ProPred1: prediction of promiscuous MHC Class-I binding sites. Harpreet Singh and G.P.S. Raghava
|
|
|
- Britton Charles
- 6 years ago
- Views:
Transcription
1 BIOINFORMATICS Vol. 19 no , pages DOI: /bioinformatics/btg108 ProPred1: prediction of promiscuous MHC Class-I binding sites Harpreet Singh and G.P.S. Raghava Institute of Microbial technology, Chandigarh , India Received on July 18, 2002; revised on July 18, 2002; accepted on January 3, 2003 ABSTRACT Summary: ProPred1 is an on-line web tool for the prediction of peptide binding to MHC class-i alleles. This is a matrix-based method that allows the prediction of MHC binding sites in an antigenic sequence for 47 MHC class-i alleles. The server represents MHC binding regions within an antigenic sequence in user-friendly formats. These formats assist user in the identification of promiscuous MHC binders in an antigen sequence that can bind to large number of alleles. ProPred1 also allows the prediction of the standard proteasome and immunoproteasome cleavage sites in an antigenic sequence. This server allows identification of MHC binders, who have the cleavage site at the C terminus. The simultaneous prediction of MHC binders and proteasome cleavage sites in an antigenic sequence leads to the identification of potential T-cell epitopes. Availability: Server is available at in/raghava/propred1/. Mirror site of this server is available at Supplementary information: Matrices and document on server are available at propred1/page2.html Contact: raghava@imtech.res.in INTRODUCTION The altered proteins of self or pathogenic origin are fragmented to small peptides by the proteasomes. Some of these peptides bind to the class-i major histocompatibility complex (MHC) molecules. The resulting MHC peptide complex is recognized by T-cells. Identification of peptides that will be processed and presented with MHC molecules is crucial for the development of peptide-based vaccines. The experimental identification of these peptides is very time consuming and costly, therefore efforts are being made to use computational methods in prediction of immunogenic regions in an antigenic sequence. The computer-based predictions can effectively reduce the labor involved in wet lab experiments in the identification To whom the correspondence should be addressed at Bioinformatics Centre, Institute of Microbial Technology, Sector 39A, Chandigarh, India. of immunogenic regions (Hagmann, 2000). Another interest of the immunologists is the identification of cross-reactive or promiscuous antigenic regions in sequence; these regions can bind to many MHC alleles (Sturniolo et al., 1999). In this direction, we have previously developed ProPred (Singh and Raghava, 2001) for predicting the promiscuous MHC class-ii binding peptides. A number of methods have already been developed for the prediction of binding site of various MHC class-i alleles e.g. (i) BIMAS for 41 alleles (Parker et al., 1994). bind/); and (ii) SYFPEITHI for 19 MHC alleles (Rammensee et al., 1999), MHCServer.dll/EpPredict.htm). The MHC class-i molecules recognize the fragments of the antigenic protein after its processing by proteasome (Rock and Goldberg, 1999). Thus, proteasome plays a vital role in identification of potential T-cell epitopes (Niedermann et al., 1999). It has been shown experimentally that proteasome generates the C terminus of MHC class-i binding peptide (Craiu et al., 1997; Stoltze et al., 1998). Recently, numerous methods have been developed for prediction of proteasome cleavage sites (Holzhutter et al., 1999; Kuttler et al., 2000; Nussbaum et al., 2001) and attempt have been made to combine those with the prediction of MHC binding peptide like MAPPP ( In this paper we have described a matrix based web server ProPred1, developed for the prediction of MHC class-i binding peptides. The matrices used in ProPred1 have been obtained from BIMAS server and from the literature. The server gives the output of predicted MHC binder for selected MHC alleles in graphical or text format. These display formats help the user in easy detection of the promiscuous MHC binding regions in their query sequence. We also have made an attempt for the simultaneous prediction of MHC binders and proteasome cleavage sites in a protein sequence. This server has implemented the matrices described by (Toes et al., 2001) for the identification of proteasome (standard/constitutive proteasome and immunoproteasome) cleavage sites in an antigenic sequence. Bioinformatics 19(8) c Oxford University Press 2003; all rights reserved. 1009
2 H.Singh and G.P.S.Raghava Table 1. The references of the quantitative matrices used in ProPred1 server Name of MHC allele Reference Comment HLA-A2.1 (Ruppert et al., 1993) Addition matrix HLA-B*0702 (Sidney et al., 1996) Addition matrix HLA-B51 (Sidney et al., 1996) Addition matrix HLA-B*5301 (Sidney et al., 1996) Addition matrix HLA-B*5401 (Sidney et al., 1996) Addition matrix All other MHC alleles Unpublished Multiplication matrices (BIMAS Server) Recently, (Kessler et al., 2001; Ayyoub et al., 2002) have demonstrated that MHC binders having proteasome cleavage site at their C terminus have high potency to become T-cell epitopes. These observations were implemented in ProPred1 in order to identify the potential T-cell epitopes. The ProPred1 allows identification of the predicted MHC binders who have predicted proteasome site at C terminus. In brief, the server assists users in identification of promiscuous MHC binders and potential T-cell epitopes in an antigenic sequence. MATERIALS AND METHODS Source of weight matrices The most of the quantitative matrices have been obtained from BIMAS server and a few matrices were obtained from literature (See Table 1). The selection of matrices was not based on performance rather easily availability in the literature. The matrices obtained from BIMAS server are multiplication matrices, where the score is calculated by multiplying scores of each position. For example, score of peptide PACDPGRAA can be calculated by following equation. Score = P(1) A(2) C(3) D(4) P(5) G(6) R(7) A(8) A(9) (1) Where P(1) is score of P at position 1. The matrices obtained from the literature are Addition Matrices, where score is calculated by summing the scores of each position. For example, score for above peptide PACDPGRAA is calculated as follows: Score = P(1) + A(2) + C(3) + D(4) + P(5) + G(6) + R(7) + A(8) + A(9). (2) Threshold score The selection of cut off threshold is crucial in matrix-based methods, because the stringency of prediction varies with the threshold. The threshold also provides the confidence to the user in their prediction. Thus, it is important to calculate the threshold score in advance for each allele so that binders and non-binders can be distinguished with confidence. In order to calculate the threshold there is a need of sufficient data of MHC binders and nonbinders. Unfortunately, due to lack of data (particularly MHC non-binders), it is practically impossible to compute the threshold score for most of the alleles. To overcome the above problem, we adopted the following uniform procedure for the calculation of threshold score for each allele. (i) All proteins were obtained from SWISSPROT databases for creating the overlapping peptides of length nine. For example, a protein of length n will have (n + 1 9) overlapping peptides. (ii) The score of all natural 9mer peptides have been calculated using weight matrix of that allele. These peptides have been sorted on the basis of score in descending order and top 1% natural peptides have been extracted. The minimum score that we called threshold score was determined from these selected peptides. Similarly, threshold scores at 2, % were calculated. (iii) Step 1 and 2 were repeated for each MHC allele in order to calculate threshold score at different percent for each allele used in ProPred1. Prediction of MHC binders First, all possible overlapping 9mer peptides were generated for a given antigen sequence. The score of these 9mer peptides were calculated using quantitative matrix of selected MHC alleles. In next step, all peptides having score greater than selected threshold score (e.g. at 4%) were assigned as predicted binders for selected MHC allele. Weight matrices for proteasomal cutters The weight matrices used in ProPred1 for the prediction of standard proteasome and immunoproteasome, have been obtained from the work of (Toes et al., 2001). The matrix for standard proteasome matrix was derived from Table 1 of (Toes et al., 2001), where each value has been divided by one thousand, in order to rationalize the score. The derived matrix is an addition matrix where score of a peptide is calculated by summing the score of each residue. Similar procedure has been adopted for deriving the matrix for immunoproteasome, from the Table 2 of (Toes et al., 2001) The major difference between proteasomes matrices and MHC matrices is that proteasomes consider the peptide of length 12 instead of nine. In case of proteasome the cutting site is at the center of 12mer peptide. 1010
3 ProPred1 Table 2. The percent coverage of binders for different MHC alleles using ProPred1 at default threshold 4%. MHC Alleles (Total binder, % Coverage) HLA-A*0201(1221, 75%) H2-Db (189, 74%) HLA-B*0702(79, 92%) HLA-A*0205(28, 61%) H2-Dd (89, 74%) HLA-B*2705(145, 98%) HLA-A*1101(116, 80%) H2-Kb (116, 78%) HLA-B*3501(254, 84%) HLA-A*3101(33, 70%) H2-Kd (277, 83%) HLA-B*5101(51, 92%) HLA-A1 (128, 77%) H2-Kk (28, 86%) HLA-B*5102(33, 94%) HLA-A2 (976, 69%) H2-Ld (113, 60%) HLA-B*5103(30, 97%) HLA-A2.1 (77, 64%) HLA-B*5401(60, 100%) HLA-B8 (130, 75%) HLA-A24 (60, 70%) HLA-B61 (22, 95%) HLA-B62 (29, 55%) HLA-A3 (191, 64%) HLA-B14 (81, 75%) HLA-Cw*0401(20,80%) HLA-B7 (134, 81%) HLA-B*5301 (64, 95%) Computation of threshold score The threshold scores for standard proteasome and immunoproteasome have been calculated at different percent by using the approach described above for calculation of threshold score for MHC alleles. The calculation of threshold score of proteasome matrices requires the 12mer overlapping peptides. The matrices and cutoff scores at different thresold 1, 2,... 10% are available at URL matrix.html. Prediction of cleavage site in antigen sequence In order to predict proteasome cleavage sites in an antigenic sequence. The overlapping 12mer peptides were generated for antigenic sequence and score of these peptides were calculated using weight matrix of proteasome. In next step, all peptides having score greater than selected threshold score (e.g. at 5%) are considered as peptides having proteasome cleavage site. The center positions of these peptides (6-position left and 6-position right) are considered as predicted proteasome cleavage site. Similar approach has been utilized for prediction of peptides having immunoproteasome cleavage site. Simultaneous prediction of MHC binders and proteasome cleavage sites The predicted MHC binders were filtered based on prediction of proteasome cleavage sites in an antigenic sequence. Firstly, the server computes the predicted MHC binders and their C terminus position for a selected MHC allele in an antigenic sequence. Secondly server predicts the cleavage sites of proteasome in an antigenic sequence at given threshold (e.g. at 5%). Finally, all predicted MHC binding peptides whose C terminal position coincides with proteasomes cleavage sites were filtered. In other words, server removes the MHC binders, who does not have cleavage site at C terminus. RESULTS Percent coverage The fundamental question in MHC prediction is whether the prediction of binders is worth or not. In other words, whether the server can distinguish between binders and non-binders with significant accuracy. Thus, it is important to evaluate the performance of various matrices. We calculated the percent coverage (percent of binders correctly predicted as binder) for each allele for which sufficient amount of data was available. The data of binders and non-binders corresponding to each MHC alleles has been extracted from MHCBN database (Bhasin et al., The number of binders varies from 20 to 1200 (See Table 2). The default threshold score 4% (score at which sensitivity and specificity are nearly the same for most of the MHC alleles used in this study) was used for the prediction of the binders. The percent coverage has been calculated from predicted results. The value of percent coverage varies from 50 to 98% as shown in Table 2. These results clearly indicate that in most of the cases percent coverage is more than 80% which is reasonably good. Almost all alleles showed reasonable percent coverage, which means threshold criteria and matrices used in ProPred1 are beneficial for experimental scientists. Performance of ProPred1 Nonetheless, the percent coverage is a useful measure to evaluate the ability of method for the identification of binders from a given sequence, but it does not provide any information about predicted false positive binders or accuracy of prediction etc. Following three parameters are commonly used to measure the performance of a method in the field of immunoinformatics TP Sensitivity = 100 (3) TP+ FN TN Specificity = 100 (4) TN+ FP TP+ TN Accuracy = (5) TP+ FP + TN+ FN The correlation coefficient (CC) is a rigorous parameter to measure the performance of a method, which can be defined as: CC = (TP TN) (FN FP) (TP+ FN) (TN+ FP) (TP+ FP) (TN+ FN). Where TP and TN are correctly predicted binders and non-binders respectively. FP and FN are wrongly predicted binders and non-binders respectively. (6) 1011
4 H.Singh and G.P.S.Raghava Table 3. The comprehensive performance of ProPred1 at different thresholds for MHC allele HLA-A*0201 (1220 binders & 56 non-binders) and H2-Kb (116 binders & 20 non-binders) Thresholds HLA-A*0201 H2-Kb Sensitivity Specificity Accuracy Correlation Senstivity Specificity Accuracy Correlation (%) (%) (%) coefficient (%) (%) (%) coefficient 1% % % % % Table 4. The performance of Propred1 for HLA-A*0201 on data set of 128 peptides (19 high-affinity & 27 intermediate-affinity binders) of protein PRAME. These peptides were obtained from Table 1 of Kessler et al. (2001) Threshold Correctly predicted high- Correctly predicted intermediate- (%) affinity binders (out of 19) affinity binders (out of 27) (21%) 1 (4%) (47%) 6 (22%) (53%) 14 (52%) (58%) 15 (56%) (63 %) 21 (77%) (68%) 22 (81%) (68%) 23 (85%) (79%) 24 (89%) (95%) 26 (96%) (100%) 27 (100%) In this study, we compute all the above parameters for ProPred1 for its comprehensive evaluation. In order to evaluate a method one need sufficient data of experimentally proven MHC binders and non-binders. Unfortunately, most of the alleles have very limited number of binders and non-binders. Thus, the comprehensive evaluation of ProPred1 was performed only for two alleles (HLA-A*0201 & H2-Kb) for which sufficient number of binders and non-binders were available. The peptides for allele HLA-A*0201 (1220 binders & 56 non-binders) and H2-Kb (300 binders & 200 non-binders) were obtained from MHCBN database (Bhasin et al., 2002). The performance of ProPred1 for these two MHC alleles at different percent threshold has been shown in Table 3. A test for Propred1 MHC binder: The purpose of development of ProPred1 is to effectively reduce number of wet lab experiments involved in the identification of potential T-cell epitopes or suitable vaccine candidates. Recently, (Kessler et al., 2001) have experimentally determined the MHC binders and T-cell epitopes from tumor associated antigenic protein, PRAME. We analyzed the performance of ProPred1 in the identification of experimentally proven MHC binders and T-cell epitopes of PRAME. The sequence of PRAME antigenic protein was obtained from SWISSPROT database. (Kessler et al., 2001) tested 128 peptides and identified 19 as high-affinity binders and 27 intermediate-affinity binders. ProPred1 was used to predict these MHC 128 peptides of PRAME at various thresholds (See Table 4). As shown in Table 4, number of correctly predicted binders (intermediate/high affinity) depends on percent threshold. The ProPred1 predicted all binders correctly at 10% threshold. These results clearly indicate that server has capability to predict the binders with significant accuracy at 4% threshold (default threshold Potential T-cell epitopes: It has been demonstrated experimentally that MHC binders having proteasomal cleavage site at C-terminus are mostly responsible for the activation of cytotoxic T lymphocytes (CTLs). (Kessler et al., 2001) experimentally identified four regions having HLA-A*201 restricted T-cell epitopes. We tested these regions using ProPred1 server. Firstly, binding regions were predicted at default threshold (4%) in protein PRAME. Secondly, all proteasome sites were predicted at various thresholds. Finally, predicted binders having proteasome cleavage sites at C-terminus were identified. The number of peptides predicted by above falls in regions identified as T-cell epitopes by (Kessler et al., 2001), is shown in Table 5. It was observed that in the presence of standard proteasome filter at 7%, the server was able to predict the 50% of binding regions that are in agreement with experimentally proven binding regions as demonstrated by (Kessler et al., 2001). Similarly, it has been observed that at 5% of threshold of immunoproteasome filter, the server was able to identify 75% of experimentally determined binding regions. The server was able to predict 75% of binding regions in simultaneous presence of either standard proteasome or immunoproteasome filters at 5% threshold. 1012
5 ProPred1 Table 5. Propred1 was tested on four regions of PRAME (A: ; B: ; C: ; D: ) which were identified as T-cell epitopes by Kessler et al. (2001). Propred1 was first used to predict the HLA- A*0201 restricted peptide at 4% threshold. The column 2 shows the number of predicted peptides and regions (in bracket), which agree with the experimentally identified epitopes Name of filter Correctly predicted T-cell epitopes in protein PRAME at different thresholds (out of 4) 2% 3% 5% 7% Standard proteasome 0 1 (A) 1 (A) 2 (A,D) Immunoproteasome 2 (A,D) 2 (A,D) 3 (A,C,D) 3 (A,C,D) Immunoproteasome or standard 2 (A,D) 2 (A,D) 3 (A,C,D) 3 (A,C,D) proteasome Hence, all the analysis clearly indicate that it is worth using ProPred1 for the identification of MHC binding regions having proteasomal cleavage site at their C terminus or potential T-cell epitopes. It is not possible to determine the default threshold in case of prediction of proteasomes cleavage site, because sufficient data is not available, however, we suggest default threshold 5% based on our experience. Mapping of predicted binders on antigen sequence One of the important aspects of MHC prediction is the representation of binding peptides found within the antigenic sequence. This can be achieved by developing a powerful web interface for the prediction method. The ProPred1 provides three major options to visualize results in user-friendly formats, including most popular tabular format. Following is the brief description of these options. Graphical display: The graphical output represents the quantitative estimation of MHC binding propensity of the antigenic sequence. The server represents results in graphical format (X Y Plot), where amino acid sequence is shown along the X-axis and peptide score is shown along the Y -axis. Each binder is represented as a peak crossing the dashed threshold line in the image. It allows user to locate the promiscuous regions in the query sequence by looking at the peaks in graphs for different MHC alleles. Text or HTML format: This option of server presents the MHC binders within antigenic sequence in text or HTML format. It has two sub-options. The first sub-option displays the predicted MHC binders in separate lines along the antigen sequence. This option uses the separate lines for representing all the predicted overlapping binders within the sequence. This suboption is very useful for viewing the predicted overlapping binders. The second sub-option of the server represents predicted binder by different color i.e. blue. The first position of each binder is shown by red color so that user can easily distinguish the overlapping peptides. This option is useful in locating promiscuous MHC binders. Tabular format: This is the most widely used option for the display of results in most of the web servers of MHC prediction. This option displays the peptides sorted in descending order of their score. The server creates a separate table corresponding to each selected allele. IMPLEMENTATION The common gateway interface (CGI) script of ProPred1 is written using PERL. It has been installed on a Sun Server (420E) under UNIX (Solaris 7) environment and launched using Apache web server. The protein sequences can be submitted to the ProPred1 by cut-and-paste technique or by directly uploading a sequence file. The server uses ReadSeq (developed by Dr Don Gilbert) to parse the input sequence, therefore it can accept most of the commonly used sequence formats. The server allows user to select the threshold for their prediction. The threshold plays a vital role in determining the stringency of prediction. Lower the threshold, higher is the stringency of prediction i.e. lowers rate of false positives and higher rate of false negatives in the prediction. In contrast, a higher threshold value (low stringency) corresponds to a higher rate of false positives and a lower rate of false negatives. Limitations of ProPred1 All the matrices used in server were obtained from various servers and from the literature. The base of selection of matrices is on its availability from single source and not on the performance. Thus, it is not necessary that we are using best matrix for an allele if more than one matrix is available in the literature. In this server only 9mer peptide length are predicted not 8mer or 10mer. Thus it is possible that ProPred1 may miss potential 8mer and 10mer binders. The matrices for predicting ProPred1 were obtained from the paper of Toes et al. (2001), where their values were obtained for enolase-i protein. We have used these values for all predictions, it is not necessary that this generalization will work for all the proteins. ACKNOWLEDGEMENTS We are grateful to the anonymous referees for their valuable suggestions. Authors are thankful to Mr Manoj Bhasin for his help in data analysis. We are also thankful to Dr Balvinder Singh and Dr G. C. Varshney for checking the manuscript. 1013
6 H.Singh and G.P.S.Raghava REFERENCES Ayyoub,M., Stevanovic,S., Sahin,U., Guillaume,P., Servis,C., Rimoldi,D., Valmori,D., Romero,P., Cerottini,J.C., Rammensee,H.G. et al. (2002) Proteasome-assisted identification of a SSX-2-derived epitope recognized by tumorreactive CTL infiltrating metastatic melanoma. J. Immunol., 168, Bhasin,M., Singh,H. and Raghava,G.P.S. (2002) MHCBN: a comprehensive database of MHC binding and non-binding peptides. Bioinformatics, 19, Craiu,A., Akopian,T., Goldberg,A. and Rock,K.L. (1997) Two distinct proteolytic processes in the generation of a major histocompatibility complex class I-presented peptide. Proc. Natl Acad. Sci. USA, 94, Hagmann,M. (2000) Computers aid vaccine design. Science, 290, Holzhutter,H.G., Frommel,C. and Kloetzel,P.M. (1999) A theoretical approach towards the identification of cleavage-determining amino acid motifs of the 20S proteasome. J. Mol. Biol., 286, Kessler,J.H., Beekman,N.J., Bres-Vloemans,S.A., Verdijk,P., vanveelen,p.a., Kloosterman-Joosten,A.M., Vissers,D.C.J., ten Bosch,G.J.A., Kester,M.G.D., Sijts,A., Drijfhout,J.W. et al. (2001) Efficient identification of novel HLA-A*0201- presented cytotoxic T lymphocyte epitopes in the widely expressed tumor antigen PRAME by proteasome-mediated digestion analysis. J. Exp. Med., 193, Kuttler,C., Nussbaum,A.K., Dick,T.P., Rammensee,H.G., Schild,H. and Hadeler,K.P. (2000) An algorithm for the prediction of proteasomal cleavages. J. Mol. Biol., 298, Niedermann,G., Geier,E., Lucchiari-Hartz,M., Hitziger,N., Ramsperger,A. and Eichmann,K. (1999) The specificity of proteasomes: impact on MHC class I processing and presentation of antigens. Immunol. Rev., 127, Nussbaum,A.K., Kuttler,C., Hadeler,K.P., Rammensee,H.G. and Schild,H. (2001) PAProC: a prediction algorithm for proteasomal cleavages available on the WWW. Immunogenetics, 53, Parker,K.C., Bednarek,M.A. and Coligan,J.E. (1994) Scheme for ranking potential HLA-A2 binding peptides based on independent binding of individual peptide side-chains. J. Immunol., 152, Rammensee,H., Bachmann,J., Emmerich,N.P., Bachor,O.A. and Stevanovic,S. (1999) SYFPEITHI: database for MHC ligands and peptide motifs. Immunogenetics, 50, Rock,K.H. and Goldberg,A.L. (1999) Degradation of cell proteins and the generation of MHC class I presented peptides. Annu. Rev. Immunol., 17, Ruppert,J., Sidney,J., Celis,E., Kubo,R.T., Grey,H.M and Sette,A. (1993) Prominent role of secondary anchor residues in peptide binding to HLA-A2.1 molecules. Cell, 74, Sidney,J., Southwood,S., Guercio,M.D., Grey,H.M., Chesnut,R.W., Kubo,R.T. and Sette,A. (1996) Specificity and degeneracy in peptide binding to HLA-B7 like class 1 molecules. J. Immunol., 157, Singh,H. and Raghava,G.P.S. (2001) ProPred: prediction of HLA- DR binding sites. Bioinformatics, 17, Stoltze,L., Dick,T.P., Deeg,M., Pommerl,B., Rammensee,H.G. and Schild,H. (1998) Generation of the vesicular stomatitis virus nucleoprotein cytotoxic T lymphocyte epitope requires proteasomedependent and independent proteolytic activities. Eur. J. Immunol., 28, Sturniolo,T., Bono,E., Ding,J., Raddrizzani,L., Tuereci,O., Sahin,U., Braxenthaler,M., Gallazzi,F., Protti,M.P., Sinigaglia,F. and Hammer,J. (1999) Generation of tissue-specific and promiscuous HLA ligand database using DNA microarrays and virtual HLA class II matrices. Nat. Biotech., 17, Toes,R.E., Nussbaum,A.K., Degermann,S., Schirle,M., Emmerich,N.P.N., Kraft,M., Laplace,C., Zwinderman,A., Dick,T.P., Muller,J. et al. (2001) Discrete cleavage motifs of constitutive and immunoproteasomes revealed by quantitative analysis of cleavage products. J. Exp. Med., 194,
The Immune Epitope Database Analysis Resource: MHC class I peptide binding predictions. Edita Karosiene, Ph.D.
 The Immune Epitope Database Analysis Resource: MHC class I peptide binding predictions Edita Karosiene, Ph.D. edita@liai.org IEDB Workshop October 29, 2015 Outline Introduction MHC-I peptide binding prediction
The Immune Epitope Database Analysis Resource: MHC class I peptide binding predictions Edita Karosiene, Ph.D. edita@liai.org IEDB Workshop October 29, 2015 Outline Introduction MHC-I peptide binding prediction
PEPVAC: a web server for multi-epitope vaccine development based on the prediction of supertypic MHC ligands
 W138 W142 doi:10.1093/nar/gki357 PEPVAC: a web server for multi-epitope vaccine development based on the prediction of supertypic MHC ligands Pedro A. Reche 1,2, * and Ellis L. Reinherz 1,2 1 Laboratory
W138 W142 doi:10.1093/nar/gki357 PEPVAC: a web server for multi-epitope vaccine development based on the prediction of supertypic MHC ligands Pedro A. Reche 1,2, * and Ellis L. Reinherz 1,2 1 Laboratory
Eur. J. Immunol : Antigen processing 2295
 Eur. J. Immunol. 2005. 35: 2295 2303 Antigen processing 2295 Antigen processing An integrative approach to CTL epitope prediction: A combined algorithm integrating MHC class I binding, TAP transport efficiency,
Eur. J. Immunol. 2005. 35: 2295 2303 Antigen processing 2295 Antigen processing An integrative approach to CTL epitope prediction: A combined algorithm integrating MHC class I binding, TAP transport efficiency,
MetaMHC: a meta approach to predict peptides binding to MHC molecules
 W474 W479 Nucleic Acids Research, 2010, Vol. 38, Web Server issue Published online 18 May 2010 doi:10.1093/nar/gkq407 MetaMHC: a meta approach to predict peptides binding to MHC molecules Xihao Hu 1, Wenjian
W474 W479 Nucleic Acids Research, 2010, Vol. 38, Web Server issue Published online 18 May 2010 doi:10.1093/nar/gkq407 MetaMHC: a meta approach to predict peptides binding to MHC molecules Xihao Hu 1, Wenjian
Definition of MHC supertypes through clustering of MHC peptide binding repertoires
 Definition of MHC supertypes through clustering of MHC peptide binding repertoires Pedro A. Reche and Ellis L. Reinherz Laboratory of Immunobiology and Department of Medical Oncology, Dana-Farber Cancer
Definition of MHC supertypes through clustering of MHC peptide binding repertoires Pedro A. Reche and Ellis L. Reinherz Laboratory of Immunobiology and Department of Medical Oncology, Dana-Farber Cancer
A HLA-DRB supertype chart with potential overlapping peptide binding function
 A HLA-DRB supertype chart with potential overlapping peptide binding function Arumugam Mohanapriya 1,2, Satish Nandagond 1, Paul Shapshak 3, Uma Kangueane 1, Pandjassarame Kangueane 1, 2 * 1 Biomedical
A HLA-DRB supertype chart with potential overlapping peptide binding function Arumugam Mohanapriya 1,2, Satish Nandagond 1, Paul Shapshak 3, Uma Kangueane 1, Pandjassarame Kangueane 1, 2 * 1 Biomedical
Identification and characterization of merozoite surface protein 1 epitope
 Identification and characterization of merozoite surface protein epitope Satarudra Prakash Singh,2, Bhartendu Nath Mishra 2* Amity Institute of Biotechnology, Amity University, Uttar Pradesh, Gomti Nagar,
Identification and characterization of merozoite surface protein epitope Satarudra Prakash Singh,2, Bhartendu Nath Mishra 2* Amity Institute of Biotechnology, Amity University, Uttar Pradesh, Gomti Nagar,
Definition of MHC Supertypes Through Clustering of MHC Peptide-Binding Repertoires
 11 Definition of MHC Supertypes Through Clustering of MHC Peptide-Binding Repertoires Pedro A. Reche and Ellis L. Reinherz Summary Identification of peptides that can bind to major histocompatibility complex
11 Definition of MHC Supertypes Through Clustering of MHC Peptide-Binding Repertoires Pedro A. Reche and Ellis L. Reinherz Summary Identification of peptides that can bind to major histocompatibility complex
SUPPLEMENTARY INFORMATION
 Supplementary Notes 1: accuracy of prediction algorithms for peptide binding affinities to HLA and Mamu alleles For each HLA and Mamu allele we have analyzed the accuracy of four predictive algorithms
Supplementary Notes 1: accuracy of prediction algorithms for peptide binding affinities to HLA and Mamu alleles For each HLA and Mamu allele we have analyzed the accuracy of four predictive algorithms
CS229 Final Project Report. Predicting Epitopes for MHC Molecules
 CS229 Final Project Report Predicting Epitopes for MHC Molecules Xueheng Zhao, Shanshan Tuo Biomedical informatics program Stanford University Abstract Major Histocompatibility Complex (MHC) plays a key
CS229 Final Project Report Predicting Epitopes for MHC Molecules Xueheng Zhao, Shanshan Tuo Biomedical informatics program Stanford University Abstract Major Histocompatibility Complex (MHC) plays a key
Prediction of proteasome cleavage motifs by neural networks
 Protein Engineering vol.15 no.4 pp.287 296, 2002 Prediction of proteasome cleavage motifs by neural networks Can Keşmir 1,2,6, Alexander K.Nussbaum 3, Hansjörg Schild 3, Vincent Detours 4,5 and Søren Brunak
Protein Engineering vol.15 no.4 pp.287 296, 2002 Prediction of proteasome cleavage motifs by neural networks Can Keşmir 1,2,6, Alexander K.Nussbaum 3, Hansjörg Schild 3, Vincent Detours 4,5 and Søren Brunak
Use of BONSAI decision trees for the identification of potential MHC Class I peptide epitope motifs.
 Use of BONSAI decision trees for the identification of potential MHC Class I peptide epitope motifs. C.J. SAVOIE, N. KAMIKAWAJI, T. SASAZUKI Dept. of Genetics, Medical Institute of Bioregulation, Kyushu
Use of BONSAI decision trees for the identification of potential MHC Class I peptide epitope motifs. C.J. SAVOIE, N. KAMIKAWAJI, T. SASAZUKI Dept. of Genetics, Medical Institute of Bioregulation, Kyushu
Profiling HLA motifs by large scale peptide sequencing Agilent Innovators Tour David K. Crockett ARUP Laboratories February 10, 2009
 Profiling HLA motifs by large scale peptide sequencing 2009 Agilent Innovators Tour David K. Crockett ARUP Laboratories February 10, 2009 HLA Background The human leukocyte antigen system (HLA) is the
Profiling HLA motifs by large scale peptide sequencing 2009 Agilent Innovators Tour David K. Crockett ARUP Laboratories February 10, 2009 HLA Background The human leukocyte antigen system (HLA) is the
A community resource benchmarking predictions of peptide binding to MHC-I molecules
 Downloaded from orbit.dtu.dk on: Dec 17, 2017 A community resource benchmarking predictions of peptide binding to MHC-I molecules Peters, B; Bui, HH; Pletscher-Frankild, Sune; Nielsen, Morten; Lundegaard,
Downloaded from orbit.dtu.dk on: Dec 17, 2017 A community resource benchmarking predictions of peptide binding to MHC-I molecules Peters, B; Bui, HH; Pletscher-Frankild, Sune; Nielsen, Morten; Lundegaard,
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data
 Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Immuno-Oncology Therapies and Precision Medicine: Personal Tumor-Specific Neoantigen Prediction by Machine Learning
 Immuno-Oncology Therapies and Precision Medicine: Personal Tumor-Specific Neoantigen Prediction by Machine Learning Yi-Hsiang Hsu, MD, SCD Sep 16, 2017 yihsianghsu@hsl.harvard.edu HSL GeneticEpi Center,
Immuno-Oncology Therapies and Precision Medicine: Personal Tumor-Specific Neoantigen Prediction by Machine Learning Yi-Hsiang Hsu, MD, SCD Sep 16, 2017 yihsianghsu@hsl.harvard.edu HSL GeneticEpi Center,
Molecular Graphics Perspective of Protein Structure and Function
 Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Chapter 6. Antigen Presentation to T lymphocytes
 Chapter 6 Antigen Presentation to T lymphocytes Generation of T-cell Receptor Ligands T cells only recognize Ags displayed on cell surfaces These Ags may be derived from pathogens that replicate within
Chapter 6 Antigen Presentation to T lymphocytes Generation of T-cell Receptor Ligands T cells only recognize Ags displayed on cell surfaces These Ags may be derived from pathogens that replicate within
IMMUNOINFORMATICS: Bioinformatics Challenges in Immunology
 Bioinformatics 1 -- Lecture 22 IMMUNOINFORMATICS: Bioinformatics Challenges in Immunology Most slides courtesy of Julia Ponomarenko, San Diego Supercomputer Center or Oliver Kohlbacher, WSI/ZBIT, Eberhard-Karls-
Bioinformatics 1 -- Lecture 22 IMMUNOINFORMATICS: Bioinformatics Challenges in Immunology Most slides courtesy of Julia Ponomarenko, San Diego Supercomputer Center or Oliver Kohlbacher, WSI/ZBIT, Eberhard-Karls-
BIOINFORMATICS. June Immunological Bioinformatics
 IMMUNOLOGICAL BIOINFORMATICS MEHTEIDHWLEFSATKLSSCDSFTSTINELNHCLSLRT YLVGNSLSLADLCVWATLKGNAAWQEQLKQKKAPVH VKRWFGFLEAQQAFQSVGTKWDVSTTKARVAPEKKQ DVGKFVELPGAEMGKVTVRFPPEASGYLHIGHAKAA LLNQHYQVNFKGKLIMRFDDTNPEKEKEDFEKVILE
IMMUNOLOGICAL BIOINFORMATICS MEHTEIDHWLEFSATKLSSCDSFTSTINELNHCLSLRT YLVGNSLSLADLCVWATLKGNAAWQEQLKQKKAPVH VKRWFGFLEAQQAFQSVGTKWDVSTTKARVAPEKKQ DVGKFVELPGAEMGKVTVRFPPEASGYLHIGHAKAA LLNQHYQVNFKGKLIMRFDDTNPEKEKEDFEKVILE
Contents. Just Classifier? Rules. Rules: example. Classification Rule Generation for Bioinformatics. Rule Extraction from a trained network
 Contents Classification Rule Generation for Bioinformatics Hyeoncheol Kim Rule Extraction from Neural Networks Algorithm Ex] Promoter Domain Hybrid Model of Knowledge and Learning Knowledge refinement
Contents Classification Rule Generation for Bioinformatics Hyeoncheol Kim Rule Extraction from Neural Networks Algorithm Ex] Promoter Domain Hybrid Model of Knowledge and Learning Knowledge refinement
Immuno-Oncology Therapies and Precision Medicine: Personal Tumor-Specific Neoantigen Prediction by Machine Learning
 Immuno-Oncology Therapies and Precision Medicine: Personal Tumor-Specific Neoantigen Prediction by Machine Learning Yi-Hsiang Hsu, MD, SCD Sep 16, 2017 yihsianghsu@hsl.harvard.edu Director & Associate
Immuno-Oncology Therapies and Precision Medicine: Personal Tumor-Specific Neoantigen Prediction by Machine Learning Yi-Hsiang Hsu, MD, SCD Sep 16, 2017 yihsianghsu@hsl.harvard.edu Director & Associate
BIOINF 3360 Computational Immunomics
 BIOINF 3360 Computational Immunomics Oliver Kohlbacher Summer 2011 11. Summary TYOTK Things you ought to know! Caveat: These are just examples, not the full list of potential ti exam questions! TYOTK:
BIOINF 3360 Computational Immunomics Oliver Kohlbacher Summer 2011 11. Summary TYOTK Things you ought to know! Caveat: These are just examples, not the full list of potential ti exam questions! TYOTK:
BIOINF 3360 Computational Immunomics
 BIOINF 3360 Computational Immunomics Oliver Kohlbacher Summer 2011 11. Summary TYOTK Things you ought to know! Caveat: These are just examples, not the full list of potential exam questions! TYOTK: Immunology
BIOINF 3360 Computational Immunomics Oliver Kohlbacher Summer 2011 11. Summary TYOTK Things you ought to know! Caveat: These are just examples, not the full list of potential exam questions! TYOTK: Immunology
Degenerate T-cell Recognition of Peptides on MHC Molecules Creates Large Holes in the T-cell Repertoire
 Degenerate T-cell Recognition of Peptides on MHC Molecules Creates Large Holes in the T-cell Repertoire Jorg J. A. Calis*, Rob J. de Boer, Can Keşmir Theoretical Biology & Bioinformatics, Utrecht University,
Degenerate T-cell Recognition of Peptides on MHC Molecules Creates Large Holes in the T-cell Repertoire Jorg J. A. Calis*, Rob J. de Boer, Can Keşmir Theoretical Biology & Bioinformatics, Utrecht University,
Epitope discovery. Using predicted MHC binding CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS
 Epitope discovery Using predicted MHC binding SARS corona virus The 2003 outbreak The disease Lung Pathology Inflammatory exudation in alveoli and interstitial spaces Monocytic and lymphocytic infiltration
Epitope discovery Using predicted MHC binding SARS corona virus The 2003 outbreak The disease Lung Pathology Inflammatory exudation in alveoli and interstitial spaces Monocytic and lymphocytic infiltration
The Major Histocompatibility Complex (MHC)
 The Major Histocompatibility Complex (MHC) An introduction to adaptive immune system before we discuss MHC B cells The main cells of adaptive immune system are: -B cells -T cells B cells: Recognize antigens
The Major Histocompatibility Complex (MHC) An introduction to adaptive immune system before we discuss MHC B cells The main cells of adaptive immune system are: -B cells -T cells B cells: Recognize antigens
Predicting Protein-Peptide Binding Affinity by Learning Peptide-Peptide Distance Functions
 Predicting Protein-Peptide Binding Affinity by Learning Peptide-Peptide Distance Functions Chen Yanover and Tomer Hertz,2 School of Computer Science and Engineering 2 The Center for Neural Computation,
Predicting Protein-Peptide Binding Affinity by Learning Peptide-Peptide Distance Functions Chen Yanover and Tomer Hertz,2 School of Computer Science and Engineering 2 The Center for Neural Computation,
LIPOPREDICT: Bacterial lipoprotein prediction server
 www.bioinformation.net Server Volume 8(8) LIPOPREDICT: Bacterial lipoprotein prediction server S Ramya Kumari, Kiran Kadam, Ritesh Badwaik & Valadi K Jayaraman* Centre for Development of Advanced Computing
www.bioinformation.net Server Volume 8(8) LIPOPREDICT: Bacterial lipoprotein prediction server S Ramya Kumari, Kiran Kadam, Ritesh Badwaik & Valadi K Jayaraman* Centre for Development of Advanced Computing
User Guide. Protein Clpper. Statistical scoring of protease cleavage sites. 1. Introduction Protein Clpper Analysis Procedure...
 User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
Vaccine Design: A Statisticans Overview
 GoBack : A Statisticans Overview. Surajit Ray sray@samsi.info Surajit Ray Samsi PostDoc Seminar: Nov 2: 2004 - slide #1 The Chinese are credited with making the observation that deliberately infecting
GoBack : A Statisticans Overview. Surajit Ray sray@samsi.info Surajit Ray Samsi PostDoc Seminar: Nov 2: 2004 - slide #1 The Chinese are credited with making the observation that deliberately infecting
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
MHC Tetramers and Monomers for Immuno-Oncology and Autoimmunity Drug Discovery
 MHC Tetramers and Monomers for Immuno-Oncology and Autoimmunity Drug Discovery Your Partner in Drug Discovery and Research MHC Tetramer Background T-Cell Receptors recognize and bind to complexes composed
MHC Tetramers and Monomers for Immuno-Oncology and Autoimmunity Drug Discovery Your Partner in Drug Discovery and Research MHC Tetramer Background T-Cell Receptors recognize and bind to complexes composed
Leukemia (2007) 21, & 2007 Nature Publishing Group All rights reserved /07 $
 REVIEW (2007) 21, 1859 1874 & 2007 Nature Publishing Group All rights reserved 0887-6924/07 $30.00 www.nature.com/leu Department of Immunohematology and Blood Transfusion, Leiden University Medical Center,
REVIEW (2007) 21, 1859 1874 & 2007 Nature Publishing Group All rights reserved 0887-6924/07 $30.00 www.nature.com/leu Department of Immunohematology and Blood Transfusion, Leiden University Medical Center,
Short peptide epitope design from hantaviruses causing HFRS
 www.bioinformation.net Volume 13(7) Hypothesis Short peptide epitope design from hantaviruses causing HFRS Sathish Sankar*, Mageshbabu Ramamurthy, Balaji Nandagopal, Gopalan Sridharan Sri Sakthi Amma Institute
www.bioinformation.net Volume 13(7) Hypothesis Short peptide epitope design from hantaviruses causing HFRS Sathish Sankar*, Mageshbabu Ramamurthy, Balaji Nandagopal, Gopalan Sridharan Sri Sakthi Amma Institute
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16
 38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
There are approximately 30,000 proteasomes in a typical human cell Each proteasome is approximately 700 kda in size The proteasome is made up of 3
 Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
NIH Public Access Author Manuscript Immunogenetics. Author manuscript; available in PMC 2014 September 01.
 NIH Public Access Author Manuscript Published in final edited form as: Immunogenetics. 2013 September ; 65(9): 655 665. doi:10.1007/s00251-013-0714-9. MHCcluster, a method for functional clustering of
NIH Public Access Author Manuscript Published in final edited form as: Immunogenetics. 2013 September ; 65(9): 655 665. doi:10.1007/s00251-013-0714-9. MHCcluster, a method for functional clustering of
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Selection of epitope-based vaccine targets of HCV genotype 1 of Asian origin: a systematic in silico approach
 www.bioinformation.net Hypothesis Volume 8(20) Selection of epitope-based vaccine targets of HCV genotype 1 of Asian origin: a systematic in silico approach Abida Shehzadi 1 *, Shahid Ur Rehman 1, 2 &
www.bioinformation.net Hypothesis Volume 8(20) Selection of epitope-based vaccine targets of HCV genotype 1 of Asian origin: a systematic in silico approach Abida Shehzadi 1 *, Shahid Ur Rehman 1, 2 &
Antigen Recognition by T cells
 Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
Integrated modeling of the major events in the MHC class I antigen processing pathway
 Integrated modeling of the major events in the MHC class I antigen processing pathway PIERRE DÖNNES AND OLIVER KOHLBACHER Department for Simulation of Biological Systems, WSI/ZBIT, Eberhard Karls University
Integrated modeling of the major events in the MHC class I antigen processing pathway PIERRE DÖNNES AND OLIVER KOHLBACHER Department for Simulation of Biological Systems, WSI/ZBIT, Eberhard Karls University
Our T cell epitope test service offers you:
 Our T cell epitope test service offers you: high throughput peptide discovery fast lead-finding maximum of epitopes in a minimum of blood saving effort and costs on developing your own platforms custom-made
Our T cell epitope test service offers you: high throughput peptide discovery fast lead-finding maximum of epitopes in a minimum of blood saving effort and costs on developing your own platforms custom-made
Workshop presentation: Development and application of Bioinformatics methods
 Workshop presentation: Development and application of Bioinformatics methods Immune system Innate fast Addaptive remembers Cellular Cytotoxic T lymphocytes (CTL) Helper T lymphocytes (HTL) Humoral B lymphocytes
Workshop presentation: Development and application of Bioinformatics methods Immune system Innate fast Addaptive remembers Cellular Cytotoxic T lymphocytes (CTL) Helper T lymphocytes (HTL) Humoral B lymphocytes
Data mining with Ensembl Biomart. Stéphanie Le Gras
 Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
DIRECT IDENTIFICATION OF NEO-EPITOPES IN TUMOR TISSUE
 DIRECT IDENTIFICATION OF NEO-EPITOPES IN TUMOR TISSUE Eustache Paramithiotis PhD Vice President, Biomarker Discovery & Diagnostics 17 March 2016 PEPTIDE PRESENTATION BY MHC MHC I Antigen presentation by
DIRECT IDENTIFICATION OF NEO-EPITOPES IN TUMOR TISSUE Eustache Paramithiotis PhD Vice President, Biomarker Discovery & Diagnostics 17 March 2016 PEPTIDE PRESENTATION BY MHC MHC I Antigen presentation by
SubLasso:a feature selection and classification R package with a. fixed feature subset
 SubLasso:a feature selection and classification R package with a fixed feature subset Youxi Luo,3,*, Qinghan Meng,2,*, Ruiquan Ge,2, Guoqin Mai, Jikui Liu, Fengfeng Zhou,#. Shenzhen Institutes of Advanced
SubLasso:a feature selection and classification R package with a fixed feature subset Youxi Luo,3,*, Qinghan Meng,2,*, Ruiquan Ge,2, Guoqin Mai, Jikui Liu, Fengfeng Zhou,#. Shenzhen Institutes of Advanced
IMPaLA tutorial.
 IMPaLA tutorial http://impala.molgen.mpg.de/ 1. Introduction IMPaLA is a web tool, developed for integrated pathway analysis of metabolomics data alongside gene expression or protein abundance data. It
IMPaLA tutorial http://impala.molgen.mpg.de/ 1. Introduction IMPaLA is a web tool, developed for integrated pathway analysis of metabolomics data alongside gene expression or protein abundance data. It
PERFORMANCE MEASURES
 PERFORMANCE MEASURES Of predictive systems DATA TYPES Binary Data point Value A FALSE B TRUE C TRUE D FALSE E FALSE F TRUE G FALSE Real Value Data Point Value a 32.3 b.2 b 2. d. e 33 f.65 g 72.8 ACCURACY
PERFORMANCE MEASURES Of predictive systems DATA TYPES Binary Data point Value A FALSE B TRUE C TRUE D FALSE E FALSE F TRUE G FALSE Real Value Data Point Value a 32.3 b.2 b 2. d. e 33 f.65 g 72.8 ACCURACY
MHC class I MHC class II Structure of MHC antigens:
 MHC class I MHC class II Structure of MHC antigens: MHC class I antigens consist of a transmembrane heavy chain (α chain) that is non-covalently associated with β2- microglobulin. Membrane proximal domain
MHC class I MHC class II Structure of MHC antigens: MHC class I antigens consist of a transmembrane heavy chain (α chain) that is non-covalently associated with β2- microglobulin. Membrane proximal domain
Machine Intelligence Approach for Development of MHC Binders and Fragment Based Peptide Vaccines from Tetrahymena pyriformis
 Global Journal of Biotechnology & Biochemistry 5 (3): 161-165, 2010 ISSN 2078-466X IDOSI Publications, 2010 Machine Intelligence Approach for Development of MHC Binders and Fragment Based Peptide Vaccines
Global Journal of Biotechnology & Biochemistry 5 (3): 161-165, 2010 ISSN 2078-466X IDOSI Publications, 2010 Machine Intelligence Approach for Development of MHC Binders and Fragment Based Peptide Vaccines
Potential cross reactions between HIV 1 specific T cells and the microbiome. Andrew McMichael Suzanne Campion
 Potential cross reactions between HIV 1 specific T cells and the microbiome Andrew McMichael Suzanne Campion Role of the Microbiome? T cell (and B cell) immune responses to HIV and Vaccines are influenced
Potential cross reactions between HIV 1 specific T cells and the microbiome Andrew McMichael Suzanne Campion Role of the Microbiome? T cell (and B cell) immune responses to HIV and Vaccines are influenced
Protein Structure & Function. University, Indianapolis, USA 3 Department of Molecular Medicine, University of South Florida, Tampa, USA
 Protein Structure & Function Supplement for article entitled MoRFpred, a computational tool for sequence-based prediction and characterization of short disorder-to-order transitioning binding regions in
Protein Structure & Function Supplement for article entitled MoRFpred, a computational tool for sequence-based prediction and characterization of short disorder-to-order transitioning binding regions in
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity
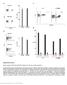 Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
SEQUENCE FEATURE VARIANT TYPES
 SEQUENCE FEATURE VARIANT TYPES DEFINITION OF SFVT: The Sequence Feature Variant Type (SFVT) component in IRD (http://www.fludb.org) is a relatively novel approach that delineates specific regions, called
SEQUENCE FEATURE VARIANT TYPES DEFINITION OF SFVT: The Sequence Feature Variant Type (SFVT) component in IRD (http://www.fludb.org) is a relatively novel approach that delineates specific regions, called
Efficacy of the Extended Principal Orthogonal Decomposition Method on DNA Microarray Data in Cancer Detection
 202 4th International onference on Bioinformatics and Biomedical Technology IPBEE vol.29 (202) (202) IASIT Press, Singapore Efficacy of the Extended Principal Orthogonal Decomposition on DA Microarray
202 4th International onference on Bioinformatics and Biomedical Technology IPBEE vol.29 (202) (202) IASIT Press, Singapore Efficacy of the Extended Principal Orthogonal Decomposition on DA Microarray
Immune Epitope Database NEWSLETTER Volume 6, Issue 2 July 2009
 Immune Epitope Database NEWSLETTER Volume 6, Issue 2 http://www.iedb.org July 2009 In response to concerns of a swine flu epidemic, the IEDB Curation curation team updated the database s collection of
Immune Epitope Database NEWSLETTER Volume 6, Issue 2 http://www.iedb.org July 2009 In response to concerns of a swine flu epidemic, the IEDB Curation curation team updated the database s collection of
ACE ImmunoID. ACE ImmunoID. Precision immunogenomics. Precision Genomics for Immuno-Oncology
 ACE ImmunoID ACE ImmunoID Precision immunogenomics Precision Genomics for Immuno-Oncology Personalis, Inc. A universal biomarker platform for immuno-oncology Patient response to cancer immunotherapies
ACE ImmunoID ACE ImmunoID Precision immunogenomics Precision Genomics for Immuno-Oncology Personalis, Inc. A universal biomarker platform for immuno-oncology Patient response to cancer immunotherapies
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Major Histocompatibility Complex Class II Prediction
 American Journal of Bioinformatics Research 2012, 2(1): 14-20 DOI: 10.5923/j.bioinformatics.20120201.03 Major Histocompatibility Complex Class II Prediction Zeinab Abd El Halim *, Amr Badr, Khaled Tawfik,
American Journal of Bioinformatics Research 2012, 2(1): 14-20 DOI: 10.5923/j.bioinformatics.20120201.03 Major Histocompatibility Complex Class II Prediction Zeinab Abd El Halim *, Amr Badr, Khaled Tawfik,
From single studies to an EBM based assessment some central issues
 From single studies to an EBM based assessment some central issues Doug Altman Centre for Statistics in Medicine, Oxford, UK Prognosis Prognosis commonly relates to the probability or risk of an individual
From single studies to an EBM based assessment some central issues Doug Altman Centre for Statistics in Medicine, Oxford, UK Prognosis Prognosis commonly relates to the probability or risk of an individual
Enhanced Asthma Management with Mobile Communication
 Enhanced Asthma Management with Mobile Communication P.S. Ngai, S. Chan, C.T. Lau, K.M. Lau Abstract In this paper, we propose a prototype system to enhance the management of asthma condition in patients
Enhanced Asthma Management with Mobile Communication P.S. Ngai, S. Chan, C.T. Lau, K.M. Lau Abstract In this paper, we propose a prototype system to enhance the management of asthma condition in patients
Chapter 7 Conclusions
 VII-1 Chapter 7 Conclusions VII-2 The development of cell-based therapies ranging from well-established practices such as bone marrow transplant to next-generation strategies such as adoptive T-cell therapy
VII-1 Chapter 7 Conclusions VII-2 The development of cell-based therapies ranging from well-established practices such as bone marrow transplant to next-generation strategies such as adoptive T-cell therapy
International Journal of Pharma and Bio Sciences ANTIGEN PROTEIN FROM SCHISTOSOMA MANSONI: NEW PARADIGM OF SYNTHETIC VACCINE DEVELOPMENT
 International Journal of Pharma and Bio Sciences ANTIGEN PROTEIN FROM SCHISTOSOMA MANSONI: NEW PARADIGM OF SYNTHETIC VACCINE DEVELOPMENT GOMASE V.S.* and CHITLANGE N.R. School of Technology, S.R.T.M. University,
International Journal of Pharma and Bio Sciences ANTIGEN PROTEIN FROM SCHISTOSOMA MANSONI: NEW PARADIGM OF SYNTHETIC VACCINE DEVELOPMENT GOMASE V.S.* and CHITLANGE N.R. School of Technology, S.R.T.M. University,
Warfarin Help Documentation
 Warfarin Help Documentation Table Of Contents Warfarin Management... 1 iii Warfarin Management Warfarin Management The Warfarin Management module is a powerful tool for monitoring INR results and advising
Warfarin Help Documentation Table Of Contents Warfarin Management... 1 iii Warfarin Management Warfarin Management The Warfarin Management module is a powerful tool for monitoring INR results and advising
Antigen Presentation and T Lymphocyte Activation. Abul K. Abbas UCSF. FOCiS
 1 Antigen Presentation and T Lymphocyte Activation Abul K. Abbas UCSF FOCiS 2 Lecture outline Dendritic cells and antigen presentation The role of the MHC T cell activation Costimulation, the B7:CD28 family
1 Antigen Presentation and T Lymphocyte Activation Abul K. Abbas UCSF FOCiS 2 Lecture outline Dendritic cells and antigen presentation The role of the MHC T cell activation Costimulation, the B7:CD28 family
TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer
 AD Award Number: W8XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins
AD Award Number: W8XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins
Analysis of Diabetic Dataset and Developing Prediction Model by using Hive and R
 Indian Journal of Science and Technology, Vol 9(47), DOI: 10.17485/ijst/2016/v9i47/106496, December 2016 ISSN (Print) : 0974-6846 ISSN (Online) : 0974-5645 Analysis of Diabetic Dataset and Developing Prediction
Indian Journal of Science and Technology, Vol 9(47), DOI: 10.17485/ijst/2016/v9i47/106496, December 2016 ISSN (Print) : 0974-6846 ISSN (Online) : 0974-5645 Analysis of Diabetic Dataset and Developing Prediction
NetMHCpan, a Method for Quantitative Predictions of Peptide Binding to Any HLA-A and -B Locus Protein of Known Sequence
 NetMHCpan, a Method for Quantitative Predictions of Peptide Binding to Any HLA-A and -B Locus Protein of Known Sequence Morten Nielsen 1 *, Claus Lundegaard 1, Thomas Blicher 1, Kasper Lamberth 2, Mikkel
NetMHCpan, a Method for Quantitative Predictions of Peptide Binding to Any HLA-A and -B Locus Protein of Known Sequence Morten Nielsen 1 *, Claus Lundegaard 1, Thomas Blicher 1, Kasper Lamberth 2, Mikkel
New technologies for studying human immunity. Lisa Wagar Postdoctoral fellow, Mark Davis lab Stanford University School of Medicine
 New technologies for studying human immunity Lisa Wagar Postdoctoral fellow, Mark Davis lab Stanford University School of Medicine New strategies: Human immunology is ideal for a systems approach We have
New technologies for studying human immunity Lisa Wagar Postdoctoral fellow, Mark Davis lab Stanford University School of Medicine New strategies: Human immunology is ideal for a systems approach We have
Top 10 Tips for Successful Searching ASMS 2003
 Top 10 Tips for Successful Searching I'd like to present our top 10 tips for successful searching with Mascot. Like any hit parade, we will, of course, count them off in reverse order 1 10. Don t specify
Top 10 Tips for Successful Searching I'd like to present our top 10 tips for successful searching with Mascot. Like any hit parade, we will, of course, count them off in reverse order 1 10. Don t specify
DEPARTMENT OF BIOTECHNOLOGY
 IDENTIFICATION OF PEPTIDE CONTAINING T CELL EPITOPE OF HEMAGGLUTININ PROTEIN IN H5N1 INFLUENZA VIRUS A thesis submitted in partial fulfillment of the requirements for the award Of the degree of MASTER
IDENTIFICATION OF PEPTIDE CONTAINING T CELL EPITOPE OF HEMAGGLUTININ PROTEIN IN H5N1 INFLUENZA VIRUS A thesis submitted in partial fulfillment of the requirements for the award Of the degree of MASTER
Antigen Presentation to T lymphocytes
 Antigen Presentation to T lymphocytes Immunology 441 Lectures 6 & 7 Chapter 6 October 10 & 12, 2016 Jessica Hamerman jhamerman@benaroyaresearch.org Office hours by arrangement Antigen processing: How are
Antigen Presentation to T lymphocytes Immunology 441 Lectures 6 & 7 Chapter 6 October 10 & 12, 2016 Jessica Hamerman jhamerman@benaroyaresearch.org Office hours by arrangement Antigen processing: How are
TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer
 AD Award Number: W8-XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins
AD Award Number: W8-XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins
Antigen Presentation to T lymphocytes
 Antigen Presentation to T lymphocytes Immunology 441 Lectures 6 & 7 Chapter 6 October 10 & 12, 2016 Jessica Hamerman jhamerman@benaroyaresearch.org Office hours by arrangement Antibodies and T cell receptors
Antigen Presentation to T lymphocytes Immunology 441 Lectures 6 & 7 Chapter 6 October 10 & 12, 2016 Jessica Hamerman jhamerman@benaroyaresearch.org Office hours by arrangement Antibodies and T cell receptors
Comparing Multifunctionality and Association Information when Classifying Oncogenes and Tumor Suppressor Genes
 000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
A general overview of Immune system and. Shuyan Li 2/4/2009
 A general overview of Immune system and Immunology Shuyan Li 2/4/2009 Definition Immune system a collection of biological processes within an organism that protects against disease by identifying and killing
A general overview of Immune system and Immunology Shuyan Li 2/4/2009 Definition Immune system a collection of biological processes within an organism that protects against disease by identifying and killing
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
1. Introduction. Department of Hematology, University of Siena, Siena, Italy 2
 Leukemia Research and Treatment Volume 2012, Article ID 150651, 4 pages doi:10.1155/2012/150651 Research Article Identification of a Novel P190-Derived Breakpoint Peptide Suitable for Peptide Vaccine Therapeutic
Leukemia Research and Treatment Volume 2012, Article ID 150651, 4 pages doi:10.1155/2012/150651 Research Article Identification of a Novel P190-Derived Breakpoint Peptide Suitable for Peptide Vaccine Therapeutic
Therapeutic Cancer Vaccines
 Therapeutic Cancer Vaccines Goal for all therapeutic cancer vaccines: To enhance the natural immune response so that it becomes an effective therapy Approaches being investigated in clinical studies: Whole
Therapeutic Cancer Vaccines Goal for all therapeutic cancer vaccines: To enhance the natural immune response so that it becomes an effective therapy Approaches being investigated in clinical studies: Whole
Inferring condition-specific mirna activity from matched mirna and mrna expression data
 Inferring condition-specific mirna activity from matched mirna and mrna expression data Junpeng Zhang 1, Thuc Duy Le 2, Lin Liu 2, Bing Liu 3, Jianfeng He 4, Gregory J Goodall 5 and Jiuyong Li 2,* 1 Faculty
Inferring condition-specific mirna activity from matched mirna and mrna expression data Junpeng Zhang 1, Thuc Duy Le 2, Lin Liu 2, Bing Liu 3, Jianfeng He 4, Gregory J Goodall 5 and Jiuyong Li 2,* 1 Faculty
Manitoba Health, Healthy Living and Seniors
 Manitoba Health, Healthy Living and Seniors Manitoba Annual Immunization Surveillance Report, 2012 and 2013 January 1, 2012 to December 31, 2013 with 5-year average comparison (January 1, 2007 to December
Manitoba Health, Healthy Living and Seniors Manitoba Annual Immunization Surveillance Report, 2012 and 2013 January 1, 2012 to December 31, 2013 with 5-year average comparison (January 1, 2007 to December
Below, we included the point-to-point response to the comments of both reviewers.
 To the Editor and Reviewers: We would like to thank the editor and reviewers for careful reading, and constructive suggestions for our manuscript. According to comments from both reviewers, we have comprehensively
To the Editor and Reviewers: We would like to thank the editor and reviewers for careful reading, and constructive suggestions for our manuscript. According to comments from both reviewers, we have comprehensively
Supplementary Information
 Supplementary Information Modelling the Yeast Interactome Vuk Janjić, Roded Sharan 2 and Nataša Pržulj, Department of Computing, Imperial College London, London, United Kingdom 2 Blavatnik School of Computer
Supplementary Information Modelling the Yeast Interactome Vuk Janjić, Roded Sharan 2 and Nataša Pržulj, Department of Computing, Imperial College London, London, United Kingdom 2 Blavatnik School of Computer
Problem-solving Test: The Mechanism of Protein Synthesis
 Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Predicting Breast Cancer Survivability Rates
 Predicting Breast Cancer Survivability Rates For data collected from Saudi Arabia Registries Ghofran Othoum 1 and Wadee Al-Halabi 2 1 Computer Science, Effat University, Jeddah, Saudi Arabia 2 Computer
Predicting Breast Cancer Survivability Rates For data collected from Saudi Arabia Registries Ghofran Othoum 1 and Wadee Al-Halabi 2 1 Computer Science, Effat University, Jeddah, Saudi Arabia 2 Computer
Overview: The immune responses of animals can be divided into innate immunity and acquired immunity.
 GUIDED READING - Ch. 43 - THE IMMUNE SYSTEM NAME: Please print out these pages and HANDWRITE the answers directly on the printouts. Typed work or answers on separate sheets of paper will not be accepted.
GUIDED READING - Ch. 43 - THE IMMUNE SYSTEM NAME: Please print out these pages and HANDWRITE the answers directly on the printouts. Typed work or answers on separate sheets of paper will not be accepted.
TWO HANDED SIGN LANGUAGE RECOGNITION SYSTEM USING IMAGE PROCESSING
 134 TWO HANDED SIGN LANGUAGE RECOGNITION SYSTEM USING IMAGE PROCESSING H.F.S.M.Fonseka 1, J.T.Jonathan 2, P.Sabeshan 3 and M.B.Dissanayaka 4 1 Department of Electrical And Electronic Engineering, Faculty
134 TWO HANDED SIGN LANGUAGE RECOGNITION SYSTEM USING IMAGE PROCESSING H.F.S.M.Fonseka 1, J.T.Jonathan 2, P.Sabeshan 3 and M.B.Dissanayaka 4 1 Department of Electrical And Electronic Engineering, Faculty
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project
 Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Downloaded by on April 28, 2018 https://pubs.acs.org Publication Date: April 24, 1984 doi: /bk
 1 Virus-Receptor Interactions BERNARD N. FIELDS Department of Microbiology and Molecular Genetics, Harvard Medical School, and Department of Medicine (Infectious Disease), Brigham and Women's Hospital,
1 Virus-Receptor Interactions BERNARD N. FIELDS Department of Microbiology and Molecular Genetics, Harvard Medical School, and Department of Medicine (Infectious Disease), Brigham and Women's Hospital,
Antigenic Epitope Prediction towards SARS Corona Virus using Bioinformatics Analysis *1
 Antigenic Epitope Prediction towards SARS Corona Virus using Bioinformatics Analysis *1 Sunitha Panigrahi, 2 Nikkitha Umesh, Deaprtment of Biotechnology, St. Mary s College, Hyderabad * Email ID: sritha17@yahoo.com
Antigenic Epitope Prediction towards SARS Corona Virus using Bioinformatics Analysis *1 Sunitha Panigrahi, 2 Nikkitha Umesh, Deaprtment of Biotechnology, St. Mary s College, Hyderabad * Email ID: sritha17@yahoo.com
CONVENTIONAL VACCINE DEVELOPMENT
 CONVENTIONAL VACCINE DEVELOPMENT PROBLEM Lethal germ Dead mouse LIVE VACCINES Related but harmless germ gives protection against lethal pathogen. Examples are the original pox vaccine and some TB vaccines
CONVENTIONAL VACCINE DEVELOPMENT PROBLEM Lethal germ Dead mouse LIVE VACCINES Related but harmless germ gives protection against lethal pathogen. Examples are the original pox vaccine and some TB vaccines
Workshop on Analysis and prediction of contacts in proteins
 Workshop on Analysis and prediction of contacts in proteins 1.09.09 Eran Eyal 1, Vladimir Potapov 2, Ronen Levy 3, Vladimir Sobolev 3 and Marvin Edelman 3 1 Sheba Medical Center, Ramat Gan, Israel; Departments
Workshop on Analysis and prediction of contacts in proteins 1.09.09 Eran Eyal 1, Vladimir Potapov 2, Ronen Levy 3, Vladimir Sobolev 3 and Marvin Edelman 3 1 Sheba Medical Center, Ramat Gan, Israel; Departments
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Epitope discovery and Rational vaccine design Morten Nielsen
 Epitope discovery and Rational vaccine design Morten Nielsen mniel@cbs.dtu.dk Bioinformatics in a nutshell List of peptides that have a given biological feature Mathematical model (neural network, hidden
Epitope discovery and Rational vaccine design Morten Nielsen mniel@cbs.dtu.dk Bioinformatics in a nutshell List of peptides that have a given biological feature Mathematical model (neural network, hidden
Tumor responses (patients responding/ patients treated)
 Table 1. ACT clinical trial tumor responses and toxicities. a Target antigen Cancer(s) Receptor type Tumor responses (patients responding/ patients treated) Immune-mediated toxicities (patients experiencing
Table 1. ACT clinical trial tumor responses and toxicities. a Target antigen Cancer(s) Receptor type Tumor responses (patients responding/ patients treated) Immune-mediated toxicities (patients experiencing
Development of the Web Users Self Efficacy scale (WUSE)
 Development of the Web Users Self Efficacy scale (WUSE) Eachus, P and Cassidy, SF Title Authors Type URL Published Date 2004 Development of the Web Users Self Efficacy scale (WUSE) Eachus, P and Cassidy,
Development of the Web Users Self Efficacy scale (WUSE) Eachus, P and Cassidy, SF Title Authors Type URL Published Date 2004 Development of the Web Users Self Efficacy scale (WUSE) Eachus, P and Cassidy,
Integrated Analysis of Copy Number and Gene Expression
 Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
