Huizhen Zhang 1*, Hongxiang Guo 2, Qingtang Fan 1 and Yiming Wu 1
|
|
|
- Sophie Turner
- 6 years ago
- Views:
Transcription
1 Life Science Journal, Vol 6, No 3, Two-dimensional Gel Electrophoresis and Matrix-assisted Laser Desorption Ionization Time of Flight Mass Spectrometry Huizhen Zhang 1*, Hongxiang Guo 2, Qingtang Fan 1 and Yiming Wu 1 1 Department of Labor and Environmental Health,College of Public Health, Zhengzhou University, Henan , PR China,2 Department of life Sciences, Henan agriculture University, Henan , PR China Received March 10, 2009 Abstract: Tumor-specific protein spots were identified, including ApolipoproteinA-I precursor (Apo-AI), Peptidyl-prolyl cis-trans isomerase A, Calgranulin B (MRP-14), Calgizzarin (S100C protein), Ras-related protein Rab-14, apolipoprotein E. Among these 6 proteins, the potential significance of the differential expressions is discussed. These findings demonstrate that differential expression analysis of proteomes may be useful for the development of new molecular markers for diagnosis and prognosis of lung cancer. [Life Science Journal. 2009; 6(3): 46 53] (ISSN: ) Keywords proteome; two-dimensional electrophoresis (2DE); matrix assisted laser desorption ionization time of flight mass spectrometry (MALDI-TOF-MS); lung cancer, tumor marker 1. Introduction because of the known heterogeneity of lung cancers. Lung cancer is the leading cause of cancer death Methods have been developed to identify the tumor worldwide, and the number of lung cancer patients has associated antigens such as molecular cloning in system been increased with the exposure to the environmental or using a biochemical strategy based on the extraction risk factors [1 4]. Prevention of lung cancer is a serious of antigenic peptides bound to major histocompatibility worldwide challenge. It is clear that the successful complex class 1 molecules from tumor cells. These prevention of lung cancer will depend on the reduction of methods have allowed the recognition of certain human risk factors along with better methods of screening and tumor antigens [7]. However, no evidence has been early detection. Tumor markers are widely used for obtained indicating that the detection of these markers screening, diagnosis, staging, prognosis, monitoring precedes clinical diagnosis of lung cancer. response to treatment, and detection of recurrent Proteome is defined as the total proteins expressed disease [5]. The analysis of proteins overexpressed in lung by a genome and the comparative analyses of proteomes cancer, and making the proteins serve as tumor markers, from cancer cells or tissues promise the discovery of new has been the subject of extensive research. biomarkers for early detection of cancers [8, 9]. In this Although many insights into the molecular proteomics approach, two-dimensional polyacrylamide pathology of lung tumors have been achieved, additional gel electrophoresis (2D-PAGE) and mass spectrometry information is critical to our understanding of the have been the most important technologies for the development and progression of these tumors as well as separation and identification of proteins respectively [10, to early diagnosis. The most commonly evaluated 11]. 2D-PAGE is a powerful research technique, which markers include neuron-specific enolase, makes it possible to simultaneously examine hundreds of carcinoembryonic antigen, cytokeratin 19 fragments polypeptides in a tissue sample. It has been widely used (CYFRA 21-1), squamous cell carcinoma antigen, cancer for the detection and identification of potential tumor antigen CA 125, and tissue polypeptide antigen [6]. markers [12].Using 2D-PAGE and matrix assisted laser It is a complex work to detect new candidate markers desorption/ionization-time of flight-mass spectrometry (MALDI-TOF-MS), we tried to identify the biomarkers *Corresponding Author Huizhen Zhang, PhD of lung cancer by the comparative proteomes analysis of Huizhenzhang@zzu.edu.cn human normal lung tissues and cancerous lung tissues, 46
2 Huizhen Zhang et al. and the study will help to elucidate the molecular mechanism of cellular events associated with cancer progression, such as cellular signaling [13,14]. estimated according to a commercial Bradford reagent. BSA was used as standard D electrophoresis and image analysis First-dimension iso-electric focusing (IEF) was 1 Materials and methods 1.1Reagents Immobiline DryStrips (ph 3 10, 17 cm), DryStrip coverfluids, immobilized ph gradient (IPG) buffer, 3-[(3-cholamidopropyl) dimethylammonio]-1-propanesulfonate(chaps), carried out on a ProteanIEF cell (Bio-Rad) using precast 17 cm ph 3 10 IPG gelstrips (Bio-Rad). 300 g total protein for silver staining gels (1000 g for Coomassie Brilliant Blue staining gels) was mixed with rehydration solution (8 M urea, 2% CHAPS, 50 mm DTT,0.2% Bio-Lyte 3/10 ampholyte, and 0.001% bromophenol blue) bromophenol blue, agarose, acrylamide, tris-base, to a total volume of 500 L. Rehydration and IEF were glycine, sodium dodecyl sulfate (SDS), carried out as follows: 12 h of passive rehydration, IEF at N,N,N',N'-tetramethylethylenediamine (TEMED), and coomassie brilliant blue R-250, dithiothreitol(dtt), iodoacetamide, acetic acid, trypsin(sequencing grade), trifluoroacetic acid were bought from Bio-Rad and Sigma. The remaining chemicals were of analytical grade. All buffers were prepared with Milli-Q water. 1.2 Tissues and extraction preparation Lung cancer tissues and corresponding adjacent noncancerous lung tissues were obtained with informed consent from patients at the First Affiliated Hospital of Zhengzhou University. Cancer samples were obtained from the core part of the tumor to avoid the adjacent noncancerous tissue. For each of the normal tissues, 500 V for 30 min, 5000 V for 3 h, and 10,000 V for 8h. The current was limited to 70 A per gel strip. All IEF steps were carried out at 17 C. After IEF separation, the gel strips were immediately equilibrated using two steps in equilibrium buffer containing 50 mm Tris-HCl (ph 8.8), 6 M urea, 30% glycerol, and 2% SDS. At the first step, 2% DTT (W/V) was included in the equilibrium buffer. 2.5% iodoacetamide (W/V) was added at the second step. The second dimension separation (13% T, 2.7% C) was carried out using Tris-glycine buffer containing 1 g/l SDS on a Protean II xi 2-D cell (Bio-Rad) according to the following procedure: 5 ma/gel for the initial 0.5 h and 18 ma/gel there after and surface epithelium was procured selectively by a temperature of 4 C, until the bromophenol blue front dissection with special care for minimal contamination of nonepithelial cells, and samples were immediately snap-frozen in liquid nitrogen. They were classified histologically according to Lauren s classification after hematoxylin and eosin staining. Fragments of normal and malignant tissues were sharp dissected and homogenized with a homogenizer in 2 ml fresh lysis buffer [7 mol/l urea, 2 mol/l thiourea, reached the bottom of the gel. For silver staining, gels were immersed in ethanol: acetic acid: water (35:7:58) for 1.5 h, followed by washed twice in deionized water for 20 min. Gels were pretreated for 1 min in a solution of 0.2 g/l Na 2 S 2 O 3 and followed by 3 of 1-min washes in deionized water. Proteins were stained in a solution containing 2 g/l AgNO 3 and 0.075% formalin (37 g/l formaldehyde in 40 mmol/l Tris,40 g/l 3-([3-cholamidopropyl] water) for 20 min, and washed twice in deionized water dimethylammonio), 1-propanesulfonate (CHAPS), for 1 min. Subsequently, gels were developed in a 100mmol/L dithiothreilol, 0.5 mmol/l PMSF, 0.5 mmol/l EDTA, 2%(V/V)NP-40, 2%(V/V) Bio-Lyte 3/10, 1%(V/V)TritonX-100], then placed into tubes on ice for dealing with ultrasonic for 10 min. The mixture was centrifuged at r/min for 5 min to remove tissue and cell debris, then centrifuged in a tableletop ultracentrifuge at g for 30 min at 4 C. The supernatant was pipetted off and stored at -80 C until use. These were used as the 2DE samples for the soluble fraction. Protein concentration of 2-DE samples was solution of 0.6 g/l formaldehyde, 20 g/l Na 2 S 2 O 3 and g/l Na 2 S 2 O 3. When the desired intensity was attained, the developer was discarded and reaction was stopped by 10 g/l EDTA-Na 2. For Coomassie Brilliant Blue staining of gels, gels were equilibrated in a solution containing 500 ml/l methanol, 50 ml/l acetic acid and 25 g/l Coomassie Brilliant Blue R-250. Gels were rinsed in 300 ml/l ethanol containing 70 ml/l acetic acid. To account for experimental variation, three batches of total proteins extracted from the experimental and 47
3 Life Science Journal, Vol 6, No 3, control sample respectively, were subjected to 2-D electrophoresis and replicate gels were simultaneously run three times. Protein patterns in the gels after staining the target and dried. Dried spots were analyzed in an REFLEX-III (Bluker) MALDI-TOF mass spectrometer. The spectrometer was run in positive ion mode and in were recorded as digitalized images using a reflector mode with the setting: accelerating voltage, 20 high-resolution scanner. Gel image matching was done with PDQuest software(version 6.2, Bio-Rad, Richmond, CA). 1.4 In-gel digestion Protein spots on Coomassie blue stained gel were performed essentially as described. After the completion of staining, the gel slab was washed twice with water for 10 min. The spots of interest were excised with a scalpel and put into 1.5 ml micro-tubes. The particles were kv; grid voltage, 76%; guide wire voltage, 0.01%; and a delay time of 150 ns. The low mass gate was set at 500 m/z. Protein identification was carried out by peptide mass fingerprinting with searching the protein databases of Swiss-Prot/TrEMBL ( and Mascot ( The following search parameters were applied: a mass tolerance of 50 ppm and one incomplete cleavage were allowed; acetylation of the washed twice with water and then twice with N-terminus, alkylation of lysine by water/acetonitrile (1:1) for 15 min. The solvent volumes were about twice that of the gel. Liquid was removed, carboxyamidomethylation were considered as possible modifications. acetonitrile was added to the gel particles and the mixture was left for 2 h. After that, liquid was removed and the particles were rehydrated in 25 mmol/l NH 4 HCO 3 for 5 min. Acetonitrile was added to produce a 1:1 mixture of 25 mmol/l NH 4 HCO 3 /acetonitrile and the mixture was incubated for 15 min. All liquid was removed. Gel particles were dried in a vacuum centrifuge, reswelled in 10 mmol/l dithiothreilol and 25 mmol/l NH 4 HCO 3, and incubated for 30 min at 56 C to reduce the peptides. After chillness of tubes to room temperature and removal of the liquid, 55 mmol/l iodoacetamide in 25 mmol/l NH 4 HCO 3 was added. The tubes were incubated for 30 min at room temperature in the dark to S-alkylate the peptides. Then iodoacetamide solution was 2 Results 2.1 Protein expression maps of paired samples and image analysis In order to evaluate the reproducibility of the soluble protein preparations and to quantify the protein extracts for 2DE gel analysis, mini 2DE gels (7 cm, ph3-10) were used at first. To ensure quality and reproducibility of results, three gels per sample were processed simultaneously with the same power supply and analyzed using PDQuest 2-D software(version 7.1, Bio-Rad, Richmond, CA). Protein extracts prepared from tissues were compared in this way and found to be highly reproducible. Protein patterns were analyzed using removed, the particles were washed with 25 mmol/l PDQuest software. Qualitative and quantitative NH 4 HCO 3 and acetonitrile, dried in a vacuum centrifuge, rehydrated in digestion buffer containing 50 mmol/l NH 4 HCO 3 and 12.5 ng/l trypsin (TPCK-treated, proteomics grade, Sigma, USA), incubated for 12 h at 37 C. After digestion, 25 mmol/l NH 4 HCO 3 was added, and the tube was incubated for 15 min. Acetonitrile was added and the tube was incubated for another 15 min. The supernatant was recovered, and the comparisons were made between replicate groups, comprising the three gels for each sample. Quantitation is based on the peak intensity and area of Gaussianfitted spots, allowing more accurate quantitation than summation of pixel intensities. Any differential spots that were not present or absent in all three gels within a replicate group were excluded. Two-dimensional gel electrophoresis maps extraction was repeated twice with 50 g/l were constructed for human normal lung TFA/acetonitrile (1:1). The three extracts were pooled and dried in a vacuum centrifuge. tissues and cancerous lung tissues proteins in fig.1. We made separate maps for the ph range 1.5 MALDI-TOF-MS identification of peptide 10. The second-dimensional gel mixtures and database searching The peptide mixtures solubilized with matrix solution CCA ( -cyano-4-hydroxycinnamic acid) was spotted on electrophoresis was performed with 12.5% SDS-PAGE. For each sample, over 800 protein spots were resolved in a 2-DE gel (20 cm 20 48
4 Huizhen Zhang et al. cm) with silver-stained by computer-aided image analysis, and over 500 protein spots were resolved with Coomassie Brilliant Blue-stained. Figure 1A (from cancer tissues) contains a total of 530 spots, whereas figure 1B (from normal tissues) contains a total of 560 spots. A total of 515 spots from cancer tissues could be matched to those from normal tissues. In total, we were able to identify cancer-specific spots in 2DE gels. The positions of the identified proteins are shown in figure 1A. All of the identified spots could be considered as abundant proteins because of Coomassie Blue staining. A B Figure 1. 2-D gel images of protein expression in lung cancer tissue and normal lung tissue A: sample from lung cancer tissue. B: sample from normal lung tissue of the same patient. Proteins were separated on ph 3-10 linear IPG strip in the first dimension and 125 g/l SDS-PAGE in the second dimension. All labeled spots were different expresses proteins. To evaluate the potential for spurious differences that are not stage-specific, we also compared replicate groups in which gels were arbitrarily assigned to one of three groups, each containing one gel from each stage. No differences were detected between these groups, which suggested that the differentially represented spots are genuinely sample-specific. 2.2 Identification of overexpression proteins in cancerous lung tissues using peptide mass fingerprinting Overexpression proteins spots were excised from the 2-D gels of cancerous lung tissues and subjected to trypsin digestion and MALDI mass spectrometry. 6 spots were identified successfully. The criteria used to accept identifications including the extent of sequence coverage, the number of peptides matched, the probability score, and whether human protein appeared as the top candidates in the first pass search where no restriction was applied to the species of origin. The data for the protein spot 1 as an example are shown in fig. 2. Fig. 2A shows the MALDI-TOF MS peptide mass fingerprint spectrum of trypsin-digested protein spot 1. Fig. 2B is the probability based mowse score. Score is -10*Log(P), where P is the probability that the observed match is a random event. Protein scores greater than 53 are significant (p<0.05). The score of two proteins is significant, we choice the protein with the highest score.fig. 2C lists the matching peptides. Twenty peptides were matched with ApolipoproteinA-I, accession-numbered as P02647 in the protein database SWISS-PROT. These 20 peptides are indicated in the protein sequence shown in fig. 2D. The results of identification are summarized in the table 1. These identified protein spots are numbered as shown in Figure
5 Life Science Journal, Vol 6, No 3, A B C APA1_HUMAN; Apolipoprotein A-I precursor (Apo-AI) (P02647) 1 MKAAVLTLAV LFLTGSQARH FWQQDEPPQS PWDRVKDLAT VYVDVLKDSG 51 RDYVSQFEGS ALGKQLNLKL LDNWDSVTST FSKLREQLGP VTQEFWDNLE 101 KETEGLRQEM SKDLEEVKAK VQPYLDDFQK KWQEEMELYR QKVEPLRAEL 151 QEGARQKLHE LQEKLSPLGE EMRDRARAHV DALRTHLAPY SDELRQRLAA 201 RLEALKENGG ARLAEYHAKA TEHLSTLSEK AKPALEDLRQ GLLPVLESFK 251 VSFLSALEEY TKKLNTQ% D Figure 2. Analysis of spot NO.1 from human lung 2-DE map (ph 3 10) by MALDI-TOF MS. (A) MALDI-TOF MS peptide mass fingerprint spectrum obtained from crude peptide mixture after in-gel tryptic digest of spot NO.1. (B) Probability based mowse score (C) The list of matching peptides between the experimental and theoretical values. (D) The sequence of ApolipoproteinA-I identified. The matched peptides are shadowed in the sequenc 50
6 Huizhen Zhang et al. Table 1. MALDI-TOF-MS identification of overexpression proteins in lung cancer tissues Spot* AC** Matched Peptide Top score Theoretic Mr Sequence*** coverage (%) Name and description 1 P ApolipoproteinA-I precursor (Apo-AI) 2 gi apolipoprotein E precursor 3 P Ras-related protein Rab-14 4 P Calgranulin B (MRP-14) 5 P Peptidyl-prolyl cis-trans isomerase 6 P Cyclophilin A * The numbering and lettering corresponding to the two-dimensional gel electrophoresis image in Fig. 1. **Accession number in NCBInr, SWISS-PROT. ***Percentage of the protein sequence covered by the matched peptides. 3 Discussion Proteomic study means the analysis of the entire protein complement expressed by a genome [15]. Classically, proteome analysis consists of three steps: Two-dimensional gel electrophoresis (2-DE) used for proteins profiling, mass spectrometry (MS) for critical confirmation of the molecular weights and bioinformatics for protein identification. Comparative proteome analysis is a good strategy to discover proteins that undergo changes in expression level and may underlie the differences of phenotype [16]. 2-DE with its recent developments has been seen as an ideal tool for proteome analyses. Immobilized ph-gradient (IPG) strips are used in the first-dimensional gel electrophoresis to provide a basis for reproducible separation according to proteins isoelectric points. Advanced computer graphics and image analysis systems have offered a possibility for quantitative detection of protein spots, in addition to edition and storage of the 2-DE images. Proteome comparison between tissues in the different situations can cast a light on some protein spots, which are differently expressed in quantity under different surroundings. Nowadays, mass spectrometry such as matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF-MS) [17], can routinely identify and characterize proteins. Peptide mass fingerprinting identifies a protein based on the molecular weights of its peptides obtained by MS after digestion by trypsin [18]. The developed bioinformatics allows the identification of most proteins [19]. In this paper, we reported comparative proteome studies between normal lung tissues and cancerous lung tissues. The ph 3 10 range in the first-dimensional gel electrophoresis separates the soluble proteins in the lung tissues. All paired samples, which would be compared each other, were run together at the same conditions. This avoids any artificial differences between samples due to the different running conditions. The 2-DE patterns of normal lung tissues and cancerous lung tissues were compared each other with the help of the software PDQest, and different expressional protein spots were found. These protein spots have a consistent increase in all 8 independent paired samples. Over-expressed protein spots were excised from the gel and digested with trypsin. Molecular weights of the tryptic peptides were determined by MALDI-TOF-MS. Obtained protein scores are significant (P < 0.05) in the protein database search. 19% 72% coverage of protein sequences was obtained in the analyses of MALDI-TOF MS. Number 4 and number 6, were identified as Calgranulin B (S100A9, MRP14) and Calgizzarin (S100A11, S100C protein), two members of the S100 family. S100 proteins comprise a family of 1 14 kda EF-hand-containing calcium binding proteins that function to transmit calcium-dependent cell regulatory 51
7 Life Science Journal, Vol 6, No 3, 2009 signals, which are specifically expressed in a variety of tissues and cell types. It is thought that they are primarily involved in Ca 2+ -mediated signal transduction. Changes in intracellular calcium levels alter the structure and function of these proteins [20]. They have been frequently described in association with cell growth and differentiation, with cell cycle regulation and with several human diseases such as cancer and skin disorders. Studies suggest that these members of the S-100 protein family have a role in cytoskeletal changes seen in various skin diseases. Calgizzarin( S100A11)is a calcium-binding protein implicated in a variety of biologic functions such as proliferation and differentiation in cancer. Calgizzarin, a new tumor marker protein in nasopharyngeal carcinoma (NPC) [21],plays a significant role in tumor suppression and it is involved in prostate cancer development and progression. Calgranulin B represents a novel molecular parameters of the early events of inflammatory reactions that reveal interesting aspects for the pathomechanism of chronic inflammatory reactions [22]. Tumor specific protein markers were identified in colon tumors as calgranulin B that is also upregulated in colorectal cancer [23,24]. Calgizzarin and Calgranulin B present in lung cancer has not be documented. Apolipoprotein A-I (apoa-i) is the major protein constituent of high density lipoproteins (HDL) and lymph hylomicrons. In human, proapoa-i is synthesized as a precursor protein, preproapoa-i, of 267 amino acids and is thought to occupy a surface position on the lipoprotein. ApoA-I activates lecithin-cholesterol acyltransferase, which is the cholesterol-esterifying enzyme of plasma involved in the production of mature circulating HDL [25]. Apolipoprotein E (apoe), a protein with three common isoforms, has a large impact on longevity and successful aging. Impairments in cognitive performance have been observed in aged apolipoprotein (apoe)-deficient mice [26]. In this study, apolipoprotein (apo)a-i and Apolipoprotein E are found to be up-regulated in lung cancer. Cyclophilins (CyPs) are a large class of highly conserved ubiquitous peptidyl-prolyl cis-trans isomerases. CyPs have also been identified as a specific receptor for the immunosuppressive drug cyclosporin A and are involved in a variety of biological functions. Cyclophilin A (CypA) is a cytosolic protein that has 52 many biological functions including immune modulation, cell growth, tumorigenesis, and vascular disease. The objective of this study was to determine the effect of CypA on cell proliferation and several gene expressions in human endothelial cells and vascular smooth muscle cells [27]. Cyclophilin A (CyPA) is overexpressed in non-small cell lung cancer (NSCLC), but it was not found that CyPA is of prognostic significance [28].This paper reported Cyclophilin A (CypA) overexprssed in lung cancer,but it s function of a biomarker for lung cancer need further study to validate. In conclusion, the differences of the proteins between normal and cancerous lung tissues are complex. To affirm proteins studying above be cancer-associated of lung cancer and to serve for the further basic and clinical investigation, we need do some other works such as enlarging sample, purifying and analyzing these differential proteins, and study the actions during the multistage process of lung cancer. Acknowledgements This work was finished in Henan Key Laboratory for Molecular Medicine. This work was supported by the key disciplines granted from the national "211 Project" and the National Natural Science Foundation of China (No ). References 1. Greenlee RT, Murray T, Bolden S, et al. Cancer statistics. CA Cancer J Clin, 2000, 50: Kamp DW, Graceffa P, Pryor WA, et al. The role of free radicals in asbestos-induced diseases. Free Radical Biol Med, 1999,12: Weitzman SA, Graceffa P. Asbestos catalyzes hydroxyl and superoxide radical generation from hydrogen peroxide. Arch Biochem Biophys, 1984,228: Bailey-Wilson JE, Pugh EW, Wiest JS, et al. Lung cancer: genetic epidemiology. In:Bertino JR,editor. Encyclopedia of cancer.san Diego, CA:Academic Press, Lindblom A, Liljegren A. Tumor markers in malignancies. Br Med J, 2000,320: Stieber P, Aronsson AC, Bialk P, et al. Tumor markers in lung cancer: EGTM recommendations. Anticancer Res, 1999,19:
8 Huizhen Zhang et al. 7. Lawrie LC, Fothergill JE, Murray GI. Spot the differences: proteomics in cancer research. Lancet Oncol, 2001,2: Simpson RJ, Dorow DS. Cancer proteomics: signaling networks to tumor markers. Trends Biotechnol, 2001,19(10 Suppl):S40 S48 9. Bichsel VE, Liotta LA, Petricoin EF 3rd. Cancer protoemics: from biomarker discovery to signal pathway profiling. Cancer J, 2001,7(1): Gorg A, Obermaier C, Boguth G, et al. The current state twodimensional electrophoresis with immobilized ph gradients. Electrophoresis, 2000,21: Lahm HW, Langen H. Mass spectrometry: for the identification of proteins separated by gels. Electrophoresis, 2000,21: Okuzawa K, Franzen B, Lindholm J, et al. Characterization of gene expression in clinical lung cancer materials by two-dimensional polyacrylamide gel electrophoresis. Electrophoresis, 1994,15: Gygi SP, Corthals GL, Zhang Y, Rochon Y, Aebersold R. Evaluation of two-dimensional gel electrophoresis-based proteome analysis technology. Proc Natl Acad Sci U S A, 2000, 97: Ni XG, Zhao P, Liu Y, et al. Application of proteomic approach for solid tumor marker discovery. Aizheng[Chinese journal of cancer], 2003, 22: Wilkins MR, Sanchez JC, Gooley AA, et al. Progress with proteome projects: why all proteins expressed by a genome should be identified and how to do it. Biotechnol Genet Eng Rev, 1996,13: O Farrel PZ. High-resolution two-dimensional electrophoresis of proteins. J Biol Chem, 1975,250: Nordhoff E, Egelhofer V, Giavalisco P, et al. Large-gel two-dimensional electrophoresis-matrix-assisted laser desorption/ionization time of flight mass spectrometry: an analytical challenge for studying complex protein mixtures. Electrophoresis, 2001,22(14): spectrometric molecular weight information to identify proteins in sequence databases. Biol Mass Spectrom, 1993,22: Tripathi KK. Bioinformatics: the foundation of present and future biotechnology. Curr Sci, 2000,79(5): Ruse M, Lambert A, Robinson N, et al. S100A7, S100A10, and S100A11 are transglutaminase substrates. Biochemistry, 2001,40(10): Fung LF, Lo AK, Yuen PW, et al. Differential gene expression in nasopharyngeal carcinoma cells. Life Sci, 2000, 67(8): Sorg C. The calcium binding proteins MRP8 and MRP14 in acute and chronic inflammation. Behring Inst Mitt, 1992,(91): Chaurand P, DaGue BB, Pearsall RS, et al. Profiling proteins from azoxymethane-induced colon tumors at the molecular level by matrix-assisted laser desorption/ionization mass spectrometry. Proteomics, 2001,1(10): Jungblut PR, Zimny-Arndt U, Zeindl-Eberhart E, et al Proteomics in human disease: cancer, heart and infectious diseases. Electrophoresis, 1999,20(10): Ghiselli G, Gotto AM, Tanenbaum S, et al. Proapolipoprotein A-I conversion kinetics in vivo in human and in rat. Proc Natl Acad Sci U S A, 1985,82: Law A, Gauthier S, Quirion R. Alteration of nitric oxide synthase activity in young and aged apolipoprotein E-deficient mice. Neurobiol Aging, 2003,24 (1): Huang LL, Zhao XM, Huang CQ, et al. Structure of recombinant human cyclophilin J, a novel member of the cyclophilin family. Acta Crystallogr D Biol Crystallogr, 2005,61(Pt 3): Howard BA, Zheng Z, Campa MJ, et al.translating biomarkers into clinical practice: prognostic implications of cyclophilin A and macrophage migratory inhibitory factor identified from protein expression profiles in non-small cell lung cancer. Lung Cancer, 2004,46(3): Mann M, Hojrup P, Roepstorff P. Use of mass 53
Supplementary material: Materials and suppliers
 Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Agilent Protein In-Gel Tryptic Digestion Kit
 Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
In-Gel Tryptic Digestion Kit
 INSTRUCTIONS In-Gel Tryptic Digestion Kit 3747 N. Meridian Road P.O. Box 117 Rockford, IL 61105 89871 1468.2 Number Description 89871 In-Gel Tryptic Digestion Kit, sufficient reagents for approximately
INSTRUCTIONS In-Gel Tryptic Digestion Kit 3747 N. Meridian Road P.O. Box 117 Rockford, IL 61105 89871 1468.2 Number Description 89871 In-Gel Tryptic Digestion Kit, sufficient reagents for approximately
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
Trypsin Digestion Mix
 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name 239PR Trypsin Digestion Mix Provides optimal buffered conditions for in gel trypsin digestion
G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name 239PR Trypsin Digestion Mix Provides optimal buffered conditions for in gel trypsin digestion
Improve Protein Analysis with the New, Mass Spectrometry- Compatible ProteasMAX Surfactant
 Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging
 Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Proteomics Grade. Protocol. Catalog # Agilent Technologies. Research Use Only. Not for use in Diagnostic Procedures. Version A, January 2010
 Proteomics Grade Trypsin Catalog #204310 Protocol Version A, January 2010 Research Use Only. Not for use in Diagnostic Procedures. Agilent Technologies Notices Agilent Technologies, Inc. 2010 No part of
Proteomics Grade Trypsin Catalog #204310 Protocol Version A, January 2010 Research Use Only. Not for use in Diagnostic Procedures. Agilent Technologies Notices Agilent Technologies, Inc. 2010 No part of
Mass Spectrometry. Mass spectrometer MALDI-TOF ESI/MS/MS. Basic components. Ionization source Mass analyzer Detector
 Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
PosterREPRINT INTRODUCTION. 2-D PAGE of Mouse Liver Samples. 2-D PAGE of E.coli Samples. Digestion / Cleanup. EXPERIMENTAL 1-D PAGE of BSA Samples
 INTRODUCTION Identification and characterization of low abundance proteins separated by 2D gel electrophoresis is complicated by two important factors; the use of suitable staining techniques for the visualization
INTRODUCTION Identification and characterization of low abundance proteins separated by 2D gel electrophoresis is complicated by two important factors; the use of suitable staining techniques for the visualization
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
Introduction. Methods RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES. TCW Poon *, HLY Chan, HWC Leung, A Lo, RHY Lau, AY Hui, JJY Sung
 RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Liver specific glycoforms of serum proteins in chronic hepatitis B infection: identification by lectin affinity chromatography and quantitative proteomic
RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Liver specific glycoforms of serum proteins in chronic hepatitis B infection: identification by lectin affinity chromatography and quantitative proteomic
Mammalian Membrane Protein Extraction Kit
 Mammalian Membrane Protein Extraction Kit Catalog number: AR0155 Boster s Mammalian Membrane Protein Extraction Kit is a simple, rapid and reproducible method to prepare cellular protein fractions highly
Mammalian Membrane Protein Extraction Kit Catalog number: AR0155 Boster s Mammalian Membrane Protein Extraction Kit is a simple, rapid and reproducible method to prepare cellular protein fractions highly
Combination of 2-D Gel and Liquid-Phase Electrophoretic Separations as Proteomic Tools in Neuroscience. Analytical 2-D. Electrophoresis.
 electrophoresis tech note 2859 Combination of 2-D Gel and Liquid-Phase Electrophoretic Separations as Proteomic Tools in Neuroscience Pia Davidsson, Department of Clinical Neuroscience, Experimental Neuroscience
electrophoresis tech note 2859 Combination of 2-D Gel and Liquid-Phase Electrophoretic Separations as Proteomic Tools in Neuroscience Pia Davidsson, Department of Clinical Neuroscience, Experimental Neuroscience
Biological Mass Spectrometry. April 30, 2014
 Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Matrix Assisted Laser Desorption Ionization Time-of-flight Mass Spectrometry
 Matrix Assisted Laser Desorption Ionization Time-of-flight Mass Spectrometry Time-of-Flight Mass Spectrometry. Basic principles An attractive feature of the time-of-flight (TOF) mass spectrometer is its
Matrix Assisted Laser Desorption Ionization Time-of-flight Mass Spectrometry Time-of-Flight Mass Spectrometry. Basic principles An attractive feature of the time-of-flight (TOF) mass spectrometer is its
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for
 Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
DetergentOUT Tween. DetergentOUT GBS10. OrgoSol DetergentOUT
 252PR 01 G-Biosciences, St Louis, MO. USA 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents
252PR 01 G-Biosciences, St Louis, MO. USA 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents
PosterREPRINT A NOVEL APPROACH TO MALDI-TOF-MS SAMPLE PREPARATION. Presented at ABRF 2002, Austin, Texas, USA, 9th - 12th March 2002.
 Introduction A NOVEL APPROACH TO MALDI-TOF-MS SAMPLE PREPARATION Ed Bouvier 2, Jeff Brown 1, Emmanuelle Claude 1, John L. Gebler 2, Weibin Chen 2, *Dominic Gostick 1, Kevin Howes 1, James Langridge 1,
Introduction A NOVEL APPROACH TO MALDI-TOF-MS SAMPLE PREPARATION Ed Bouvier 2, Jeff Brown 1, Emmanuelle Claude 1, John L. Gebler 2, Weibin Chen 2, *Dominic Gostick 1, Kevin Howes 1, James Langridge 1,
Problem-solving Test: The Mechanism of Protein Synthesis
 Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
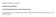 Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
Supplementary Information. Top-down/bottom-up mass spectrometry workflow using dissolvable polyacrylamide gels
 Supplementary Information Top-down/bottom-up mass spectrometry workflow using dissolvable polyacrylamide gels Nobuaki Takemori,,* Ayako Takemori,, Piriya Wongkongkathep, Michael Nshanian, Rachel R. Ogorzalek
Supplementary Information Top-down/bottom-up mass spectrometry workflow using dissolvable polyacrylamide gels Nobuaki Takemori,,* Ayako Takemori,, Piriya Wongkongkathep, Michael Nshanian, Rachel R. Ogorzalek
The Detection of Allergens in Food Products with LC-MS
 The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples:
 Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Isolation of pure cell populations from healthy and
 Direct Analysis of Laser Capture Microdissected Cells by MALDI Mass Spectrometry Baogang J. Xu and Richard M. Caprioli Department of Chemistry, Vanderbilt University, Nashville, Tennessee, USA Melinda
Direct Analysis of Laser Capture Microdissected Cells by MALDI Mass Spectrometry Baogang J. Xu and Richard M. Caprioli Department of Chemistry, Vanderbilt University, Nashville, Tennessee, USA Melinda
Mass Spectrometry. - Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications
 - Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications Adapted from Mass Spectrometry in Biotechnology Gary Siuzdak,, Academic Press 1996 1 Introduction
- Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications Adapted from Mass Spectrometry in Biotechnology Gary Siuzdak,, Academic Press 1996 1 Introduction
Microalbuminuric Diabetic patients N=18
 ONLINE APPENDIX Table A1 Clinical and laboratory features of healthy subjects, type 2 diabetic patients and NDCKD patients enrolled in the present study. Healthy Subjects N= 2 Normoalbuminuric Diabetic
ONLINE APPENDIX Table A1 Clinical and laboratory features of healthy subjects, type 2 diabetic patients and NDCKD patients enrolled in the present study. Healthy Subjects N= 2 Normoalbuminuric Diabetic
MALDI Imaging Mass Spectrometry
 MALDI Imaging Mass Spectrometry Nan Kleinholz Mass Spectrometry and Proteomics Facility The Ohio State University Mass Spectrometry and Proteomics Workshop What is MALDI Imaging? MALDI: Matrix Assisted
MALDI Imaging Mass Spectrometry Nan Kleinholz Mass Spectrometry and Proteomics Facility The Ohio State University Mass Spectrometry and Proteomics Workshop What is MALDI Imaging? MALDI: Matrix Assisted
Protocol for Gene Transfection & Western Blotting
 The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
MALDI-TOF. Introduction. Schematic and Theory of MALDI
 MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Introduction to Proteomics 1.0
 Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
TECHNICAL BULLETIN. R 2 GlcNAcβ1 4GlcNAcβ1 Asn
 GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow
 Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
(III) MALDI instrumentation
 Dr. Sanjeeva Srivastava (I) Basics of MALDI-TF (II) Sample preparation In-gel digestion Zip-tip sample clean-up Matrix and sample plating (III) MALDI instrumentation 2 1 (I) Basics of MALDI-TF Analyte
Dr. Sanjeeva Srivastava (I) Basics of MALDI-TF (II) Sample preparation In-gel digestion Zip-tip sample clean-up Matrix and sample plating (III) MALDI instrumentation 2 1 (I) Basics of MALDI-TF Analyte
2D protein gel analysis tool for studying the protein phosphorylation changes associated with contraction and relaxation of vascular smooth muscle
 2D protein gel analysis tool for studying the protein phosphorylation changes associated with contraction and relaxation of vascular smooth muscle Internship details Where: Department of Bioengineering
2D protein gel analysis tool for studying the protein phosphorylation changes associated with contraction and relaxation of vascular smooth muscle Internship details Where: Department of Bioengineering
MALDI-IMS (MATRIX-ASSISTED LASER DESORPTION/IONIZATION
 MALDI-IMS (MATRIX-ASSISTED LASER DESORPTION/IONIZATION IMAGING MASS SPECTROMETER) IN TISSUE STUDY YANXIAN CHEN MARCH 8 TH, WEDNESDAY. SEMINAR FOCUSING ON What is MALDI imaging mass spectrometer? How does
MALDI-IMS (MATRIX-ASSISTED LASER DESORPTION/IONIZATION IMAGING MASS SPECTROMETER) IN TISSUE STUDY YANXIAN CHEN MARCH 8 TH, WEDNESDAY. SEMINAR FOCUSING ON What is MALDI imaging mass spectrometer? How does
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
ProteaseMAX Surfactant, Trypsin Enhancer
 Technical Bulletin ProteaseMAX Surfactant, Trypsin Enhancer INSTRUCTIONS FOR USE OF PRODUCTS V2071 AND V2072. PRINTED IN USA. Revised 1/10 ProteaseMAX Surfactant, Trypsin Enhancer All technical literature
Technical Bulletin ProteaseMAX Surfactant, Trypsin Enhancer INSTRUCTIONS FOR USE OF PRODUCTS V2071 AND V2072. PRINTED IN USA. Revised 1/10 ProteaseMAX Surfactant, Trypsin Enhancer All technical literature
Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 2011
 Supporting Information Experimental details 1. Materials Ferric chloride (FeCl 3 6H 2 O), Ethylene glycol (EG) and Ammonium hydroxide (NH 3 H 2 O) were obtained from Beijing Chemical Regent Co. Ltd. (Beijing,
Supporting Information Experimental details 1. Materials Ferric chloride (FeCl 3 6H 2 O), Ethylene glycol (EG) and Ammonium hydroxide (NH 3 H 2 O) were obtained from Beijing Chemical Regent Co. Ltd. (Beijing,
Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g.
 Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g., casein more interestingly, in most other cases, it is a transient
Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g., casein more interestingly, in most other cases, it is a transient
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Peptide sequencing using chemically assisted fragmentation (CAF) and Ettan MALDI-ToF Pro mass spectrometry
 application note Ett MALDI-ToF Pro Peptide sequencing using chemically assisted fragmentation (CAF) d Ett MALDI-ToF Pro mass spectrometry Key words: MALDI-ToF, PSD, peptide sequencing, chemically assisted
application note Ett MALDI-ToF Pro Peptide sequencing using chemically assisted fragmentation (CAF) d Ett MALDI-ToF Pro mass spectrometry Key words: MALDI-ToF, PSD, peptide sequencing, chemically assisted
Components of a Mass Spectrometer
 Components of a Mass Spectrometer Sample Introduction Inlet GC LC Direct Insertion (Syringe/Probe) Ionization Ion Separation Ion Detection Ion Source EI,CI,,, MALDI Mass Analyzer Under vacuum TOF, Quadrupole,
Components of a Mass Spectrometer Sample Introduction Inlet GC LC Direct Insertion (Syringe/Probe) Ionization Ion Separation Ion Detection Ion Source EI,CI,,, MALDI Mass Analyzer Under vacuum TOF, Quadrupole,
Annexure III SOLUTIONS AND REAGENTS
 Annexure III SOLUTIONS AND REAGENTS A. STOCK SOLUTIONS FOR DNA ISOLATION 0.5M Ethylene-diamine tetra acetic acid (EDTA) (ph=8.0) 1M Tris-Cl (ph=8.0) 5M NaCl solution Red cell lysis buffer (10X) White cell
Annexure III SOLUTIONS AND REAGENTS A. STOCK SOLUTIONS FOR DNA ISOLATION 0.5M Ethylene-diamine tetra acetic acid (EDTA) (ph=8.0) 1M Tris-Cl (ph=8.0) 5M NaCl solution Red cell lysis buffer (10X) White cell
DetergentOUT Detergent Removal Systems
 252PR-04 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents from Peptide
252PR-04 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents from Peptide
Proteins. Amino acids, structure and function. The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka
 Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Heterogeneity Analysis of the Human Pituitary Proteome
 Clinical Chemistry 49:10 1740 1751 (2003) Oak Ridge Conference Heterogeneity Analysis of the Human Pituitary Proteome Xianquan Zhan 1 and Dominic M. Desiderio 1,2,3* Background: A human proteome is relatively
Clinical Chemistry 49:10 1740 1751 (2003) Oak Ridge Conference Heterogeneity Analysis of the Human Pituitary Proteome Xianquan Zhan 1 and Dominic M. Desiderio 1,2,3* Background: A human proteome is relatively
Universal sample preparation method for proteome analysis
 nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
Ion Source. Mass Analyzer. Detector. intensity. mass/charge
 Proteomics Informatics Overview of spectrometry (Week 2) Ion Source Analyzer Detector Peptide Fragmentation Ion Source Analyzer 1 Fragmentation Analyzer 2 Detector b y Liquid Chromatography (LC)-MS/MS
Proteomics Informatics Overview of spectrometry (Week 2) Ion Source Analyzer Detector Peptide Fragmentation Ion Source Analyzer 1 Fragmentation Analyzer 2 Detector b y Liquid Chromatography (LC)-MS/MS
The distribution of log 2 ratio (H/L) for quantified peptides. cleavage sites in each bin of log 2 ratio of quantified. peptides
 Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Supporting information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Glycan Reductive Isotope-coded Amino Acid Labeling (GRIAL) for Mass Spectrometry-based
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Glycan Reductive Isotope-coded Amino Acid Labeling (GRIAL) for Mass Spectrometry-based
How many proteins can be identified in a 2DE gel spot within an analysis of a complex human cancer tissue proteome?
 Electrophoresis 2018, 39, 965 980 965 Xianquan Zhan 1,2,3,4 Haiyan Yang 1,2,3 Fang Peng 1,2,3 Jianglin Li 5 Yun Mu 1,2,3 Ying Long 1,2,3 Tingting Cheng 1,2,3 Yuda Huang 1,2,3 Zhao Li 6 Miaolong Lu 1,2,3
Electrophoresis 2018, 39, 965 980 965 Xianquan Zhan 1,2,3,4 Haiyan Yang 1,2,3 Fang Peng 1,2,3 Jianglin Li 5 Yun Mu 1,2,3 Ying Long 1,2,3 Tingting Cheng 1,2,3 Yuda Huang 1,2,3 Zhao Li 6 Miaolong Lu 1,2,3
Application of Two-Dimensional Electrophoresis in the Research of Retinal Proteins of Diabetic Rat
 Cellular & Molecular Immunology 65 Article Application of Two-Dimensional Electrophoresis in the Research of Retinal Proteins of Diabetic Rat Shangqing Liu 1, 2, Yanyan Zhang 2, Xianyong Xie 2, Weiming
Cellular & Molecular Immunology 65 Article Application of Two-Dimensional Electrophoresis in the Research of Retinal Proteins of Diabetic Rat Shangqing Liu 1, 2, Yanyan Zhang 2, Xianyong Xie 2, Weiming
Up to now, gel electrophoresis remains the most powerful
 pubs.acs.org/ac Polyacrylamide Gel with Switchable Trypsin Activity for Analysis of Proteins Fangjie Liu,, Mingliang Ye,*, Chunli Wang,, Zhengyan Hu,, Yi Zhang,, Hongqiang Qin,, Kai Cheng,, and Hanfa Zou*,
pubs.acs.org/ac Polyacrylamide Gel with Switchable Trypsin Activity for Analysis of Proteins Fangjie Liu,, Mingliang Ye,*, Chunli Wang,, Zhengyan Hu,, Yi Zhang,, Hongqiang Qin,, Kai Cheng,, and Hanfa Zou*,
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
The Schedule and the Manual of Basic Techniques for Cell Culture
 The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
5 Identification of Binding Partners of the Annexin A2 / P11 Complex by Chemical Cross-Linking
 5 Identification of Binding Partners of the Annexin A2 / P11 Complex by Chemical Cross-Linking In the quest of the omics sciences for holistic schemes, the identification of binding partners of proteins
5 Identification of Binding Partners of the Annexin A2 / P11 Complex by Chemical Cross-Linking In the quest of the omics sciences for holistic schemes, the identification of binding partners of proteins
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
In-Solution Digestion for proteomics
 In-Solution Digestion for proteomics Guidelines for sample preparation (How to protect your samples from contamination with keratin) 1. Try to avoid any contact of samples and solutions with dust, skin
In-Solution Digestion for proteomics Guidelines for sample preparation (How to protect your samples from contamination with keratin) 1. Try to avoid any contact of samples and solutions with dust, skin
DetergentOUT GBS10 Spin Plates
 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT GBS10 Spin Plates 96-Well Plates for the Removal of Detergents from Peptide
G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT GBS10 Spin Plates 96-Well Plates for the Removal of Detergents from Peptide
Metabolomics: quantifying the phenotype
 Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
NOTES & TIPS. Evaluation of the Efficiency of In-Gel Digestion of Proteins by Peptide Isotopic Labeling and MALDI Mass Spectrometry
 All articles available online at http://www.idealibrary.com on NOTES & TIPS Evaluation of the Efficiency of In-Gel Digestion of Proteins by Peptide Isotopic Labeling and MALDI Mass Spectrometry Anna Shevchenko
All articles available online at http://www.idealibrary.com on NOTES & TIPS Evaluation of the Efficiency of In-Gel Digestion of Proteins by Peptide Isotopic Labeling and MALDI Mass Spectrometry Anna Shevchenko
Tissue and Fluid Proteomics Chao-Cheng (Sam) Wang
 Tissue and Fluid Proteomics Chao-Cheng (Sam) Wang UAB-03/09/2004 What is proteomics? A snap shot of the protein pattern!! What can proteomics do? To provide information on functional networks and/or involvement
Tissue and Fluid Proteomics Chao-Cheng (Sam) Wang UAB-03/09/2004 What is proteomics? A snap shot of the protein pattern!! What can proteomics do? To provide information on functional networks and/or involvement
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates
 HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
ASSAY OF SPHINGOMYELINASE ACTIVITY
 ASSAY OF SPHINGOMYELINASE ACTIVITY Protocol for Protein Extraction Stock Solution 1. Leupeptin/hydrochloride (FW 463.0,
ASSAY OF SPHINGOMYELINASE ACTIVITY Protocol for Protein Extraction Stock Solution 1. Leupeptin/hydrochloride (FW 463.0,
Protein MultiColor Stable, Low Range
 Product Name: DynaMarker Protein MultiColor Stable, Low Range Code No: DM670L Lot No: ******* Size: 200 μl x 3 (DM670 x 3) (120 mini-gel lanes) Storage: 4 C Stability: 12 months at 4 C Storage Buffer:
Product Name: DynaMarker Protein MultiColor Stable, Low Range Code No: DM670L Lot No: ******* Size: 200 μl x 3 (DM670 x 3) (120 mini-gel lanes) Storage: 4 C Stability: 12 months at 4 C Storage Buffer:
Systematic analysis of protein-detergent complexes applying dynamic light scattering to optimize solutions for crystallization trials
 Supporting information 1 2 3 Volume 71 (2015) Supporting information for article: 4 5 6 7 8 Systematic analysis of protein-detergent complexes applying dynamic light scattering to optimize solutions for
Supporting information 1 2 3 Volume 71 (2015) Supporting information for article: 4 5 6 7 8 Systematic analysis of protein-detergent complexes applying dynamic light scattering to optimize solutions for
Rapid Detection and Identification of Bacteria in Milk Products by Matrix-Assisted Laser Desorption/Ionization Tandem Mass Spectrometry (MALDI-MS/MS)
 Rapid Detection and Identification of Bacteria in Milk Products by Matrix-Assisted Laser Desorption/Ionization Tandem Mass Spectrometry (MALDI-MS/MS) Nyesa Enakaya It is essential to be able to identify
Rapid Detection and Identification of Bacteria in Milk Products by Matrix-Assisted Laser Desorption/Ionization Tandem Mass Spectrometry (MALDI-MS/MS) Nyesa Enakaya It is essential to be able to identify
Proteomic analysis of water insoluble proteins from normal and cataractous human lenses
 Received 22 January 2007 Accepted 30 August 2007 Published 14 September 2007 Proteomic analysis of water insoluble proteins from normal and cataractous human lenses V. Harrington, 1 O.P. Srivastava, 2
Received 22 January 2007 Accepted 30 August 2007 Published 14 September 2007 Proteomic analysis of water insoluble proteins from normal and cataractous human lenses V. Harrington, 1 O.P. Srivastava, 2
SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons
 LECTROPHORES/S Revised November 1992 SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons I NTRODUCTION Electrophoresis in polyacrylamide gels in the presence of sodium dodecyl sulfate (SDS), an anionic detergent,
LECTROPHORES/S Revised November 1992 SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons I NTRODUCTION Electrophoresis in polyacrylamide gels in the presence of sodium dodecyl sulfate (SDS), an anionic detergent,
Chapter 10apter 9. Chapter 10. Summary
 Chapter 10apter 9 Chapter 10 The field of proteomics has developed rapidly in recent years. The essence of proteomics is to characterize the behavior of a group of proteins, the system rather than the
Chapter 10apter 9 Chapter 10 The field of proteomics has developed rapidly in recent years. The essence of proteomics is to characterize the behavior of a group of proteins, the system rather than the
Determination of Tetracyclines in Chicken by Solid-Phase Extraction and High-Performance Liquid Chromatography
 Determination of Tetracyclines in Chicken by Solid-Phase Extraction and High-Performance Liquid Chromatography Application ote Food Safety Authors Chen-Hao Zhai and Yun Zou Agilent Technologies Co. Ltd.
Determination of Tetracyclines in Chicken by Solid-Phase Extraction and High-Performance Liquid Chromatography Application ote Food Safety Authors Chen-Hao Zhai and Yun Zou Agilent Technologies Co. Ltd.
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Supporting Information for:
 Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Determination of 6-Chloropicolinic Acid (6-CPA) in Crops by Liquid Chromatography with Tandem Mass Spectrometry Detection. EPL-BAS Method No.
 Page 1 of 10 Determination of 6-Chloropicolinic Acid (6-CPA) in Crops by Liquid Chromatography with Tandem Mass Spectrometry Detection EPL-BAS Method No. 205G881B Method Summary: Residues of 6-CPA are
Page 1 of 10 Determination of 6-Chloropicolinic Acid (6-CPA) in Crops by Liquid Chromatography with Tandem Mass Spectrometry Detection EPL-BAS Method No. 205G881B Method Summary: Residues of 6-CPA are
Preadipocyte factor 1 induces pancreatic ductal cell differentiation into insulin-producing cells
 Supplementary Information Preadipocyte factor 1 induces pancreatic ductal cell differentiation into insulin-producing cells Marie Rhee, Seung-Hwan Lee, Ji-Won Kim, Dong-Sik Ham, Heon-Seok Park, Hae Kyung
Supplementary Information Preadipocyte factor 1 induces pancreatic ductal cell differentiation into insulin-producing cells Marie Rhee, Seung-Hwan Lee, Ji-Won Kim, Dong-Sik Ham, Heon-Seok Park, Hae Kyung
Supporting Information
 Supporting Information Cyclic Peptidyl Inhibitors against Human Peptidyl-Prolyl Isomerase Pin1 Tao Liu, Yu Liu, Hung-Ying Kao, *,, and Dehua Pei Table of Contents: Table S1. Structures of amino acid building
Supporting Information Cyclic Peptidyl Inhibitors against Human Peptidyl-Prolyl Isomerase Pin1 Tao Liu, Yu Liu, Hung-Ying Kao, *,, and Dehua Pei Table of Contents: Table S1. Structures of amino acid building
Babu Antharavally, Ryan Bomgarden, and John Rogers Thermo Fisher Scientific, Rockford, IL
 A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
Proteomics to identify novel biomarkers and therapeutic targets in cardiovascular disease
 Proteomics to identify novel biomarkers and therapeutic targets in cardiovascular disease Markus Kubicek Laboratorios de Biologia Vascular y Proteínómica Estructural, Instituto de Biomedicina de Valencia
Proteomics to identify novel biomarkers and therapeutic targets in cardiovascular disease Markus Kubicek Laboratorios de Biologia Vascular y Proteínómica Estructural, Instituto de Biomedicina de Valencia
Structural vs. nonstructural proteins
 Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
The study of phospholipids in single cells using an integrated microfluidic device
 Supporting Information: The study of phospholipids in single cells using an integrated microfluidic device combined with matrix-assisted laser desorption/ionization mass spectrometry Weiyi Xie,, Dan Gao,
Supporting Information: The study of phospholipids in single cells using an integrated microfluidic device combined with matrix-assisted laser desorption/ionization mass spectrometry Weiyi Xie,, Dan Gao,
HDL Purification Kit (Ultracentrifugation Free)
 Product Manual HDL Purification Kit (Ultracentrifugation Free) Catalog Number STA- 607 10 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lipoproteins are submicroscopic particles
Product Manual HDL Purification Kit (Ultracentrifugation Free) Catalog Number STA- 607 10 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lipoproteins are submicroscopic particles
Quantitative chromatin proteomics reveals a dynamic histone. post-translational modification landscape that defines asexual
 Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
PHOSPHOPEPTIDE ANALYSIS USING IMAC SAMPLE PREPARATION FOLLOWED BY MALDI-MS and MALDI PSD MX
 PHOSPHOPEPTIDE ANALYSIS USING IMAC SAMPLE PREPARATION FOLLOWED BY MALDI-MS and MALDI PSD MX E. Claude 1, E. Emekwue 2, M. Snel 1, T. McKenna 1 and J. Langridge 1 1: Waters Corporation, Manchester, UK 2:
PHOSPHOPEPTIDE ANALYSIS USING IMAC SAMPLE PREPARATION FOLLOWED BY MALDI-MS and MALDI PSD MX E. Claude 1, E. Emekwue 2, M. Snel 1, T. McKenna 1 and J. Langridge 1 1: Waters Corporation, Manchester, UK 2:
Chapter 3. Structure of Enzymes. Enzyme Engineering
 Chapter 3. Structure of Enzymes Enzyme Engineering 3.1 Introduction With purified protein, Determining M r of the protein Determining composition of amino acids and the primary structure Determining the
Chapter 3. Structure of Enzymes Enzyme Engineering 3.1 Introduction With purified protein, Determining M r of the protein Determining composition of amino acids and the primary structure Determining the
Minute TM Plasma Membrane Protein Isolation and Cell Fractionation Kit User Manual (v5)
 Minute TM Plasma Membrane Protein Isolation and Cell Fractionation Kit Catalog number: SM-005 Description Minute TM plasma membrane (PM) protein isolation kit is a novel and patented native PM protein
Minute TM Plasma Membrane Protein Isolation and Cell Fractionation Kit Catalog number: SM-005 Description Minute TM plasma membrane (PM) protein isolation kit is a novel and patented native PM protein
FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE.
 User Protocol 539790 Rev. 28 March 05 JSW Page 1 of 17 ProteoExtract Subcellular Proteome Extraction Kit Cat. No. 539790 User Protocol 539790 Rev. 28 March 05 JSW Page 2 of 17 1. Introduction One major
User Protocol 539790 Rev. 28 March 05 JSW Page 1 of 17 ProteoExtract Subcellular Proteome Extraction Kit Cat. No. 539790 User Protocol 539790 Rev. 28 March 05 JSW Page 2 of 17 1. Introduction One major
Sulfate Radical-Mediated Degradation of Sulfadiazine by CuFeO 2 Rhombohedral Crystal-Catalyzed Peroxymonosulfate: Synergistic Effects and Mechanisms
 Supporting Information for Sulfate Radical-Mediated Degradation of Sulfadiazine by CuFeO 2 Rhombohedral Crystal-Catalyzed Peroxymonosulfate: Synergistic Effects and Mechanisms Submitted by Yong Feng, Deli
Supporting Information for Sulfate Radical-Mediated Degradation of Sulfadiazine by CuFeO 2 Rhombohedral Crystal-Catalyzed Peroxymonosulfate: Synergistic Effects and Mechanisms Submitted by Yong Feng, Deli
Author copy. Impact of un-polymerized acrylamide monomer residues onto protein identification by MALDI TOF MS. 1. Introduction
 Cent. Eur. J. Chem. 10(4) 2012 1073-1078 DOI: 10.2478/s11532-012-0024-3 Central European Journal of Chemistry Impact of un-polymerized acrylamide monomer residues onto protein identification by MALDI TOF
Cent. Eur. J. Chem. 10(4) 2012 1073-1078 DOI: 10.2478/s11532-012-0024-3 Central European Journal of Chemistry Impact of un-polymerized acrylamide monomer residues onto protein identification by MALDI TOF
Analysis of Triglycerides in Cooking Oils Using MALDI-TOF Mass Spectrometry and Principal Component Analysis
 Analysis of Triglycerides in Cooking Oils Using MALDI-TOF Mass Spectrometry and Principal Component Analysis Kevin Cooley Chemistry Supervisor: Kingsley Donkor 1. Abstract Triglycerides are composed of
Analysis of Triglycerides in Cooking Oils Using MALDI-TOF Mass Spectrometry and Principal Component Analysis Kevin Cooley Chemistry Supervisor: Kingsley Donkor 1. Abstract Triglycerides are composed of
Analytical Method for 2, 4, 5-T (Targeted to Agricultural, Animal and Fishery Products)
 Analytical Method for 2, 4, 5-T (Targeted to Agricultural, Animal and Fishery Products) The target compound to be determined is 2, 4, 5-T. 1. Instrument Liquid Chromatograph-tandem mass spectrometer (LC-MS/MS)
Analytical Method for 2, 4, 5-T (Targeted to Agricultural, Animal and Fishery Products) The target compound to be determined is 2, 4, 5-T. 1. Instrument Liquid Chromatograph-tandem mass spectrometer (LC-MS/MS)
Protocol for protein SDS PAGE and Transfer
 Protocol for protein SDS PAGE and Transfer According to Laemmli, (1970) Alaa El -Din Hamid Sayed, Alaa_h254@yahoo.com Serum Selection of a protein source cell cultures (bacteria, yeast, mammalian, etc.)
Protocol for protein SDS PAGE and Transfer According to Laemmli, (1970) Alaa El -Din Hamid Sayed, Alaa_h254@yahoo.com Serum Selection of a protein source cell cultures (bacteria, yeast, mammalian, etc.)
LDL/VLDL Purification Kit (Ultracentrifugation Free)
 Product Manual LDL/VLDL Purification Kit (Ultracentrifugation Free) Catalog Number STA- 606 10 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lipoproteins are submicroscopic
Product Manual LDL/VLDL Purification Kit (Ultracentrifugation Free) Catalog Number STA- 606 10 preps FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Lipoproteins are submicroscopic
Comparison of mass spectrometers performances
 Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Total Protein Extraction with TCA-Acetone
 TCA-Acetone Extraction 1 1 Total Protein Extraction with TCA-Acetone Valérie Méchin, Catherine Damerval, and Michel Zivy Summary We describe a procedure allowing extraction of total proteins that performs
TCA-Acetone Extraction 1 1 Total Protein Extraction with TCA-Acetone Valérie Méchin, Catherine Damerval, and Michel Zivy Summary We describe a procedure allowing extraction of total proteins that performs
Neosolaniol. [Methods listed in the Feed Analysis Standards]
![Neosolaniol. [Methods listed in the Feed Analysis Standards] Neosolaniol. [Methods listed in the Feed Analysis Standards]](/thumbs/86/94003482.jpg) Neosolaniol [Methods listed in the Feed Analysis Standards] 1 Simultaneous analysis of mycotoxins by liquid chromatography/ tandem mass spectrometry [Feed Analysis Standards, Chapter 5, Section 1 9.1 ]
Neosolaniol [Methods listed in the Feed Analysis Standards] 1 Simultaneous analysis of mycotoxins by liquid chromatography/ tandem mass spectrometry [Feed Analysis Standards, Chapter 5, Section 1 9.1 ]
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
