a-dS. Code assigned:
|
|
|
- Pierce Russell
- 5 years ago
- Views:
Transcription
1 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the separate document Help with completing a taxonomic proposal Please try to keep related proposals within a single document; you can copy the modules to create more than one genus within a new family, for example. MODULE 1: TITLE, AUTHORS, etc Code assigned: a-dS (to be completed by ICTV officers) Short title: Creation of three new species (Limnipivirus A, Limnipivirus B, Limnipivirus C) in a new genus (Limnipivirus). (e.g. 6 new species in the genus Zetavirus) Modules attached (modules 1 and 10 are required) Author(s): R. Zell, E. Delwart, A.E. Gorbalenya, T. Hovi, A.M.Q. King, N.J. Knowles, A.M. Lindberg, M.A. Pallansch, A.C. Palmenberg, G. Reuter, P. Simmonds, T. Skern, G. Stanway, T. Yamashita Corresponding author with address: Roland Zell (roland.zell@med.uni-jena.de) List the ICTV study group(s) that have seen this proposal: A list of study groups and contacts is provided at If in doubt, contact the appropriate subcommittee chair (fungal, invertebrate, plant, prokaryote or vertebrate viruses) Picornaviridae Study Group ICTV Study Group comments (if any) and response of the proposer: Date first submitted to ICTV: 29/06/2015 Date of this revision (if different to above): 06/07/2015 ICTV-EC comments and response of the proposer: Page 1 of 7
2 MODULE 2: NEW SPECIES creating and naming one or more new species. If more than one, they should be a group of related species belonging to the same genus. All new species must be placed in a higher taxon. This is usually a genus although it is also permissible for species to be unassigned within a subfamily or family. Wherever possible, provide sequence accession number(s) for one isolate of each new species proposed. Code aS (assigned by ICTV officers) To create 3 new species within: Genus: Limnipivirus (new) Subfamily: - Family: Picornaviridae Order: Picornavirales Name of new species: Limnipivirus A Limnipivirus B Representative isolate: (only 1 per species please) BGPV-1 bluegill/usa/04-032/2003 (bluegill picornavirus 1) CPV-1 F37/06 (carp picornavirus 1) Fill in all that apply. If the higher taxon has yet to be created (in a later module, below) write (new) after its proposed name. If no genus is specified, enter unassigned in the genus box. GenBank sequence accession number(s) JX KF3067 Limnipivirus C FHMPV-1 fhm/1/mn/usa/2010 (fathead minnow picornavirus 1) KF Reasons to justify the creation and assignment of the new species: Explain how the proposed species differ(s) from all existing species. o If species demarcation criteria (see module 3) have previously been defined for the genus, explain how the new species meet these criteria. o If criteria for demarcating species need to be defined (because there will now be more than one species in the genus), please state the proposed criteria. Further material in support of this proposal may be presented in the Appendix, Module 9 The three Limnipivirus species are new picornaviruses isolated from teleost fish of the Cyprinidae and Centrarchidae families from freshwater lakes and ponds in the USA and Germany. These viruses are most closely related to eel picornavirus 1, Avihepatovirus, Parechovirus and Pasivirus but show low amino acid identity with the orthologous proteins of these and other picornaviruses (capsid proteins 1AB, 1C, 1D: <25%, 2C Hel : <26%, 3C pro : <20%, 3D pol : <41%). The three Limnipivirus species have in common and unique genome organization: VPg+5'UTR typeiv [1AB-1C-1D-2A1 NPGP /2A2 NPGP /2B-2C Hel /3A-3B VPg -3C pro -3D pol ]3'UTR-poly(A) All limnipiviruses share a characteristic, truncated type IV IRES relative to hepatitis C virus (Asnani et al., Virology 478:61-74, 2015) and two aphthovirus-like 2A proteins with NPG P motif. Within the genus, 2B, 3A and 3B exhibit no similarity; amino acid identities of the concatenated sequences of the orthologous proteins is greater 48%: Page 2 of 7
3 Percent Identity Matrix 1AB-1C-1D-2C-3C-3D - created by Clustal2.1: JX134222_Bluegill_picornavirus_ KF3067_Carp_picornavirus_1_F37/ KC465953_Fathead_minnow_ KF4490_Fathead_minnow_fhm/20/IL/USA/ KF183915_Fathead_minnow_fhm/1/MN/USA/ KF183916_Fathead_minnow_fhm/2/MN/USA/ MODULE 3: NEW GENUS creating a new genus Ideally, a genus should be placed within a higher taxon. Code bS (assigned by ICTV officers) To create a new genus within: Subfamily: - Family: Picornaviridae Order: Picornavirales Fill in all that apply. If the higher taxon has yet to be created (in a later module, below) write (new) after its proposed name. If no family is specified, enter unassigned in the family box naming a new genus Code cS (assigned by ICTV officers) To name the new genus: Limnipivirus Assigning the type species and other species to a new genus Code dS (assigned by ICTV officers) To designate the following as the type species of the new genus Limnipivirus A Every genus must have a type species. This should be a well characterized species although not necessarily the first to be discovered The new genus will also contain any other new species created and assigned to it (Module 2) and any that are being moved from elsewhere (Module 7b). Please enter here the TOTAL number of species (including the type species) that the genus will contain: 3 Reasons to justify the creation of a new genus: Additional material in support of this proposal may be presented in the Appendix, Module 9 Limnipivirus has a unique picornavirus genome layout (3-4-4 type) and low amino acid identity to the conserved proteins of other picornaviruses. The capsid polypeptide VP0 is predicted to be uncleaved. There are two aphthovirus-like 2A proteins with a NPG P sequence motif separated by a stretch of more than 120 amino acids in all three species. The phylogenetic relationship of the capsid-encoding P1 region and the 3CD region with other picornaviruses is shown in Appendix Figures 2 and 3. Page 3 of 7
4 Origin of the new genus name: Limnipi: from Greek limne ( ), "lake", and pi from picornavirus. Reasons to justify the choice of type species: Limnipvirus A is easily isolated and frequently detected in bluegills; it was also detected in other fish of the Centrarchidae family and from common carp and channel catfish. Species demarcation criteria in the new genus: If there will be more than one species in the new genus, list the criteria being used for species demarcation and explain how the proposed members meet these criteria. According to current knowledge, members of the proposed Limnipivirus genus share less than 60% amino acid identity for the concatenated orthologous proteins 1AB-1C-1D-2C-3C-3D. Page 4 of 7
5 MODULE 10: APPENDIX: supporting material additional material in support of this proposal References: Barbknecht M, Sepsenwol S, Leis E, Tuttle-Lau M, Gaikowski M, Knowles NJ, Lasee B, Hoffman MA Characterization of a new picornavirus isolated from the freshwater fish Lepomis macrochirus. J. Gen. Virol. 95: Lange J, Groth M, Fichtner D, Granzow H, Keller B, Walther M, Platzer M, Sauerbrei A, Zell R Virus isolate from carp: genetic characterization reveals a novel picornavirus with two apthovirus 2A-like sequences. J. Gen. Virol. 95: Phelps NBD, Mor SK, Armien AG, Batts W, Goodwin AE, Hopper L, McCann R, Ng TFF, Puzach C, Waltzek TB, Delwart E, Winton J, Goyal SM Isolation and molecular characterization of a novel picornavirus from baitfish in the USA. PLoS One 9(2):e8. Asnani M, Kumar P, Hellen CUT Widespread distribution and structural diversity of type IV IRESs in members of Picornaviridae. Virology 478: Annex: Include as much information as necessary to support the proposal, including diagrams comparing the old and new taxonomic orders. The use of Figures and Tables is strongly recommended but direct pasting of content from publications will require permission from the copyright holder together with appropriate acknowledgement as this proposal will be placed on a public web site. For phylogenetic analysis, try to provide a tree where branch length is related to genetic distance. Genome Organisation: CPV-1 NPG/P BGPV-1 NPG/P FHMPV-1 NPG/P nt 1 5'-UTR 524 nt Type IV IRES nt 525 aa 1 1AB 238 aa nt 1238 aa 238 1C 229 aa CPV-1 E/S BGPV-1 E/G nt 1925 aa 467 1D 189 aa nt 2492 aa 656 nt 2645 aa 707 # 2A1 51 aa nt 3044 aa 840 # 2A2 133 aa 2B 2 aa nt 3749 aa 10 2C 337 aa CPV-1 E/G BGPV-1 Q/G FHMPV-1 E/D nt 5027 aa 1501 nt 4760 aa A 89 aa 3B 39 aa nt 5144 aa C 205 aa nt 59 aa 1745 CPV-1 E/G BGPV-1 E/G 3D 524 aa nt 7331 aa 22 nt '-UTR 301 nt CPV-1 E/A BGPV-1 Q/A CPV-1 E/A BGPV-1 Q/G FHMPV-1 Q/A CPV-1 Q/S BGPV-1 E/S CPV-1 NPG/P BGPV-1 NPG/P FHMPV-1 NPG/P CPV-1 E/A BGPV-1 Q/R FHMPV-1 E/A CPV-1 E/S BGPV-1 E/G FHMPV-1 Q/A Figure 1: Schematic depiction of the Limnipivirus genome (example CPV-1). The open reading frame is indicated by a box. Positions of putative nt and aa cleavage sites of CPV-1 and the lengths of the deduced proteins are shown. Grey boxes show a comparison of the cleavage sites. Triangles ( ) indicate the 3C pro cleavage sites; the hash (#) indicates both ribosomal skipping sites at the NPG P motif. Page 5 of 7
6 P HQ595340, unassigned, bat picornavirus 1 [NC16A] HQ595341, unassigned, bat picornavirus 1 [LMH22A] HQ595342, unassigned, bat picornavirus 2 [MH9F] HQ595343, unassigned, bat picornavirus 2 [SK17F] JN8316, unassigned, canine picornavirus 1 [dog/hong Kong/325F/2008] JQ814852, unassigned, Ia io picornavirus 1 KJ6414, unassigned, bat picornavirus [BtRs-PicoV/YN2010] KJ6413, unassigned, bat picornavirus [BtRh-PicoV/SC2013] HQ595344, unassigned, bat picornavirus 3 [TLC5F] HQ595345, unassigned, bat picornavirus 3 [TLC21F] KJ6416, unassigned, bat picornavirus [BtMf-PicoV/FJ2012] KJ6410, unassigned, bat picornavirus [BtMf-PicoV-1/GD2012] KJ6416, unassigned, bat picornavirus [BtMf-PicoV-2/SAX2011] JN2116, unassigned, feline picornavirus [127F] JN2118, unassigned, feline picornavirus [661F] JN 2115, unassigned, feline picornavirus 1 [cat/hong Kong/073F/2007] JN2117, unassigned, feline picornavirus [6F] JN2119, unassigned, feline picornavirus [1021F] KJ641689, unassigned, bat picornavirus [BtMa-PicoV/FJ2012] KJ6416, unassigned, bat picornavirus [BtVs-PicoV/SC2013] AF406813, Sapelovirus A, PSV-1 [V13] AY064708, Sapelovirus B, SSV-1 [SV2-2383] KJ6416, unassigned, bat picornavirus [BtNv-PicoV/SC2013] KJ641685, unassigned, bat picornavirus [BtRf-PicoV-1/YN2012] KJ6412, unassigned, bat picornavirus [BtRa-PicoV/JS2013] KJ641688, unassigned, bat picornavirus [BtRlep-PicoV/FJ2012] JN420368, unassigned, California sealion [11] AY563023, Avian sapelovirus, ASV-1 [TW90A] JN674502, unassigned, quail picornavirus 1 [quail/hun/2010] FR727144, unassigned, pigeon picornavirus B [pigeon/norway/03/641/2003] KC560801, unassigned, pigeon picornavirus B [GAL-7/2010/Hungary] KP2338, unassigned, rabovirus [A Berlin/Jan2011/02] KJ950883, unassigned, rat picornavirus [RPV/NYC-B10] K02121, Rhinovirus B, HRV-14 [1959] EF186077, Rhinovirus C, HRV-C3 [QPM] L24917, Rhinovirus A, HRV-16 [117] V01149, Enterovirus C, EV-C PV-1 [Mahoney] M33854, Enterovirus B, EV-B CVB3 [Nancy] AF201894, Enterovirus H, EV-H1 SEV-A1 A-2 plaque D000, Enterovirus D, EV-D70 [J670/71] U22521, Enterovirus A, EV-A71 [BrCr] AF326766, Enterovirus J, EV-J1 [SV6-1631] AF363453, Enterovirus G, EV-G1 (former PEV-9) [UKG/410/73] DQ0927, Enterovirus E, EV-E1 (former BEV-1) [LC-R4] DQ092770, Enterovirus F, EV-F1 (former BEV-2) [BEV-261 RM2] KM3707, unassigned, lesavirus 1 [Mis101308/2012] KM3708, unassigned, lesavirus 2 [Nai108015/2012] JQ941880, Hunnivirus A, hunnivirus A1 [cattle/2008/hun] AF2317, Teschovirus A, porcine teschovirus 1 [Talfan] X001, Foot-and-mouth disease virus, FMDV O1 [Kaufbeuren] JN06, Bovine rhinitis A virus, BRAV-2 [H-1] EU236594, Bovine rhinitis B virus, BRBV-1 [EC11] X0, Equine rhinitis A virus, ERAV [PERV] X1, Erbovirus A, equine rhinitis B virus 1 [P1436/71] KM5898, unassigned, bovine picornavirus [TCH6] JF36, Mosavirus A, mosavirus A1 [mouse/m-7/usa/2010] KM3616, unassigned, tortoise picornavirus [ ] JQ864242, Cardiovirus C, Boone cardiovirus [rat/usa/2010] JX683808, Cardiovirus C, Boone cardiovirus [rat/usa/2012] M81861, Cardiovirus A, Encephalomyocarditis virus 1 [R] M205, Cardiovirus B, Theiler's murine encephalomyelitis virus [GDVII] JQ814851, Mischivirus A, Miniopterus schreibersii picornavirus 1 [bat/china/2010] KP644, unassigned, African bat icavirus A [TNo13] DQ6412, Senecavirus A, Seneca Valley virus 1 [US/SVV-001] FJ438908, Cosavirus D,human cosavirus D1 [5004] FJ438902, Cosavirus A, human cosavirus A1 [0553] JN8678, Cosavirus F, human cosavirus F1 [PK5006] FJ438907, Cosavirus B, human cosavirus B1 [2263] FJ555055, Cosavirus E, human cosavirus E1 [Australia/81] JN819202, Cadicivirus A, canine picodicistrovirus [209] JF3686, Rosavirus A, rosavirus A1 [mouse/usa/m-7/2010] HM11, Melegrivirus A, turkey hepatitis virus [23D] KC6638, unassigned, duck megrivirus [LY] KC6003, unassigned, mesivirus [HK21] KC811837, unassigned, mesivirus 2 [pigeon/galii5-pimev/2011/hun] KJ415177, unassigned, tortoise rafivirus A [UF4] GU1408, Oscivirus A, oscivirus A1 [robin/hong Kong/10717/2006] GU1410, Oscivirus A, oscivirus A2 [robin/hong Kong/108/2006] KF741227, Sicinivirus A, sicinivirus A1 [chicken/ucc001/eire] KF31, Sicinivirus A, sicinivirus A1 [ChPV-1 chicken/55c/hong Kong/2008] JQ1613, Gallivirus A, gallivirus A1 [turkey/m176/2011/hun] GU1406, Passerivirus A, passerivirus A1 [thrush/hong Kong/006/2007] KF3721, Sakobuvirus A, sakobuvirus A1 [FFUP1/Portugal/2012] GQ1740, Salivirus A, salivirus A1 [NG-J1] KJ6411, unassigned, bat picornavirus [BtMf-PicoV-2/GD2012] KJ641686, unassigned, bat picornavirus [BtMr-PicoV/JX2010] AB010145, Aichivirus A, AiV-A1 [A846/88] JN3133, Aichivirus A, CKV-1 [dog/an211d/usa/2009] JF5427, Aichivirus A, MKV-1 [M-5/USA/2010] EU74590, Aichivirus C, PKV [swine/s-1-hun-2007/hungary] AB084788, Aichivirus B, BKV-1 [U-1] KF0065, Aichivirus B, ferret kobuvirus [ferret/mpkov38/nl] KF183915, Limnipivirus C*, fathead minnow picornavirus [fhm/1/mn/usa/2010] KF183916, Limnipivirus C*, fathead minnow picornavirus [fhm/2/mn/usa/201] KF4490, Limnipivirus C*, fathead minnow picornavirus [fhm/20/il/usa/2010] KC465953, Limnipivirus C*, fathead minnow picornavirus [09-283] KF3067, Limnipivirus B*, carp picornavirus 1 [F37/06] JX134222, Limnipivirus A*, bluegill picornavirus 1 [04-032] KC465954, Avisivirus A, avisivirus A1 [turkey/m176-tuasv/2011/hun] KC614703, Avisivirus A, avisivirus A1 [USA-IN1] KM203656, unassigned, chicken orivirus 1 [chicken/pf-chk1/2013/hun] DQ2492, Avihepatovirus A, duck hepatitis A virus 1 [03D] KJ0006, unassigned, aalivirus A1 [duck picornavirus GL/12] JQ316470, Pasivirus A, pasivirus A1 [swine/france/2011] AB79, unassigned, crohivirus 1 [shrew/zm54/zambia/2012] KJ6416, unassigned, bat picornavirus isolate [BtMf-PicoV-1/SAX2011] JQ814853, unassigned, Rhinolophus affinis picornavirus 1 KC79, Kunsagivirus A, kunsagivirus A1 [roller/szal6-kuv/2011/hun] unassigned, Bat Kunsagivirus EU142040, Aquamavirus A, Aquamavirus A1 (SePV-1) [HO-02-21] KC8437, Potamipivirus A*, eel picornavirus [F15-05] KF00, unassigned, ferret parechovirus [ferret/mppev1/nl] L021, Parechovirus A, human parechovirus 1 [Harris] AF327920, Parechovirus B, ljunganvirus 1 [-012] KP230449, unassigned, falcovirus A1 [kestrel/vove02/2013/hun] KJ641684, unassigned, bat picornavirus [BtRf-PicoV-2/YN2012] AJ225173, Tremovirus A, avian encephalomyelitis virus 1 [Calnek vaccine strain] M14707, Hepatovirus A, human hepatitis A virus [HM-1] Genus: Sapelovirus Sapelovirus Enterovirus Hunnivirus Teschovirus Aphthovirus Erbovirus Mosavirus Cardiovirus Mischivirus Senecavirus Cosavirus Dicipivirus Rosavirus Megrivirus Oscivirus Sicinivirus Gallivirus Passerivirus Sakobuvirus Salivirus Kobuvirus Limnipivirus* Avisivirus Avihepatovirus Pasivirus Kunsagivirus Aquamavirus Potamipivirus* Parechovirus Tremovirus Hepatovirus Figure 2: Maximum likelihood tree of picornavirus P1 gene region. 118 picornavirus sequences retrieved from GenBank were included. Presented are GenBank accession numbers, species names (in bold print and underlined) and type designations. If available, designations of isolates and sequenced specimens, respectively, are given in square brackets. Unassigned viruses are printed in blue. Proposed names are printed in red and indicated by an asterisk (*). Braces indicate acknowledged and proposed (*) genera. Numbers at nodes indicate bootstrap support obtained after 0 replicates. The scale indicates substitutions/site. Page 6 of 7
7 3CD JN2117, unassigned, feline picornavirus 1 [6F] JN2119, unassigned, feline picornavirus 1 [1021F] JN2115, unassigned, feline picornavirus 1 [cat/hong Kong/073F/2007] JN2116, unassigned, feline picornavirus [127F] JN2118, unassigned, feline picornavirus [661F] KJ6413, unassigned, bat picornavirus [BtRh-PicoV/SC2013] HQ595344, unassigned, bat picornavirus 3 [TLC5F] HQ595345, unassigned, bat picornavirus 3 [TLC21F] JN8316, unassigned, canine picornavirus 1 [dog/hong Kong/325F/2008] JQ814852, unassigned, Ia io picornavirus 1 HQ595340, unassigned, bat picornavirus 1 [NC16A] HQ595341, unassigned, bat picornavirus 1 [LMH22A] HQ595342, unassigned, bat picornavirus 2 [MH9F] HQ595343, unassigned, bat picornavirus 2 [SK17F] JN420368, unassigned, Californian sealion sapelovirus 1 [11] AF406813, Sapelovirus A, PSV-1 (former PEV8) [V13] AY064708, Sapelovirus B, SSV-1 [SV2-2383] AY563023, Avian Sapelovirus, ASV-1 [TW90A] JN674502, unassigned, quail picornavirus 1 [quail/hun/2010] FR727145, unassigned, pigeon picornavirus A [pigeon/norway/03/603-7/2003] FR727144, unassigned, pigeon picornavirus B [pigeon/norway/03/641/2003] KC560801, unassigned, pigeon picornavirus B [GAL-7/2010/Hungary] KP2338, unassigned, rabovirus A [Berlin/Jan2011/02] KJ950883, unassigned, rat picornavirus A [rat/nyc-b10/usa/2010] L24917, Rhinovirus A, HRV-16 [117] EF186077, Rhinovirus C, HRV-C3 [QPM] K02121, Rhinovirus B, HRV-14 [1959] AF201894, Enterovirus H, EV-H1 (former SEV-A1) [A-2 plaque] DQ0927, Enterovirus E, EV-E1 (former BEV-1) [LC-R4] DQ092770, Enterovirus F, EV-F1 (former BEV-2) [BEV-261 RM2] AF363453, Enterovirus G, EV-G1 (former PEV-9) [UKG/410/73] U22521, Enterovirus A, EV-A71 [BrCr] M33854, Enterovirus B, EV-B CV-B3 [Nancy] AF326766, Enterovirus J, EV-J1 (SV6) [1631] D000, Enterovirus D, EV-D70 [J670-71] V01149, Enterovirus C, EV-C PV-1 [Mahoney] AJ225173, Tremovirus A, avian encephalomyelitisvirus 1 [Calnek vaccine strain] M14707, Hepatovirus A, human hepatitis A virus [HM-1] KP230449, unassigned, falcovirus A1 [kestrel/vove02/2013/hun] KC79, Kunsagivirus A, kunsagivirus A1 [roller/szal6-kuv/2011/hun] EU142040, Aquamavirus A, aquamavirus A1 (SePV-1) [HO-02-21] KF , Limnipivirus C*,fathead minnow picornavirus [fhm/1/mn/usa/2010] KF , Limnipivirus C*,fathead minnow picornavirus [fhm/2/mn/usa/2010] KF4490, Limnipivirus C*, fathead minnow picornavirus [fhm/20/il/usa/2010] KF3067, Limnipivirus B*, carp picornavirus 1 [F37/06] JX134222, Limnipivirus A*, bluegill picornavirus 1 [04-032] KC8437, Potamipivirus A*, eel picornavirus 1 [F15/05] KC465954, Avisivirus A, Asv-A1 [turkey/m176-tuasv/2011/hun] KC614703, Avisivirus A, AsV-A1 [turkey/usa/in1/2010] KJ0006, unassigned, aalivirus A1 [duck/gl/12/china/2012] DQ2492, Avihepatovirus A, duck hepatitis A virus 1 [03D] KM203656, unassigned, chicken orivirus 1 [chicken/pf-chk1/2013/hun] KJ6416, unassigned, bat picornavirus [bat/btmf-picov-1/sax2011] JQ316470, Pasivirus A, PaV-A1 [swine/france/2011] AB79, unassigned, crohivirus 1 [shrew/zm54/zambia/2012] KF00, unassigned, ferret parechovirus [ferret/mppev1/nl] L021, Parechovirus A, human parechovirus 1 [Harris] AF327920, Parechovirus B, ljunganvirus 1 [-012] HF677705, unassigned, sebokele virus 1 [An B 1227 d] JN819202, Cadicivirus A, canine picodicistrovirus [209] JF3686, Rosavirus A, RoV-A1 [mouse/usa/m-7/2010] KF1186, Melegrivirus A, chicken megrivirus [chicken/b21-chv/2012/hun] KF11, Melegrivirus A, chicken megrivirus [chicken/chk-iv-chv/2013/hun] KJ09, Melegrivirus A, chicken proventriculitis virus [CPV/Korea/03] HM11, Melegrivirus A, turkey hepatitis virus [23D] KF36, Melegrivirus A, chicken picornavirus 5 [27C] KC6638, unassigned, duck megrivirus [LY ] KC811837, unassigned, pigeon mesivirus 2 [pigeon/galii5-pimev/2011/hun] KC6003, unassigned, mesivirus [HK21] KJ415177, unassigned, tortoise rafivirus A [UF4] GU1408, Oscivirus A, oscivirus A1 [robin/hong Kong/10717/2006] GU1410, Oscivirus A, oscivirus A2 [robin/hong Kong/108/2006] KF741227, Sicinivirus A, sicinivirus A1 [chicken/ucc001/eire] KF31 Sicinivirus A, sicinivirus A1 (ChPV-1) [chicken/55c/hong Kong/2008] GU1406, Passerivirus A, passerivirus A1 [thrush/hong Kong/006/2007] JQ1613, Gallivirus A, gallivirus A1 [turkey/hun/m176/2011] KF3721, Sakobuvirus A, sakobuvirus A1 [FFUP1/Portugal/2012] KJ641686, unassigned, bat picornavirus [BtMr-PicoV/JX2010] KJ6411, unassigned, bat picornavirus [BtMf-PicoV-2/GD2012] GQ1740, Salivirus A, salivirus A1 [hu/ng-j1/nigeria/2007] JF5427, Aichivirus A, MKV-1 [M-5/USA/2010] JN3133, Aichivirus A, CKV-1 [dog/an211d/usa/2009] AB010145, Aichivirus A, AiV-1 [A846-88] EU7450, Aichivirus C, PKV-1 [sw/s-1-hun/2007] AB084788, Aichivirus B, BKV-1 [U-1] KF0065, Aichivirus B, ferret kobuvirus [ferret/mpkov38/nl/2010]] JF36, Mosavirus A, mosavirus A1 [mouse/m-7/usa/2010] KM3616, unassigned, tortoise picornavirus [ ] KM3617, unassigned, tortoise picornavirus [5-03] KM3612, unassigned, tortoise picornavirus [5-04] KM3611, unassigned, tortoise picornavirus [14-04] KM3615, unassigned, tortoise picornavirus [2013-T4] KM3613, unassigned, tortoise picornavirus [9-05] KM3614, unassigned, tortoise picornavirus [144-10] KM3707, unassigned, lesavirus 1 [Mis101308/2012] KM3708, unassigned, lesavirus 2 [Nai108015/2012] JQ941880, Hunnivirus A, hunnivirus A1 [cattle/hun/2008] AF2317, Teschovirus A, porcine teschovirus 1 [Talfan] X1, Erbovirus A, equine rhinitis B virus 1 [P1436/71] KM5898, unassigned, bovine picornavirus [TCH6] EU236594, Bovine rhinitis B virus, BRBV-1 [EC11] JN06, Bovine rhinitis A virus, BRAV-2 [H-1] X001, Foot-and-mouth disease virus, FMDV O1 [Kaufbeuren] X0, Equine rhinitis A virus, ERAV [PERV] FJ438902, Cosavirus A, human cosavirus A1 [0553] JN8678, Cosavirus F, human cosavirus F1 [PK5006] FJ438907, Cosavirus B, human cosavirus B1 [2263] FJ438908, Cosavirus D, human cosavirus D1 [5004] FJ555055, Cosavirus E, human cosavirus E1 [Australia/81] JQ814851, Mischivirus A, Miniopterus schreibersii picornavirus 1 [bat/china/2010] KP644, unassigned, African bat icavirus A [TNo13] DQ6412, Senecavirus A, Seneca Valley virus 1 [US/SVV-001] M205, Cardiovirus B, Theiler's murine encephalomyelitis virus [GDVII] M81861, Cardiovirus A, encephalomyocarditis virus [R] 89 JQ864242, Cardiovirus C, Boone cardiovirus 1 [rat/usa/2010] JX683808, Cardiovirus C, Boone cardiovirus 2 [rat/usa/2012] KP770140, unassigned, ampivirus A1 [NEWT/2013/HUN] Genus: Sapelovirus Enterovirus Tremovirus Hepatovirus Kunsagivirus Aquamavirus Limnipivirus* Potamipivirus* Avisivirus Avihepatovirus Pasivirus Parechovirus Dicipivirus Rosavirus Megrivirus Oscivirus Sicinivirus Passerivirus Gallivirus Sakobuvirus Salivirus Kobuvirus Mosavirus Hunnivirus Teschovirus Erbovirus Aphthovirus Cosavirus Mischivirus Senecavirus Cardiovirus 0.2 Figure 3: Maximum likelihood tree of picornavirus 3CD gene region. 117 picornavirus sequences retrieved from GenBank were included. Presented are GenBank accession numbers, species names (in bold print and underlined) and type designations. If available, designations of isolates and sequenced specimens, respectively, are given in square brackets. Unassigned viruses are printed in blue. Proposed names are indicated by an asterisk (*). Braces indicate acknowledged and proposed (*) genera. Numbers at nodes indicate bootstrap support obtained after 0 replicates. The scale indicates substitutions/site. Page 7 of 7
a-dS (modules 1 and 11 are required)
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
aV. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
a-dS. Short title: Create 1 new species (Ampivirus A) in a new genus (Ampivirus) (e.g. 6 new species in the genus Zetavirus) Modules attached
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
a-dV. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Virus isolate from carp: genetic characterization reveals a novel picornavirus with two aphthovirus 2A-like sequences
 Journal of General Virology (2014), 95, 80 90 DOI 10.1099/vir.0.058172-0 Virus isolate from carp: genetic characterization reveals a novel picornavirus with two aphthovirus 2A-like sequences Jeannette
Journal of General Virology (2014), 95, 80 90 DOI 10.1099/vir.0.058172-0 Virus isolate from carp: genetic characterization reveals a novel picornavirus with two aphthovirus 2A-like sequences Jeannette
Identification and characterization of novel picornaviruses by molecular biological methods. Ph.D. Thesis. Péter Pankovics
 Identification and characterization of novel picornaviruses by molecular biological methods Ph.D. Thesis Péter Pankovics Supervisor: 1 Gábor Reuter, M.D., Ph.D., Med. Habil. Program leader: 2 Levente Emődy,
Identification and characterization of novel picornaviruses by molecular biological methods Ph.D. Thesis Péter Pankovics Supervisor: 1 Gábor Reuter, M.D., Ph.D., Med. Habil. Program leader: 2 Levente Emődy,
aD (modules 1 and 10 are required)
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
aV (modules 1 and 9 are required)
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
a-hV. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
aM. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
aP. Code assigned: Short title: Remove (abolish) the species Narcissus symptomless virus in the genus Carlavirus, family Betaflexiviridae
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
aM (modules 1 and 10 are required)
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
The Origin, Evolution and Diagnosis of Seneca Valley Virus, a New Vesicular Disease-Causing Picornavirus of Pigs
 The Origin, Evolution and Diagnosis of Seneca Valley Virus, a New Vesicular Disease-Causing Picornavirus of Pigs Nick J. Knowles, Katarzyna Bachanek-Bankowska, Jemma Wadsworth, Veronica L. Fowler & Donald
The Origin, Evolution and Diagnosis of Seneca Valley Virus, a New Vesicular Disease-Causing Picornavirus of Pigs Nick J. Knowles, Katarzyna Bachanek-Bankowska, Jemma Wadsworth, Veronica L. Fowler & Donald
a,bD. Code assigned: Short title: Two new species in the genus Orthohepadnavirus (e.g. 6 new species in the genus Zetavirus) Modules attached
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
ÁNTSZ Regional Institute of State Public Health Service, Pécs, Hungary
 Journal of eneral Virology (2013), 94, 2029 2035 DOI 10.1099/vir.0.054676-0 Short ommunication orrespondence ábor Reuter reuter.gabor@ddr.antsz.hu enetic characterization of a novel picornavirus distantly
Journal of eneral Virology (2013), 94, 2029 2035 DOI 10.1099/vir.0.054676-0 Short ommunication orrespondence ábor Reuter reuter.gabor@ddr.antsz.hu enetic characterization of a novel picornavirus distantly
aV. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
aV. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Analysis of the interaction between the 2C protein of parechoviruses and lipid droplets. Ali Aidarous Khrid
 Analysis of the interaction between the 2C protein of parechoviruses and lipid droplets Ali Aidarous Khrid A thesis submitted for the degree of Doctor of Philosophy School of Biological Sciences University
Analysis of the interaction between the 2C protein of parechoviruses and lipid droplets Ali Aidarous Khrid A thesis submitted for the degree of Doctor of Philosophy School of Biological Sciences University
aV (modules 1 and 9 are required)
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Isolation and Molecular Characterization of a Novel Picornavirus from Baitfish in the USA
 Isolation and Molecular Characterization of a Novel Picornavirus from Baitfish in the USA Nicholas B. D. Phelps 1,2 *, Sunil K. Mor 1, Anibal G. Armien 1,2, William Batts 3, Andrew E. Goodwin 4, Lacey
Isolation and Molecular Characterization of a Novel Picornavirus from Baitfish in the USA Nicholas B. D. Phelps 1,2 *, Sunil K. Mor 1, Anibal G. Armien 1,2, William Batts 3, Andrew E. Goodwin 4, Lacey
Aichi Virus 1: Environmental Occurrence and Behavior
 Pathogens 2015, 4, 256-268; doi:10.3390/pathogens4020256 Review OPEN ACCESS pathogens ISSN 2076-0817 www.mdpi.com/journal/pathogens Aichi Virus 1: Environmental Occurrence and Behavior Masaaki Kitajima,
Pathogens 2015, 4, 256-268; doi:10.3390/pathogens4020256 Review OPEN ACCESS pathogens ISSN 2076-0817 www.mdpi.com/journal/pathogens Aichi Virus 1: Environmental Occurrence and Behavior Masaaki Kitajima,
Complete Genome Analysis of Three Novel Picornaviruses from Diverse Bat Species
 JOURNAL OF VIROLOGY, Sept. 2011, p. 8819 8828 Vol. 85, No. 17 0022-538X/11/$12.00 doi:10.1128/jvi.02364-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Complete Genome Analysis
JOURNAL OF VIROLOGY, Sept. 2011, p. 8819 8828 Vol. 85, No. 17 0022-538X/11/$12.00 doi:10.1128/jvi.02364-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Complete Genome Analysis
Yip, CY; Lo, KL; Que, TL; Lee, RA; Chan, KH; Yuen, KY; Woo, PCY; Lau, SKP
 Title Author(s) Epidemiology of human parechovirus, Aichi virus and salivirus in fecal samples from hospitalized children with gastroenteritis in Hong Kong Yip, CY; Lo, KL; Que, TL; Lee, RA; Chan, KH;
Title Author(s) Epidemiology of human parechovirus, Aichi virus and salivirus in fecal samples from hospitalized children with gastroenteritis in Hong Kong Yip, CY; Lo, KL; Que, TL; Lee, RA; Chan, KH;
ICTVdB Virus Descriptions
 [Home] [Index of Viruses ] [Descriptions] [ Character List ] [ Picture Gallery ] [ Interactive Key ] [ Data Entry] [ 2002 ICTV] ICTVdB Virus Descriptions Descriptions are generated automatically from the
[Home] [Index of Viruses ] [Descriptions] [ Character List ] [ Picture Gallery ] [ Interactive Key ] [ Data Entry] [ 2002 ICTV] ICTVdB Virus Descriptions Descriptions are generated automatically from the
aP (modules 1 and 9 are required) Caulimoviridae
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
aD. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
bV. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Evaluation of Seneca Valley Virus Levels Mouse to Man
 Evaluation of Seneca Valley Virus Levels Mouse to Man EMEA/IHC Meeting on Virus Shedding Rotterdam, The Netherlands October 30, 2007 Paul Hallenbeck Neotropix, Inc. 351 Phoenixville Pike Malvern, PA 19380
Evaluation of Seneca Valley Virus Levels Mouse to Man EMEA/IHC Meeting on Virus Shedding Rotterdam, The Netherlands October 30, 2007 Paul Hallenbeck Neotropix, Inc. 351 Phoenixville Pike Malvern, PA 19380
Molecular analysis of duck hepatitis virus type 1 reveals a novel lineage close to the genus Parechovirus in the family Picornaviridae
 Journal of General Virology (2006), 87, 3307 3316 DOI 10.1099/vir.0.81804-0 Molecular analysis of duck hepatitis virus type 1 reveals a novel lineage close to the genus Parechovirus in the family Picornaviridae
Journal of General Virology (2006), 87, 3307 3316 DOI 10.1099/vir.0.81804-0 Molecular analysis of duck hepatitis virus type 1 reveals a novel lineage close to the genus Parechovirus in the family Picornaviridae
The vast majority of emerging infectious diseases (EIDs) in humans
 RESEARCH ARTICLE crossmark Discovery of a Novel Hepatovirus (Phopivirus of Seals) Related to Human Hepatitis A Virus S. J. Anthony, a,b,c J. A. St. Leger, d E. Liang, a,c A. L. Hicks, a M. D. Sanchez-Leon,
RESEARCH ARTICLE crossmark Discovery of a Novel Hepatovirus (Phopivirus of Seals) Related to Human Hepatitis A Virus S. J. Anthony, a,b,c J. A. St. Leger, d E. Liang, a,c A. L. Hicks, a M. D. Sanchez-Leon,
CELL SURFACE INTERACTIONS OF COXSACKIEVIRUS A9 AND HUMAN PARECHOVIRUS 1. Pirjo Merilahti
 CELL SURFACE INTERACTIONS OF COXSACKIEVIRUS A9 AND HUMAN PARECHOVIRUS 1 Pirjo Merilahti TURUN YLIOPISTON JULKAISUJA ANNALES UNIVERSITATIS TURKUENSIS Sarja - ser. D osa - tom. 1283 Medica - Odontologica
CELL SURFACE INTERACTIONS OF COXSACKIEVIRUS A9 AND HUMAN PARECHOVIRUS 1 Pirjo Merilahti TURUN YLIOPISTON JULKAISUJA ANNALES UNIVERSITATIS TURKUENSIS Sarja - ser. D osa - tom. 1283 Medica - Odontologica
Picornaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Picornaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=1) Diameter of 30 nm Genome Linear single-stranded RNA, positive
Picornaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=1) Diameter of 30 nm Genome Linear single-stranded RNA, positive
Complete genome sequence analysis of Seneca Valley virus-001, a novel oncolytic picornavirus
 Journal of General Virology (2008), 89, 1265 1275 DOI 10.1099/vir.0.83570-0 Complete genome sequence analysis of Seneca Valley virus-001, a novel oncolytic picornavirus Laura M. Hales, 1 3 Nick J. Knowles,
Journal of General Virology (2008), 89, 1265 1275 DOI 10.1099/vir.0.83570-0 Complete genome sequence analysis of Seneca Valley virus-001, a novel oncolytic picornavirus Laura M. Hales, 1 3 Nick J. Knowles,
Sequence analysis for VP4 of enterovirus 71 isolated in Beijing during 2007 to 2008
 16 2009 3 4 1 Journal of Microbes and Infection, March 2009, Vol. 4, No. 1 2007 2008 71 VP4 1, 2, 2, 2, 1, 2, 2, 2, 1, 2 1., 100730; 2., 100020 : 2007 2008 71 ( EV71), 2007 3 EV71( 1, 2 ) 2008 5 EV71(
16 2009 3 4 1 Journal of Microbes and Infection, March 2009, Vol. 4, No. 1 2007 2008 71 VP4 1, 2, 2, 2, 1, 2, 2, 2, 1, 2 1., 100730; 2., 100020 : 2007 2008 71 ( EV71), 2007 3 EV71( 1, 2 ) 2008 5 EV71(
Recombination and Selection in the Evolution of Picornaviruses and Other Mammalian Positive-Stranded RNA Viruses
 JOURNAL OF VIROLOGY, Nov. 2006, p. 11124 11140 Vol. 80, No. 22 0022-538X/06/$08.00 0 doi:10.1128/jvi.01076-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Recombination and Selection
JOURNAL OF VIROLOGY, Nov. 2006, p. 11124 11140 Vol. 80, No. 22 0022-538X/06/$08.00 0 doi:10.1128/jvi.01076-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Recombination and Selection
aP. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Sequencing of Porcine Enterovirus Groups II and III Reveals Unique Features of Both Virus Groups
 REFERENCES CONTENT ALERTS Sequencing of Porcine Enterovirus Groups II and III Reveals Unique Features of Both Virus Groups Andi Krumbholz, Malte Dauber, Andreas Henke, Eckhard Birch-Hirschfeld, Nick J.
REFERENCES CONTENT ALERTS Sequencing of Porcine Enterovirus Groups II and III Reveals Unique Features of Both Virus Groups Andi Krumbholz, Malte Dauber, Andreas Henke, Eckhard Birch-Hirschfeld, Nick J.
Picornaviruses (family Picornaviridae), with small, nonenveloped
 omparative omplete enome nalysis of hicken and Turkey Megriviruses (Family Picornaviridae): Long 3= ntranslated Regions with a Potential Second Open Reading Frame and Evidence for Possible Recombination
omparative omplete enome nalysis of hicken and Turkey Megriviruses (Family Picornaviridae): Long 3= ntranslated Regions with a Potential Second Open Reading Frame and Evidence for Possible Recombination
Taxonomic changes and additions for human and animal viruses,
 JCM Accepted Manuscript Posted Online 19 October 2016 J. Clin. Microbiol. doi:10.1128/jcm.01525-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 3 Taxonomic changes and additions
JCM Accepted Manuscript Posted Online 19 October 2016 J. Clin. Microbiol. doi:10.1128/jcm.01525-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 3 Taxonomic changes and additions
Enterovirus D68-US Outbreak 2014 Laboratory Perspective
 Enterovirus D68-US Outbreak 2014 Laboratory Perspective Allan Nix Team Lead Picornavirus Laboratory Polio and Picornavirus Laboratory Branch PAHO / SARInet Webinar November 6, 2014 National Center for
Enterovirus D68-US Outbreak 2014 Laboratory Perspective Allan Nix Team Lead Picornavirus Laboratory Polio and Picornavirus Laboratory Branch PAHO / SARInet Webinar November 6, 2014 National Center for
Vincent Racaniello
 Vincent Racaniello vrr1@columbia.edu www.virology.ws Poliomyelitis Polio (grey), myelon (marrow) = Greek itis (inflammation of) = Latin A common, acute viral disease characterized clinically by a brief
Vincent Racaniello vrr1@columbia.edu www.virology.ws Poliomyelitis Polio (grey), myelon (marrow) = Greek itis (inflammation of) = Latin A common, acute viral disease characterized clinically by a brief
aP. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
A highly divergent picornavirus in a marine mammal. ACCEPTED
 JVI Accepts, published online ahead of print on 17 October 2007 J. Virol. doi:10.1128/jvi.01240-07 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JVI Accepts, published online ahead of print on 17 October 2007 J. Virol. doi:10.1128/jvi.01240-07 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
University of Canberra Research Publication Collection
 - University of Canberra Research Publication Collection Faculty of Health 2012 - Peer-Reviewed Journal Article for Dr. Reena Ghildyal (Associate Professor, University of Canberra) This is the peer reviewed
- University of Canberra Research Publication Collection Faculty of Health 2012 - Peer-Reviewed Journal Article for Dr. Reena Ghildyal (Associate Professor, University of Canberra) This is the peer reviewed
NOTES. Characterization of a Canine Homolog of Human Aichivirus. Received 6 June 2011/Accepted 22 August 2011
 JOURNAL OF VIROLOGY, Nov. 2011, p. 11520 11525 Vol. 85, No. 21 0022-538X/11/$12.00 doi:10.1128/jvi.05317-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. NOTES Characterization
JOURNAL OF VIROLOGY, Nov. 2011, p. 11520 11525 Vol. 85, No. 21 0022-538X/11/$12.00 doi:10.1128/jvi.05317-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. NOTES Characterization
Susanne Johansson, 1 Bo Niklasson, 2 Jacob Maizel, 3 Alexander E. Gorbalenya, 4 * and A. Michael Lindberg 1 *
 JOURNAL OF VIROLOGY, Sept. 2002, p. 8920 8930 Vol. 76, No. 17 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.17.8920 8930.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Molecular
JOURNAL OF VIROLOGY, Sept. 2002, p. 8920 8930 Vol. 76, No. 17 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.17.8920 8930.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Molecular
Novel Picornavirus in Turkey Poults with Hepatitis, California, USA
 RESEARCH Novel Picornavirus in Turkey Poults with Hepatitis, California, USA Kirsi S. Honkavuori, H. L. Shivaprasad, Thomas Briese, Craig Street, David L. Hirschberg, Stephen K. Hutchison, and W. Ian Lipkin
RESEARCH Novel Picornavirus in Turkey Poults with Hepatitis, California, USA Kirsi S. Honkavuori, H. L. Shivaprasad, Thomas Briese, Craig Street, David L. Hirschberg, Stephen K. Hutchison, and W. Ian Lipkin
Introduction: RNA viruses
 SECTION I Introduction: RNA viruses Carol Shoshkes Reiss Viruses that infect the central nervous system may cause acute, chronic, or latent infections. In some cases, the diseases manifested are attributable
SECTION I Introduction: RNA viruses Carol Shoshkes Reiss Viruses that infect the central nervous system may cause acute, chronic, or latent infections. In some cases, the diseases manifested are attributable
First report of human salivirus/klassevirus in respiratory specimens of a child with fatal adenovirus infection
 Virus Genes (2016) 52:620 624 DOI 10.1007/s11262-016-1361-7 First report of human salivirus/klassevirus in respiratory specimens of a child with fatal adenovirus infection Na Pei 1,3 Jiaosheng Zhang 2
Virus Genes (2016) 52:620 624 DOI 10.1007/s11262-016-1361-7 First report of human salivirus/klassevirus in respiratory specimens of a child with fatal adenovirus infection Na Pei 1,3 Jiaosheng Zhang 2
Recombinaison expérimentale intra-et interespèce chez les Rhinovirus et Quantification d'arn de Rhinovirus par RT-PCR en temps réel.
 Thesis Recombinaison expérimentale intra-et interespèce chez les Rhinovirus et Quantification d'arn de Rhinovirus par RT-PCR en temps réel SCHIBLER, Manuel Abstract Les rhinovirus sont des petits virus
Thesis Recombinaison expérimentale intra-et interespèce chez les Rhinovirus et Quantification d'arn de Rhinovirus par RT-PCR en temps réel SCHIBLER, Manuel Abstract Les rhinovirus sont des petits virus
An Integrated Bioinformatics and Computational Biophysics Approach to Enterovirus Surveillance and Research.
 An Integrated Bioinformatics and Computational Biophysics Approach to Enterovirus Surveillance and Research. A thesis submitted in fulfilment of the requirements of the degree of Doctor of Philosophy Jason
An Integrated Bioinformatics and Computational Biophysics Approach to Enterovirus Surveillance and Research. A thesis submitted in fulfilment of the requirements of the degree of Doctor of Philosophy Jason
Supplementary Fig. S1: E. helvum cytochrome b haplotype Bayesian phylogeny with E. dupreanum used as an outgroup. Three main clades are identified:
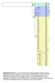 Supplementary Fig. S1: E. helvum cytochrome b haplotype Bayesian phylogeny with E. dupreanum used as an outgroup. Three main clades are identified: A, B and C. Posterior probabilities are shown on the
Supplementary Fig. S1: E. helvum cytochrome b haplotype Bayesian phylogeny with E. dupreanum used as an outgroup. Three main clades are identified: A, B and C. Posterior probabilities are shown on the
Chinese Journal of Biochemistry and Molecular Biology ; (untranslated region,utr) (internal ribosome entry site, IRES). ,.
 ISSN 100727626 CN 1123870ΠQ 2007 7 Chinese Journal of Biochemistry and Molecular Biology 23 (7) :513 518 RNA ( IRES),,,, (, 430072 ;, 430072) 3 RNA 5 (untranslated region,utr) (internal ribosome entry
ISSN 100727626 CN 1123870ΠQ 2007 7 Chinese Journal of Biochemistry and Molecular Biology 23 (7) :513 518 RNA ( IRES),,,, (, 430072 ;, 430072) 3 RNA 5 (untranslated region,utr) (internal ribosome entry
Detection of HEV in Estonia. Anna Ivanova, Irina Golovljova, Valentina Tefanova
 Detection of HEV in Estonia Anna Ivanova, Irina Golovljova, Valentina Tefanova National Institute for Health Development Department of Virology Zoonotic pathogens Tick-borne pathogens: TBEV, Borrelia sp.,
Detection of HEV in Estonia Anna Ivanova, Irina Golovljova, Valentina Tefanova National Institute for Health Development Department of Virology Zoonotic pathogens Tick-borne pathogens: TBEV, Borrelia sp.,
Mapping of the influenza A hemagglutinin serotypes evolution by the ISSCOR method
 Regular paper Supplementary Materials pp 441 452 Vol. 61, 2014 on-line at: www.actabp.pl Mapping of the influenza A hemagglutinin serotypes evolution by the ISSCOR method Jan P. Radomski 1,4 *, Piotr P.
Regular paper Supplementary Materials pp 441 452 Vol. 61, 2014 on-line at: www.actabp.pl Mapping of the influenza A hemagglutinin serotypes evolution by the ISSCOR method Jan P. Radomski 1,4 *, Piotr P.
aV. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Sequences in the 5" Non-coding Region of Human Rhinovims 14 RNA that Affect in vitro Translation
 J. gen. Virol. (1989), 70, 2799-2804. Printed in Great Britain 2799 Key words: rhinovirus, human type 14/5' non-coding region~in vitro translation Sequences in the 5" Non-coding Region of Human Rhinovims
J. gen. Virol. (1989), 70, 2799-2804. Printed in Great Britain 2799 Key words: rhinovirus, human type 14/5' non-coding region~in vitro translation Sequences in the 5" Non-coding Region of Human Rhinovims
... Department of Biological Sciences, John Tabor Laboratories, University of Essex, Colchester CO4 3SQ, UK
 Journal of General Virology (2000), 81, 201 207. Printed in Great Britain... The 2A proteins of three diverse picornaviruses are related to each other and to the H-rev107 family of proteins involved in
Journal of General Virology (2000), 81, 201 207. Printed in Great Britain... The 2A proteins of three diverse picornaviruses are related to each other and to the H-rev107 family of proteins involved in
Characterization of a Novel Circovirus from a Stranded Longman s Beaked Whale (Indopacetus pacificus)
 Characterization of a Novel Circovirus from a Stranded Longman s Beaked Whale (Indopacetus pacificus) Nelmarie Landrau Giovannetti, Kuttichantran Subramaniam, Terry Fei Fan Ng, David S. Rotstein, Kristi
Characterization of a Novel Circovirus from a Stranded Longman s Beaked Whale (Indopacetus pacificus) Nelmarie Landrau Giovannetti, Kuttichantran Subramaniam, Terry Fei Fan Ng, David S. Rotstein, Kristi
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
First Identification and Characterization of Porcine Enterovirus G in the United States
 First Identification and Characterization of Porcine Enterovirus G in the United States Srivishnupriya Anbalagan 1, Richard A. Hesse 2, Ben M. Hause 1 * 1 Newport Laboratories, Inc., Worthington, Minnesota,
First Identification and Characterization of Porcine Enterovirus G in the United States Srivishnupriya Anbalagan 1, Richard A. Hesse 2, Ben M. Hause 1 * 1 Newport Laboratories, Inc., Worthington, Minnesota,
Ch10 Classification and Nomenclature of Viruses 國立台灣海洋大學海洋生物研究所陳歷歷
 Ch10 Classification and Nomenclature of Viruses 國立台灣海洋大學海洋生物研究所陳歷歷 At a glance 10.1 History of virus classification and nomenclature Some of characteristics can be used for the purposes of classification:
Ch10 Classification and Nomenclature of Viruses 國立台灣海洋大學海洋生物研究所陳歷歷 At a glance 10.1 History of virus classification and nomenclature Some of characteristics can be used for the purposes of classification:
Change of Major Genotype of Enterovirus 71 in Outbreaks of Hand-Foot-and-Mouth Disease in Taiwan between 1998 and 2000
 JOURNAL OF CLINICAL MICROBIOLOGY, Jan. 2002, p. 10 15 Vol. 40, No. 1 0095-1137/02/$04.00 0 DOI: 10.1128/JCM.40.1.10 15.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Change
JOURNAL OF CLINICAL MICROBIOLOGY, Jan. 2002, p. 10 15 Vol. 40, No. 1 0095-1137/02/$04.00 0 DOI: 10.1128/JCM.40.1.10 15.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Change
aP (modules 1 and 9 are required)
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
ALIGNMENTS AND COMPARATIVE PROFILESOF PICORNAVIRUS GENERA
 In: Molecular Biology of Picornaviruses. Eds: B. Semler & E. Wimmer, ASM Press, pp 149-155 (2002) ALIGNMENTS AND COMPARATIVE PROFILESOF PICORNAVIRUS GENERA Ann C. Palmenberg* and Jean-Yves Sgro Insti.
In: Molecular Biology of Picornaviruses. Eds: B. Semler & E. Wimmer, ASM Press, pp 149-155 (2002) ALIGNMENTS AND COMPARATIVE PROFILESOF PICORNAVIRUS GENERA Ann C. Palmenberg* and Jean-Yves Sgro Insti.
Master of Science in Microbiology. Rhodes University. Tembisa Innocencia Jauka. May 2010
 Generation of polyclonal antibodies against Theiler s Murine Encephalomyelitis virus protein 2C, and their use in investigating localisation of the protein in infected cells Dissertation submitted in fulfillment
Generation of polyclonal antibodies against Theiler s Murine Encephalomyelitis virus protein 2C, and their use in investigating localisation of the protein in infected cells Dissertation submitted in fulfillment
Envelope e_ :------, Envelope glycoproteins e_ ~ Single-stranded RNA ----, Nucleocapsid
 Virus Antib(tdies Your Expertise, Our Antibodies, Accelerated Discovery. Envelope e_---------:------, Envelope glycoproteins e_-------1~ Single-stranded RNA ----, Nucleocapsid First identified in 1989,
Virus Antib(tdies Your Expertise, Our Antibodies, Accelerated Discovery. Envelope e_---------:------, Envelope glycoproteins e_-------1~ Single-stranded RNA ----, Nucleocapsid First identified in 1989,
on June 19, 2018 by guest
 JOURNAL OF VIROLOGY, Sept. 2001, p. 8021 8030 Vol. 75, No. 17 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.17.8021 8030.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Construction
JOURNAL OF VIROLOGY, Sept. 2001, p. 8021 8030 Vol. 75, No. 17 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.17.8021 8030.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Construction
An automated global clade classification tool for H1 hemagglutinin genes from swine influenza A viruses
 An automated global clade classification tool for H1 hemagglutinin genes from swine influenza A viruses Yun Zhang J. Craig Venter Institute June 25, 2017 Outline A web-based, automated H1 clade classification
An automated global clade classification tool for H1 hemagglutinin genes from swine influenza A viruses Yun Zhang J. Craig Venter Institute June 25, 2017 Outline A web-based, automated H1 clade classification
MIRTRIYKPTYTTTTPPTCHTPIKMEEDPREKMHPQSMWRLVRLRAQRLLSYSESTDLSTREFLEDVSKS I...R...M...D.A...I.G.
 A. FDLV iiiiiiiiiiiiiii1 AnaPV(1110RN043)iiii1 AnaPV(1110RN047)iiii1 AnaPV(1201RN001)I ii1 AnaPV(1201RN003) iii1 AnaPV(1203RN009) iii1 Biti-CA98 iiiiiiiiii1 Dasy-GER00 iiiiiiiii1 Xeno-USA99 iiiiiiiii1
A. FDLV iiiiiiiiiiiiiii1 AnaPV(1110RN043)iiii1 AnaPV(1110RN047)iiii1 AnaPV(1201RN001)I ii1 AnaPV(1201RN003) iii1 AnaPV(1203RN009) iii1 Biti-CA98 iiiiiiiiii1 Dasy-GER00 iiiiiiiii1 Xeno-USA99 iiiiiiiii1
Human parechovirus type 1, 3, 4, 5 and 6 detection in picornavirus cultures ACCEPTED. Running title: HPeV type 1, 3, 4, 5 and 6 in virus cultures
 JCM Accepts, published online ahead of print on 12 December 2007 J. Clin. Microbiol. doi:10.1128/jcm.02009-07 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions. All
JCM Accepts, published online ahead of print on 12 December 2007 J. Clin. Microbiol. doi:10.1128/jcm.02009-07 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions. All
The Bunyaviridae Study Group supports this proposal. RM Elliott, 19/06/2014
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Yuxin Zhao, 1 Song Li, 2 Yufa Zhou, 3,4 Wengang Song, 2 Yujing Tang, 1 Quanhai Pang, 3 and Zengmin Miao Introduction
 BioMed Volume 2015, Article ID 267520, 6 pages http://dx.doi.org/10.1155/2015/267520 Research Article Phylogenetic Analysis of Hemagglutinin Genes of H9N2 Avian Influenza Viruses Isolated from Chickens
BioMed Volume 2015, Article ID 267520, 6 pages http://dx.doi.org/10.1155/2015/267520 Research Article Phylogenetic Analysis of Hemagglutinin Genes of H9N2 Avian Influenza Viruses Isolated from Chickens
Investigation of the subcellular localization of HPeV 3A and 2A proteins.
 Biomedical Research 2017; 28 (20): 8639-8648 ISSN 0970-938X www.biomedres.info Investigation of the subcellular localization of HPeV 3A and 2A proteins. Arsalan Salimi *, Siamak Amirfakhri School of Biological
Biomedical Research 2017; 28 (20): 8639-8648 ISSN 0970-938X www.biomedres.info Investigation of the subcellular localization of HPeV 3A and 2A proteins. Arsalan Salimi *, Siamak Amirfakhri School of Biological
Diversity of picornaviruses in rural Bolivia
 Journal of General Virology (2013), 94, 2017 2028 DOI 10.10/vir.3827-0 Diversity of picornaviruses in rural Bolivia W. Allan Nix, 1 Nino Khetsuriani, 1 Silvia Peñaranda, 1 Kaija Maher, 1 Linda Venczel,
Journal of General Virology (2013), 94, 2017 2028 DOI 10.10/vir.3827-0 Diversity of picornaviruses in rural Bolivia W. Allan Nix, 1 Nino Khetsuriani, 1 Silvia Peñaranda, 1 Kaija Maher, 1 Linda Venczel,
Report concerning identification of the Northern European isolate of BTV (26 th August 2006)
 Report concerning identification of the Northern European isolate of BTV (26 th August 2006) Peter Mertens, Carrie Batten, Simon Anthony, Lydia Kgosana, Karin Darpel, Kasia Bankowska, Eva Veronesi, Abid
Report concerning identification of the Northern European isolate of BTV (26 th August 2006) Peter Mertens, Carrie Batten, Simon Anthony, Lydia Kgosana, Karin Darpel, Kasia Bankowska, Eva Veronesi, Abid
Viral Taxonomic Classification
 Viruses Part I Viral Taxonomic Classification Order>> -virales Family>> - viridae Subfamily>> -virinae Genus>> -virus Species Order>> Picornavirales Family>> Picornaviridae Subfamily>> Picornavirinae Genus>>
Viruses Part I Viral Taxonomic Classification Order>> -virales Family>> - viridae Subfamily>> -virinae Genus>> -virus Species Order>> Picornavirales Family>> Picornaviridae Subfamily>> Picornavirinae Genus>>
Epidemiology and dynamic of hand, foot and mouth disease in Vietnam
 Epidemiology and dynamic of hand, foot and mouth disease in Vietnam Nghia Ngu Duy To cite this version: Nghia Ngu Duy. Epidemiology and dynamic of hand, foot and mouth disease in Vietnam. Infectious diseases.
Epidemiology and dynamic of hand, foot and mouth disease in Vietnam Nghia Ngu Duy To cite this version: Nghia Ngu Duy. Epidemiology and dynamic of hand, foot and mouth disease in Vietnam. Infectious diseases.
Highly diverse population of Picornaviridae and other members of the Picornavirales, in Cameroonian fruit bats
 Yinda et al. BMC Genomics (2017) 18:249 DOI 10.1186/s12864-017-3632-7 RESEARCH ARTICLE Open Access Highly diverse population of Picornaviridae and other members of the Picornavirales, in Cameroonian fruit
Yinda et al. BMC Genomics (2017) 18:249 DOI 10.1186/s12864-017-3632-7 RESEARCH ARTICLE Open Access Highly diverse population of Picornaviridae and other members of the Picornavirales, in Cameroonian fruit
Foot-and Mouth Disease Ecological Studies In Endemic Settings: Ongoing Studies in Vietnam and Pakistan
 Foot-and Mouth Disease Ecological Studies In Endemic Settings: Ongoing Studies in Vietnam and Pakistan Luis Rodriguez (Jonathan Arzt) Research leader, USDA-ARS Plum Island 1 PAKISTAN CHARACTERIZATION OF
Foot-and Mouth Disease Ecological Studies In Endemic Settings: Ongoing Studies in Vietnam and Pakistan Luis Rodriguez (Jonathan Arzt) Research leader, USDA-ARS Plum Island 1 PAKISTAN CHARACTERIZATION OF
Framework for the evaluation of. and scientific impacts of plant viruses. technologies.
 Framework for the evaluation of biosecurity, commercial, regulatory and scientific impacts of plant viruses and viroids identified by NGS technologies. Sebastien Massart, Thierry Candresse, José Gil, Christophe
Framework for the evaluation of biosecurity, commercial, regulatory and scientific impacts of plant viruses and viroids identified by NGS technologies. Sebastien Massart, Thierry Candresse, José Gil, Christophe
Genome Sequencing and Analysis of a Porcine Delta Coronavirus from Eastern China
 ResearchArticle imedpub Journals http://www.imedpub.com/ DOI: 10.21767/2248-9215.100025 European Journal of Experimental Biology Genome Sequencing and Analysis of a Porcine Delta Coronavirus from Eastern
ResearchArticle imedpub Journals http://www.imedpub.com/ DOI: 10.21767/2248-9215.100025 European Journal of Experimental Biology Genome Sequencing and Analysis of a Porcine Delta Coronavirus from Eastern
The use of nonhuman primates in biomedical research has led to the isolation of many
 JVI Accepts, published online ahead of print on 29 September 2010 J. Virol. doi:10.1128/jvi.01928-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
JVI Accepts, published online ahead of print on 29 September 2010 J. Virol. doi:10.1128/jvi.01928-10 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
CONSTRUCTION OF PHYLOGENETIC TREE USING NEIGHBOR JOINING ALGORITHMS TO IDENTIFY THE HOST AND THE SPREADING OF SARS EPIDEMIC
 CONSTRUCTION OF PHYLOGENETIC TREE USING NEIGHBOR JOINING ALGORITHMS TO IDENTIFY THE HOST AND THE SPREADING OF SARS EPIDEMIC 1 MOHAMMAD ISA IRAWAN, 2 SITI AMIROCH 1 Institut Teknologi Sepuluh Nopember (ITS)
CONSTRUCTION OF PHYLOGENETIC TREE USING NEIGHBOR JOINING ALGORITHMS TO IDENTIFY THE HOST AND THE SPREADING OF SARS EPIDEMIC 1 MOHAMMAD ISA IRAWAN, 2 SITI AMIROCH 1 Institut Teknologi Sepuluh Nopember (ITS)
Rotavirus Genotyping and Enhanced Annotation in the Virus Pathogen Resource (ViPR) Yun Zhang J. Craig Venter Institute ASV 2016 June 19, 2016
 Rotavirus Genotyping and Enhanced Annotation in the Virus Pathogen Resource (ViPR) Yun Zhang J. Craig Venter Institute ASV 2016 June 19, 2016 Loading Virus Pathogen Database and Analysis About Resource
Rotavirus Genotyping and Enhanced Annotation in the Virus Pathogen Resource (ViPR) Yun Zhang J. Craig Venter Institute ASV 2016 June 19, 2016 Loading Virus Pathogen Database and Analysis About Resource
A mobile genetic element with unknown function found in distantly related viruses
 Tengs et al. Virology Journal 2013, 10:132 RESERH Open ccess mobile genetic element with unknown function found in distantly related viruses Torstein Tengs 1*, nja Bråthen Kristoffersen 1, Tsvetan R Bachvaroff
Tengs et al. Virology Journal 2013, 10:132 RESERH Open ccess mobile genetic element with unknown function found in distantly related viruses Torstein Tengs 1*, nja Bråthen Kristoffersen 1, Tsvetan R Bachvaroff
Characterization of a canine homolog of human Aichivirus.
 JVI Accepts, published online ahead of print on 31 August 2011 J. Virol. doi:10.1128/jvi.05317-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JVI Accepts, published online ahead of print on 31 August 2011 J. Virol. doi:10.1128/jvi.05317-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
Supplementary Online Content
 Supplementary Online Content Rodger AJ, Cambiano V, Bruun T, et al; PARTNER Study Group. Sexual activity without condoms and risk of HIV transmission in serodifferent couples when the HIVpositive partner
Supplementary Online Content Rodger AJ, Cambiano V, Bruun T, et al; PARTNER Study Group. Sexual activity without condoms and risk of HIV transmission in serodifferent couples when the HIVpositive partner
Evolution of DS-1-like human G2P[4] rotaviruses assessed by complete genome analyses
![Evolution of DS-1-like human G2P[4] rotaviruses assessed by complete genome analyses Evolution of DS-1-like human G2P[4] rotaviruses assessed by complete genome analyses](/thumbs/74/70215795.jpg) Journal of General Virology (2014), 95, 91 109 DO 10.10/vir.0.056788-0 Evolution of DS-1-like human G2P[4] rotaviruses assessed by complete genome analyses Giovanni M. Giammanco, 1 Floriana Bonura, 1 Mark
Journal of General Virology (2014), 95, 91 109 DO 10.10/vir.0.056788-0 Evolution of DS-1-like human G2P[4] rotaviruses assessed by complete genome analyses Giovanni M. Giammanco, 1 Floriana Bonura, 1 Mark
Review article. Structure, function and evolution of picornaviruses. Glyn Stanway. Introduction. Structure of picornaviruses
 Journal of General Virology (1990), 71, 2483-2501. Printed in Great Britain 2483 Review article Structure, function and evolution of picornaviruses Glyn Stanway Department of Biology, University of Essex,
Journal of General Virology (1990), 71, 2483-2501. Printed in Great Britain 2483 Review article Structure, function and evolution of picornaviruses Glyn Stanway Department of Biology, University of Essex,
Continued seasonal circulation of enterovirus D68 in the Netherlands,
 Rapid communications Continued seasonal circulation of enterovirus D68 in the Netherlands, 20 20 A Meijer (Adam.Meijer@rivm.nl), K S Benschop, G A Donker 2, H G van der Avoort. Centre for Infectious Disease
Rapid communications Continued seasonal circulation of enterovirus D68 in the Netherlands, 20 20 A Meijer (Adam.Meijer@rivm.nl), K S Benschop, G A Donker 2, H G van der Avoort. Centre for Infectious Disease
Summary of Current Respiratory Season and Genetic Analysis of Influenza Virus Circulating in Alberta
 Laboratory Bulletin Date: February 28, 2013 To: From: Re: Alberta Health, Alberta MicroNet, Communicable Disease Nurses, Infectious Diseases Physicians, Infection Prevention and Control, Medical Officers
Laboratory Bulletin Date: February 28, 2013 To: From: Re: Alberta Health, Alberta MicroNet, Communicable Disease Nurses, Infectious Diseases Physicians, Infection Prevention and Control, Medical Officers
Received 19 May 2009/Returned for modification 7 August 2009/Accepted 2 October 2009
 JOURNAL OF CLINICAL MICROBIOLOGY, Dec. 2009, p. 3958 37 Vol. 47, No. 12 0095-1137/09/$12.00 doi:10.1128/jcm.003-09 Copyright 2009, American Society for Microbiology. All Rights Reserved. Screening Respiratory
JOURNAL OF CLINICAL MICROBIOLOGY, Dec. 2009, p. 3958 37 Vol. 47, No. 12 0095-1137/09/$12.00 doi:10.1128/jcm.003-09 Copyright 2009, American Society for Microbiology. All Rights Reserved. Screening Respiratory
Translation. Host Cell Shutoff 1) Initiation of eukaryotic translation involves many initiation factors
 Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
The Fecal Viral Flora of Wild Rodents
 The Fecal Viral Flora of Wild Rodents Tung G. Phan 1,2, Beatrix Kapusinszky 1,2,3, Chunlin Wang 4, Robert K. Rose 5, Howard L. Lipton 6, Eric L. Delwart 1,2 * 1 Blood Systems Research Institute, San Francisco,
The Fecal Viral Flora of Wild Rodents Tung G. Phan 1,2, Beatrix Kapusinszky 1,2,3, Chunlin Wang 4, Robert K. Rose 5, Howard L. Lipton 6, Eric L. Delwart 1,2 * 1 Blood Systems Research Institute, San Francisco,
HAV HBV HCV HDV HEV HGV
 Viral Hepatitis HAV HBV HCV HDV HEV HGV Additional well-characterized viruses that can cause sporadic hepatitis, such as yellow fever virus, cytomegalovirus, Epstein-Barr virus, herpes simplex virus, rubella
Viral Hepatitis HAV HBV HCV HDV HEV HGV Additional well-characterized viruses that can cause sporadic hepatitis, such as yellow fever virus, cytomegalovirus, Epstein-Barr virus, herpes simplex virus, rubella
Molecular Evolution of the Human Enteroviruses: Correlation of Serotype with VP1 Sequence and Application to Picornavirus Classification
 JOURNAL OF VIROLOGY, Mar. 1999, p. 1941 1948 Vol. 73, No. 3 0022-538X/99/$04.00 0 Molecular Evolution of the Human Enteroviruses: Correlation of Serotype with VP1 Sequence and Application to Picornavirus
JOURNAL OF VIROLOGY, Mar. 1999, p. 1941 1948 Vol. 73, No. 3 0022-538X/99/$04.00 0 Molecular Evolution of the Human Enteroviruses: Correlation of Serotype with VP1 Sequence and Application to Picornavirus
Foot and Mouth Disease Control, A vaccine perspective. John Barlow, DVM PhD
 Foot and Mouth Disease Control, A vaccine perspective John Barlow, DVM PhD john.barlow@uvm.edu May 28, 2015 Countries in which FMD was reported to the OIE between 1990 and 2002 Antigenic variation is common
Foot and Mouth Disease Control, A vaccine perspective John Barlow, DVM PhD john.barlow@uvm.edu May 28, 2015 Countries in which FMD was reported to the OIE between 1990 and 2002 Antigenic variation is common
