular Pathology of Lymphoma (Part 1) Rex K.H. Au-Yeung Department of Pathology, HKU Lecture outline Time 10:00 11:00 11:15 12:10 12:20 13:15 Content Introduction to lymphoma Review of lymphocyte biology
Molecular Pathology of Lymphoma (Part 1) Rex K.H. Au-Yeung Department of Pathology, HKU Lecture outline Time 10:00 11:00 11:15 12:10 12:20 13:15 Content Introduction to lymphoma Review of lymphocyte biology
Immunopathology of Lymphoma
 Immunopathology of Lymphoma Noraidah Masir MBBCh, M.Med (Pathology), D.Phil. Department of Pathology Faculty of Medicine Universiti Kebangsaan Malaysia Lymphoma classification has been challenging to pathologists.
Immunopathology of Lymphoma Noraidah Masir MBBCh, M.Med (Pathology), D.Phil. Department of Pathology Faculty of Medicine Universiti Kebangsaan Malaysia Lymphoma classification has been challenging to pathologists.
Introduction. Introduction. Lymphocyte development (maturation)
 Introduction Abbas Chapter 8: Lymphocyte Development and the Rearrangement and Expression of Antigen Receptor Genes Christina Ciaccio, MD Children s Mercy Hospital January 5, 2009 Lymphocyte development
Introduction Abbas Chapter 8: Lymphocyte Development and the Rearrangement and Expression of Antigen Receptor Genes Christina Ciaccio, MD Children s Mercy Hospital January 5, 2009 Lymphocyte development
Molecular Diagnosis. Nucleic acid based testing in Oncology
 Molecular Diagnosis Nucleic acid based testing in Oncology Objectives Describe uses of NAT in Oncology Diagnosis, Prediction, monitoring. Genetics Screening, presymptomatic testing, diagnostic testing,
Molecular Diagnosis Nucleic acid based testing in Oncology Objectives Describe uses of NAT in Oncology Diagnosis, Prediction, monitoring. Genetics Screening, presymptomatic testing, diagnostic testing,
Nucleic Acid Testing - Oncology. Molecular Diagnosis. Gain/Loss of Nucleic Acid. Objectives. MYCN and Neuroblastoma. Molecular Diagnosis
 Nucleic Acid Testing - Oncology Molecular Diagnosis Nucleic acid based testing in Oncology Gross alterations in DNA content of tumors (ploidy) Gain/Loss of nucleic acids Markers of Clonality Oncogene/Tumor
Nucleic Acid Testing - Oncology Molecular Diagnosis Nucleic acid based testing in Oncology Gross alterations in DNA content of tumors (ploidy) Gain/Loss of nucleic acids Markers of Clonality Oncogene/Tumor
Objectives. Morphology and IHC. Flow and Cyto FISH. Testing for Heme Malignancies 3/20/2013
 Molecular Markers in Hematologic Malignancy: Ways to locate the needle in the haystack. Objectives Review the types of testing for hematologic malignancies Understand rationale for molecular testing Marcie
Molecular Markers in Hematologic Malignancy: Ways to locate the needle in the haystack. Objectives Review the types of testing for hematologic malignancies Understand rationale for molecular testing Marcie
Differential diagnosis of hematolymphoid tumors composed of medium-sized cells. Brian Skinnider B.C. Cancer Agency, Vancouver General Hospital
 Differential diagnosis of hematolymphoid tumors composed of medium-sized cells Brian Skinnider B.C. Cancer Agency, Vancouver General Hospital Lymphoma classification Lymphoma diagnosis starts with morphologic
Differential diagnosis of hematolymphoid tumors composed of medium-sized cells Brian Skinnider B.C. Cancer Agency, Vancouver General Hospital Lymphoma classification Lymphoma diagnosis starts with morphologic
Non-Hodgkin s Lymphoma: Molecular Features of B Cell Lymphoma
 Non-Hodgkin s Lymphoma: Molecular Features of B Cell Lymphoma Elizabeth Macintyre (Chair), Dennis Willerford, and Stephan W. Morris The rapid increase in the incidence of the B cell non-hodgkin s lymphomas
Non-Hodgkin s Lymphoma: Molecular Features of B Cell Lymphoma Elizabeth Macintyre (Chair), Dennis Willerford, and Stephan W. Morris The rapid increase in the incidence of the B cell non-hodgkin s lymphomas
Haematology Probes for Multiple Myeloma
 Haematology Probes for Multiple Myeloma MULTIPLE MYELOMA Multiple myeloma (MM) is a plasma cell neoplasm, characterised by the accumulation of clonal plasma cells in the bone marrow and by very complex
Haematology Probes for Multiple Myeloma MULTIPLE MYELOMA Multiple myeloma (MM) is a plasma cell neoplasm, characterised by the accumulation of clonal plasma cells in the bone marrow and by very complex
The Development of Lymphocytes: B Cell Development in the Bone Marrow & Peripheral Lymphoid Tissue Deborah A. Lebman, Ph.D.
 The Development of Lymphocytes: B Cell Development in the Bone Marrow & Peripheral Lymphoid Tissue Deborah A. Lebman, Ph.D. OBJECTIVES 1. To understand how ordered Ig gene rearrangements lead to the development
The Development of Lymphocytes: B Cell Development in the Bone Marrow & Peripheral Lymphoid Tissue Deborah A. Lebman, Ph.D. OBJECTIVES 1. To understand how ordered Ig gene rearrangements lead to the development
Classification of Hematologic Malignancies. Patricia Aoun MD MPH
 Classification of Hematologic Malignancies Patricia Aoun MD MPH Objectives Know the basic principles of the current classification system for hematopoietic and lymphoid malignancies Understand the differences
Classification of Hematologic Malignancies Patricia Aoun MD MPH Objectives Know the basic principles of the current classification system for hematopoietic and lymphoid malignancies Understand the differences
MOLECULAR PATHOGENESIS OF NON-HODGKIN LYMPHOMA: A CLINICAL PERSPECTIVE
 molecular basis of disease Haematologica 1995; 80:454-472 MOLECULAR PATHOGENESIS OF NON-HODGKIN LYMPHOMA: A CLINICAL PERSPECTIVE Gianluca Gaidano, Cristina Pastore, Gisella Volpe Laboratorio di Medicina
molecular basis of disease Haematologica 1995; 80:454-472 MOLECULAR PATHOGENESIS OF NON-HODGKIN LYMPHOMA: A CLINICAL PERSPECTIVE Gianluca Gaidano, Cristina Pastore, Gisella Volpe Laboratorio di Medicina
Generation of antibody diversity October 18, Ram Savan
 Generation of antibody diversity October 18, 2016 Ram Savan savanram@uw.edu 441 Lecture #10 Slide 1 of 30 Three lectures on antigen receptors Part 1 : Structural features of the BCR and TCR Janeway Chapter
Generation of antibody diversity October 18, 2016 Ram Savan savanram@uw.edu 441 Lecture #10 Slide 1 of 30 Three lectures on antigen receptors Part 1 : Structural features of the BCR and TCR Janeway Chapter
Activation of cellular proto-oncogenes to oncogenes. How was active Ras identified?
 Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Role of FISH in Hematological Cancers
 Role of FISH in Hematological Cancers Thomas S.K. Wan PhD,FRCPath,FFSc(RCPA) Honorary Professor, Department of Pathology & Clinical Biochemistry, Queen Mary Hospital, University of Hong Kong. e-mail: wantsk@hku.hk
Role of FISH in Hematological Cancers Thomas S.K. Wan PhD,FRCPath,FFSc(RCPA) Honorary Professor, Department of Pathology & Clinical Biochemistry, Queen Mary Hospital, University of Hong Kong. e-mail: wantsk@hku.hk
The development of clonality testing for lymphomas in the Bristol Genetics Laboratory. Dr Paula Waits Bristol Genetics Laboratory
 The development of clonality testing for lymphomas in the Bristol Genetics Laboratory Dr Paula Waits Bristol Genetics Laboratory Introduction The majority of lymphoid malignancies belong to the B cell
The development of clonality testing for lymphomas in the Bristol Genetics Laboratory Dr Paula Waits Bristol Genetics Laboratory Introduction The majority of lymphoid malignancies belong to the B cell
7 Omar Abu Reesh. Dr. Ahmad Mansour Dr. Ahmad Mansour
 7 Omar Abu Reesh Dr. Ahmad Mansour Dr. Ahmad Mansour -Leukemia: neoplastic leukocytes circulating in the peripheral bloodstream. -Lymphoma: a neoplastic process in the lymph nodes, spleen or other lymphatic
7 Omar Abu Reesh Dr. Ahmad Mansour Dr. Ahmad Mansour -Leukemia: neoplastic leukocytes circulating in the peripheral bloodstream. -Lymphoma: a neoplastic process in the lymph nodes, spleen or other lymphatic
Methods used to diagnose lymphomas
 Institut für Pathologie Institut für Pathologie Methods used to diagnose lymphomas Prof. Dr.Med. Leticia Quintanilla-Fend Molecular techniques NGS histology Cytology AS-PCR Sanger seq. MYC Immunohistochemistry
Institut für Pathologie Institut für Pathologie Methods used to diagnose lymphomas Prof. Dr.Med. Leticia Quintanilla-Fend Molecular techniques NGS histology Cytology AS-PCR Sanger seq. MYC Immunohistochemistry
Chapter 11. B cell generation, Activation, and Differentiation. Pro-B cells. - B cells mature in the bone marrow.
 Chapter B cell generation, Activation, and Differentiation - B cells mature in the bone marrow. - B cells proceed through a number of distinct maturational stages: ) Pro-B cell ) Pre-B cell ) Immature
Chapter B cell generation, Activation, and Differentiation - B cells mature in the bone marrow. - B cells proceed through a number of distinct maturational stages: ) Pro-B cell ) Pre-B cell ) Immature
Chapter 11. B cell generation, Activation, and Differentiation. Pro-B cells. - B cells mature in the bone marrow.
 Chapter B cell generation, Activation, and Differentiation - B cells mature in the bone marrow. - B cells proceed through a number of distinct maturational stages: ) Pro-B cell ) Pre-B cell ) Immature
Chapter B cell generation, Activation, and Differentiation - B cells mature in the bone marrow. - B cells proceed through a number of distinct maturational stages: ) Pro-B cell ) Pre-B cell ) Immature
Molecular Diagnostics in the Workup of Lymphomas for the General Pathologist. Outline. Role of Molecular Testing in Lymphoma Diagnosis
 Molecular Diagnostics in the Workup of Lymphomas for the General Pathologist L. Jeffrey Medeiros, M.D. M.D. Anderson Cancer Center Outline Gene rearrangement studies / Clonality Translocations / mutations
Molecular Diagnostics in the Workup of Lymphomas for the General Pathologist L. Jeffrey Medeiros, M.D. M.D. Anderson Cancer Center Outline Gene rearrangement studies / Clonality Translocations / mutations
Chapter 4 Cellular Oncogenes ~ 4.6 -
 Chapter 4 Cellular Oncogenes - 4.2 ~ 4.6 - Many retroviruses carrying oncogenes have been found in chickens and mice However, attempts undertaken during the 1970s to isolate viruses from most types of
Chapter 4 Cellular Oncogenes - 4.2 ~ 4.6 - Many retroviruses carrying oncogenes have been found in chickens and mice However, attempts undertaken during the 1970s to isolate viruses from most types of
Follicular Lymphoma. ced3 APOPTOSIS. *In the nematode Caenorhabditis elegans 131 of the organism's 1031 cells die during development.
 Harvard-MIT Division of Health Sciences and Technology HST.176: Cellular and Molecular Immunology Course Director: Dr. Shiv Pillai Follicular Lymphoma 1. Characterized by t(14:18) translocation 2. Ig heavy
Harvard-MIT Division of Health Sciences and Technology HST.176: Cellular and Molecular Immunology Course Director: Dr. Shiv Pillai Follicular Lymphoma 1. Characterized by t(14:18) translocation 2. Ig heavy
Test Name Results Units Bio. Ref. Interval. Positive
 LL - LL-ROHINI (NATIONAL REFERENCE 135091534 Age 36 Years Gender Female 1/9/2017 120000AM 1/9/2017 105316AM 2/9/2017 104147AM Ref By Final LEUKEMIA GENETIC ROFILE ANY SIX MARKERS, CR QUALITATIVE AML ETO
LL - LL-ROHINI (NATIONAL REFERENCE 135091534 Age 36 Years Gender Female 1/9/2017 120000AM 1/9/2017 105316AM 2/9/2017 104147AM Ref By Final LEUKEMIA GENETIC ROFILE ANY SIX MARKERS, CR QUALITATIVE AML ETO
membrane form secreted form 13 aa 26 aa K K V V K K 3aa
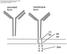 Harvard-MIT Division of Health Sciences and Technology HST.176: Cellular and Molecular Immunology Course Director: Dr. Shiv Pillai secreted form membrane form 13 aa 26 aa K K V V K K 3aa Hapten monosaccharide
Harvard-MIT Division of Health Sciences and Technology HST.176: Cellular and Molecular Immunology Course Director: Dr. Shiv Pillai secreted form membrane form 13 aa 26 aa K K V V K K 3aa Hapten monosaccharide
LYMPHOMAS an overview of some subtypes of NHLs
 One of the confusing aspects of the lymphoid neoplasms concerns the use of the descriptive terms "leukemia" and "lymphoma." LYMPHOMAS an overview of some subtypes of NHLs Leukemia is used for lymphoid
One of the confusing aspects of the lymphoid neoplasms concerns the use of the descriptive terms "leukemia" and "lymphoma." LYMPHOMAS an overview of some subtypes of NHLs Leukemia is used for lymphoid
Molecular Markers in Acute Leukemia. Dr Muhd Zanapiah Zakaria Hospital Ampang
 Molecular Markers in Acute Leukemia Dr Muhd Zanapiah Zakaria Hospital Ampang Molecular Markers Useful at diagnosis Classify groups and prognosis Development of more specific therapies Application of risk-adjusted
Molecular Markers in Acute Leukemia Dr Muhd Zanapiah Zakaria Hospital Ampang Molecular Markers Useful at diagnosis Classify groups and prognosis Development of more specific therapies Application of risk-adjusted
Test Name Results Units Bio. Ref. Interval. Positive
 LL - LL-ROHINI (NATIONAL REFERENCE 135091533 Age 28 Years Gender Male 1/9/2017 120000AM 1/9/2017 105415AM 4/9/2017 23858M Ref By Final LEUKEMIA DIAGNOSTIC COMREHENSIVE ROFILE, ANY 6 MARKERS t (1;19) (q23
LL - LL-ROHINI (NATIONAL REFERENCE 135091533 Age 28 Years Gender Male 1/9/2017 120000AM 1/9/2017 105415AM 4/9/2017 23858M Ref By Final LEUKEMIA DIAGNOSTIC COMREHENSIVE ROFILE, ANY 6 MARKERS t (1;19) (q23
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
The Adaptive Immune Response. T-cells
 The Adaptive Immune Response T-cells T Lymphocytes T lymphocytes develop from precursors in the thymus. Mature T cells are found in the blood, where they constitute 60% to 70% of lymphocytes, and in T-cell
The Adaptive Immune Response T-cells T Lymphocytes T lymphocytes develop from precursors in the thymus. Mature T cells are found in the blood, where they constitute 60% to 70% of lymphocytes, and in T-cell
Prepared by: Dr.Mansour Al-Yazji
 C L L CLL Prepared by: Abd El-Hakeem Abd El-Rahman Abu Naser Ahmed Khamis Abu Warda Ahmed Mohammed Abu Ghaben Bassel Ziad Abu Warda Nedal Mostafa El-Nahhal Dr.Mansour Al-Yazji LEUKEMIA Leukemia is a form
C L L CLL Prepared by: Abd El-Hakeem Abd El-Rahman Abu Naser Ahmed Khamis Abu Warda Ahmed Mohammed Abu Ghaben Bassel Ziad Abu Warda Nedal Mostafa El-Nahhal Dr.Mansour Al-Yazji LEUKEMIA Leukemia is a form
T cell Receptor. Chapter 9. Comparison of TCR αβ T cells
 Chapter 9 The αβ TCR is similar in size and structure to an antibody Fab fragment T cell Receptor Kuby Figure 9-3 The αβ T cell receptor - Two chains - α and β - Two domains per chain - constant (C) domain
Chapter 9 The αβ TCR is similar in size and structure to an antibody Fab fragment T cell Receptor Kuby Figure 9-3 The αβ T cell receptor - Two chains - α and β - Two domains per chain - constant (C) domain
Molecular Markers. Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC
 Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Antibodies and T Cell Receptor Genetics Generation of Antigen Receptor Diversity
 Antibodies and T Cell Receptor Genetics 2008 Peter Burrows 4-6529 peterb@uab.edu Generation of Antigen Receptor Diversity Survival requires B and T cell receptor diversity to respond to the diversity of
Antibodies and T Cell Receptor Genetics 2008 Peter Burrows 4-6529 peterb@uab.edu Generation of Antigen Receptor Diversity Survival requires B and T cell receptor diversity to respond to the diversity of
MECHANISMS OF B-CELL LYMPHOMA PATHOGENESIS
 MECHANISMS OF B-CELL LYMPHOMA PATHOGENESIS Ralf Küppers Abstract Chromosomal translocations involving the immunoglobulin loci are a hallmark of many types of B-cell. Other factors, however, also have important
MECHANISMS OF B-CELL LYMPHOMA PATHOGENESIS Ralf Küppers Abstract Chromosomal translocations involving the immunoglobulin loci are a hallmark of many types of B-cell. Other factors, however, also have important
Chapter 5. Generation of lymphocyte antigen receptors
 Chapter 5 Generation of lymphocyte antigen receptors Structural variation in Ig constant regions Isotype: different class of Ig Heavy-chain C regions are encoded in separate genes Initially, only two of
Chapter 5 Generation of lymphocyte antigen receptors Structural variation in Ig constant regions Isotype: different class of Ig Heavy-chain C regions are encoded in separate genes Initially, only two of
Development of B and T lymphocytes
 Development of B and T lymphocytes What will we discuss today? B-cell development T-cell development B- cell development overview Stem cell In periphery Pro-B cell Pre-B cell Immature B cell Mature B cell
Development of B and T lymphocytes What will we discuss today? B-cell development T-cell development B- cell development overview Stem cell In periphery Pro-B cell Pre-B cell Immature B cell Mature B cell
HIGH GRADE B-CELL LYMPHOMA DAVID NOLTE, MD (PGY-2) HUSSAM AL-KATEB, PHD, FACMG DEBORAH FUCHS, MD
 HIGH GRADE B-CELL LYMPHOMA DAVID NOLTE, MD (PGY-2) HUSSAM AL-KATEB, PHD, FACMG DEBORAH FUCHS, MD OUTLINE High grade B-cell lymphoma with MYC and BCL2 and/or BCL6 rearrangements Patient presentation 2008/2016
HIGH GRADE B-CELL LYMPHOMA DAVID NOLTE, MD (PGY-2) HUSSAM AL-KATEB, PHD, FACMG DEBORAH FUCHS, MD OUTLINE High grade B-cell lymphoma with MYC and BCL2 and/or BCL6 rearrangements Patient presentation 2008/2016
Multistep nature of cancer development. Cancer genes
 Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Molecular Pathogenesis of Multiple Myeloma:
 Molecular Pathogenesis of Multiple Myeloma: Ig translocations hyperdiploid vs non-hyperdiploid CYCLIN D dysregulation other oncogenic events Michael Kuehl MM: post-germinal center tumor of long-lived BM
Molecular Pathogenesis of Multiple Myeloma: Ig translocations hyperdiploid vs non-hyperdiploid CYCLIN D dysregulation other oncogenic events Michael Kuehl MM: post-germinal center tumor of long-lived BM
Template for Reporting Results of Biomarker Testing of Specimens From Patients With Chronic Lymphocytic Leukemia/Small Lymphocytic Lymphoma
 Template for Reporting Results of Biomarker Testing of Specimens From Patients With Chronic Lymphocytic Leukemia/Small Lymphocytic Lymphoma Version: CLLBiomarkers 1.0.0.2 Protocol Posting Date: June 2017
Template for Reporting Results of Biomarker Testing of Specimens From Patients With Chronic Lymphocytic Leukemia/Small Lymphocytic Lymphoma Version: CLLBiomarkers 1.0.0.2 Protocol Posting Date: June 2017
Introduction to Cancer Biology
 Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Molecular Hematopathology Leukemias I. January 14, 2005
 Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
Andrea s SI Session PCB Practice Test Test 3
 Practice Test Test 3 READ BEFORE STARTING PRACTICE TEST: Remember to please use this practice test as a tool to measure your knowledge, and DO NOT use it as your only tool to study for the test, since
Practice Test Test 3 READ BEFORE STARTING PRACTICE TEST: Remember to please use this practice test as a tool to measure your knowledge, and DO NOT use it as your only tool to study for the test, since
Use of MYC, BCL2 and BCL6 FISH for investigations of high grade B cell lymphoma
 Use of MYC, BCL2 and BCL6 FISH for investigations of high grade B cell lymphoma Dr Anthony Bench Haematopathology and Oncology Diagnostic Service Cambrıdge Unıversıty Hospitals NHS Foundatıon Trust Cambridge
Use of MYC, BCL2 and BCL6 FISH for investigations of high grade B cell lymphoma Dr Anthony Bench Haematopathology and Oncology Diagnostic Service Cambrıdge Unıversıty Hospitals NHS Foundatıon Trust Cambridge
Large cell immunoblastic Diffuse histiocytic (DHL) Lymphoblastic lymphoma Diffuse lymphoblastic Small non cleaved cell Burkitt s Non- Burkitt s
 Non Hodgkin s Lymphoma Introduction 6th most common cause of cancer death in United States. Increasing in incidence and mortality. Since 1970, the incidence of has almost doubled. Overview The types of
Non Hodgkin s Lymphoma Introduction 6th most common cause of cancer death in United States. Increasing in incidence and mortality. Since 1970, the incidence of has almost doubled. Overview The types of
Mixed Phenotype Acute Leukemias
 Mixed Phenotype Acute Leukemias CHEN GAO; AMY M. SANDS; JIANLAN SUN NORTH AMERICAN JOURNAL OF MEDICINE AND SCIENCE APR 2012 VOL 5 NO.2 INTRODUCTION Most cases of acute leukemia can be classified based
Mixed Phenotype Acute Leukemias CHEN GAO; AMY M. SANDS; JIANLAN SUN NORTH AMERICAN JOURNAL OF MEDICINE AND SCIENCE APR 2012 VOL 5 NO.2 INTRODUCTION Most cases of acute leukemia can be classified based
Corrigenda. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues (revised 4th edition): corrections made in second print run
 Corrigenda WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues (revised 4th edition): corrections made in second print run In addition to corrections of minor typographical errors, corrections
Corrigenda WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues (revised 4th edition): corrections made in second print run In addition to corrections of minor typographical errors, corrections
WBCs Disorders 1. Dr. Nabila Hamdi MD, PhD
 WBCs Disorders 1 Dr. Nabila Hamdi MD, PhD ILOs Compare and contrast ALL, AML, CLL, CML in terms of age distribution, cytogenetics, morphology, immunophenotyping, laboratory diagnosis clinical features
WBCs Disorders 1 Dr. Nabila Hamdi MD, PhD ILOs Compare and contrast ALL, AML, CLL, CML in terms of age distribution, cytogenetics, morphology, immunophenotyping, laboratory diagnosis clinical features
EBV infection B cells and lymphomagenesis. Sridhar Chaganti
 EBV infection B cells and lymphomagenesis Sridhar Chaganti How EBV infects B-cells How viral genes influence the infected B cell Differences and similarities between in vitro and in vivo infection How
EBV infection B cells and lymphomagenesis Sridhar Chaganti How EBV infects B-cells How viral genes influence the infected B cell Differences and similarities between in vitro and in vivo infection How
Pathology #07. Hussein Al-Sa di. Dr. Sohaib Al-Khatib. Mature B-Cell Neoplasm. 0 P a g e
 Pathology #07 Mature B-Cell Neoplasm Hussein Al-Sa di Dr. Sohaib Al-Khatib 0 P a g e Thursday 18/2/2016 Our lecture today (with the next 2 lectures) will be about lymphoid tumors This is a little bit long
Pathology #07 Mature B-Cell Neoplasm Hussein Al-Sa di Dr. Sohaib Al-Khatib 0 P a g e Thursday 18/2/2016 Our lecture today (with the next 2 lectures) will be about lymphoid tumors This is a little bit long
Significance of Chromosome Changes in Hematological Disorders and Solid Tumors
 Significance of Chromosome Changes in Hematological Disorders and Solid Tumors Size of Components of Human Genome Size of haploid genome 3.3 X 10 9 DNA basepairs Estimated genetic constitution 30,000
Significance of Chromosome Changes in Hematological Disorders and Solid Tumors Size of Components of Human Genome Size of haploid genome 3.3 X 10 9 DNA basepairs Estimated genetic constitution 30,000
Significance of Chromosome Changes in Hematological Disorders and Solid Tumors
 Significance of Chromosome Changes in Hematological Disorders and Solid Tumors Size of Components of Human Genome Size of haploid genome! Estimated genetic constitution! Size of average chromosome
Significance of Chromosome Changes in Hematological Disorders and Solid Tumors Size of Components of Human Genome Size of haploid genome! Estimated genetic constitution! Size of average chromosome
Small B-cell (Histologically Low Grade) Lymphoma
 Frequency of Lymphoid Neoplasms Small B-cell (Histologically Low Grade) Lymphoma Stephen Hamilton-Dutoit Institute of Pathology Aarhus University Hospital B-cell neoplasms 88% Diffuse large B-cell lymphoma
Frequency of Lymphoid Neoplasms Small B-cell (Histologically Low Grade) Lymphoma Stephen Hamilton-Dutoit Institute of Pathology Aarhus University Hospital B-cell neoplasms 88% Diffuse large B-cell lymphoma
B Lymphocyte Development and Activation
 Harvard-MIT Division of Health Sciences and Technology HST.176: Cellular and Molecular Immunology Course Director: Dr. Shiv Pillai 09/26/05; 9 AM Shiv Pillai B Lymphocyte Development and Activation Recommended
Harvard-MIT Division of Health Sciences and Technology HST.176: Cellular and Molecular Immunology Course Director: Dr. Shiv Pillai 09/26/05; 9 AM Shiv Pillai B Lymphocyte Development and Activation Recommended
Problem Set 8 Key 1 of 8
 7.06 2003 Problem Set 8 Key 1 of 8 7.06 2003 Problem Set 8 Key 1. As a bright MD/PhD, you are interested in questions about the control of cell number in the body. Recently, you've seen three patients
7.06 2003 Problem Set 8 Key 1 of 8 7.06 2003 Problem Set 8 Key 1. As a bright MD/PhD, you are interested in questions about the control of cell number in the body. Recently, you've seen three patients
FOLLICULARITY in LYMPHOMA
 FOLLICULARITY in LYMPHOMA Reactive Follicular Hyperplasia Follicular Hyperplasia irregular follicles Follicular Hyperplasia dark and light zones Light Zone Dark Zone Follicular hyperplasia MIB1 Follicular
FOLLICULARITY in LYMPHOMA Reactive Follicular Hyperplasia Follicular Hyperplasia irregular follicles Follicular Hyperplasia dark and light zones Light Zone Dark Zone Follicular hyperplasia MIB1 Follicular
The next lymphoma classification Luca Mazzucchelli Istituto cantonale di patologia, Locarno
 Evolution of classification The next classification Luca Mazzucchelli Istituto cantonale di patologia, Locarno The Lymphoma Forum of Excellence, Bellinzona, January 2011 Rappaport Lukes and Collins (immunophenotype)
Evolution of classification The next classification Luca Mazzucchelli Istituto cantonale di patologia, Locarno The Lymphoma Forum of Excellence, Bellinzona, January 2011 Rappaport Lukes and Collins (immunophenotype)
Variations in Chromosome Structure & Function. Ch. 8
 Variations in Chromosome Structure & Function Ch. 8 1 INTRODUCTION! Genetic variation refers to differences between members of the same species or those of different species Allelic variations are due
Variations in Chromosome Structure & Function Ch. 8 1 INTRODUCTION! Genetic variation refers to differences between members of the same species or those of different species Allelic variations are due
Problem Set 5 KEY
 2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
PhenoPath. Diagnoses you can count on B CELL NON-HODGKIN LYMPHOMA
 PhenoPath Diagnoses you can count on B CELL NON-HODGKIN LYMPHOMA C urrent diagnosis of B cell non-hodgkin lymphoma (B-NHL) is based on the 2008 WHO Classification of Tumours of Haematopoietic and Lymphoid
PhenoPath Diagnoses you can count on B CELL NON-HODGKIN LYMPHOMA C urrent diagnosis of B cell non-hodgkin lymphoma (B-NHL) is based on the 2008 WHO Classification of Tumours of Haematopoietic and Lymphoid
Molecular Hematopathology
 Molecular Hematopathology Charles E. Hill, MD, PhD Emory University School of Medicine April 2013 The Association for Molecular Pathology Education. Innovation and Improved Patient Care. Advocacy. www.amp.org
Molecular Hematopathology Charles E. Hill, MD, PhD Emory University School of Medicine April 2013 The Association for Molecular Pathology Education. Innovation and Improved Patient Care. Advocacy. www.amp.org
Review. Molecular Diagnostic Approach to Non-Hodgkin s Lymphoma. Materials and Methods. Daniel A. Arber
 Journal of Molecular Diagnostics, Vol. 2, No. 4, November 2000 Copyright American Society for Investigative Pathology and the Association for Molecular Pathology Review Molecular Diagnostic Approach to
Journal of Molecular Diagnostics, Vol. 2, No. 4, November 2000 Copyright American Society for Investigative Pathology and the Association for Molecular Pathology Review Molecular Diagnostic Approach to
Contents. vii. Preface... Acknowledgments... v xiii
 Contents Preface... Acknowledgments... v xiii SECTION I 1. Introduction... 3 Knowledge-Based Diagnosis... 4 Systematic Examination of the Lymph Node... 7 Cell Type Identification... 9 Cell Size and Cellularity...
Contents Preface... Acknowledgments... v xiii SECTION I 1. Introduction... 3 Knowledge-Based Diagnosis... 4 Systematic Examination of the Lymph Node... 7 Cell Type Identification... 9 Cell Size and Cellularity...
Aggressive B-cell lymphomas and gene expression profiling towards individualized therapy?
 Aggressive B-cell lymphomas and gene expression profiling towards individualized therapy? Andreas Rosenwald Institute of Pathology, University of Würzburg, Germany Barcelona, June 18, 2010 NEW WHO CLASSIFICATION
Aggressive B-cell lymphomas and gene expression profiling towards individualized therapy? Andreas Rosenwald Institute of Pathology, University of Würzburg, Germany Barcelona, June 18, 2010 NEW WHO CLASSIFICATION
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Defined lymphoma entities in the current WHO classification
 Defined lymphoma entities in the current WHO classification Luca Mazzucchelli Istituto cantonale di patologia, Locarno Bellinzona, January 29-31, 2016 Evolution of lymphoma classification Rappaport Lukes
Defined lymphoma entities in the current WHO classification Luca Mazzucchelli Istituto cantonale di patologia, Locarno Bellinzona, January 29-31, 2016 Evolution of lymphoma classification Rappaport Lukes
Cancer Genetics. What is Cancer? Cancer Classification. Medical Genetics. Uncontrolled growth of cells. Not all tumors are cancerous
 Session8 Medical Genetics Cancer Genetics J avad Jamshidi F a s a U n i v e r s i t y o f M e d i c a l S c i e n c e s, N o v e m b e r 2 0 1 7 What is Cancer? Uncontrolled growth of cells Not all tumors
Session8 Medical Genetics Cancer Genetics J avad Jamshidi F a s a U n i v e r s i t y o f M e d i c a l S c i e n c e s, N o v e m b e r 2 0 1 7 What is Cancer? Uncontrolled growth of cells Not all tumors
Anaplastic Large Cell Lymphoma (of T cell lineage)
 Anaplastic Large Cell Lymphoma (of T cell lineage) Definition T-cell lymphoma comprised of large cells with abundant cytoplasm and pleomorphic, often horseshoe-shaped nuclei CD30+ Most express cytotoxic
Anaplastic Large Cell Lymphoma (of T cell lineage) Definition T-cell lymphoma comprised of large cells with abundant cytoplasm and pleomorphic, often horseshoe-shaped nuclei CD30+ Most express cytotoxic
Case Report A case of EBV positive diffuse large B-cell lymphoma of the adolescent
 Int J Clin Exp Med 2014;7(1):307-311 www.ijcem.com /ISSN:1940-5901/IJCEM1311029 Case Report A case of EBV positive diffuse large B-cell lymphoma of the adolescent Qilin Ao 2, Ying Wang 1, Sanpeng Xu 2,
Int J Clin Exp Med 2014;7(1):307-311 www.ijcem.com /ISSN:1940-5901/IJCEM1311029 Case Report A case of EBV positive diffuse large B-cell lymphoma of the adolescent Qilin Ao 2, Ying Wang 1, Sanpeng Xu 2,
Ig light chain rearrangement: Rescue pathway
 B Cell Development Ig light chain rearrangement: Rescue pathway There is only a 1:3 chance of the join between the V and J region being in frame Vk Jk Ck Non-productive Rearrangement Light chain has a
B Cell Development Ig light chain rearrangement: Rescue pathway There is only a 1:3 chance of the join between the V and J region being in frame Vk Jk Ck Non-productive Rearrangement Light chain has a
Gray Zones and Double Hits Distinguishing True Burkitt Lymphoma from Other High-Grade B-NHLs Burkitt Lymphoma Burkitt-Like Lymphoma DLBCL Patrick Tres
 Gray Zones and Double Hits Distinguishing True Burkitt Lymphoma from Other High-Grade B-NHLs Burkitt Lymphoma Burkitt-Like Lymphoma DLBCL Patrick Treseler, MD, PhD University of California San Francisco
Gray Zones and Double Hits Distinguishing True Burkitt Lymphoma from Other High-Grade B-NHLs Burkitt Lymphoma Burkitt-Like Lymphoma DLBCL Patrick Treseler, MD, PhD University of California San Francisco
VIII Curso Internacional del PIRRECV. Some molecular mechanisms of cancer
 VIII Curso Internacional del PIRRECV Some molecular mechanisms of cancer Laboratorio de Comunicaciones Celulares, Centro FONDAP Estudios Moleculares de la Celula (CEMC), ICBM, Facultad de Medicina, Universidad
VIII Curso Internacional del PIRRECV Some molecular mechanisms of cancer Laboratorio de Comunicaciones Celulares, Centro FONDAP Estudios Moleculares de la Celula (CEMC), ICBM, Facultad de Medicina, Universidad
Volume 7, Issue 1 January 2012
 The Hong Kong College of Pathologists, Incorporated in Hong Kong with Limited Liability Volume 7, Issue 1 January 2012 Editorial note: Chronic lymphocytic leukaemia (CLL) is the commonest chronic lymphoproliferative
The Hong Kong College of Pathologists, Incorporated in Hong Kong with Limited Liability Volume 7, Issue 1 January 2012 Editorial note: Chronic lymphocytic leukaemia (CLL) is the commonest chronic lymphoproliferative
Chronic Lymphocytic Leukemia Mantle Cell Lymphoma Elias Campo
 Chronic Lymphocytic Leukemia Mantle Cell Lymphoma Elias Campo Hospital Clinic, University of Barcelona Small B-cell lymphomas NAIVE -B LYMPHOCYTE MEMORY CELL CLL MCL FL MZL Small cell size Low proliferation
Chronic Lymphocytic Leukemia Mantle Cell Lymphoma Elias Campo Hospital Clinic, University of Barcelona Small B-cell lymphomas NAIVE -B LYMPHOCYTE MEMORY CELL CLL MCL FL MZL Small cell size Low proliferation
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
The patient had a mild splenomegaly but no obvious lymph node enlargement. The consensus phenotype obtained from part one of the exercise was:
 Case History An 86 year old male was admitted to hospital with chest infection. Haematological examination subsequently revealed the following: Hb- 11.0 g/dl; WBC- 67.1 x 10^9/l; PLT- 99 x10^9/l; RBC-
Case History An 86 year old male was admitted to hospital with chest infection. Haematological examination subsequently revealed the following: Hb- 11.0 g/dl; WBC- 67.1 x 10^9/l; PLT- 99 x10^9/l; RBC-
Burkitt lymphoma. Sporadic Endemic in Africa associated with EBV Translocations involving MYC gene on chromosome 8
 Heme 8 Burkitt lymphoma Sporadic Endemic in Africa associated with EBV Translocations involving MYC gene on chromosome 8 Most common is t(8;14) Believed to be the fastest growing tumor in humans!!!! Morphology
Heme 8 Burkitt lymphoma Sporadic Endemic in Africa associated with EBV Translocations involving MYC gene on chromosome 8 Most common is t(8;14) Believed to be the fastest growing tumor in humans!!!! Morphology
DETERMINATION OF A LYMPHOID PROCESS
 Chapter 2 Applications of Touch Preparation Cytology to Intraoperative Consultations: Lymph Nodes and Extranodal Tissues for Evaluation of Hematolymphoid Disorders INTRODUCTION As discussed in Chap. 1,
Chapter 2 Applications of Touch Preparation Cytology to Intraoperative Consultations: Lymph Nodes and Extranodal Tissues for Evaluation of Hematolymphoid Disorders INTRODUCTION As discussed in Chap. 1,
Canadian College of Medical Geneticists (CCMG) Cytogenetics Examination. May 4, 2010
 Canadian College of Medical Geneticists (CCMG) Cytogenetics Examination May 4, 2010 Examination Length = 3 hours Total Marks = 100 (7 questions) Total Pages = 8 (including cover sheet and 2 pages of prints)
Canadian College of Medical Geneticists (CCMG) Cytogenetics Examination May 4, 2010 Examination Length = 3 hours Total Marks = 100 (7 questions) Total Pages = 8 (including cover sheet and 2 pages of prints)
CELL CYCLE MOLECULAR BASIS OF ONCOGENESIS
 CELL CYCLE MOLECULAR BASIS OF ONCOGENESIS Summary of the regulation of cyclin/cdk complexes during celll cycle Cell cycle phase Cyclin-cdk complex inhibitor activation Substrate(s) G1 Cyclin D/cdk 4,6
CELL CYCLE MOLECULAR BASIS OF ONCOGENESIS Summary of the regulation of cyclin/cdk complexes during celll cycle Cell cycle phase Cyclin-cdk complex inhibitor activation Substrate(s) G1 Cyclin D/cdk 4,6
oncogenes-and- tumour-suppressor-genes)
 Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Hepatic Lymphoma Diagnosis An Algorithmic Approach
 Hepatic Lymphoma Diagnosis An Algorithmic Approach Ryan M. Gill, M.D., Ph.D. University of California, San Francisco PLEASE TURN OFF YOUR CELL PHONES Disclosure of Relevant Financial Relationships USCAP
Hepatic Lymphoma Diagnosis An Algorithmic Approach Ryan M. Gill, M.D., Ph.D. University of California, San Francisco PLEASE TURN OFF YOUR CELL PHONES Disclosure of Relevant Financial Relationships USCAP
Introduction to Genetics
 Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
609G: Concepts of Cancer Genetics and Treatments (3 credits)
 Master of Chemical and Life Sciences Program College of Computer, Mathematical, and Natural Sciences 609G: Concepts of Cancer Genetics and Treatments (3 credits) Text books: Principles of Cancer Genetics,
Master of Chemical and Life Sciences Program College of Computer, Mathematical, and Natural Sciences 609G: Concepts of Cancer Genetics and Treatments (3 credits) Text books: Principles of Cancer Genetics,
Major Histocompatibility Complex (MHC) and T Cell Receptors
 Major Histocompatibility Complex (MHC) and T Cell Receptors Historical Background Genes in the MHC were first identified as being important genes in rejection of transplanted tissues Genes within the MHC
Major Histocompatibility Complex (MHC) and T Cell Receptors Historical Background Genes in the MHC were first identified as being important genes in rejection of transplanted tissues Genes within the MHC
number Done by Corrected by Doctor Maha Shomaf
 number 19 Done by Waseem Abo-Obeida Corrected by Abdullah Zreiqat Doctor Maha Shomaf Carcinogenesis: the molecular basis of cancer. Non-lethal genetic damage lies at the heart of carcinogenesis and leads
number 19 Done by Waseem Abo-Obeida Corrected by Abdullah Zreiqat Doctor Maha Shomaf Carcinogenesis: the molecular basis of cancer. Non-lethal genetic damage lies at the heart of carcinogenesis and leads
From Morphology to Molecular Pathology: A Practical Approach for Cytopathologists Part 1-Cytomorphology. Songlin Zhang, MD, PhD LSUHSC-Shreveport
 From Morphology to Molecular Pathology: A Practical Approach for Cytopathologists Part 1-Cytomorphology Songlin Zhang, MD, PhD LSUHSC-Shreveport I have no Conflict of Interest. FNA on Lymphoproliferative
From Morphology to Molecular Pathology: A Practical Approach for Cytopathologists Part 1-Cytomorphology Songlin Zhang, MD, PhD LSUHSC-Shreveport I have no Conflict of Interest. FNA on Lymphoproliferative
Eurekah Bioscience Collection
 in Malignant 5/31/06 12:09 PM in Malignant to Leukemia and Lymphoma Eurekah Bioscience Collection in Malignant Sarah E. enrickson Elena M. artmann German Ott Andreas Rosenwald* The practice of clinical
in Malignant 5/31/06 12:09 PM in Malignant to Leukemia and Lymphoma Eurekah Bioscience Collection in Malignant Sarah E. enrickson Elena M. artmann German Ott Andreas Rosenwald* The practice of clinical
Alternative splicing. Biosciences 741: Genomics Fall, 2013 Week 6
 Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Aggressive B-cell Lymphomas Updated WHO classification Elias Campo
 Aggressive B-cell Lymphomas Updated WHO classification Elias Campo Hospital Clinic, University of Barcelona Diffuse Large B-cell Lymphoma A Heterogeneous Category Subtypes with differing: Histology and
Aggressive B-cell Lymphomas Updated WHO classification Elias Campo Hospital Clinic, University of Barcelona Diffuse Large B-cell Lymphoma A Heterogeneous Category Subtypes with differing: Histology and
Case 3. Ann T. Moriarty,MD
 Case 3 Ann T. Moriarty,MD Case 3 59 year old male with asymptomatic cervical lymphadenopathy. These images are from a fine needle biopsy of a left cervical lymph node. Image 1 Papanicolaou Stained smear,100x.
Case 3 Ann T. Moriarty,MD Case 3 59 year old male with asymptomatic cervical lymphadenopathy. These images are from a fine needle biopsy of a left cervical lymph node. Image 1 Papanicolaou Stained smear,100x.
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
Determination Differentiation. determinated precursor specialized cell
 Biology of Cancer -Developmental Biology: Determination and Differentiation -Cell Cycle Regulation -Tumor genes: Proto-Oncogenes, Tumor supressor genes -Tumor-Progression -Example for Tumor-Progression:
Biology of Cancer -Developmental Biology: Determination and Differentiation -Cell Cycle Regulation -Tumor genes: Proto-Oncogenes, Tumor supressor genes -Tumor-Progression -Example for Tumor-Progression:
Intronic BCL-6 mutations are preferentially targeted to the translocated allele in t(3;14)(q27;q32) non-hodgkin B-cell lymphoma
 NEOPLASIA Brief report Intronic BCL-6 mutations are preferentially targeted to the translocated allele in t(3;14)(q27;q32) non-hodgkin B-cell lymphoma Fabrice Jardin, Christian Bastard, Nathalie Contentin,
NEOPLASIA Brief report Intronic BCL-6 mutations are preferentially targeted to the translocated allele in t(3;14)(q27;q32) non-hodgkin B-cell lymphoma Fabrice Jardin, Christian Bastard, Nathalie Contentin,
Genome of Hepatitis B Virus. VIRAL ONCOGENE Dr. Yahwardiah Siregar, PhD Dr. Sry Suryani Widjaja, Mkes Biochemistry Department
 Genome of Hepatitis B Virus VIRAL ONCOGENE Dr. Yahwardiah Siregar, PhD Dr. Sry Suryani Widjaja, Mkes Biochemistry Department Proto Oncogen and Oncogen Oncogen Proteins that possess the ability to cause
Genome of Hepatitis B Virus VIRAL ONCOGENE Dr. Yahwardiah Siregar, PhD Dr. Sry Suryani Widjaja, Mkes Biochemistry Department Proto Oncogen and Oncogen Oncogen Proteins that possess the ability to cause
T Cell Receptor & T Cell Development
 T Cell Receptor & T Cell Development Questions for the next 2 lectures: How do you generate a diverse T cell population with functional TCR rearrangements? How do you generate a T cell population that
T Cell Receptor & T Cell Development Questions for the next 2 lectures: How do you generate a diverse T cell population with functional TCR rearrangements? How do you generate a T cell population that
