Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69
|
|
|
- Melissa Singleton
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69 germline gene sequence. 2G1 and 8M2 acquired 14 and 18 mutations from the germline gene sequence, respectively. In 2G1, the mutations are mostly located in the HCDR1 and HCDR2 loops. Antibody residues that are in contact with HA are highlighted in red. The signature hydrophobic residues of the V H 1-69 germline, Ile53 and Phe54, are highlighted with red reverse triangles.
2 Supplementary Figure 2. Glycan binding analysis of recombinant escape mutants of H2 HA on the glycan microarray. (a) A/Japan/305+/1957 R137Q. (b) T193K. (c) G135D. (d) K156E. All error bars in the figure are indicative of standard deviation from quadruplicate measurements. Specific bindings for neutral glycans (1 and 2), α2-3- linked glycans (3 to 35), α2-6-linked sialylated glycans (36 to 56) and glycans of α2-6 and α2-3 mixed linkages (57 and 58) are colored in black, blue, red and cyan bars, respectively. The list of glycans on the array is provided in Supplemental Table 1. Note that the panels shown in this figure are on a different scale than Fig. 6B-C due to the relatively low avidity of the mutants. In general, the single mutations abolish glycan binding (or weaken glycan binding to below the detection threshold) for H2 HA, compared with the wild-type HA tested under the same conditions. Mutant R137Q retained some weak binding for one α2-6-linked glycan (#55).
3 Supplementary Figure 3. Electron density maps (2Fo-Fc) of CDR loops in the crystal structures. (a) 2G1 HCDR2, (b) 8M2 HCDR2, (c) 8F8 HCDR3. The maps are contoured at a 2σ level.
4 Supplementary Table 1. Glycans imprinted on the microarray. Glycan # 1 Galβ(1-4)-GlcNAcβ-ethyl-NH2 2 Name Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-3)-[Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-6)]-Manβ(1-4)- GlcNAcβ(1-4)-GlcNAcβ-Asn- 3 NeuAcα(2-3)-Galβ(1-4)-6-O-sulfo-GlcNAcβ-propyl- 4 NeuAcα(2-3)-Galβ(1-4)-[Fucα(1-3)]-6-O-sulfo-GlcNAcβ-propyl-NH2 5 NeuAcα(2-3)-6-O-sulfo-Galβ(1-4)-GlcNAcβ-ethyl-NH2 6 NeuAcα(2-3)-6-O-sulfo-Galβ(1-4)-[Fucα(1-3)]-GlcNAcβ-propyl-NH2 7 NeuAcα(2-3)-Galβ(1-3)-6-O-sulfo-GlcNAcβ-propyl-NH2 8 NeuAcα(2-3)-Galβ(1-4)-Glcβ-ethyl- 9 NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ-ethyl- 10 NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ-ethyl- 11 NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ-ethyl- NH2 12 NeuAcα(2-3)-GalNAcβ(1-4)-GlcNAcβ-ethyl- 13 NeuAcα(2-3)-Galβ(1-3)-GlcNAcβ-ethyl- 14 NeuAcα(2-3)-Galβ(1-3)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ-ethyl- 15 NeuAcα(2-3)-Galβ(1-3)-GlcNAcβ(1-3)-Galβ(1-3)-GlcNAcβ-ethyl- 16 NeuAcα(2-3)-Galβ(1-3)-GalNAcβ(1-3)-Galα(1-4)-Galβ(1-4)-Glcβ-ethyl- 17 NeuAcα(2-3)-Galβ(1-3)-GalNAcα-Thr- 18 NeuAcα(2-3)-Galβ(1-3)-[GlcNAcβ(1-6)]-GalNAcα-Thr- 19 NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-6)-[Galβ(1-3)]-GalNAcα-Thr- 20 NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-6)-[Galβ(1-3)]-GalNAcα-Thr- 21 NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-3)-GalNAcα-Thr- 22 NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-3)-GalNAcα-Thr- 23 NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-3)-[NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-6)]-GalNAcα- Thr- 24 NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-3)-[NeuAcα(2-3)-Galβ(1-4)- GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-6)]-GalNAcα-Thr- 25 NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-3)-[NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-2)- Manα(1-6)]-Manβ(1-4)-GlcNAcβ(1-4)-GlcNAcβ-Asn- NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-3)-[NeuAcα(2-3)- 26 Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-6)]-Manβ(1-4)-GlcNAcβ(1-4)- GlcNAcβ-Asn- NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-2)- 27 Manα(1-3)-[NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)- GlcNAcβ(1-2)-Manα(1-6)]-Manβ(1-4)-GlcNAcβ(1-4)-GlcNAcβ-Asn- 28 NeuAcα(2-3)-[GalNAcβ(1-4)]-Galβ(1-4)-GlcNAcβ-ethyl- 29 NeuAcα(2-3)-[GalNAcβ(1-4)]-Galβ(1-4)-Glcβ-ethyl- 30 Galβ(1-3)-GalNAcβ(1-4)-[NeuAcα(2-3)]-Galβ(1-4)-Glcβ-ethyl- 31 NeuAcα(2-3)-Galβ(1-4)-[Fucα(1-3)]-GlcNAcβ-propyl- 32 NeuAcα(2-3)-Galβ(1-3)-[Fucα(1-4)]-GlcNAcβ(1-3)-Galβ(1-4)-[Fucα(1-3)]-GlcNAcβ-ethyl- 33 NeuAcα(2-3)-Galβ(1-4)-[Fucα(1-3)]-GlcNAcβ(1-3)-Galβ(1-4)-[Fucα(1-3)]-GlcNAcβ-ethyl- 34 NeuAcα(2-3)-Galβ(1-4)-[Fucα(1-3)]-GlcNAcβ(1-3)-Galβ(1-4)-[Fucα(1-3)]-GlcNAcβ(1-3)- Galβ(1-4)-[Fucα(1-3)]-GlcNAcβ-ethyl- 35 NeuGcα(2-3)-Galβ(1-4)-GlcNAcβ-ethyl- 36 NeuAcα(2-6)-Galβ(1-4)-6-O-sulfo-GlcNAcβ-propyl- 37 NeuAcα(2-6)-Galβ(1-4)-Glcβ-ethyl- 38 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ-ethyl-
5 39 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ-ethyl- 40 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ-ethyl- 41 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-3)-[NeuAcα(2-6)]-Galβ(1-4)-GlcNAcβ-ethyl-NH2 42 NeuAcα(2-6)-GalNAcβ(1-4)-GlcNAcβ-ethyl- 43 NeuAcα(2-6)-[Galβ(1-3)]-GalNAcα-Thr- 44 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-6)-[Galβ(1-3)]-GalNAcα-Thr- 45 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-6)-[Galβ(1-3)]-GalNAcα-Thr- 46 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-3)-GalNAcα-Thr- 47 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-3)-GalNAcα-Thr- 48 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-3)-[NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-6)]-GalNAcα- Thr- 49 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-3)-[NeuAcα(2-6)-Galβ(1-4)- GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-6)]-GalNAcα-Thr- 50 Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-3)-[NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-6)]- Manβ(1-4)-GlcNAcβ(1-4)-GlcNAcβ-Asn- 51 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-3)-[Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-6)]- Manβ(1-4)-GlcNAcβ(1-4)-GlcNAcβ-Asn- 52 GlcNAcβ(1-2)-Manα(1-3)-[NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-6)]-Manβ(1-4)- GlcNAcβ(1-4)-GlcNAcβ-Asn- 53 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-3)-[NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-2)- Manα(1-6)]-Manβ(1-4)-GlcNAcβ(1-4)-GlcNAcβ-Asn- NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-3)-[NeuAcα(2-6)- 54 Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-6)]-Manβ(1-4)-GlcNAcβ(1-4)- GlcNAcβ-Asn- NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-2)- 55 Manα(1-3)-[NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)-GlcNAcβ(1-3)-Galβ(1-4)- GlcNAcβ(1-2)-Manα(1-6)]-Manβ(1-4)-GlcNAcβ(1-4)-GlcNAcβ-Asn- 56 NeuGcα(2-6)-Galβ(1-4)-GlcNAcβ-ethyl- 57 NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-3)-[NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-2)- Manα(1-6)]-Manβ(1-4)-GlcNAcβ(1-4)-GlcNAcβ-Asn- 58 NeuAcα(2-6)-Galβ(1-4)-GlcNAcβ(1-2)-Manα(1-3)-[NeuAcα(2-3)-Galβ(1-4)-GlcNAcβ(1-2)- Manα(1-6)]-Manβ(1-4)-GlcNAcβ(1-4)-GlcNAcβ-Asn-
Supplementary Figure 1 Weight and body temperature of ferrets inoculated with
 Supplementary Figure 1 Weight and body temperature of ferrets inoculated with A/Anhui/1/2013 (H7N9) influenza virus. (a) Body temperature and (b) weight change of ferrets after intranasal inoculation with
Supplementary Figure 1 Weight and body temperature of ferrets inoculated with A/Anhui/1/2013 (H7N9) influenza virus. (a) Body temperature and (b) weight change of ferrets after intranasal inoculation with
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12391 Table 1. Binding of A(H7N9) viruses to individual glycan structures. # Structure a Anhui1 Shanghai1 1 Neu5Acα nb nb 2 Neu5Acα ++ nb 3 Neu5Acβ nb nb 4 Neu5Acα2-3(6-O-Su)Galβ1-4GlcNAcβ
doi:10.1038/nature12391 Table 1. Binding of A(H7N9) viruses to individual glycan structures. # Structure a Anhui1 Shanghai1 1 Neu5Acα nb nb 2 Neu5Acα ++ nb 3 Neu5Acβ nb nb 4 Neu5Acα2-3(6-O-Su)Galβ1-4GlcNAcβ
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Information
 Supplementary Information Profiling of core fucosylated N-glycans using a novel bacterial lectin that specifically recognizes α1,6 fucosylated chitobiose Saulius Vainauskas 1, Rebecca M. Duke 1,2, James
Supplementary Information Profiling of core fucosylated N-glycans using a novel bacterial lectin that specifically recognizes α1,6 fucosylated chitobiose Saulius Vainauskas 1, Rebecca M. Duke 1,2, James
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
What sort of Science is Glycoscience? (Introductory lecture)
 Glycosciences: Glycobiology & Glycochemistry e-learning course What sort of Science is Glycoscience? (Introductory lecture) Paula Videira Faculdade de Ciências Médicas Nova University, Lisbon Portugal
Glycosciences: Glycobiology & Glycochemistry e-learning course What sort of Science is Glycoscience? (Introductory lecture) Paula Videira Faculdade de Ciências Médicas Nova University, Lisbon Portugal
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Information
 Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information.
 Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al.
 Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplemental Material can be found at:
 Supplemental Material can be found at: http://www.jbc.org/content/suppl/2009/05/06/m109.012807.dc1.html THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 284, NO. 27, pp. 18537 18544, July 3, 2009 Author s Choice
Supplemental Material can be found at: http://www.jbc.org/content/suppl/2009/05/06/m109.012807.dc1.html THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 284, NO. 27, pp. 18537 18544, July 3, 2009 Author s Choice
OligoTech - Human milk oligosaccharides - HMOs
 ligotech - Human milk oligosaccharides - HMs Elicityl is developing a unique offer of HMs available at quantities and cost not seen before. Human milk oligosaccharides (HMs) are recognized as natural molecules
ligotech - Human milk oligosaccharides - HMs Elicityl is developing a unique offer of HMs available at quantities and cost not seen before. Human milk oligosaccharides (HMs) are recognized as natural molecules
on Non-Consensus Protein Motifs Analytical & Formulation Sciences, Amgen. Seattle, WA
 N-Linked Glycosylation on Non-Consensus Protein Motifs Alain Balland Analytical & Formulation Sciences, Amgen. Seattle, WA CASSS - Mass Spec 2010 Marina Del Rey, CA. September 8 th, 2010 Outline 2 Consensus
N-Linked Glycosylation on Non-Consensus Protein Motifs Alain Balland Analytical & Formulation Sciences, Amgen. Seattle, WA CASSS - Mass Spec 2010 Marina Del Rey, CA. September 8 th, 2010 Outline 2 Consensus
Crystallization-grade After D After V3 cocktail. Time (s) Time (s) Time (s) Time (s) Time (s) Time (s)
 Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Supplementary Information for Structural basis for the inhibition of Polo-like kinase 1
 Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Acceptor Specificity Studies of Fucosyl- and Sialyltransferases
 Acceptor Specificity Studies of Fucosyl- and Sialyltransferases Suvi Toivonen Institute of Biotechnology and Department of Biosciences, Division of Biochemistry Faculty of Science University of Helsinki
Acceptor Specificity Studies of Fucosyl- and Sialyltransferases Suvi Toivonen Institute of Biotechnology and Department of Biosciences, Division of Biochemistry Faculty of Science University of Helsinki
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Roles of Polyvalency in
 Roles of Polyvalency in the Mechanism of Glyco Recognition. An Important Direction toward the Future Glycosciences 1 2 Albert M. Wu Albert M. Wu Glyco-Immunochemistry Research Laboratory Institute of Molecular
Roles of Polyvalency in the Mechanism of Glyco Recognition. An Important Direction toward the Future Glycosciences 1 2 Albert M. Wu Albert M. Wu Glyco-Immunochemistry Research Laboratory Institute of Molecular
PURIFICATION AND CHARACTERIZATION OF AN N-ACETYL-D- GLUCOSAMINE SPECIFIC LECTIN FROM THE AUSTRALIAN MUSHROOM PSATHYRELLA ASPEROSPORA
 PURIFICATION AND CHARACTERIZATION OF AN N-ACETYL-D- GLUCOSAMINE SPECIFIC LECTIN FROM THE AUSTRALIAN MUSHROOM PSATHYRELLA ASPEROSPORA MOHAMED ALI ABOL HASSAN 1, RAZINA ROUF 1, JOÃO P RIBEIRO 2, FABIA JÄGGIN
PURIFICATION AND CHARACTERIZATION OF AN N-ACETYL-D- GLUCOSAMINE SPECIFIC LECTIN FROM THE AUSTRALIAN MUSHROOM PSATHYRELLA ASPEROSPORA MOHAMED ALI ABOL HASSAN 1, RAZINA ROUF 1, JOÃO P RIBEIRO 2, FABIA JÄGGIN
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Supersite of immune vulnerability on the glycosylated face of HIV-1 envelope glycoprotein gp120
 Supersite of immune vulnerability on the glycosylated face of HIV-1 envelope glycoprotein gp120 Leopold Kong, Jeong Hyun Lee, Katie J. Doores, Charles D. Murin, Jean-Philippe Julien, Ryan McBride, Yan
Supersite of immune vulnerability on the glycosylated face of HIV-1 envelope glycoprotein gp120 Leopold Kong, Jeong Hyun Lee, Katie J. Doores, Charles D. Murin, Jean-Philippe Julien, Ryan McBride, Yan
Structure of the measles virus hemagglutinin bound to the CD46 receptor. César Santiago, María L. Celma, Thilo Stehle and José M.
 Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION
 Supplementary discussion of the role of C05 HCDR1 in antigen binding. Deletion of the 5 amino-acid insertion in HCDR1 del(fgest) and scanning mutagenesis of the tip of the loop had differing effects on
Supplementary discussion of the role of C05 HCDR1 in antigen binding. Deletion of the 5 amino-acid insertion in HCDR1 del(fgest) and scanning mutagenesis of the tip of the loop had differing effects on
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
SUPPLEMENTARY INFORMATION
 Supplementary Information Supplementary Figure 1. The serine paradox. Nature-designed AlaRSs pin-down the -NH3 + group of alanine with an acidic Asp that is critical for binding and activation of alanine.
Supplementary Information Supplementary Figure 1. The serine paradox. Nature-designed AlaRSs pin-down the -NH3 + group of alanine with an acidic Asp that is critical for binding and activation of alanine.
BIOSYNTHESIS OF CANCER-RELATED CARBOHYDRATE ANTIGENS. Fabio Dall Olio Department of Experimental Pathology University of Bologna, Italy
 BIOSYNTHESIS OF CANCER-RELATED CARBOHYDRATE ANTIGENS Fabio Dall Olio Department of Experimental Pathology University of Bologna, Italy TOPICS OF THE LECTURE 1. Structure and function of some representative
BIOSYNTHESIS OF CANCER-RELATED CARBOHYDRATE ANTIGENS Fabio Dall Olio Department of Experimental Pathology University of Bologna, Italy TOPICS OF THE LECTURE 1. Structure and function of some representative
Nature Medicine: doi: /nm.4322
 1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
1 2 3 4 5 6 7 8 9 10 11 Supplementary Figure 1. Predicted RNA structure of 3 UTR and sequence alignment of deleted nucleotides. (a) Predicted RNA secondary structure of ZIKV 3 UTR. The stem-loop structure
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Chapter 5. Generation of lymphocyte antigen receptors
 Chapter 5 Generation of lymphocyte antigen receptors Structural variation in Ig constant regions Isotype: different class of Ig Heavy-chain C regions are encoded in separate genes Initially, only two of
Chapter 5 Generation of lymphocyte antigen receptors Structural variation in Ig constant regions Isotype: different class of Ig Heavy-chain C regions are encoded in separate genes Initially, only two of
The Binding Mode of by Electron Crystallography
 The Binding Mode of Epothilone A on α,β-tubulin by Electron Crystallography James H. Nettles, Huilin Li, Ben Cornett, Joseph M. Krahn, James P. Snyder, Kenneth H. Downing Science, Volume 305, August 6,
The Binding Mode of Epothilone A on α,β-tubulin by Electron Crystallography James H. Nettles, Huilin Li, Ben Cornett, Joseph M. Krahn, James P. Snyder, Kenneth H. Downing Science, Volume 305, August 6,
P450 CYCLE. All P450s follow the same catalytic cycle of;
 P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
Nature Structural & Molecular Biology: doi: /nsmb.1933
 The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B
 3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
STRUCTURE OF A UBIQUITIN-LOADED HECT LIGASE REVEALS THE MOLECULAR BASIS FOR CATALYTIC PRIMING SUPPLEMENTARY INFORMATION
 STRUCTURE OF A UBIQUITINLOADED ECT LIGASE REVEALS TE MOLECULAR BASIS FOR CATALYTIC PRIMING Elena Maspero 1, Eleonora Valentini 1, Sara Mari 1, Valentina Cecatiello 2, Paolo Soffientini 1, Sebastiano Pasqualato
STRUCTURE OF A UBIQUITINLOADED ECT LIGASE REVEALS TE MOLECULAR BASIS FOR CATALYTIC PRIMING Elena Maspero 1, Eleonora Valentini 1, Sara Mari 1, Valentina Cecatiello 2, Paolo Soffientini 1, Sebastiano Pasqualato
Gerlind Sulzenbacher, Véronique Roig-Zamboni, Willy J. Peumans, Pierre Rougé, Els J.M. Van Damme, Yves Bourne
 Accepted Manuscript Crystal structure of the GalNAc/Gal-specific agglutinin from the phytopathogenic ascomycete Sclerotinia sclerotiorum reveals novel adaptation of a β-trefoil domain Gerlind Sulzenbacher,
Accepted Manuscript Crystal structure of the GalNAc/Gal-specific agglutinin from the phytopathogenic ascomycete Sclerotinia sclerotiorum reveals novel adaptation of a β-trefoil domain Gerlind Sulzenbacher,
Enzymatic synthesis of known and novel oligosaccharides
 Enzymatic synthesis of known and novel oligosaccharides Jari Natunen Institute of Biotechnology Department of Biosciences, Division of Biochemistry Faculty of Science University of Helsinki Finland Academic
Enzymatic synthesis of known and novel oligosaccharides Jari Natunen Institute of Biotechnology Department of Biosciences, Division of Biochemistry Faculty of Science University of Helsinki Finland Academic
HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation
 HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation Meng Zhang 1, River Charles 1, Huimin Tong 1, Lei Zhang 1, Mili Patel 2, Francis Wang 1, Matthew
HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation Meng Zhang 1, River Charles 1, Huimin Tong 1, Lei Zhang 1, Mili Patel 2, Francis Wang 1, Matthew
BLOOD GROUP PRODUCTS
 BLOOD GROUP PRODUCTS The discovery of the ABO blood typing system by Karl Landsteiner over 100 years ago and the subsequent elucidation of their carbohydrate structures by Walter Morgan were exceedingly
BLOOD GROUP PRODUCTS The discovery of the ABO blood typing system by Karl Landsteiner over 100 years ago and the subsequent elucidation of their carbohydrate structures by Walter Morgan were exceedingly
The Basics: A general review of molecular biology:
 The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Microwave-Mediated Analysis for Sugar, Fatty Acid and Sphingoid
 Microwave-Mediated Analysis for Sugar, Fatty Acid and Sphingoid Compositions of Glycosphingolipids Saki Itonori 1*, Masato Takahashi 1, Tomonori Kitamura 1, Kazuhiro Aoki 2, John T. Dulaney 3 and Mutsumi
Microwave-Mediated Analysis for Sugar, Fatty Acid and Sphingoid Compositions of Glycosphingolipids Saki Itonori 1*, Masato Takahashi 1, Tomonori Kitamura 1, Kazuhiro Aoki 2, John T. Dulaney 3 and Mutsumi
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
Supplementary Figure 1 Overall structure of the transmembrane pore of FraC. a, Crystal packing of the pore along the y-z, b, the z-x, and c, the y-x
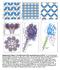 Supplementary Figure 1 Overall structure of the transmembrane pore of FraC. a, Crystal packing of the pore along the y-z, b, the z-x, and c, the y-x planes. The N-terminal and the β-core regions are depicted
Supplementary Figure 1 Overall structure of the transmembrane pore of FraC. a, Crystal packing of the pore along the y-z, b, the z-x, and c, the y-x planes. The N-terminal and the β-core regions are depicted
Glycan receptor specificity as a useful tool for characterization and surveillance of influenza A virus
 Glycan receptor specificity as a useful tool for characterization and surveillance of influenza A virus The MIT Faculty has made this article openly available. Please share how this access benefits you.
Glycan receptor specificity as a useful tool for characterization and surveillance of influenza A virus The MIT Faculty has made this article openly available. Please share how this access benefits you.
Structural analysis of fungus-derived FAD glucose dehydrogenase
 Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Johnson DB, Balko JM, Compton ML, et al. Fulminant myocarditis
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Johnson DB, Balko JM, Compton ML, et al. Fulminant myocarditis
Nature Immunology: doi: /ni Supplementary Figure 1. Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice.
 Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
A standardized method for lectin microarray-based tissue glycome. mapping
 A standardized method for lectin microarray-based tissue glycome mapping Xia Zou,2,#, Maki Yoshida,#, Chiaki Nagai-Okatani,#, Jun Iwaki, Atsushi Matsuda, Binbin Tan,2, Kozue Hagiwara, Takashi Sato, Yoko
A standardized method for lectin microarray-based tissue glycome mapping Xia Zou,2,#, Maki Yoshida,#, Chiaki Nagai-Okatani,#, Jun Iwaki, Atsushi Matsuda, Binbin Tan,2, Kozue Hagiwara, Takashi Sato, Yoko
SUPPLEMENTARY INFORMATION FOR. (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation. by COX-2
 SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10134 Supplementary Figure 1. Anti-inflammatory activity of sfc. a, Autoantibody immune complexes crosslink activating Fc receptors, promoting activation of macrophages, and WWW.NATURE.COM/NATURE
doi:10.1038/nature10134 Supplementary Figure 1. Anti-inflammatory activity of sfc. a, Autoantibody immune complexes crosslink activating Fc receptors, promoting activation of macrophages, and WWW.NATURE.COM/NATURE
Figure S1. Expression efficiency of rd2bpl3 for various expression hosts. (A) BL21(DE3), (B)
 1 Supplementary Figure S1. 2 3 4 5 6 7 8 Figure S1. Expression efficiency of rd2bpl3 for various expression hosts. (A) BL21(DE3), (B) BL21(DE3)pLysS, (C) BL21-CodonPlus(DE3)-RIL, (D) Rosetta(DE3). M, Molecular
1 Supplementary Figure S1. 2 3 4 5 6 7 8 Figure S1. Expression efficiency of rd2bpl3 for various expression hosts. (A) BL21(DE3), (B) BL21(DE3)pLysS, (C) BL21-CodonPlus(DE3)-RIL, (D) Rosetta(DE3). M, Molecular
Research Online. Edith Cowan University. Constantinos K. Wibmer. Jinal N. Bhiman. Elin S. Gray Edith Cowan University,
 Edith Cowan University Research Online ECU Publications 2013 2013 Viral Escape From HIV-1 Neutralizing Antibodies Drives Increased Plasma Neutralization Breadth through Sequential Recognition Of Multiple
Edith Cowan University Research Online ECU Publications 2013 2013 Viral Escape From HIV-1 Neutralizing Antibodies Drives Increased Plasma Neutralization Breadth through Sequential Recognition Of Multiple
Symbiotic Relationship between Human and Bifidobacteria. Kenji YAMAMOTO. Graduate School of Biostudies, Kyoto University
 Symbiotic Relationship between Human and Bifidobacteria Kenji YAMAMOTO Graduate School of Biostudies, Kyoto University Bifidobacterium longum Classification of Major Lactic Acid Bacteria Kingdom Subkingdom
Symbiotic Relationship between Human and Bifidobacteria Kenji YAMAMOTO Graduate School of Biostudies, Kyoto University Bifidobacterium longum Classification of Major Lactic Acid Bacteria Kingdom Subkingdom
Chapter 6. X-ray structure analysis of D30N tethered HIV-1 protease. dimer/saquinavir complex
 Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Holy Grail of Transfusion Medicine
 Hemolytic Transfusion Reactions ABO Blood Group System October 21, 2005 Steven L. Spitalnik, M.D. Holy Grail of Transfusion Medicine Manipulate the composition of blood: With complete control Without adverse
Hemolytic Transfusion Reactions ABO Blood Group System October 21, 2005 Steven L. Spitalnik, M.D. Holy Grail of Transfusion Medicine Manipulate the composition of blood: With complete control Without adverse
Antigen Receptor Structures October 14, Ram Savan
 Antigen Receptor Structures October 14, 2016 Ram Savan savanram@uw.edu 441 Lecture #8 Slide 1 of 28 Three lectures on antigen receptors Part 1 (Today): Structural features of the BCR and TCR Janeway Chapter
Antigen Receptor Structures October 14, 2016 Ram Savan savanram@uw.edu 441 Lecture #8 Slide 1 of 28 Three lectures on antigen receptors Part 1 (Today): Structural features of the BCR and TCR Janeway Chapter
ERK1/2/MAPK pathway-dependent regulation of the telomeric factor TRF2
 ERK1/2/MAPK pathway-dependent regulation of the telomeric factor TRF2 SUPPLEMENTARY FIGURES AND TABLE Supplementary Figure S1: Conservation of the D domain throughout evolution. Alignment of TRF2 sequences
ERK1/2/MAPK pathway-dependent regulation of the telomeric factor TRF2 SUPPLEMENTARY FIGURES AND TABLE Supplementary Figure S1: Conservation of the D domain throughout evolution. Alignment of TRF2 sequences
SUPPLEMENTAL MATERIAL. UNC119 is required for G protein trafficking in sensory neurons
 1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
CHAPTER 21: Amino Acids, Proteins, & Enzymes. General, Organic, & Biological Chemistry Janice Gorzynski Smith
 CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
Nature Immunology: doi: /ni Supplementary Figure 1. Cytokine pattern in skin in response to urushiol.
 Supplementary Figure 1 Cytokine pattern in skin in response to urushiol. Wild-type (WT) and CD1a-tg mice (n = 3 per group) were sensitized and challenged with urushiol (uru) or vehicle (veh). Quantitative
Supplementary Figure 1 Cytokine pattern in skin in response to urushiol. Wild-type (WT) and CD1a-tg mice (n = 3 per group) were sensitized and challenged with urushiol (uru) or vehicle (veh). Quantitative
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Supplementary Figure 1. Generation of a conditional allele of the Kindlin-2 gene. (A) A restriction map of the relevant genomic region of Kindlin-2 (top), the targeting construct
SUPPLEMENTARY INFORMATION Supplementary Figure 1. Generation of a conditional allele of the Kindlin-2 gene. (A) A restriction map of the relevant genomic region of Kindlin-2 (top), the targeting construct
SUPPORTING INFORMATION. Evidence for Regulation of Hemoglobin Metabolism and Intracellular Ionic Flux
 SUPPORTING INFORMATION Evidence for Regulation of Hemoglobin Metabolism and Intracellular Ionic Flux by the Plasmodium falciparum Chloroquine Resistance Transporter Andrew H. Lee, Satish K. Dhingra, Ian
SUPPORTING INFORMATION Evidence for Regulation of Hemoglobin Metabolism and Intracellular Ionic Flux by the Plasmodium falciparum Chloroquine Resistance Transporter Andrew H. Lee, Satish K. Dhingra, Ian
Supplementary Information
 Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
SUPPLEMENTARY FIG. S2. Representative counting fields used in quantification of the in vitro neural differentiation of pattern of dnscs.
 Supplementary Data SUPPLEMENTARY FIG. S1. Representative counting fields used in quantification of the in vitro neural differentiation of pattern of anpcs. A panel of lineage-specific markers were used
Supplementary Data SUPPLEMENTARY FIG. S1. Representative counting fields used in quantification of the in vitro neural differentiation of pattern of anpcs. A panel of lineage-specific markers were used
Annotation of potential isobaric and isomeric lipid species analyzed using the MxP Quant 500 Kit
 Annotation of potential isobaric and isomeric lipid species analyzed using the MxP Quant 500 Kit Introduction The MxP Quant 500 kit enables the analysis of up to 630 metabolites covering 26 compound classes
Annotation of potential isobaric and isomeric lipid species analyzed using the MxP Quant 500 Kit Introduction The MxP Quant 500 kit enables the analysis of up to 630 metabolites covering 26 compound classes
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Meaning-based guidance of attention in scenes as revealed by meaning maps
 SUPPLEMENTARY INFORMATION Letters DOI: 1.138/s41562-17-28- In the format provided by the authors and unedited. -based guidance of attention in scenes as revealed by meaning maps John M. Henderson 1,2 *
SUPPLEMENTARY INFORMATION Letters DOI: 1.138/s41562-17-28- In the format provided by the authors and unedited. -based guidance of attention in scenes as revealed by meaning maps John M. Henderson 1,2 *
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Your partner of choice for carbohydrates
 Your partner of choice for carbohydrates Contents About Dextra 5 Custom synthesis 6 cgmp production facility 9 Analytical capabilities 10 New Zealand Pharmaceuticals 12 How to place an order 15 Product
Your partner of choice for carbohydrates Contents About Dextra 5 Custom synthesis 6 cgmp production facility 9 Analytical capabilities 10 New Zealand Pharmaceuticals 12 How to place an order 15 Product
Preferential induction of cross-group influenza A hemagglutinin stem specific memory B cells after H7N9 immunization in humans
 INFECTIOUS DISEASE Preferential induction of cross-group influenza A hemagglutinin stem specific memory B cells after immunization in humans Sarah F. Andrews, 1 * M. Gordon Joyce, 1 Michael J. Chambers,
INFECTIOUS DISEASE Preferential induction of cross-group influenza A hemagglutinin stem specific memory B cells after immunization in humans Sarah F. Andrews, 1 * M. Gordon Joyce, 1 Michael J. Chambers,
Main Study: Summer Methods. Design
 Main Study: Summer 2000 Methods Design The experimental design is within-subject each participant experiences five different trials for each of the ten levels of Display Condition and for each of the three
Main Study: Summer 2000 Methods Design The experimental design is within-subject each participant experiences five different trials for each of the ten levels of Display Condition and for each of the three
The Interaction of the Gut Microbiota with the Mucus Barrier in Health and Disease in Human
 Review The Interaction of the Gut Microbiota with the Mucus Barrier in Health and Disease in Human Anthony P. Corfield Mucin Research Group, School of Clinical Sciences, Bristol Royal Infirmary, Level
Review The Interaction of the Gut Microbiota with the Mucus Barrier in Health and Disease in Human Anthony P. Corfield Mucin Research Group, School of Clinical Sciences, Bristol Royal Infirmary, Level
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
Supplementary Information for. Heavy chain-only IgG2b-llama antibody effects near-pan HIV-1 neutralization by
 Supplementary Information for Heavy chain-only IgG2b-llama antibody effects near-pan HIV-1 neutralization by recognizing a CD4-induced epitope that includes elements of co-receptor- and CD4-binding sites
Supplementary Information for Heavy chain-only IgG2b-llama antibody effects near-pan HIV-1 neutralization by recognizing a CD4-induced epitope that includes elements of co-receptor- and CD4-binding sites
Journal: Nature Methods
 Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Glycan Receptor for Influenza Virus
 26 The Open Antimicrobial Agents Journal, 2010, 2, 26-33 Glycan Receptor for Influenza Virus Open Access Kazuya I.P.J. Hidari * and Takashi Suzuki Department of Biochemistry, University of Shizuoka, School
26 The Open Antimicrobial Agents Journal, 2010, 2, 26-33 Glycan Receptor for Influenza Virus Open Access Kazuya I.P.J. Hidari * and Takashi Suzuki Department of Biochemistry, University of Shizuoka, School
The mechanism of patellamide macrocyclization revealed by study of the Prochloron sp PatG macrocyclase domain
 The mechanism of patellamide macrocyclization revealed by study of the Prochloron sp PatG macrocyclase domain Jesko Koehnke 1,2, Andrew Bent 1,2, Wael E. Houssen 2,3,4, David Zollman 1, Falk Morawitz 1,
The mechanism of patellamide macrocyclization revealed by study of the Prochloron sp PatG macrocyclase domain Jesko Koehnke 1,2, Andrew Bent 1,2, Wael E. Houssen 2,3,4, David Zollman 1, Falk Morawitz 1,
DEVELOPMENTAL CONTROL OF BACTERIAL RECEPTORS IN THE GASTROINTESTINAL TRACT. SHU-HEH W. CHU and W. ALLAN WALKER
 Old Herborn University Seminar Monograph 4: Host-microflora interactions in the first years after birth. Editors: Peter J. Heidt, Rial D. Rolfe, Volker Rusch, and Dirk van der Waaij. Institute for Microbiology
Old Herborn University Seminar Monograph 4: Host-microflora interactions in the first years after birth. Editors: Peter J. Heidt, Rial D. Rolfe, Volker Rusch, and Dirk van der Waaij. Institute for Microbiology
Intestinal mucosal barrier function in health and disease. Turner, JR Nature Rev. Immunology 9: 799 (2009)
 Chapter 9 O-GalNAc Glycans - Essentials of Glycobiology 2 nd edition Intestinal mucosal barrier function in health and disease. Turner, JR Nature Rev. Immunology 9: 799 (2009) From Mucins to Mucus Thornton
Chapter 9 O-GalNAc Glycans - Essentials of Glycobiology 2 nd edition Intestinal mucosal barrier function in health and disease. Turner, JR Nature Rev. Immunology 9: 799 (2009) From Mucins to Mucus Thornton
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Pathogenicity and transmission of a swine influenza A(H6N6) virus
 OPEN (2017) 6, e17; doi:10.1038/emi.2017.3 www.nature.com/emi ORIGINAL ARTICLE Pathogenicity and transmission of a swine influenza A(H6N6) virus Hailiang Sun 1, Bryan S Kaplan 2, Minhui Guan 1, Guihong
OPEN (2017) 6, e17; doi:10.1038/emi.2017.3 www.nature.com/emi ORIGINAL ARTICLE Pathogenicity and transmission of a swine influenza A(H6N6) virus Hailiang Sun 1, Bryan S Kaplan 2, Minhui Guan 1, Guihong
