Supplementary Information
|
|
|
- Hilary Watson
- 6 years ago
- Views:
Transcription
1 Supplementary Information Effect of head group and lipid tail oxidation in the cell membrane revealed through integrated simulations and experiments M Yusupov 1, K Wende 2, S Kupsch, E C Neyts 1, S Reuter 2 and A Bogaerts 1 1 Research Group PLASMANT, Department of Chemistry, University of Antwerp, Universiteitsplein 1, B Antwerp, Belgium 2 Leibniz Institute for Plasma Science and Technology, INP Greifswald e.v. Felix-Hausdorff-Str. 2, Greifswald, Germany Leibniz-Center for Medicine and Biosciences, Research Center Borstel, Division of Immunobiophysics, Parkallee 1-40, 2845 Borstel, Germany maksudbek.yusupov@uantwerpen.be Simulation details Reactive (DFTB) MD simulations In our reactive MD simulations we use the so-called DFTB method, which is the extended and improved version of self-consistent charge DFTB 1. It is derived from a third order expansion of the DFT total energy expression, and accurately describes the H binding energies, proton affinities and H transfer barriers without losing its speed and robustness 1. In this work, we use a parameter set (so called OB, emphasizing the use for Organic and Bio-molecules) that was specifically developed for DFTB 2, containing parameters for C/H/O/N/S/P atoms. The model system for the DFTB simulations consists of 8 DOPC (or POPC) molecules with water layers on top and at the bottom (see Fig. 1b in the main paper) that are placed in a box with dimensions of ~16 Å 72 Å 18 Å. The geometry of the structure is optimized using the conjugate gradient algorithm. The system is then equilibrated for 5 ps (i.e., 10 4 iterations) in the isothermal-isobaric (i.e., NPT) ensemble at 00 K, employing the Berendsen thermostat and barostat with a coupling constant of 100 fs and 700 fs, respectively. In all simulations (i.e., during the thermalization, as well as during the particle impact simulations) we use a time step of 0.5 fs. Periodic boundary conditions are applied in all three directions to represent a bulk system. The impinging plasma species first interact with the water layer before reaching the head group of the PLB and therefore to study their behavior in the water layer we perform some test runs by creating a single plasma species in a small box of water (including water molecules with a density of 1 g/cm ) with dimensions of 10 Å 10 Å 10 Å. The water structure (with the single particle in it) is optimized as explained above and then equilibrated in the isothermal (i.e., NVT) ensemble at 00 K, employing the Berendsen thermostat with a coupling constant of 100 fs. The model system consisting of a single PL molecule (which contains 54 atoms) is placed in a box with size of 25 Å 25 Å 25 Å. Note that the volume of the box is much larger than the system itself. This is done to randomly create the plasma particle around the structure. Again this system is optimized using the conjugate gradient algorithm and equilibrated for 10 ps in the NVT ensemble. Subsequently, a single plasma species (e.g., an OH radical) is randomly created around the structure with a minimum distance of 5 Å from the system. We perform 100 simulations for each impinging species, i.e., for OH, HO 2 and H 2O 2, and each simulation lasts for 15 ps (i.e., 10 4 iterations), which is long enough to observe the bond breaking/formation processes. For the analysis of the possible reactions that are obtained from the single PL molecule we use the real water-stabilized PLB structure (i.e., the structure with 8 PLs, presented in Fig. 1b of the main paper). Because of the high computational cost of the DFTB method, we place the species closer (at ~2 Å) to the position where they should react, based on the results obtained for the single PL molecule. Subsequently, we
2 perform again MD simulations, now for 5 ps (i.e., 10 4 iterations), because of the high computational cost of the DFTB method. However, since the plasma species are positioned close enough to the place where they should react, this time is already sufficient for the reaction to occur. Non-reactive MD simulations In our non-reactive MD simulations the model systems with OX2 oxidation products are created from the non-oxidized (i.e., native) PLBs containing 128 PLs, by replacing 12, 2, 64 and 128 DOPC (or POPC) molecules with OX2, corresponding to concentrations of 9.4%, 25%, 50% and 100%, respectively. In the case of ALD oxidation products, we keep 25% of the OX2 products and replace the other intact DOPC molecules with 16, 2, 64 and 96 of ALD products, the concentrations of which correspond to 12.5%, 25%, 50% and 75%, respectively. Thus, in the fully oxidized PLB we will have either 100% of OX2 or 25% of OX2 and 75% of ALD oxidation products. Note that the two bilayer leaflets always contain the same number of oxidized PLs in our model systems. In this way, we obtain 9 different bilayer systems (including the intact PLB) for DOPC and 5 for POPC. All 14 model systems are created using the Packmol package 4. Each system is then optimized using the steepest descent algorithm and equilibrated for 80 ns in the NPT ensemble, at 00 K and 1 atmosphere, employing the Nosé-Hoover thermostat with a coupling constant of 0.2 ps 5 as well as the semi-isotropic Parinello-Rahman barostat with a compressibility and coupling constant of 4.5x10-5 bar -1 and 1 ps, respectively 6. Subsequently, MD simulations are run for 100 ns applying again the NPT ensemble. In all simulations a time step of 2 fs is used. Periodic boundary conditions are applied in all directions. The simulations are carried out using GROMACS 4.6 software, applying the classical unitedatom GROMOS 4A1-S force field 7, which contains parameters for a wide variety of lipids, including DOPC and POPC. Table S1 shows the comparison for the average surface area per lipid and the bilayer thickness of the intact PLB, between our calculated values and the values obtained from experiments and other computational studies. It is clear that our computational results are well within the range of data reported in literature Table S1. Comparison between our calculated data and experimental or other computational data from literature, for pure DOPC and POPC bilayers. phospholipid Properties Calculated value Data from literature DOPC Surface area per lipid (nm 2 ) ± Bilayer thickness (nm).902 ± POPC Surface area per lipid (nm 2 ) 0.69 ± Bilayer thickness (nm).782 ± DFT simulations We also performed DFT calculations to obtain new parameters for the non-reactive force field (for the oxidized/modified bilayers), by carrying out a fitting of some standard potential energy functions to the DFT energy. For this purpose we used the ADF modeling suite package 17,18, employing the BLYP functional 19 with the Slater-type basis set TZ2P 20. The implemented new parameters to the force field are given in Table S2. For the calculation of bond and angle (as well as improper dihedral) force constants, we restrained the bond lengths and angles at seven different values and then fitted a harmonic potential function to the energy profile. For the proper dihedral parameters, dihedral angles were restrained at 6 different values from 0 to 60, and the corresponding dihedral function was fitted to the potential energy (see Table S2). Note that we performed the parametrization (or fitting) procedure only for those bonds or angles or dihedrals which are not included in the force field. Moreover, we used only those Lennard-Jones parameters which are present in the force field.
3 Table S2. New parameters used in the GROMOS 4A1-S force field for describing the newly formed bonds, angles and dihedrals in the oxidized PLBs. The functional forms used for the bonds, angles and dihedrals can be found in the manual of GROMACS Note that C atoms given in the table are united atoms (see Fig. S1). Bonds Bond b 0 (nm) k b (kj.mol -1.nm -4 ) functional type (see 21 ) C 1-N 4 (Fig. S1a) C -N 4 (Fig. S1a) C 2-N 4 (Fig. S1a) Angles Angle θ 0 (deg) k θ (kj.mol -1 ) functional type (see 21 ) C 1-N 4-C 2 (Fig. S1a) C -N 4-C 2 (Fig. S1a) C 1-N 4-C (Fig. S1a) H 1-O 2-C (Fig. S1b) O 1-C 2-C (Fig. S1c) H 1-O 2-C (Fig. S1d) Dihedral angles improper dihedral ξ 0 (deg) k ξ (kj.mol -1.rad -2 ) functional type (see 21 ) N 4-C 1-C -C 2 (Fig. S1a) N 4-C 2-C 1-C (Fig. S1a) N 4-C 2-C -C 1 (Fig. S1a) proper dihedral (multiple) ϕ s (deg) k ϕ (kj.mol -1 ) functional type and multiplicity (see 21 ) H 1-O 2-C -C 4 (Fig. S1b) ; 1 H 1-O 2-C -C 4 (Fig. S1b) ; 2 H 1-O 2-C -C 4 (Fig. S1b) ; P-O 1-C 2-C (Fig. S1c) ; 1 P-O 1-C 2-C (Fig. S1c) ; 2 proper dihedral (Ryckaert-Bellemans) C 0, C 1, C 2, C, C 4, C 5 ( kj.mol -1 ) functional type (see 21 ) O 2-C -C 4-O 5 (Fig. S1b) ; -5.28; ; 8.689; ; C 2-C -C 4-O 5 (Fig. S1c).91271; ; ; ; ; H 1-O 2-C -C 4 (Fig. S1d) ; ; ; ; ; O 2-C -C 4-C 5 (Fig. S1d) ; ;.1699; ; ; Partial charges atom charge atom charge C 1 (Fig. S1a) 0. C 2 (Fig. S1c) 0.2 C 2 (Fig. S1a) 0.4 C (Fig. S1c) 0.0 C (Fig. S1a) 0. C 4 (Fig. S1c) 0. N 4 (Fig. S1a) 0.0 O 5 (Fig. S1c) -0.4 H 1 (Fig. S1b) 0. C 6 (Fig. S1c) 0.5 O 2 (Fig. S1b) -0.6 O 7 (Fig. S1c) -0.5 C (Fig. S1b) 0. H 1 (Fig. S1d) 0. C 4 (Fig. S1b) 0. O 2 (Fig. S1d) -0.6 O 5 (Fig. S1b) -0.6 C (Fig. S1d) 0. P 6 (Fig. S1b) 1.7 C 4 (Fig. S1d) 0.0 O 7 (Fig. S1b) -0.9 C 5 (Fig. S1d) 0.0 O 8 (Fig. S1b) -0.9 O 9 (Fig. S1b) -0.6
4 Figure S1: Schematic picture of the fragments of an oxidized (modified) DOPC (or POPC) PLB for OX1 ((a) and (b)) as well as for OX2 ((c) and (d)), cf. Fig. 5b,c in the main paper. It should be mentioned here that initially we performed the parametrization for both OX1 and OX2 oxidation products, since our reactive (DFTB) MD simulation results showed these two oxidation products to be most abundantly formed. However, the experimental results did not reveal OX1 as a prominent oxidation product, and therefore we did not use the parameters of this oxidation product in our further non-reactive MD simulations. Subsequent reaction of a new OH radical Figure S2 illustrates the reaction energies of the two possible subsequent reactions of the head group oxidized with OX1, with a new OH radical (see also Fig. 2 in the main paper). A new OH radical can either abstract the H atom from the CH 2 group (see initial structure in the red curve of Fig. S2), forming a stable double C=C bond and water molecule (see final structure in the red curve) or it can react with the C radical (see initial structure in the black curve), forming an alcohol group (see final structure in the black curve). As is clear from Fig. S2, the calculated reaction energies employing the above described DFT method revealed that the reaction in which the alcohol group is formed (see final structure in the black curve) is energetically slightly more favorable (cf. ΔE 2 in Fig. S2). Moreover, both reactions do not have a barrier. Note that to calculate the reaction energies of these two systems we performed a constrained optimization at 25 different points. In other words, we restrained the distance between an O atom of the OH radical and an H atom of the CH 2 residue connected to the phosphate group (see initial structure in red curve) and the distance between the O atom of the OH radical and the C radical (see initial configuration in black curve), at 25 different values and calculated the energy profiles. Figure S2: Reaction energies of two systems. The calculated reaction energies show that the reaction presented by the black curve (i.e., the reaction in which a new OH group is created, instead of a double C=C bond and a water molecule) is energetically more favorable.
5 Reactive (DFTB) MD results As mentioned above, the reactions of the OH radicals with the head group of the PL were studied using reactive (DFTB) MD simulations. To get some limited statistics we performed 100 MD simulations. According to the simulation results, we observed six reaction mechanisms which are presented in Fig. S. For clarity, we named these mechanisms as OX1, OX2,, OX6. As is clear from Fig. S, all of these reaction mechanisms are initiated by H-abstraction from (different parts of) the head groups, but few of them give rise to further bond breaking. Note that the OX6 reaction mechanism can take place in both lipid tails (beneath the ester groups, see Fig. S) and the percentage given is the sum of the OX6 occurrences in both tails. A similar summation over the CH groups is also applied for OX1. We did not investigate the further reaction of a new OH radical with the modified PL (by OX1 X6) due to the computational cost of the DFTB method, except for OX1, where we performed further DFT calculations (see above) to find out the energetically most favorable reaction product. It turned out that the formation of an alcohol group was most favorable, therefore we assumed the same for the other modified PLs. This is indeed the most logical for these structures. We studied the longer-term behavior of only OX2 (as well as ALD, see the main text) in our non-reactive MD simulations and we did not study the consequences of the other mechanisms. The reason is that the experimental results suggested only OX2 together with ALD to be the most prominent oxidation products. Figure S: Schematic illustration of six (OX1 to OX6) reaction mechanisms observed in the reactive (DFTB) MD simulations, with their relative occurrences in brackets. The consequences of OX and OX4 are similar to the consequences of OX1 and OX2, respectively, whereas OX5 and OX6 do not lead to the destruction of the head group and are therefore unimportant for the longer term simulations. Non-reactive MD results The surface area per lipid and the bilayer thickness are plotted in Fig. S4 as a function of the concentration of the oxidized PLs, for the OX2 reaction mechanism on the POPC and DPPC PLBs. As is
6 clear, similar trends were obtained for the POPC and DPPC bilayers as for the DOPC PLB (see section Nonreactive MD results in the main paper for a detailed explanation). The only difference is that the relative changes (increase and decrease) in the values are more pronounced in the case of DPPC (see Fig. S4c,d). This is probably due to the difference in the melting temperature of the different PLs. At the simulated temperature (i.e., 00 K), DPPC is in the gel phase (T m ), while POPC and DOPC (T m 294 K and 274 K, respectively 22 ) are already in the liquid state (i.e., with more disordered lipids). Detachment of a lipid tail (see Fig. 5c in the main paper) has a stiffening effect on the PLB, which makes that the OX2 oxidized DPPC bilayers remain in the gel phase. Finally, it is clear from Fig. S4c,d that both the area per lipid and the bilayer thickness level off after 50% oxidation with OX2, which indicates that the lipids are already nearly perfectly structured in the DPPC system. Hence, both the area per lipid as well as the bilayer thickness cannot further decrease/increase. Figure S4: Surface area per lipid and thickness of the bilayer as a function of the concentration of the oxidized PLs, for the OX2 oxidation product, for the POPC ((a) and (b)) and DPPC ((c) and (d)) PLBs.
7 Results of high resolution mass spectrometry Figure S5: Mirror plot of DOPC control (top) and plasma treated DOPC (bottom). Data generated from direct infusion, electrospray ionization / TripleTOF5600 high resolution mass spectrometry. Data normalized to base peak count (DOPC, 20,000/s). Only minor differences were detected between treated and untreated DOPC (see text). Peaks with higher m/z belong to alkali and earth alkali adducts of DOPC. In the low m/z range in source fragments and impurities/traces from buffers occur. Figure S6: Mirror plot of DMPC control (top) and plasma treated DMPC (bottom). Data generated from direct infusion, electrospray ionization / TripleTOF5600 high resolution mass spectrometer. Data normalized to base peak count (DMPC, 40,000/s). Only minor differences were detected between treated and untreated DMPC (see text). Peaks with higher m/z belong to alkali and earth alkali adducts of DMPC. In low m/z range, major peaks belong to doubly charged ions of DMPC + magnesium or calcium.
8 References 1 Gaus, M., Goez, A. & Elstner, M. Parametrization and benchmark of DFTB for organic molecules. Journal of Chemical Theory and Computation 9, 8-54 (2012). 2 Gaus, M., Lu, X., Elstner, M. & Cui, Q. Parameterization of DFTB/OB for Sulfur and Phosphorus for chemical and biological applications. Journal of chemical theory and computation 10, (2014). Berendsen, H. J., Postma, J. P. M., van Gunsteren, W. F., DiNola, A. & Haak, J. Molecular dynamics with coupling to an external bath. The Journal of chemical physics 81, (1984). 4 Martínez, L., Andrade, R., Birgin, E. G. & Martínez, J. M. PACKMOL: a package for building initial configurations for molecular dynamics simulations. Journal of computational chemistry 0, (2009). 5 Hoover, W. G. Canonical dynamics: equilibrium phase-space distributions. Physical Review A 1, 1695 (1985). 6 Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. Journal of Applied physics 52, (1981). 7 Chiu, S.-W., Pandit, S. A., Scott, H. & Jakobsson, E. An improved united atom force field for simulation of mixed lipid bilayers. The Journal of Physical Chemistry B 11, (2009). 8 Kučerka, N., Nieh, M.-P. & Katsaras, J. Fluid phase lipid areas and bilayer thicknesses of commonly used phosphatidylcholines as a function of temperature. Biochimica et Biophysica Acta (BBA)- Biomembranes 1808, (2011). 9 Lewis, B. A. & Engelman, D. M. Lipid bilayer thickness varies linearly with acyl chain length in fluid phosphatidylcholine vesicles. Journal of molecular biology 166, (198). 10 Leekumjorn, S. & Sum, A. K. Molecular characterization of gel and liquid-crystalline structures of fully hydrated POPC and POPE bilayers. The Journal of Physical Chemistry B 111, (2007). 11 Siu, S. W., Vácha, R., Jungwirth, P. & Böckmann, R. A. Biomolecular simulations of membranes: physical properties from different force fields. The Journal of chemical physics 128, (2008). 12 Liu, Y. & Nagle, J. F. Diffuse scattering provides material parameters and electron density profiles of biomembranes. Physical Review E 69, (2004). 1 Kučerka, N. et al. Lipid bilayer structure determined by the simultaneous analysis of neutron and X- ray scattering data. Biophysical journal 95, (2008). 14 Pan, J., Tristram-Nagle, S., Kučerka, N. & Nagle, J. F. Temperature dependence of structure, bending rigidity, and bilayer interactions of dioleoylphosphatidylcholine bilayers. Biophysical journal 94, (2008). 15 Hughes, Z. E., Mark, A. E. & Mancera, R. L. Molecular dynamics simulations of the interactions of DMSO with DPPC and DOPC phospholipid membranes. The journal of physical chemistry. B 116, (2012). 16 Hyvonen, M. T. & Kovanen, P. T. Molecular dynamics simulations of unsaturated lipid bilayers: effects of varying the numbers of double bonds. European biophysics journal : EBJ 4, (2005) Te Velde, G. et al. Chemistry with ADF. Journal of Computational Chemistry 22, (2001). 19 Stephens, P., Devlin, F., Chabalowski, C. & Frisch, M. J. Ab initio calculation of vibrational absorption and circular dichroism spectra using density functional force fields. The Journal of Physical Chemistry 98, (1994). 20 Van Lenthe, E. & Baerends, E. J. Optimized Slater type basis sets for the elements Journal of computational chemistry 24, (200) Silvius, J. R. Thermotropic phase transitions of pure lipids in model membranes and their modifications by membrane proteins. Lipid-protein interactions 2, (1982).
Supporting Information: Revisiting partition in. hydrated bilayer systems
 Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
Supplementary information for Effects of Stretching Speed on. Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular
 Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Supplementary Information
 Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field
 Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Massive oxidation of phospholipid membranes. leads to pore creation and bilayer. disintegration
 Massive oxidation of phospholipid membranes leads to pore creation and bilayer disintegration Lukasz Cwiklik and Pavel Jungwirth Institute of Organic Chemistry and Biochemistry, Academy of Sciences of
Massive oxidation of phospholipid membranes leads to pore creation and bilayer disintegration Lukasz Cwiklik and Pavel Jungwirth Institute of Organic Chemistry and Biochemistry, Academy of Sciences of
Supporting material. Membrane permeation induced by aggregates of human islet amyloid polypeptides
 Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Biological Membranes. Lipid Membranes. Bilayer Permeability. Common Features of Biological Membranes. A highly selective permeability barrier
 Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes
 Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Coarse grained simulations of Lipid Bilayer Membranes
 Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Chemical Surface Transformation 1
 Chemical Surface Transformation 1 Chemical reactions at Si H surfaces (inorganic and organic) can generate very thin films (sub nm thickness up to µm): inorganic layer formation by: thermal conversion:
Chemical Surface Transformation 1 Chemical reactions at Si H surfaces (inorganic and organic) can generate very thin films (sub nm thickness up to µm): inorganic layer formation by: thermal conversion:
Molecular Structure and Permeability at the Interface between Phase-Separated Membrane Domains
 SUPPORTING INFORMATION Molecular Structure and Permeability at the Interface between Phase-Separated Membrane Domains Rodrigo M. Cordeiro Universidade Federal do ABC, Avenida dos Estados 5001, CEP 09210-580,
SUPPORTING INFORMATION Molecular Structure and Permeability at the Interface between Phase-Separated Membrane Domains Rodrigo M. Cordeiro Universidade Federal do ABC, Avenida dos Estados 5001, CEP 09210-580,
The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method
 Journal of Physics: Conference Series PAPER OPEN ACCESS The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method To cite this article: Sri
Journal of Physics: Conference Series PAPER OPEN ACCESS The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method To cite this article: Sri
Supplementary Information: A Critical. Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers
 Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study
 Biophysical Journal Volume 96 February 2009 785 797 785 Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study Charles H. Davis and Max L. Berkowitz * Department
Biophysical Journal Volume 96 February 2009 785 797 785 Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study Charles H. Davis and Max L. Berkowitz * Department
Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation
 Biophysical Journal Volume 77 October 1999 2075 2089 2075 Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation Alexander
Biophysical Journal Volume 77 October 1999 2075 2089 2075 Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation Alexander
Supplement 2 - Quantifying sample mosaicity
 Supplement 2 - Quantifying sample mosaicity Supplementary information for: Order parameters and areas in fluid-phase oriented lipid membranes using wide angle x-ray scattering Thalia T. Mills, Gilman E.
Supplement 2 - Quantifying sample mosaicity Supplementary information for: Order parameters and areas in fluid-phase oriented lipid membranes using wide angle x-ray scattering Thalia T. Mills, Gilman E.
Simulationen von Lipidmembranen
 Simulationen von Lipidmembranen Thomas Stockner thomas.stockner@meduniwien.ac.at Molecular biology Molecular modelling Membranes environment Many cellular functions occur in or around membranes: energy
Simulationen von Lipidmembranen Thomas Stockner thomas.stockner@meduniwien.ac.at Molecular biology Molecular modelling Membranes environment Many cellular functions occur in or around membranes: energy
Phase Transition Behaviours of the Supported DPPC Bilayer. Investigated by Sum Frequency Generation (SFG) and Atomic Force
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 Supporting Information for Phase Transition Behaviours of the Supported DPPC Bilayer
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 Supporting Information for Phase Transition Behaviours of the Supported DPPC Bilayer
MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE
 MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE A Thesis Presented to The Academic Faculty by Young Kyoung Kim In Partial Fulfillment of the
MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE A Thesis Presented to The Academic Faculty by Young Kyoung Kim In Partial Fulfillment of the
8 Influence of permeation modulators on the behaviour of a SC lipid model mixture
 8 Influence of permeation modulators on the behaviour of a SC lipid model mixture 8.1 Introduction In the foregoing parts of this thesis, a model membrane system of SC lipids has been developed and characterized.
8 Influence of permeation modulators on the behaviour of a SC lipid model mixture 8.1 Introduction In the foregoing parts of this thesis, a model membrane system of SC lipids has been developed and characterized.
Phospholipid Component Volumes: Determination and Application to Bilayer Structure Calculations
 734 Biophysical Journal Volume 75 August 1998 734 744 Phospholipid Component Volumes: Determination and Application to Bilayer Structure Calculations Roger S. Armen, Olivia D. Uitto, and Scott E. Feller
734 Biophysical Journal Volume 75 August 1998 734 744 Phospholipid Component Volumes: Determination and Application to Bilayer Structure Calculations Roger S. Armen, Olivia D. Uitto, and Scott E. Feller
Photochemical Applications to the Study of Complexity Phospholipid Bilayer Environments
 Virginia Commonwealth University VCU Scholars Compass Theses and Dissertations Graduate School 2006 Photochemical Applications to the Study of Complexity Phospholipid Bilayer Environments Christopher John
Virginia Commonwealth University VCU Scholars Compass Theses and Dissertations Graduate School 2006 Photochemical Applications to the Study of Complexity Phospholipid Bilayer Environments Christopher John
Biology 5357: Membranes
 s 5357 Biology 5357: s Assembly and Thermodynamics of Soft Matter Paul H. MD, PhD Department of Cell Biology and Physiology pschlesinger@.wustl.edu 362-2223 Characteristics s 5357 s are polymorphic s 5357
s 5357 Biology 5357: s Assembly and Thermodynamics of Soft Matter Paul H. MD, PhD Department of Cell Biology and Physiology pschlesinger@.wustl.edu 362-2223 Characteristics s 5357 s are polymorphic s 5357
Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes.
 Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes. Alexander Kantardjiev 1 and Pavletta Shestakova 1 1 Institute of Organic Chemistry with
Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes. Alexander Kantardjiev 1 and Pavletta Shestakova 1 1 Institute of Organic Chemistry with
Ethanol Induces the Formation of Water-Permeable Defects in Model. Bilayers of Skin Lipids
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2015 Ethanol Induces the Formation of Water-Permeable Defects in Model Bilayers of Skin
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2015 Ethanol Induces the Formation of Water-Permeable Defects in Model Bilayers of Skin
Supporting Information: Atomistic Model for Near-Quantitative. Simulations of Langmuir Monolayers
 Supporting Information: Atomistic Model for Near-Quantitative Simulations of Langmuir Monolayers Matti Javanainen,,, Antti Lamberg, Lukasz Cwiklik,, Ilpo Vattulainen,,, and Samuli Ollila,,# Laboratory
Supporting Information: Atomistic Model for Near-Quantitative Simulations of Langmuir Monolayers Matti Javanainen,,, Antti Lamberg, Lukasz Cwiklik,, Ilpo Vattulainen,,, and Samuli Ollila,,# Laboratory
Supporting Information for Origin of 1/f noise in hydration dynamics on lipid membrane surfaces
 Supporting Informati for Origin of /f noise in hydrati dynamics lipid membrane surfaces Eiji Yamamoto, Takuma kimoto, Masato Yasui, 2 and Kenji Yasuoka Department of Mechanical Engineering, Keio University,
Supporting Informati for Origin of /f noise in hydrati dynamics lipid membrane surfaces Eiji Yamamoto, Takuma kimoto, Masato Yasui, 2 and Kenji Yasuoka Department of Mechanical Engineering, Keio University,
obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and in a
 Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
Karen Doniza 1*, Hui Lin Ong 2, Al Rey Villagracia 1, Aristotle Ubando 5 Melanie David 1, Nelson Arboleda Jr. 1,3, Alvin Culaba 5, Hideaki Kasai 4
 A Molecular Dynamics Study on the Permeability of Oxygen, Carbon Dioxide and Xenon on dioleoyl-phosphatidylcholine (DOPC) and 1- palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine (POPE) Double Lipid
A Molecular Dynamics Study on the Permeability of Oxygen, Carbon Dioxide and Xenon on dioleoyl-phosphatidylcholine (DOPC) and 1- palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine (POPE) Double Lipid
Physical Cell Biology Lecture 10: membranes elasticity and geometry. Hydrophobicity as an entropic effect
 Physical Cell Biology Lecture 10: membranes elasticity and geometry Phillips: Chapter 5, Chapter 11 and Pollard Chapter 13 Hydrophobicity as an entropic effect 1 Self-Assembly of Lipid Structures Lipid
Physical Cell Biology Lecture 10: membranes elasticity and geometry Phillips: Chapter 5, Chapter 11 and Pollard Chapter 13 Hydrophobicity as an entropic effect 1 Self-Assembly of Lipid Structures Lipid
Properties of water hydrating the galactolipid and phospholipid bilayers: a molecular dynamics simulation study*
 Regular paper Vol. 62, No 3/2015 475 481 http://dx.doi.org/10.18388/abp.2015_1077 Properties of water hydrating the galactolipid and phospholipid bilayers: a molecular dynamics simulation study* Michał
Regular paper Vol. 62, No 3/2015 475 481 http://dx.doi.org/10.18388/abp.2015_1077 Properties of water hydrating the galactolipid and phospholipid bilayers: a molecular dynamics simulation study* Michał
Chapter 12: Mass Spectrometry: molecular weight of the sample
 Structure Determination: hapter 12: Mass Spectrometry- molecular weight of the sample; formula hapter 12: Infrared Spectroscopy- indicated which functional groups are present hapter 13: Nuclear Magnetic
Structure Determination: hapter 12: Mass Spectrometry- molecular weight of the sample; formula hapter 12: Infrared Spectroscopy- indicated which functional groups are present hapter 13: Nuclear Magnetic
The Influence of Ceramide Tail Length on the. Structure of Bilayers Composed of Stratum. Corneum Lipids
 The Influence of Ceramide Tail Length on the Structure of Bilayers Composed of Stratum Corneum Lipids T. C. Moore, R. Hartkamp, C. R. Iacovella, A. L. Bunge, and C. M c Cabe Condensed Title: Stratum Corneum
The Influence of Ceramide Tail Length on the Structure of Bilayers Composed of Stratum Corneum Lipids T. C. Moore, R. Hartkamp, C. R. Iacovella, A. L. Bunge, and C. M c Cabe Condensed Title: Stratum Corneum
Structural Characterization on the Gel to Liquid-Crystal Phase Transition of Fully Hydrated DSPC and DSPE Bilayers
 8114 J. Phys. Chem. B 2009, 113, 8114 8123 Structural Characterization on the Gel to Liquid-Crystal Phase Transition of Fully Hydrated DSPC and DSPE Bilayers Shan-Shan Qin and Zhi-Wu Yu* Key Lab of Bioorganic
8114 J. Phys. Chem. B 2009, 113, 8114 8123 Structural Characterization on the Gel to Liquid-Crystal Phase Transition of Fully Hydrated DSPC and DSPE Bilayers Shan-Shan Qin and Zhi-Wu Yu* Key Lab of Bioorganic
and controllable behavior - Supplementary Information
 Metastability in lipid based particles exhibits temporally deterministic and controllable behavior - Supplementary Information Guy Jacoby, Keren Cohen, Kobi Barkan, Yeshayahu Talmon, Dan Peer, Roy Beck
Metastability in lipid based particles exhibits temporally deterministic and controllable behavior - Supplementary Information Guy Jacoby, Keren Cohen, Kobi Barkan, Yeshayahu Talmon, Dan Peer, Roy Beck
Role of Surface Ligands in Nanoparticle Permeation through a Model. Membrane: A Coarse grained Molecular Dynamics Simulations Study
 Role of Surface Ligands in Nanoparticle Permeation through a Model Membrane: A Coarse grained Molecular Dynamics Simulations Study Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad * Department of
Role of Surface Ligands in Nanoparticle Permeation through a Model Membrane: A Coarse grained Molecular Dynamics Simulations Study Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad * Department of
Methods and Materials
 a division of Halcyonics GmbH Anna-Vandenhoeck-Ring 5 37081 Göttingen, Germany Application Note Micostructured lipid bilayers ANDREAS JANSHOFF 1), MAJA GEDIG, AND SIMON FAISS Fig.1: Thickness map of microstructured
a division of Halcyonics GmbH Anna-Vandenhoeck-Ring 5 37081 Göttingen, Germany Application Note Micostructured lipid bilayers ANDREAS JANSHOFF 1), MAJA GEDIG, AND SIMON FAISS Fig.1: Thickness map of microstructured
Data: The GROMOS 43A1-S3 Force Field
 Comparing Simulations of Lipid Bilayers to Scattering Data: The GROMOS 43A1-S3 Force Field Anthony R. Braun, Jonathan N. Sachs and John F. Nagle * Department of Biomedical Engineering, University of Minnesota,
Comparing Simulations of Lipid Bilayers to Scattering Data: The GROMOS 43A1-S3 Force Field Anthony R. Braun, Jonathan N. Sachs and John F. Nagle * Department of Biomedical Engineering, University of Minnesota,
Free Volume Properties of Sphingomyelin, DMPC, DPPC, and PLPC Bilayers
 PUBLISHED IN: JOURNAL OF COMPUTATIONAL AND THEORETICAL NANOSCIENCE, VOL. 2, PP. 40-43 (2005). Free Volume Properties of Sphingomyelin, DMPC, DPPC, and PLPC Bilayers M. Kupiainen, E. Falck, S. Ollila, P.
PUBLISHED IN: JOURNAL OF COMPUTATIONAL AND THEORETICAL NANOSCIENCE, VOL. 2, PP. 40-43 (2005). Free Volume Properties of Sphingomyelin, DMPC, DPPC, and PLPC Bilayers M. Kupiainen, E. Falck, S. Ollila, P.
Molecular Dynamics Simulations of the Anchoring and Tilting of the Lung-Surfactant Peptide SP-B 1-25 in Palmitic Acid Monolayers
 Biophysical Journal Volume 89 December 2005 3807 3821 3807 Molecular Dynamics Simulations of the Anchoring and Tilting of the Lung-Surfactant Peptide SP-B 1-25 in Palmitic Acid Monolayers Hwankyu Lee,*
Biophysical Journal Volume 89 December 2005 3807 3821 3807 Molecular Dynamics Simulations of the Anchoring and Tilting of the Lung-Surfactant Peptide SP-B 1-25 in Palmitic Acid Monolayers Hwankyu Lee,*
PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY
 PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY Physical and Mathematical Sciences 2018, 52(3), p. 217 221 P h y s i c s STUDY OF THE SWELLING OF THE PHOSPHOLIPID BILAYER, DEPENDING ON THE ANGLE BETWEEN THE
PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY Physical and Mathematical Sciences 2018, 52(3), p. 217 221 P h y s i c s STUDY OF THE SWELLING OF THE PHOSPHOLIPID BILAYER, DEPENDING ON THE ANGLE BETWEEN THE
Improved Coarse-Grained Modeling of Cholesterol-Containing Lipid Bilayers
 This is an open access article published under an ACS AuthorChoice License, which permits copying and redistribution of the article or any adaptations for non-commercial purposes. pubs.acs.org/jctc Improved
This is an open access article published under an ACS AuthorChoice License, which permits copying and redistribution of the article or any adaptations for non-commercial purposes. pubs.acs.org/jctc Improved
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Lessons of Slicing Membranes: Interplay of Packing, Free Area, and Lateral Diffusion in Phospholipid/Cholesterol Bilayers
 1076 Biophysical Journal Volume 87 August 2004 1076 1091 Lessons of Slicing Membranes: Interplay of Packing, Free Area, and Lateral Diffusion in Phospholipid/Cholesterol Bilayers Emma Falck,* Michael Patra,
1076 Biophysical Journal Volume 87 August 2004 1076 1091 Lessons of Slicing Membranes: Interplay of Packing, Free Area, and Lateral Diffusion in Phospholipid/Cholesterol Bilayers Emma Falck,* Michael Patra,
Molecular modeling of the pathways of vesicle membrane interaction. Tongtao Yue and Xianren Zhang
 Molecular modeling of the pathways of vesicle membrane interaction Tongtao Yue and Xianren Zhang I. ELECTRONIC SUPPLEMENTARY INFORMATION (ESI): METHODS Dissipative particle dynamics method The dissipative
Molecular modeling of the pathways of vesicle membrane interaction Tongtao Yue and Xianren Zhang I. ELECTRONIC SUPPLEMENTARY INFORMATION (ESI): METHODS Dissipative particle dynamics method The dissipative
Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale
 Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale Rakesh Gupta and Beena Rai* TATA Research Development & Design Centre, TCS Innovation labs, Pune India,
Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale Rakesh Gupta and Beena Rai* TATA Research Development & Design Centre, TCS Innovation labs, Pune India,
Asymmetry of lipid bilayers induced by monovalent salt: Atomistic molecular-dynamics study
 THE JOURNAL OF CHEMICAL PHYSICS 122, 244902 2005 Asymmetry of lipid bilayers induced by monovalent salt: Atomistic molecular-dynamics study Andrey A. Gurtovenko a Laboratory of Physics and Helsinki Institute
THE JOURNAL OF CHEMICAL PHYSICS 122, 244902 2005 Asymmetry of lipid bilayers induced by monovalent salt: Atomistic molecular-dynamics study Andrey A. Gurtovenko a Laboratory of Physics and Helsinki Institute
Diffusion and spectroscopy of water and lipids in fully hydrated dimyristoylphosphatidylcholine bilayer membranes
 Diffusion and spectroscopy of water and lipids in fully hydrated dimyristoylphosphatidylcholine bilayer membranes J. Yang, C. Calero, and J. Martí Citation: The Journal of Chemical Physics 14, 1491 (214);
Diffusion and spectroscopy of water and lipids in fully hydrated dimyristoylphosphatidylcholine bilayer membranes J. Yang, C. Calero, and J. Martí Citation: The Journal of Chemical Physics 14, 1491 (214);
Simulationen von Lipidmembranen
 Simulationen von Lipidmembranen Thomas Stockner Thomas.stockner@meduniwien.ac.at Summary Methods Force Field MD simulations Membrane simulations Application Oxidized lipids Anesthetics Molecular biology
Simulationen von Lipidmembranen Thomas Stockner Thomas.stockner@meduniwien.ac.at Summary Methods Force Field MD simulations Membrane simulations Application Oxidized lipids Anesthetics Molecular biology
7/11/17. Cell Function & Chemistry. Molecular and Cellular Biology. 2. Bio-Chemical Foundations & Key Molecules of a Cell
 Molecular and Cellular Biology Cell Function & Chemistry 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Interaction Molecular Bonds Define Cellular Functions Water H 2 O Interactions
Molecular and Cellular Biology Cell Function & Chemistry 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Interaction Molecular Bonds Define Cellular Functions Water H 2 O Interactions
Interactions of Polyethylenimines with Zwitterionic and. Anionic Lipid Membranes
 Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Supporting Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
Comparing Simulations of Lipid Bilayers to Scattering Data: The GROMOS 43A1-S3 Force Field
 pubs.acs.org/jpcb Comparing Simulations of Lipid Bilayers to Scattering Data: The GROMOS 43A1-S3 Force Field Anthony R. Braun, Jonathan N. Sachs, and John F. Nagle*, Department of Biomedical Engineering,
pubs.acs.org/jpcb Comparing Simulations of Lipid Bilayers to Scattering Data: The GROMOS 43A1-S3 Force Field Anthony R. Braun, Jonathan N. Sachs, and John F. Nagle*, Department of Biomedical Engineering,
Supplementary Material
 10.1071/CH15728_AC The Authors 2016 Australian Journal of Chemistry 2016, 69(8), 846-855 Supplementary Material Structural Diversity and Properties of Six Zn II /Cd II Coordination Polymers Based on a
10.1071/CH15728_AC The Authors 2016 Australian Journal of Chemistry 2016, 69(8), 846-855 Supplementary Material Structural Diversity and Properties of Six Zn II /Cd II Coordination Polymers Based on a
9/11/18. Cell Function & Chemistry. Molecular and Cellular Biology. 2. Bio-Chemical Foundations & Key Molecules of a Cell
 Molecular and Cellular Biology Cell Function & Chemistry 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Interaction Molecular Bonds Define Cellular Functions Water H 2 O Interactions
Molecular and Cellular Biology Cell Function & Chemistry 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Interaction Molecular Bonds Define Cellular Functions Water H 2 O Interactions
Molecular and Cellular Biology. 2. Bio-Chemical Foundations & Key Molecules of a Cell
 Molecular and Cellular Biology 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Cell Function & Chemistry Interaction 1 Molecular Bonds Define Cellular Functions Interactions
Molecular and Cellular Biology 2. Bio-Chemical Foundations & Key Molecules of a Cell Prof. Dr. Klaus Heese Cell Function & Chemistry Interaction 1 Molecular Bonds Define Cellular Functions Interactions
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Phosphatidylcholines are a class of glycerophospholipids which along with other phospholipids
 Phosphatidylcholine Phosphatidylcholines are a class of glycerophospholipids which along with other phospholipids account for more than half of the lipids in most membranes. Phosphatidylcholines can further
Phosphatidylcholine Phosphatidylcholines are a class of glycerophospholipids which along with other phospholipids account for more than half of the lipids in most membranes. Phosphatidylcholines can further
Supplementary Figure 1: Number of hydrophobic lipid tail-water contacts. Contacts were time averaged over 20 ns as a function of the distance of the
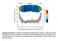 Supplementary Figure 1: Number of hydrophobic lipid tail-water contacts. Contacts were time averaged over 20 ns as a function of the distance of the lipid head group from the bilayer COM. Lipid head groups
Supplementary Figure 1: Number of hydrophobic lipid tail-water contacts. Contacts were time averaged over 20 ns as a function of the distance of the lipid head group from the bilayer COM. Lipid head groups
Neutron reflectivity in biology and medicine. Jayne Lawrence
 Neutron reflectivity in biology and medicine Jayne Lawrence Why neutron reflectivity studies? build up a detailed picture of the structure of a surface in the z direction n e u tro n s in n e u tro n s
Neutron reflectivity in biology and medicine Jayne Lawrence Why neutron reflectivity studies? build up a detailed picture of the structure of a surface in the z direction n e u tro n s in n e u tro n s
Barotropic Phase Transitions of Dilauroylphosphatidylcholine Bilayer Membrane
 High Pressure Bioscience and Biotechnology 68 Proceedings of the 4 th International Conference on High Pressure Bioscience and Biotechnology, Vol. 1, 68 72, 2007 Barotropic Phase Transitions of Dilauroylphosphatidylcholine
High Pressure Bioscience and Biotechnology 68 Proceedings of the 4 th International Conference on High Pressure Bioscience and Biotechnology, Vol. 1, 68 72, 2007 Barotropic Phase Transitions of Dilauroylphosphatidylcholine
Supporting information for: Hydration dynamics of a peripheral membrane. protein
 Supporting information for: Hydration dynamics of a peripheral membrane protein Olivier Fisette,, Christopher Päslack,,, Ryan Barnes, J. Mario Isas, Ralf Langen, Matthias Heyden, Songi Han, and Lars V.
Supporting information for: Hydration dynamics of a peripheral membrane protein Olivier Fisette,, Christopher Päslack,,, Ryan Barnes, J. Mario Isas, Ralf Langen, Matthias Heyden, Songi Han, and Lars V.
Lipids: Membranes Testing Fluid Mosaic Model of Membrane Structure: Cellular Fusion
 Models for Membrane Structure NEW MODEL (1972) Fluid Mosaic Model proposed by Singer & Nicholson Lipids form a viscous, twodimensional solvent into which proteins are inserted and integrated more or less
Models for Membrane Structure NEW MODEL (1972) Fluid Mosaic Model proposed by Singer & Nicholson Lipids form a viscous, twodimensional solvent into which proteins are inserted and integrated more or less
A: All atom molecular simulation systems
 Cholesterol level affects surface charge of lipid membranes in physiological environment Aniket Magarkar a, Vivek Dhawan b, Paraskevi Kallinteri a, Tapani Viitala c, Mohammed Elmowafy c, Tomasz Róg d,
Cholesterol level affects surface charge of lipid membranes in physiological environment Aniket Magarkar a, Vivek Dhawan b, Paraskevi Kallinteri a, Tapani Viitala c, Mohammed Elmowafy c, Tomasz Róg d,
Membrane structure correlates to function of LLP2 on the cytoplasmic tail of HIV-1 gp41 protein
 Membrane structure correlates to function of LLP2 on the cytoplasmic tail of HIV-1 gp41 protein Alexander L. Boscia 1, Kiyotaka Akabori 1, Zachary Benamram 1, Jonathan A. Michel 1, Michael S. Jablin 1,
Membrane structure correlates to function of LLP2 on the cytoplasmic tail of HIV-1 gp41 protein Alexander L. Boscia 1, Kiyotaka Akabori 1, Zachary Benamram 1, Jonathan A. Michel 1, Michael S. Jablin 1,
Molecular Dynamics Simulation of. Amphiphilic Aggregates
 Molecular Dynamics Simulation of Amphiphilic Aggregates By Lanyuan Lu A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements
Molecular Dynamics Simulation of Amphiphilic Aggregates By Lanyuan Lu A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements
1. Describe the difference between covalent and ionic bonds. What are the electrons doing?
 Exam 1 Review Bio 212: 1. Describe the difference between covalent and ionic bonds. What are the electrons doing? 2. Label each picture either a Carbohydrate, Protein, Nucleic Acid, or Fats(Lipid). a.
Exam 1 Review Bio 212: 1. Describe the difference between covalent and ionic bonds. What are the electrons doing? 2. Label each picture either a Carbohydrate, Protein, Nucleic Acid, or Fats(Lipid). a.
Supporting information for: Quantifying. lateral inhomogeneity of. cholesterol-containing membranes
 Supporting information for: Quantifying lateral inhomogeneity of cholesterol-containing membranes Celsa Díaz-Tejada, Igor Ariz-Extreme, Neha Awasthi, and Jochen S. Hub Georg-August-University Göttingen,
Supporting information for: Quantifying lateral inhomogeneity of cholesterol-containing membranes Celsa Díaz-Tejada, Igor Ariz-Extreme, Neha Awasthi, and Jochen S. Hub Georg-August-University Göttingen,
Biochemistry 1 Recitation1 Cell & Water
 Biochemistry 1 Recitation1 Cell & Water There are several important themes that transcends the chemistry and bring the importance of understanding the cell biological differences between eukaryotes and
Biochemistry 1 Recitation1 Cell & Water There are several important themes that transcends the chemistry and bring the importance of understanding the cell biological differences between eukaryotes and
Effect of Sodium and Chloride Binding on a Lecithin Bilayer. A Molecular Dynamics Study
 membranes Article Effect of Sodium and Chloride Binding on a Lecithin Bilayer. A Molecular Dynamics Study Maria M. Reif 1,2, Christopher Kallies 2 and Volker Knecht 2,3, * 1 Physics Department (T38), Technische
membranes Article Effect of Sodium and Chloride Binding on a Lecithin Bilayer. A Molecular Dynamics Study Maria M. Reif 1,2, Christopher Kallies 2 and Volker Knecht 2,3, * 1 Physics Department (T38), Technische
SUPPLEMENTARY INFORMATION
 High-speed atomic force microscopy shows that annexin V stabilizes membranes on the second timescale Atsushi Miyagi, Chris Chipot, Martina Rangl & Simon Scheuring Supplementary Movies: Supplementary Movie
High-speed atomic force microscopy shows that annexin V stabilizes membranes on the second timescale Atsushi Miyagi, Chris Chipot, Martina Rangl & Simon Scheuring Supplementary Movies: Supplementary Movie
Nanoparticle Permeation Induces Water Penetration, Ion Transport, and Lipid Flip-Flop
 pubs.acs.org/langmuir Nanoparticle Permeation Induces Water Penetration, Ion Transport, and Lipid Flip-Flop Bo Song, Huajun Yuan, Sydney V. Pham, Cynthia J. Jameson, and Sohail Murad* Department of Chemical
pubs.acs.org/langmuir Nanoparticle Permeation Induces Water Penetration, Ion Transport, and Lipid Flip-Flop Bo Song, Huajun Yuan, Sydney V. Pham, Cynthia J. Jameson, and Sohail Murad* Department of Chemical
Chapter 1 Membrane Structure and Function
 Chapter 1 Membrane Structure and Function Architecture of Membranes Subcellular fractionation techniques can partially separate and purify several important biological membranes, including the plasma and
Chapter 1 Membrane Structure and Function Architecture of Membranes Subcellular fractionation techniques can partially separate and purify several important biological membranes, including the plasma and
Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes
 Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes A. K. Rønnest, G.H. Peters and F.Y. Hansen Department of Chemistry, Technical University of Denmark,
Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes A. K. Rønnest, G.H. Peters and F.Y. Hansen Department of Chemistry, Technical University of Denmark,
Structural Change in Lipid Bilayers and Water Penetration Induced by Shock Waves: Molecular Dynamics Simulations
 2198 Biophysical Journal Volume 91 September 2006 2198 2205 Structural Change in Lipid Bilayers and Water Penetration Induced by Shock Waves: Molecular Dynamics Simulations Kenichiro Koshiyama,* Tetsuya
2198 Biophysical Journal Volume 91 September 2006 2198 2205 Structural Change in Lipid Bilayers and Water Penetration Induced by Shock Waves: Molecular Dynamics Simulations Kenichiro Koshiyama,* Tetsuya
AFM In Liquid: A High Sensitivity Study On Biological Membranes
 University of Wollongong Research Online Faculty of Science - Papers (Archive) Faculty of Science, Medicine and Health 2006 AFM In Liquid: A High Sensitivity Study On Biological Membranes Michael J. Higgins
University of Wollongong Research Online Faculty of Science - Papers (Archive) Faculty of Science, Medicine and Health 2006 AFM In Liquid: A High Sensitivity Study On Biological Membranes Michael J. Higgins
Conjugated Double Bonds in Lipid Bilayers: A. Molecular Dynamic Simulation Study
 Conjugated Double Bonds in Lipid Bilayers: A Molecular Dynamic Simulation Study Guijun Zhao,P. V. Subbaiah, See-Wing Chiu, Eric Jakobsson, and H. L. Scott Department of Biological, Chemical, and Physical
Conjugated Double Bonds in Lipid Bilayers: A Molecular Dynamic Simulation Study Guijun Zhao,P. V. Subbaiah, See-Wing Chiu, Eric Jakobsson, and H. L. Scott Department of Biological, Chemical, and Physical
Effect of Cholesterol on the Properties of Phospholipid Membranes. 2. Free Energy Profile of Small Molecules
 5322 J. Phys. Chem. B 2003, 107, 5322-5332 Effect of Cholesterol on the Properties of Phospholipid Membranes. 2. Free Energy Profile of Small Molecules Pál Jedlovszky*, Department of Colloid Chemistry,
5322 J. Phys. Chem. B 2003, 107, 5322-5332 Effect of Cholesterol on the Properties of Phospholipid Membranes. 2. Free Energy Profile of Small Molecules Pál Jedlovszky*, Department of Colloid Chemistry,
in-silico Design of Nanoparticles for Transdermal Drug Delivery Application
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 in-silico Design of Nanoparticles for Transdermal Drug Delivery Application Rakesh Gupta and Beena
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 in-silico Design of Nanoparticles for Transdermal Drug Delivery Application Rakesh Gupta and Beena
H 2 O. Liquid, solid, and vapor coexist in the same environment
 Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
Gold nanocrystals at DPPC bilayer. Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad
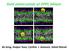 Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Biochimica et Biophysica Acta
 iochimica et iophysica cta 1828 (2013) 1259 1270 Contents lists available at SciVerse ScienceDirect iochimica et iophysica cta journal homepage: www.elsevier.com/locate/bbamem Molecular dynamics study
iochimica et iophysica cta 1828 (2013) 1259 1270 Contents lists available at SciVerse ScienceDirect iochimica et iophysica cta journal homepage: www.elsevier.com/locate/bbamem Molecular dynamics study
Simulation-Based Methods for Interpreting X-Ray Data from Lipid Bilayers
 2796 Biophysical Journal Volume 90 April 2006 2796 2807 Simulation-Based Methods for Interpreting X-Ray Data from Lipid Bilayers Jeffery B. Klauda,* Norbert Kučerka, y Bernard R. Brooks,* Richard W. Pastor,
2796 Biophysical Journal Volume 90 April 2006 2796 2807 Simulation-Based Methods for Interpreting X-Ray Data from Lipid Bilayers Jeffery B. Klauda,* Norbert Kučerka, y Bernard R. Brooks,* Richard W. Pastor,
A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state
![A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state](/thumbs/86/93673402.jpg) A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state Mingguang Pan, Min Xue* Department of Chemistry, Zhejiang University, Hangzhou 310027, P. R. China Fax:
A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state Mingguang Pan, Min Xue* Department of Chemistry, Zhejiang University, Hangzhou 310027, P. R. China Fax:
SAXS on lipid structures
 Practical Course in Biophysics, Experiment R2b SAXS on lipid structures Summer term 2015 Room: Advisor: X-ray lab at LS Rädler, NU111 Stefan Fischer Tel: +49-(0)89-2180-1459 Email: stefan.f.fischer@physik.lmu.de
Practical Course in Biophysics, Experiment R2b SAXS on lipid structures Summer term 2015 Room: Advisor: X-ray lab at LS Rädler, NU111 Stefan Fischer Tel: +49-(0)89-2180-1459 Email: stefan.f.fischer@physik.lmu.de
Chapter 7: Membranes
 Chapter 7: Membranes Roles of Biological Membranes The Lipid Bilayer and the Fluid Mosaic Model Transport and Transfer Across Cell Membranes Specialized contacts (junctions) between cells What are the
Chapter 7: Membranes Roles of Biological Membranes The Lipid Bilayer and the Fluid Mosaic Model Transport and Transfer Across Cell Membranes Specialized contacts (junctions) between cells What are the
Structure and Dynamics of Sphingomyelin Bilayer: Insight Gained through Systematic Comparison to Phosphatidylcholine
 2976 Biophysical Journal Volume 87 November 2004 2976 2989 Structure and Dynamics of Sphingomyelin Bilayer: Insight Gained through Systematic Comparison to Phosphatidylcholine Perttu Niemelä,* Marja T.
2976 Biophysical Journal Volume 87 November 2004 2976 2989 Structure and Dynamics of Sphingomyelin Bilayer: Insight Gained through Systematic Comparison to Phosphatidylcholine Perttu Niemelä,* Marja T.
Water-phospholipid interactions at the interface of lipid. membranes: comparison of different force fields
 Water-phospholipid interactions at the interface of lipid membranes: comparison of different force fields Jianjun Jiang, 1,2 Weixin Li, 1,3 Liang Zhao, 1 Yuanyan Wu, 1 Peng Xiu, 4 Guoning Tang, 3 and Yusong
Water-phospholipid interactions at the interface of lipid membranes: comparison of different force fields Jianjun Jiang, 1,2 Weixin Li, 1,3 Liang Zhao, 1 Yuanyan Wu, 1 Peng Xiu, 4 Guoning Tang, 3 and Yusong
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/12/e1601838/dc1 Supplementary Materials for General and programmable synthesis of hybrid liposome/metal nanoparticles Jin-Ho Lee, Yonghee Shin, Wooju Lee, Keumrai
advances.sciencemag.org/cgi/content/full/2/12/e1601838/dc1 Supplementary Materials for General and programmable synthesis of hybrid liposome/metal nanoparticles Jin-Ho Lee, Yonghee Shin, Wooju Lee, Keumrai
Cholesterol-Sphingomyelin Interactions: A Molecular Dynamics Simulation Study
 3756 Biophysical Journal Volume 91 November 2006 3756 3767 Cholesterol-Sphingomyelin Interactions: A Molecular Dynamics Simulation Study Tomasz Róg* y and Marta Pasenkiewicz-Gierula* * Department of Biophysics,
3756 Biophysical Journal Volume 91 November 2006 3756 3767 Cholesterol-Sphingomyelin Interactions: A Molecular Dynamics Simulation Study Tomasz Róg* y and Marta Pasenkiewicz-Gierula* * Department of Biophysics,
The Interaction between Lipid Bilayers and Biological Membranes. Chapter 18
 The Interaction between Lipid Bilayers and Biological Membranes Chapter 18 Introduction Membrane & Phospholipid Bilayer Structure Membrane Lipid bilayer 2 Introduction Forces Acting between Surfaces in
The Interaction between Lipid Bilayers and Biological Membranes Chapter 18 Introduction Membrane & Phospholipid Bilayer Structure Membrane Lipid bilayer 2 Introduction Forces Acting between Surfaces in
Fatty Acid and Phospholipid Crystalline Adsorbates: Ultrafast T-jump Dynamics
 94 Chapter 6 Fatty Acid and Phospholipid Crystalline Adsorbates: Ultrafast Tjump Dynamics To study the dynamics of the surface adsorbates of crystalline fatty acids and phospholipids, the diffraction patterns
94 Chapter 6 Fatty Acid and Phospholipid Crystalline Adsorbates: Ultrafast Tjump Dynamics To study the dynamics of the surface adsorbates of crystalline fatty acids and phospholipids, the diffraction patterns
Modification of the CHARMM Force Field for DMPC Lipid Bilayer
 Modification of the CHARMM Force Field for DMPC Lipid Bilayer CARL-JOHAN HÖGBERG, 1 ALEXEI M. NIKITIN, 1,2 ALEXANDER P. LYUBARTSEV 1 1 Division of Physical Chemistry, Arrhenius Laboratory, Stockholm University,
Modification of the CHARMM Force Field for DMPC Lipid Bilayer CARL-JOHAN HÖGBERG, 1 ALEXEI M. NIKITIN, 1,2 ALEXANDER P. LYUBARTSEV 1 1 Division of Physical Chemistry, Arrhenius Laboratory, Stockholm University,
Behaviour of small solutes and large drugs in a lipid bilayer from computer simulations
 Biochimica et Biophysica Acta 1718 (2005) 1 21 http://www.elsevier.com/locate/bba Behaviour of small solutes and large drugs in a lipid bilayer from computer simulations D. Bemporad a, C. Luttmann b, J.W.
Biochimica et Biophysica Acta 1718 (2005) 1 21 http://www.elsevier.com/locate/bba Behaviour of small solutes and large drugs in a lipid bilayer from computer simulations D. Bemporad a, C. Luttmann b, J.W.
Lipid Packing Drives the Segregation of Transmembrane Helices into Disordered Lipid Domains in Model Membranes
 Supporting Information Lipid Packing Drives the Segregation of Transmembrane Helices into Disordered Lipid Domains in Model Membranes Lars V. Schäfer, Djurre H. de Jong, Andrea Holt, Andrzej J. Rzepiela,
Supporting Information Lipid Packing Drives the Segregation of Transmembrane Helices into Disordered Lipid Domains in Model Membranes Lars V. Schäfer, Djurre H. de Jong, Andrea Holt, Andrzej J. Rzepiela,
Under the Influence of Alcohol: The Effect of Ethanol and Methanol on Lipid Bilayers
 Biophysical Journal Volume 90 February 2006 1121 1135 1121 Under the Influence of Alcohol: The Effect of Ethanol and Methanol on Lipid Bilayers Michael Patra,* y Emppu Salonen, z Emma Terama, z Ilpo Vattulainen,
Biophysical Journal Volume 90 February 2006 1121 1135 1121 Under the Influence of Alcohol: The Effect of Ethanol and Methanol on Lipid Bilayers Michael Patra,* y Emppu Salonen, z Emma Terama, z Ilpo Vattulainen,
Supporting Information
 Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
level and yield the relevant dynamical and structural information.
 Lecture 1 ATOMIC LEVEL SIMULATIONS OF MEMBRANES A fundamental understanding of the properties of biological membranes and membrane proteins from the atomic point of view is undoubtedly of great biochemical,
Lecture 1 ATOMIC LEVEL SIMULATIONS OF MEMBRANES A fundamental understanding of the properties of biological membranes and membrane proteins from the atomic point of view is undoubtedly of great biochemical,
Chemical Science EDGE ARTICLE. Effect of lipid peroxidation on membrane permeability of cancer and normal cells subjected to oxidative stress
 Chemical Science EDGE ARTICLE View Article Online View Journal View Issue Cite this: Chem. Sci., 2016, 7, 489 Received 25th June 2015 Accepted 14th October 2015 DOI: 10.1039/c5sc02311d www.rsc.org/chemicalscience
Chemical Science EDGE ARTICLE View Article Online View Journal View Issue Cite this: Chem. Sci., 2016, 7, 489 Received 25th June 2015 Accepted 14th October 2015 DOI: 10.1039/c5sc02311d www.rsc.org/chemicalscience
Lipid14: The Amber Lipid Force Field
 pubs.acs.org/jctc Lipid14: The Amber Lipid Force Field Callum J. Dickson,, Benjamin D. Madej,,±, Åge A. Skjevik,, Robin M. Betz, Knut Teigen, Ian R. Gould,*, and Ross C. Walker*,,± Department of Chemistry
pubs.acs.org/jctc Lipid14: The Amber Lipid Force Field Callum J. Dickson,, Benjamin D. Madej,,±, Åge A. Skjevik,, Robin M. Betz, Knut Teigen, Ian R. Gould,*, and Ross C. Walker*,,± Department of Chemistry
