Supporting information for: Hydration dynamics of a peripheral membrane. protein
|
|
|
- Meryl Wilcox
- 5 years ago
- Views:
Transcription
1 Supporting information for: Hydration dynamics of a peripheral membrane protein Olivier Fisette,, Christopher Päslack,,, Ryan Barnes, J. Mario Isas, Ralf Langen, Matthias Heyden, Songi Han, and Lars V. Schäfer, Center for Theoretical Chemistry, Ruhr-University, Bochum, Germany Max-Planck Institut für Kohlenforschung, Mülheim an der Ruhr, Germany Department of Chemistry and Biochemistry and Department of Chemical Engineering, University of California, Santa Barbara, U.S.A. Department of Biochemistry and Molecular Biology, Zilkha Neurogenetic Institute, Keck School of Medicine, University of Southern California, Los Angeles, U.S.A. Contributed equally to this work lars.schaefer@ruhr-uni-bochum.de S1
2 Details of MD simulations Anx coordinates were taken from an X-ray crystal structure (PDB ID 1DM5) S1 of the E105K variant; chain A from the homohexamer was retained, along with its two bound Ca 2+ ions and crystal water molecules. The structure was subjected to 500 steps of steepest-descent (SD) energy minimization. A bilayer with 512 DOPC lipids was generated from a preequilibrated 128 DOPC bilayer by replicating the bilayer four-fold in the plane of the membrane. A DOPC/DOPS (7:3) bilayer was prepared from the DOPC bilayer, replacing the three terminal N-methyl groups (C13 C15) by hydrogens and the H12B atom by a carboxyl group (thus yielding the appropriate stereoisomer at the C12 position). An equal number of DOPC molecules from each leaflet was converted to DOPS; these molecules were chosen randomly and were evenly distributed in the membrane plane. The resulting system was energy-minimized (500 steps SD). A purely hydrophobic bilayer was prepared by setting all partial charges in the lipid molecules to zero. Harmonic position restraints (force constant 1000 kj mol -1 nm -2 ) were applied to the head group phosphorus and the terminal tail carbon atoms. To test the influence of the position restraints, a standard DOPC bilayer (i.e. with normal partial charges) was simulated with the same position restraints applied to the DOPC molecules. To test the influence of protein electrostatics, position-restrained (force constant 1000 kj mol -1 nm -2 on all protein non-hydrogen atoms) Anx with partial charges set to zero was simulated. For the Anx-on-membrane systems, Anx was then positioned atop the bilayer; the approximate initial insertion depth was chosen according to experimental results. S2 After energy minimization (500 steps SD), all systems were solvated in identicallysized boxes ( Å 3 ) that accommodate the protein, bilayer, and at least 20 Å of water above the protein in the dimension normal to the membrane. The minimum distance between Anx and its periodic image never dropped below 50 Å in any of our MD simulations, and hence the simulation box is sufficiently large. Random water molecules were then replaced by ions to neutralize the system. To test the influence of the counterion gradient, 120 mm ions were added to an additional Anx-on-DOPC-bilayer system. Total system sizes S2
3 were about atoms. Simulations were carried out with GROMACS S3,S4 The Amber99SB*-ILDNP protein forcefield, S5 S10 Slipids forcefield S11,S12 and TIP4P/2005 S13 water model were used. For one control system (see Table 1, main text), the TIP3P S14 water model was used instead. The SETTLE S15 and LINCS S16 algorithms were applied to constrain the internal degrees of freedom of water molecules and the bonds in other molecules, respectively, allowing for a 2-fs integration time step. Short-range non-bonded Coulomb and Lennard-Jones 6-12 interactions were treated with a Verlet buffered pair list S17 with potentials smoothly shifted to zero at a 10 Å cut-off. Long-range Coulomb interactions were treated with the PME method S18 with a grid spacing of 1.2 Å and cubic spline interpolation. Analytical dispersion corrections were applied for energy and pressure to compensate for the truncation of the Lennard-Jones interactions. Periodic rectangular cells were used. The thermodynamic ensemble was npt, except for systems including position-restrained lipids, where nvt was used instead, and except for the simulations that were used for the computation of the vibrational density of states, which were simulated in the nve ensemble. In the npt and nvt simulations, temperature was kept constant at 298 K by a velocity-rescaling thermostat S19 with coupling time constant 0.1 ps. For constant 1.0 bar pressure, a Berendsen barostat S20 was used with coupling time constant 0.5 ps and bar 1 compressibility. The barostat was semi-isotropic for all lipid bilayer systems, and isotropic for those without lipids. Analysis of protein position and stability To investigate the binding of Anx to the membrane, we monitored the center-of-mass (COM) motion of the protein along the bilayer normal during the MD. We used the average position of the phosphate head groups of the upper leaflet to determine a reference point from which distances were measured. The distance of Anx to the membrane surface (Fig. S1A) increases until 30 ns and then fluctuates around an equilibrium value. We therefore considered the S3
4 first 30 ns of all simulations as additional equilibration time and excluded them from the analysis, unless otherwise noted. To validate the membrane binding observed in our MD simulations, we compared the equilibrium position of Anx in our simulations to EPR measurements of fractional solvent accessibility (f SA ) S21 of residues 219 to 242. To do so, we counted the average number of water oxygens within 6.0 Å of each Anx residue during the last 70 ns of the trajectory for Anx on DOPC/DOPS bilayer (Fig. S1B). Residues 219, 222, 223, 227, 234, 235, 237, 238 and 239 were characterized as buried by EPR (f SA < 0.05). These residues, except 227 located in a flexible loop, are also the least solvent-accessed ones in our simulations (below the dashed line in Fig. S1B). The general trend observed throughout the entire sequence is very similar between our MD simulations and the EPR experiment, indicating that the Anx position observed in our simulations is consistent with current knowledge. The binding mode of Anx onto phospholipid membranes is nonetheless a topic of active research, S22 and our MD simulations do not take into account possible effects due to oligomerization S23 and induced membrane curvature. S21,S24 These processes, however, are unlikely to substantially change the altered water dynamics around the peripheral membrane protein. Figure S1: Anx position on bilayer. A: Time series of the COM distance of Anx to the phosphate head groups. B: Water accessibility of Anx residues 219 to 242 from MD (water oxygen within 6.0 Å) and EPR (f SA ). S21 After equilibration, the Anx-bound calcium ions establish salt bridges with the negatively- S4
5 charged carboxylate groups of the DOPS lipids. Experimentally, membrane binding of Anx is a calcium-dependent process S25 that also requires negatively charged lipids. In our simulations, Anx remains bound even on a pure DOPC (zwitterionic lipid) bilayer because the timescale of dissociation is much longer than the simulation length. Anx is positioned on the bilayer at the start of the simulation, and thus remains stably bound for over 100 ns. For all systems, the structural integrity of the protein was assessed from C α -RMSD (Fig. S2). These were stable after 20 ns and remained essentially below 2.5 Å, with mean values ranging from 1.2 Å to 2.1 Å Anx on DOPC/DOPS bilayer Anx on DOPC bilayer Anx on hydrophobic bilayer Anx in solution RMSD [Å] Time [ns] Figure S2: C α -RMSD timeseries. Water retardation from mean squared displacements The retardation factors in Fig. 2 (main text) were computed from lateral MSDs of water molecules after 10 ps, which are plotted in Fig. S3. Retardation factors for additional control simulations are shown in Fig. S4. To assess the influence of considering 3D instead of 2D (lateral) translational motions, i.e., of including the component perpendicular to the membrane interface, water retardation factors for the Anx-on-DOPC/DOPS system were also computed from 3D MSDs (Fig. S4, magenta hexagons). The results are virtually identical to those for 2D translation (blue circles), indicating that lateral and perpendicular diffusion S5
6 are effectively slowed-down in similar ways by the membrane and the protein. Next, to investigate the influence of the water model, we repeated the Anx-on-DOPC/DOPS simulation with the TIP3P model (Fig. S4, red triangles; retardation factor computed relative to bulk TIP3P water). The trend is highly similar to that of the corresponding TIP4P/2005 simulations, although in terms of absolute numbers the diffusion of TIP3P is more than a factor of two faster than that of TIP4P/2005. Finally, to investigate whether (and, if so, by how much) the observed water dynamics depends on the position of Anx on the membrane, retardation factors were computed for the Anx on DOPC/DOPS system from trajectories initiated after only 5 ns of equilibration, i.e., before the protein COM position had stabilized (see Fig. S1A). The general trend is preserved (Fig. S4, green triangles), demonstrating that the water retardation is robust with respect to small changes in the actual position of the protein on the membrane. The protein-induced effect is smeared over a larger region and a clear plateau is harder to identify, though, since Anx position on the bilayer is not stable throughout the whole simulation. Water dynamics around individual residues Table S1 shows the water dynamics around eleven spin-labeled residues for Anx in solution measured by ODNP-enhanced NMR relaxometry. The corresponding retardation factors and their MD-derived counterparts are shown in Table S2. See also Fig. 3 in the main text. S6
7 Figure S3: Lateral MSD of water around Anx as a function of distance to the phosphate head groups. Anx position on the membrane is indicated by the grey box. Vertical bars show standard deviations, reflecting the width of the underlying distribution. The statistical errors (not shown) are very small since the values are averaged over i) 14 independent simulations, and ii) several water molecules in each slab. Retardation factor Bulk water Anx on DOPC/DOPS bilayer Anx on DOPC/DOPS bilayer, TIP3P Anx on DOPC/DOPS bilayer, 3D Anx on DOPC/DOPS bilayer, first 5 ns Anx on restrained DOPC bilayer Hydrophobic Anx in solution Anx Distance to membrane [Å] Figure S4: Water retardation around Anx as a function of distance to phosphate head groups in control simulations (see Table 1, main text). S7
8 Table S1: ODNP results. The values are given together with the standard deviations. Residue k σ [M 1 s 1 ] k ρ [M 1 s 1 ] ξ 100 τ [ps] ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± ± Table S2: Membrane distance (d) and local water retardation factors from MD* and ODNP for individual residues. The values are given together with the standard deviations. Residue d Retardation factor Anx on membrane Anx in solution Å MD (5 Å) MD (3 Å) ODNP MD (5 Å) MD (3 Å) ODNP S ± ± ± ± ± ± 1.9 R ± ± ± ± ± ± 1.5 D ± ± ± ± ± ± 3.6 L ± ± ± 3.8 N ± ± ± 0.4 H ± ± ± ± ± ± 4.9 K ± ± ± ± ± ± 3.1 K ± ± ± 0.6 K ± ± ± ± ± ± 2.8 S ± ± ± ± ± ± 2.8 S ± ± ± 0.4 D ± ± ± ± ± ± 2.1 A ± ± ± ± 0.9 L ± ± ± ± ± ± 12.6 * Retardation factors measured for both 3- and 5-Å cut-offs S8
9 References (S1) Cartailler, J. P.; Haigler, H. T.; Luecke, H. Biochemistry 2000, 39, (S2) Cheng, C.-Y.; Varkey, J.; Ambroso, M. R.; Langen, R.; Han, S. Proc. Natl. Acad. Sci. U.S.A. 2013, 110, (S3) Pronk, S.; Páll, S.; Schulz, R.; Larsson, P.; Bjelkmar, P.; Apostolov, R.; Shirts, M. R.; Smith, J. C.; Kasson, P. M.; van der Spoel, D.; Hess, B.; Lindahl, E. Bioinformatics 2013, 29, (S4) Páll, S.; Abraham, M.; Kutzner, C.; Hess, B.; Lindahl, E. In Solving Software Challenges for Exascale; Markidis, S., Laure, E., Eds.; Lecture Notes in Computer Science; Springer International Publishing, 2015; Vol. 8759; pp (S5) Cornell, W. D.; Cieplak, P.; Bayly, C. I.; Gould, I. R.; Merz, K. M.; Ferguson, D. M.; Spellmeyer, D. C.; Fox, T.; Caldwell, J. W.; Kollman, P. A. J. Am. Chem. Soc. 1995, 117, (S6) Wang, J.; Cieplak, P.; Kollman, P. A. J. Comput. Chem. 2000, 21, (S7) Hornak, V.; Abel, R.; Okur, A.; Strockbine, B.; Roitberg, A.; Simmerling, C. Proteins 2006, 65, (S8) Best, R. B.; Hummer, G. J. Phys. Chem. B 2009, 113, (S9) Lindorff-Larsen, K.; Piana, S.; Palmo, K.; Maragakis, P.; Klepeis, J. L.; Dror, R. O.; Shaw, D. E. Proteins 2010, 78, (S10) Aliev, A. E.; Kulke, M.; Khaneja, H. S.; Chudasama, V.; Sheppard, T. D.; Lanigan, R. M. Proteins 2014, 82, (S11) Jämbeck, J. P. M.; Lyubartsev, A. P. J. Phys. Chem. B 2012, 116, S9
10 (S12) Jämbeck, J. P. M.; Lyubartsev, A. P. J. Chem. Theory Comput. 2012, 8, (S13) Abascal, J. L. F.; Vega, C. J. Chem. Phys. 2005, 123, (S14) Jorgensen, W. L.; Chandrasekhar, J.; Madura, J. D.; Impey, R. W.; Klein, M. L. J. Chem. Phys. 1983, 79, (S15) Miyamoto, S.; Kollman, P. A. J. Comput. Chem. 1992, 13, (S16) Hess, B. J. Chem. Theory Comput. 2008, 4, (S17) Páll, S.; Hess, B. Comput. Phys. Commun. 2013, 184, (S18) Essmann, U.; Perera, L.; Berkowitz, M. L.; Darden, T.; Lee, H.; Pedersen, L. G. J. Chem. Phys. 1995, 103, (S19) Bussi, G.; Donadio, D.; Parrinello, M. J. Chem. Phys. 2007, 126, (S20) Berendsen, H. J. C.; Postma, J. P. M.; van Gunsteren, W. F.; DiNola, A.; Haak, J. R. J. Chem. Phys. 1984, 81, (S21) Isas, J. M.; Kim, Y. E.; Jao, C. C.; Hegde, P. B.; Haigler, H. T.; Langen, R. Biochemistry 2005, 44, (S22) Chen, Z.; Mao, Y.; Yang, J.; Zhang, T.; Zhao, L.; Yu, K.; Zheng, M.; Jiang, H.; Yang, H. Proteins 2014, 82, (S23) Langen, R.; Isas, J. M.; Luecke, H.; Haigler, H. T.; Hubbell, W. L. J. Biol. Chem. 1998, 273, (S24) Isas, J. M.; Langen, R.; Hubbell, W. L.; Haigler, H. T. J. Biol. Chem. 2004, 279, (S25) Swairjo, M. A.; Concha, N. O.; Kaetzel, M. A.; Dedman, J. R.; Seaton, B. A. Nat. Struct. Biol. 1995, 2, S10
Supplementary information for Effects of Stretching Speed on. Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular
 Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Supplementary Information
 Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field
 Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Supporting Information: Revisiting partition in. hydrated bilayer systems
 Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
Ethanol Induces the Formation of Water-Permeable Defects in Model. Bilayers of Skin Lipids
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2015 Ethanol Induces the Formation of Water-Permeable Defects in Model Bilayers of Skin
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2015 Ethanol Induces the Formation of Water-Permeable Defects in Model Bilayers of Skin
Supporting Information: Atomistic Model for Near-Quantitative. Simulations of Langmuir Monolayers
 Supporting Information: Atomistic Model for Near-Quantitative Simulations of Langmuir Monolayers Matti Javanainen,,, Antti Lamberg, Lukasz Cwiklik,, Ilpo Vattulainen,,, and Samuli Ollila,,# Laboratory
Supporting Information: Atomistic Model for Near-Quantitative Simulations of Langmuir Monolayers Matti Javanainen,,, Antti Lamberg, Lukasz Cwiklik,, Ilpo Vattulainen,,, and Samuli Ollila,,# Laboratory
Supporting Information for Origin of 1/f noise in hydration dynamics on lipid membrane surfaces
 Supporting Informati for Origin of /f noise in hydrati dynamics lipid membrane surfaces Eiji Yamamoto, Takuma kimoto, Masato Yasui, 2 and Kenji Yasuoka Department of Mechanical Engineering, Keio University,
Supporting Informati for Origin of /f noise in hydrati dynamics lipid membrane surfaces Eiji Yamamoto, Takuma kimoto, Masato Yasui, 2 and Kenji Yasuoka Department of Mechanical Engineering, Keio University,
Coarse grained simulations of Lipid Bilayer Membranes
 Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study
 Biophysical Journal Volume 96 February 2009 785 797 785 Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study Charles H. Davis and Max L. Berkowitz * Department
Biophysical Journal Volume 96 February 2009 785 797 785 Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study Charles H. Davis and Max L. Berkowitz * Department
Supporting information for: Quantifying. lateral inhomogeneity of. cholesterol-containing membranes
 Supporting information for: Quantifying lateral inhomogeneity of cholesterol-containing membranes Celsa Díaz-Tejada, Igor Ariz-Extreme, Neha Awasthi, and Jochen S. Hub Georg-August-University Göttingen,
Supporting information for: Quantifying lateral inhomogeneity of cholesterol-containing membranes Celsa Díaz-Tejada, Igor Ariz-Extreme, Neha Awasthi, and Jochen S. Hub Georg-August-University Göttingen,
Supporting material. Membrane permeation induced by aggregates of human islet amyloid polypeptides
 Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes
 Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Supplementary Information: A Critical. Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers
 Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Conformational Flexibility of the Peptide Hormone Ghrelin in Solution and Lipid Membrane Bound: A Molecular Dynamics Study
 Open Access Article The authors, the publisher, and the right holders grant the right to use, reproduce, and disseminate the work in digital form to all users. Journal of Biomolecular Structure & Dynamics,
Open Access Article The authors, the publisher, and the right holders grant the right to use, reproduce, and disseminate the work in digital form to all users. Journal of Biomolecular Structure & Dynamics,
Interactions of Polyethylenimines with Zwitterionic and. Anionic Lipid Membranes
 Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Properties of water hydrating the galactolipid and phospholipid bilayers: a molecular dynamics simulation study*
 Regular paper Vol. 62, No 3/2015 475 481 http://dx.doi.org/10.18388/abp.2015_1077 Properties of water hydrating the galactolipid and phospholipid bilayers: a molecular dynamics simulation study* Michał
Regular paper Vol. 62, No 3/2015 475 481 http://dx.doi.org/10.18388/abp.2015_1077 Properties of water hydrating the galactolipid and phospholipid bilayers: a molecular dynamics simulation study* Michał
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
SUPPLEMENTARY INFORMATION
 High-speed atomic force microscopy shows that annexin V stabilizes membranes on the second timescale Atsushi Miyagi, Chris Chipot, Martina Rangl & Simon Scheuring Supplementary Movies: Supplementary Movie
High-speed atomic force microscopy shows that annexin V stabilizes membranes on the second timescale Atsushi Miyagi, Chris Chipot, Martina Rangl & Simon Scheuring Supplementary Movies: Supplementary Movie
Molecular dynamics simulations of charged and neutral lipid bilayers: treatment of electrostatic interactions
 Vol. 50 No. 3/2003 789 798 QUARTERLY Molecular dynamics simulations of charged and neutral lipid bilayers: treatment of electrostatic interactions Tomasz Róg, Krzysztof Murzyn and Marta Pasenkiewicz-Gierula
Vol. 50 No. 3/2003 789 798 QUARTERLY Molecular dynamics simulations of charged and neutral lipid bilayers: treatment of electrostatic interactions Tomasz Róg, Krzysztof Murzyn and Marta Pasenkiewicz-Gierula
Supplementary Information
 Supplementary Information Effect of head group and lipid tail oxidation in the cell membrane revealed through integrated simulations and experiments M Yusupov 1, K Wende 2, S Kupsch, E C Neyts 1, S Reuter
Supplementary Information Effect of head group and lipid tail oxidation in the cell membrane revealed through integrated simulations and experiments M Yusupov 1, K Wende 2, S Kupsch, E C Neyts 1, S Reuter
Phase Behavior of a Phospholipid/Fatty Acid/Water Mixture Studied in Atomic Detail
 Published on Web 01/19/2006 Phase Behavior of a Phospholipid/Fatty Acid/Water Mixture Studied in Atomic Detail Volker Knecht,*, Alan E. Mark,, and Siewert-Jan Marrink Contribution from the Max Planck Institute
Published on Web 01/19/2006 Phase Behavior of a Phospholipid/Fatty Acid/Water Mixture Studied in Atomic Detail Volker Knecht,*, Alan E. Mark,, and Siewert-Jan Marrink Contribution from the Max Planck Institute
Application of Molecular Modelling and EPR Spectroscopy to Lipid Membranes a Combined Approach
 Armenian Journal of Physics, 2016, vol. 9, issue 2, pp. 159-166 Application of Molecular Modelling and EPR Spectroscopy to Lipid Membranes a Combined Approach Andrea Catte and Vasily S. Oganesyan * School
Armenian Journal of Physics, 2016, vol. 9, issue 2, pp. 159-166 Application of Molecular Modelling and EPR Spectroscopy to Lipid Membranes a Combined Approach Andrea Catte and Vasily S. Oganesyan * School
Water-phospholipid interactions at the interface of lipid. membranes: comparison of different force fields
 Water-phospholipid interactions at the interface of lipid membranes: comparison of different force fields Jianjun Jiang, 1,2 Weixin Li, 1,3 Liang Zhao, 1 Yuanyan Wu, 1 Peng Xiu, 4 Guoning Tang, 3 and Yusong
Water-phospholipid interactions at the interface of lipid membranes: comparison of different force fields Jianjun Jiang, 1,2 Weixin Li, 1,3 Liang Zhao, 1 Yuanyan Wu, 1 Peng Xiu, 4 Guoning Tang, 3 and Yusong
Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes.
 Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes. Alexander Kantardjiev 1 and Pavletta Shestakova 1 1 Institute of Organic Chemistry with
Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes. Alexander Kantardjiev 1 and Pavletta Shestakova 1 1 Institute of Organic Chemistry with
The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method
 Journal of Physics: Conference Series PAPER OPEN ACCESS The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method To cite this article: Sri
Journal of Physics: Conference Series PAPER OPEN ACCESS The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method To cite this article: Sri
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation
 www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
Massive oxidation of phospholipid membranes. leads to pore creation and bilayer. disintegration
 Massive oxidation of phospholipid membranes leads to pore creation and bilayer disintegration Lukasz Cwiklik and Pavel Jungwirth Institute of Organic Chemistry and Biochemistry, Academy of Sciences of
Massive oxidation of phospholipid membranes leads to pore creation and bilayer disintegration Lukasz Cwiklik and Pavel Jungwirth Institute of Organic Chemistry and Biochemistry, Academy of Sciences of
Molecular Dynamics Simulations of the Anchoring and Tilting of the Lung-Surfactant Peptide SP-B 1-25 in Palmitic Acid Monolayers
 Biophysical Journal Volume 89 December 2005 3807 3821 3807 Molecular Dynamics Simulations of the Anchoring and Tilting of the Lung-Surfactant Peptide SP-B 1-25 in Palmitic Acid Monolayers Hwankyu Lee,*
Biophysical Journal Volume 89 December 2005 3807 3821 3807 Molecular Dynamics Simulations of the Anchoring and Tilting of the Lung-Surfactant Peptide SP-B 1-25 in Palmitic Acid Monolayers Hwankyu Lee,*
Translocation of C 60 and Its Derivatives Across a Lipid Bilayer
 Translocation of C 60 and Its Derivatives Across a Lipid Bilayer Rui Qiao* NANO LETTERS 2007 Vol. 7, No. 3 614-619 Department of Mechanical Engineering, Clemson UniVersity, Clemson, South Carolina 29634
Translocation of C 60 and Its Derivatives Across a Lipid Bilayer Rui Qiao* NANO LETTERS 2007 Vol. 7, No. 3 614-619 Department of Mechanical Engineering, Clemson UniVersity, Clemson, South Carolina 29634
Molecular modeling of the pathways of vesicle membrane interaction. Tongtao Yue and Xianren Zhang
 Molecular modeling of the pathways of vesicle membrane interaction Tongtao Yue and Xianren Zhang I. ELECTRONIC SUPPLEMENTARY INFORMATION (ESI): METHODS Dissipative particle dynamics method The dissipative
Molecular modeling of the pathways of vesicle membrane interaction Tongtao Yue and Xianren Zhang I. ELECTRONIC SUPPLEMENTARY INFORMATION (ESI): METHODS Dissipative particle dynamics method The dissipative
A: All atom molecular simulation systems
 Cholesterol level affects surface charge of lipid membranes in physiological environment Aniket Magarkar a, Vivek Dhawan b, Paraskevi Kallinteri a, Tapani Viitala c, Mohammed Elmowafy c, Tomasz Róg d,
Cholesterol level affects surface charge of lipid membranes in physiological environment Aniket Magarkar a, Vivek Dhawan b, Paraskevi Kallinteri a, Tapani Viitala c, Mohammed Elmowafy c, Tomasz Róg d,
Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes
 Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes A. K. Rønnest, G. H. Peters, F. Y. Hansen, H. Taub, and A. Miskowiec Citation: The Journal of Chemical
Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes A. K. Rønnest, G. H. Peters, F. Y. Hansen, H. Taub, and A. Miskowiec Citation: The Journal of Chemical
obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and in a
 Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
proteins Lipid-associated aggregate formation of superoxide dismutase-1 is initiated by membrane-targeting loops
 proteins STRUCTURE O FUNCTION O BIOINFORMATICS Lipid-associated aggregate formation of superoxide dismutase-1 is initiated by membrane-targeting loops Choon-Peng Chng 1 * and Richard W. Strange 2 1 Biophysical
proteins STRUCTURE O FUNCTION O BIOINFORMATICS Lipid-associated aggregate formation of superoxide dismutase-1 is initiated by membrane-targeting loops Choon-Peng Chng 1 * and Richard W. Strange 2 1 Biophysical
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
The Influence of Ceramide Tail Length on the. Structure of Bilayers Composed of Stratum. Corneum Lipids
 The Influence of Ceramide Tail Length on the Structure of Bilayers Composed of Stratum Corneum Lipids T. C. Moore, R. Hartkamp, C. R. Iacovella, A. L. Bunge, and C. M c Cabe Condensed Title: Stratum Corneum
The Influence of Ceramide Tail Length on the Structure of Bilayers Composed of Stratum Corneum Lipids T. C. Moore, R. Hartkamp, C. R. Iacovella, A. L. Bunge, and C. M c Cabe Condensed Title: Stratum Corneum
Molecular dynamic study of MlaC protein in Gram-negative bacteria: conformational flexibility, solvent effect and proteinphospholipid
 Molecular dynamic study of MlaC protein in Gram-negative bacteria: conformational flexibility, solvent effect and proteinphospholipid binding Yu-ming M. Huang, 1 * Yinglong Miao, 2 Jason Munguia, 3 Leo
Molecular dynamic study of MlaC protein in Gram-negative bacteria: conformational flexibility, solvent effect and proteinphospholipid binding Yu-ming M. Huang, 1 * Yinglong Miao, 2 Jason Munguia, 3 Leo
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Pacific Symposium on Biocomputing 12:40-50(2007)
 PREDICTING STRUCTURE AND DYNAMICS OF LOOSELY- ORDERED PROTEIN COMPLEXES: INFLUENZA HEMAGGLUTININ FUSION PEPTIDE PETER M. KASSON Medical Scientist Training Program Stanford University, Stanford CA 94305
PREDICTING STRUCTURE AND DYNAMICS OF LOOSELY- ORDERED PROTEIN COMPLEXES: INFLUENZA HEMAGGLUTININ FUSION PEPTIDE PETER M. KASSON Medical Scientist Training Program Stanford University, Stanford CA 94305
Molecular Recognition of Agonist and Antagonist for Peroxisome Proliferator-Activated Receptor-α Studied by Molecular Dynamics Simulations
 Int. J. Mol. Sci. 2014, 15, 8743-8752; doi:10.3390/ijms15058743 Article PEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Molecular Recognition of Agonist
Int. J. Mol. Sci. 2014, 15, 8743-8752; doi:10.3390/ijms15058743 Article PEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Molecular Recognition of Agonist
Three-dimensional (3D) structure prediction of the American and African oil-palms β-ketoacyl-[acp] synthase-ii protein by comparative modelling
![Three-dimensional (3D) structure prediction of the American and African oil-palms β-ketoacyl-[acp] synthase-ii protein by comparative modelling Three-dimensional (3D) structure prediction of the American and African oil-palms β-ketoacyl-[acp] synthase-ii protein by comparative modelling](/thumbs/87/96045358.jpg) www.bioinformation.net Hypothesis Volume 10(3) Three-dimensional (3D) structure prediction of the American and African oil-palms β-ketoacyl-[acp] synthase-ii protein by comparative modelling Edina Wang,
www.bioinformation.net Hypothesis Volume 10(3) Three-dimensional (3D) structure prediction of the American and African oil-palms β-ketoacyl-[acp] synthase-ii protein by comparative modelling Edina Wang,
Biological Membranes. Lipid Membranes. Bilayer Permeability. Common Features of Biological Membranes. A highly selective permeability barrier
 Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
g_membed: Efficient Insertion of a Membrane Protein into an Equilibrated Lipid Bilayer with Minimal Perturbation
 g_membed: Efficient Insertion of a Membrane Protein into an Equilibrated Lipid Bilayer with Minimal Perturbation MAARTEN G. WOLF, 1 MARTIN HOEFLING, 2 CAMILO APONTE-SANTAMARÍA, 2 HELMUT GRUBMÜLLER, 2 GERRIT
g_membed: Efficient Insertion of a Membrane Protein into an Equilibrated Lipid Bilayer with Minimal Perturbation MAARTEN G. WOLF, 1 MARTIN HOEFLING, 2 CAMILO APONTE-SANTAMARÍA, 2 HELMUT GRUBMÜLLER, 2 GERRIT
g membed: Efficient insertion of a membrane protein into an equilibrated lipid bilayer with minimal perturbation.
 g membed: Efficient insertion of a membrane protein into an equilibrated lipid bilayer with minimal perturbation. Maarten G. Wolf, Martin Hoefling, Camilo Aponte-Santamaría, Helmut Grubmüller and Gerrit
g membed: Efficient insertion of a membrane protein into an equilibrated lipid bilayer with minimal perturbation. Maarten G. Wolf, Martin Hoefling, Camilo Aponte-Santamaría, Helmut Grubmüller and Gerrit
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
arxiv:cond-mat/ v1 [cond-mat.soft] 14 Aug 2002
![arxiv:cond-mat/ v1 [cond-mat.soft] 14 Aug 2002 arxiv:cond-mat/ v1 [cond-mat.soft] 14 Aug 2002](/thumbs/94/118067187.jpg) Hydrogen Bond Dynamics Near A Micellar Surface: Origin of the Universal Slow Relaxation at Complex Aqueous Interfaces arxiv:cond-mat/2827v [cond-mat.soft] 4 Aug 22 Sundaram Balasubramanian, Subrata Pal
Hydrogen Bond Dynamics Near A Micellar Surface: Origin of the Universal Slow Relaxation at Complex Aqueous Interfaces arxiv:cond-mat/2827v [cond-mat.soft] 4 Aug 22 Sundaram Balasubramanian, Subrata Pal
Energetics of dendrimer binding to HIV-1 gp120- CD4 complex and mechanismic aspects of its role as an entry-inhibitor
 Journal of Physics: Conference Series PAPER OPEN ACCESS Energetics of dendrimer binding to HIV-1 gp120- CD4 complex and mechanismic aspects of its role as an entry-inhibitor Related content - Charged nanoparticle
Journal of Physics: Conference Series PAPER OPEN ACCESS Energetics of dendrimer binding to HIV-1 gp120- CD4 complex and mechanismic aspects of its role as an entry-inhibitor Related content - Charged nanoparticle
MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE
 MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE A Thesis Presented to The Academic Faculty by Young Kyoung Kim In Partial Fulfillment of the
MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE A Thesis Presented to The Academic Faculty by Young Kyoung Kim In Partial Fulfillment of the
Structural Characterization on the Gel to Liquid-Crystal Phase Transition of Fully Hydrated DSPC and DSPE Bilayers
 8114 J. Phys. Chem. B 2009, 113, 8114 8123 Structural Characterization on the Gel to Liquid-Crystal Phase Transition of Fully Hydrated DSPC and DSPE Bilayers Shan-Shan Qin and Zhi-Wu Yu* Key Lab of Bioorganic
8114 J. Phys. Chem. B 2009, 113, 8114 8123 Structural Characterization on the Gel to Liquid-Crystal Phase Transition of Fully Hydrated DSPC and DSPE Bilayers Shan-Shan Qin and Zhi-Wu Yu* Key Lab of Bioorganic
Effect of Cholesterol on the Properties of Phospholipid Membranes. 2. Free Energy Profile of Small Molecules
 5322 J. Phys. Chem. B 2003, 107, 5322-5332 Effect of Cholesterol on the Properties of Phospholipid Membranes. 2. Free Energy Profile of Small Molecules Pál Jedlovszky*, Department of Colloid Chemistry,
5322 J. Phys. Chem. B 2003, 107, 5322-5332 Effect of Cholesterol on the Properties of Phospholipid Membranes. 2. Free Energy Profile of Small Molecules Pál Jedlovszky*, Department of Colloid Chemistry,
Supporting Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
John M. Leveritt III, Almudena Pino-Angeles, Themis Lazaridis*.
 The structure of a melittin-stabilized pore John M. Leveritt III, Almudena Pino-Angeles, Themis Lazaridis*. Department of Chemistry, The City College of New York, 160 Convent Ave, New York, NY, 10031,
The structure of a melittin-stabilized pore John M. Leveritt III, Almudena Pino-Angeles, Themis Lazaridis*. Department of Chemistry, The City College of New York, 160 Convent Ave, New York, NY, 10031,
Modeling the Interaction of Dodecylphosphocholine Micelles with the Anticoccidial Peptide PW2 Guided by NMR Data
 Molecules 2013, 18, 10056-10080; doi:10.3390/molecules180810056 OPEN ACCESS molecules ISSN 1420-3049 www.mdpi.com/journal/molecules Article Modeling the Interaction of Dodecylphosphocholine Micelles with
Molecules 2013, 18, 10056-10080; doi:10.3390/molecules180810056 OPEN ACCESS molecules ISSN 1420-3049 www.mdpi.com/journal/molecules Article Modeling the Interaction of Dodecylphosphocholine Micelles with
Cellular Neurophysiology I Membranes and Ion Channels
 Cellular Neurophysiology I Membranes and Ion Channels Reading: BCP Chapter 3 www.bioelectriclab All living cells maintain an electrical potential (voltage) across their membranes (V m ). Resting Potential
Cellular Neurophysiology I Membranes and Ion Channels Reading: BCP Chapter 3 www.bioelectriclab All living cells maintain an electrical potential (voltage) across their membranes (V m ). Resting Potential
Model of an Asymmetric DPPC/DPPS Membrane: Effect of Asymmetry on the Lipid Properties. A Molecular Dynamics Simulation Study
 2358 J. Phys. Chem. B 2006, 110, 2358-2363 Model of an Asymmetric DPPC/DPPS Membrane: Effect of Asymmetry on the Lipid Properties. A Molecular Dynamics Simulation Study J. J. López Cascales,*, T. F. Otero,
2358 J. Phys. Chem. B 2006, 110, 2358-2363 Model of an Asymmetric DPPC/DPPS Membrane: Effect of Asymmetry on the Lipid Properties. A Molecular Dynamics Simulation Study J. J. López Cascales,*, T. F. Otero,
Lipid Packing Drives the Segregation of Transmembrane Helices into Disordered Lipid Domains in Model Membranes
 Supporting Information Lipid Packing Drives the Segregation of Transmembrane Helices into Disordered Lipid Domains in Model Membranes Lars V. Schäfer, Djurre H. de Jong, Andrea Holt, Andrzej J. Rzepiela,
Supporting Information Lipid Packing Drives the Segregation of Transmembrane Helices into Disordered Lipid Domains in Model Membranes Lars V. Schäfer, Djurre H. de Jong, Andrea Holt, Andrzej J. Rzepiela,
Physicochemical Properties of Nanoparticles Regulate Translocation across Pulmonary. Surfactant Monolayer and Formation of Lipoprotein Corona
 Physicochemical Properties of Nanoparticles Regulate Translocation across Pulmonary Surfactant Monolayer and Formation of Lipoprotein Corona Supplementary Information Guoqing Hu 1 *, Bao Jiao 1, Xinghua
Physicochemical Properties of Nanoparticles Regulate Translocation across Pulmonary Surfactant Monolayer and Formation of Lipoprotein Corona Supplementary Information Guoqing Hu 1 *, Bao Jiao 1, Xinghua
Simulation of Self-Assembly of Ampiphiles Using Molecular Dynamics
 Simulation of Self-Assembly of Ampiphiles Using Molecular Dynamics Haneesh Kesari, Reza Banki, and Misty Davies Project Final Paper ME346 Stanford University December 15, 2005 1 Introduction Understanding
Simulation of Self-Assembly of Ampiphiles Using Molecular Dynamics Haneesh Kesari, Reza Banki, and Misty Davies Project Final Paper ME346 Stanford University December 15, 2005 1 Introduction Understanding
level and yield the relevant dynamical and structural information.
 Lecture 1 ATOMIC LEVEL SIMULATIONS OF MEMBRANES A fundamental understanding of the properties of biological membranes and membrane proteins from the atomic point of view is undoubtedly of great biochemical,
Lecture 1 ATOMIC LEVEL SIMULATIONS OF MEMBRANES A fundamental understanding of the properties of biological membranes and membrane proteins from the atomic point of view is undoubtedly of great biochemical,
CHAPTER 4. Tryptophan fluorescence quenching by brominated lipids
 CHAPTER 4 Tryptophan fluorescence quenching by brominated lipids 102 4.1 INTRODUCTION The structure and dynamics of biological macromolecules have been widely studied with fluorescence quenching. The accessibility
CHAPTER 4 Tryptophan fluorescence quenching by brominated lipids 102 4.1 INTRODUCTION The structure and dynamics of biological macromolecules have been widely studied with fluorescence quenching. The accessibility
Phospholipid Component Volumes: Determination and Application to Bilayer Structure Calculations
 734 Biophysical Journal Volume 75 August 1998 734 744 Phospholipid Component Volumes: Determination and Application to Bilayer Structure Calculations Roger S. Armen, Olivia D. Uitto, and Scott E. Feller
734 Biophysical Journal Volume 75 August 1998 734 744 Phospholipid Component Volumes: Determination and Application to Bilayer Structure Calculations Roger S. Armen, Olivia D. Uitto, and Scott E. Feller
Karen Doniza 1*, Hui Lin Ong 2, Al Rey Villagracia 1, Aristotle Ubando 5 Melanie David 1, Nelson Arboleda Jr. 1,3, Alvin Culaba 5, Hideaki Kasai 4
 A Molecular Dynamics Study on the Permeability of Oxygen, Carbon Dioxide and Xenon on dioleoyl-phosphatidylcholine (DOPC) and 1- palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine (POPE) Double Lipid
A Molecular Dynamics Study on the Permeability of Oxygen, Carbon Dioxide and Xenon on dioleoyl-phosphatidylcholine (DOPC) and 1- palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine (POPE) Double Lipid
Globular proteins Proteins globular fibrous
 Globular proteins Globular proteins Proteins are biochemical compounds consisting of one or more polypeptides typically folded into a globular or fibrous form in a biologically functional way. Globular
Globular proteins Globular proteins Proteins are biochemical compounds consisting of one or more polypeptides typically folded into a globular or fibrous form in a biologically functional way. Globular
Asymmetry of lipid bilayers induced by monovalent salt: Atomistic molecular-dynamics study
 THE JOURNAL OF CHEMICAL PHYSICS 122, 244902 2005 Asymmetry of lipid bilayers induced by monovalent salt: Atomistic molecular-dynamics study Andrey A. Gurtovenko a Laboratory of Physics and Helsinki Institute
THE JOURNAL OF CHEMICAL PHYSICS 122, 244902 2005 Asymmetry of lipid bilayers induced by monovalent salt: Atomistic molecular-dynamics study Andrey A. Gurtovenko a Laboratory of Physics and Helsinki Institute
Simulationen von Lipidmembranen
 Simulationen von Lipidmembranen Thomas Stockner Thomas.stockner@meduniwien.ac.at Summary Methods Force Field MD simulations Membrane simulations Application Oxidized lipids Anesthetics Molecular biology
Simulationen von Lipidmembranen Thomas Stockner Thomas.stockner@meduniwien.ac.at Summary Methods Force Field MD simulations Membrane simulations Application Oxidized lipids Anesthetics Molecular biology
Supplementary Figure 1. Crystallography of PI4KIIα with ADP bound. a, A
 Supplementary Figure 1. Crystallography of PI4KIIα with ADP bound. a, A stereo image of overall 2Fo-Fc electron density map of PI4KIIα contoured at 1.0 σ. b, 2FoFc electron density map (1.0 σ) around the
Supplementary Figure 1. Crystallography of PI4KIIα with ADP bound. a, A stereo image of overall 2Fo-Fc electron density map of PI4KIIα contoured at 1.0 σ. b, 2FoFc electron density map (1.0 σ) around the
Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation
 Biophysical Journal Volume 77 October 1999 2075 2089 2075 Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation Alexander
Biophysical Journal Volume 77 October 1999 2075 2089 2075 Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation Alexander
Alchembed: A Computational Method for Incorporating Multiple Proteins into Complex Lipid Geometries
 This is an open access article published under a Creative Commons Attribution (CC-BY) License, which permits unrestricted use, distribution and reproduction in any medium, provided the author and source
This is an open access article published under a Creative Commons Attribution (CC-BY) License, which permits unrestricted use, distribution and reproduction in any medium, provided the author and source
Water between membranes: Structure and Dynamics. Abstract
 Water between membranes: Structure and Dynamics. Sotiris Samatas, Carles Calero, Fausto Martelli, and Giancarlo Franzese Abstract An accurate description of the structure and dynamics of interfacial water
Water between membranes: Structure and Dynamics. Sotiris Samatas, Carles Calero, Fausto Martelli, and Giancarlo Franzese Abstract An accurate description of the structure and dynamics of interfacial water
Diffusion and spectroscopy of water and lipids in fully hydrated dimyristoylphosphatidylcholine bilayer membranes
 Diffusion and spectroscopy of water and lipids in fully hydrated dimyristoylphosphatidylcholine bilayer membranes J. Yang, C. Calero, and J. Martí Citation: The Journal of Chemical Physics 14, 1491 (214);
Diffusion and spectroscopy of water and lipids in fully hydrated dimyristoylphosphatidylcholine bilayer membranes J. Yang, C. Calero, and J. Martí Citation: The Journal of Chemical Physics 14, 1491 (214);
Calcium Binding and Head Group Dipole Angle in Phosphatidylserine#Phosphatidylcholine Bilayers
 Article Subscriber access provided by MPI FUR KOLLOID UND GRENZFLACH Calcium Binding and Head Group Dipole Angle in Phosphatidylserine#Phosphatidylcholine Bilayers P. Thomas Vernier, Matthew J. Ziegler,
Article Subscriber access provided by MPI FUR KOLLOID UND GRENZFLACH Calcium Binding and Head Group Dipole Angle in Phosphatidylserine#Phosphatidylcholine Bilayers P. Thomas Vernier, Matthew J. Ziegler,
Cell Structure and Function Exam Study Guide Part I
 Cell Structure and Function Exam Study Guide Part I 1. Which image best depicts the hot water, which the cold? 2. What is the relationship between temperature and the speed of molecular motion? 3. If a
Cell Structure and Function Exam Study Guide Part I 1. Which image best depicts the hot water, which the cold? 2. What is the relationship between temperature and the speed of molecular motion? 3. If a
Four-Scale Description of Membrane Sculpting by BAR Domains
 2806 Biophysical Journal Volume 95 September 2008 2806 2821 Four-Scale Description of Membrane Sculpting by BAR Domains Anton Arkhipov, Ying Yin, and Klaus Schulten Department of Physics and Beckman Institute,
2806 Biophysical Journal Volume 95 September 2008 2806 2821 Four-Scale Description of Membrane Sculpting by BAR Domains Anton Arkhipov, Ying Yin, and Klaus Schulten Department of Physics and Beckman Institute,
Molecular Dynamics Simulation of a Dipalmitoylphosphatidylcholine Bilayer with NaCl
 Biophysical Journal Volume 84 June 2003 3743 3750 3743 Molecular Dynamics Simulation of a Dipalmitoylphosphatidylcholine Bilayer with NaCl Sagar A. Pandit,* David Bostick, y and Max L. Berkowitz* *Department
Biophysical Journal Volume 84 June 2003 3743 3750 3743 Molecular Dynamics Simulation of a Dipalmitoylphosphatidylcholine Bilayer with NaCl Sagar A. Pandit,* David Bostick, y and Max L. Berkowitz* *Department
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale
 Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale Rakesh Gupta and Beena Rai* TATA Research Development & Design Centre, TCS Innovation labs, Pune India,
Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale Rakesh Gupta and Beena Rai* TATA Research Development & Design Centre, TCS Innovation labs, Pune India,
Role of Surface Ligands in Nanoparticle Permeation through a Model. Membrane: A Coarse grained Molecular Dynamics Simulations Study
 Role of Surface Ligands in Nanoparticle Permeation through a Model Membrane: A Coarse grained Molecular Dynamics Simulations Study Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad * Department of
Role of Surface Ligands in Nanoparticle Permeation through a Model Membrane: A Coarse grained Molecular Dynamics Simulations Study Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad * Department of
Highly efficient gene silencing activity of sirna embedded in a nanostructured gyroid cubic lipid matrix
 Highly efficient gene silencing activity of sirna embedded in a nanostructured gyroid cubic lipid matrix Cecília Leal *, Nathan F. Bouxsein, Kai K. Ewert and Cyrus R. Safinya * Physics, Materials, and
Highly efficient gene silencing activity of sirna embedded in a nanostructured gyroid cubic lipid matrix Cecília Leal *, Nathan F. Bouxsein, Kai K. Ewert and Cyrus R. Safinya * Physics, Materials, and
NIH Public Access Author Manuscript J Am Chem Soc. Author manuscript; available in PMC 2008 September 29.
 NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes
 Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes A. K. Rønnest, G.H. Peters and F.Y. Hansen Department of Chemistry, Technical University of Denmark,
Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes A. K. Rønnest, G.H. Peters and F.Y. Hansen Department of Chemistry, Technical University of Denmark,
Andrey A. Gurtovenko*, and Ilpo Vattulainen,,
 J. Phys. Chem. B 2008, 112, 1953-1962 1953 Effect of NaCl and KCl on Phosphatidylcholine and Phosphatidylethanolamine Lipid Membranes: Insight from Atomic-Scale Simulations for Understanding Salt-Induced
J. Phys. Chem. B 2008, 112, 1953-1962 1953 Effect of NaCl and KCl on Phosphatidylcholine and Phosphatidylethanolamine Lipid Membranes: Insight from Atomic-Scale Simulations for Understanding Salt-Induced
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Minicircle topoisomer generation. a, Generation of supercoiled topoisomers. Minicircles were nicked with the sequence-specific nicking endonuclease, Nb.BbvCI.
Supplementary Figures Supplementary Figure 1 Minicircle topoisomer generation. a, Generation of supercoiled topoisomers. Minicircles were nicked with the sequence-specific nicking endonuclease, Nb.BbvCI.
Changes in a Phospholipid Bilayer Induced by the Hydrolysis of a Phospholipase A 2 Enzyme: A Molecular Dynamics Simulation Study
 Biophysical Journal Volume 80 February 2001 565 578 565 Changes in a Phospholipid Bilayer Induced by the Hydrolysis of a Phospholipase A 2 Enzyme: A Molecular Dynamics Simulation Study Marja T. Hyvönen,*
Biophysical Journal Volume 80 February 2001 565 578 565 Changes in a Phospholipid Bilayer Induced by the Hydrolysis of a Phospholipase A 2 Enzyme: A Molecular Dynamics Simulation Study Marja T. Hyvönen,*
AFM In Liquid: A High Sensitivity Study On Biological Membranes
 University of Wollongong Research Online Faculty of Science - Papers (Archive) Faculty of Science, Medicine and Health 2006 AFM In Liquid: A High Sensitivity Study On Biological Membranes Michael J. Higgins
University of Wollongong Research Online Faculty of Science - Papers (Archive) Faculty of Science, Medicine and Health 2006 AFM In Liquid: A High Sensitivity Study On Biological Membranes Michael J. Higgins
Supplementary Figure 1: Number of hydrophobic lipid tail-water contacts. Contacts were time averaged over 20 ns as a function of the distance of the
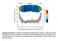 Supplementary Figure 1: Number of hydrophobic lipid tail-water contacts. Contacts were time averaged over 20 ns as a function of the distance of the lipid head group from the bilayer COM. Lipid head groups
Supplementary Figure 1: Number of hydrophobic lipid tail-water contacts. Contacts were time averaged over 20 ns as a function of the distance of the lipid head group from the bilayer COM. Lipid head groups
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Physiologically-relevant levels of sphingomyelin, but not GM1, induces a beta-sheet-rich structure in the amyloid-beta(1-42) monomer
 https://helda.helsinki.fi Physiologically-relevant levels of sphingomyelin, but not GM1, induces a beta-sheet-rich structure in the amyloid-beta(1-42) monomer Owen, Michael C. 2018-09 Owen, M C, Kulig,
https://helda.helsinki.fi Physiologically-relevant levels of sphingomyelin, but not GM1, induces a beta-sheet-rich structure in the amyloid-beta(1-42) monomer Owen, Michael C. 2018-09 Owen, M C, Kulig,
Chapter VI. Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase
 Chapter VI Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase Chapter VI Abstract We report the study of T12A Cyt c CcP complex using isothermal titration calorimetry (ITC),
Chapter VI Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase Chapter VI Abstract We report the study of T12A Cyt c CcP complex using isothermal titration calorimetry (ITC),
BIOCHEMISTRY 460 FIRST HOUR EXAMINATION FORM A (yellow) ANSWER KEY February 11, 2008
 WRITE YOUR AND I.D. NUMBER LEGIBLY ON EVERY PAGE PAGES WILL BE SEPARATED FOR GRADING! CHECK TO BE SURE YOU HAVE 6 PAGES, (print): ANSWERS INCLUDING COVER PAGE. I swear/affirm that I have neither given
WRITE YOUR AND I.D. NUMBER LEGIBLY ON EVERY PAGE PAGES WILL BE SEPARATED FOR GRADING! CHECK TO BE SURE YOU HAVE 6 PAGES, (print): ANSWERS INCLUDING COVER PAGE. I swear/affirm that I have neither given
Experimental. Crystal data. C 19 H 19 N 3 O 3 M r = Monoclinic, C2=c a = (3) Å b = (2) Å c = (3) Å = 110.
 organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 A second monoclinic polymorph of 4-(2- hydroxy-4-methoxybenzylideneamino)- 1,5-dimethyl-2-phenyl-1H-pyrazol- 3(2H)-one
organic compounds Acta Crystallographica Section E Structure Reports Online ISSN 1600-5368 A second monoclinic polymorph of 4-(2- hydroxy-4-methoxybenzylideneamino)- 1,5-dimethyl-2-phenyl-1H-pyrazol- 3(2H)-one
Bioinformatics for molecular biology
 Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
Effects of Epicholesterol on the Phosphatidylcholine Bilayer: A Molecular Simulation Study
 1818 Biophysical Journal Volume 84 March 2003 1818 1826 Effects of Epicholesterol on the Phosphatidylcholine Bilayer: A Molecular Simulation Study Tomasz Róg and Marta Pasenkiewicz-Gierula Department of
1818 Biophysical Journal Volume 84 March 2003 1818 1826 Effects of Epicholesterol on the Phosphatidylcholine Bilayer: A Molecular Simulation Study Tomasz Róg and Marta Pasenkiewicz-Gierula Department of
Effect of Sodium and Chloride Binding on a Lecithin Bilayer. A Molecular Dynamics Study
 membranes Article Effect of Sodium and Chloride Binding on a Lecithin Bilayer. A Molecular Dynamics Study Maria M. Reif 1,2, Christopher Kallies 2 and Volker Knecht 2,3, * 1 Physics Department (T38), Technische
membranes Article Effect of Sodium and Chloride Binding on a Lecithin Bilayer. A Molecular Dynamics Study Maria M. Reif 1,2, Christopher Kallies 2 and Volker Knecht 2,3, * 1 Physics Department (T38), Technische
Molecular Structure and Permeability at the Interface between Phase-Separated Membrane Domains
 SUPPORTING INFORMATION Molecular Structure and Permeability at the Interface between Phase-Separated Membrane Domains Rodrigo M. Cordeiro Universidade Federal do ABC, Avenida dos Estados 5001, CEP 09210-580,
SUPPORTING INFORMATION Molecular Structure and Permeability at the Interface between Phase-Separated Membrane Domains Rodrigo M. Cordeiro Universidade Federal do ABC, Avenida dos Estados 5001, CEP 09210-580,
Interactions between Bisphosphate. Geminis and Sodium Lauryl Ether
 Chapter 5 Interactions between Bisphosphate Geminis and Sodium Lauryl Ether Sulphate 110 5.1 Introduction The physiochemical and surface active properties of mixed surfactants are of more interest and
Chapter 5 Interactions between Bisphosphate Geminis and Sodium Lauryl Ether Sulphate 110 5.1 Introduction The physiochemical and surface active properties of mixed surfactants are of more interest and
Membrane Binding and Self-Association of the Epsin N-Terminal Homology Domain
 http://dx.doi.org/10.1016/j.jmb.2012.08.010 J. Mol. Biol. (2012) 423, 800 817 Contents lists available at www.sciencedirect.com Journal of Molecular Biology journal homepage: http://ees.elsevier.com.jmb
http://dx.doi.org/10.1016/j.jmb.2012.08.010 J. Mol. Biol. (2012) 423, 800 817 Contents lists available at www.sciencedirect.com Journal of Molecular Biology journal homepage: http://ees.elsevier.com.jmb
