Exome sequencing identifies frequent mutation of ARID1A in molecular subtypes of gastric cancer
|
|
|
- Karen Suzanna West
- 5 years ago
- Views:
Transcription
1 Exome sequencing identifies frequent mutation of ARID1A in molecular subtypes of gastric cancer Kai Wang 1,7, Junsuo Kan 2,7, Siu Tsan Yuen 2, Stephanie T Shi 3, Kent Man Chu 4, Simon Law 4, Tsun Leung Chan 2, Zhengyan Kan 1, Annie S Y Chan 2, Wai Yin Tsui 2, Siu Po Lee 2, Siu Lun Ho 2, Anthony K W Chan 2, Grace H W Cheng 2, Peter C Roberts 5, Paul A Rejto 1, Neil W Gibson 1,6, David J Pocalyko 1, Mao Mao 1, Jiangchun Xu 1 & Suet Yi Leung 2 Gastric cancer is a heterogeneous disease with multiple environmental etiologies and alternative pathways of carcinogenesis 1,2. Beyond mutations in TP53, alterations in other genes or pathways account for only small subsets of the disease. We performed exome sequencing of 22 gastric cancer samples and identified previously unreported mutated genes and pathway alterations; in particular, we found genes involved in chromatin modification to be commonly mutated. A downstream validation study confirmed frequent inactivating mutations or protein deficiency of ARID1A, which encodes a member of the SWI-SNF chromatin remodeling family, in 83% of gastric cancers with microsatellite instability (MSI), 73% of those with Epstein-Barr virus (EBV) infection and 11% of those that were not infected with EBV and microsatellite stable (MSS). The mutation spectrum for ARID1A differs between molecular subtypes of gastric cancer, and mutation prevalence is negatively associated with mutations in TP53. Clinically, ARID1A alterations were associated with better prognosis in a stage-independent manner. These results reveal the genomic landscape, and highlight the importance of chromatin remodeling, in the molecular taxonomy of gastric cancer. Recent studies using next-generation sequencing (NGS) have revealed an extensive repertoire of potential cancer-driving genes in several cancer types To further explore the genetic basis of gastric cancer, we performed whole-exome capture using the Agilent SureSelect Human All Exon kit followed by NGS on the Illumina Genome Analyzer II or HiSeq 2000 platforms to identify somatic mutations in 22 matched pairs of gastric cancer and normal tissue (Supplementary Table 1), with a mean depth of 116 and 91.4% of bases covered to at least 10 (Supplementary Table 2). Somatic mutations were predicted using algorithms described in the Supplementary Note, and an extensive subset was confirmed by Sequenom MassARRAY genotyping (Supplementary Table 3), giving a positive prediction rate of 96.8% (95% confidence interval (CI), %) for somatic single-nucleotide variation (SNV) and 97.9% (95% CI, %) for somatic insertion or deletion (indel) alterations. Overall, 7,036 somatic mutations were detected in the 22 gastric cancer samples, of which 4,653 occurred in coding regions or essential splice sites (2,513 missense, 137 nonsense, 5 stop codon loss, 90 splice site, 855 indel and 1,053 synonymous mutations) (Table 1 and Supplementary Table 4). Consistent with the known consequences of mismatch repair deficiency, the gastric cancer samples with MSI had an average of somatic mutations (including both SNVs and indels) per megabase of DNA, whereas the MSS gastric cancer samples had an average of 3.29, a difference of approximately tenfold. Likewise, MSI gastric cancer samples had an average of 620 protein-altering somatic mutations, which was tenfold higher than the the number found in MSS gastric cancer samples (mean of 62). Although there was no significant difference in the nonsynonymous-to-synonymous (NS/S) ratio between the MSI and the MSS gastric cancers, we found a significantly higher NS/S ratio in EBV-infected gastric cancer samples (4.55 ± 0.97 compared to 2.76 ± 0.72 in non-ebv infected gastric cancer samples, P < 0.012) and in poorly differentiated gastric cancers (3.52 ± 0.94 compared to 2.56 ± 0.78 in well- and moderately differentiated gastric cancers, P < 0.014), suggestive of a higher positive selection pressure for driver mutations in these subgroups 18,19. Of note, the NS/S ratio for the EBV-infected gastric cancer samples (range ) is among the highest for solid organ cancers (range ) reported to date 3,8 10,12,17. C-to-T transitions were the most common mutation (51%) across all gastric cancers, with 68% involving CG dinucleotides. The MSI gastric cancers also had a distinctly high rate of T-to-C transitions (30%) (Supplementary Table 5). The observed mutation incidence for MSS gastric cancers (3.29 per megabase) is higher than our previous results from sequencing the kinome of most other cancers ( per megabase for renal, colorectal and ovarian cancers) but lower than in lung cancers (4.21 per megabase) 20. The observed mutation spectrum 1 Oncology Research Unit, Pfizer Worldwide Research and Development, La Jolla, California, USA. 2 Department of Pathology, The University of Hong Kong, Queen Mary Hospital, Pokfulam, Hong Kong. 3 External Research Solutions, Pfizer Worldwide Research and Development, La Jolla, California, USA. 4 Department of Surgery, The University of Hong Kong, Queen Mary Hospital, Pokfulam, Hong Kong. 5 Research Embedded Business Technology, Pfizer Worldwide Research and Development, La Jolla, California, USA. 6 Present address: Regulus Therapeutics, San Diego, California, USA. 7 These authors contributed equally to this work. Correspondence should be addressed to S.Y.L. (suetyi@hku.hk) or J.X. (jiangchun.xu@pfizer.com). Received 18 May; accepted 23 September; published online 30 October 2011; doi: /ng.982 Nature Genetics VOLUME 43 NUMBER 12 DECEMBER
2 Table 1 Summary of somatic mutation types and prevalence in 22 gastric cancers SNV Sample Synonymous Missense Stop gained Stop lost MSS samples Coding regions and essential splice sites is similar to our previous report for both MSS and MSI gastric cancers 20. In the current study, we have generated a list of 2,890 genes harboring protein-altering somatic mutations in gastric cancer, 59 of which were reported in the kinome study 20 and in the Catalogue of Somatic Mutations in Cancer (COSMIC) 21. Thus, 2,831 new genes have been discovered to be mutated in gastric cancer (Supplementary Table 6). Eighty-two genes were mutated in three or more samples and 447 genes in two or more samples (Supplementary Table 7). Using a driver-gene score 22,23 (described in the Supplementary Note), 20 genes were identified as candidate drivers in our gastric cancer cohort at a false discovery rate (FDR) of 0.2 (Table 2). These genes included known drivers of gastric carcinogenesis, such as TP53 (ref. 24; mutated in 36% of samples), PTEN 25 (27%) and CTNNB1 (ref. 26; 9%), as well as genes reported in COSMIC to be mutated in gastric cancer, such as TTK 27 (mutated in 18% of samples) and ACVR2A 28 (18%). In addition, we identified many genes previously known to associate with gastric cancer with slightly higher FDRs, including PIK3CA 29 (mutated in 14% of samples), APC 30 (14%) and CDH1 (refs. 31,32; 9%) (Supplementary Table 7). Our candidate drivers also included highly penetrant variations in genes not previously reported to be mutated in gastric cancer, including ARID1A (6 of 22 gastric cancer samples or 27%) and other genes with diverse cellular functions of potential importance in cancer, such as FMN2 (involved in cytoskeletal organization and cell polarity) and SEMA3E (involved in axon guidance) 33. Essential splice site Indel Total Other non-coding regions Mutation per Mb DNA NS/S SNV Indel pfg001t pfg002t pfg003t pfg005t pfg006t pfg007t pfg009t a pfg010t pfg011t pfg014t pfg015t pfg018t a pfg020t pfg021t pfg022t a pfg024t pfg025t pfg029t Total MSS , MSI samples pfg008t , pfg016t , pfg017t pfg019t Total MSI 697 1, , ,408 Overall total 1,053 2, , ,462 Mutation per Mb DNA was calculated by dividing the total number of somatic SNVs and indels overlapping with the coding regions and essential splice sites by the total number of coding bases sufficiently covered ( 3 in tumor and 10 in matched normal samples) by sequencing data. a EBV-infected gastric cancer samples. We searched for over-represented molecular pathways among genes with protein-altering somatic mutations of biological significance (Supplementary Tables 8, 9 and Supplementary Note). Chromatin modification and cell junction organization were the pathways with the most significant enrichment of mutated genes. Frequent mutations of chromatin remodeling genes, such as ARID1A, PBRM1, MEN1 and DAXX, were recently discovered in other cancers, but their involvement in gastric cancer had not been reported 4,5,7,11. We found that, in addition to ARID1A, other members of the SWI-SNF complex (ARID1B, PBRM1 and SMARCC1), ISWI complex (SMARCA1) and NuRD complex (CHD3, CHD4 and MBD2), as well as other genes encoding histone-modifying proteins (SIRT1 and SETD2), were also mutated, in total affecting 59% of gastric cancers. Moreover, a single gastric cancer sample could have mutations in multiple chromatin remodeling genes involving different complexes (Supplementary Table 10). Overall, 59% of gastric cancer samples had mutations in at least one gene affecting cell junction organization, including CDH1 and other cadherin family members, consistent with the infiltrative nature of gastric cancer and its great tendency toward the loss of cellto-cell adhesion. Genes involved in cell cycle regulation were mutated in 77% of gastric cancers, including TP53, PTEN and TTK. Other core signaling pathways frequently mutated in gastric cancer included the Wnt-BMP-TGFβ, axon guidance, MAPK, DNA replication, focal adhesion, ERBB, ATR-BRCA and Rb pathways, many of which may have therapeutic implications VOLUME 43 NUMBER 12 DECEMBER 2011 Nature Genetics
3 Table 2 Candidate driver genes in gastric cancer predicted from exome sequencing of 22 gastric cancers Gene symbol Name Size N S N T N SNV MSS N SNV MSI N indel MSS N indel MSI P Q value Q score TP53 Cellular tumor antigen p53 1, PTEN PIP 3 phosphatase and dual-specificity protein phosphatase (PTEN) The frequent mutation of genes encoding chromatin remodelers in gastric cancer and, specifically, the discovery of ARID1A as one of the top candidate driver genes in gastric cancer prompted us to study ARID1A in greater detail. ARID1A was recently discovered to be a driver for ovarian clear cell carcinoma 7,11. Because of the incomplete coverage of the ARID1A gene in exome sequencing, we resequenced all coding exons of the discovery cohort as well as an additional 87 gastric cancer samples by Sanger sequencing. The two cohorts together encompass 109 gastric cancers, including 23 MSI 34,35, 15 MSS, EBV-infected 36,37 and 71 MSS, non-ebv infected samples (Supplementary Tables 1 and 11). We found a striking difference in the somatic mutation rate of ARID1A across the different molecular subtypes of gastric cancer, with very high incidence in the MSI types (78%, 18/23, P < compared to MSS, non-ebv infected) and the MSS, EBV-infected gastric cancers (47%, 7/15, P = compared to MSS, non-ebv infected) and lower incidence in MSS, non-ebv infected gastric cancers (10%, 7/71) (Fig. 1 and Supplementary Table 12). Among the 32 gastric cancer samples containing ARID1A mutations, 14 samples had two mutations and 18 had a single mutation, giving a total of 46 ARID1A mutations, out of which 39 (85%) were truncating mutations (Fig. 2, Supplementary Table 13 and Supplementary Fig. 1). The majority (75%, 24/32) of gastric cancers with ARID1A mutations had either a loss of or substantially lower protein expression compared to gastric cancers without ARID1A mutation (P < 0.001), as determined by immunostaining (Fig. 1, Supplementary Tables 12, 13 and Supplementary Fig. 2). Of note, six gastric cancer samples showed absent or weak protein expression despite the lack of detectable ARID1A mutations, suggesting that other mechanisms may contribute to ARID1A inactivation. Combining mutation and protein expression data, ARID1A alterations (either through mutation or reduced protein levels) were detected in 73% (11/15) of EBV-infected gastric cancer samples, at levels comparable to the alteration rate in MSI gastric cancers (83%, 19/23) and substantially higher than in the MSS, non-ebv infected subtype (11%, 8/71) (Fig. 1 and Supplementary Table 12). Gastric cancers with ARID1A alterations 1, ARID1A AT-rich interactive domain containing protein 1A 6, RPL22 60S ribosomal protein L TTK Dual specificity protein kinase TTK 2, FMN2 Formin-2 5, SPRR2B Small proline-rich protein 2B PTN Pleiotrophin precursor ACVR2A Activin receptor type-2a precursor 1, PMS2L3 Postmeiotic segregation increased 2-like protein DNAH7 Dynein heavy chain 7, axonemal 12, TTN Titin isoform novex-3 82, FSCB Fibrous sheath CABYR-binding protein 2, CTNNB1 Catenin beta-1 2, SEMA3E Semaphorin-3E precursor 2, MCHR1 Melanin-concentrating hormone receptor 1 1, SPANXN2 Sperm protein associated with the nucleus on the X chromosome N METTL3 N6-adenosine-methyltransferase 70 kda subunit 1, EIF3A Eukaryotic translation initiation factor 3 subunit A 4, EPB41L3 Band 4.1-like protein 3 3, Numbers of somatic mutations shown include only protein-altering mutations. Size, interrogated size of a gene, obtained by counting the coding bases targeted by the SureSelect capture baits; N S, number of cancer samples in which a gene is mutated; N T, total number of mutations; N SNV MSS, number of SNV mutations in MSS cancers; N SNV MSI, number of SNV mutations in MSI cancers; N indel MSS, number of indel mutations in MSS cancers; N indel MSI, number of indel mutations in MSI cancers. The derivation of driver gene statistics (P values, Q values and Q scores) is described in the Supplementary Note. were less likely to harbor mutations in the TP53 gene. TP53 mutations were observed in 21% (8/38) of gastric cancer samples with ARID1A alterations but in 52% (37/71) of samples without alterations (P = 0.002; Fig. 1 and Supplementary Table 12). Of note, TP53 mutation was uncommon in both MSI (17.4%, 4/23) and EBV-infected gastric cancers (6.7%, 1/15). Gastric cancer samples with ARID1A alterations showed a trend of prolonged, recurrence-free survival (P = 0.058, log-rank test) (Supplementary Fig. 3). More importantly, in multivariate analysis incorporating tumor stage, Lauren s tumor type, MSI status and ARID1A alteration status, only advanced tumor stage a Number of samples % mutated 83% altered *** 47% mutated 73% altered ** 10% mutated 11% altered MSI MSS EBV MSS non-ebv ARID1A wild type ARID1A protein deficient ARID1A mutated b Number of samples ** 52% mutated ARID1A no alteration 21% mutated ARID1A altered TP53 wild type TP53 mutated Figure 1 Relationship of ARID1A alterations (mutation or protein deficiency) with molecular subtypes of gastric cancer. (a) Incidence of ARID1A mutation and protein deficiency in different molecular subtypes of gastric cancer. Blue color indicates samples with ARID1A mutation, and shaded area indicates samples with protein deficiency, as determined by immunohistochemistry. **P < 0.01, ***P < for both ARID1A mutation and alteration (see Supplementary Table 12 for detail). (b) TP53 mutation rate in gastric cancer samples with or without ARID1A alterations. Nature Genetics VOLUME 43 NUMBER 12 DECEMBER
4 Figure 2 Difference in the mutation spectrum of ARID1A between molecular subtypes of gastric cancer. Individual exons of the ARID1A gene are represented as numbered gray boxes. Mutations detected by sequencing the coding region of ARID1A in 23 MSI and 86 MSS gastric cancers are shown. cdna and peptide positions are based on the ENST transcript. Colors of text correspond to functional impact prediction by SIFT, as indicated at the bottom. (adjusted hazard ratio stage IV compared to stage I, 13.8; stage III compared to stage I, 9.25; P = 0.001) and absence of ARID1A alterations (adjusted hazard ratio 3.09, P = 0.029) were independent variables that could predict early recurrence (Supplementary Table 14). The mutation spectrum of ARID1A was strikingly different between the MSI and MSS subtypes (Fig. 2). In the MSI gastric cancer samples, 97% (28/29) of the mutations were indels, mostly involving short mononucleotide repeats of C or G (89%, 25/28). Specifically, one single G7 tract located in exon 20 was mutated in 26% of MSI gastric cancers. For the MSS gastric cancer samples (both EBV infected and non EBV infected), 59% (10/17) of the mutations were SNVs with 6 nonsense and 4 missense mutations. Only 7 mutations were indels, with 1 involving a mononucleotide repeat sequence. ARID1A contains many short repeats of 4 7 mononucleotides in its coding region. The high rate of somatic indels at these repeats in MSI gastric cancer samples is not likely to be caused simply by a high background mutation rate. Compared with the global background mutation rate of somatic indels at mononucleotide tracts of similar length in MSI gastric cancers, the mutation rate in ARID1A is 12- to 61-fold higher (P < ; Supplementary Fig. 4 and Supplementary Table 15), consistent with a driver gene being targeted by the MSI mechanism 38. The overall mutation rate of ARID1A in MSI gastric cancer (78%) is comparable to that of well-established and functionally validated driver genes inactivated by MSI, such as TGFBR2 (ref. 39). This study revealed for the first time the diverse gene and pathway alterations in gastric cancer and the importance of chromatin remodeling genes, with frequent mutations in ARID1A and other associated factors affected 59% of gastric cancer samples in total. Consistent with our findings, loss of protein expression of ARID1A has been noted in 11 14% of gastric cancer samples by immunostaining 40,41, and homozygous deletion of ARID1A has been detected in a gastric cancer cell line 4. Interestingly, we noted an inverse relationship between the two most frequently mutated gastric cancer genes ARID1A and TP53, suggesting that they may drive alternative subsets of gastric cancer. Mutation of TP53 is rare in other cancers occurring in the ovary 7,11, endometrium 41, kidney 4 or endocrine pancreas 5, in which mutation of chromatin-modifying genes is common. Likewise, TP53 mutation is uncommon in both MSI and EBV-infected gastric cancers, two subtypes with frequent ARID1A mutation. Thus, mutation of genes encoding chromatin-remodeling enzymes may constitute an alternative pathway of carcinogenesis independent of TP53 that drives cancer development through epigenetic modification. Comparison of the ARID1A mutation spectrum between gastric and ovarian cancers 7 revealed ten overlapping mutations, all of which are indels involving G7 or C6 repeats. These mutations are potentially caused by MSI, as a meta-analysis showed that 11% of clear cell ovarian cancers are mismatch repair deficient 42 ; thus, mutation of ARID1A may be prevalent in MSI tumors arising in other organ sites. The high c.16_17insc 5 UTR 373 nt 5 UTR 373 nt c.275_291delgcggg CCCGGCGCGGAG c.490_497delgccgcggc c.666delc c.822delg c.822delg c.833g>c G278A c.885_888delcaac c.1029_1043deiagc TGCGGCGGCGGC c.1060c>t Q354* c.2268delc c.2268delc c.2290_2291insc c.2253delt c.2802_2803insaata ARID c.3148g>a D1050N c.3156t>a Y1052* ARID c.3972delc c.3978_3980delgca MSI gastric cancers c.3598c>t Q1200* c.3773a>g D1258G c.3826c>t R1276* c.3971_3972insc c.4336c>t R1446* c.4336c>t R1446* c.4350delc c.4550delc c.4656_4657insc c.4683_4684insc c.4852delc c.5542delg c.5542delg c.5542delg c.5689delc c.6303delc c.6415delc c.6415delc Nonsense mutation or frame shift c.2772delg c.5643_5652deltgtg ACAACA MSS gastric cancers Damaging (low confidence) rate of ARID1A alteration (73%) in gastric cancers with EBV infection is intriguing. It would be interesting to examine other EBV-associated cancer types to see if the association with ARID1A mutation is gastric specific. As in ovarian clear cell carcinoma 7, we noted frequent twohit inactivation of ARID1A, supporting its role as a tumor suppressor gene. For gastric cancer samples with a single mutation, protein loss was commonly observed, indicating that other mechanisms may contribute to inactivation. Moreover, haploinsufficiency of ARID1A may already contribute to carcinogenesis, as mice heterozygous for ARID1A mutation are embryonic lethal 43. ARID1A functions as a component of the SWI-SNF complex and can repress gene expression, including that of MYC and of genes regulated by the E2F transcription factors More interestingly, a previous shrna screen revealed that knockdown of ARID1A in Jurkat cells led to inhibition of Fas-mediated apoptosis 47. One histological feature unique to MSI 35,48 and EBV-infected gastric cancers 2,36,37 is abundant tumor-infiltrating lymphocytes. Thus, it is tempting to speculate that selection for mutated ARID1A in these gastric cancer subtypes may be due to the ability of this mutation to confer resistance to Fas-mediated apoptosis and thereby promote immune evasion. Our preliminary analysis shows that the observed better prognosis in individuals with gastric cancer with ARID1A alterations may not be attributable solely to a link between ARID1A alteration and MSI status. ARID1A alteration is predictive of disease-free survival even after controlling for other clinicopathological variables, including tumor stage and MSI status, suggesting that ARID1A alterations may define a molecular subgroup of gastric cancer with a unique mechanism of carcinogenesis leading to distinct clinical behavior. Finally, our discovery of frequent ARID1A alterations in two specific molecular subtypes of gastric cancer underscores the importance of a comprehensive understanding of pathwayspecific molecular changes to support the development of targeted molecular therapy and raises the possibility that specific epigenetic therapy targeting alterations in ARID1A or other chromatin-modifying enzymes may be useful in treating gastric cancer. URLs. European Genome-Phenome Archive, COSMIC, Methods Methods and any associated references are available in the online version of the paper at c.6260g>a G2087E Tolerated c.6641g>c S2214T c.6709_6710insc 3 UTR 1,354 nt 3 UTR 1,354 nt 1222 VOLUME 43 NUMBER 12 DECEMBER 2011 Nature Genetics
5 Accession numbers. Exome sequencing data has been deposited in the European Genome-Phenome Archive with the study ID EGAS Note: Supplementary information is available on the Nature Genetics website. Acknowledgments We thank L. Ng and M.K. Ng for technical assistance and clinicians in the Hong Kong Hospital Authority for clinical care. We thank Illumina for performing whole-exome sequencing. AUTHOR CONTRIBUTIONS D.J.P., P.A.R., N.W.G., M.M., J.X., S.T.Y. and S.Y.L. conceived of the study. S.T.Y., S.Y.L., J.X. and M.M. directed the study. K.W., S.T.S., P.A.R., J.X., M.M., S.T.Y. and S.Y.L. contributed to the project design. K.W., Z.K. and G.H.W.C. performed the bioinformatics data analysis. J.K., T.L.C., A.S.Y.C., W.Y.T., S.P.L., S.L.H. and A.K.W.C. performed experiments on ARID1A mutation and other molecular analysis. K.M.C. and S.L. contributed samples, data and comments on the manuscript. P.C.R. contributed to data management. K.W., J.K., S.T.Y., M.M., J.X. and S.Y.L. analyzed and interpreted data and wrote the manuscript with the assistance and final approval from all authors. COMPETING FINANCIAL INTERESTS The authors declare competing financial interests: details accompany the full-text HTML version of the paper at Published online at Reprints and permissions information is available online at reprints/index.html. 1. Lauwers, G.Y. et al. Gastric carcinoma. in WHO Classification of Tumours of the Digestive System (eds. Bosman, F.T., Carneiro, F., Hruban, R.H. & Theise, N.D.) (IARC, Lyon, 2010). 2. Osato, T. & Imai, S. Epstein-Barr virus and gastric carcinoma. Semin. Cancer Biol. 7, (1996). 3. Wei, X. et al. Exome sequencing identifies GRIN2A as frequently mutated in melanoma. Nat. Genet. 43, (2011). 4. Varela, I. et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469, (2011). 5. Jiao, Y. et al. DAXX/ATRX, MEN1, and mtor pathway genes are frequently altered in pancreatic neuroendocrine tumors. Science 331, (2011). 6. Chapman, M.A. et al. Initial genome sequencing and analysis of multiple myeloma. Nature 471, (2011). 7. Wiegand, K.C. et al. ARID1A mutations in endometriosis-associated ovarian carcinomas. N. Engl. J. Med. 363, (2010). 8. Pleasance, E.D. et al. A small-cell lung cancer genome with complex signatures of tobacco exposure. Nature 463, (2010). 9. Pleasance, E.D. et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 463, (2010). 10. Lee, W. et al. The mutation spectrum revealed by paired genome sequences from a lung cancer patient. Nature 465, (2010). 11. Jones, S. et al. Frequent mutations of chromatin remodeling gene ARID1A in ovarian clear cell carcinoma. Science 330, (2010). 12. Ding, L. et al. Genome remodelling in a basal-like breast cancer metastasis and xenograft. Nature 464, (2010). 13. Shah, S.P. et al. Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature 461, (2009). 14. Mardis, E.R. et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N. Engl. J. Med. 361, (2009). 15. Ley, T.J. et al. DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature 456, (2008). 16. Campbell, P.J. et al. Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nat. Genet. 40, (2008). 17. Totoki, Y. et al. High-resolution characterization of a hepatocellular carcinoma genome. Nat. Genet. 43, (2011). 18. Hughes, A.L. & Nei, M. Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 335, (1988). 19. Li, W.H. & Gojobori, T. Rapid evolution of goat and sheep globin genes following gene duplication. Mol. Biol. Evol. 1, (1983). 20. Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, (2007). 21. Forbes, S.A. et al. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 39, D945 D950 (2011). 22. Kan, Z. et al. Diverse somatic mutation patterns and pathway alterations in human cancers. Nature 466, (2010). 23. Sjoblom, T. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, (2006). 24. Uchino, S. et al. p53 mutation in gastric cancer: a genetic model for carcinogenesis is common to gastric and colorectal cancer. Int. J. Cancer 54, (1993). 25. Wang, J.Y. et al. Mutation analysis of the putative tumor suppressor gene PTEN/MMAC1 in advanced gastric carcinomas. Virchows Arch. 442, (2003). 26. Park, W.S. et al. Frequent somatic mutations of the beta-catenin gene in intestinaltype gastric cancer. Cancer Res. 59, (1999). 27. Ahn, C.H., Kim, Y.R., Kim, S.S., Yoo, N.J. & Lee, S.H. Mutational analysis of TTK gene in gastric and colorectal cancers with microsatellite instability. Cancer Res. Treat. 41, (2009). 28. Hempen, P.M. et al. Evidence of selection for clones having genetic inactivation of the activin A type II receptor (ACVR2) gene in gastrointestinal cancers. Cancer Res. 63, (2003). 29. Li, V.S. et al. Mutations of PIK3CA in gastric adenocarcinoma. BMC Cancer 5, 29 (2005). 30. Nakatsuru, S. et al. Somatic mutation of the APC gene in gastric cancer: frequent mutations in very well differentiated adenocarcinoma and signet-ring cell carcinoma. Hum. Mol. Genet. 1, (1992). 31. Becker, K.F. et al. E-cadherin gene mutations provide clues to diffuse type gastric carcinomas. Cancer Res. 54, (1994). 32. Guilford, P. et al. E-cadherin germline mutations in familial gastric cancer. Nature 392, (1998). 33. Chedotal, A., Kerjan, G. & Moreau-Fauvarque, C. The brain within the tumor: new roles for axon guidance molecules in cancers. Cell Death Differ. 12, (2005). 34. Leung, S.Y. et al. hmlh1 promoter methylation and lack of hmlh1 expression in sporadic gastric carcinomas with high-frequency microsatellite instability. Cancer Res. 59, (1999). 35. Leung, S.Y. et al. Microsatellite instability, Epstein-Barr virus, mutation of type II transforming growth factor beta receptor and BAX in gastric carcinomas in Hong Kong Chinese. Br. J. Cancer 79, (1999). 36. Yuen, S.T. et al. In situ detection of Epstein-Barr virus in gastric and colorectal adenocarcinomas. Am. J. Surg. Pathol. 18, (1994). 37. Chen, X. et al. Variation in gene expression patterns in human gastric cancers. Mol. Biol. Cell 14, (2003). 38. Woerner, S.M., Kloor, M., von Knebel Doeberitz, M. & Gebert, J.F. Microsatellite instability in the development of DNA mismatch repair deficient tumors. Cancer Biomark. 2, (2006). 39. Markowitz, S. et al. Inactivation of the type II TGF-beta receptor in colon cancer cells with microsatellite instability. Science 268, (1995). 40. Wiegand, K.C. et al. Loss of BAF250a (ARID1A) is frequent in high-grade endometrial carcinomas. J. Pathol. 224, (2011). 41. Guan, B. et al. Mutation and Loss of Expression of ARID1A in Uterine Low-grade Endometrioid Carcinoma. Am. J. Surg. Pathol. 35, (2011). 42. Murphy, M.A. & Wentzensen, N. Frequency of mismatch repair deficiency in ovarian cancer: a systematic review. Int. J. Cancer 129, (2011). 43. Gao, X. et al. ES cell pluripotency and germ-layer formation require the SWI/SNF chromatin remodeling component BAF250a. Proc. Natl. Acad. Sci. USA 105, (2008). 44. Nagl, N.G. Jr., Zweitzig, D.R., Thimmapaya, B., Beck, G.R. Jr. & Moran, E. The c-myc gene is a direct target of mammalian SWI/SNF-related complexes during differentiation-associated cell cycle arrest. Cancer Res. 66, (2006). 45. Nagl, N.G. Jr., Wang, X., Patsialou, A., Van Scoy, M. & Moran, E. Distinct mammalian SWI/SNF chromatin remodeling complexes with opposing roles in cell-cycle control. EMBO J. 26, (2007). 46. Inoue, H. et al. Target genes of the largest human SWI/SNF complex subunit control cell growth. Biochem. J. 434, (2011). 47. Luo, B. et al. Highly parallel identification of essential genes in cancer cells. Proc. Natl. Acad. Sci. USA 105, (2008). 48. dos Santos, N.R., Seruca, R., Constancia, M., Seixas, M. & Sobrinho-Simoes, M. Microsatellite instability at multiple loci in gastric carcinoma: clinicopathologic implications and prognosis. Gastroenterology 110, (1996). Nature Genetics VOLUME 43 NUMBER 12 DECEMBER
6 ONLINE METHODS Sample preparation, MSI and EBV testing. The use of archival frozen human gastric samples for this study was approved by the Institutional Review Board of the University of Hong Kong and the Hospital Authority Hong Kong West Cluster. Because samples were archival in nature, the IRBs waived the need for informed consent, in compliance with international guidelines. Frozen tumor and non-neoplastic gastric tissues were collected from gastrectomy specimens from Queen Mary Hospital, The University of Hong Kong. DNA was extracted from frozen tumor tissue after macrodissection, and the tumor content was confirmed to be above 70% by cryostat sectioning for samples submitted for exome sequencing and above 50% for the validation cohort. DNA was also extracted from paired unaffected gastric tissue distant from the tumor site that was confirmed to be tumor free by prior histological examination of cryostat sections. Microsatellite instability analysis was performed using the standard method as previously described 35,49, with the addition of several loci, including SLC7A8, TMPPB5 and ZNF2. At least five loci were analyzed in each case, including both dinucleotide and mononucleotide repeats. Those with high-level microsatellite instability (at least 40% of markers positive) were considered MSI. Those with low-level or no microsatellite instability (<40% of markers positive) were considered MSS. The presence of EBV in cancer cells was detected by in situ hybridization for EBV-encoded RNA (EBER) as described previously 36,37 or using the Ventana INFORM EBER Probe (Ventana Medical Systems) according to the manufacturer s protocol. The RNA Positive Control Probe (Ventana Medical Systems) was used to ensure RNA integrity. Helicobacter pylori infection status was defined by its presence in the gastrectomy specimen or endoscopic gastric biopsy before surgery. Immunohistochemistry for ARID1A. Immunostaining was performed using a rabbit polyclonal antibody recognizing human ARID1A (Sigma-Aldrich HPA005456, dilution 1 in 200). After heat-mediated antigen retrieval and primary antibody incubation, signal was detected using the LabVision UltraVision kit (Thermo Scientific) and developed using diaminobenzidine counterstained with Mayer s hematoxylin. Staining was performed on a tissue microarray containing the gastric cancer tissues, and histological images were obtained with the ScanScope CS digital scanning system (Aperio Technologies). Nuclear staining of cancer cells was graded in comparison to normal cells (including infiltrating lymphocytes, fibroblasts and normal gastric glandular cells). ARID1A is strongly and uniformly expressed in normal cells, including gastric epithelial cells, lymphocytes and fibroblasts. ARID1A expression in gastric cancer cells was graded as normal if staining intensity was similar to that in normal cells, weak if staining intensity was substantially weaker than in normal cells or loss if nuclear staining was absent, in which case the normal cells served as an internal positive control. Grading was performed independently by two pathologists (S.T.Y. and S.L.H.) who were blinded to the mutation results. Concordance rates between the two pathologists were high, and the few discrepancies were reviewed and consensus grading was assigned. Exome capture, library construction and sequencing. Exome capture was performed by Illumina using Agilent SureSelect in-solution target enrichment technology (Agilent Technologies). Libraries were constructed following the Illumina Paired-End Sequencing Library Preparation Protocol version from the SureSelect Human All Exon kit, with an added gel purification step for insert size selection. The kit contains a pool of RNA-based 120-mer capture oligomers (or baits) targeting 37,640,396 bases of 165,637 consensus coding sequence exons and their flanking regions. Sequencing was performed in two phases. In the first phase, libraries were constructed for ten pairs of tumor and matched normal samples with an insert size of ~300 bp and sequenced using two lanes on an Illumina Genome Analyzer II sequencer. In the second phase, 12 additional pairs of samples were sequenced on one lane of the newer HiSeq 2000 sequencer and with a targeted insert size of ~180 bp. All sequencing was run with paired-end 75-bp reads and was performed according to Illumina s standard protocol. Library construction and sequencing of DNA from subject pfg011 were done in duplicate to assess the reproducibility of the technology and to serve as controls for the somatic mutation calling algorithm (see Supplementary Note). Reads from both replicates were combined in the final analysis. On average, ~158.5 million purity-filtered reads were generated for each sample. The mean percentage of duplicate reads due to PCR and optical artifacts was 6.3% in our data set. After removing these duplicate reads, ~133.5 million uniquely mapped reads were obtained for each sample. On average, 59.3% of reads in each sample had at least 50% overlap with any targeted region ± 100 bp in the SureSelect whole-exome bait library. The targeted regions in each sample were sequenced to an average depth of 115.8, with ~97.8% of the targeted regions covered 1, ~91.4% 10, ~77.6% 30 and ~65.7% 50. We have also genotyped ~1.1 million markers using Illumina HumanOmni1-Quad BeadChips on the same set of samples. The average concordance between the array-based and sequencing-based genotyping calls was 99.5%. A detailed summary of sequencing statistics for all samples can be found in Supplementary Table 2. Mutation confirmation using the Sequenom MassARRAY system. Sequenom MassARRAY assays were performed by Sequenom. Assays were designed by using ProxSNP and PreXTEND analysis and validated by finding successful PCR and primer extension in Coriell DNA. Sample analysis was carried out using Sequenom iplex PRO chemistry. All Typer calls were manually confirmed by examining the spectra for each assay and sample. Peaks for two alleles were checked against the background of each well, and a mutation call was confirmed if the peak was unique to that allele. Mutation analysis for ARID1A and TP53 in gastric cancers and matched normal tissues by Sanger sequencing. The coding exons of ARID1A were assessed by Sanger sequencing using primers similar to those previously published 11 with minor modifications (Supplementary Table 16). The discovery cohort of 22 gastric cancer samples was also resequenced because of incomplete coverage in some regions of ARID1A in exome sequencing. An additional 87 gastric cancers were also sequenced. All mutations identified in tumors were confirmed by independent PCR and Sanger sequencing in the specific tumors and their paired normal tissue to determine their somatic nature. Sequencing was performed using the ABI BigDye Terminator v3.1 Cycle Sequencing kit (Applied Biosystems). The sequence chromatograms were visually inspected with assistance from Mutation Surveyor. For analysis of the TP53 gene, exons 5 to 9 that include the mutation hotspots were sequenced using the primers listed in Supplementary Table 16. All mutations identified in tumors were confirmed by independent PCR and Sanger sequencing in the specific tumors and their paired normal tissue. Statistical analysis of clinico-pathological correlations. Comparison of the NS/S ratio with clinico-pathological parameters was performed using the Mann-Whitney U test. Analysis of ARID1A mutation or alteration frequency with clinico-pathological parameters was performed using either the chisquared test or Fisher s exact test, when appropriate. Sixty-two individuals with gastric cancer who had undergone curative resection were analyzed for factors related to disease-free survival. Sex, age, tumor stage, site and differentiation, Lauren s tumor type, H. pylori infection, MSI status, EBV status, TP53 mutation and ARID1A alterations were analyzed for their prognostic value using Kaplan-Meier survival analysis with log-ranked tests as well as Univariate Cox regression analysis. Significant factors in Univariate analysis (tumor stage, Lauren s tumor type, MSI status and ARID1A alterations) were further subjected to a multivariate Cox regression analysis in a forward stepwise manner. Mononucleotide repeat mutation frequency analysis in MSI gastric cancers. The MSI cancers tend to have frequent random mutations involving mononucleotide repeats, and the mutation frequency increases with the length of the mononucleotide repeat. True driver genes targeted by the MSI mechanism generally show a substantially elevated mutation rate in their coding repeats compared to the background mutation rate for the same repeat length 50. Thus, to address whether the observed high mutation rate of ARID1A in MSI gastric cancers is simply due to a high background rate of mutations in these tumors, we first estimated the background mutation rate for coding mononucleotide repeats in relation to repeat length across the entire exome for the four MSI gastric cancers in the discovery cohort (for which whole-exome sequencing data were available). Specifically, for each mononucleotide repeat length we Nature Genetics doi: /ng.982
7 calculated the total number of coding repeats mutated in the four MSI gastric cancers and determined the total number of coding repeats that were adequately covered among all targeted regions by SureSelect (including 16,568 genes). Mutation frequency for each mononucleotide repeat in ARID1A in all 23 MSI gastric cancers was then compared to the global background mutation rate using the chi-squared test (Supplementary Table 15). Driver gene analysis. Please refer to Supplementary Note and Supplementary Table Yuen, S.T. et al. Similarity of the phenotypic patterns associated with BRAF and KRAS mutations in colorectal neoplasia. Cancer Res. 62, (2002). 50. Woerner, S.M. et al. Pathogenesis of DNA repair-deficient cancers: a statistical metaanalysis of putative Real Common Target genes. Oncogene 22, (2003). doi: /ng.982 Nature Genetics
Case presentation 04/13/2017. Genomic/morphological classification of endometrial carcinoma
 Genomic/morphological classification of endometrial carcinoma Robert A. Soslow, MD soslowr@mskcc.org architecture.about.com Case presentation 49 year old woman with vaginal bleeding Underwent endometrial
Genomic/morphological classification of endometrial carcinoma Robert A. Soslow, MD soslowr@mskcc.org architecture.about.com Case presentation 49 year old woman with vaginal bleeding Underwent endometrial
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Characterisation of structural variation in breast. cancer genomes using paired-end sequencing on. the Illumina Genome Analyser
 Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
IPATIMUP Translational Research Unit Matching IPATIMUP s Expertise with Company s Research/Clinical Strategy
 IPATIMUP Translational Research Unit Matching IPATIMUP s Expertise with Company s Research/Clinical Strategy Innovative Projects Questions & Needs André Albergaria INDUSTRY Pharma R&D Clinical Research
IPATIMUP Translational Research Unit Matching IPATIMUP s Expertise with Company s Research/Clinical Strategy Innovative Projects Questions & Needs André Albergaria INDUSTRY Pharma R&D Clinical Research
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Development of Carcinoma Pathways
 The Construction of Genetic Pathway to Colorectal Cancer Moriah Wright, MD Clinical Fellow in Colorectal Surgery Creighton University School of Medicine Management of Colon and Diseases February 23, 2019
The Construction of Genetic Pathway to Colorectal Cancer Moriah Wright, MD Clinical Fellow in Colorectal Surgery Creighton University School of Medicine Management of Colon and Diseases February 23, 2019
NGS in tissue and liquid biopsy
 NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
MSI positive MSI negative
 Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
BWA alignment to reference transcriptome and genome. Convert transcriptome mappings back to genome space
 Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Computer Science, Biology, and Biomedical Informatics (CoSBBI) Outline. Molecular Biology of Cancer AND. Goals/Expectations. David Boone 7/1/2015
 Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
AD (Leave blank) TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients
 AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium
 Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium Ulrike Peters Fred Hutchinson Cancer Research Center University of Washington U01-CA137088-05, PI: Peters
Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium Ulrike Peters Fred Hutchinson Cancer Research Center University of Washington U01-CA137088-05, PI: Peters
DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK
 CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
Transform genomic data into real-life results
 CLINICAL SUMMARY Transform genomic data into real-life results Biomarker testing and targeted therapies can drive improved outcomes in clinical practice New FDA-Approved Broad Companion Diagnostic for
CLINICAL SUMMARY Transform genomic data into real-life results Biomarker testing and targeted therapies can drive improved outcomes in clinical practice New FDA-Approved Broad Companion Diagnostic for
The mutations that drive cancer. Paul Edwards. Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge
 The mutations that drive cancer Paul Edwards Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge Previously on Cancer... hereditary predisposition Normal Cell Slightly
The mutations that drive cancer Paul Edwards Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge Previously on Cancer... hereditary predisposition Normal Cell Slightly
Next generation histopathological diagnosis for precision medicine in solid cancers
 Next generation histopathological diagnosis for precision medicine in solid cancers from genomics to clinical application Aldo Scarpa ARC-NET Applied Research on Cancer Department of Pathology and Diagnostics
Next generation histopathological diagnosis for precision medicine in solid cancers from genomics to clinical application Aldo Scarpa ARC-NET Applied Research on Cancer Department of Pathology and Diagnostics
Clonal evolution of human cancers
 Clonal evolution of human cancers -Pathology-based microdissection and genetic analysis precisely demonstrates molecular evolution of neoplastic clones- Hiroaki Fujii, MD Ageo Medical Laboratories, Yashio
Clonal evolution of human cancers -Pathology-based microdissection and genetic analysis precisely demonstrates molecular evolution of neoplastic clones- Hiroaki Fujii, MD Ageo Medical Laboratories, Yashio
Molecular Subtyping of Endometrial Cancer: A ProMisE ing Change
 Molecular Subtyping of Endometrial Cancer: A ProMisE ing Change Charles Matthew Quick, M.D. Associate Professor of Pathology Director of Gynecologic Pathology University of Arkansas for Medical Sciences
Molecular Subtyping of Endometrial Cancer: A ProMisE ing Change Charles Matthew Quick, M.D. Associate Professor of Pathology Director of Gynecologic Pathology University of Arkansas for Medical Sciences
Biology of cancer development in the GI tract
 1 Genesis and progression of GI cancer a genetic disease Colorectal cancer Fearon and Vogelstein proposed a genetic model to explain the stepwise formation of colorectal cancer (CRC) from normal colonic
1 Genesis and progression of GI cancer a genetic disease Colorectal cancer Fearon and Vogelstein proposed a genetic model to explain the stepwise formation of colorectal cancer (CRC) from normal colonic
Frequency(%) KRAS G12 KRAS G13 KRAS A146 KRAS Q61 KRAS K117N PIK3CA H1047 PIK3CA E545 PIK3CA E542K PIK3CA Q546. EGFR exon19 NFS-indel EGFR L858R
 Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Refining Prognosis of Early Stage Lung Cancer by Molecular Features (Part 2): Early Steps in Molecularly Defined Prognosis
 5/17/13 Refining Prognosis of Early Stage Lung Cancer by Molecular Features (Part 2): Early Steps in Molecularly Defined Prognosis Johannes Kratz, MD Post-doctoral Fellow, Thoracic Oncology Laboratory
5/17/13 Refining Prognosis of Early Stage Lung Cancer by Molecular Features (Part 2): Early Steps in Molecularly Defined Prognosis Johannes Kratz, MD Post-doctoral Fellow, Thoracic Oncology Laboratory
Molecular Pathology of Ovarian Carcinoma with Morphological Correlation
 Molecular athology of Ovarian Carcinoma with Morphological Correlation Kathleen R. Cho, M.D. Comprehensive Cancer Center and Departments of athology and Internal Medicine University of Michigan Medical
Molecular athology of Ovarian Carcinoma with Morphological Correlation Kathleen R. Cho, M.D. Comprehensive Cancer Center and Departments of athology and Internal Medicine University of Michigan Medical
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
The Role of Next Generation Sequencing in Solid Tumor Mutation Testing
 The Role of Next Generation Sequencing in Solid Tumor Mutation Testing Allie H. Grossmann MD PhD Department of Pathology, University of Utah Division of Anatomic Pathology & Oncology, ARUP Laboratories
The Role of Next Generation Sequencing in Solid Tumor Mutation Testing Allie H. Grossmann MD PhD Department of Pathology, University of Utah Division of Anatomic Pathology & Oncology, ARUP Laboratories
Tumor mutational burden and its transition towards the clinic
 Tumor mutational burden and its transition towards the clinic G C C A T C A C Wolfram Jochum Institute of Pathology Kantonsspital St.Gallen CH-9007 St.Gallen wolfram.jochum@kssg.ch 30th European Congress
Tumor mutational burden and its transition towards the clinic G C C A T C A C Wolfram Jochum Institute of Pathology Kantonsspital St.Gallen CH-9007 St.Gallen wolfram.jochum@kssg.ch 30th European Congress
UNIVERSITY OF TORINO DEPARTMENT OF ONCOLOGY. Giorgio V. Scagliotti University of Torino Dipartment of Oncology
 Giorgio V. Scagliotti University of Torino Dipartment of Oncology giorgio.scagliotti@unito.it Molecular landscape of MM not fully characterized to allow personalized treatment Recurrent genetic alterations
Giorgio V. Scagliotti University of Torino Dipartment of Oncology giorgio.scagliotti@unito.it Molecular landscape of MM not fully characterized to allow personalized treatment Recurrent genetic alterations
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies
 Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep
 Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
Molecular Testing Updates. Karen Rasmussen, PhD, FACMG Clinical Molecular Genetics Spectrum Medical Group, Pathology Division Portland, Maine
 Molecular Testing Updates Karen Rasmussen, PhD, FACMG Clinical Molecular Genetics Spectrum Medical Group, Pathology Division Portland, Maine Keeping Up with Predictive Molecular Testing in Oncology: Technical
Molecular Testing Updates Karen Rasmussen, PhD, FACMG Clinical Molecular Genetics Spectrum Medical Group, Pathology Division Portland, Maine Keeping Up with Predictive Molecular Testing in Oncology: Technical
iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers).
 Supplementary Materials Supplementary Methods iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers). IDH1 R132H and R132L Forward:
Supplementary Materials Supplementary Methods iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers). IDH1 R132H and R132L Forward:
MEDICAL GENOMICS LABORATORY. Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG)
 Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
The molecular genetics of endometrial cancer
 The molecular genetics of endometrial cancer Lora Hedrick Ellenson, M.D. Department of Pathology and Laboratory Medicine Weill Medical College of Cornell University Introduction Classification of endometrial
The molecular genetics of endometrial cancer Lora Hedrick Ellenson, M.D. Department of Pathology and Laboratory Medicine Weill Medical College of Cornell University Introduction Classification of endometrial
Multistep nature of cancer development. Cancer genes
 Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Genomic Medicine: What every pathologist needs to know
 Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Supplementary Table 1. PIK3CA mutation in colorectal cancer
 Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Identification of Potential Therapeutic Targets by Molecular and Genomic Profiling of 628 Cases of Uterine Serous Carcinoma
 Identification of Potential Therapeutic Targets by Molecular and Genomic Profiling of 628 Cases of Uterine Serous Carcinoma Nathaniel L Jones 1, Joanne Xiu 2, Sandeep K. Reddy 2, Ana I. Tergas 1, William
Identification of Potential Therapeutic Targets by Molecular and Genomic Profiling of 628 Cases of Uterine Serous Carcinoma Nathaniel L Jones 1, Joanne Xiu 2, Sandeep K. Reddy 2, Ana I. Tergas 1, William
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/22278 holds various files of this Leiden University dissertation. Author: Cunha Carvalho de Miranda, Noel Filipe da Title: Mismatch repair and MUTYH deficient
Cover Page The handle http://hdl.handle.net/1887/22278 holds various files of this Leiden University dissertation. Author: Cunha Carvalho de Miranda, Noel Filipe da Title: Mismatch repair and MUTYH deficient
Case Presentation Diana Lim, MBBS, FRCPA, FRCPath Senior Consultant Department of Pathology, National University Health System, Singapore Assistant Pr
 Case Presentation Diana Lim, MBBS, FRCPA, FRCPath Senior Consultant Department of Pathology, National University Health System, Singapore Assistant Professor Yong Loo Lin School of Medicine, National University
Case Presentation Diana Lim, MBBS, FRCPA, FRCPath Senior Consultant Department of Pathology, National University Health System, Singapore Assistant Professor Yong Loo Lin School of Medicine, National University
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Accel-Amplicon Panels
 Accel-Amplicon Panels Amplicon sequencing has emerged as a reliable, cost-effective method for ultra-deep targeted sequencing. This highly adaptable approach is especially applicable for in-depth interrogation
Accel-Amplicon Panels Amplicon sequencing has emerged as a reliable, cost-effective method for ultra-deep targeted sequencing. This highly adaptable approach is especially applicable for in-depth interrogation
Lynch Syndrome Screening for Endometrial Cancer: Basic Concepts 1/16/2017
 1 Hi, my name is Sarah Kerr. I m a pathologist at Mayo Clinic, where I participate in our high volume Lynch syndrome tumor testing practice. Today I hope to cover some of the basics needed to understand
1 Hi, my name is Sarah Kerr. I m a pathologist at Mayo Clinic, where I participate in our high volume Lynch syndrome tumor testing practice. Today I hope to cover some of the basics needed to understand
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies
 Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer
 RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer J Yu *, KF To, QY Liang K e y M e s s a g e s 1. Somatostatin receptor 1
RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer J Yu *, KF To, QY Liang K e y M e s s a g e s 1. Somatostatin receptor 1
Germline Testing for Hereditary Cancer with Multigene Panel
 Germline Testing for Hereditary Cancer with Multigene Panel Po-Han Lin, MD Department of Medical Genetics National Taiwan University Hospital 2017-04-20 Disclosure No relevant financial relationships with
Germline Testing for Hereditary Cancer with Multigene Panel Po-Han Lin, MD Department of Medical Genetics National Taiwan University Hospital 2017-04-20 Disclosure No relevant financial relationships with
Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder Labs August 30, 2012
 Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Cellecta Overview. Started Operations in 2007 Headquarters: Mountain View, CA
 Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Evolution of Pathology
 1 Traditional pathology Molecular pathology 2 Evolution of Pathology Gross Pathology Cellular Pathology Morphologic Pathology Molecular/Predictive Pathology Antonio Benivieni (1443-1502): First autopsy
1 Traditional pathology Molecular pathology 2 Evolution of Pathology Gross Pathology Cellular Pathology Morphologic Pathology Molecular/Predictive Pathology Antonio Benivieni (1443-1502): First autopsy
Insights from Sequencing the Melanoma Exome
 Insights from Sequencing the Melanoma Exome Michael Krauthammer, MD PhD, December 2 2015 Yale University School Yof Medicine 1 2012 Exome Screens and Results Exome Sequencing of 108 sun-exposed melanomas
Insights from Sequencing the Melanoma Exome Michael Krauthammer, MD PhD, December 2 2015 Yale University School Yof Medicine 1 2012 Exome Screens and Results Exome Sequencing of 108 sun-exposed melanomas
Enterprise Interest Thermo Fisher Scientific / Employee
 Enterprise Interest Thermo Fisher Scientific / Employee A next-generation sequencing assay to estimate tumor mutation load from FFPE research samples Fiona Hyland. Director of R&D, Bioinformatics Clinical
Enterprise Interest Thermo Fisher Scientific / Employee A next-generation sequencing assay to estimate tumor mutation load from FFPE research samples Fiona Hyland. Director of R&D, Bioinformatics Clinical
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Cancer genetics
 Cancer genetics General information about tumorogenesis. Cancer induced by viruses. The role of somatic mutations in cancer production. Oncogenes and Tumor Suppressor Genes (TSG). Hereditary cancer. 1
Cancer genetics General information about tumorogenesis. Cancer induced by viruses. The role of somatic mutations in cancer production. Oncogenes and Tumor Suppressor Genes (TSG). Hereditary cancer. 1
Identifying Mutations Responsible for Rare Disorders Using New Technologies
 Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery
 CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery PacBio CoLab Session October 20, 2017 For Research Use Only. Not for use in diagnostics procedures. Copyright 2017 by Pacific Biosciences
CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery PacBio CoLab Session October 20, 2017 For Research Use Only. Not for use in diagnostics procedures. Copyright 2017 by Pacific Biosciences
(a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable
 Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Familial and Hereditary Colon Cancer
 Familial and Hereditary Colon Cancer Aasma Shaukat, MD, MPH, FACG, FASGE, FACP GI Section Chief, Minneapolis VAMC Associate Professor, Division of Gastroenterology, Department of Medicine, University of
Familial and Hereditary Colon Cancer Aasma Shaukat, MD, MPH, FACG, FASGE, FACP GI Section Chief, Minneapolis VAMC Associate Professor, Division of Gastroenterology, Department of Medicine, University of
Serrated Polyps and a Classification of Colorectal Cancer
 Serrated Polyps and a Classification of Colorectal Cancer Ian Chandler June 2011 Structure Serrated polyps and cancer Molecular biology The Jass classification The familiar but oversimplified Vogelsteingram
Serrated Polyps and a Classification of Colorectal Cancer Ian Chandler June 2011 Structure Serrated polyps and cancer Molecular biology The Jass classification The familiar but oversimplified Vogelsteingram
Androgen Receptor Expression in Renal Cell Carcinoma: A New Actionable Target?
 Androgen Receptor Expression in Renal Cell Carcinoma: A New Actionable Target? New Frontiers in Urologic Oncology Juan Chipollini, MD Clinical Fellow Department of Genitourinary Oncology Moffitt Cancer
Androgen Receptor Expression in Renal Cell Carcinoma: A New Actionable Target? New Frontiers in Urologic Oncology Juan Chipollini, MD Clinical Fellow Department of Genitourinary Oncology Moffitt Cancer
Supplementary Online Content
 Supplementary Online Content Fumagalli D, Venet D, Ignatiadis M, et al. RNA Sequencing to predict response to neoadjuvant anti-her2 therapy: a secondary analysis of the NeoALTTO randomized clinical trial.
Supplementary Online Content Fumagalli D, Venet D, Ignatiadis M, et al. RNA Sequencing to predict response to neoadjuvant anti-her2 therapy: a secondary analysis of the NeoALTTO randomized clinical trial.
Fluxion Biosciences and Swift Biosciences Somatic variant detection from liquid biopsy samples using targeted NGS
 APPLICATION NOTE Fluxion Biosciences and Swift Biosciences OVERVIEW This application note describes a robust method for detecting somatic mutations from liquid biopsy samples by combining circulating tumor
APPLICATION NOTE Fluxion Biosciences and Swift Biosciences OVERVIEW This application note describes a robust method for detecting somatic mutations from liquid biopsy samples by combining circulating tumor
Anatomic Molecular Pathology: An Emerging Field
 Anatomic Molecular Pathology: An Emerging Field Antonia R. Sepulveda M.D., Ph.D. University of Pennsylvania asepu@mail.med.upenn.edu 2008 ASIP Annual Meeting Anatomic pathology (U.S.) is a medical specialty
Anatomic Molecular Pathology: An Emerging Field Antonia R. Sepulveda M.D., Ph.D. University of Pennsylvania asepu@mail.med.upenn.edu 2008 ASIP Annual Meeting Anatomic pathology (U.S.) is a medical specialty
The use of diagnostic FFPE material in cancer epidemiology research
 The use of diagnostic FFPE material in cancer epidemiology research Neil O Callaghan Genetic Epidemiology Laboratory Department of Pathology The University of Melbourne www.pedigree.org.au Overview Who
The use of diagnostic FFPE material in cancer epidemiology research Neil O Callaghan Genetic Epidemiology Laboratory Department of Pathology The University of Melbourne www.pedigree.org.au Overview Who
LESSON 3.2 WORKBOOK. How do normal cells become cancer cells? Workbook Lesson 3.2
 For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
WHAT SHOULD WE DO WITH TUMOUR BUDDING IN EARLY COLORECTAL CANCER?
 CANCER STAGING TNM and prognosis in CRC WHAT SHOULD WE DO WITH TUMOUR BUDDING IN EARLY COLORECTAL CANCER? Alessandro Lugli, MD Institute of Pathology University of Bern Switzerland Maastricht, June 19
CANCER STAGING TNM and prognosis in CRC WHAT SHOULD WE DO WITH TUMOUR BUDDING IN EARLY COLORECTAL CANCER? Alessandro Lugli, MD Institute of Pathology University of Bern Switzerland Maastricht, June 19
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Clinical Grade Genomic Profiling: The Time Has Come
 Clinical Grade Genomic Profiling: The Time Has Come Gary Palmer, MD, JD, MBA, MPH Senior Vice President, Medical Affairs Foundation Medicine, Inc. Oct. 22, 2013 1 Why We Are Here A Shared Vision At Foundation
Clinical Grade Genomic Profiling: The Time Has Come Gary Palmer, MD, JD, MBA, MPH Senior Vice President, Medical Affairs Foundation Medicine, Inc. Oct. 22, 2013 1 Why We Are Here A Shared Vision At Foundation
IntelliGENSM. Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community.
 IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
Tumor suppressor genes D R. S H O S S E I N I - A S L
 Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing.
 Supplementary Figure 1 CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing. Whole-exome sequencing of six plasmacytoid-variant bladder
Supplementary Figure 1 CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing. Whole-exome sequencing of six plasmacytoid-variant bladder
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
The diagnostic and prognostic value of genetic aberrations in resectable distal bile duct cancer Rijken, A.M.
 UvA-DARE (Digital Academic Repository) The diagnostic and prognostic value of genetic aberrations in resectable distal bile duct cancer Rijken, A.M. Link to publication Citation for published version (APA):
UvA-DARE (Digital Academic Repository) The diagnostic and prognostic value of genetic aberrations in resectable distal bile duct cancer Rijken, A.M. Link to publication Citation for published version (APA):
September 20, Submitted electronically to: Cc: To Whom It May Concern:
 History Study (NOT-HL-12-147), p. 1 September 20, 2012 Re: Request for Information (RFI): Building a National Resource to Study Myelodysplastic Syndromes (MDS) The MDS Cohort Natural History Study (NOT-HL-12-147).
History Study (NOT-HL-12-147), p. 1 September 20, 2012 Re: Request for Information (RFI): Building a National Resource to Study Myelodysplastic Syndromes (MDS) The MDS Cohort Natural History Study (NOT-HL-12-147).
Hepatitis B virus X gene in the development of hepatocellular carcinoma. Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p.
 Title Hepatitis B virus X gene in the development of hepatocellular carcinoma Author(s) Ma, NF; Lau, SH; Hu, L; Dong, SS; Guan, XY Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p. 44-47
Title Hepatitis B virus X gene in the development of hepatocellular carcinoma Author(s) Ma, NF; Lau, SH; Hu, L; Dong, SS; Guan, XY Citation Hong Kong Medical Journal, 2011, v. 17 n. 6, suppl. 6, p. 44-47
Familial and Hereditary Colon Cancer
 Familial and Hereditary Colon Cancer Aasma Shaukat, MD, MPH, FACG, FASGE, FACP GI Section Chief, Minneapolis VAMC Associate Professor, Division of Gastroenterology, Department of Medicine, University of
Familial and Hereditary Colon Cancer Aasma Shaukat, MD, MPH, FACG, FASGE, FACP GI Section Chief, Minneapolis VAMC Associate Professor, Division of Gastroenterology, Department of Medicine, University of
CANCER. Inherited Cancer Syndromes. Affects 25% of US population. Kills 19% of US population (2nd largest killer after heart disease)
 CANCER Affects 25% of US population Kills 19% of US population (2nd largest killer after heart disease) NOT one disease but 200-300 different defects Etiologic Factors In Cancer: Relative contributions
CANCER Affects 25% of US population Kills 19% of US population (2nd largest killer after heart disease) NOT one disease but 200-300 different defects Etiologic Factors In Cancer: Relative contributions
Current Status of Biomarkers (including DNA Tumor Markers and Immunohistochemistry in the Laboratory Diagnosis of Tumors)
 Current Status of Biomarkers (including DNA Tumor Markers and Immunohistochemistry in the Laboratory Diagnosis of Tumors) Kael Mikesell, DO McKay-Dee Hospital May 14, 2015 Outline Update to DNA Testing
Current Status of Biomarkers (including DNA Tumor Markers and Immunohistochemistry in the Laboratory Diagnosis of Tumors) Kael Mikesell, DO McKay-Dee Hospital May 14, 2015 Outline Update to DNA Testing
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Supplemental Data: Detailed Characteristics of Patients with MKRN3. Patient 1 was born after an uneventful pregnancy. She presented in our
 1 2 Supplemental Data: Detailed Characteristics of Patients with MKRN3 Mutations 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Patient 1 was born after an uneventful pregnancy. She presented
1 2 Supplemental Data: Detailed Characteristics of Patients with MKRN3 Mutations 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Patient 1 was born after an uneventful pregnancy. She presented
oncogenes-and- tumour-suppressor-genes)
 Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Biomarker development in the era of precision medicine. Bei Li, Interdisciplinary Technical Journal Club
 Biomarker development in the era of precision medicine Bei Li, 23.08.2016 Interdisciplinary Technical Journal Club The top ten highest-grossing drugs in the United States help between 1 in 25 and 1 in
Biomarker development in the era of precision medicine Bei Li, 23.08.2016 Interdisciplinary Technical Journal Club The top ten highest-grossing drugs in the United States help between 1 in 25 and 1 in
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Accepted Manuscript. Pancreatic Cancer Subtypes: Beyond Lumping and Splitting. Andrew J. Aguirre
 Accepted Manuscript Pancreatic Cancer Subtypes: Beyond Lumping and Splitting Andrew J. Aguirre PII: S0016-5085(18)35213-2 DOI: https://doi.org/10.1053/j.gastro.2018.11.004 Reference: YGAST 62235 To appear
Accepted Manuscript Pancreatic Cancer Subtypes: Beyond Lumping and Splitting Andrew J. Aguirre PII: S0016-5085(18)35213-2 DOI: https://doi.org/10.1053/j.gastro.2018.11.004 Reference: YGAST 62235 To appear
Biochemistry of Carcinogenesis. Lecture # 35 Alexander N. Koval
 Biochemistry of Carcinogenesis Lecture # 35 Alexander N. Koval What is Cancer? The term "cancer" refers to a group of diseases in which cells grow and spread unrestrained throughout the body. It is difficult
Biochemistry of Carcinogenesis Lecture # 35 Alexander N. Koval What is Cancer? The term "cancer" refers to a group of diseases in which cells grow and spread unrestrained throughout the body. It is difficult
Case Study. Overview. Deleterious MLH1 mutation detected on sequencing 10/16/2014
 The Role of Next Generation Sequencing for Hereditary Cancer Syndromes: A Focus on Endometrial Cancer Laura J. Tafe, MD Assistant Professor of Pathology Assistant Director, Molecular Pathology Dartmouth-Hitchcock
The Role of Next Generation Sequencing for Hereditary Cancer Syndromes: A Focus on Endometrial Cancer Laura J. Tafe, MD Assistant Professor of Pathology Assistant Director, Molecular Pathology Dartmouth-Hitchcock
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Colorectal cancer Chapelle, J Clin Oncol, 2010
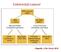 Colorectal cancer Chapelle, J Clin Oncol, 2010 Early-Stage Colorectal cancer: Microsatellite instability, multigene assay & emerging molecular strategy Asit Paul, MD, PhD 11/24/15 Mr. X: A 50 yo asymptomatic
Colorectal cancer Chapelle, J Clin Oncol, 2010 Early-Stage Colorectal cancer: Microsatellite instability, multigene assay & emerging molecular strategy Asit Paul, MD, PhD 11/24/15 Mr. X: A 50 yo asymptomatic
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
MEDICAL GENOMICS LABORATORY. Peripheral Nerve Sheath Tumor Panel by Next-Gen Sequencing (PNT-NG)
 Peripheral Nerve Sheath Tumor Panel by Next-Gen Sequencing (PNT-NG) Ordering Information Acceptable specimen types: Blood (3-6ml EDTA; no time limitations associated with receipt) Saliva (OGR-575 DNA Genotek;
Peripheral Nerve Sheath Tumor Panel by Next-Gen Sequencing (PNT-NG) Ordering Information Acceptable specimen types: Blood (3-6ml EDTA; no time limitations associated with receipt) Saliva (OGR-575 DNA Genotek;
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
The Biology and Genetics of Cells and Organisms The Biology of Cancer
 The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
Molecular Testing in Lung Cancer
 Molecular Testing in Lung Cancer Pimpin Incharoen, M.D. Assistant Professor, Thoracic Pathology Department of Pathology, Ramathibodi Hospital Genetic alterations in lung cancer Source: Khono et al, Trans
Molecular Testing in Lung Cancer Pimpin Incharoen, M.D. Assistant Professor, Thoracic Pathology Department of Pathology, Ramathibodi Hospital Genetic alterations in lung cancer Source: Khono et al, Trans
Gene expression profiling predicts clinical outcome of prostate cancer. Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M.
 SUPPLEMENTARY DATA Gene expression profiling predicts clinical outcome of prostate cancer Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M. Hoffman, William L. Gerald Table of Contents
SUPPLEMENTARY DATA Gene expression profiling predicts clinical outcome of prostate cancer Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M. Hoffman, William L. Gerald Table of Contents
GENETICS OF COLORECTAL CANCER: HEREDITARY ASPECTS By. Magnitude of the Problem. Magnitude of the Problem. Cardinal Features of Lynch Syndrome
 GENETICS OF COLORECTAL CANCER: HEREDITARY ASPECTS By HENRY T. LYNCH, M.D. 1 Could this be hereditary Colon Cancer 4 Creighton University School of Medicine Omaha, Nebraska Magnitude of the Problem Annual
GENETICS OF COLORECTAL CANCER: HEREDITARY ASPECTS By HENRY T. LYNCH, M.D. 1 Could this be hereditary Colon Cancer 4 Creighton University School of Medicine Omaha, Nebraska Magnitude of the Problem Annual
Lynch Syndrome. Angie Strang, PGY2
 Lynch Syndrome Angie Strang, PGY2 Background Previously hereditary nonpolyposis colorectal cancer Autosomal dominant inherited cancer susceptibility syndrome Caused by defects in the mismatch repair system
Lynch Syndrome Angie Strang, PGY2 Background Previously hereditary nonpolyposis colorectal cancer Autosomal dominant inherited cancer susceptibility syndrome Caused by defects in the mismatch repair system
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits
 AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) cancer DNA
 www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
Neoplasia 2018 lecture 11. Dr H Awad FRCPath
 Neoplasia 2018 lecture 11 Dr H Awad FRCPath Clinical aspects of neoplasia Tumors affect patients by: 1. their location 2. hormonal secretions 3. paraneoplastic syndromes 4. cachexia Tumor location Even
Neoplasia 2018 lecture 11 Dr H Awad FRCPath Clinical aspects of neoplasia Tumors affect patients by: 1. their location 2. hormonal secretions 3. paraneoplastic syndromes 4. cachexia Tumor location Even
