Retroviruses must gain access to the nucleus to replicate.
|
|
|
- Meagan Blair
- 5 years ago
- Views:
Transcription
1 Nuclear entry and CRM1-dependent nuclear export of the Rous sarcoma virus Gag polyprotein Lisa Z. Scheifele*, Rachel A. Garbitt, Jonathan D. Rhoads, and Leslie J. Parent* *Cell and Molecular Biology Program and Departments of Microbiology and Immunology and Medicine, Pennsylvania State University College of Medicine, 500 University Drive, Hershey, PA Edited by John M. Coffin, Tufts University School of Medicine, Boston, MA, and approved January 17, 2002 (received for review December 6, 2001) The retroviral Gag polyprotein directs budding from the plasma membrane of infected cells. Until now, it was believed that Gag proteins of type C retroviruses, including the prototypic oncoretrovirus Rous sarcoma virus, were synthesized on cytosolic ribosomes and targeted directly to the plasma membrane. Here we reveal a previously unknown step in the subcellular trafficking of the Gag protein, that of transient nuclear localization. We have identified a targeting signal within the N-terminal matrix domain that facilitates active nuclear import of the Gag polyprotein. We also found that Gag is transported out of the nucleus through the CRM1 nuclear export pathway, based on observations that treatment of virus-expressing cells with leptomycin B resulted in the redistribution of Gag proteins from the cytoplasm to the nucleus. Internal deletion of the C-terminal portion of the Gag p10 region resulted in the nuclear sequestration of Gag and markedly diminished budding, suggesting that the nuclear export signal might reside within p10. Finally, we observed that a previously described matrix mutant, Myr1E, was insensitive to the effects of leptomycin B, apparently bypassing the nuclear compartment during virus assembly. Myr1E has a defect in genomic RNA packaging, implying that nuclear localization of Gag might be involved in viral RNA interactions. Taken together, these findings provide evidence that nuclear entry and egress of the Gag polyprotein are intrinsic components of the Rous sarcoma virus assembly pathway. Retroviruses must gain access to the nucleus to replicate. After receptor binding, entry, and reverse transcription, the integration-competent nucleoprotein complex (called the preintegration complex or PIC) enters the nucleus. For oncoretroviruses like Rous sarcoma virus (RSV) that primarily infect dividing cells, the PIC awaits breakdown of the nuclear envelope during mitosis for nuclear entry. Lentiviruses including HIV-1 infect nondividing cells, and PICs are transported through intact nuclear envelopes. HIV-1 nuclear entry is complex, and redundant signals have been identified in the viral matrix (MA), integrase, and Vpr proteins (reviewed in ref. 1). The recent report that RSV can replicate at low levels in quiescent cells does raise the possibility that a viral protein might mediate active nuclear targeting of the RSV PIC (2). After nuclear entry of the PIC and proviral integration, viral RNA is transcribed, and unspliced genome-length viral mrnas must exit the nucleus for translation into viral structural proteins and encapsidation into virions. The nuclear export of introncontaining mrnas is normally inhibited by cellular mechanisms, so retroviruses must circumvent this obstacle. Lentiviruses encode trans-acting factors such as the HIV-1 Rev protein to mediate nuclear export of intron-containing viral RNAs (3). Oncoretroviruses including RSV lack Rev-like transport factors and instead have cis-acting constitutive transport elements to facilitate the export of unspliced viral RNA (4). The regulation of nuclear transport is mediated by components of the nuclear pore complex (NPC) and the superfamily of nuclear transport receptors called importins. Although passive diffusion of small molecules can occur through NPCs, nucleocytoplasmic transport of nearly all proteins and RNAs is mediated by an active mechanism with directionality provided by the Ran-GTPase system (reviewed in refs. 3 and 5). Facilitated nuclear import of cytosolic proteins relies on specific interactions between importins and nuclear localization signals (NLSs). Classical examples of NLSs are the highly basic motifs of simian virus 40 T antigen and nucleoplasmin, although many additional nonclassical sequences have been shown to function as NLSs (1, 5, 6). The nuclear export of proteins and RNAs is also a signal-dependent process mediated by soluble receptors called exportins (3, 5). The best-characterized nuclear export signals (NESs) were initially identified in the HIV-1 Rev protein (7) and the protein kinase A inhibitor (8). These leucine-rich NESs consist of short peptide sequences with four closely spaced hydrophobic residues, a motif recognized by the CRM1 nuclear export receptor (9, 10). Leptomycin B (LMB) attaches to the central domain of CRM1 to disrupt its interaction with the NES, making LMB a specific tool for studying CRM1-mediated nuclear export (9, 11). After nuclear export of full-length viral mrnas, Gag and Gag-Pol proteins are synthesized on free ribosomes. Initial Gag Gag contacts occur between I (interaction) domains found within the NC region (12 14). RNA is essential early in assembly as scaffolding for the assembly of Gag multimers (15, 16). Gag RNA interactions also mediate selective encapsidation of the viral genome, a noncovalently linked RNA dimer, through association of NC and the cis-acting packaging element (17, 18). Assembly intermediates are directed to the plasma membrane via the M (membrane-binding) domain in the N-terminal MA sequence of Gag (19, 20). Because the RSV M domain has suboptimal membrane-binding activity, I domains provide cooperative protein protein interactions to stabilize membrane association (14). Final release of the virus particle is controlled by the late (L) domain (21, 22). After budding, the RSV Gag polyprotein precursor is proteolytically cleaved into the structural proteins MA, p2a, p2b, p10, CA, NC, and PR. In this report, we describe the unexpected finding that the RSV Gag protein enters the nucleus via a nuclear-targeting sequence in the MA domain. We show that Gag is subsequently transported into the cytoplasm by using a CRM1-mediated nuclear export pathway. Elimination of this nuclear step correlates with a defect in RNA packaging. These findings demonstrate a previously unknown step in the assembly pathway of RSV. Materials and Methods Viruses, Cells, and Plasmids. Proviral constructs were derived from prcv8 containing the RSV Prague C gag gene of patv8 (23, 24). Plasmids prc.myr1e, p.rc.myr1e, prc.myr2.hb12, This paper was submitted directly (Track II) to the PNAS office. Abbreviations: RSV, Rous sarcoma virus; LMB, leptomycin B; PIC, preintegration complex; NPC, nuclear pore complex; NLS, nuclear localization signal; NES, nuclear export signal; GFP, green fluorescent protein; MA, matrix. To whom reprint requests should be addressed. lparent@psu.edu. The publication costs of this article were defrayed in part by page charge payment. This article must therefore be hereby marked advertisement in accordance with 18 U.S.C solely to indicate this fact PNAS March 19, 2002 vol. 99 no. 6 cgi doi pnas
2 Fig. 1. The RSV Gag protein and derivatives. The RSV Gag protein (Pr76) is depicted at the top, with cleavage proteins MA, p2a, p2b, p10, CA, NC, and PR indicated. RC.Myr0 is a derivative of prcv8 and expresses the wild-type Prague C MA protein (45). Gag-GFP fusion proteins and truncations of MA fused to GFP are depicted below. Note that GFP replaces the PR sequence in Gag-GFP ( PR). Gag-GFP substitution mutants are shown in the bottom panel. The box labeled Src contains the N-terminal 10 residues of the Src oncoprotein. Myristic acid is shown as a zigzag line. pma-green fluorescent protein (GFP) (25), pgag-gfp (26), and psv.myr0.bgbs (12) have been described. QT6 cells, chemically transformed quail fibroblasts, were maintained as described (27, 28). Plasmids encoding C-terminal truncations of the RSV Gag protein fused to GFP (Fig. 1) were made by PCR amplification of prcv8 by using primer USP (25) and a set of downstream primers each containing an ApaI site, for cloning into SacI ApaI sites of pegfp.n2 (CLONTECH). MA truncations included amino acids 1 24, 1 43, 1 66, 1 88, 1 119, and Gag truncations consisted of MA-p2a (amino acid 1 155), MA-p2 (amino acid 1 177), and MA-p2-p10 (amino acid 1 239). Deletions involving p10 were derived from prc. p10.31, prc. p10.52, and prc. QM1 [(29), kind gifts of Becky Craven and John Wills (Pennsylvania State University College of Medicine)] by PCR using primers USP and 5 -TCAGTAT- AGGGGCCCCGAGTCGGCAGGTGGCTCA. pma-flag was made by ligating fragments from pma-gfp (ApaI-Klenow BglII) with pcmv.flag5b (Sigma) (EcoRV- Klenow BglII). Mutations of the gag sequence (myr2.hb12, myr1e, and myr1e ) were cloned into pgag-gfp by SstI BspEI fragment exchange. The IKB GFP expression vector was a kind gift from Thomas Hope (University of Illinois, Chicago). Confocal Microscopy. QT6 cells were examined 18 h after transfection by using a Zeiss LSM 10 BioMed confocal microscope (25). In indicated experiments, cells were grown in medium augmented with LMB at a final concentration of 10 ng ml (18 nm) for 2 h before imaging. For indirect immunofluorescence, cells were seeded in LabTek chamber slides (Nunc), transfected, and fixed in 2% paraformaldeyde or in 3:1 methanol:acetone at 20 C. Cells were rehydrated in PBS, blocked with 1.5% BSA, incubated with polyclonal anti-rsv or anti-ma (24, 27), washed, stained with goat anti-rabbit IgG-FITC (Sigma) or sheep anti-rabbit IgG-Cy3 (Sigma), mounted with SlowFade (Molecular Probes), and analyzed by confocal microscopy. Viral Protein Detection and Budding Assay. For immunoblotting, transfected cells were lysed in radioimmunoprecipitation assay buffer (50 mm Tris, ph mm NaCl 1% Nonidet P % sodium deoxycholate 0.1% SDS), proteins were separated by electrophoresis, transferred to nitrocellulose, probed with anti-gfp Living Colors Peptide Ab (CLONTECH) and antimouse IgG-conjugated HRP (Sigma), and detected by chemiluminescence. Radioimmunoprecipitation assays were performed as described (30, 31). To calculate particle release, Gag protein bands were quantitated by using a PhosphoImager (Molecular Dynamics), and the amount of Gag protein in the medium was divided by the total Gag protein in lysates and medium. Statistical analysis was performed by using a twosample t test. Results We have described mutants of the RSV MA sequence that interfere with viral RNA packaging and dimerization (24, 25). One such mutant, Myr1E (Fig. 1), contains the Src membranebinding domain as an N-terminal extension of the Gag protein. Although Myr1E produces viral particles more efficiently than wild type, it has undetectable infectivity and packages only 40% of wild-type levels of viral RNA. The Myr1E.MA protein is mislocalized, demonstrating greatly enhanced plasma membrane accumulation compared to the wild-type MA protein (25). We hypothesized that the defect in genomic RNA incorporation results from transporting the Myr1E.Gag protein too quickly to the plasma membrane, omitting a cellular compartment or a trafficking pathway required for Gag RNA interactions. This idea led us to further examine the subcellular localization of wild-type and mutant RSV MA and Gag proteins to dissect the targeting determinants within their sequences. Subcellular Distribution of the MA Protein. To study the subcellular localizations of MA and Gag in living cells, C-terminal fusions were made with GFP (Fig. 1) (25, 26). As analyzed by confocal microscopy, MA-GFP was observed throughout the cytosol with strong accumulation inside the nucleus (Fig. 2A) (25). In contrast, the distribution of unconjugated GFP was diffuse through the cell. To determine whether the presence of the foreign GFP protein altered the cellular distribution of MA, the FLAG epitope was fused to the C terminus of MA, and cells were examined by using indirect immunofluorescence with an anti-ma Ab. Similarly to MA-GFP, MA-FLAG also exhibited nuclear staining. Even a small sequence like FLAG could affect protein localization, so wild-type MA was expressed in the absence of other viral proteins by using a proviral vector, and cells were analyzed by indirect immunofluorescence. In these fixed cells, MA appeared within nuclei, confirming our previous results (Fig. 2A, RC.Myr0). Because MA and MA-FLAG are small proteins, they could enter the nucleus by passive diffusion, because in theory proteins up to kda can diffuse through the NPC. In reality, few proteins actually traverse the NPC passively because the directional active transport system maintains strict compartmentalization (3, 5). However, even if MA and MA-FLAG entered by diffusion, they would be unlikely to accumulate in the nucleus under steady state conditions. Furthermore, MA-GFP was concentrated in the nucleus despite its larger molecular weight (52 kda) compared to GFP (33 kda), which was distributed uniformly in the cell. The nuclear concentration of MA-GFP is distinct from the MICROBIOLOGY Scheifele et al. PNAS March 19, 2002 vol. 99 no
3 Fig. 3. Identification of an LMB-sensitive NES within the Gag protein. Live cells transfected with plasmids encoding C-terminal truncations of Gag fused to GFP were examined by confocal microscopy. LMB-treated cells were incubated with 10 ng ml LMB for 2 h before imaging. Fig. 2. Subcellular localization of the RSV MA protein. (A Upper) Subcellular localizations of GFP, MA-GFP, and Gag-GFP fusion proteins were analyzed in live cells using confocal microscopy 18 h after transfection. (Lower) Transfected QT6 cells were fixed and incubated with polyclonal anti-ma Ab and Cy3-conjugated secondary Ab. (B) Immunoblot analysis of intracellular expression levels of indicated proteins from lysates of transfected cells by using an anti-gfp Ab. Molecular weight markers are indicated to the left. (C) Mapping a nuclear-targeting sequence within the MA domain. Plasmids encoding truncations of the MA sequence fused to GFP were transfected and cells fixed, and epifluorescence was detected. The cells were stained with propidium iodide to visualize nuclear DNA, and identical fields are shown. localization of Gag-GFP, which excludes the nucleus and accumulates at focal patches along the plasma membrane (26) (Fig. 2 A). A priori, we expected the MA protein to be localized at the plasma membrane given that MA and Gag share identical N-terminal sequences and therefore contain the same M domain. The unexpected difference in subcellular localization suggested that there must be additional targeting information located either within MA or Gag to account for their distinctive distributions. A conformational change resulting from cleavage of the Gag precursor during maturation might reveal new targeting information within MA, similar in principle to the myristyl-switch mechanism proposed to explain different conformations of the HIV-1 MA and Gag proteins (32 34). Alternatively, cleavage of the MA protein might remove downstreamtargeting signals that are dominant over nuclear targeting in the context of the Gag polyprotein. Nuclear Localization of MA Requires the Presence of the Four -Helix Domain. The localization of MA within the nucleus suggested the presence of an NLS, but sequences resembling a classical motif could not be identified by database searches (35, 36). However, MA is lysine- and arginine-rich, so perhaps the basic residues form an NLS in the three-dimensional conformation of the protein. To ascertain whether a simple nuclear-targeting sequence could be found, deletions were made based on the NMR structure of the N-terminal half of the MA protein, which consists of four overlapping helices and a 3 10 helix joined by flexible loops (37). C-terminal truncations of MA were fused to GFP (Fig. 1). Immunoblot analysis revealed that each fusion protein was stably expressed, and there was no detectable free GFP except in the MA 1 88 construct (Fig. 2B). Examination by confocal microscopy revealed that addition of the first helix (MA 1 24), the first two helices (MA 1 43), or the first three helices (MA 1 66) to GFP resulted in accumulation of the proteins in the cytoplasm (Fig. 2C). Fusion of the first 88 residues of MA (all four helixes) to GFP restored nuclear concentration of the protein, and further extension into the second half of MA maintained nuclear localization (MA and MA 1 140). Identification of a CRM1-Dependent NES Within the Gag Sequence. To identify the sequence required for the change from nuclear to plasma membrane localization, GFP fusion proteins including the p2a, p2 (p2a plus p2b), p10, and CA domains of Gag were examined (Fig. 1). MA-p2a-GFP (data not shown) and MA-p2- GFP fusion proteins displayed the same subcellular localization as MA-GFP with nuclear accumulation (Fig. 3). However, extending through the p10 and CA domains of Gag produced a dramatic alteration in localization: MA-p2-p10-GFP and MAp2-p10-CA-GFP were present exclusively in the cytoplasm. The nuclear exclusion of Gag-GFP fusion proteins including p10 and CA could be explained by either increased molecular weight, precluding passive nuclear entry, or by the presence of a nuclear export activity. To determine whether there is a CRM1-dependent NES in Gag, cells expressing GFP fusion proteins were treated with LMB (9, 11). As a control for LMB activity in QT6 cells, I B, a cellular protein containing a known NES (38), was expressed as a GFP fusion protein, and the minimal concentration of LMB (10 ng ml) that blocked its nuclear export was used in subsequent experiments (Fig. 3). The distributions of GFP, MA-GFP, and MA-p2-GFP were unaffected by LMB. Strikingly, both GFP fusion proteins that contain p10 showed marked sensitivity to LMB, with retention of the cgi doi pnas Scheifele et al.
4 Fig. 4. Effects of p10 deletions on subcellular localization. (A) Schematic diagram of the p10 sequence with hydrophobic residues that could function as an NES highlighted in red. Gag mutants with p10 deletions are shown below with heavy black lines indicating residues present in the mutant protein. (B) Confocal micrographs of cells expressing Gag-GFP p10 mutants reveal a putative NES. At 18 h after transfection, cells were fixed in paraformaldehyde. (Left) GFP epifluorescence. (Right) Propidium iodide-stained nuclei. MA-p2 p10-gfp and MA-p2-p10-CA-GFP proteins in the nucleus. Besides demonstrating the presence of a CRM1- dependent NES within the MA-p2-p10 region, this finding also confirmed that the nuclear-targeting activity within MA acts through an active process because it mediates nuclear entry of the 81 kda MA-p2-p10-CA-GFP protein. CRM1-dependent NESs are typically leucine-rich sequences, although other large hydrophobic amino acids may be substituted (39, 40). Two clusters of hydrophobic residues in p10 were potential candidates for an NES, as shown in Fig. 4A. To map the putative export signal, Gag-GFP mutants with deletions involving p10 were studied. Deletion of the N-terminal region of p10 did not alter the typical plasma membrane localization observed for Gag-GFP ( QM1, Fig. 4B). In contrast, an internal deletion that removes six hydrophobic residues was completely trapped in the nucleus, indicating that removal of this sequence interferes with nuclear export of Gag ( p10.52). A smaller deletion that eliminates L219 and W222 from the C-terminal portion of p10 also abrogates nuclear export ( p10.31). These results suggest that the NES is likely to be located in the second half of p10; although it remains possible that addition of the p10 sequence reveals a conformation-dependent NES within the MA or p2 sequences. Additionally, despite the existence of an LMB sensitive NES in this region of Gag, there might also be a cellular factor that mediates nuclear export of the protein. LMB Sensitivity of Gag and Mutants with Altered MA Sequences. We next examined the effect of LMB on localization of the fulllength Gag protein because previous constructs extended only through CA. We found that Gag was trapped in the nucleus after treatment with LMB, indicating that the trafficking pathway of Gag includes a nuclear phase (Fig. 5A). We reasoned that Gag mutants with strong membrane-binding domains might be targeted so rapidly to the plasma membrane that they fail to enter the nucleus. For example, Myr1E (Fig. 1) has the Src membranebinding domain extended from the N terminus of Gag and efficiently associates with the plasma membrane, even without the contribution of I domains (25). In the presence of LMB, Myr1E.Gag remained localized to the plasma membrane with no evidence of nuclear retention (Fig. 5A). This is intriguing because Myr1E is noninfectious because of a defect in genomic RNA packaging and dimerization (24). In Myr1E, the Src membrane-targeting activity might override the NLS NES activities normally found in Gag, bypassing the nuclear compartment. Thus, elimination of nuclear transport might explain the RNA packaging defect of Myr1E if nuclear localization of Gag is involved in genomic RNA incorporation. In support of this idea, we found that Myr1E, which is identical to Myr1E except for a G2A substitution that eliminates Src myristylation, demonstrated nuclear localization in the presence of LMB. Myr1E has normal infectivity and packages wild-type levels of RNA (24). A third mutant, Myr2.HB12, also remained in the nucleus Fig. 5. LMB sensitivity of mutant Gag proteins with defects in RNA packaging and dimerization. (A) Live cells expressing the indicated Gag-GFP proteins were visualized without (Left) and with (Right) LMB treatment, revealing that wildtype Gag and mutants Myr2.HB12 and Myr1E become trapped in the nucleus after treatment whereas Myr1E is insensitive to the effects of the drug. (B) Gag localization was analyzed in cells stably expressing wild-type or mutant RSV proviral genomes by indirect immunofluorescence in fixed cells either untreated or treated with LMB. Note that patterns of subcellular localization were the same as in A, indicating that GFP had not influenced the distribution of Gag. (C) Particle release in response to LMB treatment. The amount of immunoprecipitated Gag protein released from transfected cells during a 2.5-h-labeling period was divided by the total Gag protein detected in cells and medium to calculate budding percentage. Budding percentages from untreated (hatched bars) and treated (solid gray bars) cells were compared. Treated cells were incubated with LMB for 1 2 h followed by metabolic radiolabeling for 2.5 h in the presence of LMB. Each bar represents the average of three independent experiments with SDs indicated. Inhibition of particle assembly with LMB treatment was determined by a twosample t test; *, P and **, P MICROBIOLOGY Scheifele et al. PNAS March 19, 2002 vol. 99 no
5 with LMB treatment, and it packages nearly normal levels of genomic RNA (86% of wild-type levels, data not shown), although it is noninfectious. This mutant contains a myristic acid addition site at the Gag N terminus (designated Myr2) and a cluster of basic residues substituted for amino acids in MA (25). Interestingly, Myr2.HB12 packages only monomers of RNA (25), suggesting that packaging and dimerization of viral RNA are not directly linked, and dimer formation might occur in a postnuclear compartment. Thus, there is a correlation between the nuclear localization of Gag proteins in response to LMB treatment and genomic RNA incorporation. Effect of LMB Treatment on Gag Localization in Virus-Expressing Cells. The subcellular trafficking of Gag proteins and their derivatives in response to LMB treatment was tested in cells expressing wild-type or mutant proviruses to determine whether GFP might have influenced localization. In cells expressing the wild-type RSV genome, Gag proteins were seen in the cytoplasm with focal concentration at the plasma membrane when analyzed by indirect immunofluorescence (Fig. 5B). When cells were incubated with LMB, Gag proteins became sequestered in the nucleus. Thus, during infection Gag undergoes nuclear entry and export independently of GFP and localization is not altered by the expression of other viral gene products. In cells expressing the myr1e genome, Myr1E. Gag was seen at the plasma membrane both in untreated and treated cells, indicating that the nuclear compartment was bypassed. Particle Assembly in the Presence of LMB. Because LMB treatment of infected cells resulted in nuclear accumulation of Gag proteins, we expected that fewer Gag molecules would be available for transport to the plasma membrane, and virus budding might be diminished. Cells were transfected with wild-type or mutant Gag-GFP constructs, pretreated with LMB for 1 2 h, metabolically labeled for 2.5 h in the presence of LMB, and RSV Gag proteins were immunoprecipitated from cell lysates and medium. Budding directed by Gag-GFP was significantly inhibited in the presence of LMB (P ; mean of 49% reduction, 95% confidence interval ; Fig. 5C). Myr2.HB12.Gag- GFP, which showed LMB-dependent nuclear localization, also had a significant reduction in extracellular particle release (P 0.014; mean of 49% reduction, 95% confidence interval ). In contrast, budding of Myr1E.Gag-GFP was not significantly altered with LMB treatment (P 0.28). Analysis of budding for the p10 deletion mutants revealed that QM1.Gag-GFP, which had a subcellular localization similar to wild-type Gag, showed a trend toward reduction in budding because of LMB, but the effect was not statistically significant (P 0.097, 45% reduction; data not shown). Mutants with deletions affecting hydrophobic residues in the C-terminal portion of p10 released particles at a much lower rate that wild-type ( p10.31.gag-gfp, 69% reduced and p10.52.gag-gfp, 64% reduced, data not shown). Budding efficiency was not further diminished with LMB treatment (P 0.22 and 0.43, respectively). Taken together, these results indicate that inhibition of nuclear export, either pharmacologically or by mutagenesis of p10, limits the amount of Gag in the cytoplasm thereby interfering with the normal trafficking pathway required for particle assembly. Discussion In this report, we describe the discovery of an unexpected step in the RSV assembly pathway. Previously, Gag proteins were believed to be located exclusively within the cytoplasm after synthesis, with subsequent targeting directly to the plasma membrane. Now we demonstrate that RSV Gag has a nuclear phase that is mediated by two targeting signals: one for nuclear import and the other for CRM1-dependent nuclear export. Fig. 6. Model for the role of the nuclear localization of Gag during RSV replication. Hypothesis 1 (steps 1 4): The nuclear-targeting signal within MA delivers the cleaved MA protein to the nucleus during viral entry. Nuclear Gag proteins are exported via the CRM1-dependent NES so they are available in the cytoplasm to direct particle assembly. Hypothesis 2: (steps 5 7) The nucleartargeting signal carries the Gag protein into the nucleus where Gag might interact with unspliced viral RNA to begin the packaging process. Cytoplasmic relocalization of Gag occurs via the NES; Gag RNA complexes provide a nucleation point for Gag multimerization, and assembly intermediates are transported to the plasma membrane where budding occurs. Notably, hypotheses 1 and 2 are not mutually exclusive. Although the reason for the transient nuclear localization of the RSV Gag protein is unknown, we propose two main hypotheses (Fig. 6). First, we suggest that the NES in Gag counteracts the nuclear import signal in MA to keep Gag out of the nucleus. In this scenario, RSV MA plays a role early in infection to facilitate nuclear entry of the viral PIC. Although a requirement for facilitated nuclear targeting of the PIC has not been demonstrated for RSV, the report that RSV can infect growtharrested cells makes it possible that the nuclear-targeting sequence in MA might be involved (2). Because the nuclear import signal in MA also is present on the Gag polyprotein, Gag, too, would be transported into the nucleus but this would certainly be detrimental to virus assembly. To counteract nuclear import, the NES would return Gag to the cytoplasm for virus particle production. Besides import of the PIC, other roles could be imagined for the MA protein in the nucleus, including regulation of reverse transcription, integration, or viral transcription; however, mutants of MA that affect these processes have not yet been found. Whatever the function of MA might be in the nucleus, clearly its N-terminal helical domain is sufficient for nuclear import of the MA and Gag proteins. The MA nucleartargeting signal is large and complex compared to classical NLSs, and further experiments may reveal specific residues critical for interaction with nuclear import machinery or with a host factor that mediates the nuclear entry of MA and Gag. In the second model, we propose that Gag enters and exits the nucleus to fulfill a role during virion assembly (Fig. 6). After proviral integration, unspliced viral RNA transcripts are synthesized and exported out of the nucleus for translation into structural and enzymatic proteins. Export of unspliced viral transcripts must elude cellular mechanisms designed to retain intron-containing mrna in the nucleus. For RSV, it was shown that cytoplasmic accumulation of unspliced viral RNA is mediated by cis-acting DR elements in a CRM1-independent fashion, although neither Gag proteins nor the RSV genome was present in those experiments (41, 42). It is possible that the fraction of unspliced RNA exported for translation is distinct from the cgi doi pnas Scheifele et al.
6 fraction used for genome encapsidation. In our model, Gag proteins enter the nucleus where they might interact with unspliced viral RNA transcripts. The Gag-RNA nucleoprotein complex is then transported through the NPC via the CRM1 export pathway. In the cytoplasm, the Gag-RNA complex would be a nucleation point for the multimerization of additional Gag proteins that are then targeted to the plasma membrane. Importantly, these two models are not mutually exclusive, as MA might have a nuclear role early in infection and Gag might also enter the nucleus during assembly. It is also possible that Gag might have some other as yet unknown function in the nucleus unrelated to genomic RNA encapsidation. Although the nuclear entry of oncoviral Gag proteins is unique, other viruses have well-described replication strategies that take advantage of nuclear machinery to transport viral genomes, structural proteins and regulatory proteins (reviewed in ref. 1). For example, HIV-1 uses multiple signals to deliver the PIC to the nucleus of nondividing cells (1), and a CRM1- dependent NES in MA influences the nuclear export of fulllength viral RNAs (43). Alteration of the NES results in accumulation of HIV-1 Gag in the nucleus, although Gag itself was not shown to be sensitive to LMB treatment. Whether there are functional correlates between nuclear transport signals in RSV and HIV-1 Gag remains to be determined. As well, it is intriguing to consider possible parallels between the RSV Gag and influenza M1 proteins because both proteins are present in the nucleus and direct particle assembly at the plasma membrane. Influenza M1 associates with viral ribonucleoprotein complexes in the nucleus, and the viral genome is exported in a CRM1-dependent fashion (1, 44). In this report, we demonstrated that the MA domain is involved in import of the Gag polyprotein into the nucleus and that an NES in Gag mediates its cytoplasmic relocalization to facilitate virus assembly. We found that Gag mutants that package normal amounts of genomic RNA also have nuclear entry and export phases (Myr2.HB12 and Myr1E ) whereas a mutant with a reduced level of viral RNA incorporation does not enter the nucleus (Myr1E). The observation that RSV replication includes transient nuclear localization of the Gag protein raises intriguing questions about early Gag RNA interactions and the intracellular trafficking pathways taken by assembling virions. We thank Becky Craven, Tina Cairns, Carol Wilson, and John Wills for reagents and for sharing unpublished results, and John Wills for thoughtful review of the manuscript. Tom Hope generously provided plasmids and helpful advice. We thank Dr. Vernon Chincilli (Pennsylvania State University College of Medicine) for help with statistical analysis. This work was supported by a grant from the National Institutes of Health to L.J.P (R01 CA76534) and a Graduate Student Fellowship from the National Science Foundation to L.Z.S. 1. Whittaker, G. R. & Helenius, A. (1998) Virology 246, Hatzoiioannou, T. G. & Goff, S. P. (2001) J. Virol. 75, Cullen, B. R. (2000) Mol. Cell Biol. 20, Ogert, R. A. & Beemon, K. L. (1998) J. Virol. 72, Mattaj, I. W. & Englmeier, L. (1998) Annu. Rev. Biochem. 67, Christophe, D., Christophe-Hobertus, C. & Pichon, B. (2000) Cell. Signalling 12, Fischer, U., Huber, J., Boelens, W. C., Mattaj, I. M. & Luhrmann, R. (1995) Cell 82, Wen, W., Meinkoth, J. L., Tsien, R. Y. & Taylor, S. S. (1995) Cell 82, Fukuda, M., Asano, S., Nakamura, T., Adachi, M., Yoshida, M., Yanagida, M. & Nishida, E. (1997) Nature (London) 390, Fornerod, M., Ohno, M., Yoshida, M. & Mattaj, I. W. (1997) Cell 90, Kudo, N., Matsumori, N., Taoka, H., Fujiwara, D., Schreiner, E. P., Wolff, B., Yoshida, M. & Horinouchi, S. (1999) Proc. Natl. Acad. Sci. USA 96, Weldon, R. A. & Wills, J. W. (1993) J. Virol. 67, Bowzard, J. B., Bennett, R. P., Krishna, N. K., Ernst, S. M., Rein, A. & Wills, J. W. (1998) J. Virol. 72, Swanstrom, R. & Wills, J. W. (1997) in Retroviruses, eds. Coffin, J. M., Hughes, S. H. & Varmus, H. E. (Cold Spring Harbor Lab. Press, Plainview, NY), pp Campbell, S. & Vogt, V. M. (1995) J. Virol. 69, Muriaux, D., Mirro, J., Harvin, D. & Rein, A. (2001) Proc. Natl. Acad. Sci. USA 98, Shields, A., Witte, W. N., Rothenberg, E. & Baltimore, D. (1978) Cell 14, Vogt, V. M. (1997) in Retroviruses, eds. Coffin, J. M., Hugher, S. H. & Varmus, H. E. (Cold Spring Harbor Lab. Press, Plainview, NY), pp Nelle, T. D. & Wills, J. W. (1996) J. Virol. 70, Verderame, M. F., Nelle, T. D. & Wills, J. W. (1996) J. Virol. 70, Wills, J. W., Cameron, C. E., Wilson, C. B., Xiang, Y., Bennett, R. P. & Leis, J. (1994) J. Virol. 68, Parent, L. J., Bennett, R. P., Craven, R. C., Nelle, T. D., Krishna, N. K., Bowzard, J. B., Wilson, C. B., Puffer, B. A., Montelaro, R. C. & Wills, J. W. (1995) J. Virol. 69, Craven, R. C., Leure-duPree, A. E., Erdie, C. R., Wilson, C. B. & Wills, J. W. (1993) J. Virol. 67, Parent, L. J., Cairns, T. M., Albert, J. A., Wilson, C. B., Wills, J. W. & Craven, R. C. (2000) J. Virol. 74, Garbitt, R. A., Albert, J. A., Kessler, M. D. & Parent, L. J. (2001) J. Virol. 75, Callahan, E. M. & Wills, J. W. (2000) J. Virol. 74, Craven, R. C., Leure-duPree, A. E., Weldon, R. A., Jr. & Wills, J. W. (1995) J. Virol. 69, Moscovici, C., Moscovici, M. G., Jimenez, H., Lai, M. M. C., Hayman, M. J. & Vogt, P. K. (1977) Cell 11, Krishna, N. K., Campbell, S., Vogt, V. M. & Wills, J. W. (1998) J. Virol. 72, Parent, L. J., Wilson, C. B., Resh, M. D. & Wills, J. W. (1996) J. Virol. 70, Weldon, R. A., Erdie, C. R., Oliver, M. G. & Wills, J. W. (1990) J. Virol. 64, Hermida-Matsumoto, L. & Resh, M. D. (1999) J. Virol. 73, Paillart, J. C. & Gottlinger, H. G. (1999) J. Virol. 73, Spearman, P., Horton, R., Ratner, L. & Kuli-Zade, I. (1997) J. Virol. 71, Hofmann, K., Bucher, P., Falquet, L. & Bairoch, A. (1999) Nucleic Acids Res. 27, Cokol, M., Nair, R. & Rost, B. (2000) EMBO Rep. 1, McDonnell, J. M., Fushman, D., Cahill, S. M., Zhou, W., Wolven, A., Wilson, C. B., Nelle, T. D., Resh, M. D., Wills, J. & Cowburn, D. (1998) J. Mol. Biol. 279, Johnson, C., Van Antwerp, D. & Hope, T. J. (1999) EMBO J. 18, Hope, T. J. (1999) Arch. Biochem. Biophys. 365, Gorlich, D. & Kutay, U. (1999) Annu. Rev. Cell Dev. Biol. 15, Ogert, R. A., Lee, L. H. & Beemon, K. L. (1996) J. Virol. 70, Paca, R. E., Ogert, R. A., Hibbert, C. S., Izaurralde, E. & Beemon, K. L. (2000) J. Virol. 74, Dupont, S., Sharova, N., DeHoratius, C., Virbasius, C.-M. A., Zhu, X., Bukrinskaya, A. G., Stevenson, M. & Green, M. R. (1999) Nature (London) 402, Ma, K., Roy, A. M. M. & Whittaker, G. R. (2001) Virology 282, Wills, J. W., Craven, R. C., Weldon, R. A., Jr., Nelle, T. D. & Erdie, C. R. (1991) J. Virol. 65, MICROBIOLOGY Scheifele et al. PNAS March 19, 2002 vol. 99 no
MOLECULAR CELL BIOLOGY
 1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
ACCEPTED. Overlapping roles of the Rous sarcoma virus Gag p10. domain in nuclear export and virion core morphology
 JVI Accepts, published online ahead of print on 1 July 00 J. Virol. doi:./jvi.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved. 1 1 1 1 1
JVI Accepts, published online ahead of print on 1 July 00 J. Virol. doi:./jvi.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved. 1 1 1 1 1
Fayth K. Yoshimura, Ph.D. September 7, of 7 HIV - BASIC PROPERTIES
 1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Regulation of a nuclear export signal by an adjacent inhibitory sequence: The effector domain of the influenza virus NS1 protein
 Proc. Natl. Acad. Sci. USA Vol. 95, pp. 4864 4869, April 1998 Biochemistry Regulation of a nuclear export signal by an adjacent inhibitory sequence: The effector domain of the influenza virus NS1 protein
Proc. Natl. Acad. Sci. USA Vol. 95, pp. 4864 4869, April 1998 Biochemistry Regulation of a nuclear export signal by an adjacent inhibitory sequence: The effector domain of the influenza virus NS1 protein
Last time we talked about the few steps in viral replication cycle and the un-coating stage:
 Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Bio 401 Sp2014. Multiple Choice (Circle the letter corresponding to the correct answer)
 MIDTERM EXAM KEY (60 pts) You should have 5 pages. Please put your name on every page. You will have 70 minutes to complete the exam. You may begin working as soon as you receive the exam. NOTE: the RED
MIDTERM EXAM KEY (60 pts) You should have 5 pages. Please put your name on every page. You will have 70 minutes to complete the exam. You may begin working as soon as you receive the exam. NOTE: the RED
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
Supplementary information. MARCH8 inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins
 Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Dr. Ahmed K. Ali Attachment and entry of viruses into cells
 Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
Retroviruses. ---The name retrovirus comes from the enzyme, reverse transcriptase.
 Retroviruses ---The name retrovirus comes from the enzyme, reverse transcriptase. ---Reverse transcriptase (RT) converts the RNA genome present in the virus particle into DNA. ---RT discovered in 1970.
Retroviruses ---The name retrovirus comes from the enzyme, reverse transcriptase. ---Reverse transcriptase (RT) converts the RNA genome present in the virus particle into DNA. ---RT discovered in 1970.
JOURNAL OF VIROLOGY, July 1999, p Vol. 73, No. 7. Copyright 1999, American Society for Microbiology. All Rights Reserved.
 JOURNAL OF VIROLOGY, July 1999, p. 5654 5662 Vol. 73, No. 7 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Formation of Virus Assembly Intermediate Complexes
JOURNAL OF VIROLOGY, July 1999, p. 5654 5662 Vol. 73, No. 7 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Formation of Virus Assembly Intermediate Complexes
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Fayth K. Yoshimura, Ph.D. September 7, of 7 RETROVIRUSES. 2. HTLV-II causes hairy T-cell leukemia
 1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
In Vivo Interference of Rous Sarcoma Virus Budding by cis Expression of a WW Domain
 JOURNAL OF VIROLOGY, Mar. 2002, p. 2789 2795 Vol. 76, No. 6 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.6.2789 2795.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. In Vivo Interference
JOURNAL OF VIROLOGY, Mar. 2002, p. 2789 2795 Vol. 76, No. 6 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.6.2789 2795.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. In Vivo Interference
ACCEPTED. Intermolecular Interactions between Retroviral Gag. Proteins in the Nucleus. [Running title: Intranuclear Gag-Gag interactions]
![ACCEPTED. Intermolecular Interactions between Retroviral Gag. Proteins in the Nucleus. [Running title: Intranuclear Gag-Gag interactions] ACCEPTED. Intermolecular Interactions between Retroviral Gag. Proteins in the Nucleus. [Running title: Intranuclear Gag-Gag interactions]](/thumbs/91/105499942.jpg) JVI Accepts, published online ahead of print on 1 October 00 J. Virol. doi:./jvi.00-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved. 1 1 1
JVI Accepts, published online ahead of print on 1 October 00 J. Virol. doi:./jvi.00-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved. 1 1 1
Subcellular Localization of Feline Immunodeficiency Virus Integrase and Mapping of Its Karyophilic Determinant
 JOURNAL OF VIROLOGY, Apr. 2003, p. 4516 4527 Vol. 77, No. 8 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.8.4516 4527.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Subcellular
JOURNAL OF VIROLOGY, Apr. 2003, p. 4516 4527 Vol. 77, No. 8 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.8.4516 4527.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Subcellular
VIROLOGY. Engineering Viral Genomes: Retrovirus Vectors
 VIROLOGY Engineering Viral Genomes: Retrovirus Vectors Viral vectors Retrovirus replicative cycle Most mammalian retroviruses use trna PRO, trna Lys3, trna Lys1,2 The partially unfolded trna is annealed
VIROLOGY Engineering Viral Genomes: Retrovirus Vectors Viral vectors Retrovirus replicative cycle Most mammalian retroviruses use trna PRO, trna Lys3, trna Lys1,2 The partially unfolded trna is annealed
Zool 3200: Cell Biology Exam 4 Part II 2/3/15
 Name:Key Trask Zool 3200: Cell Biology Exam 4 Part II 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
Name:Key Trask Zool 3200: Cell Biology Exam 4 Part II 2/3/15 Answer each of the following questions in the space provided, explaining your answers when asked to do so; circle the correct answer or answers
7.012 Problem Set 6 Solutions
 Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
Section 6. Junaid Malek, M.D.
 Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Translation. Host Cell Shutoff 1) Initiation of eukaryotic translation involves many initiation factors
 Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
SUPPLEMENTARY INFORMATION
 Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Julianne Edwards. Retroviruses. Spring 2010
 Retroviruses Spring 2010 A retrovirus can simply be referred to as an infectious particle which replicates backwards even though there are many different types of retroviruses. More specifically, a retrovirus
Retroviruses Spring 2010 A retrovirus can simply be referred to as an infectious particle which replicates backwards even though there are many different types of retroviruses. More specifically, a retrovirus
Human Immunodeficiency Virus
 Human Immunodeficiency Virus Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Viruses and hosts Lentivirus from Latin lentis (slow), for slow progression of disease
Human Immunodeficiency Virus Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Viruses and hosts Lentivirus from Latin lentis (slow), for slow progression of disease
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
LESSON 4.6 WORKBOOK. Designing an antiviral drug The challenge of HIV
 LESSON 4.6 WORKBOOK Designing an antiviral drug The challenge of HIV In the last two lessons we discussed the how the viral life cycle causes host cell damage. But is there anything we can do to prevent
LESSON 4.6 WORKBOOK Designing an antiviral drug The challenge of HIV In the last two lessons we discussed the how the viral life cycle causes host cell damage. But is there anything we can do to prevent
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Packaging and Abnormal Particle Morphology
 JOURNAL OF VIROLOGY, OCt. 1990, p. 5230-5234 0022-538X/90/105230-05$02.00/0 Copyright 1990, American Society for Microbiology Vol. 64, No. 10 A Mutant of Human Immunodeficiency Virus with Reduced RNA Packaging
JOURNAL OF VIROLOGY, OCt. 1990, p. 5230-5234 0022-538X/90/105230-05$02.00/0 Copyright 1990, American Society for Microbiology Vol. 64, No. 10 A Mutant of Human Immunodeficiency Virus with Reduced RNA Packaging
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Role of the C terminus Gag protein in human immunodefieieney virus type 1 virion assembly and maturation
 Journal of General Virology (1995), 76, 3171-3179. Printed in Great Britain 3171 Role of the C terminus Gag protein in human immunodefieieney virus type 1 virion assembly and maturation X.-F. Yu, ~ Z.
Journal of General Virology (1995), 76, 3171-3179. Printed in Great Britain 3171 Role of the C terminus Gag protein in human immunodefieieney virus type 1 virion assembly and maturation X.-F. Yu, ~ Z.
LESSON 4.4 WORKBOOK. How viruses make us sick: Viral Replication
 DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Overview of virus life cycle
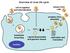 Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Coronaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Supplementary Material
 Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
Chapter 6- An Introduction to Viruses*
 Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
Running Head: AN UNDERSTANDING OF HIV- 1, SYMPTOMS, AND TREATMENTS. An Understanding of HIV- 1, Symptoms, and Treatments.
 Running Head: AN UNDERSTANDING OF HIV- 1, SYMPTOMS, AND TREATMENTS An Understanding of HIV- 1, Symptoms, and Treatments Benjamin Mills Abstract HIV- 1 is a virus that has had major impacts worldwide. Numerous
Running Head: AN UNDERSTANDING OF HIV- 1, SYMPTOMS, AND TREATMENTS An Understanding of HIV- 1, Symptoms, and Treatments Benjamin Mills Abstract HIV- 1 is a virus that has had major impacts worldwide. Numerous
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Supplementary figure legends
 Supplementary figure legends Supplementary Figure 1. Exposure of CRT occurs independently from the apoptosisassociated loss of the mitochondrial membrane potential (MMP). (A) HeLa cells treated with MTX
Supplementary figure legends Supplementary Figure 1. Exposure of CRT occurs independently from the apoptosisassociated loss of the mitochondrial membrane potential (MMP). (A) HeLa cells treated with MTX
Gammaretroviral pol sequences act in cis to direct polysome loading and NXF1/NXT-dependent protein production by gag-encoded RNA
 University of Massachusetts Medical School escholarship@umms Open Access Articles Open Access Publications by UMMS Authors 9-12-2014 Gammaretroviral pol sequences act in cis to direct polysome loading
University of Massachusetts Medical School escholarship@umms Open Access Articles Open Access Publications by UMMS Authors 9-12-2014 Gammaretroviral pol sequences act in cis to direct polysome loading
Structural vs. nonstructural proteins
 Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Herpesviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Herpesviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped icosahedral capsid (T=16), diameter 125 nm Diameter of enveloped virion 200 nm Capsid
Herpesviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped icosahedral capsid (T=16), diameter 125 nm Diameter of enveloped virion 200 nm Capsid
Name Section Problem Set 6
 Name Section 7.012 Problem Set 6 Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed of lipids
Name Section 7.012 Problem Set 6 Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed of lipids
Nuclear Pore Tutorial
 Michael Brenner, Harvard January 7, 2010 Nuclear Pore Tutorial Outline (1) What is the nuclear pore/what does it look like? What are the proteins that make it up? (2) How does the nuclear pore work? The
Michael Brenner, Harvard January 7, 2010 Nuclear Pore Tutorial Outline (1) What is the nuclear pore/what does it look like? What are the proteins that make it up? (2) How does the nuclear pore work? The
Problem-solving Test: The Mechanism of Protein Synthesis
 Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
SUPPLEMENTARY INFORMATION
 sirna pool: Control Tetherin -HA-GFP HA-Tetherin -Tubulin Supplementary Figure S1. Knockdown of HA-tagged tetherin expression by tetherin specific sirnas. HeLa cells were cotransfected with plasmids expressing
sirna pool: Control Tetherin -HA-GFP HA-Tetherin -Tubulin Supplementary Figure S1. Knockdown of HA-tagged tetherin expression by tetherin specific sirnas. HeLa cells were cotransfected with plasmids expressing
Protein Trafficking in the Secretory and Endocytic Pathways
 Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
number Done by Corrected by Doctor Ashraf
 number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
Interaction of the Influenza Virus Nucleoprotein with the Cellular CRM1-Mediated Nuclear Export Pathway
 JOURNAL OF VIROLOGY, Jan. 2001, p. 408 419 Vol. 75, No. 1 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.1.408 419.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Interaction of
JOURNAL OF VIROLOGY, Jan. 2001, p. 408 419 Vol. 75, No. 1 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.1.408 419.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Interaction of
Problem Set 5 KEY
 2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
2006 7.012 Problem Set 5 KEY ** Due before 5 PM on THURSDAY, November 9, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You are studying the development
Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication
 Manuscript EMBO-2010-74756 Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication Tsai-Ling Liao, Chung-Yi Wu, Wen-Chi Su, King-Song Jeng and Michael Lai Corresponding
Manuscript EMBO-2010-74756 Ubiquitination and deubiquitination of NP protein regulates influenza A virus RNA replication Tsai-Ling Liao, Chung-Yi Wu, Wen-Chi Su, King-Song Jeng and Michael Lai Corresponding
MCB130 Midterm. GSI s Name:
 1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
Influenza viruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Virus Entry/Uncoating
 Virus Entry/Uncoating Delivery of genome to inside of a cell Genome must be available for first step of replication The Problem--barriers to infection Virion Barriers: Non-enveloped viruses capsid Enveloped
Virus Entry/Uncoating Delivery of genome to inside of a cell Genome must be available for first step of replication The Problem--barriers to infection Virion Barriers: Non-enveloped viruses capsid Enveloped
Chapter 3. Expression of α5-megfp in Mouse Cortical Neurons. on the β subunit. Signal sequences in the M3-M4 loop of β nachrs bind protein factors to
 22 Chapter 3 Expression of α5-megfp in Mouse Cortical Neurons Subcellular localization of the neuronal nachr subtypes α4β2 and α4β4 depends on the β subunit. Signal sequences in the M3-M4 loop of β nachrs
22 Chapter 3 Expression of α5-megfp in Mouse Cortical Neurons Subcellular localization of the neuronal nachr subtypes α4β2 and α4β4 depends on the β subunit. Signal sequences in the M3-M4 loop of β nachrs
Mutations of the Human Immunodeficiency Virus Type 1 p6 Gag Domain Result in Reduced Retention of Pol Proteins during Virus Assembly
 JOURNAL OF VIROLOGY, Apr. 1998, p. 3412 3417 Vol. 72, No. 4 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology Mutations of the Human Immunodeficiency Virus Type 1 p6 Gag Domain Result
JOURNAL OF VIROLOGY, Apr. 1998, p. 3412 3417 Vol. 72, No. 4 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology Mutations of the Human Immunodeficiency Virus Type 1 p6 Gag Domain Result
Fine Mapping of a cis-acting Sequence Element in Yellow Fever Virus RNA That Is Required for RNA Replication and Cyclization
 JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
AND JEREMY LUBAN 1,2 *
 JOURNAL OF VIROLOGY, Oct. 1999, p. 8527 8540 Vol. 73, No. 10 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency Virus Type 1 Gag Polyprotein
JOURNAL OF VIROLOGY, Oct. 1999, p. 8527 8540 Vol. 73, No. 10 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency Virus Type 1 Gag Polyprotein
Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection
 Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection Melissa Mihelidakis May 6, 2004 7.340 Research Proposal Introduction Apoptosis, or programmed cell
Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection Melissa Mihelidakis May 6, 2004 7.340 Research Proposal Introduction Apoptosis, or programmed cell
Nuclear Import and Export of Venezuelan Equine Encephalitis Virus Nonstructural Protein 2
 JOURNAL OF VIROLOGY, Oct. 2007, p. 10268 10279 Vol. 81, No. 19 0022-538X/07/$08.00 0 doi:10.1128/jvi.00371-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Nuclear Import and
JOURNAL OF VIROLOGY, Oct. 2007, p. 10268 10279 Vol. 81, No. 19 0022-538X/07/$08.00 0 doi:10.1128/jvi.00371-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Nuclear Import and
RECAP FROM MONDAY AND TUESDAY LECTURES
 RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
Charged Amino Acid Residues of Human Immunodeficiency Virus Type 1 Nucleocapsid p7 Protein Involved in RNA Packaging and Infectivity
 JOURNAL OF VIROLOGY, Oct. 1996, p. 6607 6616 Vol. 70, No. 10 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Charged Amino Acid Residues of Human Immunodeficiency Virus Type 1 Nucleocapsid
JOURNAL OF VIROLOGY, Oct. 1996, p. 6607 6616 Vol. 70, No. 10 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Charged Amino Acid Residues of Human Immunodeficiency Virus Type 1 Nucleocapsid
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Edens and Levy, http://www.jcb.org/cgi/content/full/jcb.201406004/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Nuclear shrinking does not depend on the cytoskeleton
Supplemental material Edens and Levy, http://www.jcb.org/cgi/content/full/jcb.201406004/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Nuclear shrinking does not depend on the cytoskeleton
Reverse Genetics of RNA Viruses
 Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Peptide hydrolysis uncatalyzed half-life = ~450 years HIV protease-catalyzed half-life = ~3 seconds
 Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Received 30 January 2002/Accepted 18 December 2002
 JOURNAL OF VIROLOGY, Mar. 2003, p. 3634 3646 Vol. 77, No. 6 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.6.3634 3646.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Rational Site-Directed
JOURNAL OF VIROLOGY, Mar. 2003, p. 3634 3646 Vol. 77, No. 6 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.6.3634 3646.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Rational Site-Directed
Viral structure م.م رنا مشعل
 Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Role of ESCRT-I in Retroviral Budding
 JOURNAL OF VIROLOGY, Apr. 2003, p. 4794 4804 Vol. 77, No. 8 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.8.4794 4804.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Role of ESCRT-I
JOURNAL OF VIROLOGY, Apr. 2003, p. 4794 4804 Vol. 77, No. 8 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.8.4794 4804.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Role of ESCRT-I
III. What are the requirements for taking and passing this course?
 1 Molecular Virology Lecture # 1: Course Introduction I. Instructor and Background Dr. Richard Kuhn rjkuhn@bragg.bio.purdue.edu B-129 Lilly Hall 494-1164 Office Hours - Wednesday 10:30-11:30 II. Objective:
1 Molecular Virology Lecture # 1: Course Introduction I. Instructor and Background Dr. Richard Kuhn rjkuhn@bragg.bio.purdue.edu B-129 Lilly Hall 494-1164 Office Hours - Wednesday 10:30-11:30 II. Objective:
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
There are approximately 30,000 proteasomes in a typical human cell Each proteasome is approximately 700 kda in size The proteasome is made up of 3
 Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
VIRUSES. 1. Describe the structure of a virus by completing the following chart.
 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
/07/$15.00/0 Molecular Endocrinology 21(12): Printed in U.S.A.
 0888-8809/07/$15.00/0 Molecular Endocrinology 21(12):3087 3099 Printed in U.S.A. Copyright 2007 by The Endocrine Society doi: 10.1210/me.2006-0476 The Glucose Transporter 4 FQQI Motif Is Necessary for
0888-8809/07/$15.00/0 Molecular Endocrinology 21(12):3087 3099 Printed in U.S.A. Copyright 2007 by The Endocrine Society doi: 10.1210/me.2006-0476 The Glucose Transporter 4 FQQI Motif Is Necessary for
Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles
 Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles Amin Feizpour Reinhard Lab Department of Chemistry and the Photonics Center, Boston University, Boston, MA May 2014
Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles Amin Feizpour Reinhard Lab Department of Chemistry and the Photonics Center, Boston University, Boston, MA May 2014
MCB Chapter 11. Topic E. Splicing mechanism Nuclear Transport Alternative control modes. Reading :
 MCB Chapter 11 Topic E Splicing mechanism Nuclear Transport Alternative control modes Reading : 419-449 Topic E Michal Linial 14 Jan 2004 1 Self-splicing group I introns were the first examples of catalytic
MCB Chapter 11 Topic E Splicing mechanism Nuclear Transport Alternative control modes Reading : 419-449 Topic E Michal Linial 14 Jan 2004 1 Self-splicing group I introns were the first examples of catalytic
7.06 Cell Biology EXAM #3 April 24, 2003
 7.06 Spring 2003 Exam 3 Name 1 of 8 7.06 Cell Biology EXAM #3 April 24, 2003 This is an open book exam, and you are allowed access to books and notes. Please write your answers to the questions in the
7.06 Spring 2003 Exam 3 Name 1 of 8 7.06 Cell Biology EXAM #3 April 24, 2003 This is an open book exam, and you are allowed access to books and notes. Please write your answers to the questions in the
Problem Set #5 4/3/ Spring 02
 Question 1 Chloroplasts contain six compartments outer membrane, intermembrane space, inner membrane, stroma, thylakoid membrane, and thylakoid lumen each of which is populated by specific sets of proteins.
Question 1 Chloroplasts contain six compartments outer membrane, intermembrane space, inner membrane, stroma, thylakoid membrane, and thylakoid lumen each of which is populated by specific sets of proteins.
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Identification of Mutation(s) in. Associated with Neutralization Resistance. Miah Blomquist
 Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
Host Cell Nuclear Proteins Are Recruited to Cytoplasmic Vaccinia Virus Replication Complexes
 JOURNAL OF VIROLOGY, Oct. 2005, p. 12852 12860 Vol. 79, No. 20 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.20.12852 12860.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Host
JOURNAL OF VIROLOGY, Oct. 2005, p. 12852 12860 Vol. 79, No. 20 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.20.12852 12860.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Host
Reverse transcription and integration
 Reverse transcription and integration Lecture 9 Biology 3310/4310 Virology Spring 2018 One can t believe impossible things, said Alice. I dare say you haven t had much practice, said the Queen. Why, sometimes
Reverse transcription and integration Lecture 9 Biology 3310/4310 Virology Spring 2018 One can t believe impossible things, said Alice. I dare say you haven t had much practice, said the Queen. Why, sometimes
TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer
 Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplementary Information TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer Yabing Mu, Reshma Sundar, Noopur Thakur, Maria Ekman, Shyam Kumar Gudey, Mariya
Supplemental Information
 Cell Host & Microbe, Volume 14 Supplemental Information HIV-1 Induces the Formation of Stable Microtubules to Enhance Early Infection Yosef Sabo, Derek Walsh, Denis S. Barry, Sedef Tinaztepe, Kenia de
Cell Host & Microbe, Volume 14 Supplemental Information HIV-1 Induces the Formation of Stable Microtubules to Enhance Early Infection Yosef Sabo, Derek Walsh, Denis S. Barry, Sedef Tinaztepe, Kenia de
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
Cell Quality Control. Peter Takizawa Department of Cell Biology
 Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Sestrin2 and BNIP3 (Bcl-2/adenovirus E1B 19kDa-interacting. protein3) regulate autophagy and mitophagy in renal tubular cells in. acute kidney injury
 Sestrin2 and BNIP3 (Bcl-2/adenovirus E1B 19kDa-interacting protein3) regulate autophagy and mitophagy in renal tubular cells in acute kidney injury by Masayuki Ishihara 1, Madoka Urushido 2, Kazu Hamada
Sestrin2 and BNIP3 (Bcl-2/adenovirus E1B 19kDa-interacting protein3) regulate autophagy and mitophagy in renal tubular cells in acute kidney injury by Masayuki Ishihara 1, Madoka Urushido 2, Kazu Hamada
MedChem 401~ Retroviridae. Retroviridae
 MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
SD-1 SD-1: Cathepsin B levels in TNF treated hch
 SD-1 SD-1: Cathepsin B levels in TNF treated hch. A. RNA and B. protein extracts from TNF treated and untreated human chondrocytes (hch) were analyzed via qpcr (left) and immunoblot analyses (right) for
SD-1 SD-1: Cathepsin B levels in TNF treated hch. A. RNA and B. protein extracts from TNF treated and untreated human chondrocytes (hch) were analyzed via qpcr (left) and immunoblot analyses (right) for
Evidence for specific nucleocytoplasmic transport pathways used by leucine-rich nuclear export signals
 Proc. Natl. Acad. Sci. USA Vol. 96, pp. 6229 6234, May 1999 Cell Biology Evidence for specific nucleocytoplasmic transport pathways used by leucine-rich nuclear export signals CLAUDIA ELFGANG*, OLAF ROSORIUS,
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 6229 6234, May 1999 Cell Biology Evidence for specific nucleocytoplasmic transport pathways used by leucine-rich nuclear export signals CLAUDIA ELFGANG*, OLAF ROSORIUS,
