Ginkgo Interactive analysis and quality assessment of single-cell CNV data
|
|
|
- Gervais Page
- 5 years ago
- Views:
Transcription
1 Ginkgo Interactive analysis and quality assessment of single-cell CNV Robert Aboukhalil, Tyler Garvin, Jude Kendall, Timour Baslan, Gurinder S. Atwal, Jim Hicks, Michael Wigler, Michael C. Schatz
2 Outline Introduction Comparison of WGA methods Ginkgo
3 Outline Introduction Comparison of WGA methods Ginkgo
4 Single-cell sequencing Circulating tumor cells Neuronal mosaicism Clonal evolution in tumors Recombination/ crossover in germ cells
5 Single-cell vs. bulk sequencing Brian Owens, Nature News 2012
6 Single-cell vs. bulk sequencing Brian Owens, Nature News 2012
7 Copy-number variant analysis Low coverage allows us to study copy-number variants <1X$coverage,$o-en$<0.1X$
8 Copy-number variant analysis NNNNNN" Low coverage allows us to study copy-number variants <1X$coverage,$o-en$<0.1X$
9 Copy-number variant analysis NNNNNN" Divide genome into bins with ~ reads / bin Baslan et al., Nature Protocols 2012
10 Copy-number variant analysis NNNNNN"
11 Copy-number variant analysis NNNNNN"
12 Copy-number variant analysis NNNNNN" Circular Binary Segmentation (CBS) to reduce noise in data
13 Copy-number variant analysis We can estimate integer copy-number states by scaling the profile and minimizing the sum of squares error
14 Outline Introduction Comparison of WGA methods Ginkgo
15 Copy-number variant analysis
16 Analysis Pipeline Bin reads Map reads GC bias correction.fastq$ Remove duplicates BAM to BED.BED$ Segment bins and call integer copy-number Galaxy Interactive visualization Ginkgo
17 Ginkgo Demo
18 Sample dataset Triple-negative (ER, PR, HER2 ) ductal carcinoma Navin et al., Nature 2011
19 QB.CSHL.EDU/GINKGO
20 QB.CSHL.EDU/GINKGO
21 QB.CSHL.EDU/GINKGO
22 QB.CSHL.EDU/GINKGO
23 Diploids from both primary and metastasis cluster QB.CSHL.EDU/GINKGO
24 Diploids from both primary and metastasis cluster QB.CSHL.EDU/GINKGO
25 Non-diploids from primary and metastasis segregate QB.CSHL.EDU/GINKGO
26 Non-diploids from primary and metastasis segregate QB.CSHL.EDU/GINKGO
27 Non-diploids from primary and metastasis segregate QB.CSHL.EDU/GINKGO
28 Non-diploids from primary and metastasis segregate QB.CSHL.EDU/GINKGO
29 QB.CSHL.EDU/GINKGO
30 QB.CSHL.EDU/GINKGO
31 QB.CSHL.EDU/GINKGO
32 QB.CSHL.EDU/GINKGO
33 QB.CSHL.EDU/GINKGO
34 QB.CSHL.EDU/GINKGO Although primary and metastasis cells are similar, they are slightly different from each other
35 QB.CSHL.EDU/GINKGO Heatmap also tells us there are 3 sub populations: 1 diploid, 1 primary, 1 metastasis
36 QB.CSHL.EDU/GINKGO Can click on cells to get indepth view
37 QB.CSHL.EDU/GINKGO
38 QB.CSHL.EDU/GINKGO Interactive profile viewer allows you to zoom into a region of interest
39 QB.CSHL.EDU/GINKGO
40 QB.CSHL.EDU/GINKGO We automatically upload amplification and deletion tracks to the UCSC browser
41 QB.CSHL.EDU/GINKGO We can also annotate the CNV profile with genes of interest
42 QB.CSHL.EDU/GINKGO
43 QB.CSHL.EDU/GINKGO
44 QB.CSHL.EDU/GINKGO
45 QB.CSHL.EDU/GINKGO
46 QB.CSHL.EDU/GINKGO
47 QB.CSHL.EDU/GINKGO
48 QB.CSHL.EDU/GINKGO
49 QB.CSHL.EDU/GINKGO Static CNV profile
50 QB.CSHL.EDU/GINKGO
51 Original read count data before analysis QC metrics QB.CSHL.EDU/GINKGO
52 QB.CSHL.EDU/GINKGO GC bias, before and after correction
53 QB.CSHL.EDU/GINKGO How we infer integer copy-number state using sum of squares error
54 Outline Introduction Comparison of WGA methods Ginkgo
55 Whole Genome Amplification (WGA) methods DOP-PCR (Degenerate Oligonucleotide Primed PCR) MDA (Multiple Displacement Amplification) MALBAC (Multiple Annealing and Looping Based Amplification Cycles) Paul Blainey, FEMS Microbiol Rev. 2013
56 Comparison of WGA methods Paper WGA Method Tissue Navin et al., 2011 DOP-PCR Breast (T10) Navin et al., 2011 DOP-PCR Breast (T16P/M) McConnnell et al., 2013 DOP-PCR Neuron Lu et al., 2012 MALBAC Sperm Ni et al., 2013 MALBAC Lung Hou et al., 2013 MALBAC Oocyte Kirkness et al., 2013 MDA Sperm Wang et al., 2012 MDA Sperm Evrony et al., 2012 MDA Neuron Explore the effects of WGA method on data quality: 1) GC bias 2) Coverage dispersion Garvin and Aboukhalil et al., Nature Methods, 2015
57 GC Bias Garvin and Aboukhalil et al., Nature Methods, 2015
58 Coverage Dispersion DOP-PCR MALBAC MDA Garvin and Aboukhalil et al., Nature Methods, 2015
59 Coverage Dispersion DOP-PCR MALBAC MDA Garvin and Aboukhalil et al., Nature Methods, 2015
60 Summary Ginkgo is a platform for single-cell CNV analysis and visualization For copy-number analysis, we recommend DOP-PCR Check out Ginkgo and give us feedback qb.cshl.edu/ginkgo Garvin and Aboukhalil et al., Nature Methods, 2015 > 1K USERS 21,500 PAGEVIEWS$ 10 MIN TIME SPENT Nov 3, San Francisco
61 Thanks Ginkgo Team Tyler Garvin Jude Kendall Timour Baslan Jim Hicks Gurinder S. Atwal Michael Wigler Michael C. Schatz qb.cshl.edu/ginkgo
Interactive analysis and quality assessment of single-cell copy-number variations
 Interactive analysis and quality assessment of single-cell copy-number variations Tyler Garvin, Robert Aboukhalil, Jude Kendall, Timour Baslan, Gurinder S. Atwal, James Hicks, Michael Wigler, Michael C.
Interactive analysis and quality assessment of single-cell copy-number variations Tyler Garvin, Robert Aboukhalil, Jude Kendall, Timour Baslan, Gurinder S. Atwal, James Hicks, Michael Wigler, Michael C.
MALBAC Technology and Its Application in Non-invasive Chromosome Screening (NICS)
 MALBAC Technology and Its Application in Non-invasive Chromosome Screening (NICS) The Power of One Adapted from Internet Single Cell Genomic Studies Ultra Low Sample Input Advances and applications of
MALBAC Technology and Its Application in Non-invasive Chromosome Screening (NICS) The Power of One Adapted from Internet Single Cell Genomic Studies Ultra Low Sample Input Advances and applications of
Investigating Breast Cancer Evolution with Single Cell Genomics
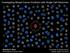 Investigating Breast Cancer Evolution with Single Cell Genomics AACR Drug Sensitivity & Resistance June 21 st, 2014 Nicholas E. Navin MD Anderson Cancer Center Innovation in Breast Cancer Madrid, Spain
Investigating Breast Cancer Evolution with Single Cell Genomics AACR Drug Sensitivity & Resistance June 21 st, 2014 Nicholas E. Navin MD Anderson Cancer Center Innovation in Breast Cancer Madrid, Spain
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Analysis with SureCall 2.1
 Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
MPS for translocations
 MPS for translocations Filip Van Nieuwerburgh, Ph.D. Lab of Pharmaceutical Biotechnology NXTGNT massively parallel sequencing facility, Ghent University In collaboration with: Center for Medical Genetics,
MPS for translocations Filip Van Nieuwerburgh, Ph.D. Lab of Pharmaceutical Biotechnology NXTGNT massively parallel sequencing facility, Ghent University In collaboration with: Center for Medical Genetics,
Global variation in copy number in the human genome
 Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Single-Cell Sequencing in Cancer. Peter A. Sims, Columbia University G4500: Cellular & Molecular Biology of Cancer October 22, 2018
 Single-Cell Sequencing in Cancer Peter A. Sims, Columbia University G4500: Cellular & Molecular Biology of Cancer October 22, 2018 Lecture will focus on technology for and applications of single-cell RNA-seq
Single-Cell Sequencing in Cancer Peter A. Sims, Columbia University G4500: Cellular & Molecular Biology of Cancer October 22, 2018 Lecture will focus on technology for and applications of single-cell RNA-seq
Clonal Evolution in Breast Cancer Revealed by Single Nucleus Genome Sequencing
 Clonal Evolution in Breast Cancer Revealed by Single Nucleus Genome Sequencing The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters.
Clonal Evolution in Breast Cancer Revealed by Single Nucleus Genome Sequencing The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters.
Genome. Institute. GenomeVIP: A Genomics Analysis Pipeline for Cloud Computing with Germline and Somatic Calling on Amazon s Cloud. R. Jay Mashl.
 GenomeVIP: the Genome Institute at Washington University A Genomics Analysis Pipeline for Cloud Computing with Germline and Somatic Calling on Amazon s Cloud R. Jay Mashl October 20, 2014 Turnkey Variant
GenomeVIP: the Genome Institute at Washington University A Genomics Analysis Pipeline for Cloud Computing with Germline and Somatic Calling on Amazon s Cloud R. Jay Mashl October 20, 2014 Turnkey Variant
The Comparison of the Performance of Four Whole Genome Amplification Kits on Ion Proton Platform in Copy Number Variation Detection
 Bioscience Reports: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
Bioscience Reports: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University of Cambridge, Clara East, The Old Schools, Cambridge CB2 1TN
 AWARD NUMBER: W81XWH-14-1-0110 TITLE: A Molecular Framework for Understanding DCIS PRINCIPAL INVESTIGATOR: Gregory Hannon CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University
AWARD NUMBER: W81XWH-14-1-0110 TITLE: A Molecular Framework for Understanding DCIS PRINCIPAL INVESTIGATOR: Gregory Hannon CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University
Understanding DNA Copy Number Data
 Understanding DNA Copy Number Data Adam B. Olshen Department of Epidemiology and Biostatistics Helen Diller Family Comprehensive Cancer Center University of California, San Francisco http://cc.ucsf.edu/people/olshena_adam.php
Understanding DNA Copy Number Data Adam B. Olshen Department of Epidemiology and Biostatistics Helen Diller Family Comprehensive Cancer Center University of California, San Francisco http://cc.ucsf.edu/people/olshena_adam.php
Sensitivity to sequencing depth in singlecell cancer genomics
 Alves and Posada Genome Medicine (208) :29 https://doi.org/.86/s3073-08-037-2 RESEARCH Sensitivity to sequencing depth in singlecell cancer genomics João M. Alves,2,3* and David Posada,2,3* Open Access
Alves and Posada Genome Medicine (208) :29 https://doi.org/.86/s3073-08-037-2 RESEARCH Sensitivity to sequencing depth in singlecell cancer genomics João M. Alves,2,3* and David Posada,2,3* Open Access
CNV PCA Search Tutorial
 CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
Alper Sarikaya 1, Michael Correll 2, Jorge M. Dinis 1, David H. O Connor 1,3, and Michael Gleicher 1
 Alper Sarikaya 1, Michael Correll 2, Jorge M. Dinis 1, David H. O Connor 1,3, and Michael Gleicher 1 1 University of Wisconsin-Madison 2 University of Washington 3 Wisconsin National Primate Center @yelperalp
Alper Sarikaya 1, Michael Correll 2, Jorge M. Dinis 1, David H. O Connor 1,3, and Michael Gleicher 1 1 University of Wisconsin-Madison 2 University of Washington 3 Wisconsin National Primate Center @yelperalp
Detection of copy number variations in PCR-enriched targeted sequencing data
 Detection of copy number variations in PCR-enriched targeted sequencing data German Demidov Parseq Lab, Saint-Petersburg University of Russian Academy of Sciences, current: Center for Genomic Regulation
Detection of copy number variations in PCR-enriched targeted sequencing data German Demidov Parseq Lab, Saint-Petersburg University of Russian Academy of Sciences, current: Center for Genomic Regulation
SCALPEL MICRO-ASSEMBLY APPROACH TO DETECT INDELS WITHIN EXOME-CAPTURE DATA. Giuseppe Narzisi, PhD Schatz Lab
 SCALPEL MICRO-ASSEMBLY APPROACH TO DETECT INDELS WITHIN EXOME-CAPTURE DATA Giuseppe Narzisi, PhD Schatz Lab November 14, 2013 Micro-Assembly Approach to detect INDELs 2 Outline Scalpel micro-assembly pipeline
SCALPEL MICRO-ASSEMBLY APPROACH TO DETECT INDELS WITHIN EXOME-CAPTURE DATA Giuseppe Narzisi, PhD Schatz Lab November 14, 2013 Micro-Assembly Approach to detect INDELs 2 Outline Scalpel micro-assembly pipeline
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS
 Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Agilent s Copy Number Variation (CNV) Portfolio
 Technical Overview Agilent s Copy Number Variation (CNV) Portfolio Abstract Copy Number Variation (CNV) is now recognized as a prevalent form of structural variation in the genome contributing to human
Technical Overview Agilent s Copy Number Variation (CNV) Portfolio Abstract Copy Number Variation (CNV) is now recognized as a prevalent form of structural variation in the genome contributing to human
Illuminating the genetics of complex human diseases
 Illuminating the genetics of complex human diseases Michael Schatz Sept 27, 2012 Beyond the Genome @mike_schatz / #BTG2012 Outline 1. De novo mutations in human diseases 1. Autism Spectrum Disorder 2.
Illuminating the genetics of complex human diseases Michael Schatz Sept 27, 2012 Beyond the Genome @mike_schatz / #BTG2012 Outline 1. De novo mutations in human diseases 1. Autism Spectrum Disorder 2.
Computer Science, Biology, and Biomedical Informatics (CoSBBI) Outline. Molecular Biology of Cancer AND. Goals/Expectations. David Boone 7/1/2015
 Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University of Cambridge, Clara East, The Old Schools, Cambridge CB2 1TN
 AWARD NUMBER: W81XWH-14-1-0110 TITLE: A Molecular Framework for Understanding DCIS PRINCIPAL INVESTIGATOR: Gregory Hannon CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University
AWARD NUMBER: W81XWH-14-1-0110 TITLE: A Molecular Framework for Understanding DCIS PRINCIPAL INVESTIGATOR: Gregory Hannon CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit
 APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
Reducing INDEL calling errors in whole genome and exome sequencing data.
 Reducing INDEL calling errors in whole genome and exome sequencing data. Han Fang November 8, 2014 CSHL Biological Data Science Meeting Acknowledgments Lyon Lab Yiyang Wu Jason O Rawe Laura J Barron Max
Reducing INDEL calling errors in whole genome and exome sequencing data. Han Fang November 8, 2014 CSHL Biological Data Science Meeting Acknowledgments Lyon Lab Yiyang Wu Jason O Rawe Laura J Barron Max
Integrated Analysis of Copy Number and Gene Expression
 Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer
 Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
Genome-wide copy-number calling (CNAs not CNVs!) Dr Geoff Macintyre
 Genome-wide copy-number calling (CNAs not CNVs!) Dr Geoff Macintyre Structural variation (SVs) Copy-number variations C Deletion A B C Balanced rearrangements A B A B C B A C Duplication Inversion Causes
Genome-wide copy-number calling (CNAs not CNVs!) Dr Geoff Macintyre Structural variation (SVs) Copy-number variations C Deletion A B C Balanced rearrangements A B A B C B A C Duplication Inversion Causes
Molecular Characterization of Tumors Using Next-Generation Sequencing
 Molecular Characterization of Tumors Using Next-Generation Sequencing Using BaseSpace to visualize molecular changes in cancer. Tumor-Normal Sequencing Data in BaseSpace To enable researchers new to next-generation
Molecular Characterization of Tumors Using Next-Generation Sequencing Using BaseSpace to visualize molecular changes in cancer. Tumor-Normal Sequencing Data in BaseSpace To enable researchers new to next-generation
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Complete Genomics, Inc.
 Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Introduction to Systems Biology of Cancer Lecture 2
 Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
MODULE 4: SPLICING. Removal of introns from messenger RNA by splicing
 Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
The Chancellor, Masters and Scholars of the University of Cambridge, Clara East, The Old Schools, Cambridge CB2 1TN
 AWARD NUMBER: W81XWH-14-1-0110 TITLE: A Molecular Framework for Understanding DCIS PRINCIPAL INVESTIGATOR: Gregory Hannon CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University
AWARD NUMBER: W81XWH-14-1-0110 TITLE: A Molecular Framework for Understanding DCIS PRINCIPAL INVESTIGATOR: Gregory Hannon CONTRACTING ORGANIZATION: The Chancellor, Masters and Scholars of the University
Chip Seq Peak Calling in Galaxy
 Chip Seq Peak Calling in Galaxy Chris Seward PowerPoint by Pei-Chen Peng Chip-Seq Peak Calling in Galaxy Chris Seward 2018 1 Introduction This goals of the lab are as follows: 1. Gain experience using
Chip Seq Peak Calling in Galaxy Chris Seward PowerPoint by Pei-Chen Peng Chip-Seq Peak Calling in Galaxy Chris Seward 2018 1 Introduction This goals of the lab are as follows: 1. Gain experience using
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
Characterisation of structural variation in breast. cancer genomes using paired-end sequencing on. the Illumina Genome Analyser
 Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Carcinome du sein Biologie moléculaire. Thomas McKee Service de Pathologie Clinique Genève
 Carcinome du sein Biologie moléculaire Thomas McKee Service de Pathologie Clinique Genève Pathology Diagnostic Prognostic information Predictive information The information provided depends on the available
Carcinome du sein Biologie moléculaire Thomas McKee Service de Pathologie Clinique Genève Pathology Diagnostic Prognostic information Predictive information The information provided depends on the available
DETECTION OF LOW FREQUENCY CXCR4-USING HIV-1 WITH ULTRA-DEEP PYROSEQUENCING. John Archer. Faculty of Life Sciences University of Manchester
 DETECTION OF LOW FREQUENCY CXCR4-USING HIV-1 WITH ULTRA-DEEP PYROSEQUENCING John Archer Faculty of Life Sciences University of Manchester HIV Dynamics and Evolution, 2008, Santa Fe, New Mexico. Overview
DETECTION OF LOW FREQUENCY CXCR4-USING HIV-1 WITH ULTRA-DEEP PYROSEQUENCING John Archer Faculty of Life Sciences University of Manchester HIV Dynamics and Evolution, 2008, Santa Fe, New Mexico. Overview
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Data mining with Ensembl Biomart. Stéphanie Le Gras
 Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
The Cancer Genome Atlas & International Cancer Genome Consortium
 The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
Comprehensive Genome and Transcriptome Structural Analysis of a Breast Cancer Cell Line using PacBio Long Read Sequencing
 Comprehensive Genome and Transcriptome Structural Analysis of a Breast Cancer Cell Line using PacBio Long Read Sequencing Maria Nattestad Schatz + McCombie + Hicks at Cold Spring Harbor Laboratory McPherson
Comprehensive Genome and Transcriptome Structural Analysis of a Breast Cancer Cell Line using PacBio Long Read Sequencing Maria Nattestad Schatz + McCombie + Hicks at Cold Spring Harbor Laboratory McPherson
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Introduction to genetic variation. He Zhang Bioinformatics Core Facility 6/22/2016
 Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
PG-Seq NGS Kit for Preimplantation Genetic Screening
 Application Note: PG-Seq Validation Study PG-Seq NGS Kit for Preimplantation Genetic Screening Validation using Multi (5-10) Cells and Single Cells from euploid and aneuploid cell lines Introduction Advances
Application Note: PG-Seq Validation Study PG-Seq NGS Kit for Preimplantation Genetic Screening Validation using Multi (5-10) Cells and Single Cells from euploid and aneuploid cell lines Introduction Advances
Sexual Reproduction & Inheritance
 Sexual Reproduction & Sexual Reproduction & Overview Asexual vs Sexual Reproduction Meiosis Genetic Diversity Mendel & The Laws of Sexual Reproduction Sexual Reproduction Asexual Reproduction Prokaryotes
Sexual Reproduction & Sexual Reproduction & Overview Asexual vs Sexual Reproduction Meiosis Genetic Diversity Mendel & The Laws of Sexual Reproduction Sexual Reproduction Asexual Reproduction Prokaryotes
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Below, we included the point-to-point response to the comments of both reviewers.
 To the Editor and Reviewers: We would like to thank the editor and reviewers for careful reading, and constructive suggestions for our manuscript. According to comments from both reviewers, we have comprehensively
To the Editor and Reviewers: We would like to thank the editor and reviewers for careful reading, and constructive suggestions for our manuscript. According to comments from both reviewers, we have comprehensively
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Copy Number Varia/on Detec/on. Alex Mawla UCD Genome Center Bioinforma5cs Core Tuesday June 16, 2015
 Copy Number Varia/on Detec/on Alex Mawla UCD Genome Center Bioinforma5cs Core Tuesday June 16, 2015 Today s Goals Understand the applica5on and capabili5es of using targe5ng sequencing and CNV calling
Copy Number Varia/on Detec/on Alex Mawla UCD Genome Center Bioinforma5cs Core Tuesday June 16, 2015 Today s Goals Understand the applica5on and capabili5es of using targe5ng sequencing and CNV calling
Core Technology Development Team Meeting
 Core Technology Development Team Meeting To hear the meeting, you must call in Toll-free phone number: 1-866-740-1260 Access Code: 2201876 For international call in numbers, please visit: https://www.readytalk.com/account-administration/international-numbers
Core Technology Development Team Meeting To hear the meeting, you must call in Toll-free phone number: 1-866-740-1260 Access Code: 2201876 For international call in numbers, please visit: https://www.readytalk.com/account-administration/international-numbers
Nature Genetics: doi: /ng Supplementary Figure 1. Rates of different mutation types in CRC.
 Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Investigating rare diseases with Agilent NGS solutions
 Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Targeted qpcr. Debate on PGS Technology: Targeted vs. Whole genome approach. Discolsure Stake shareholder of GENETYX S.R.L
 Antonio Capalbo, PhD Laboratory Director GENETYX, reproductive genetics laboratory, Italy PGT responsible GENERA centers for reproductive medicine, Italy Debate on PGS Technology: Targeted vs. Whole genome
Antonio Capalbo, PhD Laboratory Director GENETYX, reproductive genetics laboratory, Italy PGT responsible GENERA centers for reproductive medicine, Italy Debate on PGS Technology: Targeted vs. Whole genome
Network-assisted data analysis
 Network-assisted data analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Protein identification in shotgun proteomics Protein digestion LC-MS/MS Protein
Network-assisted data analysis Bing Zhang Department of Biomedical Informatics Vanderbilt University bing.zhang@vanderbilt.edu Protein identification in shotgun proteomics Protein digestion LC-MS/MS Protein
Nature Biotechnology: doi: /nbt.1904
 Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
HBV. Next Generation Sequencing, data analysis and reporting. Presenter Leen-Jan van Doorn
 HBV Next Generation Sequencing, data analysis and reporting Presenter Leen-Jan van Doorn HBV Forum 3 October 24 th, 2017 Marriott Marquis, Washington DC www.forumresearch.org HBV Biomarkers HBV biomarkers:
HBV Next Generation Sequencing, data analysis and reporting Presenter Leen-Jan van Doorn HBV Forum 3 October 24 th, 2017 Marriott Marquis, Washington DC www.forumresearch.org HBV Biomarkers HBV biomarkers:
White Paper. Copy number variant detection. Sample to Insight. August 19, 2015
 White Paper Copy number variant detection August 19, 2015 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 Fax: +45 86 20 12 22 www.clcbio.com
White Paper Copy number variant detection August 19, 2015 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 Fax: +45 86 20 12 22 www.clcbio.com
Comprehensive Chromosome Screening Is NextGen Likely to be the Final Best Platform and What are its Advantages and Quirks?
 Comprehensive Chromosome Screening Is NextGen Likely to be the Final Best Platform and What are its Advantages and Quirks? Embryo 1 Embryo 2 combine samples for a single sequencing chip Barcode 1 CTAAGGTAAC
Comprehensive Chromosome Screening Is NextGen Likely to be the Final Best Platform and What are its Advantages and Quirks? Embryo 1 Embryo 2 combine samples for a single sequencing chip Barcode 1 CTAAGGTAAC
Research Strategy: 1. Background and Significance
 Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Figure S4. 15 Mets Whole Exome. 5 Primary Tumors Cancer Panel and WES. Next Generation Sequencing
 Figure S4 Next Generation Sequencing 15 Mets Whole Exome 5 Primary Tumors Cancer Panel and WES Get coverage of all variant loci for all three Mets Variant Filtering Sequence Alignments Index and align
Figure S4 Next Generation Sequencing 15 Mets Whole Exome 5 Primary Tumors Cancer Panel and WES Get coverage of all variant loci for all three Mets Variant Filtering Sequence Alignments Index and align
Titrations in Cytobank
 The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
ncounter Data Analysis Guidelines for Copy Number Variation (CNV) Molecules That Count NanoString Technologies, Inc.
 ncounter Data Analysis Guidelines for Copy Number Variation (CNV) NanoString Technologies, Inc. 530 Fairview Ave N Suite 2000 Seattle, Washington 98109 www.nanostring.com Tel: 206.378.6266 888.358.6266
ncounter Data Analysis Guidelines for Copy Number Variation (CNV) NanoString Technologies, Inc. 530 Fairview Ave N Suite 2000 Seattle, Washington 98109 www.nanostring.com Tel: 206.378.6266 888.358.6266
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
No evidence of clonally selected somatic genomic alterations in cancer associated
 Supplementary Data Resource No evidence of clonally selected somatic genomic alterations in cancer associated fibroblasts from human breast and ovarian carcinomas Wen Qiu, Min Hu, Anita Sridhar, Ken Opeskin,
Supplementary Data Resource No evidence of clonally selected somatic genomic alterations in cancer associated fibroblasts from human breast and ovarian carcinomas Wen Qiu, Min Hu, Anita Sridhar, Ken Opeskin,
Performance Characteristics BRCA MASTR Plus Dx
 Performance Characteristics BRCA MASTR Plus Dx with drmid Dx for Illumina NGS systems Manufacturer Multiplicom N.V. Galileïlaan 18 2845 Niel Belgium Table of Contents 1. Workflow... 4 2. Performance Characteristics
Performance Characteristics BRCA MASTR Plus Dx with drmid Dx for Illumina NGS systems Manufacturer Multiplicom N.V. Galileïlaan 18 2845 Niel Belgium Table of Contents 1. Workflow... 4 2. Performance Characteristics
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: W81XWH-12-1-0298 TITLE: MTHFR Functional Polymorphism C677T and Genomic Instability in the Etiology of Idiopathic Autism in Simplex Families PRINCIPAL INVESTIGATOR: Xudong Liu, PhD CONTRACTING
AD Award Number: W81XWH-12-1-0298 TITLE: MTHFR Functional Polymorphism C677T and Genomic Instability in the Etiology of Idiopathic Autism in Simplex Families PRINCIPAL INVESTIGATOR: Xudong Liu, PhD CONTRACTING
Associating Copy Number and SNP Variation with Human Disease. Autism Segmental duplication Neurobehavioral, includes social disability
 Technical Note Associating Copy Number and SNP Variation with Human Disease Abstract The Genome-Wide Human SNP Array 6.0 is an affordable tool to examine the role of copy number variation in disease by
Technical Note Associating Copy Number and SNP Variation with Human Disease Abstract The Genome-Wide Human SNP Array 6.0 is an affordable tool to examine the role of copy number variation in disease by
Clinical evaluation of microarray data
 Clinical evaluation of microarray data David Amor 19 th June 2011 Single base change Microarrays 3-4Mb What is a microarray? Up to 10 6 bits of Information!! Highly multiplexed FISH hybridisations. Microarray
Clinical evaluation of microarray data David Amor 19 th June 2011 Single base change Microarrays 3-4Mb What is a microarray? Up to 10 6 bits of Information!! Highly multiplexed FISH hybridisations. Microarray
Trinity: Transcriptome Assembly for Genetic and Functional Analysis of Cancer [U24]
![Trinity: Transcriptome Assembly for Genetic and Functional Analysis of Cancer [U24] Trinity: Transcriptome Assembly for Genetic and Functional Analysis of Cancer [U24]](/thumbs/87/95271099.jpg) Trinity: Transcriptome Assembly for Genetic and Functional Analysis of Cancer [U24] ITCR meeting, June 2016 The Cancer Transcriptome A window into the (expressed) genetic and epigenetic state of a tumor
Trinity: Transcriptome Assembly for Genetic and Functional Analysis of Cancer [U24] ITCR meeting, June 2016 The Cancer Transcriptome A window into the (expressed) genetic and epigenetic state of a tumor
Single-Cell, Genome-wide Sequencing Identifies Clonal Somatic Copy-Number Variation in the Human Brain
 Cell Reports Report Single-Cell, Genome-wide Sequencing Identifies Clonal Somatic Copy-Number Variation in the Human Brain Xuyu Cai, 1,2,3,4,6 Gilad D. Evrony, 1,2,3,4 Hillel S. Lehmann, 1,2,3 Princess
Cell Reports Report Single-Cell, Genome-wide Sequencing Identifies Clonal Somatic Copy-Number Variation in the Human Brain Xuyu Cai, 1,2,3,4,6 Gilad D. Evrony, 1,2,3,4 Hillel S. Lehmann, 1,2,3 Princess
New technologies reaching the clinic
 New technologies reaching the clinic Martin Däumer May 31, 2018 Deep-sequencing Standard Sanger-sequencing...PQIYMDDHTRE... Ultra-deep-sequencing...PQIYMDDHTRE......PQIYMDDHTRE......PQIYVDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE...
New technologies reaching the clinic Martin Däumer May 31, 2018 Deep-sequencing Standard Sanger-sequencing...PQIYMDDHTRE... Ultra-deep-sequencing...PQIYMDDHTRE......PQIYMDDHTRE......PQIYVDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE......PQIYMDDHTRE...
A METHOD FOR FINDING NOVEL ASSOCIATIONS BETWEEN GENOME-WIDE COPY NUMBER AND DNA METHYLATION PATTERNS
 A METHOD FOR FINDING NOVEL ASSOCIATIONS BETWEEN GENOME-WIDE COPY NUMBER AND DNA METHYLATION PATTERNS Man-Hung Eric Tang 1), Vinay Varadan 2), Sid Kamalakaran 2), Michael Q. Zhang 3), Nevenka Dimitrova
A METHOD FOR FINDING NOVEL ASSOCIATIONS BETWEEN GENOME-WIDE COPY NUMBER AND DNA METHYLATION PATTERNS Man-Hung Eric Tang 1), Vinay Varadan 2), Sid Kamalakaran 2), Michael Q. Zhang 3), Nevenka Dimitrova
SINGLE-CELL SEQUENCING
 SINGLE-CELL SEQUENCING Jaena Han Professor: Doug Brutlag, PhD : Genomics, Bioinformatics, and Medicine Introduction Jaena Han Once considered a technical feat only possible for a few specialized labs,
SINGLE-CELL SEQUENCING Jaena Han Professor: Doug Brutlag, PhD : Genomics, Bioinformatics, and Medicine Introduction Jaena Han Once considered a technical feat only possible for a few specialized labs,
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
PSSV User Manual (V2.1)
 PSSV User Manual (V2.1) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
PSSV User Manual (V2.1) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
The feasibility of circulating tumour DNA as an alternative to biopsy for mutational characterization in Stage III melanoma patients
 The feasibility of circulating tumour DNA as an alternative to biopsy for mutational characterization in Stage III melanoma patients ASSC Scientific Meeting 13 th October 2016 Prof Andrew Barbour UQ SOM
The feasibility of circulating tumour DNA as an alternative to biopsy for mutational characterization in Stage III melanoma patients ASSC Scientific Meeting 13 th October 2016 Prof Andrew Barbour UQ SOM
v Feature Stamping SMS 12.0 Tutorial Prerequisites Requirements TABS model Map Module Mesh Module Scatter Module Time minutes
 v. 12.0 SMS 12.0 Tutorial Objectives In this lesson will teach how to use conceptual modeling techniques to create numerical models that incorporate flow control structures into existing bathymetry. The
v. 12.0 SMS 12.0 Tutorial Objectives In this lesson will teach how to use conceptual modeling techniques to create numerical models that incorporate flow control structures into existing bathymetry. The
Statistical Analysis of Single Nucleotide Polymorphism Microarrays in Cancer Studies
 Statistical Analysis of Single Nucleotide Polymorphism Microarrays in Cancer Studies Stanford Biostatistics Workshop Pierre Neuvial with Henrik Bengtsson and Terry Speed Department of Statistics, UC Berkeley
Statistical Analysis of Single Nucleotide Polymorphism Microarrays in Cancer Studies Stanford Biostatistics Workshop Pierre Neuvial with Henrik Bengtsson and Terry Speed Department of Statistics, UC Berkeley
Networks of gene expression and brain function
 Networks of gene expression and brain function OHBM 25 June 2017 VILAS MENON HHMI JANELIA RESEARCH CAMPUS Introduction to Imaging Genetics A brief history of imaging genetics JB Poline The modern day endophenotype
Networks of gene expression and brain function OHBM 25 June 2017 VILAS MENON HHMI JANELIA RESEARCH CAMPUS Introduction to Imaging Genetics A brief history of imaging genetics JB Poline The modern day endophenotype
Same Day, Cost-Effective Aneuploidy Detection with Agilent Oligonucleotide array CGH and MDA Single Cell Amplification Method
 Same Day, Cost-Effective Aneuploidy Detection with Agilent Oligonucleotide array CGH and MDA Single Cell Amplification Method Presenter: Dr. Ali Hellani, Founder, Viafet Genomic Center, Dubai Wednesday,
Same Day, Cost-Effective Aneuploidy Detection with Agilent Oligonucleotide array CGH and MDA Single Cell Amplification Method Presenter: Dr. Ali Hellani, Founder, Viafet Genomic Center, Dubai Wednesday,
Identification of regions with common copy-number variations using SNP array
 Identification of regions with common copy-number variations using SNP array Agus Salim Epidemiology and Public Health National University of Singapore Copy Number Variation (CNV) Copy number alteration
Identification of regions with common copy-number variations using SNP array Agus Salim Epidemiology and Public Health National University of Singapore Copy Number Variation (CNV) Copy number alteration
CNV Detection and Interpretation in Genomic Data
 CNV Detection and Interpretation in Genomic Data Benjamin W. Darbro, M.D., Ph.D. Assistant Professor of Pediatrics Director of the Shivanand R. Patil Cytogenetics and Molecular Laboratory Overview What
CNV Detection and Interpretation in Genomic Data Benjamin W. Darbro, M.D., Ph.D. Assistant Professor of Pediatrics Director of the Shivanand R. Patil Cytogenetics and Molecular Laboratory Overview What
- 1 - Reviewers' comments: Reviewer #1 (Remarks to the Author):
 Reviewers' comments: Reviewer #1 (Remarks to the Author): Summary: In this manuscript, the authors characterized cellular and transcriptional heterogeneity of breast cancer using single-cell RNA sequencing.
Reviewers' comments: Reviewer #1 (Remarks to the Author): Summary: In this manuscript, the authors characterized cellular and transcriptional heterogeneity of breast cancer using single-cell RNA sequencing.
v Feature Stamping SMS 13.0 Tutorial Prerequisites Requirements Map Module Mesh Module Scatter Module Time minutes
 v. 13.0 SMS 13.0 Tutorial Objectives Learn how to use conceptual modeling techniques to create numerical models which incorporate flow control structures into existing bathymetry. The flow control structures
v. 13.0 SMS 13.0 Tutorial Objectives Learn how to use conceptual modeling techniques to create numerical models which incorporate flow control structures into existing bathymetry. The flow control structures
underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
 Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
To open a CMA file > Download and Save file Start CMA Open file from within CMA
 Example name Effect size Analysis type Level Tamiflu Hospitalized Risk ratio Basic Basic Synopsis The US government has spent 1.4 billion dollars to stockpile Tamiflu, in anticipation of a possible flu
Example name Effect size Analysis type Level Tamiflu Hospitalized Risk ratio Basic Basic Synopsis The US government has spent 1.4 billion dollars to stockpile Tamiflu, in anticipation of a possible flu
Ch. 15 The Chromosomal Basis of Inheritance
 Ch. 15 The Chromosomal Basis of Inheritance Nov 12 12:58 PM 1 Essential Question: Are chromosomes the basis of inheritance? Nov 12 1:00 PM 2 1902 Walter S. Sutton, Theodor Boveri, et al Chromosome Theory
Ch. 15 The Chromosomal Basis of Inheritance Nov 12 12:58 PM 1 Essential Question: Are chromosomes the basis of inheritance? Nov 12 1:00 PM 2 1902 Walter S. Sutton, Theodor Boveri, et al Chromosome Theory
BIOLOGY - CLUTCH CH.15 - CHROMOSOMAL THEORY OF INHERITANCE
 !! www.clutchprep.com Chromosomal theory of inheritance: chromosomes are the carriers of genetic material. Independent Assortment alleles for different characters sort independently of each other during
!! www.clutchprep.com Chromosomal theory of inheritance: chromosomes are the carriers of genetic material. Independent Assortment alleles for different characters sort independently of each other during
Part-II: Statistical analysis of ChIP-seq data
 Part-II: Statistical analysis of ChIP-seq data Outline ChIP-seq data, features, detailed modeling aspects (today). Other ChIP-seq related problems - overview (next lecture). IDR (next lecture) Stat 877
Part-II: Statistical analysis of ChIP-seq data Outline ChIP-seq data, features, detailed modeling aspects (today). Other ChIP-seq related problems - overview (next lecture). IDR (next lecture) Stat 877
Actions in the Eye: Dynamic Gaze Datasets and Learnt Saliency Models for Visual Recognition
 Actions in the Eye: Dynamic Gaze Datasets and Learnt Saliency Models for Visual Recognition Stefan Mathe, Cristian Sminchisescu Presented by Mit Shah Motivation Current Computer Vision Annotations subjectively
Actions in the Eye: Dynamic Gaze Datasets and Learnt Saliency Models for Visual Recognition Stefan Mathe, Cristian Sminchisescu Presented by Mit Shah Motivation Current Computer Vision Annotations subjectively
A general framework for analyzing tumor subclonality using SNP array and DNA sequencing data
 Li and Li Genome Biology 2014, 15:473 METHOD A general framework for analyzing tumor subclonality using SNP array and DNA sequencing data Bo Li 1 and Jun Z Li 2 Open Access Abstract Intra-tumor heterogeneity
Li and Li Genome Biology 2014, 15:473 METHOD A general framework for analyzing tumor subclonality using SNP array and DNA sequencing data Bo Li 1 and Jun Z Li 2 Open Access Abstract Intra-tumor heterogeneity
Broad GDAC. Lung Adenocarcinoma AWG Run 2013_02_07. Dan DiCara Hailei Zhang Michael Noble
 Broad GDAC Lung Adenocarcinoma AWG Run 2013_02_07 Dan DiCara Hailei Zhang Michael Noble Copyright 2013 Broad Institute. All rights reserved. gdac@broadinstitute.org http://gdac.broadinstitute.org GDAC
Broad GDAC Lung Adenocarcinoma AWG Run 2013_02_07 Dan DiCara Hailei Zhang Michael Noble Copyright 2013 Broad Institute. All rights reserved. gdac@broadinstitute.org http://gdac.broadinstitute.org GDAC
