Nature Neuroscience: doi: /nn Supplementary Figure 1. Missense damaging predictions as a function of allele frequency
|
|
|
- Melvyn O’Neal’
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 Missense damaging predictions as a function of allele frequency Percentage of missense variants classified as damaging by eight different classifiers and a classifier consisting of the intersection of classifiers SIFT, PolyPhen-2 HDIV, PolyPhen-2 HVAR, LRT, Mutation Taster, Mutation Assessor, and PROVEAN as a function of minor allele count across 12,332 unrelated individuals from Sweden. Gray and black colors indicate, respectively, variants never observed and variants already observed in the ExAC cohort of 45,376 individuals.
2
3 Supplementary Figure 2 Correlations across ultra-rare variant types counts Relationships between four separate types of coding-sequence and splice-site URV counts across 12,332 unrelated individuals from Sweden: (a) disruptive vs. damaging; (b) disruprive vs. missense non-damaging; (c) damaging vs. missense non-damaging; (d) disruptive vs. synonymous; (e) damaging vs. synonymous; (f) missense non-damaging vs synonymous. Black dots indicate individuals with less than or equal to 100 URVs detected, and red crosses indicate individuals with more than 100 URVs detected. Pearson correlation coefficients are indicated on the top left of each panel.
4 Supplementary Figure 3 Enrichment in schizophrenia cases across ultra-rare variant types Observed enrichment in 4,877 schizophrenia cases compared to 6,203 controls for (a) URVs across all annotations, with non-coding including both intronic and untranslated region variants, (b) missense URVs classified as damaging by classifiers PolyPhen-2 HDIV, PolyPhen-2 HVAR, SIFT, LRT, PROVEAN, FATHMM, Mutation Taster, and Mutation Assessor, as well as missense damaging, inframe indel, protein-protein-contact, splice-acceptor, splice-donor, stop-gained, and frameshift URVs. Enrichment and P values were computed using a linear regression model (left panels) and a logistic regression model (right panels) and correcting for covariates (a) with and (b) without the exclusion of URV individual count. Horizontal bars indicate 95% confidence intervals.
5
6 Supplementary Figure 4 Single gene burden tests for disruptive and damaging variants at different allele frequency thresholds Quantile-quantile plots for burden tests for association with schizophrenia of all Ensembl genes for disruptive and damaging variants. Burden test for association was performed using SKAT software.
7 Supplementary Figure 5 Enrichment of durvs in schizophrenia cases across selected gene sets stratified per variant types and cohorts Burden of durvs across selected gene sets analyzed in this study stratified across disruptive and damaging URVs (left panel) and across previously analyzed exomes and newly available exomes (right panel). Enrichment and P values were computed using a logistic regression model using exome-wide durv count as a covariate to correct for average exome-wide burden (dot-dashed line). Horizontal bars indicate 95% confidence intervals.
8 Supplementary Figure 6 Enrichment of durvs in schizophrenia cases across constrained and unconstrained genes Burden of durvs across loss-of-function intolerant (LoF-intolerant) genes, missense constrained genes and their complementary sets, stratified across disruptive and damaging URVs. Enrichment and P values were computed using a linear regression model (left panels) and a logistic regression model (right panels) and, contrary to most other analyses in this study, without using exome-wide durv count to correct for average exome-wide burden. Horizontal bars indicate 95% confidence intervals. As LoF-intolerant and missense constrained genes were defined outside this study, different enrichments observed within and outside these gene sets cannot be attributed to differential false positive rates between cases and controls.
9 Supplementary Figure 7 Enrichment of durvs in schizophrenia cases across gene sets from different tissues Burden of durvs across 27 tissue expression specific gene sets. Each gene set was generated selecting genes for which expression in a given tissue was at least 5 times the median expression across 27 different human organs and tissues ascertained from 95 individual. Enrichment and P values were computed using a logistic regression model using exome-wide durv count as a covariate to correct for average exome-wide burden (dot-dashed line). Horizontal bars indicate 95% confidence intervals. Brain specific genes were significantly more enriched for durvs in schizophrenia cases than the average gene.
10 Supplementary Figure 8 Enrichment of durvs in schizophrenia cases across gene sets from different brain and neuronal cell types Burden analysis for durvs across genes defined from (a) 11 brain cell types and (b) 3 neuron cell types: excitatory pyramidal neurons (Exc), parvalbumin (PV)-expressing fast-spiking interneurons, vasoactive intestinal peptide (VIP)-expressing interneurons, both for expressed genes and cell-type specific genes. Brain cell type gene sets were generated selecting genes for which log-expression in a given cell type was 0.5 greater than the median log-expression across 11 central nervous system cell types ascertained from developing and mature mouse forebrain. Neuron cell type expressed gene sets were defined as those with more than 50 observed transcripts per million (TPM). Neuron cell type specific gene sets were defined as those observed more than 5 times the minimum expression across the 3 different cell types. Expression profiles for neurons were ascertained from nuclei isolated from adult (8 11 weeks) mouse neocortex. Enrichment and P values were computed using a logistic regression model using exome-wide durv count as a covariate to correct for average exome-wide burden (dot-dashed line). Horizontal bars indicate 95% confidence intervals.
11 Supplementary Figure 9 Enrichment of durvs in schizophrenia cases across different X linked intellectual disability gene sets Burden analysis for durvs across X linked intellectual disability (XLID) genes and developmental disorder genes. X linked ID genes correspond to the union of X linked ID sets of genes as defined from OMIM, GCC, and Chicago. Enrichment was computed separately for males, females, and both groups together. Enrichment and P values were computed using a logistic regression model using exomewide durv count as a covariate to correct for average exome-wide burden (dot-dashed line). Horizontal bars indicate 95% confidence intervals. Larger confidence intervals for males reflect the smaller number of variants observed in males due to carrying half as much X chromosome DNA.
12 Supplementary Figure 10 Average ultra-rare variant types count across sequencing waves Average number of URVs detected in controls and in schizophrenia cases used in this analysis across several classes of variants and sequencing waves. Numbers of controls and cases within each wave is indicated in parentheses. Wave 1 is denoted in red as for this batch, accounting for approximately 1% of the whole cohort, an earlier version of the hybrid-capture procedure was used which captured approximately 10% less of the exome. Vertical bars indicate 95% confidence intervals.
13 Supplementary Figure 11 Duplicate and first degree relationship estimates across cohort Estimates of percentage of genome shared IBD (PI_HAT) and percentage of genome shared IBD1 (Z1) among all pairs from 12,384 samples for which PI_HAT>.35. Estimates were generated with plink command --genome full.
14
15 Supplementary Figure 12 Ultra-rare variants counts relationships with population stratification Covariates within the Swedish exome cohort: (a) ultra-rare SNP and indel counts across cohort 12,334 individuals; (b) distribution of birth year across cohort individuals; (c) relationship of cohort individuals (in black) with 1000 Genomes project phase 1 individuals with respect to the first two principal components with removed outliers (in red); (d) relationship of cohort individuals with respect to the two main Swedish principal components; (e) relationship between principal components tracking Finnish ancestry and URV count; (f) relationship between principal components tracking Northern-Southern Swedish ancestry and URV count. Black crosses indicate individuals with less than or equal to 100 URVs detected, and red crosses indicate individuals with more than 100 URVs detected.
16 Supplementary Figure 13 Gender estimates from genotypes X chromosome inbreeding coefficient (F) and Y-chromosome non-missing genotype calls (YCOUNT) for the entire cohort of 12,384 samples. Male individuals inferred with 47,XXY karyotype (Klinefelter syndrome) are indicated in green and individuals with mismatching reported and genotyped sex are indicated as yellow crosses. Estimates were generated with plink command --check-sex ycount.
17 Supplementary Figure 14 Manhattan plot for association with schizophrenia of common exome variants Manhattan plot for association of exonic variants with schizophrenia phenotype using a logistic regression model with sex and first five principal components as covariates. Variants with P value less than 10-6 include variant rs on chromosome 2 in the UTR5 of genes TYW5 and C2orf47, and seven variants in the MHC region around the HLA genes.
18 Supplementary Figure 15 Missense predictors comparisons for association with schizophrenia P values for enrichment of missense URVs classified as damaging by all 256 possible predictors defined as the combination of damaging definitions from any subset of Polyphen2_HDIV, Polyphen2_HVAR, SIFT, LRT, PROVEAN, FATHMM, Mutation Taster, and Mutation Assessor algorithms. P values were computed using a linear regression model and correcting for covariates including total URV count. Predictors including FATHMM performed significantly worse than predictors excluding FATHMM while 25 predictors performed better than the predictor chosen for the analyses in this manuscript (bright red). Both predictors including all algorithms but FATHMM (bright red) and including LRT, MutationTaster, PolyPhen2 HDIV, PolyPhen2 HVAR, and SIFT (green) performed better than each predictor based on a single algorithm.
19 Covariate URV variance explained (%) durv variance explained (%) URV (Not used) Sex 0.02 NA Birth year Exome capturing kit 0.22 NA 1 st principal component 0.01 NA 2 nd principal component 0.15 NA 3 rd principal component 7.22 NA 4 th principal component 0.39 NA 5 th principal component NA 6 th principal component 1.10 NA 7 th principal component 0.28 NA 8 th principal component NA NA 9 th principal component NA NA 10 th principal component NA NA 11 th principal component 0.12 NA 12 th principal component NA NA 13 th principal component NA NA 14 th principal component NA NA 15 th principal component NA NA 16 th principal component 0.03 NA 17 th principal component 0.01 NA 18 th principal component NA NA 19 th principal component 0.02 NA 20 th principal component NA NA Supplementary Table 1 Variance in number of ultra-rare variants uniquely explained by each covariate Variance of the total number of URVs and durvs across individuals uniquely explained by each covariate used for linear regression and logistic regression analyses. All covariates combined explained 34.68% and 28.87% of the overall variability in, respectively, URV and durv count. Principal components 3 and 5 correlate with, respectively, the amount of Finnish ancestry and the amount of Northern Swedish ancestry and accounted alone for 31.21% of the variability in URV count and 8.49% of the variability in durv count. Including year of birth accounted for an additional 0.25% of the variability in URV count, possibly because younger individuals tended to have more exotic ancestry or because younger individuals recent variants were less likely to be shared with individuals from the rest of the cohort due to slightly longer coalescent paths.
20 Gene symbol P value durvs in cases durvs in controls bp length KL ,039 STXBP ,782 PCYT1A ,104 CPVL ,431 KDM5B ,635 HEPHL ,480 AP5Z ,424 TTN ,053 CSNK1G ,368 KCNH ,591 ZDBF ,065 ITPR ,232 PRSS ,545 CCDC ,352 LAMB ,397 PCSK ,910 PRDM ,483 KIAA ,522 NDST ,652 TMPO ,365 KIAA ,272 CDO OGDHL ,033 SORCS ,540 KDM5C ,683 PLEKHG ,140 JARID ,741 ARFGEF ,550 ASAH ,188 PMS ,589 TRIM ,235 TRAF3IP ,076 GOLGA ,693 MPP ,914 KLHL ,770 WBSCR QRICH ,331 ISYNA ,677 VSTM2A DBNL ,320 THG1L LY ,169 LY75-CD ,622 DNM1L ,211 CHRNE ,482 MTAP TBC1D2B ,892 Supplementary Table 2 Genes with strongest results for association of durvs with schizophrenia List of genes with durvs burden test statistically significant at a = Number of durvs observed in schizophrenia cases (4,877 individuals) and controls (6,203 individuals), as well as coding base pairs length of the gene, are reported.
21 Class Gene set URVs Disruptive m.a.c.=1 (no URVs) m.a.c. 5 (no URVs) m.a.c. 10 (no URVs) m.a.f.<0.1% (no URVs) m.a.f.<0.5% (no URVs) LoF-intolerant Potentially Synaptic All LoF-intolerant Damaging Potentially Synaptic All LoF-intolerant Disruptive and damaging Potentially Synaptic All Supplementary Table 8 Results for non ultra-rare variants association with schizophrenia across key gene sets P value for burden tests using SKAT software across variation classes disruptive, damaging, and combined, across genes that are loss-of-function intolerant (LoF-intolerant), potentially synaptic, and all genes, and across URVs and non-ultra-rare variants up to minor allele count (m.a.c.) 1, 5, and 10 and up to minor allele frequency (m.a.f.) 0.1% and 0.5%. Differently to other gene sets analysis in the rest of the manuscript, these analyses were not corrected for exome-wide enrichment of the same class of variants.
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Nature Genetics: doi: /ng Supplementary Figure 1. PCA for ancestry in SNV data.
 Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
6/12/2018. Disclosures. Clinical Genomics The CLIA Lab Perspective. Outline. COH HopeSeq Heme Panels
 Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Variant Annotation and Functional Prediction
 Variant Annotation and Functional Prediction Copyrighted 2018 Isabelle Schrauwen and Suzanne M. Leal This exercise touches on several functionalities of the program ANNOVAR to annotate and interpret candidate
Variant Annotation and Functional Prediction Copyrighted 2018 Isabelle Schrauwen and Suzanne M. Leal This exercise touches on several functionalities of the program ANNOVAR to annotate and interpret candidate
Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical
 Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical representation using all SNPs (n= 13,515,798) including the region on chromosome 1 including SORT1 which was previously
Supplementary Figure 1. Quantile-quantile (Q-Q) plots. (Panel A) Q-Q plot graphical representation using all SNPs (n= 13,515,798) including the region on chromosome 1 including SORT1 which was previously
Using large-scale human genetic variation to inform variant prioritization in neuropsychiatric disorders
 Using large-scale human genetic variation to inform variant prioritization in neuropsychiatric disorders Kaitlin E. Samocha Hurles lab, Wellcome Trust Sanger Institute ACGS Summer Scientific Meeting 27
Using large-scale human genetic variation to inform variant prioritization in neuropsychiatric disorders Kaitlin E. Samocha Hurles lab, Wellcome Trust Sanger Institute ACGS Summer Scientific Meeting 27
Tutorial on Genome-Wide Association Studies
 Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
Rare Variant Burden Tests. Biostatistics 666
 Rare Variant Burden Tests Biostatistics 666 Last Lecture Analysis of Short Read Sequence Data Low pass sequencing approaches Modeling haplotype sharing between individuals allows accurate variant calls
Rare Variant Burden Tests Biostatistics 666 Last Lecture Analysis of Short Read Sequence Data Low pass sequencing approaches Modeling haplotype sharing between individuals allows accurate variant calls
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.
 Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Nature Genetics: doi: /ng Supplementary Figure 1. Country distribution of GME samples and designation of geographical subregions.
 Supplementary Figure 1 Country distribution of GME samples and designation of geographical subregions. GME samples collected across 20 countries and territories from the GME. Pie size corresponds to the
Supplementary Figure 1 Country distribution of GME samples and designation of geographical subregions. GME samples collected across 20 countries and territories from the GME. Pie size corresponds to the
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Epigenetics. Jenny van Dongen Vrije Universiteit (VU) Amsterdam Boulder, Friday march 10, 2017
 Epigenetics Jenny van Dongen Vrije Universiteit (VU) Amsterdam j.van.dongen@vu.nl Boulder, Friday march 10, 2017 Epigenetics Epigenetics= The study of molecular mechanisms that influence the activity of
Epigenetics Jenny van Dongen Vrije Universiteit (VU) Amsterdam j.van.dongen@vu.nl Boulder, Friday march 10, 2017 Epigenetics Epigenetics= The study of molecular mechanisms that influence the activity of
New Enhancements: GWAS Workflows with SVS
 New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Interrogation of rare functional variation within bipolar disorder and suicidal behavior cohorts
 University of Iowa Iowa Research Online Theses and Dissertations Spring 2018 Interrogation of rare functional variation within bipolar disorder and suicidal behavior cohorts Eric Thayne Monson University
University of Iowa Iowa Research Online Theses and Dissertations Spring 2018 Interrogation of rare functional variation within bipolar disorder and suicidal behavior cohorts Eric Thayne Monson University
5/2/18. After this class students should be able to: Stephanie Moon, Ph.D. - GWAS. How do we distinguish Mendelian from non-mendelian traits?
 corebio II - genetics: WED 25 April 2018. 2018 Stephanie Moon, Ph.D. - GWAS After this class students should be able to: 1. Compare and contrast methods used to discover the genetic basis of traits or
corebio II - genetics: WED 25 April 2018. 2018 Stephanie Moon, Ph.D. - GWAS After this class students should be able to: 1. Compare and contrast methods used to discover the genetic basis of traits or
Supplementary Figure 1. Quantile-quantile (Q-Q) plot of the log 10 p-value association results from logistic regression models for prostate cancer
 Supplementary Figure 1. Quantile-quantile (Q-Q) plot of the log 10 p-value association results from logistic regression models for prostate cancer risk in stage 1 (red) and after removing any SNPs within
Supplementary Figure 1. Quantile-quantile (Q-Q) plot of the log 10 p-value association results from logistic regression models for prostate cancer risk in stage 1 (red) and after removing any SNPs within
De novo mutational profile in RB1 clarified using a mutation rate modeling algorithm
 Aggarwala et al. BMC Genomics (2017) 18:155 DOI 10.1186/s12864-017-3522-z RESEARCH ARTICLE Open Access De novo mutational profile in RB1 clarified using a mutation rate modeling algorithm Varun Aggarwala
Aggarwala et al. BMC Genomics (2017) 18:155 DOI 10.1186/s12864-017-3522-z RESEARCH ARTICLE Open Access De novo mutational profile in RB1 clarified using a mutation rate modeling algorithm Varun Aggarwala
Supplementary Information
 1 Supplementary Information MATERIALS AND METHODS Study subjects Exome-sequenced HBOC patients Clinical characteristics of 24 exome-sequenced HBOC patients are presented in Table 1. Table 1. Clinical characteristics
1 Supplementary Information MATERIALS AND METHODS Study subjects Exome-sequenced HBOC patients Clinical characteristics of 24 exome-sequenced HBOC patients are presented in Table 1. Table 1. Clinical characteristics
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Lecture 20. Disease Genetics
 Lecture 20. Disease Genetics Michael Schatz April 12 2018 JHU 600.749: Applied Comparative Genomics Part 1: Pre-genome Era Sickle Cell Anaemia Sickle-cell anaemia (SCA) is an abnormality in the oxygen-carrying
Lecture 20. Disease Genetics Michael Schatz April 12 2018 JHU 600.749: Applied Comparative Genomics Part 1: Pre-genome Era Sickle Cell Anaemia Sickle-cell anaemia (SCA) is an abnormality in the oxygen-carrying
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Replicability of blood eqtl effects in ileal biopsies from the RISK study. eqtls detected in the vicinity of SNPs associated with IBD tend to show concordant effect size and direction
Supplementary Figure 1 Replicability of blood eqtl effects in ileal biopsies from the RISK study. eqtls detected in the vicinity of SNPs associated with IBD tend to show concordant effect size and direction
MEDICAL GENOMICS LABORATORY. Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG)
 Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
Home Brewed Personalized Genomics
 Home Brewed Personalized Genomics The Quest for Meaningful Analysis Results of a 23andMe Exome Pilot Trio of Myself, Wife, and Son February 22, 2013 Gabe Rudy, Vice President of Product Development Exome
Home Brewed Personalized Genomics The Quest for Meaningful Analysis Results of a 23andMe Exome Pilot Trio of Myself, Wife, and Son February 22, 2013 Gabe Rudy, Vice President of Product Development Exome
Quality Control Analysis of Add Health GWAS Data
 2018 Add Health Documentation Report prepared by Heather M. Highland Quality Control Analysis of Add Health GWAS Data Christy L. Avery Qing Duan Yun Li Kathleen Mullan Harris CAROLINA POPULATION CENTER
2018 Add Health Documentation Report prepared by Heather M. Highland Quality Control Analysis of Add Health GWAS Data Christy L. Avery Qing Duan Yun Li Kathleen Mullan Harris CAROLINA POPULATION CENTER
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE
 CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
Welcome to the Genetic Code: An Overview of Basic Genetics. October 24, :00pm 3:00pm
 Welcome to the Genetic Code: An Overview of Basic Genetics October 24, 2016 12:00pm 3:00pm Course Schedule 12:00 pm 2:00 pm Principles of Mendelian Genetics Introduction to Genetics of Complex Disease
Welcome to the Genetic Code: An Overview of Basic Genetics October 24, 2016 12:00pm 3:00pm Course Schedule 12:00 pm 2:00 pm Principles of Mendelian Genetics Introduction to Genetics of Complex Disease
Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) cancer DNA
 www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
Nature Neuroscience: doi: /nn Supplementary Figure 1. Trial structure for go/no-go behavior
 Supplementary Figure 1 Trial structure for go/no-go behavior a, Overall timeline of experiments. Day 1: A1 mapping, injection of AAV1-SYN-GCAMP6s, cranial window and headpost implantation. Water restriction
Supplementary Figure 1 Trial structure for go/no-go behavior a, Overall timeline of experiments. Day 1: A1 mapping, injection of AAV1-SYN-GCAMP6s, cranial window and headpost implantation. Water restriction
Supplementary Figures
 Supplementary Figures Supplementary Fig 1. Comparison of sub-samples on the first two principal components of genetic variation. TheBritishsampleisplottedwithredpoints.The sub-samples of the diverse sample
Supplementary Figures Supplementary Fig 1. Comparison of sub-samples on the first two principal components of genetic variation. TheBritishsampleisplottedwithredpoints.The sub-samples of the diverse sample
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
The contribution of rare variants to risk of schizophrenia in individuals with and without intellectual disability
 The contribution of rare variants to risk of schizophrenia in individuals with and without intellectual disability Tarjinder Singh 1, James T R Walters 2, Mandy Johnstone 3, David Curtis 4,5, Jaana Suvisaari
The contribution of rare variants to risk of schizophrenia in individuals with and without intellectual disability Tarjinder Singh 1, James T R Walters 2, Mandy Johnstone 3, David Curtis 4,5, Jaana Suvisaari
BWA alignment to reference transcriptome and genome. Convert transcriptome mappings back to genome space
 Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Supplementary Figure 1: Classification scheme for non-synonymous and nonsense germline MC1R variants. The common variants with previously established
 Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
MutationTaster & RegulationSpotter
 MutationTaster & RegulationSpotter Pathogenicity Prediction of Sequence Variants: Past, Present and Future Dr. rer. nat. Jana Marie Schwarz Klinik für Pädiatrie m. S. Neurologie Exzellenzcluster NeuroCure
MutationTaster & RegulationSpotter Pathogenicity Prediction of Sequence Variants: Past, Present and Future Dr. rer. nat. Jana Marie Schwarz Klinik für Pädiatrie m. S. Neurologie Exzellenzcluster NeuroCure
Genome-wide Association Analysis Applied to Asthma-Susceptibility Gene. McCaw, Z., Wu, W., Hsiao, S., McKhann, A., Tracy, S.
 Genome-wide Association Analysis Applied to Asthma-Susceptibility Gene McCaw, Z., Wu, W., Hsiao, S., McKhann, A., Tracy, S. December 17, 2014 1 Introduction Asthma is a chronic respiratory disease affecting
Genome-wide Association Analysis Applied to Asthma-Susceptibility Gene McCaw, Z., Wu, W., Hsiao, S., McKhann, A., Tracy, S. December 17, 2014 1 Introduction Asthma is a chronic respiratory disease affecting
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
GENOME-WIDE ASSOCIATION STUDIES
 GENOME-WIDE ASSOCIATION STUDIES SUCCESSES AND PITFALLS IBT 2012 Human Genetics & Molecular Medicine Zané Lombard IDENTIFYING DISEASE GENES??? Nature, 15 Feb 2001 Science, 16 Feb 2001 IDENTIFYING DISEASE
GENOME-WIDE ASSOCIATION STUDIES SUCCESSES AND PITFALLS IBT 2012 Human Genetics & Molecular Medicine Zané Lombard IDENTIFYING DISEASE GENES??? Nature, 15 Feb 2001 Science, 16 Feb 2001 IDENTIFYING DISEASE
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
SUPPLEMENTARY FIGURES: Supplementary Figure 1
 SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
Analysis with SureCall 2.1
 Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Supplementary Figure 1. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations.
 Supplementary Figure. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations. a Eigenvector 2.5..5.5. African Americans European Americans e
Supplementary Figure. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations. a Eigenvector 2.5..5.5. African Americans European Americans e
Investigating rare diseases with Agilent NGS solutions
 Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Nature Neuroscience: doi: /nn Supplementary Figure 1. Lick response during the delayed Go versus No-Go task.
 Supplementary Figure 1 Lick response during the delayed Go versus No-Go task. Trial-averaged lick rate was averaged across all mice used for pyramidal cell imaging (n = 9). Different colors denote different
Supplementary Figure 1 Lick response during the delayed Go versus No-Go task. Trial-averaged lick rate was averaged across all mice used for pyramidal cell imaging (n = 9). Different colors denote different
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Case presentations from diagnostic exome sequencing results in ion channels M. Koko, U. Hedrich, H. Lerche DASNE meeting 2017, Eisenach
 Case presentations from diagnostic exome sequencing results in ion channels M. Koko, U. Hedrich, H. Lerche DASNE meeting 2017, Eisenach CASE 1: Clinical picture Patient s presentation: 38 year old jewish
Case presentations from diagnostic exome sequencing results in ion channels M. Koko, U. Hedrich, H. Lerche DASNE meeting 2017, Eisenach CASE 1: Clinical picture Patient s presentation: 38 year old jewish
Ct=28.4 WAT 92.6% Hepatic CE (mg/g) P=3.6x10-08 Plasma Cholesterol (mg/dl)
 a Control AAV mtm6sf-shrna8 Ct=4.3 Ct=8.4 Ct=8.8 Ct=8.9 Ct=.8 Ct=.5 Relative TM6SF mrna Level P=.5 X -5 b.5 Liver WAT Small intestine Relative TM6SF mrna Level..5 9.6% Control AAV mtm6sf-shrna mtm6sf-shrna6
a Control AAV mtm6sf-shrna8 Ct=4.3 Ct=8.4 Ct=8.8 Ct=8.9 Ct=.8 Ct=.5 Relative TM6SF mrna Level P=.5 X -5 b.5 Liver WAT Small intestine Relative TM6SF mrna Level..5 9.6% Control AAV mtm6sf-shrna mtm6sf-shrna6
Finding subtle mutations with the Shannon human mrna splicing pipeline
 Finding subtle mutations with the Shannon human mrna splicing pipeline Presentation at the CLC bio Medical Genomics Workshop American Society of Human Genetics Annual Meeting November 9, 2012 Peter K Rogan
Finding subtle mutations with the Shannon human mrna splicing pipeline Presentation at the CLC bio Medical Genomics Workshop American Society of Human Genetics Annual Meeting November 9, 2012 Peter K Rogan
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Identifying Mutations Responsible for Rare Disorders Using New Technologies
 Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Supplementary Information. Data Identifies FAN1 at 15q13.3 as a Susceptibility. Gene for Schizophrenia and Autism
 Supplementary Information A Scan-Statistic Based Analysis of Exome Sequencing Data Identifies FAN1 at 15q13.3 as a Susceptibility Gene for Schizophrenia and Autism Iuliana Ionita-Laza 1,, Bin Xu 2, Vlad
Supplementary Information A Scan-Statistic Based Analysis of Exome Sequencing Data Identifies FAN1 at 15q13.3 as a Susceptibility Gene for Schizophrenia and Autism Iuliana Ionita-Laza 1,, Bin Xu 2, Vlad
CS2220 Introduction to Computational Biology
 CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
Shashikant Kulkarni, M.S (Medicine)., Ph.D., FACMG Associate Professor of Pathology & Immunology Associate Professor of Pediatrics and Genetics
 Shashikant Kulkarni, M.S (Medicine)., Ph.D., FACMG Associate Professor of Pathology & Immunology Associate Professor of Pediatrics and Genetics Director of Cytogenomics and Molecular Pathology Evidence-based
Shashikant Kulkarni, M.S (Medicine)., Ph.D., FACMG Associate Professor of Pathology & Immunology Associate Professor of Pediatrics and Genetics Director of Cytogenomics and Molecular Pathology Evidence-based
Genome Summary. Sequencing Coverage: Variation Counts: Known Phenotype Summary: 131 disease or trait variations are found in this genome
 Report Date: August 2, 215 Software Annotation Version: 8 Genome Summary Name: Greg Mendel Genome ID: genome_greg_mendel_dad Sequencing Provider: 23andMe Sequencing Type: Genotyping SNP Array Sequencing
Report Date: August 2, 215 Software Annotation Version: 8 Genome Summary Name: Greg Mendel Genome ID: genome_greg_mendel_dad Sequencing Provider: 23andMe Sequencing Type: Genotyping SNP Array Sequencing
Nature Genetics: doi: /ng Supplementary Figure 1. Somatic coding mutations identified by WES/WGS for 83 ATL cases.
 Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Supplementary Materials
 1 Supplementary Materials Rotger et al. Table S1A: Demographic characteristics of study participants. VNP RP EC CP (n=6) (n=66) (n=9) (n=5) Male gender, n(%) 5 (83) 54 (82) 5 (56) 3 (60) White ethnicity,
1 Supplementary Materials Rotger et al. Table S1A: Demographic characteristics of study participants. VNP RP EC CP (n=6) (n=66) (n=9) (n=5) Male gender, n(%) 5 (83) 54 (82) 5 (56) 3 (60) White ethnicity,
Supplementary Figure 1 Information on transgenic mouse models and their recording and optogenetic equipment. (a) 108 (b-c) (d) (e) (f) (g)
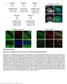 Supplementary Figure 1 Information on transgenic mouse models and their recording and optogenetic equipment. (a) In four mice, cre-dependent expression of the hyperpolarizing opsin Arch in pyramidal cells
Supplementary Figure 1 Information on transgenic mouse models and their recording and optogenetic equipment. (a) In four mice, cre-dependent expression of the hyperpolarizing opsin Arch in pyramidal cells
Numerous hypothesis tests were performed in this study. To reduce the false positive due to
 Two alternative data-splitting Numerous hypothesis tests were performed in this study. To reduce the false positive due to multiple testing, we are not only seeking the results with extremely small p values
Two alternative data-splitting Numerous hypothesis tests were performed in this study. To reduce the false positive due to multiple testing, we are not only seeking the results with extremely small p values
Studying Alternative Splicing
 Studying Alternative Splicing Meelis Kull PhD student in the University of Tartu supervisor: Jaak Vilo CS Theory Days Rõuge 27 Overview Alternative splicing Its biological function Studying splicing Technology
Studying Alternative Splicing Meelis Kull PhD student in the University of Tartu supervisor: Jaak Vilo CS Theory Days Rõuge 27 Overview Alternative splicing Its biological function Studying splicing Technology
Functional analysis of DNA variants
 Functional analysis of DNA variants GS011143, Introduction to Bioinformatics The University of Texas GSBS program, Fall 2012 Ken Chen, Ph.D. Department of Bioinformatics and Computational Biology UT MD
Functional analysis of DNA variants GS011143, Introduction to Bioinformatics The University of Texas GSBS program, Fall 2012 Ken Chen, Ph.D. Department of Bioinformatics and Computational Biology UT MD
ACE ImmunoID Biomarker Discovery Solutions ACE ImmunoID Platform for Tumor Immunogenomics
 ACE ImmunoID Biomarker Discovery Solutions ACE ImmunoID Platform for Tumor Immunogenomics Precision Genomics for Immuno-Oncology Personalis, Inc. ACE ImmunoID When one biomarker doesn t tell the whole
ACE ImmunoID Biomarker Discovery Solutions ACE ImmunoID Platform for Tumor Immunogenomics Precision Genomics for Immuno-Oncology Personalis, Inc. ACE ImmunoID When one biomarker doesn t tell the whole
Nature Medicine: doi: /nm.4439
 Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
Genética del Feocromocitoma/Paraganglioma.
 Genética del Feocromocitoma/Paraganglioma. Mercedes Robledo, PhD Head of the Hereditary Endocrine Cancer Group Human Cancer Genetics Programme CNIO, Madrid, Spain. mrobledo@cnio.es High susceptibility
Genética del Feocromocitoma/Paraganglioma. Mercedes Robledo, PhD Head of the Hereditary Endocrine Cancer Group Human Cancer Genetics Programme CNIO, Madrid, Spain. mrobledo@cnio.es High susceptibility
Large-scale identity-by-descent mapping discovers rare haplotypes of large effect. Suyash Shringarpure 23andMe, Inc. ASHG 2017
 Large-scale identity-by-descent mapping discovers rare haplotypes of large effect Suyash Shringarpure 23andMe, Inc. ASHG 2017 1 Why care about rare variants of large effect? Months from randomization 2
Large-scale identity-by-descent mapping discovers rare haplotypes of large effect Suyash Shringarpure 23andMe, Inc. ASHG 2017 1 Why care about rare variants of large effect? Months from randomization 2
Numerous hypothesis tests were performed in this study. To reduce the false positive due to
 Two alternative data-splitting Numerous hypothesis tests were performed in this study. To reduce the false positive due to multiple testing, we are not only seeking the results with extremely small p values
Two alternative data-splitting Numerous hypothesis tests were performed in this study. To reduce the false positive due to multiple testing, we are not only seeking the results with extremely small p values
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first
 Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
AD (Leave blank) TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients
 AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
Global variation in copy number in the human genome
 Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Frequency(%) KRAS G12 KRAS G13 KRAS A146 KRAS Q61 KRAS K117N PIK3CA H1047 PIK3CA E545 PIK3CA E542K PIK3CA Q546. EGFR exon19 NFS-indel EGFR L858R
 Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Nature Neuroscience: doi: /nn Supplementary Figure 1 Density plots of sequence coverage in the UK10K, INTERVAL and DDD data sets.
 Supplementary Figure 1 Density plots of sequence coverage in the UK10K, INTERVAL and DDD data sets. Per-sample sequence coverage was calculated and summarised from exome sequencing data generated in the
Supplementary Figure 1 Density plots of sequence coverage in the UK10K, INTERVAL and DDD data sets. Per-sample sequence coverage was calculated and summarised from exome sequencing data generated in the
Genetic Testing and Analysis. (858) MRN: Specimen: Saliva Received: 07/26/2016 GENETIC ANALYSIS REPORT
 GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
MSI positive MSI negative
 Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name and description (please provide any alternative names you wish listed) (A)-Testing
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name and description (please provide any alternative names you wish listed) (A)-Testing
Genomic structural variation
 Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
# For the GWAS stage, B-cell NHL cases which small numbers (N<20) were excluded from analysis.
 Supplementary Table 1a. Subtype Breakdown of all analyzed samples Stage GWAS Singapore Validation 1 Guangzhou Validation 2 Guangzhou Validation 3 Beijing Total No. of B-Cell Cases 253 # 168^ 294^ 713^
Supplementary Table 1a. Subtype Breakdown of all analyzed samples Stage GWAS Singapore Validation 1 Guangzhou Validation 2 Guangzhou Validation 3 Beijing Total No. of B-Cell Cases 253 # 168^ 294^ 713^
Supplementary Figure 1
 Supplementary Figure 1 An example of the gene-term-disease network automatically generated by Phenolyzer web server for 'autism'. The largest word represents the user s input term, Autism. The pink round
Supplementary Figure 1 An example of the gene-term-disease network automatically generated by Phenolyzer web server for 'autism'. The largest word represents the user s input term, Autism. The pink round
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
SCALPEL MICRO-ASSEMBLY APPROACH TO DETECT INDELS WITHIN EXOME-CAPTURE DATA. Giuseppe Narzisi, PhD Schatz Lab
 SCALPEL MICRO-ASSEMBLY APPROACH TO DETECT INDELS WITHIN EXOME-CAPTURE DATA Giuseppe Narzisi, PhD Schatz Lab November 14, 2013 Micro-Assembly Approach to detect INDELs 2 Outline Scalpel micro-assembly pipeline
SCALPEL MICRO-ASSEMBLY APPROACH TO DETECT INDELS WITHIN EXOME-CAPTURE DATA Giuseppe Narzisi, PhD Schatz Lab November 14, 2013 Micro-Assembly Approach to detect INDELs 2 Outline Scalpel micro-assembly pipeline
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
CONTENT SUPPLEMENTARY FIGURE E. INSTRUMENTAL VARIABLE ANALYSIS USING DESEASONALISED PLASMA 25-HYDROXYVITAMIN D. 7
 CONTENT FIGURES 3 SUPPLEMENTARY FIGURE A. NUMBER OF PARTICIPANTS AND EVENTS IN THE OBSERVATIONAL AND GENETIC ANALYSES. 3 SUPPLEMENTARY FIGURE B. FLOWCHART SHOWING THE SELECTION PROCESS FOR DETERMINING
CONTENT FIGURES 3 SUPPLEMENTARY FIGURE A. NUMBER OF PARTICIPANTS AND EVENTS IN THE OBSERVATIONAL AND GENETIC ANALYSES. 3 SUPPLEMENTARY FIGURE B. FLOWCHART SHOWING THE SELECTION PROCESS FOR DETERMINING
Field wide development of analytic approaches for sequence data
 Benjamin Neale Field wide development of analytic approaches for sequence data Cohort Allelic Sum Test (CAST; Hobbs, Cohen and others) Li and Leal (AJHG) Madsen and Browning (PLoS Genetics) C alpha and
Benjamin Neale Field wide development of analytic approaches for sequence data Cohort Allelic Sum Test (CAST; Hobbs, Cohen and others) Li and Leal (AJHG) Madsen and Browning (PLoS Genetics) C alpha and
Supplementary Table 1. Association of rs with risk of obesity among participants in NHS and HPFS
 Supplementary Table 1. Association of rs3826795 with risk of obesity among participants in NHS and HPFS Case/control NHS (1990) HPFS (1996) OR (95% CI) P- value Case/control OR (95% CI) P- value Obesity
Supplementary Table 1. Association of rs3826795 with risk of obesity among participants in NHS and HPFS Case/control NHS (1990) HPFS (1996) OR (95% CI) P- value Case/control OR (95% CI) P- value Obesity
Whole-genome detection of disease-associated deletions or excess homozygosity in a case control study of rheumatoid arthritis
 HMG Advance Access published December 21, 2012 Human Molecular Genetics, 2012 1 13 doi:10.1093/hmg/dds512 Whole-genome detection of disease-associated deletions or excess homozygosity in a case control
HMG Advance Access published December 21, 2012 Human Molecular Genetics, 2012 1 13 doi:10.1093/hmg/dds512 Whole-genome detection of disease-associated deletions or excess homozygosity in a case control
Nature Genetics: doi: /ng Supplementary Figure 1. Rates of different mutation types in CRC.
 Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Molecular Diagnosis of Genetic Diseases: From 1 Gene to 1000s
 ROMA, 23 GIUGNO 2011 Molecular Diagnosis of Genetic Diseases: From 1 Gene to 1000s USSD Lab Genetica Medica Ospedali Riuniti Bergamo The problem What does the clinician/patient want to know?? Diagnosis,
ROMA, 23 GIUGNO 2011 Molecular Diagnosis of Genetic Diseases: From 1 Gene to 1000s USSD Lab Genetica Medica Ospedali Riuniti Bergamo The problem What does the clinician/patient want to know?? Diagnosis,
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
Self reported ethnicity
 Self reported ethnicity Supplementary Figure 1 Ancestry stratifies patterns of human genetic variations. PCA plots (1 st, 2 nd and 3 rd components) estimated from human genotypes. Individuals are coloured
Self reported ethnicity Supplementary Figure 1 Ancestry stratifies patterns of human genetic variations. PCA plots (1 st, 2 nd and 3 rd components) estimated from human genotypes. Individuals are coloured
Supplementary Information. Supplementary Figures
 Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
Supplementary Material
 Supplementary Material 2 4 6 Single-locus gene screening methods We screened for single nucleotide polymorphisms (SNPs) at 16 candidate immune genes (Table S1) by initially sequencing three captive-bred
Supplementary Material 2 4 6 Single-locus gene screening methods We screened for single nucleotide polymorphisms (SNPs) at 16 candidate immune genes (Table S1) by initially sequencing three captive-bred
DNA is the genetic material that provides instructions for what our bodies look like and how they function. DNA is packaged into structures called
 DNA is the genetic material that provides instructions for what our bodies look like and how they function. DNA is packaged into structures called chromosomes. We have 23 pairs of chromosomes (for a total
DNA is the genetic material that provides instructions for what our bodies look like and how they function. DNA is packaged into structures called chromosomes. We have 23 pairs of chromosomes (for a total
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Epileptic encephalopathy, early infantile 4. OMIM number for disease 612164 Disease
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Epileptic encephalopathy, early infantile 4. OMIM number for disease 612164 Disease
Evaluating Classifiers for Disease Gene Discovery
 Evaluating Classifiers for Disease Gene Discovery Kino Coursey Lon Turnbull khc0021@unt.edu lt0013@unt.edu Abstract Identification of genes involved in human hereditary disease is an important bioinfomatics
Evaluating Classifiers for Disease Gene Discovery Kino Coursey Lon Turnbull khc0021@unt.edu lt0013@unt.edu Abstract Identification of genes involved in human hereditary disease is an important bioinfomatics
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Statistical Tests for X Chromosome Association Study. with Simulations. Jian Wang July 10, 2012
 Statistical Tests for X Chromosome Association Study with Simulations Jian Wang July 10, 2012 Statistical Tests Zheng G, et al. 2007. Testing association for markers on the X chromosome. Genetic Epidemiology
Statistical Tests for X Chromosome Association Study with Simulations Jian Wang July 10, 2012 Statistical Tests Zheng G, et al. 2007. Testing association for markers on the X chromosome. Genetic Epidemiology
