Association of the calpain-10 gene with type 2 diabetes in Europeans: Results of pooled and meta-analyses
|
|
|
- Merry Burke
- 6 years ago
- Views:
Transcription
1 Molecular Genetics and Metabolism 89 (2006) Association of the calpain-10 gene with type 2 diabetes in Europeans: Results of pooled and meta-analyses Takafumi Tsuchiya a, Peter E.H. Schwarz b, Laura del Bosque-Plata a, M. GeoVrey Hayes a, Christian Dina c, Philippe Froguel c, G. Wayne Towers b,d, Sabine Fischer b, Theodora Temelkova-Kurktschiev e, Hannes Rietzsch b, Juergen Graessler b, Josef Vcelák f, Daniela Palyzová g, Thomas Selisko b, Bela Bendlová f, Jan Schulze b, Ulrich Julius b, Markolf Hanefeld e, Michael N. Weedon h, Julie C. Evans h, Timothy M. Frayling h, Andrew T. Hattersley h, Marju Orho-Melander i, Leif Groop i, Maciej T. Malecki j, Torben Hansen k, Oluf Pedersen k, Tasha E. Fingerlin l, Michael Boehnke l, Craig L. Hanis m, Nancy J. Cox a, Graeme I. Bell a, a Departments of Medicine and Human Genetics, The University of Chicago, 5841 S. Maryland Ave., MC1027, Chicago, IL 60637, USA b Department of Endocrinopathies and Metabolic Diseases, Medical Faculty Carl-Gustav-Carus of the Technical University Dresden, Dresden, Germany c CNRS UMR 8090, Genomics and Molecular Physiology of Metabolic Diseases, Pasteur Institute, Lille, France d Centre for Genome Research, North-West University (Potchefstroom Campus), Pretoria, South Africa e Center for Clinical Studies, GWT-TU Dresden, Dresden, Germany f Institute of Endocrinology, Prague, Czech Republic g Clinic of Children and Adolescents, and 3rd Faculty of Medicine, Charles University, Prague, Czech Republic h Institute of Biomedical and Clinical Science, Peninsula Medical School, Exeter, UK i Department of Clinical Sciences, Diabetes and Endocrinology, Lund University, CRC, University Hospital, Malmö, Sweden j Department of Metabolic Diseases, Medical College, Jagiellonian University, Krakow, Poland k Steno Diabetes Center and Hagedorn Research Institute, Gentofte, Denmark l Department of Biostatistics, School of Public Health, University of Michigan, Ann Arbor, MI, USA m Human Genetics Center, The University of Texas Health Science Center at Houston, Houston, TX, USA Received 24 March 2006; received in revised form 24 May 2006; accepted 24 May 2006 Available online 11 July 2006 Abstract We conducted pooled and meta-analyses of the association of the calpain-10 gene (CAPN10) polymorphisms SNP-43, Indel-19 and SNP- 63 individually and as haplotypes with type 2 diabetes (T2D) in 3237 patients and 2935 controls of European ancestry. In the pooled analyses, the common SNP-43 G allele was associated with modest but statistically signiwcant increased risk of T2D (odds ratio (OR) D 1.11 (95% con- Wdence interval (CI), ), P D 0.01). Two haplotype combinations were associated with increased risk of T2D (1-2-1/1-2-1, OR D 1.20 ( ), P D 0.02; and 1-1-2/1-2-1, OR D 1.26 ( ), P D 0.04) and one with decreased risk (1-1-1/2-2-1, OR D 0.86 ( ), P D 0.03). The meta-analysis also showed a signiwcant evect of the 1-2-1/1-2-1 haplogenotype on risk (OR D 1.25 ( ), P D 0.01). However, there was evidence for heterogeneity with respect to this evect (P D 0.06). The heterogeneity appeared to be due to data sets in which the cases were selected from samples used in linkage studies of T2D. Using only the population-based case-control samples removed the heterogeneity (P D 0.89) and strengthened the evidence for association with T2D in both the pooled (SNP-43 G, OR D 1.19 ( ), P D 0.001; 1-2-1/1-2-1 haplogenotype, OR D 1.46 ( ), P D ; 1-1-2/1-2-1 haplogenotype, OR D 1.52 ( ), P D 0.007; and 1-1-1/2-2-1 haplogenotype, OR D 0.83 ( ), P D 0.03) and the meta-analysis (SNP-43 G, OR D 1.18 ( ), P D 0.005; 1-2-1/1-2-1 haplogenotype, OR D 1.68 ( ), P D ). The pooled and meta-analyses as well as the linkage disequilibrium and haplotype diversity studies suggest a role for genetic variation in CAPN10 avecting risk of T2D in Europeans. * Corresponding author. Fax: address: g-bell@uchicago.edu (G.I. Bell) /$ - see front matter 2006 Elsevier Inc. All rights reserved. doi: /j.ymgme
2 2006 Elsevier Inc. All rights reserved. Keywords: Calpain-10; Association study; Diabetes mellitus T. Tsuchiya et al. / Molecular Genetics and Metabolism 89 (2006) Introduction Our genes determine, at least in part, our risk of developing type 2 diabetes (T2D) [1]. It is commonly believed that the identiwcation of the genes and associated variants that modify risk (increase or decrease) will lead to rational approaches for preventing and treating diabetes that take into account the individual s genetic risk prowle. We have been studying the genetics of T2D in the Mexican American population of Starr County, Texas [2 4]. A genomewide search for T2D susceptibility loci mapped a gene(s) with a major evect on risk, designated NIDDM1, to the distal long arm of chromosome 2 (band 2q37.3) [2]. This locus was estimated to have a sibling recurrence risk (λ s ) of 1.37 (1-lod conwdence interval for λ s, ) for an additive and 1.36 ( ) for a non-additive model. NIDDM1 could account for 21 30% of the familial clustering of T2D in the population studied, depending on the model (additive or multiplicative) for the interaction between NIDDM1 and other susceptibility loci. We positionally cloned a gene in the NIDDM1 region that showed both linkage and association with T2D [4]. This gene, calpain-10 (CAPN10), 1 encoded a nonlysosomal cysteine protease that was expressed in all tissues examined, including tissues such as pancreatic islets, liver, skeletal muscle and fat that play an important role in the regulation of glucose homeostasis [4,5]. We showed in this Wrst report that risk could be most readily dewned in terms of speciwc haplotypes of three polymorphisms: SNP-43, Indel-19 and SNP-63. The association of these SNPs and their haplotypes with T2D has not been consistently replicated [6 11]. The inconsistency has been a troubling feature of genetic studies of CAPN10 and T2D and has raised doubts that genetic variation in CAPN10 is indeed associated with T2D. However, two recent metaand pooled-analyses in large groups of cases and controls (>3500 each) have shown association of two SNPs in calpain-10 (SNP-43 and -44) with T2D [12,13]. Thus, the data look more convincing as larger sample sizes have been assembled, although as noted by McCarthy et al. [14] the combined evidence is not, as yet, incontrovertible. This is especially true of the genetic model in which Mexican Americans with the haplogenotype (SNP-43, Indel-19, SNP-63) designated 1-1-2/1-2-1 had the highest risk and those homozygous for either the or haplotype 1 Abbreviations used: CAPN10, symbol for the human calpain-10 gene; Indel, insertion/deletion polymorphism; CI, conwdence interval; Ctrl, controls; Het, heterogeneity; HWE, Hardy Weinberg equilibrium; LD, linkage disequilibrium; MAF, minor allele frequency; NGT, normal glucose tolerance; OR, odds ratio; SNP, single nucleotide polymorphism; T2D, type 2 diabetes. were not at increased risk. However, Mexican Americans are an admixed population (the ancestral contributions to the contemporary gene pool of the Mexican American population of Starr County population are 61% Spanish, 31% Native American and 8% African [15]) and this may be a confounding factor in both the original and in subsequent studies. Thus, small samples sizes and the unique nature of the Mexican American population may be responsible for the failure to obtain consistent replication. In order to gain a better understanding of the contribution of genetic variation in CAPN10 to risk of T2D, we have begun to carry out large association studies in diverse populations with the view that diverent variants and haplotypes may avect risk in diverent populations. Here, we report the association testing of SNP-43, Indel-19 and SNP-63, individually and by haplotype with T2D in individuals of European ancestry supplemented by an examination of linkage disequilibrium and haplotype diversity of CAPN10 in this population. Materials and methods Subjects We considered all published case-control studies that examined the association of the CAPN10 polymorphisms SNP-43, Indel-19 (or SNP-56 which is in perfect linkage disequilibrium (LD) with Indel-19) and SNP-63 with T2D in populations of European (Caucasian) ancestry [4,6 11,16]. We did not include SNP-44 which has been independently associated with T2D [9] as it was not typed in four of the datasets representing 53.8% of cases and 33.3% of controls: Denmark, Finland (FUSION), Poland and France (Table 1). We excluded the study of Elbein et al. [8] which was a family-based study of approximately 700 members of 63 families in which CAPN10 variants were tested for linkage and association (transmission disequilibrium test) with T2D rather than a case-control study. We also excluded the study of Berger et al. [16] because the cases were ascertained for both T2D and endstage kidney disease. We contacted the primary investigator of each study and obtained the raw genotype data so that we could carry out both pooled and meta-analyses. We also genotyped SNP-43, Indel-19 and SNP-63 in an additional 2,514 subjects from Wve case-control studies (one Czech, one French and three German). There was overlap between the Finnish and German subjects reported in Horikawa et al. [4] and the Botnia-1 and Germany-1 cohorts, respectively, and we therefore used only the larger data sets (i.e., Botnia-1 and Germany-1) which removed all overlap. The clinical characteristics of the study populations are summarized in Table 1. The genotype and haplotype distributions for all the study populations are provided in Tables 1 and 2 in the online supplementary material. Statistical analyses We tested each polymorphism (in each data set and in the pooled sample) for deviation from HWE. We then tested for diverences in allele, genotype, haplotype and haplogenotype frequencies between groups (within study and pooled) using a chi-squared test. Previous studies have shown that there are only four common SNP-43 Indel-19 SNP-63 haplotypes observed in Mexican Americans, Japanese and Europeans rather than the eight expected permitting unambiguous haplotype assignment. The four observed haplotypes are described as 1-1-1, 1-1-2, and [4]. We
3 176 T. Tsuchiya et al. / Molecular Genetics and Metabolism 89 (2006) removed subjects with missing genotypes (86 cases and 49 controls) or the rare haplotypes 1-2-2, 2-2-2, and (an indication of possible genotyping error) (19 cases and 14 controls) from the haplotype-based analyses. We used the odds ratio as a measure of risk. In the meta-analysis, summary odds ratios and 95% conwdence intervals were estimated under both Wxed evects (Mantel Haenszel) and random evects models (DerSimonian Laird). We tested for heterogeneity across comparisons using the chi-squared-based Q statistic and heterogeneity was considered signiwcant for P < Publication bias was assessed using Begg s adjusted rank correlation test and Egger s regression asymmetry test using the STATA software. All P-values are two-sided. Linkage disequilibrium and haplotype blocks To place the pooled and meta-analyses of SNP-43, Indel-19 and SNP- 63 in the context of the overall pattern of linkage disequilibrium (LD) in the CAPN10 region, we typed 30 polymorphisms (SNPs and Indels) in this region in a group of 72 unrelated healthy individuals of European ancestry residing in San Francisco, CA using Taqman-based assays (for SNPs) and agarose gel electrophoresis (for Indels). The genotype distributions of all polymorphisms were in Hardy Weinberg equilibrium (HWE), P >0.05. We calculated pairwise measures of LD, visualized the patterns of LD using Haploview [ [17], and dewned haplotype blocks using the conwdence interval algorithm [18] and the four-gamete rule [19]. The conwdence interval dewnes haplotype blocks as follows. For each marker pair, if the one-sided 95% conwdence interval for D had an upper bound >0.98 and lower bound >0.7, the marker pairs were deemed to be in strong LD. If the upper bound was <0.9, the marker pair was considered to have undergone historic recombination. All other marker pairs were labeled inconclusive. A block was created if 95% of informative (i.e., non-inconclusive) comparisons of marker pairs with minor allele frequencies >0.05 were considered to be in strong LD. The four-gamete rule dewnes haplotype blocks by the absence of all four two-marker haplotypes for each marker pair occurring at frequency >0.01. We used Tagger [20] in Haploview [17] to determine the proportion of markers in the block that could be captured (having r 2 greater than the particular threshold set) by forcing SNP-44, SNP-43, Indel-19, and SNP- 63 as tag SNPs. LD patterns were also compared with those observed in the Europeans in HapMap [ [21]. We determined frequencies of haplotypes spanning the 15 marker block surrounding CAPN10 in the 63 individuals missing less than 20% of their genotype calls using PHASE [ stephens/software.html] [22,23] with the default number of iterations, burn-in, and thinning interval. Consistency of phase assignment was assessed through Wve runs from diverent random seeds. Results Association of SNPs in CAPN10 with type 2 diabetes We performed both pooled and meta-analyses of 12 data sets including 3237 cases and 2935 controls (Table 1). We carried out both types of analysis because pooling increases the power to detect and quantify an evect and the metaanalysis provides a control for population diverences that could lead to spurious association if there are diverences in allele frequency among groups. The power of detecting an evect in a sample of this size in the overall pooled analyses for SNP-43, Indel-19 and SNP-63 was 0.71, 0.06 and 0.69, respectively; and 0.22, 0.61, 0.68 and 0.65 for the 1-1-1, , and haplotypes described below [24]. Power Table 1 Clinical characteristics of European study populations Reference Population Selection/characteristics of cases (mean age-at-diagnosis) and controls (mean age) Eligible subjects T2D Ctrl T2D Ctrl Horikawa et al. [4] Germany (Germany-1) T2D, Probands of sibs, ( Random, ( Finland (Botnia-1) T2D, age-at-diagnosis 740 years, NGT per WHO criteria, ( ( Evans et al. [6] UK (UK-1) T2D, Probands of trios, ( Population, birth cohort ( UK (UK-2) T2D, Probands of sibs, age-atdiagnosis years, ( Orho-Melander et al. [11] Finland (Botnia-2) T2D, ( NGT per WHO criteria, ( Rasmussen et al. [7] Denmark T2D, ( NGT per WHO criteria, ( Fingerlin et al. [9] Finland (FUSION) T2D, Probands of sibs from the Wrst NGT per WHO criteria, UnaVected and second sets, ( spouses, ( Finland (FUSION) NGT per WHO criteria, Elderly 223 controls, ( Malecki et al. [10] Poland T2D, ( Non-diabetic, ( Unpublished Czech Republic T2D, ( NGT per WHO criteria ( Unpublished France T2D, ( Normoglycemic, ( Unpublished France T2D, Probands of sibs, ( Unpublished Germany (Germany-2) T2D, ( NGT per WHO criteria, ( Unpublished Germany (Germany-3) T2D, ( NGT per WHO criteria, ( SNP-56 was typed instead of Indel-19 in the FUSION samples. Ctrl, controls; T2D, type 2 diabetes; NGT, normal glucose tolerance; WHO, World Health Organization. For the meta-analysis, we combined the FUSION ctrl groups and the French case (in the overall comparison) groups.
4 Table 2 CAPN10 polymorphisms and risk of type 2 diabetes in Europeans stratiwed based on ascertainment of cases (population-based vs. linkage-based) Marker Genotype/allele Overall Population-based cases n or frequency Pooled Meta-analysis n or frequency Pooled Meta-analysis Fixed evects Random evects Het Fixed evects Random evects Het Ctrl T2D OR (95% CI) P OR (95% CI) P OR (95% CI) P P Ctrl T2D OR (95% CI) P OR (95% CI) P OR (95% CI) P P SNP-43 G/G G/A A/A G ( ) ( ) ( ) ( ) ( ) ( ) A ( ) ( ) ( ) ( ) ( ) ( ) Indel-19 2R/2R R/3R R/3R R R ( ) ( ) ( ) ( ) ( ) ( ) SNP-63 C/C C/T T/T C T ( ) ( ) ( ) ( ) ( ) ( ) The UK-1 patient group has been excluded as described in the text. The number of studies for the meta-analysis are as follows: 11 in the overall and 8 in the population-based cases. In the pooled data, all genotypic distributions were in HWE. P-values were calculated by genotype or allele. Heterogeneity was considered to be signiwcant if P < 0.10 by the Q statistic. Ctrl, controls; T2D, type 2 diabetes; OR, odds ratio; CI, conwdence interval; Het, heterogeneity. P-value for comparison of genotype distribution between cases and controls. T. Tsuchiya et al. / Molecular Genetics and Metabolism 89 (2006)
5 178 T. Tsuchiya et al. / Molecular Genetics and Metabolism 89 (2006) calculations for meta-analyses are controversial and were not carried out [25,26]. We Wrst examined each polymorphism individually including testing each for deviation from HWE as well as association with T2D. The UK-1 cases (n D 153) displayed a signiwcant departure from HWE at Indel-19 (P D 0.02) which led the entire pooled sample to show a signiwcant departure from HWE (P D 0.04) and we, thus, excluded this sample (ascertained as a trio sample with living parents) from all further analyses. All of the other case and control groups were in HWE for each variant studied here. In the pooled European sample (excluding the UK-1 cases), the common SNP-43 G allele was associated with increased risk of T2D (OR D 1.11 ( ), P D 0.01) in the overall data set (Table 2; the OR and P-value were the same if the UK-1 cases were included). The association of the SNP- 43 G allele with T2D also approached signiwcance by the meta-analysis (both Wxed and random evects models: OR D 1.08 ( ), P D 0.09) without evidence for heterogeneity (P D 0.43). The less common SNP-63 T allele (allele 2 in the haplotype) was associated with increased risk of T2D: (OR D 1.14 ( ), P D 0.05) in the pooled analysis of the overall data set (Table 2). Association of CAPN10 haplotypes with type 2 diabetes The analysis of the pooled data sets showed that three SNP-43 Indel-19 SNP-63 haplogenotypes were associated with T2D: 1-1-1/2-2-1 (OR D 0.86 ( ), P D 0.03); 1-1-2/1-2-1 (OR D 1.26 ( ), P D 0.04); and 1-2-1/1-2-1 (OR D 1.20 ( ), P D 0.02) (Table 3). The association of the 1-2-1/1-2-1 haplogenotype with T2D was also observed by meta-analysis (Wxed evects OR D 1.25 ( ), P D 0.01; random evects OR D 1.28 ( ), P D 0.02) but with evidence for heterogeneity (P D 0.06) (Table 3; the evect of inclusion of the UK-1 cases on the haplogenotype analysis are described in the legend to Table 3). There was no evidence for publication bias with the 1-2-1/1-2-1 haplogenotype by the standard funnel plot (Begg s adjusted rank correlation test, P D 0.94; Egger s regression asymmetry test, P D 0.27). However, the power of statistical tests of publication bias are low when there are relatively few studies as in the case of CAPN10 and T2D. In this case, a nonsigniwcant result indicates no evidence for substantial bias. We considered two sources that could be responsible for the observed heterogeneity with respect to the 1-2-1/1-2-1 haplogenotype amongst the data sets: (1) diverences in ageat-diagnosis of T2D in diverent study populations; and (2) the method of ascertainment of cases; i.e., population-based vs. linkage-based. To examine the possible confounding evect of age-at-diagnosis, we split the T2D group based on median age-at-diagnosis (52 in the pooled sample and repeated the comparison but using only those cases for whom age-at-diagnosis was available. Splitting on the basis of median age did not resolve the heterogeneity (P D 0.09 and 0.11 in younger and elder subsets, respectively). To examine the evect of ascertainment, we examined only studies using cases selected for case-control studies and which we call population-based cases for the purposes of this article although they may not represent a true random sample of cases or controls; i.e., we excluded studies in which the cases were selected from samples used for linkage mapping studies (linkage-based cases): French (probands of sibs), FUSION, UK-2 and Germany-1. Comparing population-based cases (n D 1627) and controls (n D 2027), removed the heterogeneity with respect to the 1-2-1/1-2-1 haplogenotype (P D 0.89) (Table 3 and Fig. 2). Moreover, the strength of the association increased for both the pooled and meta-analyses: OR D 1.46 ( ), P D and OR D 1.68 ( ), P D , respectively. By contrast, comparing linkage-based cases (n D 1632) and controls (n D 1128) did not resolve the heterogeneity with respect to the 1-2-1/1-2-1 haplogenotype (P D 0.07). There was, moreover, no signiwcant evect of the 1-2-1/1-2-1 haplogenotype on T2D risk in either the pooled (OR D 0.97 ( ), P D 0.80) or meta-analyses (Wxed evects OR D 0.98 ( ), P D 0.85; random evects OR D 0.98 ( ), P D 0.92). We repeated all the analyses using just the populationbased cases. The SNP-43 G allele and the and haplotypes showed signiwcant association with T2D in both the pooled and meta-analyses (Tables 2 and 4, respectively) and the rare haplotype reached signiwcance in only the pooled analysis. The haplogenotypes 1-1-2/1-2-1 and 1-2-1/1-2-1 were associated with signiwcantly increased risk and 1-1-1/2-2-1 with reduced risk in the pooled analysis (Table 3). Although the 1-1-2/1-1-2 haplogenotype (Table 3) was also associated with increased risk, it is suyciently rare that the risk is not signiwcant in sample of this size. Although diverent inclusion criteria among the controls may also contribute to the heterogeneity, the results suggest that population-based cases may have a diverent genetic background from linkage-based cases [27]. In this regard, combining the population-based and linkage-based cases contributed a large amount to the observed heterogeneity (62% of the Q statistic). Haplotype structure across the calpain-10 gene region in Europeans To construct regional LD context in which to place SNP-43, Indel-19 and SNP-63, we typed 30 polymorphic markers with a mean intermarker distance of 8.8 kb (range D kb) across the CAPN10 region in a panel of 72 unrelated healthy Americans of European ancestry from San Francisco, California. These markers, selected prior to knowledge of the patterns of variation in the CEPH Europeans in the HapMap, were chosen based on informativeness (minor allele frequency (MAF) and diverent LD patterns) in our original studies in Mexican Americans [4] and physical coverage of the region. Analysis of the patterns of LD revealed two regions of
6 Table 3 CAPN10 haplogenotype and risk of type 2 diabetes in Europeans stratiwed based on ascertainment of cases (population-based vs. linkage-based) Haplogenotype Overall Population-based cases n Pooled Meta-analysis n Pooled Meta-analysis Fixed evects Random evects Het Fixed evects Random evects Het Ctrl T2D OR (95% CI) P OR (95% CI) P OR (95% CI) P P Ctrl T2D OR (95% CI) P OR (95% CI) P OR (95% CI) P P 1-1-1/ ( ) ( ) ( ) ( ) ( ) ( ) / ( ) ( ) ( ) ( ) ( ) ( ) / ( ) ( ) ( ) ( ) ( ) ( ) / ( ) ( ) ( ) ( ) ( ) ( ) / ( ) ( ) ( ) ( ) ( ) ( ) / ( ) ( ) ( ) ( ) ( ) ( ) / ( ) ( ) ( ) ( ) ( ) ( ) / ( ) ( ) ( ) ( ) ( ) ( ) / ( ) ( ) ( ) ( ) ( ) ( ) / ( ) ( ) ( ) ( ) ( ) ( ) The haplotypes are those dewned by SNP-43, Indel-19 and SNP-63 and the speciwc alleles are: SNP-43, allele 1, G and allele 2, A; Indel-19, allele 1, 2 repeats of 32-bp sequence, and allele 2, 3 repeats; and SNP-63, allele 1, C, and allele 2, T. The number of studies is 11 for the meta-analysis in the overall group except 1-1-2/1-1-2 (n D 6) and 1-1-2/1-2-1 (n D 10) haplogenotypes. The number of studies is 8 for the meta-analysis of the studies using only population-based cases except 1-1-2/1-1-2 (n D 4) and 1-1-2/1-2-1 (n D 7) haplogenotypes. Heterogeneity was considered to be signiwcant if P < 0.10 by the Q statistic. Ctrl, controls; T2D, type 2 diabetes; OR, odds ratio; CI, conwdence interval; Het, heterogeneity. Note that the results were similar in the overall analysis if the UK-1 cases were included except that the evect of the 1-1-2/1-2-1 haplogenotype was no longer signiwcant (OR D 1.25 ( ), P D 0.06) in the pooled analysis, and in the meta-analysis, the heterogeneity in the 1-2-1/1-2-1 haplogenotype group was slightly strengthened (P D 0.05). In the meta-analysis, the P-values for heterogeneity in the 1-2-1/1-2-1 haplogenotype group in population-based and linkage-based cases were 0.89 (see above) and 0.07, respectively. Table 4 CAPN10 haplogenotype and risk of type 2 diabetes in Europeans stratiwed based on ascertainment of cases (population-based vs. linkage-based) Haplotype Overall Population-based cases Frequency Pooled Meta-analysis Frequency Pooled Meta-analysis Fixed evects Random evects Het Fixed evects Random evects Het Ctrl T2D OR (95% CI) P OR (95% CI) P OR (95% CI) P P Ctrl T2D OR (95% CI) P OR (95% CI) P OR (95% CI) P P ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) ( ) The number of studies is 11 for the meta-analysis in the overall group. The number of studies is 8 for the meta-analysis using population-based cases. Ctrl, controls; T2D, type 2 diabetes; OR, odds ratio; CI, conwdence interval; Het, heterogeneity. T. Tsuchiya et al. / Molecular Genetics and Metabolism 89 (2006)
7 180 T. Tsuchiya et al. / Molecular Genetics and Metabolism 89 (2006) strong LD or haplotype blocks as dewned by Gabriel et al. (D conwdence intervals) [18] and Wang (four-gamete rule) [19] (Fig. 1). The Wrst spans a region of 36.9 kb (nt 7235 (relative to the ATG of CAPN10; SNP-66) to nt (SNP-32)) following the four-gamete rule, or 39.3 kb (nt 7235 to nt (SNP-42)) following the D conwdence interval dewnition, that includes CAPN10 as well as the four polymorphisms that have been examined in many studies of this locus (SNP-44, SNP-43, Indel-19 and SNP- 63). The second region dewned by the four-gamete rule spans 9 kb and is located on the 3 -side of GPR35 (nt (SNP-35) to nt (SNP-31)). Near identical LD patterns are observed in Europeans in HapMap [21]. As we observed in the Mexican American population [28], the LD decays rapidly on both sides of the CAPN10-associated haplotype block reducing the likelihood that association between a variant in CAPN10 and T2D is due to LD with a marker outside this block. This is conwrmed by the observation that in the HapMap Europeans [21] no polymorphism within the 5 Mb telomeric end of chromosome 2, other than those within the CAPN10 block itself which is 1.3 Mb from the telomeric end of chromosome 2, have r 2 values 70.3 Fig. 1. Linkage disequilibrium and haplotype block structure for the region surrounding CAPN10 in Europeans. Informative pairwise measures indicating evidence for strong linkage disequilibrium (black), or historical recombination (white) are color-coded in the triangular matrix. Inconclusive pairwise comparisons are grey. Above the LD matrix, polymorphisms (SNP or Indel) are labeled by their dbsnp reference (rs) number followed by their UCSNP designation (in brackets) from the original study [4]. The position of each polymorphism and the refseq genes mapped within the 255 kb region of 2q37 analysed are also labeled along the diagonal. To the left of the matrix are the minor allele frequencies of each polymorphism. The red line identiwes the haplotype block using the D conwdence intervals [18], and the blue line indicates haplotype blocks following the four-gamete rule [19]. The haplotype frequencies (71%) of the D conwdence interval dewned block are shown at the top. Minor alleles are indicated in black squares, major alleles in white squares, and the haplotype frequencies are noted to the right. Haplotypes are sorted by SNP-44, SNP-43, Indel-19 and SNP-63 polymorphisms.
8 T. Tsuchiya et al. / Molecular Genetics and Metabolism 89 (2006) with SNP-44, -43 or -48 (used as a proxy for Indel-19 since r 2 D 1 between SNP-48 and Indel-19 in both the Europeans and Mexican Americans). SNP-63 could not be tested since neither it nor a SNP highly correlated with it is included in the HapMap. We next calculated the haplotype frequencies by Wrst phasing the genotypes for the 15 polymorphisms in the CAPN10 block in 63 individuals with genotypes for at least 12 of the 15 polymorphisms. The block is comprised of 6 haplotypes of 75% frequency that together capture 93% of the haplotypic variation at this locus (9 haplotypes 71% account for 98% of the variation). We are conwdent these 6 haplotypes estimated to occur at a frequency of 75% are real since they are each observed in homozygous form in at least one of the 63 individuals. What is quite clear from this pattern of haplotype diversity is that SNP-44, -43, and - 63, and Indel-19 are not suycient to tag all of the variation observed in the block. Forcing these four polymorphisms to tag the remaining eleven polymorphisms, only 4/11 are tagged at r and 8/11 are tagged at r The r 2 threshold must be reduced to for SNP-44, SNP-43, Indel-19 and SNP-63 to successfully tag all 11 SNPs in the LD block. Restricting our analyses to common polymorphisms (MAF ), eight polymorphisms (SNP-66, rs , SNP-44, SNP-43, Indel-19, SNP-69, SNP-32 and SNP-42; SNP-63 was not considered because MAF D 0.068) are required to tag the diversity in this region at r and Wve polymorphisms (SNP-66, SNP-44, SNP-43, Indel-19 and SNP-32) at r At the same MAF threshold, nine SNPs are required to tag 21 SNPs in HapMap spanning the same region (SNP-66 to SNP-42) at r including rs , SNP-44, SNP-43, SNP-77, Indel-19, SNP-47, rs , rs and SNP-32. Discussion The results of pooled and meta-analyses of 3084 cases (excluding UK-1) and 2935 controls presented here show that genetic variation in CAPN10 is associated with T2D in Europeans as suggested previously in some studies involving smaller numbers of cases and controls [4,10,11] but not others [6,7,9]. The results are also in agreement with the meta-analysis of Song et al. [13]; however, here we have removed the duplication in some of the data sets examined in their report. The strength of the disease association is similar for SNP-43 and the SNP-43 Indel-19 SNP-63 haplotypes and (Tables 2 and 4) consistent with SNP-43 playing role in determining risk as suggested previously [4]. However, not all haplotypes harboring the SNP- 43 G allele are associated with increased risk; e.g., With regard to the haplotype, SNP-44 dewnes a subset of haplotypes associated with increased risk and this SNP is itself an independent risk factor for T2D [12]. Thus, haplotypes better describe risk than a single SNP, consistent with either cis-interaction or the tagging of a rarer variant. Since two diverent haplotypes are associated with increased risk, the former seems more likely than the latter. In Europeans, haplogenotypes containing both and haplotypes are associated with highest risk including the 1-1-2/1-2-1 combination found to be associated with T2D in Mexican Americans although in Mexican Americans neither homozygous combination appeared to be associated with increased risk. The 95% conwdence intervals for the odds ratios associated with the homozygous haplogenotypes do, however, overlap in the Europeans and Mexican Americans suggesting that they could have similar evects on risk in both populations. In a Japanese study involving 827 cases and 825 controls, we found no association of the CAPN10 variants described here with T2D including the 1-2-1/1-2-1 haplogenotype [29]. Moreover, there is no overlap in the 95% conwdence interval for 1-2-1/ odds ratio in Japanese and Europeans raising the possibility there is additional variation that diverentiates risk and non-risk forms of the haplotype. Such heterogeneity might have contributed to the low risk associated with the haplotype in the studies in Mexican Americans [4]. Our studies and those of others indicate that the three variants examined here (SNP-43, Indel-19 and SNP-63) and the SNP-44/Thr504Ala polymorphism do not capture all of the variation at CAPN10 that may be relevant to risk of T2D and related phenotypes. It is also possible that the heterogeneity in risks for the haplotype observed between European and Japanese samples is attributable to unlinked genetic variants or environmental factors that modify the evects of CAPN10. The results presented here suggest that the method by which cases are selected may confound the results of a casecontrol study. Our data suggest that cases selected from a previous linkage study (e.g., an avected sibpair study) may diver with respect to genetic risk factors compared to population-based cases although as shown in Fig. 2, the conwdence intervals for the OR overlap implying that diverences are not signiwcant, at least with these samples. However, the sizes of the groups studied are still relatively small and the diverences may disappear when larger numbers of population- and linkage-based cases are examined. CAPN10 was also identiwed as a diabetes-susceptibility gene by positional cloning and thus is not a typical candidate gene like PPARG and KCNJ11 which show no evidence for linkage with T2D. In general, we would expect cases from families with at least two avected siblings to generate higher OR for genotypes increasing risk of disease than cases unselected for family history [27]. However, when there are many genetic risk factors that can avect susceptibility, the evidence for linkage can impact the expected OR for a susceptibility locus in that region. For example, we expect the OR for a susceptibility locus estimated from cases drawn from families generating strong evidence for linkage at the location of the susceptibility locus to be higher than the OR in a set of cases unselected for positive family history or linkage evidence. The higher expectation rexects both the general tendency for cases with positive family history (e.g., at least one avected sib) to be enriched for alleles increasing risk of disease and the evects of the winner s curse in linkage
9 182 T. Tsuchiya et al. / Molecular Genetics and Metabolism 89 (2006) Fig. 2. Meta-analysis plot. The odds ratio with 95% CI for the association of the 1-2-1/1-2-1 haplogenotype with T2D for the 12 studies described here are shown. * The result was derived from French data set excluding probands of sibs. ** The result was derived from French data set including probands of sibs. Note that the size of the plotting symbol does not indicate the weight given to the trial in the analysis. and positional cloning studies. The regions most likely to be subject to positional cloning studies are those with the strongest evidence for linkage. The strongest linkage signals, in turn, overestimate the actual evect size of the susceptibility gene at that location. To the extent that this overestimate rexects a chance oversampling (in the linkage sample) of families segregating for a susceptibility allele at this location, the OR estimates from these same cases will be substantially larger than the OR estimated from cases unselected for linkage to this region. In addition for some genetic models, family history positive but linkage discordant datasets can generate a lower OR than observed in unselected case samples (B. Voight and NJC, unpublished). There may be other explanations for the heterogeneity in OR among the diverent samples including overall sample size and diverences in the ascertainment of controls (see Table 1). What is the magnitude of the evect of variation in CAPN10 on T2D risk in Europeans? The results presented here show the evect depends on the combination of speciwc alleles or haplotypes inherited with some variants increasing risk by perhaps as much as 168% (e.g., meta-analysis and 1-2-1/1-2-1 haplogenotype, Table 3) and other combinations decreasing risk by up to 15% (e.g., SNP-43 A allele acting in a dominant manner, Table 2). Overall, the evects of variants in CAPN10 on T2D risk appear similar to those reported for PPARG and KCNJ11, two generally accepted T2D susceptibility loci, in similar large studies of Europeans [30,31]. The estimates of the evect of CAPN10 on T2D risk in Europeans are less than noted in our original report which overestimated the evects of this locus (OR for SNP- 43 and 1-1-2/1-2-1 haplogenotype D 1.54 and 2.80, respectively) [4]. This may be because we studied cases from families showing evidence for linkage with CAPN10 or may rexect the instability of point estimates with wide conwdence intervals due to small sizes. The genetic evidence suggests that variation in CAPN10 avects risk of T2D in Europeans. Variants in CAPN10 (SNP-44 and SNP-43) and PPARG (Pro12Ala mutation) predict T2D in the Botnia study from Finland [32]. Future studies need to dewne the nature of the cis-interactions in CAPN10 that distinguish haplotypes that increase risk from those that decrease and determine the best combinations of polymorphisms for dewning risk alleles for prospective studies, including evorts using genetic markers to predict future T2D. Acknowledgments This study was supported in part by U.S. Public Health Service (Grants DK-20595, , and ), a grant from the Technical University Dresden (Med Drive), a grant from the Czech government (IGA MH CR NR/ ), and a gift from the Kovler Family Foundation. M.G.H. was supported by a Mentor-based Fellowship from the American Diabetes Association. G.W.T. was supported by a postdoctoral fellowship from the North-West University (Potchefstroom Campus).
10 T. Tsuchiya et al. / Molecular Genetics and Metabolism 89 (2006) Appendix A. Supplementary data Supplementary data associated with this article can be found, in the online version, at doi: / j.ymgme References [1] M.I. McCarthy, Progress in dewning the molecular basis of type 2 diabetes mellitus through susceptibility gene identiwcation, Hum. Mol. Genet. (2004) R33 R41. [2] C.L. Hanis, E. Boerwinkle, R. Chakraborty, D.L. Ellsworth, P. Concannon, B. Stirling, V.A. Morrison, B. Wapelhorst, R.S. Spielman, K.J. Gogolin-Ewens, J.M. Shepard, S.R. Williams, N. Risch, D. Hinds, N. Iwasaki, M. Ogata, Y. Omori, C. Petzold, H. Rietzch, H.E. Schroder, J. Schulze, N.J. Cox, S. Menzel, V.V. Boriraj, X. Chen, L.R. Lim, T. Lindner, L.E. Mereu, Y.Q. Wang, K. Xiang, K. Yamagata, Y. Yang, G.I. Bell, A genome-wide search for human non-insulindependent (type 2) diabetes genes reveals a major susceptibility locus on chromosome 2, Nat. Genet. 13 (1996) [3] N.J. Cox, M. Frigge, D.L. Nicolae, P. Concannon, C.L. Hanis, G.I. Bell, A. Kong, Loci on chromosomes 2 (NIDDM1) and 15 interact to increase susceptibility to diabetes in Mexican Americans, Nat. Genet. 21 (1999) [4] Y. Horikawa, N. Oda, N.J. Cox, X. Li, M. Orho-Melander, M. Hara, Y. Hinokio, T.H. Lindner, H. Mashima, P.E. Schwarz, L. del Bosque- Plata, Y. Oda, I. Yoshiuchi, S. Colilla, K.S. Polonsky, S. Wei, P. Concannon, N. Iwasaki, J. Schulze, L.J. Baier, C. Bogardus, L. Groop, E. Boerwinkle, C.L. Hanis, G.I. Bell, Genetic variation in the gene encoding calpain-10 is associated with type 2 diabetes mellitus, Nat. Genet. 26 (2000) [5] E. Carlsson, P. Poulsen, H. Storgaard, P. Almgren, C. Ling, C.B. Jensen, S. Madsbad, L. Groop, A. Vaag, M. Ridderstrale, Genetic and nongenetic regulation of CAPN10 mrna in skeletal muscle, Diabetes 54 (2005) [6] J.C. Evans, T.M. Frayling, P.G. Cassell, P.J. Saker, G.A. Hitman, M. Walker, J.C. Levy, S. O Rahilly, P.V. Rao, A.J. Bennett, E.C. Jones, S. Menzel, P. Prestwich, N. Simecek, M. Wishart, R. Dhillon, C. Fletcher, A. Millward, A. Demaine, T. Wilkin, Y. Horikawa, N.J. Cox, G.I. Bell, S. Ellard, M.I. McCarthy, A.T. Hattersley, Studies of association between the gene for calpain-10 and type 2 diabetes mellitus in the United Kingdom, Am. J. Hum. Genet. 69 (2001) [7] S.K. Rasmussen, S.A. Urhammer, L. Berglund, J.N. Jensen, L. Hansen, S.M. Echwald, K. Borch-Johnsen, Y. Horikawa, H. Mashima, H. Lithell, N.J. Cox, T. Hansen, G.I. Bell, O. Pedersen, Variants within the calpain-10 gene on chromosome 2q37 (NIDDM1) and relationships to type 2 diabetes, insulin resistance, and impaired acute insulin secretion among Scandinavian Caucasians, Diabetes 51 (2002) [8] S.C. Elbein, W. Chu, Q. Ren, C. Hemphill, J. Schay, N.J. Cox, C.L. Hanis, S.J. Hasstedt, Role of calpain-10 gene variants in familial type 2 diabetes in Caucasians, J. Clin. Endocrinol. Metab. 87 (2002) [9] T.E. Fingerlin, M.R. Erdos, R.M. Watanabe, K.R. Wiles, H.M. Stringham, K.L. Mohlke, K. Silander, T.T. Valle, T.A. Buchanan, J. Tuomilehto, R.N. Bergman, M. Boehnke, F.S. Collins, Variation in three single nucleotide polymorphisms in the calpain-10 gene not associated with type 2 diabetes in a large Finnish cohort, Diabetes 51 (2002) [10] M.T. Malecki, D.K. Moczulski, T. Klupa, K. Wanic, K. Cyganek, J. Frey, J. Sieradzki, Homozygous combination of calpain 10 gene haplotypes is associated with type 2 diabetes mellitus in a Polish population, Eur. J. Endocrinol. 146 (2002) [11] M. Orho-Melander, M. Klannemark, M.K. Svensson, M. Ridderstrale, C.M. Lindgren, L. Groop, Variants in the calpain-10 gene predispose to insulin resistance and elevated free fatty acid levels, Diabetes 51 (2002) [12] M.N. Weedon, P.E. Schwarz, Y. Horikawa, N. Iwasaki, T. Illig, R. Holle, W. Rathmann, T. Selisko, J. Schulze, K.R. Owen, J. Evans, L. del Bosque-Plata, G. Hitman, M. Walker, J.C. Levy, M. Sampson, G.I. Bell, M.I. McCarthy, A.T. Hattersley, T.M. Frayling, Meta-analysis and a large association study conwrm a role for calpain-10 variation in type 2 diabetes susceptibility, Am. J. Hum. Genet. 73 (2003) [13] Y. Song, T. Niu, J.E. Manson, D.J. Kwiatkowski, S. Liu, Are variants in the CAPN10 gene related to risk of type 2 diabetes? A quantitative assessment of population and family-based association studies, Am. J. Hum. Genet. 74 (2004) [14] M.I. McCarthy, P.H. Groop, T. Hansen, Making the right associations, Diabetologia 48 (2005) [15] C.L. Hanis, D. Hewett-Emmett, T.K. Bertin, W.J. Schull, Origins of U.S. Hispanics. Implications for diabetes, Diabetes Care 14 (1991) [16] M. Berger, H.H. Stassen, K. Kohler, V. Krane, D. Monks, C. Wanner, K. HoVmann, M.M. HoVmann, M. Zimmer, H. Bickeboller, T.H. Lindner, Hidden population substructures in an apparently homogeneous population bias association studies, Eur. J. Hum. Genet. 14 (2006) [17] J.C. Barrett, B. Fry, J. Maller, M.J. Daly, Haploview: analysis and visualization of LD and haplotype maps, Bioinformatics 21 (2005) [18] S.B. Gabriel, S.F. SchaVner, H. Nguyen, J.M. Moore, J. Roy, B. Blumenstiel, J. Higgins, M. DeFelice, A. Lochner, M. Faggart, S.N. Liu- Cordero, C. Rotimi, A. Adeyemo, R. Cooper, R. Ward, E.S. Lander, M.J. Daly, D. Altshuler, The structure of haplotype blocks in the human genome, Science 296 (2002) [19] N. Wang, J.M. Akey, K. Zhang, R. Chakraborty, L. Jin, Distribution of recombination crossovers and the origin of haplotype blocks: the interplay of population history, recombination, and mutation, Am. J. Hum. Genet. 71 (2002) [20] P.I.W. de Bakker, R. Yelensky, I. Pe er, S.B. Gabriel, M.J. Daly, D. Altshuler, EYciency and power in genetic association studies, Nat. Genet. 37 (2005) [21] The International HapMap Consortium, The International HapMap Project, Nature 426 (2003) [22] M. Stephens, P. Donnelly, A comparison of bayesian methods for haplotype reconstruction from population genotype data, Am. J. Hum. Genet. 73 (2003) [23] M. Stephens, N.J. Smith, P. Donnelly, A new statistical method for haplotype reconstruction from population data, Am. J. Hum. Genet. 68 (2001) [24] S. Purcell, S.S. Cherny, P.C. Sham, Genetic Power Calculator: design of linkage and association genetic mapping studies of complex traits, Bioinformatics 19 (2003) [25] S. Muncer, S. Taylor, M. Craigie, Power dressing and meta-analysis: incorporating power analysis into meta-analysis, J. Adv. Nurs. 38 (2002) [26] R. Driver, Response to: Power dressing and meta-analysis: incorporating power analysis into meta-analysis by S. Muncer, Journal of Advanced Nursing 38 (2002) ; J. Adv. Nurs. 38 (2002) [27] N. Risch, Implications of multilocus inheritance for gene-disease association studies, Theor. Popul. Biol. 60 (2001) [28] M.G. Hayes, L. del Bosque-Plata, T. Tsuchiya, C.L. Hanis, G.I. Bell, N.J. Cox, Patterns of linkage disequilibrium in the type 2 diabetes gene calpain-10, Diabetes 54 (2005) [29] N. Iwasaki, Y. Horikawa, T. Tsuchiya, Y. Kitamura, T. Nakamura, Y. Tanizawa, Y. Oka, K. Hara, T. Kadowaki, T. Awata, M. Honda, K. Yamashita, N. Oda, L. Yu, N. Yamada, M. Ogata, N. Kamatani, Y. Iwamoto, L. del Bosque-Plata, M.G. Hayes, N.J. Cox, G.I. Bell, Genetic variants in the calpain-10 gene and the development of type 2 diabetes in the Japanese population, J. Hum. Genet. 50 (2005) [30] D. Altshuler, J.N. Hirschhorn, M. Klannemark, C.M. Lindgren, M.C. Vohl, J. Nemesh, C.R. Lane, S.F. SchaVner, S. Bolk, C. Brewer,
11 184 T. Tsuchiya et al. / Molecular Genetics and Metabolism 89 (2006) T. Tuomi, D. Gaudet, T.J. Hudson, M. Daly, L. Groop, E.S. Lander, The common PPARgamma Pro12Ala polymorphism is associated with decreased risk of type 2 diabetes, Nat. Genet. 26 (2000) [31] J.C. Florez, N. Burtt, P.I. de Bakker, P. Almgren, T. Tuomi, J. Holmkvist, D. Gaudet, T.J. Hudson, S.F. SchaVner, M.J. Daly, J.N. Hirschhorn, L. Groop, D. Altshuler, Haplotype structure and genotype-phenotype correlations of the sulfonylurea receptor and the islet ATP-sensitive potassium channel gene region, Diabetes 53 (2004) [32] V. Lyssenko, P. Almgren, D. Anevski, M. Orho-Melander, M. Sjögren, C. Saloranta, T. Tuomi, L. Groop, The Botnia Study Group, Genetic prediction of future type 2 diabetes, PLoS Med. 2 (2005) e345.
In 1996, Hanis et al. (1) reported that a genome-wide
 Brief Genetics Report Variation in Three Single Nucleotide Polymorphisms in the Calpain-10 Gene Not Associated With Type 2 Diabetes in a Large Finnish Cohort Tasha E. Fingerlin, 1,2 Michael R. Erdos, 3
Brief Genetics Report Variation in Three Single Nucleotide Polymorphisms in the Calpain-10 Gene Not Associated With Type 2 Diabetes in a Large Finnish Cohort Tasha E. Fingerlin, 1,2 Michael R. Erdos, 3
Homozygous combination of calpain 10 gene haplotypes is associated with type 2 diabetes mellitus in a Polish population
 European Journal of Endocrinology (2002) 146 695 699 ISSN 0804-4643 CLINICAL STUDY Homozygous combination of calpain 10 gene haplotypes is associated with type 2 diabetes mellitus in a Polish population
European Journal of Endocrinology (2002) 146 695 699 ISSN 0804-4643 CLINICAL STUDY Homozygous combination of calpain 10 gene haplotypes is associated with type 2 diabetes mellitus in a Polish population
TCF7L2 polymorphisms are associated with type 2 diabetes in northern Sweden
 (2007) 15, 342 346 & 2007 Nature Publishing Group All rights reserved 1018-4813/07 $30.00 ARTICLE www.nature.com/ejhg TCF7L2 polymorphisms are associated with type 2 diabetes in northern Sweden Sofia Mayans
(2007) 15, 342 346 & 2007 Nature Publishing Group All rights reserved 1018-4813/07 $30.00 ARTICLE www.nature.com/ejhg TCF7L2 polymorphisms are associated with type 2 diabetes in northern Sweden Sofia Mayans
FTO Polymorphisms Are Associated with Obesity But Not with Diabetes in East Asian Populations: A Meta analysis
 BIOMEDICAL AND ENVIRONMENTAL SCIENCES 22, 449 457 (2009) www.besjournal.com FTO Polymorphisms Are Associated with Obesity But Not with Diabetes in East Asian Populations: A Meta analysis BO XI #, + AND
BIOMEDICAL AND ENVIRONMENTAL SCIENCES 22, 449 457 (2009) www.besjournal.com FTO Polymorphisms Are Associated with Obesity But Not with Diabetes in East Asian Populations: A Meta analysis BO XI #, + AND
/02/$15.00/0 The Journal of Clinical Endocrinology & Metabolism 87(6): Copyright 2002 by The Endocrine Society
 0013-7227/02/$15.00/0 The Journal of Clinical Endocrinology & Metabolism 87(6):2606 2610 Printed in U.S.A. Copyright 2002 by The Endocrine Society Variation within the Type 2 Diabetes Susceptibility Gene
0013-7227/02/$15.00/0 The Journal of Clinical Endocrinology & Metabolism 87(6):2606 2610 Printed in U.S.A. Copyright 2002 by The Endocrine Society Variation within the Type 2 Diabetes Susceptibility Gene
Type 2 diabetes accounts for the majority of
 Brief Genetics Report Linkage but Not Association of Calpain-10 to Type 2 Diabetes Replicated in Northern Sweden Elisabet Einarsdottir, 1 Sofia Mayans, 1 Karin Ruikka, 2 Stefan A. Escher, 1 Petter Lindgren,
Brief Genetics Report Linkage but Not Association of Calpain-10 to Type 2 Diabetes Replicated in Northern Sweden Elisabet Einarsdottir, 1 Sofia Mayans, 1 Karin Ruikka, 2 Stefan A. Escher, 1 Petter Lindgren,
Hepatocyte nuclear factor (HNF)-4 is an excellent
 Brief Genetics Report Common Variants of the Hepatocyte Nuclear Factor-4 P2 Promoter Are Associated With Type 2 Diabetes in the U.K. Population Michael N. Weedon, 1 Katharine R. Owen, 1 Beverley Shields,
Brief Genetics Report Common Variants of the Hepatocyte Nuclear Factor-4 P2 Promoter Are Associated With Type 2 Diabetes in the U.K. Population Michael N. Weedon, 1 Katharine R. Owen, 1 Beverley Shields,
Supplementary Figure 1. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations.
 Supplementary Figure. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations. a Eigenvector 2.5..5.5. African Americans European Americans e
Supplementary Figure. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations. a Eigenvector 2.5..5.5. African Americans European Americans e
Introduction to Genetics and Genomics
 2016 Introduction to enetics and enomics 3. ssociation Studies ggibson.gt@gmail.com http://www.cig.gatech.edu Outline eneral overview of association studies Sample results hree steps to WS: primary scan,
2016 Introduction to enetics and enomics 3. ssociation Studies ggibson.gt@gmail.com http://www.cig.gatech.edu Outline eneral overview of association studies Sample results hree steps to WS: primary scan,
During the hyperinsulinemic-euglycemic clamp [1], a priming dose of human insulin (Novolin,
![During the hyperinsulinemic-euglycemic clamp [1], a priming dose of human insulin (Novolin, During the hyperinsulinemic-euglycemic clamp [1], a priming dose of human insulin (Novolin,](/thumbs/72/66792834.jpg) ESM Methods Hyperinsulinemic-euglycemic clamp procedure During the hyperinsulinemic-euglycemic clamp [1], a priming dose of human insulin (Novolin, Clayton, NC) was followed by a constant rate (60 mu m
ESM Methods Hyperinsulinemic-euglycemic clamp procedure During the hyperinsulinemic-euglycemic clamp [1], a priming dose of human insulin (Novolin, Clayton, NC) was followed by a constant rate (60 mu m
Association between the CYP11B2 gene 344T>C polymorphism and coronary artery disease: a meta-analysis
 Association between the CYP11B2 gene 344T>C polymorphism and coronary artery disease: a meta-analysis Y. Liu, H.L. Liu, W. Han, S.J. Yu and J. Zhang Department of Cardiology, The General Hospital of the
Association between the CYP11B2 gene 344T>C polymorphism and coronary artery disease: a meta-analysis Y. Liu, H.L. Liu, W. Han, S.J. Yu and J. Zhang Department of Cardiology, The General Hospital of the
Protein tyrosine phosphatase 1B is not a major susceptibility gene for type 2 diabetes mellitus or obesity among Pima Indians
 Diabetologia (2007) 50:985 989 DOI 10.1007/s00125-007-0611-6 SHORT COMMUNICATION Protein tyrosine phosphatase 1B is not a major susceptibility gene for type 2 diabetes mellitus or obesity among Pima Indians
Diabetologia (2007) 50:985 989 DOI 10.1007/s00125-007-0611-6 SHORT COMMUNICATION Protein tyrosine phosphatase 1B is not a major susceptibility gene for type 2 diabetes mellitus or obesity among Pima Indians
Introduction to the Genetics of Complex Disease
 Introduction to the Genetics of Complex Disease Jeremiah M. Scharf, MD, PhD Departments of Neurology, Psychiatry and Center for Human Genetic Research Massachusetts General Hospital Breakthroughs in Genome
Introduction to the Genetics of Complex Disease Jeremiah M. Scharf, MD, PhD Departments of Neurology, Psychiatry and Center for Human Genetic Research Massachusetts General Hospital Breakthroughs in Genome
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.
 Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Human population sub-structure and genetic association studies
 Human population sub-structure and genetic association studies Stephanie A. Santorico, Ph.D. Department of Mathematical & Statistical Sciences Stephanie.Santorico@ucdenver.edu Global Similarity Map from
Human population sub-structure and genetic association studies Stephanie A. Santorico, Ph.D. Department of Mathematical & Statistical Sciences Stephanie.Santorico@ucdenver.edu Global Similarity Map from
BST227 Introduction to Statistical Genetics. Lecture 4: Introduction to linkage and association analysis
 BST227 Introduction to Statistical Genetics Lecture 4: Introduction to linkage and association analysis 1 Housekeeping Homework #1 due today Homework #2 posted (due Monday) Lab at 5:30PM today (FXB G13)
BST227 Introduction to Statistical Genetics Lecture 4: Introduction to linkage and association analysis 1 Housekeeping Homework #1 due today Homework #2 posted (due Monday) Lab at 5:30PM today (FXB G13)
Effects of environment and genetic interactions on chronic metabolic diseases
 22 1 2010 1 Chinese Bulletin of Life Sciences Vol. 22, No. 1 Jan., 2010 1004-0374(2010)01-0001-06 ( 200031) 2 2 20 2 2 2 R151; R589; R587.1; R363.16 A Effects of environment and genetic interactions on
22 1 2010 1 Chinese Bulletin of Life Sciences Vol. 22, No. 1 Jan., 2010 1004-0374(2010)01-0001-06 ( 200031) 2 2 20 2 2 2 R151; R589; R587.1; R363.16 A Effects of environment and genetic interactions on
Ascertainment Through Family History of Disease Often Decreases the Power of Family-based Association Studies
 Behav Genet (2007) 37:631 636 DOI 17/s10519-007-9149-0 ORIGINAL PAPER Ascertainment Through Family History of Disease Often Decreases the Power of Family-based Association Studies Manuel A. R. Ferreira
Behav Genet (2007) 37:631 636 DOI 17/s10519-007-9149-0 ORIGINAL PAPER Ascertainment Through Family History of Disease Often Decreases the Power of Family-based Association Studies Manuel A. R. Ferreira
SUPPLEMENTARY DATA. 1. Characteristics of individual studies
 1. Characteristics of individual studies 1.1. RISC (Relationship between Insulin Sensitivity and Cardiovascular disease) The RISC study is based on unrelated individuals of European descent, aged 30 60
1. Characteristics of individual studies 1.1. RISC (Relationship between Insulin Sensitivity and Cardiovascular disease) The RISC study is based on unrelated individuals of European descent, aged 30 60
Assessing Accuracy of Genotype Imputation in American Indians
 Assessing Accuracy of Genotype Imputation in American Indians Alka Malhotra*, Sayuko Kobes, Clifton Bogardus, William C. Knowler, Leslie J. Baier, Robert L. Hanson Phoenix Epidemiology and Clinical Research
Assessing Accuracy of Genotype Imputation in American Indians Alka Malhotra*, Sayuko Kobes, Clifton Bogardus, William C. Knowler, Leslie J. Baier, Robert L. Hanson Phoenix Epidemiology and Clinical Research
The insulin-degrading enzyme (IDE or insulysin,
 Brief Genetics Report High-Density Haplotype Structure and Association Testing of the Insulin-Degrading Enzyme (IDE) Gene With Type 2 Diabetes in 4,206 People Jose C. Florez, 1,2,3,4 Steven Wiltshire,
Brief Genetics Report High-Density Haplotype Structure and Association Testing of the Insulin-Degrading Enzyme (IDE) Gene With Type 2 Diabetes in 4,206 People Jose C. Florez, 1,2,3,4 Steven Wiltshire,
Association between the -77T>C polymorphism in the DNA repair gene XRCC1 and lung cancer risk
 Association between the -77T>C polymorphism in the DNA repair gene XRCC1 and lung cancer risk B.B. Sun, J.Z. Wu, Y.G. Li and L.J. Ma Department of Respiratory Medicine, People s Hospital Affiliated to
Association between the -77T>C polymorphism in the DNA repair gene XRCC1 and lung cancer risk B.B. Sun, J.Z. Wu, Y.G. Li and L.J. Ma Department of Respiratory Medicine, People s Hospital Affiliated to
Common human diseases, like diabetes, cancer,
 Original Article Evaluation of Common Variants in the Six Known Maturity-Onset Diabetes of the Young (MODY) Genes for Association With Type 2 Diabetes Wendy Winckler, 1,2,3 Michael N. Weedon, 4 Robert
Original Article Evaluation of Common Variants in the Six Known Maturity-Onset Diabetes of the Young (MODY) Genes for Association With Type 2 Diabetes Wendy Winckler, 1,2,3 Michael N. Weedon, 4 Robert
LTA Analysis of HapMap Genotype Data
 LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
Results. Introduction
 (2009) 10, S95 S120 & 2009 Macmillan Publishers Limited All rights reserved 1466-4879/09 $32.00 www.nature.com/gene ORIGINAL ARTICLE Analysis of 55 autoimmune disease and type II diabetes loci: further
(2009) 10, S95 S120 & 2009 Macmillan Publishers Limited All rights reserved 1466-4879/09 $32.00 www.nature.com/gene ORIGINAL ARTICLE Analysis of 55 autoimmune disease and type II diabetes loci: further
Supplementary Figure 1 Forest plots of genetic variants in GDM with all included studies. (A) IGF2BP2
 Supplementary Figure 1 Forest plots of genetic variants in GDM with all included studies. (A) IGF2BP2 rs4402960, (B) MTNR1B rs10830963, (C) TCF7L2 rs7903146, (D) IRS1 rs1801278, (E) PPARG rs1801282, (F)
Supplementary Figure 1 Forest plots of genetic variants in GDM with all included studies. (A) IGF2BP2 rs4402960, (B) MTNR1B rs10830963, (C) TCF7L2 rs7903146, (D) IRS1 rs1801278, (E) PPARG rs1801282, (F)
Dan Koller, Ph.D. Medical and Molecular Genetics
 Design of Genetic Studies Dan Koller, Ph.D. Research Assistant Professor Medical and Molecular Genetics Genetics and Medicine Over the past decade, advances from genetics have permeated medicine Identification
Design of Genetic Studies Dan Koller, Ph.D. Research Assistant Professor Medical and Molecular Genetics Genetics and Medicine Over the past decade, advances from genetics have permeated medicine Identification
Hepatocyte nuclear factor 4 (HNF4A) is a transcription
 Brief Report P2 Promoter Variants of the Hepatocyte Nuclear Factor 4 Gene Are Associated With Type 2 Diabetes in Mexican Americans Donna M. Lehman, 1 Dawn K. Richardson, 2 Chris P. Jenkinson, 2 Kelly J.
Brief Report P2 Promoter Variants of the Hepatocyte Nuclear Factor 4 Gene Are Associated With Type 2 Diabetes in Mexican Americans Donna M. Lehman, 1 Dawn K. Richardson, 2 Chris P. Jenkinson, 2 Kelly J.
FTO gene variants are strongly associated with type 2 diabetes in South Asian Indians
 Diabetologia (2009) 52:247 252 DOI 10.1007/s00125-008-1186-6 SHORT COMMUNICATION FTO gene variants are strongly associated with type 2 diabetes in South Asian Indians C. S. Yajnik & C. S. Janipalli & S.
Diabetologia (2009) 52:247 252 DOI 10.1007/s00125-008-1186-6 SHORT COMMUNICATION FTO gene variants are strongly associated with type 2 diabetes in South Asian Indians C. S. Yajnik & C. S. Janipalli & S.
CS2220 Introduction to Computational Biology
 CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
SHORT COMMUNICATION. K. Lukacs & N. Hosszufalusi & E. Dinya & M. Bakacs & L. Madacsy & P. Panczel
 Diabetologia (2012) 55:689 693 DOI 10.1007/s00125-011-2378-z SHORT COMMUNICATION The type 2 diabetes-associated variant in TCF7L2 is associated with latent autoimmune diabetes in adult Europeans and the
Diabetologia (2012) 55:689 693 DOI 10.1007/s00125-011-2378-z SHORT COMMUNICATION The type 2 diabetes-associated variant in TCF7L2 is associated with latent autoimmune diabetes in adult Europeans and the
ASSOCIATION OF KCNJ1 VARIATION WITH CHANGE IN FASTING GLUCOSE AND NEW ONSET DIABETES DURING HCTZ TREATMENT
 ONLINE SUPPLEMENT ASSOCIATION OF KCNJ1 VARIATION WITH CHANGE IN FASTING GLUCOSE AND NEW ONSET DIABETES DURING HCTZ TREATMENT Jason H Karnes, PharmD 1, Caitrin W McDonough, PhD 1, Yan Gong, PhD 1, Teresa
ONLINE SUPPLEMENT ASSOCIATION OF KCNJ1 VARIATION WITH CHANGE IN FASTING GLUCOSE AND NEW ONSET DIABETES DURING HCTZ TREATMENT Jason H Karnes, PharmD 1, Caitrin W McDonough, PhD 1, Yan Gong, PhD 1, Teresa
White Paper Guidelines on Vetting Genetic Associations
 White Paper 23-03 Guidelines on Vetting Genetic Associations Authors: Andro Hsu Brian Naughton Shirley Wu Created: November 14, 2007 Revised: February 14, 2008 Revised: June 10, 2010 (see end of document
White Paper 23-03 Guidelines on Vetting Genetic Associations Authors: Andro Hsu Brian Naughton Shirley Wu Created: November 14, 2007 Revised: February 14, 2008 Revised: June 10, 2010 (see end of document
Introduction to linkage and family based designs to study the genetic epidemiology of complex traits. Harold Snieder
 Introduction to linkage and family based designs to study the genetic epidemiology of complex traits Harold Snieder Overview of presentation Designs: population vs. family based Mendelian vs. complex diseases/traits
Introduction to linkage and family based designs to study the genetic epidemiology of complex traits Harold Snieder Overview of presentation Designs: population vs. family based Mendelian vs. complex diseases/traits
Genetic association analysis of LARS2 with type 2 diabetes
 Diabetologia (2010) 53:103 110 DOI 10.1007/s00125-009-1557-7 ARTICLE Genetic association analysis of LARS2 with type 2 diabetes E. Reiling & B. Jafar-Mohammadi & E. van t Riet & M. N. Weedon & J. V. van
Diabetologia (2010) 53:103 110 DOI 10.1007/s00125-009-1557-7 ARTICLE Genetic association analysis of LARS2 with type 2 diabetes E. Reiling & B. Jafar-Mohammadi & E. van t Riet & M. N. Weedon & J. V. van
UNIVERSITY OF CALIFORNIA, LOS ANGELES
 UNIVERSITY OF CALIFORNIA, LOS ANGELES BERKELEY DAVIS IRVINE LOS ANGELES MERCED RIVERSIDE SAN DIEGO SAN FRANCISCO UCLA SANTA BARBARA SANTA CRUZ DEPARTMENT OF EPIDEMIOLOGY SCHOOL OF PUBLIC HEALTH CAMPUS
UNIVERSITY OF CALIFORNIA, LOS ANGELES BERKELEY DAVIS IRVINE LOS ANGELES MERCED RIVERSIDE SAN DIEGO SAN FRANCISCO UCLA SANTA BARBARA SANTA CRUZ DEPARTMENT OF EPIDEMIOLOGY SCHOOL OF PUBLIC HEALTH CAMPUS
Lack of association between IL-6-174G>C polymorphism and lung cancer: a metaanalysis
 Lack of association between IL-6-174G>C polymorphism and lung cancer: a metaanalysis Y. Liu, X.L. Song, G.L. Zhang, A.M. Peng, P.F. Fu, P. Li, M. Tan, X. Li, M. Li and C.H. Wang Department of Respiratory
Lack of association between IL-6-174G>C polymorphism and lung cancer: a metaanalysis Y. Liu, X.L. Song, G.L. Zhang, A.M. Peng, P.F. Fu, P. Li, M. Tan, X. Li, M. Li and C.H. Wang Department of Respiratory
Supplementary Figure S1A
 Supplementary Figure S1A-G. LocusZoom regional association plots for the seven new cross-cancer loci that were > 1 Mb from known index SNPs. Genes up to 500 kb on either side of each new index SNP are
Supplementary Figure S1A-G. LocusZoom regional association plots for the seven new cross-cancer loci that were > 1 Mb from known index SNPs. Genes up to 500 kb on either side of each new index SNP are
Does the Aspartic Acid to Asparagine Substitution at Position 76 in the Pancreas Duodenum Homeobox Gene (PDX1) Cause Late-Onset Type 2 Diabetes?
 Pathophysiology/Complications O R I G I N A L A R T I C L E Does the Aspartic Acid to Asparagine Substitution at Position 76 in the Pancreas Duodenum Homeobox Gene (PDX1) Cause Late-Onset Type 2 Diabetes?
Pathophysiology/Complications O R I G I N A L A R T I C L E Does the Aspartic Acid to Asparagine Substitution at Position 76 in the Pancreas Duodenum Homeobox Gene (PDX1) Cause Late-Onset Type 2 Diabetes?
Field wide development of analytic approaches for sequence data
 Benjamin Neale Field wide development of analytic approaches for sequence data Cohort Allelic Sum Test (CAST; Hobbs, Cohen and others) Li and Leal (AJHG) Madsen and Browning (PLoS Genetics) C alpha and
Benjamin Neale Field wide development of analytic approaches for sequence data Cohort Allelic Sum Test (CAST; Hobbs, Cohen and others) Li and Leal (AJHG) Madsen and Browning (PLoS Genetics) C alpha and
Summary. Introduction. Atypical and Duplicated Samples. Atypical Samples. Noah A. Rosenberg
 doi: 10.1111/j.1469-1809.2006.00285.x Standardized Subsets of the HGDP-CEPH Human Genome Diversity Cell Line Panel, Accounting for Atypical and Duplicated Samples and Pairs of Close Relatives Noah A. Rosenberg
doi: 10.1111/j.1469-1809.2006.00285.x Standardized Subsets of the HGDP-CEPH Human Genome Diversity Cell Line Panel, Accounting for Atypical and Duplicated Samples and Pairs of Close Relatives Noah A. Rosenberg
Supplementary Figures
 Supplementary Figures Supplementary Fig 1. Comparison of sub-samples on the first two principal components of genetic variation. TheBritishsampleisplottedwithredpoints.The sub-samples of the diverse sample
Supplementary Figures Supplementary Fig 1. Comparison of sub-samples on the first two principal components of genetic variation. TheBritishsampleisplottedwithredpoints.The sub-samples of the diverse sample
The Role of HNF4A Variants in the Risk of Type 2 Diabetes
 The Role of HNF4A Variants in the Risk of Type 2 Diabetes Karen L. Mohlke, PhD,* and Michael Boehnke, PhD Address *Department of Genetics, University of North Carolina, 103 Mason Farm Drive, CB 7264, Chapel
The Role of HNF4A Variants in the Risk of Type 2 Diabetes Karen L. Mohlke, PhD,* and Michael Boehnke, PhD Address *Department of Genetics, University of North Carolina, 103 Mason Farm Drive, CB 7264, Chapel
Tutorial on Genome-Wide Association Studies
 Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
Common variants in WFS1 confer risk of type 2 diabetes
 Europe PMC Funders Group Author Manuscript Published in final edited form as: Nat Genet. 2007 August ; 39(8): 951 953. doi:10.1038/ng2067. Common variants in WFS1 confer risk of type 2 diabetes Manjinder
Europe PMC Funders Group Author Manuscript Published in final edited form as: Nat Genet. 2007 August ; 39(8): 951 953. doi:10.1038/ng2067. Common variants in WFS1 confer risk of type 2 diabetes Manjinder
Genome-wide Association Analysis Applied to Asthma-Susceptibility Gene. McCaw, Z., Wu, W., Hsiao, S., McKhann, A., Tracy, S.
 Genome-wide Association Analysis Applied to Asthma-Susceptibility Gene McCaw, Z., Wu, W., Hsiao, S., McKhann, A., Tracy, S. December 17, 2014 1 Introduction Asthma is a chronic respiratory disease affecting
Genome-wide Association Analysis Applied to Asthma-Susceptibility Gene McCaw, Z., Wu, W., Hsiao, S., McKhann, A., Tracy, S. December 17, 2014 1 Introduction Asthma is a chronic respiratory disease affecting
Association between interleukin-17a polymorphism and coronary artery disease susceptibility in the Chinese Han population
 Association between interleukin-17a polymorphism and coronary artery disease susceptibility in the Chinese Han population G.B. Su, X.L. Guo, X.C. Liu, Q.T. Cui and C.Y. Zhou Department of Cardiothoracic
Association between interleukin-17a polymorphism and coronary artery disease susceptibility in the Chinese Han population G.B. Su, X.L. Guo, X.C. Liu, Q.T. Cui and C.Y. Zhou Department of Cardiothoracic
Role of the FOXC2-512C>T polymorphism in type 2 diabetes: possible association with the dysmetabolic syndrome.
 Role of the FOXC2-512C>T polymorphism in type 2 diabetes: possible association with the dysmetabolic syndrome. Nilsson, Emma A; Groop, Leif; Ridderstråle, Martin Published in: DOI: 10.1038/sj.ijo.0802876
Role of the FOXC2-512C>T polymorphism in type 2 diabetes: possible association with the dysmetabolic syndrome. Nilsson, Emma A; Groop, Leif; Ridderstråle, Martin Published in: DOI: 10.1038/sj.ijo.0802876
Lack of association of IL-2RA and IL-2RB polymorphisms with rheumatoid arthritis in a Han Chinese population
 Lack of association of IL-2RA and IL-2RB polymorphisms with rheumatoid arthritis in a Han Chinese population J. Zhu 1 *, F. He 2 *, D.D. Zhang 2 *, J.Y. Yang 2, J. Cheng 1, R. Wu 1, B. Gong 2, X.Q. Liu
Lack of association of IL-2RA and IL-2RB polymorphisms with rheumatoid arthritis in a Han Chinese population J. Zhu 1 *, F. He 2 *, D.D. Zhang 2 *, J.Y. Yang 2, J. Cheng 1, R. Wu 1, B. Gong 2, X.Q. Liu
Global variation in copy number in the human genome
 Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Insulin resistance is a major feature of type 2 diabetes
 BRIEF REPORT The ENPP1 K121Q Polymorphism Is Associated With Type 2 Diabetes in European Populations Evidence From an Updated Meta-Analysis in 42,042 Subjects Jarred B. McAteer, 1,2 Sabrina Prudente, 3,4
BRIEF REPORT The ENPP1 K121Q Polymorphism Is Associated With Type 2 Diabetes in European Populations Evidence From an Updated Meta-Analysis in 42,042 Subjects Jarred B. McAteer, 1,2 Sabrina Prudente, 3,4
ARTICLE. A. Dahlgren & B. Zethelius & K. Jensevik & A.-C. Syvänen & C. Berne
 Diabetologia (2007) 50:1852 1857 DOI 10.1007/s00125-007-0746-5 ARTICLE Variants of the TCF7L2 gene are associated with beta cell dysfunction and confer an increased risk of type 2 diabetes mellitus in
Diabetologia (2007) 50:1852 1857 DOI 10.1007/s00125-007-0746-5 ARTICLE Variants of the TCF7L2 gene are associated with beta cell dysfunction and confer an increased risk of type 2 diabetes mellitus in
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Genetics and Pharmacogenetics in Human Complex Disorders (Example of Bipolar Disorder)
 Genetics and Pharmacogenetics in Human Complex Disorders (Example of Bipolar Disorder) September 14, 2012 Chun Xu M.D, M.Sc, Ph.D. Assistant professor Texas Tech University Health Sciences Center Paul
Genetics and Pharmacogenetics in Human Complex Disorders (Example of Bipolar Disorder) September 14, 2012 Chun Xu M.D, M.Sc, Ph.D. Assistant professor Texas Tech University Health Sciences Center Paul
Using ancestry estimates as tools to better understand group or individual differences in disease risk or disease outcomes
 Using ancestry estimates as tools to better understand group or individual differences in disease risk or disease outcomes Jill Barnholtz-Sloan, Ph.D. Assistant Professor Case Comprehensive Cancer Center
Using ancestry estimates as tools to better understand group or individual differences in disease risk or disease outcomes Jill Barnholtz-Sloan, Ph.D. Assistant Professor Case Comprehensive Cancer Center
GENOME-WIDE ASSOCIATION STUDIES
 GENOME-WIDE ASSOCIATION STUDIES SUCCESSES AND PITFALLS IBT 2012 Human Genetics & Molecular Medicine Zané Lombard IDENTIFYING DISEASE GENES??? Nature, 15 Feb 2001 Science, 16 Feb 2001 IDENTIFYING DISEASE
GENOME-WIDE ASSOCIATION STUDIES SUCCESSES AND PITFALLS IBT 2012 Human Genetics & Molecular Medicine Zané Lombard IDENTIFYING DISEASE GENES??? Nature, 15 Feb 2001 Science, 16 Feb 2001 IDENTIFYING DISEASE
Genetics and Genomics in Medicine Chapter 8 Questions
 Genetics and Genomics in Medicine Chapter 8 Questions Linkage Analysis Question Question 8.1 Affected members of the pedigree above have an autosomal dominant disorder, and cytogenetic analyses using conventional
Genetics and Genomics in Medicine Chapter 8 Questions Linkage Analysis Question Question 8.1 Affected members of the pedigree above have an autosomal dominant disorder, and cytogenetic analyses using conventional
The angiotensin-converting enzyme (ACE) I/D polymorphism in Parkinson s disease
 4432JRA0010.1177/1470320313494432Journal of the Renin-Angiotensin-Aldosterone SystemSu et al Original Article The angiotensin-converting enzyme (ACE) I/D polymorphism in Parkinson s disease Journal of
4432JRA0010.1177/1470320313494432Journal of the Renin-Angiotensin-Aldosterone SystemSu et al Original Article The angiotensin-converting enzyme (ACE) I/D polymorphism in Parkinson s disease Journal of
Imaging Genetics: Heritability, Linkage & Association
 Imaging Genetics: Heritability, Linkage & Association David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July 17, 2011 Memory Activation & APOE ε4 Risk
Imaging Genetics: Heritability, Linkage & Association David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July 17, 2011 Memory Activation & APOE ε4 Risk
Sian Ellard, Michael P. Bulman, Timothy M. Frayling, Maggie Shepherd, and Andrew T. Hattersley
 HUMAN MUTATION Mutation in Brief #360 (2000) Online MUTATION IN BRIEF Proposed Mechanism for a Novel Insertion/Deletion Frameshift Mutation (I414G415ATCG CCA) in the Hepatocyte Nuclear Factor 1 Alpha (HNF-1α)
HUMAN MUTATION Mutation in Brief #360 (2000) Online MUTATION IN BRIEF Proposed Mechanism for a Novel Insertion/Deletion Frameshift Mutation (I414G415ATCG CCA) in the Hepatocyte Nuclear Factor 1 Alpha (HNF-1α)
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Meta-analysis of the Gly482Ser variant in PPARGC1A in type 2 diabetes and related phenotypes
 Diabetologia (2006) 49: 501 505 DOI 10.1007/s00125-005-0130-2 SHORT COMMUNICATION I. Barroso. J. Luan. M. S. Sandhu. P. W. Franks. V. Crowley. A. J. Schafer. S. O Rahilly. N. J. Wareham Meta-analysis of
Diabetologia (2006) 49: 501 505 DOI 10.1007/s00125-005-0130-2 SHORT COMMUNICATION I. Barroso. J. Luan. M. S. Sandhu. P. W. Franks. V. Crowley. A. J. Schafer. S. O Rahilly. N. J. Wareham Meta-analysis of
Familial aggregation and concordance rates of
 Variants Within the Calpain-10 Gene on Chromosome 2q37 (NIDDM1) and Relationships to Type 2 Diabetes, Insulin Resistance, and Impaired Acute Insulin Secretion Among Scandinavian Caucasians Søren K. Rasmussen,
Variants Within the Calpain-10 Gene on Chromosome 2q37 (NIDDM1) and Relationships to Type 2 Diabetes, Insulin Resistance, and Impaired Acute Insulin Secretion Among Scandinavian Caucasians Søren K. Rasmussen,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13908 Supplementary Tables Supplementary Table 1: Families in this study (.xlsx) All families included in the study are listed. For each family, we show: the genders of the probands and
doi:10.1038/nature13908 Supplementary Tables Supplementary Table 1: Families in this study (.xlsx) All families included in the study are listed. For each family, we show: the genders of the probands and
CONTENT SUPPLEMENTARY FIGURE E. INSTRUMENTAL VARIABLE ANALYSIS USING DESEASONALISED PLASMA 25-HYDROXYVITAMIN D. 7
 CONTENT FIGURES 3 SUPPLEMENTARY FIGURE A. NUMBER OF PARTICIPANTS AND EVENTS IN THE OBSERVATIONAL AND GENETIC ANALYSES. 3 SUPPLEMENTARY FIGURE B. FLOWCHART SHOWING THE SELECTION PROCESS FOR DETERMINING
CONTENT FIGURES 3 SUPPLEMENTARY FIGURE A. NUMBER OF PARTICIPANTS AND EVENTS IN THE OBSERVATIONAL AND GENETIC ANALYSES. 3 SUPPLEMENTARY FIGURE B. FLOWCHART SHOWING THE SELECTION PROCESS FOR DETERMINING
Pooled analysis of association between a genetic variant in the 3'-untranslated region of Toll-like receptor 4 and cancer risk
 Pooled analysis of association between a genetic variant in the 3'-untranslated region of Toll-like receptor 4 and cancer risk X. Wang*, Z. Xu* and C.H. Miao Department of Anesthesiology, Shanghai Cancer
Pooled analysis of association between a genetic variant in the 3'-untranslated region of Toll-like receptor 4 and cancer risk X. Wang*, Z. Xu* and C.H. Miao Department of Anesthesiology, Shanghai Cancer
ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES. Irina Durbală
 ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES Summary Irina Durbală CELL AND MOLECULAR BIOLOGY DEPARTMENT FACULTY OF MEDICINE, OVIDIUS UNIVERSITY CONSTANŢA Class II
ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES Summary Irina Durbală CELL AND MOLECULAR BIOLOGY DEPARTMENT FACULTY OF MEDICINE, OVIDIUS UNIVERSITY CONSTANŢA Class II
Diabetologia 9 Springer-Verlag 1993
 Diabetologia (1993) 36:234-238 Diabetologia 9 Springer-Verlag 1993 HLA-associated susceptibility to Type 2 (non-insulin-dependent) diabetes mellitus: the Wadena City Health Study S. S. Rich 1, L. R. French
Diabetologia (1993) 36:234-238 Diabetologia 9 Springer-Verlag 1993 HLA-associated susceptibility to Type 2 (non-insulin-dependent) diabetes mellitus: the Wadena City Health Study S. S. Rich 1, L. R. French
Biostatistics Faculty Publications
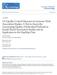 University of Kentucky UKnowledge Biostatistics Faculty Publications Biostatistics 7-24-2009 On Quality Control Measures in Genome-Wide Association Studies: A Test to Assess the Genotyping Quality of Individual
University of Kentucky UKnowledge Biostatistics Faculty Publications Biostatistics 7-24-2009 On Quality Control Measures in Genome-Wide Association Studies: A Test to Assess the Genotyping Quality of Individual
Relationship between vitamin D (1,25-dihydroxyvitamin D3) receptor gene polymorphisms and primary biliary cirrhosis risk: a meta-analysis
 Relationship between vitamin D (1,25-dihydroxyvitamin D3) receptor gene polymorphisms and primary biliary cirrhosis risk: a meta-analysis F. Fang, J. Wang, J. Pan, G.H. Su, L.X. Xu and G. Li Institute
Relationship between vitamin D (1,25-dihydroxyvitamin D3) receptor gene polymorphisms and primary biliary cirrhosis risk: a meta-analysis F. Fang, J. Wang, J. Pan, G.H. Su, L.X. Xu and G. Li Institute
Recently, a DNA sequence variant of the transcription
 Brief Report Haplotypes of Transcription Factor 7 Like 2 (TCF7L2) Gene and Its Upstream Region Are Associated With Type 2 Diabetes and Age of Onset in Mexican Americans Donna M. Lehman, 1 Kelly J. Hunt,
Brief Report Haplotypes of Transcription Factor 7 Like 2 (TCF7L2) Gene and Its Upstream Region Are Associated With Type 2 Diabetes and Age of Onset in Mexican Americans Donna M. Lehman, 1 Kelly J. Hunt,
Whole-genome detection of disease-associated deletions or excess homozygosity in a case control study of rheumatoid arthritis
 HMG Advance Access published December 21, 2012 Human Molecular Genetics, 2012 1 13 doi:10.1093/hmg/dds512 Whole-genome detection of disease-associated deletions or excess homozygosity in a case control
HMG Advance Access published December 21, 2012 Human Molecular Genetics, 2012 1 13 doi:10.1093/hmg/dds512 Whole-genome detection of disease-associated deletions or excess homozygosity in a case control
A total of 2,822 Mexican dyslipidemic cases and controls were recruited at INCMNSZ in
 Supplemental Material The N342S MYLIP polymorphism is associated with high total cholesterol and increased LDL-receptor degradation in humans by Daphna Weissglas-Volkov et al. Supplementary Methods Mexican
Supplemental Material The N342S MYLIP polymorphism is associated with high total cholesterol and increased LDL-receptor degradation in humans by Daphna Weissglas-Volkov et al. Supplementary Methods Mexican
CDKAL1 and type 2 diabetes: a global meta-analysis
 CDKAL1 and type 2 diabetes: a global meta-analysis M.A.S. Dehwah, M. Wang and Q.-Y. Huang Hubei Key Lab of Genetic Regulation and Integrative Biology, College of Life Sciences, Huazhong Normal University,
CDKAL1 and type 2 diabetes: a global meta-analysis M.A.S. Dehwah, M. Wang and Q.-Y. Huang Hubei Key Lab of Genetic Regulation and Integrative Biology, College of Life Sciences, Huazhong Normal University,
MULTIFACTORIAL DISEASES. MG L-10 July 7 th 2014
 MULTIFACTORIAL DISEASES MG L-10 July 7 th 2014 Genetic Diseases Unifactorial Chromosomal Multifactorial AD Numerical AR Structural X-linked Microdeletions Mitochondrial Spectrum of Alterations in DNA Sequence
MULTIFACTORIAL DISEASES MG L-10 July 7 th 2014 Genetic Diseases Unifactorial Chromosomal Multifactorial AD Numerical AR Structural X-linked Microdeletions Mitochondrial Spectrum of Alterations in DNA Sequence
Association between the interleukin-1β gene -511C/T polymorphism and ischemic stroke: an updated meta-analysis
 Association between the interleukin-1β gene -511C/T polymorphism and ischemic stroke: an updated meta-analysis W. Yan, Z.Y. Chen, J.Q. Chen and H.M. Chen Department of Neurology, Ningbo No. 2 Hospital,
Association between the interleukin-1β gene -511C/T polymorphism and ischemic stroke: an updated meta-analysis W. Yan, Z.Y. Chen, J.Q. Chen and H.M. Chen Department of Neurology, Ningbo No. 2 Hospital,
Supplementary Online Content
 Supplementary Online Content Lotta LA, Stewart ID, Sharp SJ, et al. Association of genetically enhanced lipoprotein lipase mediated lipolysis and low-density lipoprotein cholesterol lowering alleles with
Supplementary Online Content Lotta LA, Stewart ID, Sharp SJ, et al. Association of genetically enhanced lipoprotein lipase mediated lipolysis and low-density lipoprotein cholesterol lowering alleles with
Genetic variants on 17q21 are associated with asthma in a Han Chinese population
 Genetic variants on 17q21 are associated with asthma in a Han Chinese population F.-X. Li 1 *, J.-Y. Tan 2 *, X.-X. Yang 1, Y.-S. Wu 1, D. Wu 3 and M. Li 1 1 School of Biotechnology, Southern Medical University,
Genetic variants on 17q21 are associated with asthma in a Han Chinese population F.-X. Li 1 *, J.-Y. Tan 2 *, X.-X. Yang 1, Y.-S. Wu 1, D. Wu 3 and M. Li 1 1 School of Biotechnology, Southern Medical University,
T he prevalence of type 2 diabetes is higher in populations
 1of9 ELECTRONIC LETTER Relation of type 2 diabetes to individual admixture and candidate gene polymorphisms in the Hispanic American population of San Luis Valley, Colorado E J Parra, C J Hoggart, C Bonilla,
1of9 ELECTRONIC LETTER Relation of type 2 diabetes to individual admixture and candidate gene polymorphisms in the Hispanic American population of San Luis Valley, Colorado E J Parra, C J Hoggart, C Bonilla,
Statistical power and significance testing in large-scale genetic studies
 STUDY DESIGNS Statistical power and significance testing in large-scale genetic studies Pak C. Sham 1 and Shaun M. Purcell 2,3 Abstract Significance testing was developed as an objective method for summarizing
STUDY DESIGNS Statistical power and significance testing in large-scale genetic studies Pak C. Sham 1 and Shaun M. Purcell 2,3 Abstract Significance testing was developed as an objective method for summarizing
Chapter 2. Linkage Analysis. JenniferH.BarrettandM.DawnTeare. Abstract. 1. Introduction
 Chapter 2 Linkage Analysis JenniferH.BarrettandM.DawnTeare Abstract Linkage analysis is used to map genetic loci using observations on relatives. It can be applied to both major gene disorders (parametric
Chapter 2 Linkage Analysis JenniferH.BarrettandM.DawnTeare Abstract Linkage analysis is used to map genetic loci using observations on relatives. It can be applied to both major gene disorders (parametric
New evidence of TERT rs polymorphism and cancer risk: an updated meta-analysis
 JBUON 2016; 21(2): 491-497 ISSN: 1107-0625, online ISSN: 2241-6293 www.jbuon.com E-mail: editorial_office@jbuon.com ORIGINAL ARTICLE New evidence of TERT rs2736098 polymorphism and risk: an updated meta-analysis
JBUON 2016; 21(2): 491-497 ISSN: 1107-0625, online ISSN: 2241-6293 www.jbuon.com E-mail: editorial_office@jbuon.com ORIGINAL ARTICLE New evidence of TERT rs2736098 polymorphism and risk: an updated meta-analysis
Common obesity-related genetic variants and papillary thyroid cancer risk
 Common obesity-related genetic variants and papillary thyroid cancer risk Cari M. Kitahara 1, Gila Neta 1, Ruth M. Pfeiffer 1, Deukwoo Kwon 2, Li Xu 3, Neal D. Freedman 1, Amy A. Hutchinson 4, Stephen
Common obesity-related genetic variants and papillary thyroid cancer risk Cari M. Kitahara 1, Gila Neta 1, Ruth M. Pfeiffer 1, Deukwoo Kwon 2, Li Xu 3, Neal D. Freedman 1, Amy A. Hutchinson 4, Stephen
Association between the 8q24 rs T/G polymorphism and prostate cancer risk: a meta-analysis
 Association between the 8q24 rs6983267 T/G polymorphism and prostate cancer risk: a meta-analysis H.S. Zhu 1 *, J.F. Zhang 1 *, J.D. Zhou 2, M.J. Zhang 1 and H.X. Hu 1 1 Clinical Laboratory, East Suzhou
Association between the 8q24 rs6983267 T/G polymorphism and prostate cancer risk: a meta-analysis H.S. Zhu 1 *, J.F. Zhang 1 *, J.D. Zhou 2, M.J. Zhang 1 and H.X. Hu 1 1 Clinical Laboratory, East Suzhou
The genetic architecture of type 2 diabetes appears
 ORIGINAL ARTICLE A 100K Genome-Wide Association Scan for Diabetes and Related Traits in the Framingham Heart Study Replication and Integration With Other Genome-Wide Datasets Jose C. Florez, 1,2,3 Alisa
ORIGINAL ARTICLE A 100K Genome-Wide Association Scan for Diabetes and Related Traits in the Framingham Heart Study Replication and Integration With Other Genome-Wide Datasets Jose C. Florez, 1,2,3 Alisa
Functional Variation in VEGF Is not Associated with Type 2 Diabetes in a United Kingdom Caucasian Population
 ORIGINAL ARTICLE Functional Variation in VEGF Is not Associated with Type 2 Diabetes in a United Kingdom Caucasian Population Rachel M Freathy 1, Michael N Weedon 1, Beverley Shields 1, Graham A Hitman
ORIGINAL ARTICLE Functional Variation in VEGF Is not Associated with Type 2 Diabetes in a United Kingdom Caucasian Population Rachel M Freathy 1, Michael N Weedon 1, Beverley Shields 1, Graham A Hitman
A Large Sample of Finnish Diabetic Sib-Pairs Reveals No Evidence for a Non Insulin-dependent Diabetes Mellitus Susceptibility Locus at 2qter
 A Large Sample of Finnish Diabetic Sib-Pairs Reveals No Evidence for a Non Insulin-dependent Diabetes Mellitus Susceptibility Locus at 2qter Soumitra Ghosh,* Elizabeth R. Hauser, Victoria L. Magnuson,*
A Large Sample of Finnish Diabetic Sib-Pairs Reveals No Evidence for a Non Insulin-dependent Diabetes Mellitus Susceptibility Locus at 2qter Soumitra Ghosh,* Elizabeth R. Hauser, Victoria L. Magnuson,*
Supplementary Online Content
 Supplementary Online Content Hartwig FP, Borges MC, Lessa Horta B, Bowden J, Davey Smith G. Inflammatory biomarkers and risk of schizophrenia: a 2-sample mendelian randomization study. JAMA Psychiatry.
Supplementary Online Content Hartwig FP, Borges MC, Lessa Horta B, Bowden J, Davey Smith G. Inflammatory biomarkers and risk of schizophrenia: a 2-sample mendelian randomization study. JAMA Psychiatry.
Exclusion of Polymorphisms in Carnosinase Genes (CNDP1 and CNDP2) as a Cause of Diabetic Nephropathy in Type 1 Diabetes
 Exclusion of Polymorphisms in Carnosinase Genes (CNDP1 and CNDP2) as a Cause of Diabetic Nephropathy in Type 1 Diabetes The Harvard community has made this article openly available. Please share how this
Exclusion of Polymorphisms in Carnosinase Genes (CNDP1 and CNDP2) as a Cause of Diabetic Nephropathy in Type 1 Diabetes The Harvard community has made this article openly available. Please share how this
Letter to the Editor. Association of TCF7L2 and GCG Gene Variants with Insulin Secretion, Insulin Resistance, and Obesity in New-onset Diabetes *
 814 Biomed Environ Sci, 2016; 29(11): 814-817 Letter to the Editor Association of TCF7L2 and GCG Gene Variants with Insulin Secretion, Insulin Resistance, and Obesity in New-onset Diabetes * ZHANG Lu 1,^,
814 Biomed Environ Sci, 2016; 29(11): 814-817 Letter to the Editor Association of TCF7L2 and GCG Gene Variants with Insulin Secretion, Insulin Resistance, and Obesity in New-onset Diabetes * ZHANG Lu 1,^,
Statistical Genetics : Gene Mappin g through Linkag e and Associatio n
 Statistical Genetics : Gene Mappin g through Linkag e and Associatio n Benjamin M Neale Manuel AR Ferreira Sarah E Medlan d Danielle Posthuma About the editors List of contributors Preface Acknowledgements
Statistical Genetics : Gene Mappin g through Linkag e and Associatio n Benjamin M Neale Manuel AR Ferreira Sarah E Medlan d Danielle Posthuma About the editors List of contributors Preface Acknowledgements
Association between FTO, MC4R, SLC30A8, and KCNQ1 gene variants and type 2 diabetes in Saudi population
 Association between FTO, MC4R, SLC30A8, and KCNQ1 gene variants and type 2 diabetes in Saudi population M.D. Bazzi 1, F.A. Nasr 1, M.S. Alanazi 1, A. Alamri 1, A.A. Turjoman 2, A.S. Moustafa 2, A.A. Alfadda
Association between FTO, MC4R, SLC30A8, and KCNQ1 gene variants and type 2 diabetes in Saudi population M.D. Bazzi 1, F.A. Nasr 1, M.S. Alanazi 1, A. Alamri 1, A.A. Turjoman 2, A.S. Moustafa 2, A.A. Alfadda
New Enhancements: GWAS Workflows with SVS
 New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
Corresponding author: F.Q. Wen
 Association of a disintegrin and metalloproteinase 33 (ADAM33) gene polymorphisms with chronic obstructive pulmonary disease in the Chinese population: A meta-analysis D.D. Li 1,2, S.J. Guo 1,2, L.Q. Jia
Association of a disintegrin and metalloproteinase 33 (ADAM33) gene polymorphisms with chronic obstructive pulmonary disease in the Chinese population: A meta-analysis D.D. Li 1,2, S.J. Guo 1,2, L.Q. Jia
A role for coding functional variants in HNF4A in type 2 diabetes susceptibility
 Diabetologia (2011) 54:111 119 DOI 10.1007/s00125-010-1916-4 ARTICLE A role for coding functional variants in HNF4A in type 2 diabetes susceptibility B. Jafar-Mohammadi & C. J. Groves & A. P. Gjesing &
Diabetologia (2011) 54:111 119 DOI 10.1007/s00125-010-1916-4 ARTICLE A role for coding functional variants in HNF4A in type 2 diabetes susceptibility B. Jafar-Mohammadi & C. J. Groves & A. P. Gjesing &
Systems of Mating: Systems of Mating:
 8/29/2 Systems of Mating: the rules by which pairs of gametes are chosen from the local gene pool to be united in a zygote with respect to a particular locus or genetic system. Systems of Mating: A deme
8/29/2 Systems of Mating: the rules by which pairs of gametes are chosen from the local gene pool to be united in a zygote with respect to a particular locus or genetic system. Systems of Mating: A deme
Mendelian Inheritance. Jurg Ott Columbia and Rockefeller Universities New York
 Mendelian Inheritance Jurg Ott Columbia and Rockefeller Universities New York Genes Mendelian Inheritance Gregor Mendel, monk in a monastery in Brünn (now Brno in Czech Republic): Breeding experiments
Mendelian Inheritance Jurg Ott Columbia and Rockefeller Universities New York Genes Mendelian Inheritance Gregor Mendel, monk in a monastery in Brünn (now Brno in Czech Republic): Breeding experiments
An Introduction to Quantitative Genetics I. Heather A Lawson Advanced Genetics Spring2018
 An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
An association analysis of the HLA gene region in latent autoimmune diabetes in adults
 Diabetologia (2007) 50:68 73 DOI 10.1007/s00125-006-0513-z SHORT COMMUNICATION An association analysis of the HLA gene region in latent autoimmune diabetes in adults M. Desai & E. Zeggini & V. A. Horton
Diabetologia (2007) 50:68 73 DOI 10.1007/s00125-006-0513-z SHORT COMMUNICATION An association analysis of the HLA gene region in latent autoimmune diabetes in adults M. Desai & E. Zeggini & V. A. Horton
SNPrints: Defining SNP signatures for prediction of onset in complex diseases
 SNPrints: Defining SNP signatures for prediction of onset in complex diseases Linda Liu, Biomedical Informatics, Stanford University Daniel Newburger, Biomedical Informatics, Stanford University Grace
SNPrints: Defining SNP signatures for prediction of onset in complex diseases Linda Liu, Biomedical Informatics, Stanford University Daniel Newburger, Biomedical Informatics, Stanford University Grace
