Supplementary Tables 1
|
|
|
- Godfrey Cook
- 6 years ago
- Views:
Transcription
1 Supplementary Tables 1
2 Supplementary Table 1: The 11 genes in the geneset for Case Study 1 At1g62570 At1g09350 At1g60470 At2g47180 At4g17090 At5g20830 At2g16890 At3g51240 At3g55120 At4g27560 At5g08640 flavin-containing monooxygenase family protein / FMO family protein galactinol synthase, putative galactinol synthase, putative galactinol synthase, putative beta-amylase (CT-BMY) / 1,4-alpha-D-glucan maltohydrolase sucrose synthase / sucrose-udp glucosyltransferase (SUS1) UDP-glucuronosyl/UDP-glucosyl transferase family protein naringenin 3-dioxygenase / flavanone 3-hydroxylase (F3H) chalcone-flavanone isomerase / chalcone isomerase (CHI) glycosyltransferase family protein flavonol synthase 1 (FLS1) 2
3 Supplementary Table 2: The 14 genes in the geneset for Case Study 2 At5g18170 At1g28670 At2g35690 At1g55920 At4g22690 At1g10360 At1g16410 At2g22330 At2g23600 At2g29340 At4g15550 At5g24160 At5g14740 At3g01500 glutamate dehydrogenase 1 (GDH1) lipase acyl-coa oxidase, putative serine O-acetyltransferase, putative cytochrome P450 family protein glutathione S-transferase, putative cytochrome P450, putative cytochrome P450, putative hydrolase, alpha/beta fold family protein short-chain dehydrogenase/reductase (SDR) family protein UDP-glucose:indole-3-acetate beta-d-glucosyltransferase (IAGLU) squalene monooxygenase 1,2 / squalene epoxidase 1,2 (SQP1,2) carbonic anhydrase 2 / carbonate dehydratase 2 (CA2) (CA18) carbonic anhydrase 1, chloroplast / carbonate dehydratase 1 (CA1) 3
4 Supplementary Table 3: The 113 senescent-responsive genes in the geneset for Case Study 3 At1g07600 At1g09500 At1g13990 At1g14400 At1g18210 At1g19570 At1g20620 At1g21670 At1g30420 At1g30460 At1g35160 At1g47128 At1g51200 At1g52040 At1g52880 At1g53580 At1g53750 At1g58360 At1g59870 At1g63010 At1g67840 At1g68820 At1g70900 At1g71950 At1g73260 At1g73325 At1g73750 metallothionein-like protein 1A (MT-1A) (MT-Q) (MT-2) cinnamyl-alcohol dehydrogenase family / CAD family ubiquitin-conjugating enzyme 1 (UBC1) calcium-binding protein, putative dehydroascorbate reductase, putative catalase 3 (SEN2) ATP-binding cassette transport protein, putative zinc finger (CCCH-type) family protein / YT521-B-like family protein protein GF14 phi (GRF4) cysteine proteinase (RD21A) / thiol protease zinc finger (AN1-like) family protein jacalin lectin family protein no apical meristem (NAM) family protein hydroxyacylglutathione hydrolase, putative / glyoxalase II, putative 26S proteasome AAA-ATPase subunit (RPT1a) amino acid permease I (AAP1) ABC transporter family protein SPX (SYG1/Pho81/XPR1) domain-containing protein ATP-binding region, ATPase-like domain-containing protein membrane protein, putative trypsin and protease inhibitor family protein / Kunitz family protein trypsin and protease inhibitor family protein / Kunitz family protein Continued on next page 4
5 Supplementary Table 3: Continued At1g75280 At1g75380 At1g78080 At1g78380 At2g05540 At2g20860 At2g21660 At2g21950 At2g22240 At2g23980 At2g25450 At2g26560 At2g39900 At2g42690 At2g43290 At2g43770 At2g45210 At3g02040 At3g03720 At3g07590 At3g09390 isoflavone reductase, putative wound-responsive protein-related AP2 domain-containing transcription factor RAP2.4 glutathione S-transferase, putative glycine-rich protein lipoic acid synthase (LIP1) glycine-rich RNA-binding protein (GRP7) SKP1 interacting partner 6 (SKIP6) inositol-3-phosphate synthase isozyme 2 / myo-inositol-1-phosphate synthase 2 / MI-1-P synthase 2 / IPS 2 cyclic nucleotide-regulated ion channel / cyclic nucleotide-gated channel (CNGC6) 2-oxoglutarate-dependent dioxygenase, putative patatin, putative LIM domain-containing protein lipase, putative calmodulin-like protein (MSS3) transducin family protein / WD-40 repeat family protein auxin-responsive protein-related glycerophosphoryl diester phosphodiesterase family protein amino acid permease family protein small nuclear ribonucleoprotein D1, putative / snrnp core protein D1, putative / Sm protein D1, putative metallothionein protein, putative (MT2A) At3g12120 omega-6 fatty acid desaturase, endoplasmic reticulum (FAD2) / delta-12 desaturase At3g15580 autophagy 8i (APG8i) At3g15730 phospholipase D alpha 1 / PLD alpha 1 (PLDALPHA1) (PLD1) / choline phosphatase 1 At3g16640 translationally controlled tumor family protein Continued on next page 5
6 Supplementary Table 3: Continued At3g17790 At3g22600 At3g23920 At3g25480 At3g25760 At3g26100 At3g44100 At3g44300 At3g44720 At3g46000 At3g48000 At3g51130 At3g51730 At3g52800 At3g55430 At3g62550 At4g01610 At4g03280 At4g11360 At4g11600 At4g11910 At4g12290 At4g13250 At4g16190 At4g16740 At4g18280 acid phosphatase type 5 (ACP5) protease inhibitor/seed storage/lipid transfer protein (LTP) family protein beta-amylase, putative / 1,4-alpha-D-glucan maltohydrolase, putative rhodanese-like domain-containing protein early-responsive to dehydration stress protein (ERD12) regulator of chromosome condensation (RCC1) family protein MD-2-related lipid recognition domain-containing protein / ML domain-containing protein nitrilase 2 (NIT2) prephenate dehydratase family protein actin-depolymerizing factor, putative (ADF2) aldehyde dehydrogenase (ALDH2) saposin B domain-containing protein zinc finger (AN1-like) family protein glycosyl hydrolase family 17 protein / beta-1,3-glucanase, putative universal stress protein (USP) family protein cathepsin B-like cysteine protease, putative cytochrome B6-F complex iron-sulfur subunit, chloroplast / Rieske iron-sulfur protein / plastoquinol-plastocyanin reductase (petc) zinc finger (C3HC4-type RING finger) family protein (RHA1b) glutathione peroxidase, putative copper amine oxidase, putative short-chain dehydrogenase/reductase (SDR) family protein cysteine proteinase, putative terpene synthase/cyclase family protein glycine-rich cell wall protein-related Continued on next page 6
7 Supplementary Table 3: Continued At4g24220 At4g27020 At4g27830 glycosyl hydrolase family 1 protein At4g30210 NADPH-cytochrome p450 reductase, putative / NADPH-ferrihemoprotein reductase, putative At4g30270 At4g30440 At4g32940 At4g35750 At4g37390 At4g37990 At4g39090 At4g39670 At5g01600 At5g02380 At5g08410 At5g10650 At5g10860 At5g11520 At5g13170 At5g17220 At5g18100 At5g19540 At5g24160 At5g24770 At5g24780 At5g34850 MERI-5 protein (MERI-5) (MERI5B) / endo-xyloglucan transferase / xyloglucan endo-1,4-beta-d-glucanase (SEN4) NAD-dependent epimerase/dehydratase family protein vacuolar processing enzyme gamma / gamma-vpe Rho-GTPase-activating protein-related auxin-responsive GH3 family protein mannitol dehydrogenase, putative (ELI3-2) cysteine proteinase RD19a (RD19A) / thiol protease ferritin 1 (FER1) metallothionein protein 2B (MT-2B) ferredoxin-thioredoxin reductase, putative zinc finger (C3HC4-type RING finger) family protein CBS domain-containing protein aspartate aminotransferase, chloroplast / transaminase A (ASP3) (YLS4) nodulin MtN3 family protein glutathione S-transferase, putative superoxide dismutase [Cu-Zn] / copper/zinc superoxide dismutase (CSD3) squalene monooxygenase 1,2 / squalene epoxidase 1,2 (SQP1,2) vegetative storage protein 2 (VSP2) vegetative storage protein 1 (VSP1) calcineurin-like phosphoesterase family protein Continued on next page 7
8 At5g43850 At5g48380 At5g53350 Supplementary Table 3: Continued acireductone dioxygenase (ARD/ARD ) family protein leucine-rich repeat family protein / protein kinase family protein ATP-dependent Clp protease ATP-binding subunit ClpX1 (CLPX) At5g53730 harpin-induced family protein / HIN1 family protein / harpin-responsive family protein At5g54250 At5g57655 At5g59310 At5g60360 At1g05340 cyclic nucleotide-regulated ion channel / cyclic nucleotide-gated channel (CNGC4) xylose isomerase family protein lipid transfer protein 4 (LTP4) cysteine proteinase, putative / AALP protein (AALP) Concluded 8
WT siz1-2 siz1-3. WT siz1-2 siz1-3. -Pi, 0.05 M IAA. -Pi, 2.5 M NPA
 A WT siz1-2 siz1-3 B WT siz1-2 siz1-3 -Pi C +Pi WT siz1-2 siz1-3 D WT siz1-2 siz1-3 E +Pi, 0.05 M IAA -Pi, 0.05 M IAA WT siz1-2 siz1-3 WT siz1-2 siz1-3 F +Pi, 2.5 M NPA -Pi, 2.5 M NPA Supplemental Figure
A WT siz1-2 siz1-3 B WT siz1-2 siz1-3 -Pi C +Pi WT siz1-2 siz1-3 D WT siz1-2 siz1-3 E +Pi, 0.05 M IAA -Pi, 0.05 M IAA WT siz1-2 siz1-3 WT siz1-2 siz1-3 F +Pi, 2.5 M NPA -Pi, 2.5 M NPA Supplemental Figure
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function
 Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
Supplemental Table S1 Up-regulated genes in irjazh plants compared to WT plants by microarray.
 Supplemental Table S1 Up-regulated genes in irjazh plants compared to WT plants by microarray. Classification Fold Change Bincode (TAIR) Annotation Probe number ACC number Cell wall. Degradation 3.29 10.6.2
Supplemental Table S1 Up-regulated genes in irjazh plants compared to WT plants by microarray. Classification Fold Change Bincode (TAIR) Annotation Probe number ACC number Cell wall. Degradation 3.29 10.6.2
BIOL 158: BIOLOGICAL CHEMISTRY II
 BIOL 158: BIOLOGICAL CHEMISTRY II Lecture 5: Vitamins and Coenzymes Lecturer: Christopher Larbie, PhD Introduction Cofactors bind to the active site and assist in the reaction mechanism Apoenzyme is an
BIOL 158: BIOLOGICAL CHEMISTRY II Lecture 5: Vitamins and Coenzymes Lecturer: Christopher Larbie, PhD Introduction Cofactors bind to the active site and assist in the reaction mechanism Apoenzyme is an
SUPPLEMENTAL TABLE I. Identified Proteins in Bovine Testicular Hyaluronidase Type I-S via LC-MS/MS
 SUPPLEMENTAL TABLE I. Identified Proteins in Bovine Testicular Hyaluronidase Type I-S via LC-MS/MS No. Protein 1 serum albumin precursor gi 30794280 2 annexin A2 gi 27807289 3 Phosphatidylethanolamine-binding
SUPPLEMENTAL TABLE I. Identified Proteins in Bovine Testicular Hyaluronidase Type I-S via LC-MS/MS No. Protein 1 serum albumin precursor gi 30794280 2 annexin A2 gi 27807289 3 Phosphatidylethanolamine-binding
0.5. Normalized 95% gray value interval h
 Normalized 95% gray value interval.5.4.3.2.1 h Supplemental Figure 1: Symptom score of root samples used in the proteomics study. For each time point, the normalized 95% gray value interval is an averaged
Normalized 95% gray value interval.5.4.3.2.1 h Supplemental Figure 1: Symptom score of root samples used in the proteomics study. For each time point, the normalized 95% gray value interval is an averaged
Biological processes. Mitochondrion Metabolic process Catalytic activity Oxidoreductase
 Full name Glyceraldehyde3 phosphate dehydrogenase Succinatesemialdehyde Glutamate dehydrogenase 1, Alcohol dehydrogenase [NADP+] 2',3'cyclicnucleotide 3' phosphodiesterase Dihydropyrimidinaserelated 2
Full name Glyceraldehyde3 phosphate dehydrogenase Succinatesemialdehyde Glutamate dehydrogenase 1, Alcohol dehydrogenase [NADP+] 2',3'cyclicnucleotide 3' phosphodiesterase Dihydropyrimidinaserelated 2
Figure S1: Abundance of Fe related proteins in oceanic Synechococcus sp. strain WH8102 in. Ferredoxin. Ferredoxin. Fe ABC transporter
 0.004 SW nm 0.04 0.03 nm nm 0.16 0.1 nm nm 0.40 0.3 nm nm 0.80 0.5 nm 1.6 1 nm 16 10 nm 160 100 nm 0 nm 0.04 nm 0.16 nm 0.40 nm 0.80 nm 1.6 nm 16 nm 160 nm Spectral counts 0 nm 0.04 nm 0.16 nm 0.40 nm
0.004 SW nm 0.04 0.03 nm nm 0.16 0.1 nm nm 0.40 0.3 nm nm 0.80 0.5 nm 1.6 1 nm 16 10 nm 160 100 nm 0 nm 0.04 nm 0.16 nm 0.40 nm 0.80 nm 1.6 nm 16 nm 160 nm Spectral counts 0 nm 0.04 nm 0.16 nm 0.40 nm
Supplementary Data AtMYB12
 Supplementary Data AtMYB12 expression in tomato leads to large scale differential modulation in transcriptome and flavonoid content in leaf and fruit tissues Ashutosh Pandey 1,$,#, Prashant Misra 1,$,#,
Supplementary Data AtMYB12 expression in tomato leads to large scale differential modulation in transcriptome and flavonoid content in leaf and fruit tissues Ashutosh Pandey 1,$,#, Prashant Misra 1,$,#,
Bacterial cellulose synthesis mechanism of facultative
 1 2 3 4 Bacterial cellulose synthesis mechanism of facultative anaerobe Kaihua Ji 1 +, Wei Wang 2, 3 +, Bing Zeng 1, Sibin Chen 1, Qianqian Zhao 4, Yueqing Chen 1, Guoqiang Li 1* and Ting Ma 1* 5 6 7 Figure
1 2 3 4 Bacterial cellulose synthesis mechanism of facultative anaerobe Kaihua Ji 1 +, Wei Wang 2, 3 +, Bing Zeng 1, Sibin Chen 1, Qianqian Zhao 4, Yueqing Chen 1, Guoqiang Li 1* and Ting Ma 1* 5 6 7 Figure
Figure S3 Differentially expressed fungal and plant genes organised by catalytic activity ontology Organisation of E. festucae (A) and L.
 sak WT Number of branches 3 WT sak Figure S1 Vasculature of plants infected with the E. festucae sak mutant. Light micrographs of perennial ryegrass blade tissue showing branching between the vasculature
sak WT Number of branches 3 WT sak Figure S1 Vasculature of plants infected with the E. festucae sak mutant. Light micrographs of perennial ryegrass blade tissue showing branching between the vasculature
Coenzymes, vitamins and trace elements 209. Petr Tůma Eva Samcová
 Coenzymes, vitamins and trace elements 209 Petr Tůma Eva Samcová History and nomenclature of enzymes 1810, Gay-Lussac made an experiment with yeats alter saccharide to ethanol and CO 2 Fermentation From
Coenzymes, vitamins and trace elements 209 Petr Tůma Eva Samcová History and nomenclature of enzymes 1810, Gay-Lussac made an experiment with yeats alter saccharide to ethanol and CO 2 Fermentation From
Mineral Nutrition in Plants. Plant nutrition: essentiality, mechanism of absorption, role in plant metabolism.
 Mineral Nutrition in Plants Plant nutrition: essentiality, mechanism of absorption, role in plant metabolism. Mineral : An inorganic element Nutrient : A substance needed to survive or necessary for the
Mineral Nutrition in Plants Plant nutrition: essentiality, mechanism of absorption, role in plant metabolism. Mineral : An inorganic element Nutrient : A substance needed to survive or necessary for the
Glycogen Metabolism Dr. Mohammad Saadeh
 Glycogen Metabolism Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry II Philadelphia University Faculty of pharmacy I. overview Glucose is energy source for Brain.
Glycogen Metabolism Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry II Philadelphia University Faculty of pharmacy I. overview Glucose is energy source for Brain.
Lecture 16. Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III
 Lecture 16 Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III The Powertrain of Human Metabolism (verview) CARBHYDRATES PRTEINS
Lecture 16 Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III The Powertrain of Human Metabolism (verview) CARBHYDRATES PRTEINS
Electron Transport Chain and Oxidative phosphorylation
 Electron Transport Chain and Oxidative phosphorylation So far we have discussed the catabolism involving oxidation of 6 carbons of glucose to CO 2 via glycolysis and CAC without any oxygen molecule directly
Electron Transport Chain and Oxidative phosphorylation So far we have discussed the catabolism involving oxidation of 6 carbons of glucose to CO 2 via glycolysis and CAC without any oxygen molecule directly
number Done by Corrected by Doctor Nayef Karadsheh
 number 17 Done by Abdulrahman Alhanbali Corrected by Lara Abdallat Doctor Nayef Karadsheh 1 P a g e Pentose Phosphate Pathway (PPP) Or Hexose Monophosphate Shunt In this lecture We will talk about the
number 17 Done by Abdulrahman Alhanbali Corrected by Lara Abdallat Doctor Nayef Karadsheh 1 P a g e Pentose Phosphate Pathway (PPP) Or Hexose Monophosphate Shunt In this lecture We will talk about the
Electron transport chain, oxidative phosphorylation, mitochondrial transport systems
 Electron transport chain, oxidative phosphorylation, mitochondrial transport systems JAN ILLNER Respiratory chain & oxidative phosphorylation INTERMEMBRANE SPACE ubiquinone cytochrome c ATPase Production
Electron transport chain, oxidative phosphorylation, mitochondrial transport systems JAN ILLNER Respiratory chain & oxidative phosphorylation INTERMEMBRANE SPACE ubiquinone cytochrome c ATPase Production
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
* * Days after darkness transition
 Protein (mg g -1 FW) 20 15 10 5 * * * * * * WT ivdh-1 d2hgdh1-2 etfqo-1 etfqo-2 35S:IVDH 35S:D2HGDH * * * * 0 0 3 7 10 Days after darkness transition Supplemental Figure 1. Protein content in Arabidopsis
Protein (mg g -1 FW) 20 15 10 5 * * * * * * WT ivdh-1 d2hgdh1-2 etfqo-1 etfqo-2 35S:IVDH 35S:D2HGDH * * * * 0 0 3 7 10 Days after darkness transition Supplemental Figure 1. Protein content in Arabidopsis
Supporting Information
 Comparative Proteomic Study of Fatty Acid-treated Myoblasts Reveals Role of Cox-2 in Palmitate-induced Insulin Resistance Supporting Information Xiulan Chen 1#, Shimeng Xu 2,3#, Shasha Wei 1,3, Yaqin Deng
Comparative Proteomic Study of Fatty Acid-treated Myoblasts Reveals Role of Cox-2 in Palmitate-induced Insulin Resistance Supporting Information Xiulan Chen 1#, Shimeng Xu 2,3#, Shasha Wei 1,3, Yaqin Deng
Table Ia. Differentially regulated genes in the arr2 loss-of-function mutant compared to wildtype
 i ANHANG Table Ia. Differentially regulated genes in the arr2 loss-of-function mutant compared to wildtype AGI No. Putative Function Fold Change a Abscisic acid-related AT4G24960 abscisic acid-induced
i ANHANG Table Ia. Differentially regulated genes in the arr2 loss-of-function mutant compared to wildtype AGI No. Putative Function Fold Change a Abscisic acid-related AT4G24960 abscisic acid-induced
Introduction to Detoxification Enzymes. Evolutionary Response to Chemicals in the Environment
 Evolutionary Response to Chemicals in the Environment Introduction to Detoxification Enzymes 1. Introduction to Detoxification enzymes (focusing on cytochrome P450s) 2. Evolutionary response to toxins
Evolutionary Response to Chemicals in the Environment Introduction to Detoxification Enzymes 1. Introduction to Detoxification enzymes (focusing on cytochrome P450s) 2. Evolutionary response to toxins
Supplementary Data Transcriptome analysis in response to UV-B stress in soybean [Glycine max (L.) Merr.]
![Supplementary Data Transcriptome analysis in response to UV-B stress in soybean [Glycine max (L.) Merr.] Supplementary Data Transcriptome analysis in response to UV-B stress in soybean [Glycine max (L.) Merr.]](/thumbs/85/91947206.jpg) AJCS 6(9):195-1400 (2012) ISSN:185-27 Supplementary Data Transcriptome analysis in response to UV-B stress in soybean [Glycine max (L.) Merr.] Jun-Cheol Moon 1,2, Won Cheol Yim 1, Jae-Eun Lee, Young-Up
AJCS 6(9):195-1400 (2012) ISSN:185-27 Supplementary Data Transcriptome analysis in response to UV-B stress in soybean [Glycine max (L.) Merr.] Jun-Cheol Moon 1,2, Won Cheol Yim 1, Jae-Eun Lee, Young-Up
MITOCHONDRIA LECTURES OVERVIEW
 1 MITOCHONDRIA LECTURES OVERVIEW A. MITOCHONDRIA LECTURES OVERVIEW Mitochondrial Structure The arrangement of membranes: distinct inner and outer membranes, The location of ATPase, DNA and ribosomes The
1 MITOCHONDRIA LECTURES OVERVIEW A. MITOCHONDRIA LECTURES OVERVIEW Mitochondrial Structure The arrangement of membranes: distinct inner and outer membranes, The location of ATPase, DNA and ribosomes The
Lecture: 26 OXIDATION OF FATTY ACIDS
 Lecture: 26 OXIDATION OF FATTY ACIDS Fatty acids obtained by hydrolysis of fats undergo different oxidative pathways designated as alpha ( ), beta ( ) and omega ( ) pathways. -oxidation -Oxidation of fatty
Lecture: 26 OXIDATION OF FATTY ACIDS Fatty acids obtained by hydrolysis of fats undergo different oxidative pathways designated as alpha ( ), beta ( ) and omega ( ) pathways. -oxidation -Oxidation of fatty
MBBS First Professional (Part-I Examination) BIOCHEMISTRY Model Paper
 MBBS FIRST PROFESSIONAL (PART I) Biochemistry (MCQs) Maximum Marks: 35 Time Allowed: 45 minutes Q 1. What is not correct about cell membrane? A. Contains two layers of phospholipids B. May contain receptor
MBBS FIRST PROFESSIONAL (PART I) Biochemistry (MCQs) Maximum Marks: 35 Time Allowed: 45 minutes Q 1. What is not correct about cell membrane? A. Contains two layers of phospholipids B. May contain receptor
FREE ENERGY Reactions involving free energy: 1. Exergonic 2. Endergonic
 BIOENERGETICS FREE ENERGY It is the portion of the total energy change in a system that is available for doing work at constant temperature and pressure; it is represented as ΔG. Reactions involving free
BIOENERGETICS FREE ENERGY It is the portion of the total energy change in a system that is available for doing work at constant temperature and pressure; it is represented as ΔG. Reactions involving free
Cornstarch
 Electronic Supplementary Material (ESI) for Food & Function. This journal is The Royal Society of Chemistry 2018 Supplementary data : Supplementary Table 1: Diet composition (g/kg) on the basis of the
Electronic Supplementary Material (ESI) for Food & Function. This journal is The Royal Society of Chemistry 2018 Supplementary data : Supplementary Table 1: Diet composition (g/kg) on the basis of the
Table S1 Differentially expressed genes showing > 2 fold changes and p <0.01 for 0.01 mm NS-398, 0.1mM ibuprofen and COX-2 RNAi.
 Table S1 Differentially expressed genes showing > 2 fold changes and p
Table S1 Differentially expressed genes showing > 2 fold changes and p
LIPID METABOLISM
 LIPID METABOLISM LIPOGENESIS LIPOGENESIS LIPOGENESIS FATTY ACID SYNTHESIS DE NOVO FFA in the blood come from :- (a) Dietary fat (b) Dietary carbohydrate/protein in excess of need FA TAG Site of synthesis:-
LIPID METABOLISM LIPOGENESIS LIPOGENESIS LIPOGENESIS FATTY ACID SYNTHESIS DE NOVO FFA in the blood come from :- (a) Dietary fat (b) Dietary carbohydrate/protein in excess of need FA TAG Site of synthesis:-
BCM 221 LECTURES OJEMEKELE O.
 BCM 221 LECTURES BY OJEMEKELE O. OUTLINE INTRODUCTION TO LIPID CHEMISTRY STORAGE OF ENERGY IN ADIPOCYTES MOBILIZATION OF ENERGY STORES IN ADIPOCYTES KETONE BODIES AND KETOSIS PYRUVATE DEHYDROGENASE COMPLEX
BCM 221 LECTURES BY OJEMEKELE O. OUTLINE INTRODUCTION TO LIPID CHEMISTRY STORAGE OF ENERGY IN ADIPOCYTES MOBILIZATION OF ENERGY STORES IN ADIPOCYTES KETONE BODIES AND KETOSIS PYRUVATE DEHYDROGENASE COMPLEX
Biochemistry: A Short Course
 Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 28 Fatty Acid Synthesis 2013 W. H. Freeman and Company Chapter 28 Outline 1. The first stage of fatty acid synthesis is transfer
Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 28 Fatty Acid Synthesis 2013 W. H. Freeman and Company Chapter 28 Outline 1. The first stage of fatty acid synthesis is transfer
1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled
 Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Biologic Oxidation BIOMEDICAL IMPORTAN
 Biologic Oxidation BIOMEDICAL IMPORTAN Chemically, oxidation is defined as the removal of electrons and reduction as the gain of electrons. Thus, oxidation is always accompanied by reduction of an electron
Biologic Oxidation BIOMEDICAL IMPORTAN Chemically, oxidation is defined as the removal of electrons and reduction as the gain of electrons. Thus, oxidation is always accompanied by reduction of an electron
number Done by Corrected by Doctor
 number 19 Done by حسام ابو عوض Corrected by وسيم ابو عبيدة Doctor د.نايف 1 P a g e GAGs and Glycoproteins: GAGs: long, unbranched heteropolysaccharides, made from زunits repeating disaccharide [Acidic
number 19 Done by حسام ابو عوض Corrected by وسيم ابو عبيدة Doctor د.نايف 1 P a g e GAGs and Glycoproteins: GAGs: long, unbranched heteropolysaccharides, made from زunits repeating disaccharide [Acidic
Biosynthesis of Fatty Acids
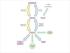 Biosynthesis of Fatty Acids Fatty acid biosynthesis takes place in the cytosol rather than the mitochondria and requires a different activation mechanism and different enzymes and coenzymes than fatty
Biosynthesis of Fatty Acids Fatty acid biosynthesis takes place in the cytosol rather than the mitochondria and requires a different activation mechanism and different enzymes and coenzymes than fatty
SUPPLEMENTARY INFORMATION
 Supplementary igure 1. The expression level of PP4 (At4g16860) is diminished in the rpp4 mutant (SK017569). The mna was extracted to analyze the expression level of PP4 using quantitative PC (qpc). Gene
Supplementary igure 1. The expression level of PP4 (At4g16860) is diminished in the rpp4 mutant (SK017569). The mna was extracted to analyze the expression level of PP4 using quantitative PC (qpc). Gene
BIO th November 2002 KEY EXAM III
 1 BIO 451 15 th November 2002 KEY EXAM III This exam may be taken apart for grading. Please PRINT your name on each page. If you do not have sufficient room for your answer in the space provided, please
1 BIO 451 15 th November 2002 KEY EXAM III This exam may be taken apart for grading. Please PRINT your name on each page. If you do not have sufficient room for your answer in the space provided, please
PCB 3023 Exam 4 - Form A First and Last Name
 PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
ANSC/NUTR 618 LIPIDS & LIPID METABOLISM. Fatty Acid Elongation and Desaturation
 ANSC/NUTR 618 LIPIDS & LIPID METABOLISM I. Fatty acid elongation A. General 1. At least 60% of fatty acids in triacylglycerols are C18. 2. Free palmitic acid (16:0) synthesized in cytoplasm is elongated
ANSC/NUTR 618 LIPIDS & LIPID METABOLISM I. Fatty acid elongation A. General 1. At least 60% of fatty acids in triacylglycerols are C18. 2. Free palmitic acid (16:0) synthesized in cytoplasm is elongated
number Done by Corrected by Doctor Dr.Diala
 number 32 Done by Mousa Salah Corrected by Bahaa Najjar Doctor Dr.Diala 1 P a g e In the last lecture we talked about the common processes between all amino acids which are: transamination, deamination,
number 32 Done by Mousa Salah Corrected by Bahaa Najjar Doctor Dr.Diala 1 P a g e In the last lecture we talked about the common processes between all amino acids which are: transamination, deamination,
Protein Class/Name. Pathways. bat00250, bat00280, bat00410, bat00640, bat bat00270, bat00330, bat00410, bat00480
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2017 Supplemental Table 1. Proteins Increased in Either Soil or Laboratory media Table 1A. Proteins Increased
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2017 Supplemental Table 1. Proteins Increased in Either Soil or Laboratory media Table 1A. Proteins Increased
Energy storage in cells
 Energy storage in cells Josef Fontana EC - 58 Overview of the lecture Introduction to the storage substances of human body Overview of storage compounds in the body Glycogen metabolism Structure of glycogen
Energy storage in cells Josef Fontana EC - 58 Overview of the lecture Introduction to the storage substances of human body Overview of storage compounds in the body Glycogen metabolism Structure of glycogen
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
f(x) = x R² = RPKM (M8.MXB) f(x) = x E-014 R² = 1 RPKM (M31.
 14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
Protein Class/Name KEGG Pathways
 Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2017 Supplemental Table 4. Proteins Increased in Either Soil medium at 37 o C and 25 o C Table 4A. Proteins
Electronic Supplementary Material (ESI) for Analyst. This journal is The Royal Society of Chemistry 2017 Supplemental Table 4. Proteins Increased in Either Soil medium at 37 o C and 25 o C Table 4A. Proteins
Propagation of the Signal
 OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Supplementary Information
 Supplementary Information Figure S1 Sequencing quality. Sample screenshot of re-sequencing raw file quality generated by FastQC. For both the graphs, scores were based on the Illumina 1.5 quality score.
Supplementary Information Figure S1 Sequencing quality. Sample screenshot of re-sequencing raw file quality generated by FastQC. For both the graphs, scores were based on the Illumina 1.5 quality score.
number Done by Corrected by Doctor Faisal Al-Khatibe
 number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
Chapter 9: Biochemical Mechanisms for Information Storage at the Cellular Level. From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D.
 Chapter 9: Biochemical Mechanisms for Information Storage at the Cellular Level From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D. Chapter 9: Dendritic Spine Figure 1 Summary: Three Primary
Chapter 9: Biochemical Mechanisms for Information Storage at the Cellular Level From Mechanisms of Memory, second edition By J. David Sweatt, Ph.D. Chapter 9: Dendritic Spine Figure 1 Summary: Three Primary
Validated Mouse Quantitative RT-PCR Genes Gene Gene Bank Accession Number Name
 5HT1a NM_008308.4 Serotonin Receptor 1a 5HT1b NM_010482.1 Serotonin Receptor 1b 5HT2a NM_172812.2 Serotonin Receptor 2a ACO-2 NM_080633.2 Aconitase 2 Adora2A NM_009630.02 Adenosine A2a Receptor Aif1 (IbaI)
5HT1a NM_008308.4 Serotonin Receptor 1a 5HT1b NM_010482.1 Serotonin Receptor 1b 5HT2a NM_172812.2 Serotonin Receptor 2a ACO-2 NM_080633.2 Aconitase 2 Adora2A NM_009630.02 Adenosine A2a Receptor Aif1 (IbaI)
Plant Cell, Development & Ultrastructure
 PCDU Lecture Plant Cell, Development & Ultrastructure Plant Cell Biology Labs Download at: http://goo.gl/111tha Peroxisomes (Microbodies/) Catabolism of Fatty Acids Catabolism of Fatty Acids Glyoxysomes
PCDU Lecture Plant Cell, Development & Ultrastructure Plant Cell Biology Labs Download at: http://goo.gl/111tha Peroxisomes (Microbodies/) Catabolism of Fatty Acids Catabolism of Fatty Acids Glyoxysomes
MBB 115:511 and 694:407 Final Exam Niederman/Deis
 MBB 115:511 and 694:407 Final Exam Niederman/Deis Tue. Dec. 19, 2006 Name Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth two points.
MBB 115:511 and 694:407 Final Exam Niederman/Deis Tue. Dec. 19, 2006 Name Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth two points.
بسم هللا الرحمن الرحيم
 بسم هللا الرحمن الرحيم -Please refer to the slides from (4-20) -Slides (4, 5) -Oxidative phosphorylation consists of 2 parts: 1.electron transport chain (series of electron transport proteins much filled
بسم هللا الرحمن الرحيم -Please refer to the slides from (4-20) -Slides (4, 5) -Oxidative phosphorylation consists of 2 parts: 1.electron transport chain (series of electron transport proteins much filled
Signal Transduction: G-Protein Coupled Receptors
 Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Dental Students Biochemistry Exam V Questions ( Note: In all cases, the only correct answer is the best answer)
 Dental Students Biochemistry Exam V Questions - 2006 ( Note: In all cases, the only correct answer is the best answer) 1. Essential fatty acids are: A. precursors of biotin B. precursors of tyrosine C.
Dental Students Biochemistry Exam V Questions - 2006 ( Note: In all cases, the only correct answer is the best answer) 1. Essential fatty acids are: A. precursors of biotin B. precursors of tyrosine C.
BIOCHEMISTRY - CLUTCH REVIEW 6.
 !! www.clutchprep.com CONCEPT: AMINO ACID OXIDATION Urea cycle occurs in liver, removes amino groups from amino acids so they may enter the citric acid cycle 2 nitrogen enter the cycle to ultimately leave
!! www.clutchprep.com CONCEPT: AMINO ACID OXIDATION Urea cycle occurs in liver, removes amino groups from amino acids so they may enter the citric acid cycle 2 nitrogen enter the cycle to ultimately leave
Oxidative Phosphorylation
 Oxidative Phosphorylation Energy from Reduced Fuels Is Used to Synthesize ATP in Animals Carbohydrates, lipids, and amino acids are the main reduced fuels for the cell. Electrons from reduced fuels are
Oxidative Phosphorylation Energy from Reduced Fuels Is Used to Synthesize ATP in Animals Carbohydrates, lipids, and amino acids are the main reduced fuels for the cell. Electrons from reduced fuels are
Lecture 10 - Protein Turnover and Amino Acid Catabolism
 Lecture 10 - Protein Turnover and Amino Acid Catabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction 2 Proteins are degraded into amino acids. Protein
Lecture 10 - Protein Turnover and Amino Acid Catabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction 2 Proteins are degraded into amino acids. Protein
BIOLOGY 103 Spring 2001 MIDTERM LAB SECTION
 BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
Lecture 16. Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III
 Lecture 16 Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III The Powertrain of Human Metabolism (verview) CARBHYDRATES PRTEINS
Lecture 16 Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III The Powertrain of Human Metabolism (verview) CARBHYDRATES PRTEINS
Hind Abu Tawileh. Moh Tarek & Razi Kittaneh. Ma moun
 26 Hind Abu Tawileh Moh Tarek & Razi Kittaneh... Ma moun Cofactors are non-protein compounds, they are divided into 3 types: Protein-based. Metals: if they are bounded tightly (covalently) to the enzyme
26 Hind Abu Tawileh Moh Tarek & Razi Kittaneh... Ma moun Cofactors are non-protein compounds, they are divided into 3 types: Protein-based. Metals: if they are bounded tightly (covalently) to the enzyme
Chapter 14. Energy conversion: Energy & Behavior
 Chapter 14 Energy conversion: Energy & Behavior Why do you Eat and Breath? To generate ATP Foods, Oxygen, and Mitochodria Cells Obtain Energy by the Oxidation of Organic Molecules Food making ATP making
Chapter 14 Energy conversion: Energy & Behavior Why do you Eat and Breath? To generate ATP Foods, Oxygen, and Mitochodria Cells Obtain Energy by the Oxidation of Organic Molecules Food making ATP making
Supporting Information
 t t l l t l Supporting Information Zemach et al. 1.173/pnas.1969517.8 TE > 1 bp Leaf n o a Embryo h y C H H m e Endosperm Seedling root Seedling shoot -3-2 -1 1 2 3 4 3 2 1-1 -2-3 Fig. S1. Distribution
t t l l t l Supporting Information Zemach et al. 1.173/pnas.1969517.8 TE > 1 bp Leaf n o a Embryo h y C H H m e Endosperm Seedling root Seedling shoot -3-2 -1 1 2 3 4 3 2 1-1 -2-3 Fig. S1. Distribution
Protein Name. IFLENVIR,DSVTYTEHAK,TV TALDVVYALK,KTVTALDVV YALK,TVTALDVVYALKR,IF LENVIRDSVTYTEHAK gi Histone H2B
 Table 1. A functional category list of proteins (Lentinula edodes) identified by 1-DGE and nesi-lc-ms/ms. The table lists indicated fraction numbers, matching peptides, scores, accession numbers, protein
Table 1. A functional category list of proteins (Lentinula edodes) identified by 1-DGE and nesi-lc-ms/ms. The table lists indicated fraction numbers, matching peptides, scores, accession numbers, protein
Fatty acids synthesis
 Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
Chemistry 107 Exam 4 Study Guide
 Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
Biochemistry 2 Recita0on Amino Acid Metabolism
 Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001
 BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
Figure 1 Original Advantages of biological reactions being catalyzed by enzymes:
 Enzyme basic concepts, Enzyme Regulation I III Carmen Sato Bigbee, Ph.D. Objectives: 1) To understand the bases of enzyme catalysis and the mechanisms of enzyme regulation. 2) To understand the role of
Enzyme basic concepts, Enzyme Regulation I III Carmen Sato Bigbee, Ph.D. Objectives: 1) To understand the bases of enzyme catalysis and the mechanisms of enzyme regulation. 2) To understand the role of
Chapter 10. Introduction to Nutrition and Metabolism, 3 rd edition David A Bender Taylor & Francis Ltd, London 2002
 Chapter 10 Introduction to Nutrition and Metabolism, 3 rd edition David A Bender Taylor & Francis Ltd, London 2002 Chapter 10: Integration and Control of Metabolism Press the space bar or click the mouse
Chapter 10 Introduction to Nutrition and Metabolism, 3 rd edition David A Bender Taylor & Francis Ltd, London 2002 Chapter 10: Integration and Control of Metabolism Press the space bar or click the mouse
Supplementary Table 8. Genes from supplementary table 4 ordered in the way in which they occur in the heatmap in Figure 4b of the main text.
 Supplementary Table 8. Genes from supplementary table 4 ordered in the way in which they occur in the heatmap in Figure 4b of the main text. gene name protein name reference name strand start positend
Supplementary Table 8. Genes from supplementary table 4 ordered in the way in which they occur in the heatmap in Figure 4b of the main text. gene name protein name reference name strand start positend
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/ MCQs. Location : 102, 105, 106, 301, 302
 FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
Anabolism of Fatty acids (Anabolic Lynen spiral) Glycerol and Triglycerides
 Anabolism of Fatty acids (Anabolic Lynen spiral) Glycerol and Triglycerides Anabolism of fatty acids Fatty acids are not stored in the body free. They are a source of energy in the form of triglycerides
Anabolism of Fatty acids (Anabolic Lynen spiral) Glycerol and Triglycerides Anabolism of fatty acids Fatty acids are not stored in the body free. They are a source of energy in the form of triglycerides
Catabolism of Carbon skeletons of Amino acids. Amino acid metabolism
 Catabolism of Carbon skeletons of Amino acids Amino acid metabolism Carbon skeleton Carbon Skeleton a carbon skeleton is the internal structure of organic molecules. Carbon Arrangements The arrangement
Catabolism of Carbon skeletons of Amino acids Amino acid metabolism Carbon skeleton Carbon Skeleton a carbon skeleton is the internal structure of organic molecules. Carbon Arrangements The arrangement
Table S1 Differential genes repressed in copt2-1 seedlings. MIPS code Ratio gene annotation gene name -Fe-Cu/ -Fe+Cu/ +Fe+Cu -Fe-Cu/ -Fe+Cu
 Table S1. Differential genes repressed in copt2-1 seedlings. MIPS code, the microarray values under -Fe-Cu, -Fe+Cu and +Fe+Cu, gene annotation and gene name are indicated. MIPS code Ratio gene annotation
Table S1. Differential genes repressed in copt2-1 seedlings. MIPS code, the microarray values under -Fe-Cu, -Fe+Cu and +Fe+Cu, gene annotation and gene name are indicated. MIPS code Ratio gene annotation
MG132. GRXS17:V5-His MG132 GRXS17:HA. RuBisCo. Supplemental Data. Walton et al. (2015). Plant Cell /tpc
 MG132 GRXS17:V5-His MG132 GRXS17:HA RuBisCo Supplemental Figure 1. Degradation of GRXS17 by the 26S proteasome. (A) Cell-free degradation assay. Recombinant protein GRXS17:V5-HIS was incubated with total
MG132 GRXS17:V5-His MG132 GRXS17:HA RuBisCo Supplemental Figure 1. Degradation of GRXS17 by the 26S proteasome. (A) Cell-free degradation assay. Recombinant protein GRXS17:V5-HIS was incubated with total
Schmid et al.: A gene expression map of Arabidopsis Development
 Supplementary Table 4. Coexpression of gene families. Average of Pearson correlation coefficients (PCC) and number of members of families shown in Fig. 5b. Gene family (above 95 th percentile) # Genes
Supplementary Table 4. Coexpression of gene families. Average of Pearson correlation coefficients (PCC) and number of members of families shown in Fig. 5b. Gene family (above 95 th percentile) # Genes
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
BIOSYNTHESIS OF FATTY ACIDS. doc. Ing. Zenóbia Chavková, CSc.
 BIOSYNTHESIS OF FATTY ACIDS doc. Ing. Zenóbia Chavková, CSc. The pathway for the of FAs is not the reversal of the oxidation pathway Both pathways are separated within different cellular compartments In
BIOSYNTHESIS OF FATTY ACIDS doc. Ing. Zenóbia Chavková, CSc. The pathway for the of FAs is not the reversal of the oxidation pathway Both pathways are separated within different cellular compartments In
SDR families. SDR10E Fatty acyl-coa reductase FACR1_HUMAN 544 x x SDR11E 3 beta-hydroxysteroid dehydrogenase 3BHS1_HUMAN 254 x x
 SDR1E UDP-glucose 4-epimerase GALE_HUMAN 5305 x x x x SDR2E dtdp-d-glucose 4,6-dehydratase TGDS_HUMAN 4194 x x x x SDR3E GDP-mannose 4,6 dehydratase GMDS_HUMAN 2314 x x x x SDR4E GDP-L-fucose synthetase
SDR1E UDP-glucose 4-epimerase GALE_HUMAN 5305 x x x x SDR2E dtdp-d-glucose 4,6-dehydratase TGDS_HUMAN 4194 x x x x SDR3E GDP-mannose 4,6 dehydratase GMDS_HUMAN 2314 x x x x SDR4E GDP-L-fucose synthetase
Biochemical networks related to carbon accumulation in sugarcane
 Biochemical networks related to carbon accumulation in sugarcane Marcos Buckeridge Department of Botany, Laboratory of Plant Physiological Ecology IB USP How are we approaching the sugarcane metabolic
Biochemical networks related to carbon accumulation in sugarcane Marcos Buckeridge Department of Botany, Laboratory of Plant Physiological Ecology IB USP How are we approaching the sugarcane metabolic
Cell wall components:
 Main differences between plant and animal cells: Plant cells have: cell walls, a large central vacuole, plastids and turgor pressure. The Cell Wall The primary cell wall is capable of rapid expansion during
Main differences between plant and animal cells: Plant cells have: cell walls, a large central vacuole, plastids and turgor pressure. The Cell Wall The primary cell wall is capable of rapid expansion during
Main differences between plant and animal cells: Plant cells have: cell walls, a large central vacuole, plastids and turgor pressure.
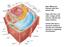 Main differences between plant and animal cells: Plant cells have: cell walls, a large central vacuole, plastids and turgor pressure. Animal cells have a lysosome (related to vacuole) and centrioles (function
Main differences between plant and animal cells: Plant cells have: cell walls, a large central vacuole, plastids and turgor pressure. Animal cells have a lysosome (related to vacuole) and centrioles (function
Chapter 3. Biological Molecules Great and Small
 Chapter 3 Biological Molecules Great and Small Chapter Goal: Understanding how cells use small building blocks to build larger molecules and how some of those molecules then fold into 3-D shapes Key Questions
Chapter 3 Biological Molecules Great and Small Chapter Goal: Understanding how cells use small building blocks to build larger molecules and how some of those molecules then fold into 3-D shapes Key Questions
BCMB 3100 Introduction to Coenzymes & Vitamins
 BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
Leen Alsahele. Razan Al-zoubi ... Faisal
 25 Leen Alsahele Razan Al-zoubi... Faisal last time we started talking about regulation of fatty acid synthesis and degradation *regulation of fatty acid synthesis by: 1- regulation of acetyl CoA carboxylase
25 Leen Alsahele Razan Al-zoubi... Faisal last time we started talking about regulation of fatty acid synthesis and degradation *regulation of fatty acid synthesis by: 1- regulation of acetyl CoA carboxylase
Biochemistry - I SPRING Mondays and Wednesdays 9:30-10:45 AM (MR-1307) Lectures Based on Profs. Kevin Gardner & Reza Khayat
 Biochemistry - I Mondays and Wednesdays 9:30-10:45 AM (MR-1307) SPRING 2017 Lectures 23-24 Based on Profs. Kevin Gardner & Reza Khayat 1 Outline Anatomy of the mitochondrion Electron transfer Fractionation
Biochemistry - I Mondays and Wednesdays 9:30-10:45 AM (MR-1307) SPRING 2017 Lectures 23-24 Based on Profs. Kevin Gardner & Reza Khayat 1 Outline Anatomy of the mitochondrion Electron transfer Fractionation
19 Oxidative Phosphorylation and Photophosphorylation W. H. Freeman and Company
 19 Oxidative Phosphorylation and Photophosphorylation 2013 W. H. Freeman and Company CHAPTER 19 Oxidative Phosphorylation and Photophosphorylation Key topics: Electron transport chain in mitochondria Capture
19 Oxidative Phosphorylation and Photophosphorylation 2013 W. H. Freeman and Company CHAPTER 19 Oxidative Phosphorylation and Photophosphorylation Key topics: Electron transport chain in mitochondria Capture
Cellular functions of protein degradation
 Protein Degradation Cellular functions of protein degradation 1. Elimination of misfolded and damaged proteins: Environmental toxins, translation errors and genetic mutations can damage proteins. Misfolded
Protein Degradation Cellular functions of protein degradation 1. Elimination of misfolded and damaged proteins: Environmental toxins, translation errors and genetic mutations can damage proteins. Misfolded
Biochemistry Sheet 27 Fatty Acid Synthesis Dr. Faisal Khatib
 Page1 بسم رلاهللا On Thursday, we discussed the synthesis of fatty acids and its regulation. We also went on to talk about the synthesis of Triacylglycerol (TAG). Last time, we started talking about the
Page1 بسم رلاهللا On Thursday, we discussed the synthesis of fatty acids and its regulation. We also went on to talk about the synthesis of Triacylglycerol (TAG). Last time, we started talking about the
Coenzymes. Coenzymes 9/11/2018. BCMB 3100 Introduction to Coenzymes & Vitamins
 BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
Functional mechanisms of drought tolerance in maize
 Functional mechanisms of drought tolerance in maize Nepolean Thirunavukkarasu Division of Genetics Indian Agricultural Research Institute New Delhi-110012 tnepolean@iari.resi.in tnepolean@gmail.com Mt
Functional mechanisms of drought tolerance in maize Nepolean Thirunavukkarasu Division of Genetics Indian Agricultural Research Institute New Delhi-110012 tnepolean@iari.resi.in tnepolean@gmail.com Mt
Oxidative Phosphorylation
 Electron Transport Chain (overview) The NADH and FADH 2, formed during glycolysis, β- oxidation and the TCA cycle, give up their electrons to reduce molecular O 2 to H 2 O. Electron transfer occurs through
Electron Transport Chain (overview) The NADH and FADH 2, formed during glycolysis, β- oxidation and the TCA cycle, give up their electrons to reduce molecular O 2 to H 2 O. Electron transfer occurs through
Supporting Information
 Supporting Information Supplemental Figure Legends Supplementary Figure 1. Experimental Design. A. To benchmark the performance of software parameter sets proteins were extracted from a pool of Arabidopsis
Supporting Information Supplemental Figure Legends Supplementary Figure 1. Experimental Design. A. To benchmark the performance of software parameter sets proteins were extracted from a pool of Arabidopsis
SP GPI Found in exosomes
 Accession /Name/Functional group Spectr. counts SP GPI Found in exosomes Description Metabolism and energy production (21) CNAG_06770 fructose-bisphosphate aldolase 0.94 - - + gluconeogenesis and glycolysis
Accession /Name/Functional group Spectr. counts SP GPI Found in exosomes Description Metabolism and energy production (21) CNAG_06770 fructose-bisphosphate aldolase 0.94 - - + gluconeogenesis and glycolysis
BOT 6516 Plant Metabolism
 BOT 6516 Plant Metabolism Lecture 19 Nitrate & Sulfur Assimilation Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bot6516/index.html Overview of Nitrate Assimilation 16.34 What does Δψ
BOT 6516 Plant Metabolism Lecture 19 Nitrate & Sulfur Assimilation Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bot6516/index.html Overview of Nitrate Assimilation 16.34 What does Δψ
Revision. camp pathway
 االله الرحمن الرحيم بسم Revision camp pathway camp pathway Revision camp pathway Adenylate cyclase Adenylate Cyclase enzyme Adenylate cyclase catalyses the formation of camp from ATP. Stimulation or inhibition
االله الرحمن الرحيم بسم Revision camp pathway camp pathway Revision camp pathway Adenylate cyclase Adenylate Cyclase enzyme Adenylate cyclase catalyses the formation of camp from ATP. Stimulation or inhibition
