Nature Biotechnology: doi: /nbt Supplementary Figure 1. Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity
|
|
|
- Candice Gallagher
- 6 years ago
- Views:
Transcription
1 Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode) compared to fluorescent labeled MHC multimers (-barcode). DNA-barcoded MHC multimer reagents were assembled with two identical DNA barcodes attached to each multimerization backbone and subsequent co-attachment of pmhc molecules. Non-barcoded multimers were generated similarly without prior attachment of DNA barcodes (i.e. -barcode and +barcode were both assembled on a dextramer backbone). Reagents assembled with HLA-A*0201 CMV pp65 NLV were applied for staining of (a) healthy donor PBMCs, and reagents assembled with HLA-A*0201 htert p988 (YLQVNSLQTV) were applied for staining of (b) expanded TILs from a melanoma patient. The binding capacity is evaluated in terms of the stain index of the multimer fluorescent intensity of T cells stained with non-barcoded or DNA-
2 barcoded MHC multimers respectively, along with the frequencies of the given multimer positive cell population of CD8 T cells, (a) % and (b) 3.8%-4.3%. Bar plots show mean stain index values of three stainings ± SD. (c) Fluorescent-based analysis of antigen specific T cells stained with relevant virus pmhc multimers and excess of irrelevant pmhc multimers. An equimolar (1:1) mixture of individually barcode labeled HLA-B*0702 CMV pp65 TPR -multimers and HLA-A*0201 HIV Pol ILK -multimers, or a mixture with 998 additional fluorescent labeled pmhc multimers, were used for staining of healthy donor PBMCs. The 998 additional MHC multimers comprised equal amounts of irrelevant-peptide HLA-A*0201 and HLA-B*0702 multimers, i.e. multimers carrying MHC molecules refolded with UV-sensitive ligand (Online Methods). Using either reagent mixture, the concentrations of each equimolar pmhc were 23 nm in the final staining volume, i.e. for staining with the 1:1:998 equimolar reagents, the volume of the MHC multimer pool were reduced 50x. Percentages of the multimer positive population of CD8 T cells are given in dot plots. (d) The multimer positive populations from (c) were sorted by FACS and DNA barcodes associated with the sorted cell population were subjected to qpcr with fluorescent reporter probes targeting each individual DNA barcode. The experiments were performed with reagent mixtures with DNA barcodes inverted between the CMV and HIV pmhc multimers (indicated in orange and purple respectively). Cross threshold (Ct) values of DNA barcodes recovered from qpcr with approximately 200 and 600 cells in separate reactions (derived from staining with 1:1 and 1:1:998 reagent mixtures respectively) are shown in bar plots (mean±range of duplicate samples). Hashtag indicate that a given barcode was not detected.
3 Supplementary Figure 2 Dynamic range and detection limit using a panel of >1000 DNA-barcoded pmhc multimers (a) Heatmap representing the pmhc multimer analysis of duplicates of seven samples with various proportions of HLA-B*0702 CMV TPR-specific T cells: 5%, 1%, 0.2%, 0.04%, 0.008%, % and % of CD8 T cells. The 5% sample corresponds to 100%
4 BC260 PBMCs. Each sample was screened with a panel of 1031 pmhc multimers, all carrying individual DNA barcodes. The heatmap shows changes in read proportions compared to background levels, as in Figure 2b. Each column represents the reads associated with a given sample. Epitopes are grouped based on their antigen origin. Within each antigen group, rows are sorted based on Log2FC, highest to lowest, compared to baseline samples. Orange-red coloring in the heatmap represents a statistically significant number of DNA barcode reads, FDR < 0.1%, defined as antigen-specific T cell responses. The gray scaling indicates non-significant number of barcode reads, i.e. any antigen-responsive T cells associated with such barcode would be present in numbers too low to discern from background. Duplicate samples are grouped side-by-side, indicated with a. and b. respectively. (b) Magnification of the panel of virusderived peptides. Rows representing antigen specificities are sorted based on Log2FC and duplicate samples a grouped as in (a). The first row represents the target 5% titrated specificity, B*0702 CMV TPR. The rows below are two T cell responses present in the HLA- B*0702 negative PBMC sample (BC262), followed by at least two lower-frequent responses that are present in the 5% donor (BC260). Dark gray scaling, i.e. barcode reads that that are non-significant but with Log2FC>1, may represent T cell responses just below the detection limit. All pmhc multimers are dextramers.
5
6 Supplementary Figure 3 Comparing the dynamic range and detection limit using panels of 110 and 1031 DNA-barcoded pmhc multimers (a) Heatmap representing the pmhc multimer analysis of duplicates of seven samples with various proportions of HLA-B*0702 CMV TPR -specific T cells: 5%, 1%, 0.2%, 0.04%, 0.008%, % and % of CD8 T cells. The 5% sample corresponds to 100% BC260 PBMCs. Each sample was screened with a panel of 110 pmhc multimers, all carrying individual DNA barcodes. The heatmap is organized as in Figure 2b, each column represents one donor. Duplicate samples are grouped side-by-side, indicated with a. and b. respectively. (b) Magnification of the panel of virus-derived peptides. Rows representing antigen specificities are sorted based on Log2FC and duplicate samples a grouped as in (a). The first row represents the target 5% titrated specificity, B*0702 CMV TPR. The rows below are T cell responses present in the HLA-B*0702 negative PBMC sample (BC262) or lower-frequent responses present in the 5% donor (BC260). Dark gray scaling, i.e. barcode reads that are non-significant but with Log2FC>1, may represent T cell responses just below the detection limit. (c) Correlations between the frequency of HLA-B*0702 CMV TPR -specific T cells determined by analyzing the same samples using combinatorial fluorescent labeled pmhc multimers or a panel of 110 DNA barcode labeled pmhc multimers. (d, e) Correlation between results obtained from screening the same samples of 2x10 6 PBMCs with the panel of 1031 or 110 DNA-barcoded pmhc multimers represented as (d) the estimated frequency of HLA-B*0702 CMV TPR -specific T cells or (e) the number of clonally reduced barcode reads associated with this pmhc multimer. Error bars represent range of duplicates (SD). The accumulated number of non-reduced read counts that mapped to any of the DNA barcodes among the 14 samples screened were 2.7x10 6 and 1.4 x10 6 reads for the 110 and 1031 pmhc multimer library respectively. All pmhc multimers are dextramers.
7 Supplementary Figure 4 T cell reactivity in healthy donor samples analyzed by a panel of 110 DNA-barcoded pmhc multimers (a) Analysis of PBMCs (1-2x10 6 ) from six different healthy donors using 110 pmhc multimers each carrying individual barcodes. The heatmap is organized as in figure 2b, each column represents one donor. Epitopes are grouped based on their antigen origin. Significant responses are shown in orange-red colors. Significance was defined as FDR<5% since the number of reads within a given sample were compared with only one baseline sample (opposed to three baseline samples in other experiments). (b) Magnification of the panel of virus-derived peptides (26 epitopes). Rows representing antigen specificities are grouped according to HLA-type and sorted within each group based on Log2FC. HLA types of donors can be seen in Supplementary Table 6. (c) Correlations between the
8 frequencies of antigen-specific T cells determined by analyzing the same samples with combinatorially encoded fluorescently labeled pmhc multimers or with 110 DNA-barcoded pmhc multimers. Each dot represents one specificity. T cell populations with FDR<5% in DNA-barcode MHC multimer analysis or 10 events and >0.002% of CD8 T cells in combinatorial encoding analysis were plotted. All specificities included in the plot were tested using both a combinatorial encoding analysis and DNA-barcoded MHC multimers. Dots plotted on the axes are nonsignificant for one of the methods. (d, e) Correlation between results obtained from screening the same samples with the panel of 1031 or 110 DNA-barcoded pmhc multimers represented as (d) the estimated frequency of antigen-specific T cells or (e) the number of clonally reduced barcode reads associated with the given pmhc multimers. Each dot represents one specificity. Only T cell populations that fulfilled the significance criteria for DNA barcode assessment (FDR<0.1% and 5% for the 1031 and 110 library, respectively) in at least one of the analyses were plotted. Dots plotted on the axes are nonsignificant for one of the library screenings. The accumulated number of non-reduced read counts that mapped to any of the DNA barcodes among the six screened samples were 2.6x10 5 and 4.3x10 5 reads for the 110 and 1031 pmhc multimer library respectively.
9 Supplementary Figure 5 T cell reactivity assessed independently from fluorescent-based separation of MHC multimer binding T cells (a) PBMCs (2x10 6 ) from one healthy donor was stained with varying amounts of DNA barcoded-pmhc multimers, i.e. a titration from 23 nm to nm in respect to each pmhc as indicated above each dot plot. This corresponds to 100%-0.016% of the amount used elsewhere in this study. Samples were stained in duplicates and either the full CD8 population (all cells in the dot plots) or only the MHC multimer positive populations (cells indicated in black) were sorted. (b) Bar plot representing the distribution of DNA barcodes in the isolated cells from (a). Left side is based on the full CD8 population, right side is based on the MHC multimer positive population. Each bar represents the -Log10(p) value in respect to the pmhc associated DNA barcode. Dotted line at y=3 (-Log10(0.001)) represent the
10 threshold of FDR < 0.1%. The donor BC035 is HLA-A*0101 and B*0801 positive and A*0201 negative (see full HLA-type in Supplementary Table 6). BC035 was previously found to carry antigen-specific T cells restricted to HLA-A*0101, CMV pp50 VTE (2.2% of CD8 T cells) and CMV pp65 YSE (0.6% of CD8 T cells). HLA- A*0201 restricted epitopes functions as negative controls. Irrespectively of sorting the full CD8 or only the multimer positive population the same responses are detected after sequencing of the DNA barcodes. When the MHC multimer binding T cells could not be separated from the full CD8 T cell population based on their fluorescent intensity, i.e. when applying the lowest amount of MHC multimer reagent, the DNA barcodes associated with positive control reagents were still recovered after sorting the full CD8 population indicating that T cell reactivity can be assessed independent on fluorescent separation of the MHC multimer binding T cells.
11 Supplementary Table 1. Oligo As. Related to Figure 1 Blue, green, black and turquoise coloring indicates primer region, UMI, 25mer barcode region and annealing region respectively. Oligo A A1 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNCGAGGGCAATGGTTAACTGACACGTGGTCAGCA A2 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNCAGAAAGCAGTCTCGTCGGTTCGAAGGTCAGCA A3 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNTAAGTAGCGGGCATAATGTACGCTCGGTCAGCA A4 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNGGATCCAGTAAGCTACTGCGTTTATGGTCAGCA A5 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNGGGCTGCGGAGCGTTTACTCTGTATGGTCAGCA A6 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNAAACGTATGTGCTTTGTCGGATGCCGGTCAGCA A7 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNATATCATCATAGGCTTAGCGACGTAGGTCAGCA A8 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNAGGAAAATCTGCTACCGCCAATGATGGTCAGCA A9 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNCTGATTGACTGCATGGAGGCTATACGGTCAGCA A10 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNGTGGCGACTTCACGATTATCTGAACGGTCAGCA A11 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNCCTGTATTGAAGGTTCAGTCCTGTTGGTCAGCAT CATTTCC A12 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNGGCTCTATAAGGTTTCCTCAAAGGTGGTCAGCA A13 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNTTGGGAGCTTTCCTATGTACAGTCCGGTCAGCA A14 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNAGAGAATATGTCGCTCCCGTTATGTGGTCAGCA A15 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNGCAGTTAGATATGCAGTTACCTGACGGTCAGCA A16 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNCTTCACCCGAACATGCAGTGTTATTGGTCAGCA A17 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNAAAGCCGTTGCAGTATCGTCTGAGCGGTCAGCA A18 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNGCTGGATGTTAATAACTGCGGTCCGGGTCAGCA A19 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNACGAGTTGACATGGACGGATCCCTCGGTCAGCA A20 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNTTCATCACTCATTGTTCTGAGTAGGGGTCAGCAT CATTTCC A21 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNATGTTTAATCTAACTTGATGCCTCCGGTCAGCAT CATTTCC A22 BiotinGAAGTTCCAGCCAGCGTCACAGTTTNNNNNNTAATACGCCTGAGGTGTTGGGTTGCGGTCAGCA
12 Supplementary Table 2. Oligo Bs. Related to Figure 1 Blue, green, black and turquoise coloring indicates primer region, UMI, 25mer barcode region and annealing region respectively. See enclosed excel file Supplementary Table 3. Forward primers with Sample-identification barcodes (SampleID) and Ion Torrent adaptor (A-Key) and reverse primer with Ion Torrent adaptor (P1-Key). Related to Figure 1 Black and blue coloring indicates primer regions and sampl ID, respectively. Purple and grey coloring indicates the adaptor regions, and the red nucleotides are linkers between the adaptor and the sample ID. See enclosed excel file Supplementary Table 4. Example of calculation of estimated frequency from barcode reads and comparison with p- value and LogFC. Related to Figures 2-4. BC254 Total clonality reduced readcounts mapped to SampleID Frequency of total multimer + cells of CD8 + (%) 3.63 Epitope Clonality reduced read counts Estimated frequency (%) HLA-A*0201 CMV pp65 NLV a 7.22E HLA-A*0201 EBV LMP2 CLG E HLA-A*0201 EBV LMP2 FLY E HLA-A*0201 EBV BMF1 GLC E HLA-A*0201 EBV BRLF1 YVL E HLA-A*0201 FLU MP GIL E MM01 Total clonality reduced readcounts mapped to SampleID Frequency of total multimer + cells of CD8 + (%) Epitope Clonality reduced read counts Estimated frequency (%) HLA-A*0201 gp100 KTW b 4.85E HLA-A*0201 gp100 AML E HLA-A*0201 MART-1 ELA E HLA-A*0201 tyrosinase CLL E HLA-A*0201 gp100 ITD E HLA-A*0201 MART-1 EAA E HLA-A*0201 gp100 IMD E a Specific calculation of estimated frequency: (16441 reads/33939 reads) x 3.63 % = 1.8 % b Specific calculation of estimated frequency: (4814 specific reads/10145 total reads) x % =12.0 % p-val p-val logfc logfc Supplementary Table barcode-labeled MHC multimer panel. Related to Figure 2, Figure 3, Supplementary Figure 2 and Supplementary Figure 3 See enclosed excel file
13 Supplementary Table 6. HLA-types of healthy donors and melanoma patients. Related to Figures 2-7 and Supplementary Figures 2-5 Healthy donor HLA-A* HLA-B* BC BC BC BC BC BC BC BC BC BC BC BC262 a, b N.A. N.A. BC BC Melanoma patient MM MM02 c MM03 a 02 N.A. N.A. N.A. MM04 a 02 N.A. N.A. N.A. MM05 a 02 N.A. N.A. N.A. MM06 c MM07 a 02 N.A. N.A. N.A. MM08 a 02 N.A. N.A. N.A. MM MM MM NSCLC patient L L N.A. = Not analyzed. a HLA-A tissue typed with PCR. b Tissue typed HLA-B*0702 negative by flow cytometry. c MM02 and MM06 are derived from tumor material taken at six months interval from the same patient.
14 Supplementary Table barcode-labeled MHC multimer panel. Related to Supplementary Figure 2 and 3. See enclosed excel file Supplementary Table barcode-labeled MHC multimer panel. Related to Figure 4 and Figure 5. See enclosed excel file Supplementary Table barcode-labeled MHC multimer panel. Related to Figure 7. See enclosed excel file
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
MHC MULTIMER PROFICIENCY PANEL 2017
 MHC MULTIMER PROFICIENCY PANEL 2017 August 2017 CONTACT Charlotte Halgreen ProficiencyPanel@immudex.com FOR MORE INFORMATION www.proficiencypanel.com MHC MULTIMER PROFICIENCY PANEL 2017 This report summarizes
MHC MULTIMER PROFICIENCY PANEL 2017 August 2017 CONTACT Charlotte Halgreen ProficiencyPanel@immudex.com FOR MORE INFORMATION www.proficiencypanel.com MHC MULTIMER PROFICIENCY PANEL 2017 This report summarizes
MHC MULTIMER PROFICIENCY PANEL 2015
 MHC MULTIMER PROFICIENCY PANEL 2015 July 2015 CONTACT Charlotte Halgreen ProficiencyPanel@immudex.com FOR MORE INFORMATION www.proficiencypanel.com MHC MULTIMER PROFICIENCY PANEL 2015 This report summarizes
MHC MULTIMER PROFICIENCY PANEL 2015 July 2015 CONTACT Charlotte Halgreen ProficiencyPanel@immudex.com FOR MORE INFORMATION www.proficiencypanel.com MHC MULTIMER PROFICIENCY PANEL 2015 This report summarizes
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in
 Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Low Avidity CMV + T Cells accumulate in Old Humans
 Supplementary Figure Legends Supplementary Figure 1. CD45RA expressing CMVpp65-specific T cell populations accumulate within HLA-A*0201 and HLA-B*0701 individuals Pooled data showing the size of the NLV/HLA-A*0201-specific
Supplementary Figure Legends Supplementary Figure 1. CD45RA expressing CMVpp65-specific T cell populations accumulate within HLA-A*0201 and HLA-B*0701 individuals Pooled data showing the size of the NLV/HLA-A*0201-specific
HD1 (FLU) HD2 (EBV) HD2 (FLU)
 ramer staining + anti-pe beads ramer staining a HD1 (FLU) HD2 (EBV) HD2 (FLU).73.11.56.46.24 1.12 b CD127 + c CD127 + d CD127 - e CD127 - PD1 - PD1 + PD1 + PD1-1 1 1 1 %CD127 + PD1-8 6 4 2 + anti-pe %CD127
ramer staining + anti-pe beads ramer staining a HD1 (FLU) HD2 (EBV) HD2 (FLU).73.11.56.46.24 1.12 b CD127 + c CD127 + d CD127 - e CD127 - PD1 - PD1 + PD1 + PD1-1 1 1 1 %CD127 + PD1-8 6 4 2 + anti-pe %CD127
Dissecting therapy-induced T-cell responses in melanoma
 Dissecting therapy-induced T-cell responses in melanoma Tumor-infiltrating lymphocyte (TIL) therapy of melanoma TIL are grown from melanoma tumors Rapid Expansion Infusion of TIL + IL-2 Patient pretreated
Dissecting therapy-induced T-cell responses in melanoma Tumor-infiltrating lymphocyte (TIL) therapy of melanoma TIL are grown from melanoma tumors Rapid Expansion Infusion of TIL + IL-2 Patient pretreated
Supplemental materials
 Supplemental materials 1 Supplemental Fig. 1 Immunogram This immunogram summarizes patient clinical data and immune parameters at corresponding time points for Patient IMF-32. The top panel illustrates
Supplemental materials 1 Supplemental Fig. 1 Immunogram This immunogram summarizes patient clinical data and immune parameters at corresponding time points for Patient IMF-32. The top panel illustrates
Supplementary Fig. 1: Ex vivo tetramer enrichment with anti-c-myc beads
 Supplementary Fig. 1: Ex vivo tetramer enrichment with anti-c-myc beads Representative example of comparative ex vivo tetramer enrichment performed in three independent experiments with either conventional
Supplementary Fig. 1: Ex vivo tetramer enrichment with anti-c-myc beads Representative example of comparative ex vivo tetramer enrichment performed in three independent experiments with either conventional
T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting
 T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting Julia Ekeruche-Makinde 1*, Mathew Clement 1*, David K Cole 1*, Emily S J Edwards 1, Kristin Ladell 1, John
T Cell Receptor Optimized Peptide Skewing of the T-Cell Repertoire Can Enhance Antigen Targeting Julia Ekeruche-Makinde 1*, Mathew Clement 1*, David K Cole 1*, Emily S J Edwards 1, Kristin Ladell 1, John
Supplementary Figure 1
 Supplementary Figure 1 Identification of IFN-γ-producing CD8 + and CD4 + T cells with naive phenotype by alternative gating and sample-processing strategies. a. Contour 5% probability plots show definition
Supplementary Figure 1 Identification of IFN-γ-producing CD8 + and CD4 + T cells with naive phenotype by alternative gating and sample-processing strategies. a. Contour 5% probability plots show definition
Supplementary Figures
 Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Supplementary Figures Supplementary Figure 1 Characterization of stable expression of GlucB and sshbira in the CT26 cell line (a) Live cell imaging of stable CT26 cells expressing green fluorescent protein
Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to
 Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
EBV Infection and Immunity. Andrew Hislop Institute for Cancer Studies University of Birmingham
 EBV Infection and Immunity Andrew Hislop Institute for Cancer Studies University of Birmingham EBV Introduction Large ds DNA virus Spread by saliva contact Lifelong infection Predominantly B-lymphotropic
EBV Infection and Immunity Andrew Hislop Institute for Cancer Studies University of Birmingham EBV Introduction Large ds DNA virus Spread by saliva contact Lifelong infection Predominantly B-lymphotropic
New technologies for studying human immunity. Lisa Wagar Postdoctoral fellow, Mark Davis lab Stanford University School of Medicine
 New technologies for studying human immunity Lisa Wagar Postdoctoral fellow, Mark Davis lab Stanford University School of Medicine New strategies: Human immunology is ideal for a systems approach We have
New technologies for studying human immunity Lisa Wagar Postdoctoral fellow, Mark Davis lab Stanford University School of Medicine New strategies: Human immunology is ideal for a systems approach We have
Supplementary Figure 1. Enhanced detection of CTLA-4 on the surface of HIV-specific
 SUPPLEMENTARY FIGURE LEGEND Supplementary Figure 1. Enhanced detection of CTLA-4 on the surface of HIV-specific CD4 + T cells correlates with intracellular CTLA-4 levels. (a) Comparative CTLA-4 levels
SUPPLEMENTARY FIGURE LEGEND Supplementary Figure 1. Enhanced detection of CTLA-4 on the surface of HIV-specific CD4 + T cells correlates with intracellular CTLA-4 levels. (a) Comparative CTLA-4 levels
Supplementary Information
 Supplementary Information High throughput epitope discovery reveals frequent recognition of neo-antigens by CD4 + T-cells in human melanoma Carsten Linnemann, Marit M. van Buuren, Laura Bies, Els M.E.Verdegaal,
Supplementary Information High throughput epitope discovery reveals frequent recognition of neo-antigens by CD4 + T-cells in human melanoma Carsten Linnemann, Marit M. van Buuren, Laura Bies, Els M.E.Verdegaal,
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
Supplementary Figures Supplementary Figure 1. Confirmation of Dnmt1 conditional knockout out mice. a, Representative images of sorted stem (Lin - CD49f high CD24 + ), luminal (Lin - CD49f low CD24 + )
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Nature Immunology: doi: /ni Supplementary Figure 1. RNA-Seq analysis of CD8 + TILs and N-TILs.
 Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Nature Immunology: doi: /ni Supplementary Figure 1
 Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Supplementary Figure 1 A β-strand positions consistently places the residues at CDR3β P6 and P7 within human and mouse TCR-peptide-MHC interfaces. (a) E8 TCR containing V β 13*06 carrying with an 11mer
Supporting Information
 Supporting Information Sui et al..7/pnas.997 Pre-CLP CM9 LA9 SL Tat# Pol Vif % Tetramer + CD + CD + Vac+IL- +IL- Vac Fig. S. Frequencies of six different CD + CD + Mamu-A*-tetramer + cells were measured
Supporting Information Sui et al..7/pnas.997 Pre-CLP CM9 LA9 SL Tat# Pol Vif % Tetramer + CD + CD + Vac+IL- +IL- Vac Fig. S. Frequencies of six different CD + CD + Mamu-A*-tetramer + cells were measured
10/18/2012. A primer in HLA: The who, what, how and why. What?
 A primer in HLA: The who, what, how and why What? 1 First recognized in mice during 1930 s and 1940 s. Mouse (murine) experiments with tumors Independent observations were made in humans with leukoagglutinating
A primer in HLA: The who, what, how and why What? 1 First recognized in mice during 1930 s and 1940 s. Mouse (murine) experiments with tumors Independent observations were made in humans with leukoagglutinating
Supplemental Figures Supplemental Figure 1:
 Supplemental Figures Supplemental Figure 1: Representative FACS data showing Concurrent Brain cell type Acquisition using either Percoll PLUS (top row) or myelin removal beads (bottom two rows). Debris
Supplemental Figures Supplemental Figure 1: Representative FACS data showing Concurrent Brain cell type Acquisition using either Percoll PLUS (top row) or myelin removal beads (bottom two rows). Debris
NC bp. b 1481 bp
 Kcna3 NC 11 178 p 1481 p 346 p c *** *** d relative expression..4.3.2.1 CD4 T cells Kcna1 Kcna2 Kcna3 Kcna4 Kcna Kcna6 Kcna7 Kcnn4 relative expression..4.3.2.1 CD8 T cells Kcna1 Kcna2 Kcna3 Kcna4 Kcna
Kcna3 NC 11 178 p 1481 p 346 p c *** *** d relative expression..4.3.2.1 CD4 T cells Kcna1 Kcna2 Kcna3 Kcna4 Kcna Kcna6 Kcna7 Kcnn4 relative expression..4.3.2.1 CD8 T cells Kcna1 Kcna2 Kcna3 Kcna4 Kcna
Immunocore Ltd isbtc Washington 30 th October 2009
 Ltd isbtc Washington 30 th October 2009 Bent Jakobsen Chief Scientific Officer Breaking immune tolerance to cancer The immune system has more cytotoxic potential and versatility than can be achieved with
Ltd isbtc Washington 30 th October 2009 Bent Jakobsen Chief Scientific Officer Breaking immune tolerance to cancer The immune system has more cytotoxic potential and versatility than can be achieved with
Supplementary Figure 1. Example of gating strategy
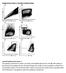 Supplementary Figure 1. Example of gating strategy Legend Supplementary Figure 1: First, gating is performed to include only single cells (singlets) (A) and CD3+ cells (B). After gating on the lymphocyte
Supplementary Figure 1. Example of gating strategy Legend Supplementary Figure 1: First, gating is performed to include only single cells (singlets) (A) and CD3+ cells (B). After gating on the lymphocyte
SUPPLEMENTARY INFORMATION
 Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
Our T cell epitope test service offers you:
 Our T cell epitope test service offers you: high throughput peptide discovery fast lead-finding maximum of epitopes in a minimum of blood saving effort and costs on developing your own platforms custom-made
Our T cell epitope test service offers you: high throughput peptide discovery fast lead-finding maximum of epitopes in a minimum of blood saving effort and costs on developing your own platforms custom-made
Cytotoxicity assays. Rory D. de Vries, PhD 1. Viroscience lab, Erasmus MC, Rotterdam, the Netherlands
 Cytotoxicity assays Rory D. de Vries, PhD 1 1 Viroscience lab, Erasmus MC, Rotterdam, the Netherlands Anti-influenza immunity Humoral / CD4+ / CD8+ / NK? Function of CTL Elimination of virus-infected cells?
Cytotoxicity assays Rory D. de Vries, PhD 1 1 Viroscience lab, Erasmus MC, Rotterdam, the Netherlands Anti-influenza immunity Humoral / CD4+ / CD8+ / NK? Function of CTL Elimination of virus-infected cells?
Surface plasmon resonance (SPR) analysis
 Surface plasmon resonance (SPR) analysis Soluble CD8αα and was manufactured as described previously. 1,2 The C12C heterodimeric TCR was produced using an engineered disulfide bridge between the constant
Surface plasmon resonance (SPR) analysis Soluble CD8αα and was manufactured as described previously. 1,2 The C12C heterodimeric TCR was produced using an engineered disulfide bridge between the constant
Supplementary Information. Novel lentiviral vectors with mutated reverse transcriptase for mrna delivery of TALE nucleases
 Supplementary Information Novel lentiviral vectors with mutated reverse transcriptase for mrna delivery of TALE nucleases Ulrike Mock 1, Kristoffer Riecken 1, Belinda Berdien 1, Waseem Qasim 2, Emma Chan
Supplementary Information Novel lentiviral vectors with mutated reverse transcriptase for mrna delivery of TALE nucleases Ulrike Mock 1, Kristoffer Riecken 1, Belinda Berdien 1, Waseem Qasim 2, Emma Chan
SUPPLEMENTARY INFORMATION
 DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
DOI:.38/ncb3399 a b c d FSP DAPI 5mm mm 5mm 5mm e Correspond to melanoma in-situ Figure a DCT FSP- f MITF mm mm MlanaA melanoma in-situ DCT 5mm FSP- mm mm mm mm mm g melanoma in-situ MITF MlanaA mm mm
08/02/59. Tumor Immunotherapy. Development of Tumor Vaccines. Types of Tumor Vaccines. Immunotherapy w/ Cytokine Gene-Transfected Tumor Cells
 Tumor Immunotherapy Autologous virus Inactivation Inactivated virus Lymphopheresis Culture? Monocyte s Dendritic cells Immunization Autologous vaccine Development of Tumor Vaccines Types of Tumor Vaccines
Tumor Immunotherapy Autologous virus Inactivation Inactivated virus Lymphopheresis Culture? Monocyte s Dendritic cells Immunization Autologous vaccine Development of Tumor Vaccines Types of Tumor Vaccines
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary information. Early development of broad neutralizing antibodies in HIV-1 infected infants
 Supplementary information Early development of broad neutralizing antibodies in HIV-1 infected infants Leslie Goo, Vrasha Chohan, Ruth Nduati, Julie Overbaugh Supplementary Figure 1. Neutralization profile
Supplementary information Early development of broad neutralizing antibodies in HIV-1 infected infants Leslie Goo, Vrasha Chohan, Ruth Nduati, Julie Overbaugh Supplementary Figure 1. Neutralization profile
Nature Immunology: doi: /ni Supplementary Figure 1. Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice.
 Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Supplementary Figure 1 Gene expression profile of CD4 + T cells and CTL responses in Bcl6-deficient mice. (a) Gene expression profile in the resting CD4 + T cells were analyzed by an Affymetrix microarray
Herpesviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Herpesviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped icosahedral capsid (T=16), diameter 125 nm Diameter of enveloped virion 200 nm Capsid
Herpesviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped icosahedral capsid (T=16), diameter 125 nm Diameter of enveloped virion 200 nm Capsid
TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer
 AD Award Number: W8-XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke,. CONTRACTING ORGANIZATION: Johns Hopkins
AD Award Number: W8-XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke,. CONTRACTING ORGANIZATION: Johns Hopkins
CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads (Miltenyi Biotec) for depletion of CD25 +
 Supplements Supplemental Materials and Methods Depletion of CD25 + T-cells from PBMC. Fresh or HD precultured PBMC were stained with the conjugate CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads
Supplements Supplemental Materials and Methods Depletion of CD25 + T-cells from PBMC. Fresh or HD precultured PBMC were stained with the conjugate CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads
Supplementary Figure 1 a
 Supplementary Figure 1 a b d c Supplementary Figure 1. Poor sensitivity for mutation detection using established single-cell RNA-sequencing methods. (a) Representative bioanalyzer trace showing expected
Supplementary Figure 1 a b d c Supplementary Figure 1. Poor sensitivity for mutation detection using established single-cell RNA-sequencing methods. (a) Representative bioanalyzer trace showing expected
Human Immunodeficiency Virus Type-1 Myeloid Derived Suppressor Cells Inhibit Cytomegalovirus Inflammation through Interleukin-27 and B7-H4
 Human Immunodeficiency Virus Type-1 Myeloid Derived Suppressor Cells Inhibit Cytomegalovirus Inflammation through Interleukin-27 and B7-H4 Ankita Garg, Rodney Trout and Stephen A. Spector,,* Department
Human Immunodeficiency Virus Type-1 Myeloid Derived Suppressor Cells Inhibit Cytomegalovirus Inflammation through Interleukin-27 and B7-H4 Ankita Garg, Rodney Trout and Stephen A. Spector,,* Department
Supplementary Materials for
 www.sciencemag.org/content/348/6241/aaa825/suppl/dc1 Supplementary Materials for A mucosal vaccine against Chlamydia trachomatis generates two waves of protective memory T cells Georg Stary,* Andrew Olive,
www.sciencemag.org/content/348/6241/aaa825/suppl/dc1 Supplementary Materials for A mucosal vaccine against Chlamydia trachomatis generates two waves of protective memory T cells Georg Stary,* Andrew Olive,
Crystallization-grade After D After V3 cocktail. Time (s) Time (s) Time (s) Time (s) Time (s) Time (s)
 Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Supplementary Table 1 Clinicopathological characteristics of 35 patients with CRCs
 Supplementary Table Clinicopathological characteristics of 35 patients with CRCs Characteristics Type-A CRC Type-B CRC P value Sex Male / Female 9 / / 8.5 Age (years) Median (range) 6. (9 86) 6.5 (9 76).95
Supplementary Table Clinicopathological characteristics of 35 patients with CRCs Characteristics Type-A CRC Type-B CRC P value Sex Male / Female 9 / / 8.5 Age (years) Median (range) 6. (9 86) 6.5 (9 76).95
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region
 Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplementary information
 Supplementary information Lipid peroxidation causes endosomal antigen release for cross-presentation Ilse Dingjan 1, Daniëlle RJ Verboogen 1, Laurent M Paardekooper 1, Natalia H Revelo 1, Simone P Sittig
Supplementary information Lipid peroxidation causes endosomal antigen release for cross-presentation Ilse Dingjan 1, Daniëlle RJ Verboogen 1, Laurent M Paardekooper 1, Natalia H Revelo 1, Simone P Sittig
Direct ex vivo characterization of human antigen-specific CD154 + CD4 + T cells Rapid antigen-reactive T cell enrichment (Rapid ARTE)
 Direct ex vivo characterization of human antigen-specific CD154 + CD4 + T cells Rapid antigen-reactive T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central
Direct ex vivo characterization of human antigen-specific CD154 + CD4 + T cells Rapid antigen-reactive T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central
Why are validated immunogenicity assays important for HIV vaccine development?
 Why are validated immunogenicity assays important for HIV vaccine development? There is a need to compare immunogenicity of products in the pipeline, when similar or different in class when developed by
Why are validated immunogenicity assays important for HIV vaccine development? There is a need to compare immunogenicity of products in the pipeline, when similar or different in class when developed by
Potential cross reactions between HIV 1 specific T cells and the microbiome. Andrew McMichael Suzanne Campion
 Potential cross reactions between HIV 1 specific T cells and the microbiome Andrew McMichael Suzanne Campion Role of the Microbiome? T cell (and B cell) immune responses to HIV and Vaccines are influenced
Potential cross reactions between HIV 1 specific T cells and the microbiome Andrew McMichael Suzanne Campion Role of the Microbiome? T cell (and B cell) immune responses to HIV and Vaccines are influenced
Lecture 6. Burr BIO 4353/6345 HIV/AIDS. Tetramer staining of T cells (CTL s) Andrew McMichael seminar: Background
 Lecture 6 Burr BIO 4353/6345 HIV/AIDS Andrew McMichael seminar: Background Tetramer staining of T cells (CTL s) 1. Vβ 19: There are 52 T cell receptor (TCR) Vβ gene segments in germ line DNA (See following
Lecture 6 Burr BIO 4353/6345 HIV/AIDS Andrew McMichael seminar: Background Tetramer staining of T cells (CTL s) 1. Vβ 19: There are 52 T cell receptor (TCR) Vβ gene segments in germ line DNA (See following
10/3/2016. Immunotherapy of human cancer can be highly effective: TIL therapy. What T cells See on Human Cancer. Anti-PD-1. Anti-PD-1 and anti-ctla-4
 Immunotherapy of human cancer can be highly effective: TIL therapy Tumor-infiltrating lymphocytes (TIL) are grown from melanoma tumors Rapid Expansion Infusion of TIL + IL-2 What T cells See on Human Cancer
Immunotherapy of human cancer can be highly effective: TIL therapy Tumor-infiltrating lymphocytes (TIL) are grown from melanoma tumors Rapid Expansion Infusion of TIL + IL-2 What T cells See on Human Cancer
TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer
 AD Award Number: W8XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins
AD Award Number: W8XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins
Rapid antigen-specific T cell enrichment (Rapid ARTE)
 Direct ex vivo characterization of human antigen-specific CD154+CD4+ T cell Rapid antigen-specific T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central role
Direct ex vivo characterization of human antigen-specific CD154+CD4+ T cell Rapid antigen-specific T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central role
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/3/114/ra23/dc1 Supplementary Materials for Regulation of Zap70 Expression During Thymocyte Development Enables Temporal Separation of CD4 and CD8 Repertoire Selection
www.sciencesignaling.org/cgi/content/full/3/114/ra23/dc1 Supplementary Materials for Regulation of Zap70 Expression During Thymocyte Development Enables Temporal Separation of CD4 and CD8 Repertoire Selection
Supplementary Figure 1 Chemokine and chemokine receptor expression during muscle regeneration (a) Analysis of CR3CR1 mrna expression by real time-pcr
 Supplementary Figure 1 Chemokine and chemokine receptor expression during muscle regeneration (a) Analysis of CR3CR1 mrna expression by real time-pcr at day 0, 1, 4, 10 and 21 post- muscle injury. (b)
Supplementary Figure 1 Chemokine and chemokine receptor expression during muscle regeneration (a) Analysis of CR3CR1 mrna expression by real time-pcr at day 0, 1, 4, 10 and 21 post- muscle injury. (b)
Nature Medicine: doi: /nm.2109
 HIV 1 Infects Multipotent Progenitor Cells Causing Cell Death and Establishing Latent Cellular Reservoirs Christoph C. Carter, Adewunmi Onafuwa Nuga, Lucy A. M c Namara, James Riddell IV, Dale Bixby, Michael
HIV 1 Infects Multipotent Progenitor Cells Causing Cell Death and Establishing Latent Cellular Reservoirs Christoph C. Carter, Adewunmi Onafuwa Nuga, Lucy A. M c Namara, James Riddell IV, Dale Bixby, Michael
Trends in molecular diagnostics
 Trends in molecular diagnostics Detection of target genes of interest Quantification Infectious diseases HIV Hepatitis C & B TB / MAC Cytomegalovirus Herpes simplex Varicella zoster CT/GC HPV Profiling
Trends in molecular diagnostics Detection of target genes of interest Quantification Infectious diseases HIV Hepatitis C & B TB / MAC Cytomegalovirus Herpes simplex Varicella zoster CT/GC HPV Profiling
SUPPLEMENTARY INFORMATION. Divergent TLR7/9 signaling and type I interferon production distinguish
 SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Supplementary Table 1. Functional avidities of survivin-specific T-cell clones against LML-peptide
 Supplementary Table 1. Functional avidities of survivin-specific T-cell clones against LML-peptide pulsed T2 cells. clone avidity by 4-hour 51 Cr-release assay 50% lysis at E:T 10:1 [LML peptide, M] #24
Supplementary Table 1. Functional avidities of survivin-specific T-cell clones against LML-peptide pulsed T2 cells. clone avidity by 4-hour 51 Cr-release assay 50% lysis at E:T 10:1 [LML peptide, M] #24
TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer
 AD Award Number: W8-XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins
AD Award Number: W8-XWH-5-- TITLE: Development of Antigen Presenting Cells for adoptive immunotherapy in prostate cancer PRINCIPAL INVESTIGATOR: Mathias Oelke, Ph.D. CONTRACTING ORGANIZATION: Johns Hopkins
Supplementary Figure 1. Overview of steps in the construction of photosynthetic protocellular systems
 Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature22976 Supplementary Discussion The adaptive immune system uses a highly diverse population of T lymphocytes to selectively recognize and respond to antigenic
SUPPLEMENTARY INFORMATION doi:10.1038/nature22976 Supplementary Discussion The adaptive immune system uses a highly diverse population of T lymphocytes to selectively recognize and respond to antigenic
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Other Clinical Diagnoses
 Donor Case ID 14 year old fe npod6342 24 year old npod6367 12 year old fe npod6268 6 year old fe npod69 22 year old fe npod6323 20 year old T1D.6 27 year old T1D.7 30 year old T1D.8 22 year old T1D.9 Duration
Donor Case ID 14 year old fe npod6342 24 year old npod6367 12 year old fe npod6268 6 year old fe npod69 22 year old fe npod6323 20 year old T1D.6 27 year old T1D.7 30 year old T1D.8 22 year old T1D.9 Duration
Antigen capture and presentation to T lymphocytes
 Antigen capture and presentation to T lymphocytes What T lymphocytes see Innate Immunity Immediately available or Very broad specificity rapidly recruited Adaptive Immunity Rare and naïve cells require
Antigen capture and presentation to T lymphocytes What T lymphocytes see Innate Immunity Immediately available or Very broad specificity rapidly recruited Adaptive Immunity Rare and naïve cells require
Supporting Information
 Supporting Information Chapuis et al. 10.1073/pnas.1113748109 SI Methods Selection of Patients, Targets, Isolation, and Expansion of Melanoma- Specific CTL Clones. Patients were HLA-typed, and their tumors
Supporting Information Chapuis et al. 10.1073/pnas.1113748109 SI Methods Selection of Patients, Targets, Isolation, and Expansion of Melanoma- Specific CTL Clones. Patients were HLA-typed, and their tumors
Clonotype Selection and Composition of Human CD8 T Cells Specific for Persistent Herpes Viruses Varies with Differentiation but Is Stable Over Time
 This information is current as of July 25, 2017. References Subscription Permissions Email Alerts Clonotype Selection and Composition of Human CD8 T Cells Specific for Persistent Herpes Viruses Varies
This information is current as of July 25, 2017. References Subscription Permissions Email Alerts Clonotype Selection and Composition of Human CD8 T Cells Specific for Persistent Herpes Viruses Varies
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10134 Supplementary Figure 1. Anti-inflammatory activity of sfc. a, Autoantibody immune complexes crosslink activating Fc receptors, promoting activation of macrophages, and WWW.NATURE.COM/NATURE
doi:10.1038/nature10134 Supplementary Figure 1. Anti-inflammatory activity of sfc. a, Autoantibody immune complexes crosslink activating Fc receptors, promoting activation of macrophages, and WWW.NATURE.COM/NATURE
Modulation of Host Learning in Aedes aegypti Mosquitoes
 Current Biology, Volume 28 Supplemental Information Modulation of Host Learning in Aedes aegypti Mosquitoes Clément Vinauger, Chloé Lahondère, Gabriella H. Wolff, Lauren T. Locke, Jessica E. Liaw, Jay
Current Biology, Volume 28 Supplemental Information Modulation of Host Learning in Aedes aegypti Mosquitoes Clément Vinauger, Chloé Lahondère, Gabriella H. Wolff, Lauren T. Locke, Jessica E. Liaw, Jay
Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h)
 Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h) after feeding. A small slice (~5-1 mm 3 ) was taken
Online Appendix Material and Methods: Pancreatic RNA isolation and quantitative real-time (q)rt-pcr. Mice were fasted overnight and killed 1 hour (h) after feeding. A small slice (~5-1 mm 3 ) was taken
Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples.
 Supplementary files Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples. Table S3. Specificity of AGO1- and AGO4-preferred 24-nt
Supplementary files Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples. Table S3. Specificity of AGO1- and AGO4-preferred 24-nt
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
Supplemental Figure 1. Gating strategies for flow cytometry and intracellular cytokinestaining
 Supplemental Figure 1. Gating strategies for flow cytometry and intracellular cytokinestaining of PBMCs. Forward scatter area (FSC-A) versus side scatter area (SSC-A) was used to select lymphocytes followed
Supplemental Figure 1. Gating strategies for flow cytometry and intracellular cytokinestaining of PBMCs. Forward scatter area (FSC-A) versus side scatter area (SSC-A) was used to select lymphocytes followed
INTEGRATION OF GENERAL AMINO ACID CONTROL AND TOR REGULATORY PATHWAYS IN NITROGEN ASSIMILATION IN YEAST
 INTEGRATION OF GENERAL AMINO ACID CONTROL AND TOR REGULATORY PATHWAYS IN NITROGEN ASSIMILATION IN YEAST Kirk A. Staschke 1, Souvik Dey 1, John M. Zaborske 2, Lakshmi Reddy Palam 1, Jeanette N. McClintick
INTEGRATION OF GENERAL AMINO ACID CONTROL AND TOR REGULATORY PATHWAYS IN NITROGEN ASSIMILATION IN YEAST Kirk A. Staschke 1, Souvik Dey 1, John M. Zaborske 2, Lakshmi Reddy Palam 1, Jeanette N. McClintick
Acute lung injury in children : from viral infection and mechanical ventilation to inflammation and apoptosis Bern, R.A.
 UvA-DARE (Digital Academic Repository) Acute lung injury in children : from viral infection and mechanical ventilation to inflammation and apoptosis Bern, R.A. Link to publication Citation for published
UvA-DARE (Digital Academic Repository) Acute lung injury in children : from viral infection and mechanical ventilation to inflammation and apoptosis Bern, R.A. Link to publication Citation for published
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION TITLE: Structural Basis of Signal Sequence Surveillance and Selection by the SRP-SR Complex AUTHORS and AFFILIATIONS Ottilie von Loeffelholz 1,2, Kèvin Knoops 1,2,6, Aileen Ariosa
SUPPLEMENTARY INFORMATION TITLE: Structural Basis of Signal Sequence Surveillance and Selection by the SRP-SR Complex AUTHORS and AFFILIATIONS Ottilie von Loeffelholz 1,2, Kèvin Knoops 1,2,6, Aileen Ariosa
Supplementary figure 1
 Supplementary figure 1 Nature Medicine: doi:1.138/nm.275 CLUSTER BY SELF-ORGANIZING MAPS SELECTED PATHWAY ANALISYS TERMS Cluster : up-regulated genes in acute patients Cell cycle/dna repair Fatty acid
Supplementary figure 1 Nature Medicine: doi:1.138/nm.275 CLUSTER BY SELF-ORGANIZING MAPS SELECTED PATHWAY ANALISYS TERMS Cluster : up-regulated genes in acute patients Cell cycle/dna repair Fatty acid
Mouse Clec9a ORF sequence
 Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Mouse Clec9a gene LOCUS NC_72 13843 bp DNA linear CON 1-JUL-27 DEFINITION Mus musculus chromosome 6, reference assembly (C57BL/6J). ACCESSION NC_72 REGION: 129358881-129372723 Mouse Clec9a ORF sequence
Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary Materials and Methods
 Silva et al. PTEN posttranslational inactivation and hyperactivation of the PI3K/Akt pathway sustain primary T cell leukemia viability Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary
Silva et al. PTEN posttranslational inactivation and hyperactivation of the PI3K/Akt pathway sustain primary T cell leukemia viability Supplementary Table; Supplementary Figures and legends S1-S21; Supplementary
CONTRACTING ORGANIZATION: Johns Hopkins University School of Medicine Baltimore, MD 21205
 AD Award Number: DAMD7---7 TITLE: Development of Artificial Antigen Presenting Cells for Prostate Cancer Immunotherapy PRINCIPAL INVESTIGATOR: Jonathan P. Schneck, M.D., Ph.D. Mathias Oelke, Ph.D. CONTRACTING
AD Award Number: DAMD7---7 TITLE: Development of Artificial Antigen Presenting Cells for Prostate Cancer Immunotherapy PRINCIPAL INVESTIGATOR: Jonathan P. Schneck, M.D., Ph.D. Mathias Oelke, Ph.D. CONTRACTING
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure 1
 Supplementary Figure 1 FMO control (no HLA-DR antibody) FMO control (no HLA-DR antibody) Supplementary Figure 1. Gating strategy for HLA-DR+ T-cells. A gate was drawn around lymphocytes based on cell size
Supplementary Figure 1 FMO control (no HLA-DR antibody) FMO control (no HLA-DR antibody) Supplementary Figure 1. Gating strategy for HLA-DR+ T-cells. A gate was drawn around lymphocytes based on cell size
AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification
 BD AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification Overview of BD AbSeq antibody-oligonucleotide conjugates. High-throughput
BD AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification Overview of BD AbSeq antibody-oligonucleotide conjugates. High-throughput
CONVENTIONAL VACCINE DEVELOPMENT
 CONVENTIONAL VACCINE DEVELOPMENT PROBLEM Lethal germ Dead mouse LIVE VACCINES Related but harmless germ gives protection against lethal pathogen. Examples are the original pox vaccine and some TB vaccines
CONVENTIONAL VACCINE DEVELOPMENT PROBLEM Lethal germ Dead mouse LIVE VACCINES Related but harmless germ gives protection against lethal pathogen. Examples are the original pox vaccine and some TB vaccines
Nature Immunology: doi: /ni.3412
 Supplementary Figure 1 Gata1 expression in heamatopoietic stem and progenitor populations. (a) Unsupervised clustering according to 100 top variable genes across single pre-gm cells. The two main cell
Supplementary Figure 1 Gata1 expression in heamatopoietic stem and progenitor populations. (a) Unsupervised clustering according to 100 top variable genes across single pre-gm cells. The two main cell
Nature Genetics: doi: /ng Supplementary Figure 1. Workflow of CDR3 sequence assembly from RNA-seq data.
 Supplementary Figure 1 Workflow of CDR3 sequence assembly from RNA-seq data. Paired-end short-read RNA-seq data were mapped to human reference genome hg19, and unmapped reads in the TCR regions were extracted
Supplementary Figure 1 Workflow of CDR3 sequence assembly from RNA-seq data. Paired-end short-read RNA-seq data were mapped to human reference genome hg19, and unmapped reads in the TCR regions were extracted
Supplemental Information. Checkpoint Blockade Immunotherapy. Induces Dynamic Changes. in PD-1 CD8 + Tumor-Infiltrating T Cells
 Immunity, Volume 50 Supplemental Information Checkpoint Blockade Immunotherapy Induces Dynamic Changes in PD-1 CD8 + Tumor-Infiltrating T Cells Sema Kurtulus, Asaf Madi, Giulia Escobar, Max Klapholz, Jackson
Immunity, Volume 50 Supplemental Information Checkpoint Blockade Immunotherapy Induces Dynamic Changes in PD-1 CD8 + Tumor-Infiltrating T Cells Sema Kurtulus, Asaf Madi, Giulia Escobar, Max Klapholz, Jackson
Evolution of MHC-based technologies used for detection of antigen-responsive T cells
 Cancer Immunol Immunother (2017) 66:657 666 DOI 10.1007/s00262-017-1971-5 FOCUSSED RESEARCH REVIEW Evolution of MHC-based technologies used for detection of antigen-responsive T cells Amalie Kai Bentzen
Cancer Immunol Immunother (2017) 66:657 666 DOI 10.1007/s00262-017-1971-5 FOCUSSED RESEARCH REVIEW Evolution of MHC-based technologies used for detection of antigen-responsive T cells Amalie Kai Bentzen
