I. Setup. - Note that: autohgpec_v1.0 can work on Windows, Ubuntu and Mac OS.
|
|
|
- Barrie Hutchinson
- 5 years ago
- Views:
Transcription
1 autohgpec: Automated prediction of novel disease-gene and diseasedisease associations and evidence collection based on a random walk on heterogeneous network Duc-Hau Le 1,*, Trang T.H. Tran 1 1 School of Computer Science and Engineering, Thuyloi University, 175 Tay Son, Dong Da, Hanoi, Vietnam. * To whom correspondence should be addressed. User Manual 1
2 Table of Contents I. Setup... 3 II. Overview of autohgpec... 4 III. Case study: Prediction of novel breast cancer-associated genes and diseases Run autohgpec in Cytoscape... 5 Step 1: Construct a heterogeneous network... 5 Step 2: Select a disease of interest... 5 Step 3: Select candidate sets... 6 Step 4: Prioritize... 6 Step 5: Examine ranked genes and diseases... 7 Visualization... 7 Search Evidences Automate autohgpec using CyREST Command API Step 1: Construct a heterogeneous network Step 2: Select a disease of interest Step 3: Select candidate sets Step 4: Prioritize Step 5: Examine ranked genes and diseases Visualize Search Evidences Automate autohgpec from R Step 1: Construct a heterogeneous network Step 2: Select a disease of interest Step 3: Select candidate sets Step 4: Prioritize Step 5: Examine ranked genes and diseases Visualize Search Evidences IV. Reference
3 I. Setup - autohgpec 1.0 can only run on Cytoscape 3.6 (or later) platform, which have Automation features, therefore user should download this version at - Cytoscape need JRE to run, therefore download JRE version 7.x or later from and install it. - Install Cytoscape to the root folder (e.g., /Applications/Cytoscape_v3.6.0). - Download autohgpec_v1.0.jar file from or Then, install it by going to Apps à App Manager. After that, choose Install from file, then select the downloaded autohgpec_v1.0.jar file. - Create folders Data in the root folder of Cytoscape (e.g., /Applications/Cytoscape_v3.6.0). - Download GO annotation data at ftp.ncbi.nlm.nih.gov/gene/data/gene2go.gz, then extract and store in the Data folder (e.g., /Applications/Cytoscape_v3.6.0/Data). - Note that: autohgpec_v1.0 can work on Windows, Ubuntu and Mac OS. 3
4 II. Overview of autohgpec After installing, autohgpec will be automatically loaded in the App menu of Cytoscape The main tasks (Prediction of Genes and Diseases, and Evidence Collection) of autohgpec are completed after five steps: - Step 1: Construct a Heterogeneous network - Step 2: Select a disease of interest (including 2 sub steps) o 1. Select a disease o 2. Create training list - Step 3: Provide Candidate Gene Set (including 4 options) o All remaining genes in the Gene network o Neighbors of training genes in Chromosome o Neighbors of training genes in Gene network o Susceptible Chromosome Regions/Bands - Step 4: Prioritize (candidate genes and diseases) - Step 5: Examine Ranked Genes and Diseases o Search Evidences o Visualize These five steps can be performed - In Cytoscape like HGPEC (Le and Pham, 2017) - Using CyREST Command API - From R statistics ( 4
5 III. Case study: Prediction of novel breast cancer-associated genes and diseases In the following section, we show the ability of autohpec in identifying novel breast cancer-associated genes and diseases. 1. Run autohgpec in Cytoscape Step 1: Construct a heterogeneous network To this end, we select a phenotypic disease similarity network containing 5,080 diseases and 19,729 interactions (i.e., Disease_Similarity_Network_5) and a human protein interaction network containing 10,486 genes and 50,791 interactions (i.e., Default_Human_PPI_Network). Then, we connect them by known disease-gene associations from either OMIM (Amberger, et al., 2009) or DisGeNET (Piñero, et al., 2017) to construct a heterogeneous network of diseases and genes by clicking Apps à autohgpec à Step 1: Construct a Heterogeneous Network To construct a heterogeneous network: 1. Select a disease similarity network. 2. Select known disease-gene associations 3. Select a network of genes/proteins (e.g., the preinstalled one or one imported from Cytoscape). 4. Click OK to connect these two networks by the known disease-gene associations. Note that: - For disease similarity network: We pre-installed 3 networks corresponding to 5, 10 or 15 nearest neighbors, which were extracted from a phenotypic disease similarity matrix data collected from (van Driel, et al., 2006) - For gene/protein interaction network: o We pre-installed a human physical protein interaction network collected from ftp://ftp.ncbi.nlm.nih.gov/gene/generif/interactions.gz. o However, user can use other protein/gene interaction networks by importing them to Cytoscape (File à Import à Network from table (Text/MS Excel) ). Genes/Proteins in the network must be identified by Gene Entrez ID. - For known disease-gene associations: User can select from either OMIM or DisGeNET Step 2: Select a disease of interest We select breast cancer (OMIM ID: ), then create training list by click menu Apps à autohgpec à Step 2: Select a disease of interest - Step 2.1. Select a disease: Apps à autohgpec à Step 2: Select a disease of interest à 1. Select a disease Enter disease keyword to retrieve a list of disease phenotypes from OMIM. Here are 4 phenotypes from OMIM related to breast cancer - Step 2.2. Create Training List: Apps à autohgpec à Step 2: Select a disease of interest à 2. Create Training List 5
6 Here are the training lists include the disease of interest (OMIM ID: ) and its 21 known associated genes. The disease of interest (OMIM ID: ) A total of 21 known associated genes Step 3: Select candidate sets For candidate diseases, all remaining diseases are specified as candidate diseases by default. Therefore, there are 5,079 diseases in this set. For candidate genes, select menu Apps à autohgpec à Step 3: Provide Candidate Gene Set, then we select option All remaining genes in Gene Network. As a result, a total of 10,465 remaining genes were selected as candidate genes. With option All remaining genes in Gene Network, 10,465 remaining genes were selected as candidate genes Four ways to construct a candidate gene set: - Neighbors of Training Genes in Gene Network o User must define distance of neighbors to training genes - Neighbors Of Training Genes in Chromosome (also known as Artificial Linkage Interval) o User must define number of neighbors of each training gene in the same chromosome. - All remaining genes in Gene Network - Susceptible Chromosome Regions/Bands o User selects candidate genes from susceptible chromosome regions/bands. Step 4: Prioritize We set three parameters (i.e., back-probability (γ), jumping probability (l) and subnetwork (Disease/Gene) importance (h)) of RWRH algorithm to 0.5, 0.6 and 0.7, respectively. Please refer to (Li and Patra, 2010) for best parameter setting. Select menu Select menu Apps à autohgpec à Step 4: Prioritize 6
7 Then click OK to rank all candidate genes and diseases in the heterogeneous network. All genes and diseases are ranked and listed in two data tables Note that, not only candidate genes and diseases are ranked, but all genes and diseases in the heterogeneous network are also ranked. Therefore, user can visualize them in one view to exploit their topologically relationships. Ranked Genes Ranked Diseases Step 5: Examine ranked genes and diseases Visualize and search evidences for highly ranked genes and diseases shown in two above data can be done by selecting menu Apps à autohgpec à Step 5: Examine Ranked Genes and Diseases Visualization Not only candidate genes and diseases are ranked, but all genes and diseases in the heterogeneous network are also ranked. Therefore, user can visualize them in one view to exploit their topologically relationships. - Visualize the topological relationships between highly ranked candidate genes and the disease of interest. For example: If we focus on topological relationships between highly ranked candidate genes and disease of interest and its associated genes, we selected top 20 ranked candidate genes, 21 training genes as above and the training disease (i.e., OMIM ID: ) for visualization o Select top 20 ranked candidate genes and 21 training genes. o Select the training disease (i.e., the disease of interest, OMIM ID: ) 7
8 o Select sub-menu Apps à autohgpec à Step 5: Examine Ranked Genes and Diseases à 2. Visualize o Select Layout à Group Attributes Layout à Role Node in rhombus shape is the disease of interest. Nodes with high rankings are in red, relative high are in pink, medium are in white and light green, low are in green. We found that the sub-network is mostly connected. In other words, highly ranked genes are directly connected to known/training genes - Visualize the topological relationships between highly ranked candidate diseases and the disease of interest. In this case, we selected top 20 ranked candidate diseases, 21 training genes and the disease of interest (i.e., OMIM ID: ) for visualization. o Select top 21 ranked candidate genes o Select top 20 ranked candidate diseases and the disease of interest 8
9 o Select sub-menu Apps à autohgpec à Step 5: Examine Ranked Genes and Diseases à 2. Visualize o Select Layout à Group Attributes Layout à Role Node in rhombus shape is the disease of interest. Nodes in rectangle shape are candidate diseases. Nodes with high rankings are in red, relative high are in pink, medium are in white and light green, low are in green Similarly, we found that the sub-network is connected. In other words, highly ranked candidate diseases are directly connected to either known/training genes or the disease of interest. This means that candidate diseases which have connections to the disease of interest or associated with training genes are highly ranked. 9
10 Search Evidences This function is to collect evidences and annotations for associations between highly ranked candidate genes/diseases and the disease of interest. Ranked genes Select a set of 20 ranked candidate genes Then, select menu Apps à autohgpec à Step 5: Examine Ranked Genes and Diseases à 1. Search Evidences. Here are the genes with annotations and evidences Ranked diseases Select a set of top 20 ranked candidate diseases for annotation and evidence collection Then, select menu Apps à autohgpec à Step 5: Examine Ranked Genes and Diseases à 1. Search Evidences Here are the diseases with annotations and evidences 10
11 2. Automate autohgpec using CyREST Command API Select menu Help à Automation à CyREST Command API Here is list of commands to run autohgpec To predict novel breast cancer-associated genes and diseases, we need to perform 5 following steps: Step 1: Construct a heterogeneous network Use Example Value then press Try it out! to create a heterogeneous network of diseases and genes, including a phenotypic disease similarity network containing 5,080 diseases and 19,729 interactions (i.e., Disease_Similarity_Network_5), a human protein interaction network containing 10,486 genes and 50,791 interactions (i.e., Default_Human_PPI_Network) and known disease-gene associations from either OMIM (Amberger, et al., 2009). 11
12 Step 2: Select a disease of interest Input breast cancer by using Example Value then press Try it out! to retrieve a list of disease phenotypes from OMIM 12
13 Here are 4 phenotypes from OMIM related to breast cancer 13
14 Select OMIM ID: by using Example Value then pressing Try it out! to create training lists Retrieve a list of 21 training genes Here are the training lists include the disease of interest (OMIM ID: ) and its 21 known associated genes in Cytoscape. 14
15 The disease of interest (OMIM ID: ) A total of 21 known associated genes Step 3: Select candidate sets Four ways to construct a candidate gene set: Op Candidate Set t 1 Neighbors of Training Genes in Gene Network User must define distance of neighbors to training genes 2 Neighbors Of Training Genes in Chromosome (also known as Artificial Linkage Interval) User must define number of neighbors of each training gene in the same chromosome. 3 All remaining genes in Gene Network 4 Susceptible Chromosome Regions/Bands User selects candidate genes from susceptible chromosome regions/bands. For this case study, we selected option 3: 15
16 16
17 à A total of 10,465 remaining genes were selected as candidate genes in Cytoscape Step 4: Prioritize Use Example Value, then press Try it out! to rank all genes and disease phenotypes in the heterogeneous network 17
18 Ranked Genes in Cytoscape Ranked Diseases in Cytoscape 18
19 Step 5: Examine ranked genes and diseases Visualize - Select ranked genes and diseases in the two above data tables to visualize, then press Try it out! To visualize selected genes and diseases in the heterogeneous network. See the results in the section Visualize in Step 5 of Run autohgpec in Cytoscape 19
20 Search Evidences - Select highly ranked candidate genes and diseases in the two above data tables, then press Try it out! to search evidences See the results in the section Search Evidences in Step 5 of Run autohgpec in Cytoscape 3. Automate autohgpec from R Make sure appropriate libraries are installed and they are functional. - Please run check-library-installation.r for libs and tests: - Please run check-cytoscape-connection-autohgpec.r for tests and initial demo: - List available commands of autohgpec in R: > commandhelp('autohgpec') [1] "Available commands for 'autohgpec':" [1] "step1_construct_network" "step2_1_select_disease" "step2_2_create_training_list" "step3_pcg_allremaining" 20
21 [5] "step3_pcg_nbchromosome" "step3_pcg_nbnetwork" "step3_pcg_suscepchromo" "step4_prioritize" [9] "step5_1_search_evidences" "step5_2_visualize" To predict novel breast cancer-associated genes and diseases, we need to perform 5 following steps in R: Step 1: Construct a heterogeneous network Use command step1_construct_network to create a heterogeneous network of diseases and genes, including a phenotypic disease similarity network containing 5,080 diseases and 19,729 interactions (i.e., Disease_Similarity_Network_5), a human protein interaction network containing 10,486 genes and 50,791 interactions (i.e., Default_Human_PPI_Network) and known disease-gene associations from either OMIM (Amberger, et al., 2009). - List available arguments of command step1_construct_network of autohgpec in R > commandhelp('autohgpec step1_construct_network') [1] "Available arguments for 'autohgpec step1_construct_network':" [1] "DiseaseGene" "diseasenetwork" "genenetwork" - Run the command to build a heterogeneous network > commandrun('autohgpec step1_construct_network DiseaseGene="Disease-gene from OMIM" diseasenetwork="disease_similarity_network_5" genenetwork="default_human_ppi_network"') [1] "Build Heterogeneous Network successfully" Step 2: Select a disease of interest Step 2.1: Select a disease Use command step2_1_select_disease - List available arguments of command step2_1_select_disease of autohgpec in R > commandhelp('autohgpec step2_1_select_disease') [1] "Available arguments for 'autohgpec step2_1_select_disease':" [1] "diseasename" - Run the command to retrieve a list of disease phenotypes from OMIM. It will return 4 phenotypes from OMIM related to breast cancer > commandrun('autohgpec step2_1_select_disease diseasename="breast cancer"') Step 2.2: Create training lists Use command step2_2_create_training_list - List available arguments of command step2_2_create_training_list of autohgpec in R > commandhelp('autohgpec step2_2_create_training_list') [1] "Available arguments for 'autohgpec step2_2_create_training_list':" [1] "diseasetraining" - Run the command to retrieve a list of training genes and disease phenotypes (OMIM ID: ) > commandrun('autohgpec step2_2_create_training_list diseasetraining="mim114480"') Here are the training lists include the disease of interest (OMIM ID: ) and its 21 known associated genes. The disease of interest (OMIM ID: ) A total of 21 known associated genes Step 3: Select candidate sets Four ways to construct a candidate gene set: Opt Candidate Set R Commands 1 Neighbors of Training Genes in Gene Network > commandhelp('autohgpec step3_pcg_nbnetwork') [1] "Available arguments for 'autohgpec 21
22 User must define distance of neighbors to training genes 2 Neighbors Of Training Genes in Chromosome (also known as Artificial Linkage Interval) User must define number of neighbors of each training gene in the same chromosome. 3 All remaining genes in Gene Network 4 Susceptible Chromosome Regions/Bands User selects candidate genes from susceptible chromosome regions/bands. step3_pcg_nbnetwork':" [1] "distance" > commandrun('autohgpec step3_pcg_nbnetwork distance=1') > commandhelp('autohgpec step3_pcg_nbchromosome') [1] "Available arguments for 'autohgpec step3_pcg_nbchromosome':" [1] "distance" "seedgene" > commandrun('autohgpec step3_pcg_nbchromosome distance=99 seedgene="all') > commandhelp('autohgpec step3_pcg_allremaining') > commandrun('autohgpec step3_pcg_allremaining') > commandhelp('autohgpec step3_pcg_suscepchromo') > commandrun('autohgpec step3_pcg_suscepchromo') For this case study, we selected option 3: > commandrun('autohgpec step3_pcg_allremaining') à A total of 10,465 remaining genes were selected as candidate genes Step 4: Prioritize Use command step4_prioritize - List available arguments of command step4_prioritize of autohgpec in R > commandhelp('autohgpec step4_prioritize') [1] "Available arguments for 'autohgpec step4_prioritize':" [1] "backprob" "jumpprob" "subnetweight" - Run the command with parameters to rank all genes and disease phenotypes in the heterogeneous network > commandrun('autohgpec step4_prioritize backprob=0.5 jumpprob=0.6 subnetweight=0.7') Ranked Genes Ranked Diseases 22
23 Step 5: Examine ranked genes and diseases Visualize - Select ranked candidate genes and diseases in the two above data tables to visualize, then use command step5_2_visualize > commandrun('autohgpec step5_2_visualize') See the results in the section Visualize in Step 5 of Run autohgpec in Cytoscape Search Evidences - Select highly ranked candidate genes and diseases in the two above data tables to search evidences, then use command step5_1_search_evidences > commandrun('autohgpec step5_1_search_evidences') See the results in the section Search Evidences in Step 5 of Run autohgpec in Cytoscape 23
24 IV. Reference Amberger, J., et al. McKusick's Online Mendelian Inheritance in Man (OMIM ). Nucleic Acids Research 2009;37(suppl 1):D793-D796. Le, D.-H. and Pham, V.-H. HGPEC: a Cytoscape app for prediction of novel disease-gene and disease-disease associations and evidence collection based on a random walk on heterogeneous network. BMC Systems Biology 2017;11(1):61. Li, Y. and Patra, J.C. Genome-wide inferring gene-phenotype relationship by walking on the heterogeneous network. Bioinformatics 2010;26(9): Piñero, J., et al. DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Research 2017;45(D1):D833-D839. van Driel, M.A., et al. A text-mining analysis of the human phenome. Eur J Hum Genet 2006;14(5):
Duc-Hau Le 1,2 and Van-Huy Pham 3*
 Le and Pham BMC Systems Biology (2017) 11:61 DOI 10.1186/s12918-017-0437-x SOFTWARE HGPEC: a Cytoscape app for prediction of novel disease-gene and disease-disease associations and evidence collection
Le and Pham BMC Systems Biology (2017) 11:61 DOI 10.1186/s12918-017-0437-x SOFTWARE HGPEC: a Cytoscape app for prediction of novel disease-gene and disease-disease associations and evidence collection
Supplementary Figure 1
 Supplementary Figure 1 An example of the gene-term-disease network automatically generated by Phenolyzer web server for 'autism'. The largest word represents the user s input term, Autism. The pink round
Supplementary Figure 1 An example of the gene-term-disease network automatically generated by Phenolyzer web server for 'autism'. The largest word represents the user s input term, Autism. The pink round
Clay Tablet Connector for hybris. User Guide. Version 1.5.0
 Clay Tablet Connector for hybris User Guide Version 1.5.0 August 4, 2016 Copyright Copyright 2005-2016 Clay Tablet Technologies Inc. All rights reserved. All rights reserved. This document and its content
Clay Tablet Connector for hybris User Guide Version 1.5.0 August 4, 2016 Copyright Copyright 2005-2016 Clay Tablet Technologies Inc. All rights reserved. All rights reserved. This document and its content
Data mining with Ensembl Biomart. Stéphanie Le Gras
 Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Evaluating Classifiers for Disease Gene Discovery
 Evaluating Classifiers for Disease Gene Discovery Kino Coursey Lon Turnbull khc0021@unt.edu lt0013@unt.edu Abstract Identification of genes involved in human hereditary disease is an important bioinfomatics
Evaluating Classifiers for Disease Gene Discovery Kino Coursey Lon Turnbull khc0021@unt.edu lt0013@unt.edu Abstract Identification of genes involved in human hereditary disease is an important bioinfomatics
P-B-54.30/141. Instrument Cluster SCN Coding for Component Replacement or Dealer Installed Accessories:
 Date: August 2005 Order No.: Supersedes: Group: 54 P-B-54.30/141 SUBJECT: Model 171.454/456/473 All Model Years A. Introduction Instrument Cluster SCN Coding for Component Replacement or Dealer Installed
Date: August 2005 Order No.: Supersedes: Group: 54 P-B-54.30/141 SUBJECT: Model 171.454/456/473 All Model Years A. Introduction Instrument Cluster SCN Coding for Component Replacement or Dealer Installed
Exercises: Differential Methylation
 Exercises: Differential Methylation Version 2018-04 Exercises: Differential Methylation 2 Licence This manual is 2014-18, Simon Andrews. This manual is distributed under the creative commons Attribution-Non-Commercial-Share
Exercises: Differential Methylation Version 2018-04 Exercises: Differential Methylation 2 Licence This manual is 2014-18, Simon Andrews. This manual is distributed under the creative commons Attribution-Non-Commercial-Share
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Fully Automated IFA Processor LIS User Manual
 Fully Automated IFA Processor LIS User Manual Unless expressly authorized, forwarding and duplication of this document is not permitted. All rights reserved. TABLE OF CONTENTS 1 OVERVIEW... 4 2 LIS SCREEN...
Fully Automated IFA Processor LIS User Manual Unless expressly authorized, forwarding and duplication of this document is not permitted. All rights reserved. TABLE OF CONTENTS 1 OVERVIEW... 4 2 LIS SCREEN...
Managing and Taking Notes
 When scheduling a meeting, the host can specify the default note-taking options that take effect once the meeting starts. During a meeting, the presenter can change the default note-taking options at any
When scheduling a meeting, the host can specify the default note-taking options that take effect once the meeting starts. During a meeting, the presenter can change the default note-taking options at any
The application can be accessed from the pull down menu by selecting ODOT > Drafting Apps > Signs, or by the following key-in command:
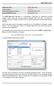 Application Name: Current version: Required MicroStation Version: Required GEOPAK Version: ODOT_Signs.mvba V14.07.18 MicroStation XM or V8i GEOPAK XM or V8i The ODOT_Signs.mvba application is a MicroStation
Application Name: Current version: Required MicroStation Version: Required GEOPAK Version: ODOT_Signs.mvba V14.07.18 MicroStation XM or V8i GEOPAK XM or V8i The ODOT_Signs.mvba application is a MicroStation
TMWSuite. DAT Interactive interface
 TMWSuite DAT Interactive interface DAT Interactive interface Using the DAT Interactive interface Using the DAT Interactive interface... 1 Setting up the system to use the DAT Interactive interface... 1
TMWSuite DAT Interactive interface DAT Interactive interface Using the DAT Interactive interface Using the DAT Interactive interface... 1 Setting up the system to use the DAT Interactive interface... 1
CSDplotter user guide Klas H. Pettersen
 CSDplotter user guide Klas H. Pettersen [CSDplotter user guide] [0.1.1] [version: 23/05-2006] 1 Table of Contents Copyright...3 Feedback... 3 Overview... 3 Downloading and installation...3 Pre-processing
CSDplotter user guide Klas H. Pettersen [CSDplotter user guide] [0.1.1] [version: 23/05-2006] 1 Table of Contents Copyright...3 Feedback... 3 Overview... 3 Downloading and installation...3 Pre-processing
Managing and Taking Notes
 When scheduling a meeting, the host can specify the default note-taking options that take effect once the meeting starts. During a meeting, the presenter can change the default note-taking options at any
When scheduling a meeting, the host can specify the default note-taking options that take effect once the meeting starts. During a meeting, the presenter can change the default note-taking options at any
NeuroLink by Applied Neuroscience, Inc. Help Manual Applied Neuroscience, Inc. Applied Neuroscience, Inc.
 NeuroLink by Applied Neuroscience, Inc. Help Manual Applied Neuroscience, Inc. 8200 Bryan Dairy Road, Suite 315 Largo, FL 33777-1355 For More Information Contact: info@anineurolink.com For Support Contact:
NeuroLink by Applied Neuroscience, Inc. Help Manual Applied Neuroscience, Inc. 8200 Bryan Dairy Road, Suite 315 Largo, FL 33777-1355 For More Information Contact: info@anineurolink.com For Support Contact:
Module 3: Pathway and Drug Development
 Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
User Guide. Association analysis. Input
 User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
Chapter 9. Tests, Procedures, and Diagnosis Codes The McGraw-Hill Companies, Inc. All rights reserved.
 Chapter 9 Tests, Procedures, and Diagnosis Codes Chapter 9 Content: Overview Ordering A Test SpringLabsTM & Reference Lab Results Managing and Charting Tests Creating A New Test Documenting and Activating
Chapter 9 Tests, Procedures, and Diagnosis Codes Chapter 9 Content: Overview Ordering A Test SpringLabsTM & Reference Lab Results Managing and Charting Tests Creating A New Test Documenting and Activating
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Phenotype analysis in humans using OMIM
 Outline: 1) Introduction to OMIM 2) Phenotype similarity map 3) Exercises Phenotype analysis in humans using OMIM Rosario M. Piro Molecular Biotechnology Center University of Torino, Italy 1 MBC, Torino
Outline: 1) Introduction to OMIM 2) Phenotype similarity map 3) Exercises Phenotype analysis in humans using OMIM Rosario M. Piro Molecular Biotechnology Center University of Torino, Italy 1 MBC, Torino
Qualys PC/SCAP Auditor
 Qualys PC/SCAP Auditor Getting Started Guide November 15, 2017 COPYRIGHT 2011-2017 BY QUALYS, INC. ALL RIGHTS RESERVED. QUALYS AND THE QUALYS LOGO ARE REGISTERED TRADEMARKS OF QUALYS, INC. ALL OTHER TRADEMARKS
Qualys PC/SCAP Auditor Getting Started Guide November 15, 2017 COPYRIGHT 2011-2017 BY QUALYS, INC. ALL RIGHTS RESERVED. QUALYS AND THE QUALYS LOGO ARE REGISTERED TRADEMARKS OF QUALYS, INC. ALL OTHER TRADEMARKS
Vega: Variational Segmentation for Copy Number Detection
 Vega: Variational Segmentation for Copy Number Detection Sandro Morganella Luigi Cerulo Giuseppe Viglietto Michele Ceccarelli Contents 1 Overview 1 2 Installation 1 3 Vega.RData Description 2 4 Run Vega
Vega: Variational Segmentation for Copy Number Detection Sandro Morganella Luigi Cerulo Giuseppe Viglietto Michele Ceccarelli Contents 1 Overview 1 2 Installation 1 3 Vega.RData Description 2 4 Run Vega
Lionbridge Connector for Hybris. User Guide
 Lionbridge Connector for Hybris User Guide Version 2.1.0 November 24, 2017 Copyright Copyright 2017 Lionbridge Technologies, Inc. All rights reserved. Published in the USA. March, 2016. Lionbridge and
Lionbridge Connector for Hybris User Guide Version 2.1.0 November 24, 2017 Copyright Copyright 2017 Lionbridge Technologies, Inc. All rights reserved. Published in the USA. March, 2016. Lionbridge and
Pathway Exercises Metabolism and Pathways
 1. Find the metabolic pathway for glycolysis. For this exercise use PlasmoDB.org Pathway Exercises Metabolism and Pathways a. Navigate to the search page for Identify Metabolic Pathways based on Pathway
1. Find the metabolic pathway for glycolysis. For this exercise use PlasmoDB.org Pathway Exercises Metabolism and Pathways a. Navigate to the search page for Identify Metabolic Pathways based on Pathway
ImageJ plugin for semiautomatic measurement of roentgenological attachment level
 ImageJ plugin for semiautomatic measurement of roentgenological attachment level User manual and installation instructions v. 1.1, June 2015 The plugin is written by Gerald R. Torgersen 1 based on the
ImageJ plugin for semiautomatic measurement of roentgenological attachment level User manual and installation instructions v. 1.1, June 2015 The plugin is written by Gerald R. Torgersen 1 based on the
IBRIDGE 1.0 USER MANUAL
 IBRIDGE 1.0 USER MANUAL Jaromir Krizek CONTENTS 1 INTRODUCTION... 3 2 INSTALLATION... 4 2.1 SYSTEM REQUIREMENTS... 5 2.2 STARTING IBRIDGE 1.0... 5 3 MAIN MENU... 6 3.1 MENU FILE... 6 3.2 MENU SETTINGS...
IBRIDGE 1.0 USER MANUAL Jaromir Krizek CONTENTS 1 INTRODUCTION... 3 2 INSTALLATION... 4 2.1 SYSTEM REQUIREMENTS... 5 2.2 STARTING IBRIDGE 1.0... 5 3 MAIN MENU... 6 3.1 MENU FILE... 6 3.2 MENU SETTINGS...
Quick guide for 3shape order form
 Quick guide for 3shape order form Elos Accurate Library Elos Accurate Library December 2017 Content: Introduction 2 Elos Accurate Hybrid Base Kit order form 3 Elos Accurate Hybrid Base Bridge order form
Quick guide for 3shape order form Elos Accurate Library Elos Accurate Library December 2017 Content: Introduction 2 Elos Accurate Hybrid Base Kit order form 3 Elos Accurate Hybrid Base Bridge order form
CNV PCA Search Tutorial
 CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
Hour 2: lm (regression), plot (scatterplots), cooks.distance and resid (diagnostics) Stat 302, Winter 2016 SFU, Week 3, Hour 1, Page 1
 Agenda for Week 3, Hr 1 (Tuesday, Jan 19) Hour 1: - Installing R and inputting data. - Different tools for R: Notepad++ and RStudio. - Basic commands:?,??, mean(), sd(), t.test(), lm(), plot() - t.test()
Agenda for Week 3, Hr 1 (Tuesday, Jan 19) Hour 1: - Installing R and inputting data. - Different tools for R: Notepad++ and RStudio. - Basic commands:?,??, mean(), sd(), t.test(), lm(), plot() - t.test()
Dosimeter Setting Device
 Instruction Manual Dosimeter Setting Device For Electronic Personal Dosimeter Dose-i (Unit:Sv, Version:1.05 English) WTA529748 a 1 / 38 Foreword Thank you for purchasing the Dosimeter Setting Device; a
Instruction Manual Dosimeter Setting Device For Electronic Personal Dosimeter Dose-i (Unit:Sv, Version:1.05 English) WTA529748 a 1 / 38 Foreword Thank you for purchasing the Dosimeter Setting Device; a
Set Up SOS Video Chat and Screen-Sharing
 Set Up SOS Video Chat and Screen-Sharing Salesforce, Spring 17 @salesforcedocs Last updated: March 11, 2017 Copyright 2000 2017 salesforce.com, inc. All rights reserved. Salesforce is a registered trademark
Set Up SOS Video Chat and Screen-Sharing Salesforce, Spring 17 @salesforcedocs Last updated: March 11, 2017 Copyright 2000 2017 salesforce.com, inc. All rights reserved. Salesforce is a registered trademark
Variant Classification. Author: Mike Thiesen, Golden Helix, Inc.
 Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Cardiac Agatston Scoring
 NA-MIC Cardiac Agatston Scoring Jessica Forbes, Hans Johnson University of Iowa Jessica-Forbes@uiowa.edu NA-MIC Tutorial Contest: Summer 2014 2014, All Rights Reserved Learning Objective This tutorial
NA-MIC Cardiac Agatston Scoring Jessica Forbes, Hans Johnson University of Iowa Jessica-Forbes@uiowa.edu NA-MIC Tutorial Contest: Summer 2014 2014, All Rights Reserved Learning Objective This tutorial
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16
 38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
NERVE ACTION POTENTIAL SIMULATION version 2013 John Cornell
 NERVE ACTION POTENTIAL SIMULATION version 2013 John Cornell http://www.jccornell.net In 1963 Alan Hodgkin and Andrew Huxley received the Nobel Prize in Physiology and Medicine for their work on the mechanism
NERVE ACTION POTENTIAL SIMULATION version 2013 John Cornell http://www.jccornell.net In 1963 Alan Hodgkin and Andrew Huxley received the Nobel Prize in Physiology and Medicine for their work on the mechanism
BlueBayCT - Warfarin User Guide
 BlueBayCT - Warfarin User Guide December 2012 Help Desk 0845 5211241 Contents Getting Started... 1 Before you start... 1 About this guide... 1 Conventions... 1 Notes... 1 Warfarin Management... 2 New INR/Warfarin
BlueBayCT - Warfarin User Guide December 2012 Help Desk 0845 5211241 Contents Getting Started... 1 Before you start... 1 About this guide... 1 Conventions... 1 Notes... 1 Warfarin Management... 2 New INR/Warfarin
Modularized Random Walk with Restart for Candidate Disease Genes Prioritization
 The Third International Symposium on Optimization and Systems Biology (OSB 09) Zhangjiajie, China, September 20 22, 2009 Copyright 2009 ORSC & APORC, pp. 353 360 Modularized Random Walk with Restart for
The Third International Symposium on Optimization and Systems Biology (OSB 09) Zhangjiajie, China, September 20 22, 2009 Copyright 2009 ORSC & APORC, pp. 353 360 Modularized Random Walk with Restart for
A framework for the study of diseases and adverse drug reactions
 A framework for the study of diseases and adverse drug reactions Laura I. Furlong IBI group Research Programme on Biomedical Informatics (GRIB) Hospital del Mar Research Institute (IMIM) Information on
A framework for the study of diseases and adverse drug reactions Laura I. Furlong IBI group Research Programme on Biomedical Informatics (GRIB) Hospital del Mar Research Institute (IMIM) Information on
NMR. Sample preparation. and Analysis
 NMR Sample preparation and Analysis Preparing the NMR Sample The NMR solvent is Chloroform-d ( CDCl 3 ) The solvent is stored in the fridge, please return after use The 5mm NMR tubes and caps are at a
NMR Sample preparation and Analysis Preparing the NMR Sample The NMR solvent is Chloroform-d ( CDCl 3 ) The solvent is stored in the fridge, please return after use The 5mm NMR tubes and caps are at a
Appendix B. Nodulus Observer XT Instructional Guide. 1. Setting up your project p. 2. a. Observation p. 2. b. Subjects, behaviors and coding p.
 1 Appendix B Nodulus Observer XT Instructional Guide Sections: 1. Setting up your project p. 2 a. Observation p. 2 b. Subjects, behaviors and coding p. 3 c. Independent variables p. 4 2. Carry out an observation
1 Appendix B Nodulus Observer XT Instructional Guide Sections: 1. Setting up your project p. 2 a. Observation p. 2 b. Subjects, behaviors and coding p. 3 c. Independent variables p. 4 2. Carry out an observation
USER GUIDE: NEW CIR APP. Technician User Guide
 USER GUIDE: NEW CIR APP. Technician User Guide 0 Table of Contents 1 A New CIR User Interface Why?... 3 2 How to get started?... 3 3 Navigating the new CIR app. user interface... 6 3.1 Introduction...
USER GUIDE: NEW CIR APP. Technician User Guide 0 Table of Contents 1 A New CIR User Interface Why?... 3 2 How to get started?... 3 3 Navigating the new CIR app. user interface... 6 3.1 Introduction...
Literature databases OMIM
 Literature databases OMIM Online Mendelian Inheritance in Man OMIM OMIM is a database that catalogues all the known diseases with a genetic component, and when possible links them to the relevant genes
Literature databases OMIM Online Mendelian Inheritance in Man OMIM OMIM is a database that catalogues all the known diseases with a genetic component, and when possible links them to the relevant genes
Instructions for the ECN201 Project on Least-Cost Nutritionally-Adequate Diets
 Instructions for the ECN201 Project on Least-Cost Nutritionally-Adequate Diets John P. Burkett October 15, 2015 1 Overview For this project, each student should (a) determine her or his nutritional needs,
Instructions for the ECN201 Project on Least-Cost Nutritionally-Adequate Diets John P. Burkett October 15, 2015 1 Overview For this project, each student should (a) determine her or his nutritional needs,
DENTRIX ENTERPRISE 8.0.5
 DENTRIX ENTERPRISE 8.0. GETTING STARTED WITH THE CURRENT CLINICAL NOTES www.dentrixenterprise.com -800-DSCHEIN Getting Started with the Current Clinical Notes Working with Clinical Notes Keeping accurate
DENTRIX ENTERPRISE 8.0. GETTING STARTED WITH THE CURRENT CLINICAL NOTES www.dentrixenterprise.com -800-DSCHEIN Getting Started with the Current Clinical Notes Working with Clinical Notes Keeping accurate
Allergy Basics. This handout describes the process for adding and removing allergies from a patient s chart.
 Allergy Basics This handout describes the process for adding and removing allergies from a patient s chart. Accessing Allergy Information Page 1 Recording No Known Medication Allergies Page 2 Recording
Allergy Basics This handout describes the process for adding and removing allergies from a patient s chart. Accessing Allergy Information Page 1 Recording No Known Medication Allergies Page 2 Recording
Guide to Use of SimulConsult s Phenome Software
 Guide to Use of SimulConsult s Phenome Software Page 1 of 52 Table of contents Welcome!... 4 Introduction to a few SimulConsult conventions... 5 Colors and their meaning... 5 Contextual links... 5 Contextual
Guide to Use of SimulConsult s Phenome Software Page 1 of 52 Table of contents Welcome!... 4 Introduction to a few SimulConsult conventions... 5 Colors and their meaning... 5 Contextual links... 5 Contextual
LabVIEW PROFIBUS VISA Driver DP-Master
 LabVIEW PROFIBUS VISA Driver DP-Master Getting Started V1.35 27.04.2017 Project No.: 5303 Doc-ID.: LabVIEW PROFIBUS VISA Driver KUNBUS d:\project\5302_df_profi_ii\anwenderdoku\labview\version 1.35\gettingstarted_win_dp-master_e.doc
LabVIEW PROFIBUS VISA Driver DP-Master Getting Started V1.35 27.04.2017 Project No.: 5303 Doc-ID.: LabVIEW PROFIBUS VISA Driver KUNBUS d:\project\5302_df_profi_ii\anwenderdoku\labview\version 1.35\gettingstarted_win_dp-master_e.doc
University of Alaska Connected! FAQs
 University of Alaska Connected! FAQs 1. What is Connected? Connected! allows employees and spouses/fips to connect a fitness device or app to Healthyroads.com. This will allow additional tracking options
University of Alaska Connected! FAQs 1. What is Connected? Connected! allows employees and spouses/fips to connect a fitness device or app to Healthyroads.com. This will allow additional tracking options
OECD QSAR Toolbox v.4.2. An example illustrating RAAF scenario 6 and related assessment elements
 OECD QSAR Toolbox v.4.2 An example illustrating RAAF scenario 6 and related assessment elements Outlook Background Objectives Specific Aims Read Across Assessment Framework (RAAF) The exercise Workflow
OECD QSAR Toolbox v.4.2 An example illustrating RAAF scenario 6 and related assessment elements Outlook Background Objectives Specific Aims Read Across Assessment Framework (RAAF) The exercise Workflow
BIOL 458 BIOMETRY Lab 7 Multi-Factor ANOVA
 BIOL 458 BIOMETRY Lab 7 Multi-Factor ANOVA PART 1: Introduction to Factorial ANOVA ingle factor or One - Way Analysis of Variance can be used to test the null hypothesis that k or more treatment or group
BIOL 458 BIOMETRY Lab 7 Multi-Factor ANOVA PART 1: Introduction to Factorial ANOVA ingle factor or One - Way Analysis of Variance can be used to test the null hypothesis that k or more treatment or group
CURRICULUM VITA OF Xiaowen Chen
 Xiaowen Chen College of Bioinformatics Science and Technology Harbin Medical University Harbin, 150086, P. R. China Mobile Phone:+86 13263501862 E-mail: hrbmucxw@163.com Education Experience 2008-2011
Xiaowen Chen College of Bioinformatics Science and Technology Harbin Medical University Harbin, 150086, P. R. China Mobile Phone:+86 13263501862 E-mail: hrbmucxw@163.com Education Experience 2008-2011
Titrations in Cytobank
 The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
Demo Mode. Once you have taken the time to navigate your RPM 2 app in "Demo mode" you should be ready to pair, connect, and try your inserts.
 Demo Mode RPM 2 is supported with a "demonstration (Demo) mode" that easily allows you to navigate the app. Demo mode is intended for navigation purposes only. Data in Demo mode are simply random data
Demo Mode RPM 2 is supported with a "demonstration (Demo) mode" that easily allows you to navigate the app. Demo mode is intended for navigation purposes only. Data in Demo mode are simply random data
Getting Started.
 Getting Started www.scientificbraintrainingpro.com Summary 1. First steps... 2 2. Log in... 2 3. Create an account for a patient... 3 4. Access an exercise with this patient... 4 5. Viewing the results
Getting Started www.scientificbraintrainingpro.com Summary 1. First steps... 2 2. Log in... 2 3. Create an account for a patient... 3 4. Access an exercise with this patient... 4 5. Viewing the results
Exercise Pro Getting Started Guide
 Exercise Pro Getting Started Guide Table Of Contents Installation... 1 Overview... 1 Tutorial... 1 The Exercise Pro 6 Interface... 1 Searching and Selecting Exercises... 2 Printing the Exercise Program...
Exercise Pro Getting Started Guide Table Of Contents Installation... 1 Overview... 1 Tutorial... 1 The Exercise Pro 6 Interface... 1 Searching and Selecting Exercises... 2 Printing the Exercise Program...
Unitron Remote Plus app
 Unitron Remote Plus app User Guide A Sonova brand Getting started Intended use The Unitron Remote Plus app is intended for hearing aids users to adjust certain aspects of Unitron hearing aids through Android
Unitron Remote Plus app User Guide A Sonova brand Getting started Intended use The Unitron Remote Plus app is intended for hearing aids users to adjust certain aspects of Unitron hearing aids through Android
A Quick-Start Guide for rseqdiff
 A Quick-Start Guide for rseqdiff Yang Shi (email: shyboy@umich.edu) and Hui Jiang (email: jianghui@umich.edu) 09/05/2013 Introduction rseqdiff is an R package that can detect differential gene and isoform
A Quick-Start Guide for rseqdiff Yang Shi (email: shyboy@umich.edu) and Hui Jiang (email: jianghui@umich.edu) 09/05/2013 Introduction rseqdiff is an R package that can detect differential gene and isoform
Content Part 2 Users manual... 4
 Content Part 2 Users manual... 4 Introduction. What is Kleos... 4 Case management... 5 Identity management... 9 Document management... 11 Document generation... 15 e-mail management... 15 Installation
Content Part 2 Users manual... 4 Introduction. What is Kleos... 4 Case management... 5 Identity management... 9 Document management... 11 Document generation... 15 e-mail management... 15 Installation
Pyxis MedStation System. Guide for Managing Patient-Specific Medication
 Pyxis MedStation System Guide for Managing Patient-Specific Medication April 2012 CareFusion, Pyxis, Pyxis MedStation, and the CareFusion logo are trademarks or registered trademarks of CareFusion Corporation
Pyxis MedStation System Guide for Managing Patient-Specific Medication April 2012 CareFusion, Pyxis, Pyxis MedStation, and the CareFusion logo are trademarks or registered trademarks of CareFusion Corporation
DICOM Conformance Statement
 DICOM Conformance Statement Document Version 2.0 Revision Summary Date Revision Changes 30 th December 2016 2.0 Update to OrthoView 7.0 Specification 20 October 2005 1.0 Release of OrthoView 3.1 WK3 build
DICOM Conformance Statement Document Version 2.0 Revision Summary Date Revision Changes 30 th December 2016 2.0 Update to OrthoView 7.0 Specification 20 October 2005 1.0 Release of OrthoView 3.1 WK3 build
Hanwell Instruments Ltd. Instruction Manual
 Hanwell Instruments Ltd Instruction Manual Document Title RL5000 Sensors - User Guide Document No. IM4177 Issue No. 3 Hanwell Instruments Ltd 12 Mead Business Centre Mead Lane Hertford SG13 7BJ UNITED
Hanwell Instruments Ltd Instruction Manual Document Title RL5000 Sensors - User Guide Document No. IM4177 Issue No. 3 Hanwell Instruments Ltd 12 Mead Business Centre Mead Lane Hertford SG13 7BJ UNITED
Managing Immunizations
 Managing Immunizations In this chapter: Viewing Immunization Information Entering Immunizations Editing Immunizations Entering a Lead Test Action Editing a Lead Test Action Entering Opt-Out Immunizations
Managing Immunizations In this chapter: Viewing Immunization Information Entering Immunizations Editing Immunizations Entering a Lead Test Action Editing a Lead Test Action Entering Opt-Out Immunizations
Network-assisted data analysis
 Network-assisted data analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Protein identification in shotgun proteomics Protein digestion LC-MS/MS Protein
Network-assisted data analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Protein identification in shotgun proteomics Protein digestion LC-MS/MS Protein
Knowledge networks of biological and medical data An exhaustive and flexible solution to model life sciences domains
 Knowledge networks of biological and medical data An exhaustive and flexible solution to model life sciences domains Dr. Sascha Losko, Dr. Karsten Wenger, Dr. Wenzel Kalus, Dr. Andrea Ramge, Dr. Jens Wiehler,
Knowledge networks of biological and medical data An exhaustive and flexible solution to model life sciences domains Dr. Sascha Losko, Dr. Karsten Wenger, Dr. Wenzel Kalus, Dr. Andrea Ramge, Dr. Jens Wiehler,
Medtech32 Diabetes Get Checked II Advanced Form Release Notes
 Medtech32 Diabetes Get Checked II Advanced Form Release Notes These Release Notes contain important information for all Medtech32 Users. Please ensure that they are circulated amongst all your staff. We
Medtech32 Diabetes Get Checked II Advanced Form Release Notes These Release Notes contain important information for all Medtech32 Users. Please ensure that they are circulated amongst all your staff. We
QuantiPhi for RL78 and MICON Racing RL78
 QuantiPhi for RL78 and MICON Racing RL78 Description: Using cutting-edge model-based design tools, you will design a strategy for a Renesas MICON car, a miniature, autonomous electric vehicle. You will
QuantiPhi for RL78 and MICON Racing RL78 Description: Using cutting-edge model-based design tools, you will design a strategy for a Renesas MICON car, a miniature, autonomous electric vehicle. You will
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types. Epigenome Informatics Workshop Bioinformatics Research Laboratory
 Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
Bowel Cancer Screening for Scotland
 Vision 3 Bowel Cancer Screening for Scotland (BoSS) Copyright INPS Ltd 2014 The Bread Factory, 1A Broughton Street, Battersea, London, SW8 3QJ T: +44 (0) 207 501700 F:+44 (0) 207 5017100 W: www.inps.co.uk
Vision 3 Bowel Cancer Screening for Scotland (BoSS) Copyright INPS Ltd 2014 The Bread Factory, 1A Broughton Street, Battersea, London, SW8 3QJ T: +44 (0) 207 501700 F:+44 (0) 207 5017100 W: www.inps.co.uk
Sleep Apnea Therapy Software Clinician Manual
 Sleep Apnea Therapy Software Clinician Manual Page ii Sleep Apnea Therapy Software Clinician Manual Notices Revised Notice Trademark Copyright Sleep Apnea Therapy Software Clinician Manual 103391 Rev A
Sleep Apnea Therapy Software Clinician Manual Page ii Sleep Apnea Therapy Software Clinician Manual Notices Revised Notice Trademark Copyright Sleep Apnea Therapy Software Clinician Manual 103391 Rev A
LabVIEW Profibus VISA Driver DP-Slave
 DP-Slave Getting Started V1.29 25.09.2007 Project No.: 5303 Doc-ID.: COMSOFT d:\windoc\icp\doku\os\lv-visa\version 1.22\gettingstarted_win_dp-slave_e1.29.doc Revision History Version Date Description V1.11
DP-Slave Getting Started V1.29 25.09.2007 Project No.: 5303 Doc-ID.: COMSOFT d:\windoc\icp\doku\os\lv-visa\version 1.22\gettingstarted_win_dp-slave_e1.29.doc Revision History Version Date Description V1.11
IMPaLA tutorial.
 IMPaLA tutorial http://impala.molgen.mpg.de/ 1. Introduction IMPaLA is a web tool, developed for integrated pathway analysis of metabolomics data alongside gene expression or protein abundance data. It
IMPaLA tutorial http://impala.molgen.mpg.de/ 1. Introduction IMPaLA is a web tool, developed for integrated pathway analysis of metabolomics data alongside gene expression or protein abundance data. It
Cortex Gateway 2.0. Administrator Guide. September Document Version C
 Cortex Gateway 2.0 Administrator Guide September 2015 Document Version C Version C of the Cortex Gateway 2.0 Administrator Guide had been updated with editing changes. Contents Preface... 1 About Cortex
Cortex Gateway 2.0 Administrator Guide September 2015 Document Version C Version C of the Cortex Gateway 2.0 Administrator Guide had been updated with editing changes. Contents Preface... 1 About Cortex
SmartVA-Analyze 2.0 Help
 SmartVA-Analyze 2.0 Help An instruction manual for using SmartVA-Analyze 2.0, which implements the Tariff 2.0 Method for computer certification of verbal autopsy (VA). Contents Contents Overview System
SmartVA-Analyze 2.0 Help An instruction manual for using SmartVA-Analyze 2.0, which implements the Tariff 2.0 Method for computer certification of verbal autopsy (VA). Contents Contents Overview System
Section D. Identification of serotype-specific amino acid positions in DENV NS1. Objective
 Section D. Identification of serotype-specific amino acid positions in DENV NS1 Objective Upon completion of this exercise, you will be able to use the Virus Pathogen Resource (ViPR; http://www.viprbrc.org/)
Section D. Identification of serotype-specific amino acid positions in DENV NS1 Objective Upon completion of this exercise, you will be able to use the Virus Pathogen Resource (ViPR; http://www.viprbrc.org/)
Installing and Testing JMonkeyEngine (jme)
 Installing and Testing JMonkeyEngine (jme) Andrew Davison, ad@fivedots.coe.psu.ac.th July 31st 2014 This document is to help students in 242-515 Animation and Game Development (AGD) install the jmonkeyengine
Installing and Testing JMonkeyEngine (jme) Andrew Davison, ad@fivedots.coe.psu.ac.th July 31st 2014 This document is to help students in 242-515 Animation and Game Development (AGD) install the jmonkeyengine
Principles of phylogenetic analysis
 Principles of phylogenetic analysis Arne Holst-Jensen, NVI, Norway. Fusarium course, Ås, Norway, June 22 nd 2008 Distance based methods Compare C OTUs and characters X A + D = Pairwise: A and B; X characters
Principles of phylogenetic analysis Arne Holst-Jensen, NVI, Norway. Fusarium course, Ås, Norway, June 22 nd 2008 Distance based methods Compare C OTUs and characters X A + D = Pairwise: A and B; X characters
Table of Contents Index Next. See inside for a complete description of program functions >> Link to the Table of Contents >> Link to the Index
 OneTouch Diabetes Management Software User Manual Next User Manual See inside for a complete description of program functions >> Link to the Table of Contents >> Link to the Index Information in this document
OneTouch Diabetes Management Software User Manual Next User Manual See inside for a complete description of program functions >> Link to the Table of Contents >> Link to the Index Information in this document
15.053x. OpenSolver (http://opensolver.org/)
 15.053x OpenSolver (http://opensolver.org/) 1 Table of Contents Introduction to OpenSolver slides 3-4 Example 1: Diet Problem, Set-Up slides 5-11 Example 1: Diet Problem, Dialog Box slides 12-17 Example
15.053x OpenSolver (http://opensolver.org/) 1 Table of Contents Introduction to OpenSolver slides 3-4 Example 1: Diet Problem, Set-Up slides 5-11 Example 1: Diet Problem, Dialog Box slides 12-17 Example
VMA Demo Unit. Introduction. This document provides information on how to set up and operate the VMA Demo Unit. Figure 1: VMA Demo Unit
 Application Specific Controllers Technical Manual 636.3 VMA Controller Section Technical Bulletin Issue Date 0199 VMA Demo Unit Introduction This document provides information on how to set up and operate
Application Specific Controllers Technical Manual 636.3 VMA Controller Section Technical Bulletin Issue Date 0199 VMA Demo Unit Introduction This document provides information on how to set up and operate
JEFIT ios Manual Version 1.0 USER MANUAL. JEFIT Workout App Version 1.0 ios Device
 USER MANUAL JEFIT Workout App Version 1.0 ios Device Jefit, Inc Copyright 2010-2011 All Rights Reserved http://www.jefit.com 1 Table Of Contents 1.) WELCOME - 5-2.) INSTALLATION - 6-2.1 Downloading from
USER MANUAL JEFIT Workout App Version 1.0 ios Device Jefit, Inc Copyright 2010-2011 All Rights Reserved http://www.jefit.com 1 Table Of Contents 1.) WELCOME - 5-2.) INSTALLATION - 6-2.1 Downloading from
Sleep Apnea Therapy Software User Manual
 Sleep Apnea Therapy Software User Manual Page ii Notices Revised Notice Trademark Copyright 103392 Rev B Published February 8, 2013 and supersedes all previous versions. The information contained in this
Sleep Apnea Therapy Software User Manual Page ii Notices Revised Notice Trademark Copyright 103392 Rev B Published February 8, 2013 and supersedes all previous versions. The information contained in this
GridMAT-MD: A Grid-based Membrane Analysis Tool for use with Molecular Dynamics
 GridMAT-MD: A Grid-based Membrane Analysis Tool for use with Molecular Dynamics William J. Allen, Justin A. Lemkul, and David R. Bevan Department of Biochemistry, Virginia Tech User s Guide Version 1.0.2
GridMAT-MD: A Grid-based Membrane Analysis Tool for use with Molecular Dynamics William J. Allen, Justin A. Lemkul, and David R. Bevan Department of Biochemistry, Virginia Tech User s Guide Version 1.0.2
EHS QUICKSTART GUIDE RTLAB / CPU SECTION EFPGASIM TOOLBOX.
 EHS QUICKSTART GUIDE EFPGASIM TOOLBOX RTLAB / CPU SECTION www.opal-rt.com 1751 Richardson, suite 2525 Montréal (Québec) Canada H3K 1G6 www.opal-rt.com 2017 All rights reserved Printed in Canada Contents
EHS QUICKSTART GUIDE EFPGASIM TOOLBOX RTLAB / CPU SECTION www.opal-rt.com 1751 Richardson, suite 2525 Montréal (Québec) Canada H3K 1G6 www.opal-rt.com 2017 All rights reserved Printed in Canada Contents
Anticoagulation Manager - Getting Started
 Vision 3 Anticoagulation Manager - Getting Started Copyright INPS Ltd 2014 The Bread Factory, 1A Broughton Street, Battersea, London, SW8 3QJ T: +44 (0) 207 501700 F:+44 (0) 207 5017100 W: www.inps.co.uk
Vision 3 Anticoagulation Manager - Getting Started Copyright INPS Ltd 2014 The Bread Factory, 1A Broughton Street, Battersea, London, SW8 3QJ T: +44 (0) 207 501700 F:+44 (0) 207 5017100 W: www.inps.co.uk
Software Version 2.0. User s Guide
 Software Version 2.0 User s Guide Table of Contents Contents Contents Important Information About Your FreeStyle Auto-Assist Software...1 Intended Use...1 System Requirements...1 Connecting to your Abbott
Software Version 2.0 User s Guide Table of Contents Contents Contents Important Information About Your FreeStyle Auto-Assist Software...1 Intended Use...1 System Requirements...1 Connecting to your Abbott
AudioConsole. User Guide. Doc. No EN/01 Part No EN
 AudioConsole Doc. No. 7-50-2180-EN/01 Part No. 7-50-21800-EN Copyright notice [2003], 2018 Inmedico A/S. All rights reserved. Oscilla is aregistered trademark of Inmedico A/S in the U.S.A. and/or other
AudioConsole Doc. No. 7-50-2180-EN/01 Part No. 7-50-21800-EN Copyright notice [2003], 2018 Inmedico A/S. All rights reserved. Oscilla is aregistered trademark of Inmedico A/S in the U.S.A. and/or other
Diabetes Management Software V1.3 USER S MANUAL
 Diabetes Management Software V1.3 Manufacturer: BIONIME CORPORATION No. 100, Sec. 2, Daqing St., South Dist., Taichung City 40242, Taiwan http: //www.bionime.com E-mail: info@bionime.com Made in Taiwan
Diabetes Management Software V1.3 Manufacturer: BIONIME CORPORATION No. 100, Sec. 2, Daqing St., South Dist., Taichung City 40242, Taiwan http: //www.bionime.com E-mail: info@bionime.com Made in Taiwan
ATLANTIS WebOrder. ATLANTIS ISUS User guide
 ATLANTIS WebOrder ATLANTIS ISUS User guide Contents ATLANTIS WebOrder Entering an ATLANTIS ISUS order 3 ATLANTIS ISUS implant suprastructures 4 ATLANTIS ISUS Bar 5 ATLANTIS ISUS Bridge 7 ATLANTIS ISUS
ATLANTIS WebOrder ATLANTIS ISUS User guide Contents ATLANTIS WebOrder Entering an ATLANTIS ISUS order 3 ATLANTIS ISUS implant suprastructures 4 ATLANTIS ISUS Bar 5 ATLANTIS ISUS Bridge 7 ATLANTIS ISUS
You can use this app to build a causal Bayesian network and experiment with inferences. We hope you ll find it interesting and helpful.
 icausalbayes USER MANUAL INTRODUCTION You can use this app to build a causal Bayesian network and experiment with inferences. We hope you ll find it interesting and helpful. We expect most of our users
icausalbayes USER MANUAL INTRODUCTION You can use this app to build a causal Bayesian network and experiment with inferences. We hope you ll find it interesting and helpful. We expect most of our users
VMMC Installation Guide (Windows NT) Version 2.0
 VMMC Installation Guide (Windows NT) Version 2.0 The Shrimp Project Department of Computer Science Princeton University February 1999 About this Document Welcome to VMMC! This document describes how to
VMMC Installation Guide (Windows NT) Version 2.0 The Shrimp Project Department of Computer Science Princeton University February 1999 About this Document Welcome to VMMC! This document describes how to
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Preparations. Planmeca Romexis Smile Design Quick guide. Capture 2D photo(s) Start Romexis Smile Design software
 Planmeca Romexis Smile Design Quick guide Preparations Capture D photo(s) A face photo with smile A retractor photo (if gum line is not showing when smiling) Start Romexis Smile Design software From desktop
Planmeca Romexis Smile Design Quick guide Preparations Capture D photo(s) A face photo with smile A retractor photo (if gum line is not showing when smiling) Start Romexis Smile Design software From desktop
PSSV User Manual (V2.1)
 PSSV User Manual (V2.1) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
PSSV User Manual (V2.1) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
Steps to Creating a New Workout Program
 Steps to Creating a New Workout Program Step 1: Log into lab website: https://fitnessandhealthpromotion.ca/ a. If you have never logged in, use your FOL username without the @fanshaweonline.ca portion
Steps to Creating a New Workout Program Step 1: Log into lab website: https://fitnessandhealthpromotion.ca/ a. If you have never logged in, use your FOL username without the @fanshaweonline.ca portion
LabVIEW PROFIBUS VISA Driver DP-Slave
 LabVIEW PROFIBUS VISA Driver DP-Slave Getting Started V1.35 27.04.2017 Project No.: 5303 Doc-ID.: LabVIEW PROFIBUS VISA Driver KUNBUS d:\project\5302_df_profi_ii\anwenderdoku\labview\version 1.35\gettingstarted_win_dp-slave_e.doc
LabVIEW PROFIBUS VISA Driver DP-Slave Getting Started V1.35 27.04.2017 Project No.: 5303 Doc-ID.: LabVIEW PROFIBUS VISA Driver KUNBUS d:\project\5302_df_profi_ii\anwenderdoku\labview\version 1.35\gettingstarted_win_dp-slave_e.doc
Lipid annotation with MS2Analyzer. Yan Ma 10/24/2013
 Lipid annotation with MS2Analyzer Yan Ma 10/24/2013 Checklist before you start You need to have: 1.A computer with Java environment and Office(2003 or higher) 2.MS/MS spectra in MGF files 3.Latest version
Lipid annotation with MS2Analyzer Yan Ma 10/24/2013 Checklist before you start You need to have: 1.A computer with Java environment and Office(2003 or higher) 2.MS/MS spectra in MGF files 3.Latest version
OMIM The Online Mendelian Inheritance in Man Knowledgebase: A Wardrobe Full of Genes. Ada Hamosh, MD, MPH
 OMIM The Online Mendelian Inheritance in Man Knowledgebase: A Wardrobe Full of Genes Ada Hamosh, MD, MPH OMIM THE ONLINE MENDELIAN INHERITANCE IN MAN KNOWLEDGEBASE: A WARDROBE FULL OF GENES The OMIM knowledgebase
OMIM The Online Mendelian Inheritance in Man Knowledgebase: A Wardrobe Full of Genes Ada Hamosh, MD, MPH OMIM THE ONLINE MENDELIAN INHERITANCE IN MAN KNOWLEDGEBASE: A WARDROBE FULL OF GENES The OMIM knowledgebase
CUCM Mixed Mode with Tokenless CTL
 CUCM Mixed Mode with Tokenless CTL Document ID: 118893 Contributed by Milosz Zajac, Michal Myszor, and Leszek Wojnarski, Cisco TAC Engineers. Apr 08, 2015 Contents Introduction Prerequisites Requirements
CUCM Mixed Mode with Tokenless CTL Document ID: 118893 Contributed by Milosz Zajac, Michal Myszor, and Leszek Wojnarski, Cisco TAC Engineers. Apr 08, 2015 Contents Introduction Prerequisites Requirements
SFARI Gene 2.0 User Guide
 1 SFARI Gene 2.0 User Guide This document is designed to acquaint the new user with SFARI Gene release 4.0, an integrated resource for autism research, and to provide enough information to allow the user
1 SFARI Gene 2.0 User Guide This document is designed to acquaint the new user with SFARI Gene release 4.0, an integrated resource for autism research, and to provide enough information to allow the user
