Regulating Transmembrane Signaling Through Plexin- Neuropilin Oligomerization
|
|
|
- Jack Ramsey
- 5 years ago
- Views:
Transcription
1 Regulating Transmembrane Signaling Through Plexin- Neuropilin Oligomerization Bryan W. Berger Department of Chemical Engineering Program in Bioengineering
2 Components of Cell Membranes Approximately 30-40% by volume of membrane is occupied by membrane proteins IMPs account for nearly 30% of the human genome, yet less than 1% of available protein structures in the Protein Data Bank (PDB)
3 Integral Membrane Proteins: The Next Frontier of Structural Biology Integral membrane proteins (IMPs) are central to a wide range of membrane-mediated processes such as signal transduction, fusion and transport MC Wiener (2004) Methods (Review) 34, 364
4 A Bottom-Up Chemical Engineering Process Ethanol separation from a binary mixture: H-O-H UNIQUAC: C chemical (VdW SA, V) R residual (empirical, experimental) Use molecular properties to estimate thermodynamic parameters Apply relevant models to predict phase behavior of system (binary or otherwise) Design equipment to separate components
5 A Bottom-Up Biological Process Platelet activation:
6 Proposed Cx43-Sema3d Pathway Fin length Segment length Cell proliferation Cx43 levels Sof Decreased Decreased Decreased Decreased Alf Increased Increased Increased Increased
7 Membrane Co-Receptor Oligomerization and Signaling: Neuropilins, Plexins and Vascular Endothelial Growth Factor Receptors Neurogenesis (Axon Guidance) Angiogenesis (Cell Proliferation)
8 Signal Transduction as an On-Off Switch OFF: - gate, - light ON: + gate, + light
9 Ligand Binding and Dimerization: 2 On-Off Switches During Signal Trasduction Sema 3D Plexin A3 OFF: - ligand - dimer - signal OFF: - ligand + dimer - signal ON: + ligand + dimer + signal
10 Ligand Binding and Dimerization: 2 On-Off Switches During Signal Trasduction Plexin A3 dimerizatio n Sema 3D binding + = Reduced Joint formation OFF: - ligand - dimer - signal OFF: - ligand + dimer - signal ON: + ligand + dimer + signal
11 What is a Dimer? A complex formed by 2 individual protein molecules, which are typically bound through non-covalent association Homodimer: A + A = A 2 Heterodimer: A + B = AB
12 How Do Proteins Dimerize? A protein dimer depends on its quarternary structure, and is stabilized through molecular interactions between each protein monomer A protein domain is a region within a protein that has a defined secondary or tertiary structure
13 Identifying individual domains within plexins and neuropilins Nrp2a PlexinA3 Sema3d MAM = conserved juxtamembrane domain CYTO CYTO MBP-Nrp2a-MAM MBP-Nrp2a-CYTO MBP-PlexinA3 CYTO GST-Sema3d His-Sema3d MBP-Sema3d
14 Linking Primary Sequence Patterns to Protein Structure: Domains and Motifs Proteins are comprised of amino acids, and the chemical identity of each amino acid in a protein sequence determines its secondary and tertiary structure There are 20 standard amino acids
15 Protein Sequence Probability: A Coin Toss Probability (P) = Number of possible outcomes Total number of outcomes P = 0.5 P = 0.5 x 0.5 x 0.5 = (0.5) 3
16 Linking Sequence Patterns to Protein Structure Plexin A3 Transmembrane Sequence AIIGIGGGGGVLLIAIIAVLIA Glycine (G): P(G) = 1/20 P(GGGGG) = (1/20) 5 = 3 x 10-7 But this is true for any 5-mer sequence, not just GGGGG! So how do we know this is actually a unique sequence motif??? We instead use a positional sampling approach (Gibbs Sampling Algorithim) that compares the motif across a family of sequences
17 gi MGSAAILSRQSPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi SP--LGRQQPPHPKR--PFITFTGDQTEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi TVSP--LGRQQPPHSKR--PFITFTGDQTEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQSASLSQK-LSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi MGSAALLSRQPPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi TVSP--LGRQQPPHPKR--PFITFTGEQAEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi LGSSMLLPRQPSPLSQK-RSFITFRGEPTEG-FNHLVVDERTGHIYLGAINRIYKLSSDL gi MGSSTLLPRQSASLSQK-LSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLSRQPPPLSQK-RSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi MGSSTLLPRQPPQMSQK-PSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPPSQKQRSFVTFQGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSAALLSRQPPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi LGSSMLLPRQPSPLSQK-RSFITFRGEPTEG-FNHLVVDERTGHIYLGAINRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLSRQPPPLSQK-RSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi gi VGAVG-----SSRP--FPAFLV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi gi VGAVG-----SSRP--FPAFLV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----NSRP--FRAFMV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----S-RP--FRAFVV--TDTKLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----S-RP--FRAFVV--TDTKLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----N-RP--FRAFMV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL
18 gi MGSAAILSRQSPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi SP--LGRQQPPHPKR--PFITFTGDQTEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi TVSP--LGRQQPPHSKR--PFITFTGDQTEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQSASLSQK-LSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi MGSAALLSRQPPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi TVSP--LGRQQPPHPKR--PFITFTGEQAEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi LGSSMLLPRQPSPLSQK-RSFITFRGEPTEG-FNHLVVDERTGHIYLGAINRIYKLSSDL gi MGSSTLLPRQSASLSQK-LSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLSRQPPPLSQK-RSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi MGSSTLLPRQPPQMSQK-PSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPPSQKQRSFVTFQGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSAALLSRQPPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi LGSSMLLPRQPSPLSQK-RSFITFRGEPTEG-FNHLVVDERTGHIYLGAINRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLSRQPPPLSQK-RSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi gi VGAVG-----SSRP--FPAFLV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi gi VGAVG-----SSRP--FPAFLV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----NSRP--FRAFMV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----S-RP--FRAFVV--TDTKLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----S-RP--FRAFVV--TDTKLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----N-RP--FRAFMV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL
19 gi MGSAAILSRQSPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi SP--LGRQQPPHPKR--PFITFTGDQTEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi TVSP--LGRQQPPHSKR--PFITFTGDQTEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQSASLSQK-LSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi MGSAALLSRQPPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi TVSP--LGRQQPPHPKR--PFITFTGEQAEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi LGSSMLLPRQPSPLSQK-RSFITFRGEPTEG-FNHLVVDERTGHIYLGAINRIYKLSSDL gi MGSSTLLPRQSASLSQK-LSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLSRQPPPLSQK-RSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi MGSSTLLPRQPPQMSQK-PSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL Positional probability: gi MGSSTLLTRQPAPPSQKQRSFVTFQGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi Frequency of MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL amino acid X occurring at position N gi MGSAALLSRQPPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi Probability of finding MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL amino acid by chance at position N gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi LGSSMLLPRQPSPLSQK-RSFITFRGEPTEG-FNHLVVDERTGHIYLGAINRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLSRQPPPLSQK-RSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi gi VGAVG-----SSRP--FPAFLV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi gi VGAVG-----SSRP--FPAFLV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----NSRP--FRAFMV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----S-RP--FRAFVV--TDTKLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----S-RP--FRAFVV--TDTKLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----N-RP--FRAFMV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL
20 gi MGSAAILSRQSPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi SP--LGRQQPPHPKR--PFITFTGDQTEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi TVSP--LGRQQPPHSKR--PFITFTGDQTEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQSASLSQK-LSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi MGSAALLSRQPPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi TVSP--LGRQQPPHPKR--PFITFTGEQAEGNFNHLVVDERTGHIYLGAVNRIYKLSSDL gi LGSSMLLPRQPSPLSQK-RSFITFRGEPTEG-FNHLVVDERTGHIYLGAINRIYKLSSDL gi MGSSTLLPRQSASLSQK-LSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLSRQPPPLSQK-RSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLPRQPPQLSQK-PSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi MGSSTLLPRQPPQMSQK-PSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPPSQKQRSFVTFQGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSAALLSRQPPSSAQK-RSFLTFTAEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPTEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi LGSSMLLPRQPSPLSQK-RSFITFRGEPTEG-FNHLVVDERTGHIYLGAINRIYKLSSDL gi MGSSTLLTRQPAPLSQKQRSFVTFRGESAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi MGSSTLLSRQPPPLSQK-RSFVTFRGEPAEG-FNHLVVDERTGHIYLGAVNRIYKLSSDL gi gi gi VGAVG-----SSRP--FPAFLV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi gi VGAVG-----SSRP--FPAFLV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----NSRP--FRAFMV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----S-RP--FRAFVV--TDTKLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----S-RP--FRAFVV--TDTKLTHLAVHRVTGEVFVGAVNRVFKLAPNL gi GGALG-----N-RP--FRAFMV--TDTTLTHLAVHRVTGEVFVGAVNRVFKLAPNL Higher Lower
21 Plexin A3 TMCY Hypothesis: PlexinA3 interacts with itself and Nrp2a through its CYTO and/or TM domains due to unique sequence motifs found in TM and CYTO. Previous work (our group and (2009) PNAS 106: ) has suggested this region (primarily CYTO) is important for signal transduction. Transmembrane: Coiled-coil: Wild-type: AIIGIGAGGGVLLIAIIAVLIAYKRKTRDADRTLKRLQLQMDNLESRV Glycine-rich region Heptad-repeat Oligomerization Hydrophobic residues pack together
22 Primary (1 0 ) and Secondary (2 0 ) Interactions That Stabilize Coiled-Coil Dimers Rho-associated kinase (PDB 1UIX) a..de.g a..de. KLEHLIENKDRMEDEVKNLNTLQL Hydrophobic (a, d) residues introduce a 1 o interaction to drive dimerization. Strength of association is modulated through 2 o (ionic) interactions
23 PDB 3IG3 Human: ZF: Error prone TMCY mutants Transmembrane: Coiled-coil: GLAAGGGLLLLAITAVLVAYK_RKTQDADRTLKRLQLQMDNLESRVALECKEAFAELQTDINELTNHMDEVQ AIIGIGAGGGVLLIAIIAVLIAYKRKTRDADRTLKRLQLQMDNLESRVALECKEAFAELQTDIQELTNDMDGVK AIIGIGAGGGALLIAIIAELIAYKRKTRDTDRSLKRLQLQMDNLESRVALECKEAFAELQTDIQELTNDMDGVK AIIGIGAGGGVLLIAIIAVLIAYKRKTRDADRTLKRLQLQMDNLESRFALECKEAFAELQTDIQELTNDMDGVK AIIGIGAGGGVLLIAIIAVLIAYKRKTRDADRTLKRLQLQMDNLESRVALECKEAFAELQTYIQELTNDMDGVK AIIGIGAGGGVLLIAIIAVLIAYKRKTRDADRTLKRLQLQMDNLESRVALECKEAFAELQTDIQELSNDMAGVK
24 Receptor Oligomerization in Live Cells Measured by BRET Plexin A3 GFP Rluc GFP Rluc 450 nm 520 nm BRET (heterooligomer) Fusions to C-terminal domains of full-length NRP2 and plexin A3 with complementary donor (Rluc) and acceptor (GFP) Addition of exogenous substrate (coelenterazine) to catalyze resonance energy transfer from Rluc to GFP Efficiency ratio (R) gives a proportionality constant to measure extent of energy transfer, which is proportional to separation distance
25 Signal Transduction as an Dimmer Switch Increasing light
26 Ligand Binding and Dimerization: 2 On-Off Switches During Signal Transduction Sema 3D Plexin A3 OFF: - ligand - dimer - signal OFF: - ligand + dimer - signal ON: + ligand + dimer + signal
27 Receptor Surface Density: A Dimmer Switch to Regulate Signal Intensity During Signal Transduction versus versus Increasing signal
28 Endocytosis Regulate Receptor Surface Density During Signal Transduction
29 Defining the Mechanism of PlexinA3 Activation by Sema3d Iovine/Berger Team BDSI 2012 Presentation Iovine/Berger Team (title) 7/26/2012
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras
 Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
Tyrosine kinases. Cell surface receptors ligand binding. Producer cell RNA. Target cell
 Tyrosine kinases http://msbl.helsinki.fi/tkseminar roducer cell Signaling molecules Receptor binding Signal transduction Target cell Activation of Gene expression RNA Biological responses proliferation,
Tyrosine kinases http://msbl.helsinki.fi/tkseminar roducer cell Signaling molecules Receptor binding Signal transduction Target cell Activation of Gene expression RNA Biological responses proliferation,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Lesson 5 Proteins Levels of Protein Structure
 Lesson 5 Proteins Levels of Protein Structure Primary 1º Structure The primary structure is simply the sequence of amino acids in a protein. Chains of amino acids are written from the amino terminus (N-terminus)
Lesson 5 Proteins Levels of Protein Structure Primary 1º Structure The primary structure is simply the sequence of amino acids in a protein. Chains of amino acids are written from the amino terminus (N-terminus)
Signal-Transduction Cascades - 2. The Phosphoinositide Cascade
 Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
Cbl ubiquitin ligase: Lord of the RINGs
 Cbl ubiquitin ligase: Lord of the RINGs Not just quite interesting - really interesting! A cell must be able to degrade proteins when their activity is no longer required. Many eukaryotic proteins are
Cbl ubiquitin ligase: Lord of the RINGs Not just quite interesting - really interesting! A cell must be able to degrade proteins when their activity is no longer required. Many eukaryotic proteins are
Reading Assignments: Chapter 16: Cell Communication Pgs ; 545 & Figure 16-15; ; work Q-1, 3, 4, 10, 12, 15, 16, 17, 20 & 23
 Biol 205 Signal Transduction, the Social Contract and Rogue Cancer Cells Inside cancer web site http://www.insidecancer.org/ National Cancer Institute http://www.cancer.gov/cancerinfo/ Reading Assignments:
Biol 205 Signal Transduction, the Social Contract and Rogue Cancer Cells Inside cancer web site http://www.insidecancer.org/ National Cancer Institute http://www.cancer.gov/cancerinfo/ Reading Assignments:
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression
 Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression Question No. 1 of 10 1. Which statement about cell signaling is correct? Question #1 (A) Cell signaling involves receiving
Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression Question No. 1 of 10 1. Which statement about cell signaling is correct? Question #1 (A) Cell signaling involves receiving
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Signaling. Dr. Sujata Persad Katz Group Centre for Pharmacy & Health research
 Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Ras and Cell Signaling Exercise
 Ras and Cell Signaling Exercise Learning Objectives In this exercise, you will use, a protein 3D- viewer, to explore: the structure of the Ras protein the active and inactive state of Ras and the amino
Ras and Cell Signaling Exercise Learning Objectives In this exercise, you will use, a protein 3D- viewer, to explore: the structure of the Ras protein the active and inactive state of Ras and the amino
Antigen Recognition by T cells
 Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
LQB383 Testbank. Week 8 Cell Communication and Signaling Mechanisms
 LQB383 Testbank Week 8 Cell Communication and Signaling Mechanisms Terms to learn match the terms to the definitions --------------------------------------------------------------------------------------------------------------------------
LQB383 Testbank Week 8 Cell Communication and Signaling Mechanisms Terms to learn match the terms to the definitions --------------------------------------------------------------------------------------------------------------------------
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Sheet #5 Dr. Mamoun Ahram 8/7/2014
 P a g e 1 Protein Structure Quick revision - Levels of protein structure: primary, secondary, tertiary & quaternary. - Primary structure is the sequence of amino acids residues. It determines the other
P a g e 1 Protein Structure Quick revision - Levels of protein structure: primary, secondary, tertiary & quaternary. - Primary structure is the sequence of amino acids residues. It determines the other
P450 CYCLE. All P450s follow the same catalytic cycle of;
 P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
1 (b) (i) ACCEPT how long it took. time / time taken ; PhysicsAndMathsTutor.com
 Question Answer Mark Guidance 1 (a) (works) outside cells ; 1 ACCEPT secreted / AW, from cells ACCEPT works in named extracellular environment e.g. digestive tract IGNORE doesn t work in cells 1 (b) (i)
Question Answer Mark Guidance 1 (a) (works) outside cells ; 1 ACCEPT secreted / AW, from cells ACCEPT works in named extracellular environment e.g. digestive tract IGNORE doesn t work in cells 1 (b) (i)
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
Chapter 11. Cell Communication. Signal Transduction Pathways
 Chapter 11 Cell Communication Signal Transduction Pathways Signal-Transduction Pathway Signal on a cell s surface is converted into a specific cellular response Local signaling (short distance) - Paracrine
Chapter 11 Cell Communication Signal Transduction Pathways Signal-Transduction Pathway Signal on a cell s surface is converted into a specific cellular response Local signaling (short distance) - Paracrine
Statin inhibition of HMG-CoA reductase: a 3-dimensional view
 Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Bio 111 Study Guide Chapter 11 Cell Communication
 Bio 111 Study Guide Chapter 11 Cell Communication BEFORE CLASS: Reading: Read the introduction on p. 210, and for Concept 11.1, read from the first full paragraph on p. 212. Read all of Concept 11.2. Pay
Bio 111 Study Guide Chapter 11 Cell Communication BEFORE CLASS: Reading: Read the introduction on p. 210, and for Concept 11.1, read from the first full paragraph on p. 212. Read all of Concept 11.2. Pay
The three important structural features of proteins:
 The three important structural features of proteins: a. Primary (1 o ) The amino acid sequence (coded by genes) b. Secondary (2 o ) The interaction of amino acids that are close together or far apart in
The three important structural features of proteins: a. Primary (1 o ) The amino acid sequence (coded by genes) b. Secondary (2 o ) The interaction of amino acids that are close together or far apart in
AP Biology Summer Assignment Chapter 3 Quiz
 AP Biology Summer Assignment Chapter 3 Quiz 2016-17 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. All of the following are found in a DNA nucleotide
AP Biology Summer Assignment Chapter 3 Quiz 2016-17 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. All of the following are found in a DNA nucleotide
G-Protein Signaling. Introduction to intracellular signaling. Dr. SARRAY Sameh, Ph.D
 G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
Receptors. Dr. Sanaa Bardaweel
 Receptors Types and Theories Dr. Sanaa Bardaweel Some terms in receptor-drug interactions Agonists: drugs that mimic the natural messengers and activate receptors. Antagonist: drugs that block receptors.
Receptors Types and Theories Dr. Sanaa Bardaweel Some terms in receptor-drug interactions Agonists: drugs that mimic the natural messengers and activate receptors. Antagonist: drugs that block receptors.
Alternative splicing. Biosciences 741: Genomics Fall, 2013 Week 6
 Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Alternative splicing Biosciences 741: Genomics Fall, 2013 Week 6 Function(s) of RNA splicing Splicing of introns must be completed before nuclear RNAs can be exported to the cytoplasm. This led to early
Chapter 9. Cellular Signaling
 Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Propagation of the Signal
 OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Biological Molecules B Lipids, Proteins and Enzymes. Triglycerides. Glycerol
 Glycerol www.biologymicro.wordpress.com Biological Molecules B Lipids, Proteins and Enzymes Lipids - Lipids are fats/oils and are present in all cells- they have different properties for different functions
Glycerol www.biologymicro.wordpress.com Biological Molecules B Lipids, Proteins and Enzymes Lipids - Lipids are fats/oils and are present in all cells- they have different properties for different functions
Project Manual Bio3055. Cholesterol Homeostasis: HMG-CoA Reductase
 Project Manual Bio3055 Cholesterol Homeostasis: HMG-CoA Reductase Bednarski 2003 Funded by HHMI Cholesterol Homeostasis: HMG-CoA Reductase Introduction: HMG-CoA Reductase is an enzyme in the cholesterol
Project Manual Bio3055 Cholesterol Homeostasis: HMG-CoA Reductase Bednarski 2003 Funded by HHMI Cholesterol Homeostasis: HMG-CoA Reductase Introduction: HMG-CoA Reductase is an enzyme in the cholesterol
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
Protein structure. Dr. Mamoun Ahram Summer semester,
 Protein structure Dr. Mamoun Ahram Summer semester, 2017-2018 Overview of proteins Proteins have different structures and some have repeating inner structures, other do not. A protein may have gazillion
Protein structure Dr. Mamoun Ahram Summer semester, 2017-2018 Overview of proteins Proteins have different structures and some have repeating inner structures, other do not. A protein may have gazillion
KEY CONCEPT QUESTIONS IN SIGNAL TRANSDUCTION
 Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Comprehensive and Easy Course Notes for BIOL1040 Exams and Assessment
 Comprehensive and Easy Course Notes for BIOL1040 Exams and Assessment MODULE 1: PRINCIPLES OF CELL FUNCTION Membrane Structure & Function Cellular membranes are fluid mosaics of lipids and proteins Phospholipids
Comprehensive and Easy Course Notes for BIOL1040 Exams and Assessment MODULE 1: PRINCIPLES OF CELL FUNCTION Membrane Structure & Function Cellular membranes are fluid mosaics of lipids and proteins Phospholipids
Cells communicate with each other via signaling ligands which interact with receptors located on the surface or inside the target cell.
 BENG 100 Frontiers of Biomedical Engineering Professor Mark Saltzman Chapter 6 SUMMARY In this chapter, cell signaling was presented within the context of three physiological systems that utilize communication
BENG 100 Frontiers of Biomedical Engineering Professor Mark Saltzman Chapter 6 SUMMARY In this chapter, cell signaling was presented within the context of three physiological systems that utilize communication
Signal Transduction: Information Metabolism. Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire
 Signal Transduction: Information Metabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire Introduction Information Metabolism How cells receive, process and respond
Signal Transduction: Information Metabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire Introduction Information Metabolism How cells receive, process and respond
Cell Biology (BIOL 4374 and BCHS 4313) Third Exam 4/24/01
 Cell Biology (BIOL 4374 and BCHS 4313) Third Exam 4/24/01 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. For multiple choice questions,
Cell Biology (BIOL 4374 and BCHS 4313) Third Exam 4/24/01 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. For multiple choice questions,
Membrane associated receptor transfers the information. Second messengers relay information
 Membrane associated receptor transfers the information Most signals are polar and large Few of the signals are nonpolar Receptors are intrinsic membrane proteins Extracellular and intracellular domains
Membrane associated receptor transfers the information Most signals are polar and large Few of the signals are nonpolar Receptors are intrinsic membrane proteins Extracellular and intracellular domains
Chapter 11. Cell Communication
 Chapter 11 Cell Communication Overview: The Cellular Internet Cell-to-cell communication Is absolutely essential for multicellular organisms Concept 11.1: External signals are converted into responses
Chapter 11 Cell Communication Overview: The Cellular Internet Cell-to-cell communication Is absolutely essential for multicellular organisms Concept 11.1: External signals are converted into responses
Protein Secondary Structure
 Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Structure and Function of Antigen Recognition Molecules
 MICR2209 Structure and Function of Antigen Recognition Molecules Dr Allison Imrie allison.imrie@uwa.edu.au 1 Synopsis: In this lecture we will examine the major receptors used by cells of the innate and
MICR2209 Structure and Function of Antigen Recognition Molecules Dr Allison Imrie allison.imrie@uwa.edu.au 1 Synopsis: In this lecture we will examine the major receptors used by cells of the innate and
Practice Exam 2 MCBII
 1. Which feature is true for signal sequences and for stop transfer transmembrane domains (4 pts)? A. They are both 20 hydrophobic amino acids long. B. They are both found at the N-terminus of the protein.
1. Which feature is true for signal sequences and for stop transfer transmembrane domains (4 pts)? A. They are both 20 hydrophobic amino acids long. B. They are both found at the N-terminus of the protein.
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia
 Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Honors Biology Chapter 3: The Molecules of Cells Name Amatuzzi Carbohydrates pp Homework
 Honors Biology Chapter 3: The Molecules of Cells Name Amatuzzi Carbohydrates pp. 37-39 1. Which elements make up carbohydrates? a. In which ratio? 2. How do living things use most of their carbohydrates?
Honors Biology Chapter 3: The Molecules of Cells Name Amatuzzi Carbohydrates pp. 37-39 1. Which elements make up carbohydrates? a. In which ratio? 2. How do living things use most of their carbohydrates?
biochem480 [Autumn2014] Enzyme bio-informatics project
![biochem480 [Autumn2014] Enzyme bio-informatics project biochem480 [Autumn2014] Enzyme bio-informatics project](/thumbs/91/104514013.jpg) biochem480 [Autumn2014] Enzyme bio-informatics project Edit out any stuff in italics for the final version Student Name: IUBSystematic Name: 3-hydroxy-3-methylglutaryl-CoA reductase Other protein Names:
biochem480 [Autumn2014] Enzyme bio-informatics project Edit out any stuff in italics for the final version Student Name: IUBSystematic Name: 3-hydroxy-3-methylglutaryl-CoA reductase Other protein Names:
Lysosomes and endocytic pathways 9/27/2012 Phyllis Hanson
 Lysosomes and endocytic pathways 9/27/2012 Phyllis Hanson General principles Properties of lysosomes Delivery of enzymes to lysosomes Endocytic uptake clathrin, others Endocytic pathways recycling vs.
Lysosomes and endocytic pathways 9/27/2012 Phyllis Hanson General principles Properties of lysosomes Delivery of enzymes to lysosomes Endocytic uptake clathrin, others Endocytic pathways recycling vs.
Life Science 1A Final Exam. January 19, 2006
 ame: TF: Section Time Life Science 1A Final Exam January 19, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use non-erasable
ame: TF: Section Time Life Science 1A Final Exam January 19, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use non-erasable
Patrick, An Introduction to Medicinal Chemistry 4e Chapter 5 Receptors and signal transduction
 atrick, An Introduction to Medicinal Chemistry 4e Answers to end-of-chapter questions 1) The diagram in the question shows two important hydrogen bonding interactions where AT acts both as a hydrogen bond
atrick, An Introduction to Medicinal Chemistry 4e Answers to end-of-chapter questions 1) The diagram in the question shows two important hydrogen bonding interactions where AT acts both as a hydrogen bond
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Development Team. Head, Department of Zoology, University of Delhi. Department of Zoology, University of Delhi
 Paper No.: 10: Module : 03: cells, BCR, TCR, Co-receptors, properties of antigen recognized by Development Team Principal Investigator: Co-Principal Investigator: Paper Coordinator: Content Writer: Content
Paper No.: 10: Module : 03: cells, BCR, TCR, Co-receptors, properties of antigen recognized by Development Team Principal Investigator: Co-Principal Investigator: Paper Coordinator: Content Writer: Content
Insulin Resistance. Biol 405 Molecular Medicine
 Insulin Resistance Biol 405 Molecular Medicine Insulin resistance: a subnormal biological response to insulin. Defects of either insulin secretion or insulin action can cause diabetes mellitus. Insulin-dependent
Insulin Resistance Biol 405 Molecular Medicine Insulin resistance: a subnormal biological response to insulin. Defects of either insulin secretion or insulin action can cause diabetes mellitus. Insulin-dependent
Protein kinases are enzymes that add a phosphate group to proteins according to the. ATP + protein OH > Protein OPO 3 + ADP
 Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Lecture 36: Review of membrane function
 Chem*3560 Lecture 36: Review of membrane function Membrane: Lipid bilayer with embedded or associated proteins. Bilayers: 40-70% neutral phospholipid 10-20% negative phospholipid 10-30% cholesterol 10-30%
Chem*3560 Lecture 36: Review of membrane function Membrane: Lipid bilayer with embedded or associated proteins. Bilayers: 40-70% neutral phospholipid 10-20% negative phospholipid 10-30% cholesterol 10-30%
Peptide hydrolysis uncatalyzed half-life = ~450 years HIV protease-catalyzed half-life = ~3 seconds
 Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
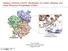
Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1
 Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1 β-lactamase (BLA) in blue. (a) c4 model and the close-up view
Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1 β-lactamase (BLA) in blue. (a) c4 model and the close-up view
Chapter 20. Cell - Cell Signaling: Hormones and Receptors. Three general types of extracellular signaling. endocrine signaling. paracrine signaling
 Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
Quaternary structure / Protein interfaces JBB2026H. Lecture 2: Protein Structure II. Most proteins are oligomers
 G. Privé Sept 2014 JBB2026H Lecture 2: Protein Structure II 1 Quaternary structure / Protein interfaces Most proteins are oligomers Common 3 structure does not imply common 4 structure Most oligomeric
G. Privé Sept 2014 JBB2026H Lecture 2: Protein Structure II 1 Quaternary structure / Protein interfaces Most proteins are oligomers Common 3 structure does not imply common 4 structure Most oligomeric
Problem Set 5 KEY
 2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION
 CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION What is Cytokine? Secreted popypeptide (protein) involved in cell-to-cell signaling. Acts in paracrine or autocrine fashion through specific cellular receptors.
CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION What is Cytokine? Secreted popypeptide (protein) involved in cell-to-cell signaling. Acts in paracrine or autocrine fashion through specific cellular receptors.
Hemoglobin & Sickle Cell Anemia Exercise
 Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Learning Objectives In this exercise, you will use StarBiochem, a protein 3D viewer, to explore the structure of the normal hemoglobin protein
Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Learning Objectives In this exercise, you will use StarBiochem, a protein 3D viewer, to explore the structure of the normal hemoglobin protein
Cellular Signaling Pathways. Signaling Overview
 Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia. Dec 14 & 19, 2006 Prof. Erin O Shea Prof. Dan Kahne
 Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne 1 Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne 1 Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
The T cell receptor for MHC-associated peptide antigens
 1 The T cell receptor for MHC-associated peptide antigens T lymphocytes have a dual specificity: they recognize polymporphic residues of self MHC molecules, and they also recognize residues of peptide
1 The T cell receptor for MHC-associated peptide antigens T lymphocytes have a dual specificity: they recognize polymporphic residues of self MHC molecules, and they also recognize residues of peptide
Lecture 4: 8/26. CHAPTER 4 Protein Three Dimensional Structure
 Lecture 4: 8/26 CHAPTER 4 Protein Three Dimensional Structure Summary of the Lecture 3 There are 20 amino acids and only the L isomer amino acid exist in proteins Each amino acid consists of a central
Lecture 4: 8/26 CHAPTER 4 Protein Three Dimensional Structure Summary of the Lecture 3 There are 20 amino acids and only the L isomer amino acid exist in proteins Each amino acid consists of a central
Amino Acids and Proteins (2) Professor Dr. Raid M. H. Al-Salih
 Amino Acids and Proteins (2) Professor Dr. Raid M. H. Al-Salih 1 Some important biologically active peptides 2 Proteins The word protein is derived from Greek word, proteios which means primary. As the
Amino Acids and Proteins (2) Professor Dr. Raid M. H. Al-Salih 1 Some important biologically active peptides 2 Proteins The word protein is derived from Greek word, proteios which means primary. As the
Chapter 3b Cells Membrane transport - Student Notes
 Chapter 3b Cells Membrane transport - Student Notes 1 Transport are permeable Some molecules the membrane; others do 2 Types of Membrane Transport processes No cellular required Substance its processes
Chapter 3b Cells Membrane transport - Student Notes 1 Transport are permeable Some molecules the membrane; others do 2 Types of Membrane Transport processes No cellular required Substance its processes
Complexity DNA. Genome RNA. Transcriptome. Protein. Proteome. Metabolites. Metabolome
 DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
Hemoglobin & Sickle Cell Anemia Exercise
 Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Background Hemoglobin is the protein in red blood cells responsible for carrying oxygen from the lungs to the rest of the body and for returning
Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Background Hemoglobin is the protein in red blood cells responsible for carrying oxygen from the lungs to the rest of the body and for returning
PHSI3009 Frontiers in Cellular Physiology 2017
 Overview of PHSI3009 L2 Cell membrane and Principles of cell communication L3 Signalling via G protein-coupled receptor L4 Calcium Signalling L5 Signalling via Growth Factors L6 Signalling via small G-protein
Overview of PHSI3009 L2 Cell membrane and Principles of cell communication L3 Signalling via G protein-coupled receptor L4 Calcium Signalling L5 Signalling via Growth Factors L6 Signalling via small G-protein
CHAPTER 1 INTRODUCTION
 CHAPTER 1 INTRODUCTION Understanding the mechanism of sweetness has been a challenging problem. It is necessary to understand this in order to design safer sweet molecules, for in many diseases (like diabetes,
CHAPTER 1 INTRODUCTION Understanding the mechanism of sweetness has been a challenging problem. It is necessary to understand this in order to design safer sweet molecules, for in many diseases (like diabetes,
Cell Communication. Chapter 11. PowerPoint Lectures for Biology, Seventh Edition. Lectures by Chris Romero. Neil Campbell and Jane Reece
 Chapter 11 Cell Communication PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: The Cellular Internet Cell-to-cell communication Is absolutely
Chapter 11 Cell Communication PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: The Cellular Internet Cell-to-cell communication Is absolutely
Proteins consist of joined amino acids They are joined by a Also called an Amide Bond
 Lecture Two: Peptide Bond & Protein Structure [Chapter 2 Berg, Tymoczko & Stryer] (Figures in Red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Proteins consist of joined amino acids
Lecture Two: Peptide Bond & Protein Structure [Chapter 2 Berg, Tymoczko & Stryer] (Figures in Red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Proteins consist of joined amino acids
INTERACTION DRUG BODY
 INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
Genetics of Cancer Lecture 32 Cancer II. Prof. Bevin Engelward, MIT Biological Engineering Department
 Genetics of Cancer Lecture 32 Cancer II rof. Bevin Engelward, MIT Biological Engineering Department Why Cancer Matters New Cancer Cases in 1997 Cancer Deaths in 1997 Genetics of Cancer: Today: What types
Genetics of Cancer Lecture 32 Cancer II rof. Bevin Engelward, MIT Biological Engineering Department Why Cancer Matters New Cancer Cases in 1997 Cancer Deaths in 1997 Genetics of Cancer: Today: What types
MCB*4010 Midterm Exam / Winter 2008
 MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
Structure and Activation of the Visual Pigment Rhodopsin
 Seminar 02.12.2011 Structure and Activation of the Visual Pigment Rhodopsin by Steven O.Smith, 2010 Constantin Schneider FB Physik, FU Berlin GPCR: Overview Rhodopsin is a G protein-coupled receptor (GPCR)
Seminar 02.12.2011 Structure and Activation of the Visual Pigment Rhodopsin by Steven O.Smith, 2010 Constantin Schneider FB Physik, FU Berlin GPCR: Overview Rhodopsin is a G protein-coupled receptor (GPCR)
Chemistry 135, First Exam. September 23, Chem 135, Exam 1 SID:
 Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
Sorting and trafficking of proteins in oligodendrocytes during myelin membrane biogenesis Klunder, Lammert
 University of Groningen Sorting and trafficking of proteins in oligodendrocytes during myelin membrane biogenesis Klunder, Lammert IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's
University of Groningen Sorting and trafficking of proteins in oligodendrocytes during myelin membrane biogenesis Klunder, Lammert IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's
a) The statement is true for X = 400, but false for X = 300; b) The statement is true for X = 300, but false for X = 200;
 1. Consider the following statement. To produce one molecule of each possible kind of polypeptide chain, X amino acids in length, would require more atoms than exist in the universe. Given the size of
1. Consider the following statement. To produce one molecule of each possible kind of polypeptide chain, X amino acids in length, would require more atoms than exist in the universe. Given the size of
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor!
 Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
- Biosignaling: Signal transduction. References: chapter 8 of Lippincots chapter 1 3 of Lehningers
 Basic concepts of Metabolism Metabolism and metabolic pathway Metabolic Map Catabolism Anabolism - Regulation of Metabolism Signals from within the cell (Intracellular) Communication between cells. - Biosignaling:
Basic concepts of Metabolism Metabolism and metabolic pathway Metabolic Map Catabolism Anabolism - Regulation of Metabolism Signals from within the cell (Intracellular) Communication between cells. - Biosignaling:
Lecture 7: Signaling Through Lymphocyte Receptors
 Lecture 7: Signaling Through Lymphocyte Receptors Questions to Consider After recognition of its cognate MHC:peptide, how does the T cell receptor activate immune response genes? What are the structural
Lecture 7: Signaling Through Lymphocyte Receptors Questions to Consider After recognition of its cognate MHC:peptide, how does the T cell receptor activate immune response genes? What are the structural
Heat Shock Proteins in Targeted Cancer Chemotherapy
 Heat Shock Proteins in Targeted Cancer Chemotherapy Aykut ÖZGÜR 1 and Yusuf TUTAR 2,* 1 Gaziosmanpaşa University, Faculty of Natural Sciences and Engineering, Department of Bioengineering, Tokat, Turkey
Heat Shock Proteins in Targeted Cancer Chemotherapy Aykut ÖZGÜR 1 and Yusuf TUTAR 2,* 1 Gaziosmanpaşa University, Faculty of Natural Sciences and Engineering, Department of Bioengineering, Tokat, Turkey
Chapter 3. Structure of Enzymes. Enzyme Engineering
 Chapter 3. Structure of Enzymes Enzyme Engineering 3.1 Introduction With purified protein, Determining M r of the protein Determining composition of amino acids and the primary structure Determining the
Chapter 3. Structure of Enzymes Enzyme Engineering 3.1 Introduction With purified protein, Determining M r of the protein Determining composition of amino acids and the primary structure Determining the
BIOLOGY 103 Spring 2001 MIDTERM LAB SECTION
 BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
Structural biology of viruses
 Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Molecular biology, isotopic labeling, and refolding of membrane proteins in phospholipid bilayers.
 Molecular biology, isotopic labeling, and refolding of membrane proteins in phospholipid bilayers. 1 Preparation of membrane proteins for NMR experiments. Established bacterial overexpression systems.
Molecular biology, isotopic labeling, and refolding of membrane proteins in phospholipid bilayers. 1 Preparation of membrane proteins for NMR experiments. Established bacterial overexpression systems.
Cell Communication. Chapter 11. Biology Eighth Edition Neil Campbell and Jane Reece. PowerPoint Lecture Presentations for
 Chapter 11 Cell Communication PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Chapter 11 Cell Communication PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp
Basics of Signal Transduction. Ebaa M Alzayadneh, PhD
 Basics of Signal Transduction Ebaa M Alzayadneh, PhD What is signal transduction? Cell signaling The science of understanding how individual cells sense their environments and respond to stimuli... how
Basics of Signal Transduction Ebaa M Alzayadneh, PhD What is signal transduction? Cell signaling The science of understanding how individual cells sense their environments and respond to stimuli... how
Solutions to Quiz I
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz I Class Average = 72 Median = 73 Grade Range % A 82-100 22 B 70-81 39 C 57 69 28 D 41 56 10 F 0 40 0 Question 1
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Solutions to 7.014 Quiz I Class Average = 72 Median = 73 Grade Range % A 82-100 22 B 70-81 39 C 57 69 28 D 41 56 10 F 0 40 0 Question 1
Biol220 Cellular Signalling. Non-receptor tyrosine kinases
 Biol220 Cellular Signalling Non-receptor tyrosine kinases The 7TM receptors initiate signal transducton pathways through changes in tertiary structure that are induced by ligand binding. A fundamentally
Biol220 Cellular Signalling Non-receptor tyrosine kinases The 7TM receptors initiate signal transducton pathways through changes in tertiary structure that are induced by ligand binding. A fundamentally
Discovery and Optimization of Inhibitors of STAT3 Activation for the Treatment of Squamous Cell Carcinoma of the Head and Neck
 Discovery and ptimization of Inhibitors of STAT3 Activation for the Treatment of Squamous Cell Carcinoma of the Head and Neck Feng Zhang Wipf Group Research Topic Seminar 02-09-2013 1 Feng Zhang @ Wipf
Discovery and ptimization of Inhibitors of STAT3 Activation for the Treatment of Squamous Cell Carcinoma of the Head and Neck Feng Zhang Wipf Group Research Topic Seminar 02-09-2013 1 Feng Zhang @ Wipf
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Chemical Mechanism of Enzymes
 Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
االمتحان النهائي لعام 1122
 االمتحان النهائي لعام 1122 Amino Acids : 1- which of the following amino acid is unlikely to be found in an alpha-helix due to its cyclic structure : -phenylalanine -tryptophan -proline -lysine 2- : assuming
االمتحان النهائي لعام 1122 Amino Acids : 1- which of the following amino acid is unlikely to be found in an alpha-helix due to its cyclic structure : -phenylalanine -tryptophan -proline -lysine 2- : assuming
Chemistry 106: Drugs in Society Lecture 19: How do Drugs Elicit an Effect? Interactions between Drugs and Macromolecular Targets 11/02/17
 Chemistry 106: Drugs in Society Lecture 19: How do Drugs Elicit an Effect? Interactions between Drugs and Macromolecular Targets 11/02/17 Targets for Therapeutic Intervention: A Comparison of Enzymes to
Chemistry 106: Drugs in Society Lecture 19: How do Drugs Elicit an Effect? Interactions between Drugs and Macromolecular Targets 11/02/17 Targets for Therapeutic Intervention: A Comparison of Enzymes to
The clathrin adaptor Numb regulates intestinal cholesterol. absorption through dynamic interaction with NPC1L1
 The clathrin adaptor Numb regulates intestinal cholesterol absorption through dynamic interaction with NPC1L1 Pei-Shan Li 1, Zhen-Yan Fu 1,2, Ying-Yu Zhang 1, Jin-Hui Zhang 1, Chen-Qi Xu 1, Yi-Tong Ma
The clathrin adaptor Numb regulates intestinal cholesterol absorption through dynamic interaction with NPC1L1 Pei-Shan Li 1, Zhen-Yan Fu 1,2, Ying-Yu Zhang 1, Jin-Hui Zhang 1, Chen-Qi Xu 1, Yi-Tong Ma
