Supplementary Information. Supplementary Figures
|
|
|
- Joy Lyons
- 5 years ago
- Views:
Transcription
1 Supplementary Information Supplementary Figures.8 57 essential gene density LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA Supplementary Figure 1. Locations and minor allele frequencies of all structural variants in curated data set. Each of the three chromosomes is indicated by black bar, with scale (in megabases) at bottom. From top (same data as Fig 1): density of essential genes (blue), locations of Tf-type retrotransposons (green), and diversity (π, average pairwise diversity from SNPs, purple). Bar heights for deletions and duplications are proportional to minor allele frequency, the scale for retrotransposons is the frequency of the insertion in the 57 non-clonal strains. Diversity and retrotransposon were calculated from 57 non-clonal strains as described in Jeffares, et al. 1. Below, we show different types of SVs: deletions (black), duplications (red), inversions (green) and translocations (blue). The vertical lines terminating with open circles above dotted lines emit from the mid-point of each SV and indicate the minor allele frequencies in the population of 161 strains.
2 distance from c/s end gene overlap: DUP distance from chromosome end proportion transcript coverage all genes essential CNV rand REA rand DUP rand DUP rand gene overlap: DEL gene overlap: INV/TRA proportion transcript coverage proportion transcript coverage DEL rand DEL rand IN/TR rand IN/TR rand Supplementary Figure 2. Structural variations are biased towards chromosome ends and to low gene density regions. Top left panel, both CNVs and rearrangements are biased towards the ends of chromosomes. CNVs; median distance to chromosome ends 236 kb compared to chromosome- and sizematched random sites 944 kb, Wilcoxon rank sum test P = 1.3 x 1-11, rearrangements median distance 569 kb vs, matched random 863 kb, Wilcoxon test P =.3). All other panels, calculated proportion of each duplication and deletion that contained all protein-coding or essential genes. Box plots show the distributions of these proportions for all genes (grey), and proportion of coverage by essential genes (red), compared to the null distribution (rand). All comparisons were significantly less than the null distributions (Wilcoxon rank sum test, P-values <1.6 x 1-4 ). The same analysis was performed with the junctions of inversions and translocations, by calculating the transcript coverage in the region 5 bp upand down-stream of the predicted start and end junctions. These rearrangements are slightly biased away from genes (P = 1.9x1-3 ), but not significantly biased away from essential genes (P >.5). The null distributions were determined by selecting 1 regions for each actual variant/junction that were the same size, and were placed in random s on the same chromosome and calculating the gene coverage of these regions. Essential genes were those with the Fission Yeast Phenotype Ontology term defined as FYPO:261 ( inviable ) in PomBase.
3 DUP.I: length: 84 DUP.II: length: 13 DUP.II: length: e+5 6e+5 7e+5 8e DUP.II: length: 2 DUP.II: length: 18 DUP.III: length: DUP.III: length: 46 DUP.III: length: Supplementary Figure 3. Duplications that segregate within closely related strains. Plots show the average coverage in 1 kb non-overlapping windows for strains with a duplication (red) and all closely related strains without duplication (green); all these strains differ by <15 SNPs. The coverage of the two standard (h + and h - ) is shown in black. Top row, from left: variant DUP.I: (cluster 12, from Japan in 57), DUP.II: (cluster 12), DUP.II: (cluster 2, unknown origin), second row DUP.II: (cluster 2), DUP.II: (cluster 1, includes reference strain from French grapes in 1947), DUP.III: (cluster 2, various locations ). Bottom row: DUP.III: (cluster 6, Jamaica/USA), and DUP.III: (cluster 5, Sicily 1966). Genes are shown on top of plots with exons as red rectangles and retrotransposon LTRs as blue rectangles.
4 (a) Normalised standard deviation of copy number (b) Normalised standard deviation of copy number R =.15 p =.18 Normalised branch length of SNP-based tree R =.9 p <.1 Normalised branch length of CNV NJ tree (c) DUP.I DUP.I DUP.III DUP.III DUP.III DUP.III DUP.III DUP.III DEL.III DEL.III DUP.III DUP.III DUP.III DUP.III DUP.III DUP.III DUP.III DUP.III DUP.III DUP.III DEL.III DEL.III DUP.III DUP.III DUP.III DUP.III DUP.III DUP.III DUP.III DUP.III DUP.III DUP.I DUP.I DUP.I DEL.I DUP.I DEL.I DUP.I DUP.I DUP.I DUP.I DEL.I DUP.I DUP.I DEL.I DEL.I DUP.I DUP.I DEL.I DUP.I DEL.I DUP.I DUP.I DUP.I DUP.I DUP.I DEL.I DUP.I DUP.I DUP.I DUP.MT DUP.II DEL.II DEL.II DUP.II DEL.II DUP.II DUP.II DEL.II DUP.II DUP.II DUP.II DUP.II DUP.II DUP.II DUP.II DEL.II DUP.II DEL.II DUP.II DEL.II DEL.II DUP.II DEL.II DEL.II DEL.II DUP.II cluster value Supplementary Figure 4. Relative standard deviation of copy number variation within clusters. (a) Standard deviation of copy number for a CNV across the dataset is only weakly correlated with the total branch length of SNP-based phylogeny from the 2kb up- and down-stream phylogeny. (b) Standard deviation is highly correlated with the branch length of a CNV-based neighbour-joining tree. (c) The relative standard deviation of each CNV within each identified cluster of strains (<15 SNPs apart) relative to its change in rest of the dataset. For clarity, all relative standard deviations <1 are shown as white.
5 all clusters clusters with 1 15 strains cluster non ref allele freq cluster non ref allele freq DUP all clusters DEL all clusters cluster non ref allele freq cluster non ref allele freq INV all clusters (non REF) TRA all clusters (non REF) cluster non ref allele freq cluster non ref allele freq INV all clusters (MAF) TRA all clusters (MAF) cluster minor allele freq cluster minor allele freq Supplementary Figure 5. Copy number variants are usually rare alleles within clonal populations. Clonal clusters, or clonal populations all differ by < 15 SNPs. In rows, from top left; we show the within-cluster frequency of the non-reference allele for all SVs, which is skewed to rare alleles. Limiting this analysis to cluster with 1 to 15 strains highlights the low frequency of non-reference alleles. Second row; CNVs (duplications and deletions) are skewed to rare alleles, because the non-reference allele is usually the derived allele. Third row; inversions and translocations are not skewed to the non-reference allele, but here non-reference alleles are not necessarily the derived allele. Bottom row; the minor allele of inversions and translocations, however, is skewed to rare alleles.
6 Supplementary Figure 6. Chromosome-scale view of gene expression changes. The relative gene expression levels (strain1/strain2) for arrays 2 and 3, and arrays 7 and 8 are shown with their s on the three chromosomes. Filled circles indicates genes that we consider to be up-regulated (red) or repressed (green). Those highlighted with open red circles are consistently altered in both arrays (either 2+3, or 7+8). The blue lines show where the segregating duplications are. Box plots at right show the spread of data.
7 array 1 DUP.II within DUP, P= 9.5e 23 near DUP, P=.42 array 1 DUP.I within DUP, P= 8.6e 21 near DUP, P=.77 array 2 DUP.II within DUP, P= 7.2e 16 near DUP, P=.16 array 2 DUP.II within DUP, P= 3.3e 6 near DUP, P= array 3 DUP.II within DUP, P= 1.4e 13 near DUP, P= array 3 DUP.II within DUP, P= 3.6e 6 near DUP, P= array 4 DUP.II within DUP, P=.99 near DUP, P= array 5 DUP.III within DUP, P= 4.1e 17 near DUP, P= array 6 DUP.III within DUP, P= 6.2e 5 near DUP, P= array 7 DUP.III within DUP, P=.13 near DUP, P= array 8 DUP.III within DUP, P= 7.4e 5 near DUP, P=.99 expression ratio all duplications Spearman rank P= copy number ratio Supplementary Figure 7. No significant increase in gene expression immediately adjacent to duplications. For each duplication examined with DNA arrays, we show the relative expression (strain 1 vs strain 2) near the duplication. P-values show the support for the genes within the duplication (red vertical lines), or the 5 kb adjacent to the duplication (green vertical lines) being more highly expressed than all other genes in the chromosome (one-sided Wilcoxon rank sum tests). The grey horizontal lines show the 5 th, 5 th and 95 th percentiles for gene expression data on the chromosome. The bottom right panel shows that the median increase in expression level within a duplication correlates with the increase in genomic copy number. The solid back line shows the expected increase for the 1:1 correspondence between genomic copy number and relative expression (the line y=x), and the dashed line shows the linear model for the data. Copy number and relative expression change are correlated (Spearman rank correlation ρ =.71 and P =.14).
8 heritability 1. h contribution SNP REA CNV real data heritability simulated: SNPs only determine traits heritability simulated: CNV only determine traits heritability simulated: REA only determine traits Supplementary Figure 8. Contributions of SNPs, CNVs and rearrangements to traits. Top panel: for 227 traits, we show the total heritability estimated by the combination of 243,289 SNPs (green), 87 CNVs (red), and 26 rearrangements (grey). We then simulated data that was entirely due to the effects of SNPs (second panel), entirely due to the effects of CNVs (next panel) or entirely due to the effects of rearrangements (lower). In the second panel (entirely due to the effects of SNPs), the contribution of CNVs and rearrangements are artefacts, but these are relatively minor. This analysis indicates that the estimates are not strongly affected by linkage.
9 viable offspring (%) τ =.35 P=.16 Jeffares 215 Avelar SNP distance (% unshared SNPs) viable offspring (%) τ =.26 P= SV distance (unshared variants) viable offspring (%) τ =.36 P= 2e INV/TRA distance (unshared variants) viable offspring (%) τ =.15 P= CNV distance (unshared variants) viable offspring (%) τ =.15 P= merged CNV distance (unshared variants) SV distance τ =.6 P= 2.1e SNP distance Supplementary Figure 9. Correlations between spore viability, parental SNP-genetic distance and parental SV-genetic distance. Spore viability was measured for 58 crosses in total, including data from both Jeffares, et al. 1 (black) and Avelar, et al. 2 (red), with each circle representing one cross. Clockwise from top left plots show correlation of spore viability with various measures of genetic distances between parents; SN distance, all SV distances, all rearrangement differences (inversions/translocations), unmerged CNV distances, merged CNV distances, and finally the relationship between SNP distance and SV distance. Unmerged CNV differences count any CNV as being different between parents when either start or end coordinates are more than 1 kb apart. Because this definition can cause us to count largely overlapping events as different, we also counted merged differences where two CNVs were considered different only if their overlap was >5% of the total of both variants. This approach will exclude nested CNVs. CNV-genetic distance is not significantly correlated with viability in either case.
10 Supplementary Figure 1. SVs are enriched close to retrotransposon LTRs. For all SVs, we computed the closest distance of start or end coordinates to any LTR discovered previously 1. As a control, we compute the closest distance of 1 random coordinates on the same chromosome. Left: the distributions of distances for real SVs (grey), those that are within 2nt (black) or random coordinates (red). Right: using the same analysis, we show the closest distance of real SVs (CL: real), and random coordinates (C: random). We also show that both start and end coordinates of SVs (ST:real, EN:real) are closer than random s (POS:random). all SVs nearest LTR (nt) Density Wilcox test P = 1.3e 25 CL:real CL:random ST:real EN:real POS:random
11 Supplementary Figure 11. Chromosome-scale read coverage plots for three chromosomes of strain JB127. Coverage is calculated relative to the reference strain (JB22 in our collection). Two large duplications that did not satisfy the criteria used to detect CNVs with cn.mops are indicated with blue arrows (DUP.I: , 24kb and DUP.I: , 51kb).
12 Supplementary References 1 Jeffares, D. C. et al. The genomic and phenotypic diversity of Schizosaccharomyces pombe. Nat Genet 47, , doi:1.138/ng.3215 (215). 2 Avelar, A. T., Perfeito, L., Gordo, I. & Ferreira, M. G. Genome architecture is a selectable trait that can be maintained by antagonistic pleiotropy. Nat Commun 4, 2235, doi:1.138/ncomms3235 (213).
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Global variation in copy number in the human genome
 Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
November 9, Johns Hopkins School of Medicine, Baltimore, MD,
 Fast detection of de-novo copy number variants from case-parent SNP arrays identifies a deletion on chromosome 7p14.1 associated with non-syndromic isolated cleft lip/palate Samuel G. Younkin 1, Robert
Fast detection of de-novo copy number variants from case-parent SNP arrays identifies a deletion on chromosome 7p14.1 associated with non-syndromic isolated cleft lip/palate Samuel G. Younkin 1, Robert
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Genomic structural variation
 Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first
 Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Nature Genetics: doi: /ng Supplementary Figure 1. Somatic coding mutations identified by WES/WGS for 83 ATL cases.
 Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Figure S2. Distribution of acgh probes on all ten chromosomes of the RIL M0022
 96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
Supplementary Figure 1 Effect of HSP90 inhibition on expression of endogenous retroviruses. (a) Inducible shrna-mediated Hsp90 silencing in mouse ESCs. Immunoblots of total cell extract expressing the
LTA Analysis of HapMap Genotype Data
 LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
SVIM: Structural variant identification with long reads DAVID HELLER MAX PLANCK INSTITUTE FOR MOLECULAR GENETICS, BERLIN JUNE 2O18, SMRT LEIDEN
 SVIM: Structural variant identification with long reads DAVID HELLER MAX PLANCK INSTITUTE FOR MOLECULAR GENETICS, BERLIN JUNE 2O18, SMRT LEIDEN Structural variation (SV) Variants larger than 50bps Affect
SVIM: Structural variant identification with long reads DAVID HELLER MAX PLANCK INSTITUTE FOR MOLECULAR GENETICS, BERLIN JUNE 2O18, SMRT LEIDEN Structural variation (SV) Variants larger than 50bps Affect
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Identification of regions with common copy-number variations using SNP array
 Identification of regions with common copy-number variations using SNP array Agus Salim Epidemiology and Public Health National University of Singapore Copy Number Variation (CNV) Copy number alteration
Identification of regions with common copy-number variations using SNP array Agus Salim Epidemiology and Public Health National University of Singapore Copy Number Variation (CNV) Copy number alteration
Variant Classification. Author: Mike Thiesen, Golden Helix, Inc.
 Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Nature Genetics: doi: /ng Supplementary Figure 1. Rates of different mutation types in CRC.
 Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Nature Biotechnology: doi: /nbt.1904
 Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
p.r623c p.p976l p.d2847fs p.t2671 p.d2847fs p.r2922w p.r2370h p.c1201y p.a868v p.s952* RING_C BP PHD Cbp HAT_KAT11
 ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
Chromosome Structure & Recombination
 Chromosome Structure & Recombination (CHAPTER 8- Brooker Text) April 4 & 9, 2007 BIO 184 Dr. Tom Peavy Genetic variation refers to differences between members of the same species or those of different
Chromosome Structure & Recombination (CHAPTER 8- Brooker Text) April 4 & 9, 2007 BIO 184 Dr. Tom Peavy Genetic variation refers to differences between members of the same species or those of different
The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0
 The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0 Introduction Loss of erozygosity (LOH) represents the loss of allelic differences. The SNP markers on the SNP Array 6.0 can be used
The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0 Introduction Loss of erozygosity (LOH) represents the loss of allelic differences. The SNP markers on the SNP Array 6.0 can be used
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
CNV PCA Search Tutorial
 CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
Next Generation Sequencing as a tool for breakpoint analysis in rearrangements of the globin-gene clusters
 Next Generation Sequencing as a tool for breakpoint analysis in rearrangements of the globin-gene clusters XXXth International Symposium on Technical Innovations in Laboratory Hematology Honolulu, Hawaii
Next Generation Sequencing as a tool for breakpoint analysis in rearrangements of the globin-gene clusters XXXth International Symposium on Technical Innovations in Laboratory Hematology Honolulu, Hawaii
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities
 Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Mosaic loss of chromosome Y in peripheral blood is associated with shorter survival and higher risk of cancer
 Supplementary Information Mosaic loss of chromosome Y in peripheral blood is associated with shorter survival and higher risk of cancer Lars A. Forsberg, Chiara Rasi, Niklas Malmqvist, Hanna Davies, Saichand
Supplementary Information Mosaic loss of chromosome Y in peripheral blood is associated with shorter survival and higher risk of cancer Lars A. Forsberg, Chiara Rasi, Niklas Malmqvist, Hanna Davies, Saichand
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
Supplementary Information
 Supplementary Information 5-hydroxymethylcytosine-mediated epigenetic dynamics during postnatal neurodevelopment and aging By Keith E. Szulwach 1,8, Xuekun Li 1,8, Yujing Li 1, Chun-Xiao Song 2, Hao Wu
Supplementary Information 5-hydroxymethylcytosine-mediated epigenetic dynamics during postnatal neurodevelopment and aging By Keith E. Szulwach 1,8, Xuekun Li 1,8, Yujing Li 1, Chun-Xiao Song 2, Hao Wu
Supplementary note: Comparison of deletion variants identified in this study and four earlier studies
 Supplementary note: Comparison of deletion variants identified in this study and four earlier studies Here we compare the results of this study to potentially overlapping results from four earlier studies
Supplementary note: Comparison of deletion variants identified in this study and four earlier studies Here we compare the results of this study to potentially overlapping results from four earlier studies
GENOME-WIDE ASSOCIATION STUDIES
 GENOME-WIDE ASSOCIATION STUDIES SUCCESSES AND PITFALLS IBT 2012 Human Genetics & Molecular Medicine Zané Lombard IDENTIFYING DISEASE GENES??? Nature, 15 Feb 2001 Science, 16 Feb 2001 IDENTIFYING DISEASE
GENOME-WIDE ASSOCIATION STUDIES SUCCESSES AND PITFALLS IBT 2012 Human Genetics & Molecular Medicine Zané Lombard IDENTIFYING DISEASE GENES??? Nature, 15 Feb 2001 Science, 16 Feb 2001 IDENTIFYING DISEASE
Nature Immunology: doi: /ni Supplementary Figure 1. DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells.
 Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Supplementary Figure 1 DNA-methylation machinery is essential for silencing of Cd4 in cytotoxic T cells. (a) Scheme for the retroviral shrna screen. (b) Histogram showing CD4 expression (MFI) in WT cytotoxic
Nature Genetics: doi: /ng Supplementary Figure 1. PCA for ancestry in SNV data.
 Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supporting Information
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 Supporting Information Variances and biases of absolute distributions were larger in the 2-line
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 Supporting Information Variances and biases of absolute distributions were larger in the 2-line
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
Challenges of CGH array testing in children with developmental delay. Dr Sally Davies 17 th September 2014
 Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
CHROMOSOMAL MICROARRAY (CGH+SNP)
 Chromosome imbalances are a significant cause of developmental delay, mental retardation, autism spectrum disorders, dysmorphic features and/or birth defects. The imbalance of genetic material may be due
Chromosome imbalances are a significant cause of developmental delay, mental retardation, autism spectrum disorders, dysmorphic features and/or birth defects. The imbalance of genetic material may be due
Unit 5 Review Name: Period:
 Unit 5 Review Name: Period: 1 4 5 6 7 & give an example of the following. Be able to apply their meanings: Homozygous Heterozygous Dominant Recessive Genotype Phenotype Haploid Diploid Sex chromosomes
Unit 5 Review Name: Period: 1 4 5 6 7 & give an example of the following. Be able to apply their meanings: Homozygous Heterozygous Dominant Recessive Genotype Phenotype Haploid Diploid Sex chromosomes
High Throughput Sequence (HTS) data analysis. Lei Zhou
 High Throughput Sequence (HTS) data analysis Lei Zhou (leizhou@ufl.edu) High Throughput Sequence (HTS) data analysis 1. Representation of HTS data. 2. Visualization of HTS data. 3. Discovering genomic
High Throughput Sequence (HTS) data analysis Lei Zhou (leizhou@ufl.edu) High Throughput Sequence (HTS) data analysis 1. Representation of HTS data. 2. Visualization of HTS data. 3. Discovering genomic
MIR retrotransposon sequences provide insulators to the human genome
 Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Investigating rare diseases with Agilent NGS solutions
 Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Nature Genetics: doi: /ng Supplementary Figure 1. Country distribution of GME samples and designation of geographical subregions.
 Supplementary Figure 1 Country distribution of GME samples and designation of geographical subregions. GME samples collected across 20 countries and territories from the GME. Pie size corresponds to the
Supplementary Figure 1 Country distribution of GME samples and designation of geographical subregions. GME samples collected across 20 countries and territories from the GME. Pie size corresponds to the
Chromatin marks identify critical cell-types for fine-mapping complex trait variants
 Chromatin marks identify critical cell-types for fine-mapping complex trait variants Gosia Trynka 1-4 *, Cynthia Sandor 1-4 *, Buhm Han 1-4, Han Xu 5, Barbara E Stranger 1,4#, X Shirley Liu 5, and Soumya
Chromatin marks identify critical cell-types for fine-mapping complex trait variants Gosia Trynka 1-4 *, Cynthia Sandor 1-4 *, Buhm Han 1-4, Han Xu 5, Barbara E Stranger 1,4#, X Shirley Liu 5, and Soumya
Whole-genome detection of disease-associated deletions or excess homozygosity in a case control study of rheumatoid arthritis
 HMG Advance Access published December 21, 2012 Human Molecular Genetics, 2012 1 13 doi:10.1093/hmg/dds512 Whole-genome detection of disease-associated deletions or excess homozygosity in a case control
HMG Advance Access published December 21, 2012 Human Molecular Genetics, 2012 1 13 doi:10.1093/hmg/dds512 Whole-genome detection of disease-associated deletions or excess homozygosity in a case control
Nature Genetics: doi: /ng Supplementary Figure 1. HOX fusions enhance self-renewal capacity.
 Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Theta sequences are essential for internally generated hippocampal firing fields.
 Theta sequences are essential for internally generated hippocampal firing fields. Yingxue Wang, Sandro Romani, Brian Lustig, Anthony Leonardo, Eva Pastalkova Supplementary Materials Supplementary Modeling
Theta sequences are essential for internally generated hippocampal firing fields. Yingxue Wang, Sandro Romani, Brian Lustig, Anthony Leonardo, Eva Pastalkova Supplementary Materials Supplementary Modeling
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the
 Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
Nature Genetics: doi: /ng Supplementary Figure 1. Clinical timeline for the discovery WES cases.
 Supplementary Figure 1 Clinical timeline for the discovery WES cases. This illustrates the timeline of the disease events during the clinical course of each patient s disease, further indicating the available
Supplementary Figure 1 Clinical timeline for the discovery WES cases. This illustrates the timeline of the disease events during the clinical course of each patient s disease, further indicating the available
Introduction to genetic variation. He Zhang Bioinformatics Core Facility 6/22/2016
 Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
CCM6+7+ Unit 12 Data Collection and Analysis
 Page 1 CCM6+7+ Unit 12 Packet: Statistics and Data Analysis CCM6+7+ Unit 12 Data Collection and Analysis Big Ideas Page(s) What is data/statistics? 2-4 Measures of Reliability and Variability: Sampling,
Page 1 CCM6+7+ Unit 12 Packet: Statistics and Data Analysis CCM6+7+ Unit 12 Data Collection and Analysis Big Ideas Page(s) What is data/statistics? 2-4 Measures of Reliability and Variability: Sampling,
indicated in shaded lowercase letters (hg19, Chr2: 217,955, ,957,266).
 Legend for Supplementary Figures Figure S1: Sequence of 2q35 encnv. The DNA sequence of the 1,375bp 2q35 encnv is indicated in shaded lowercase letters (hg19, Chr2: 217,955,892-217,957,266). Figure S2:
Legend for Supplementary Figures Figure S1: Sequence of 2q35 encnv. The DNA sequence of the 1,375bp 2q35 encnv is indicated in shaded lowercase letters (hg19, Chr2: 217,955,892-217,957,266). Figure S2:
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Nature Medicine: doi: /nm.4439
 Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
EXPression ANalyzer and DisplayER
 EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
Epigenetics. Jenny van Dongen Vrije Universiteit (VU) Amsterdam Boulder, Friday march 10, 2017
 Epigenetics Jenny van Dongen Vrije Universiteit (VU) Amsterdam j.van.dongen@vu.nl Boulder, Friday march 10, 2017 Epigenetics Epigenetics= The study of molecular mechanisms that influence the activity of
Epigenetics Jenny van Dongen Vrije Universiteit (VU) Amsterdam j.van.dongen@vu.nl Boulder, Friday march 10, 2017 Epigenetics Epigenetics= The study of molecular mechanisms that influence the activity of
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
MSI positive MSI negative
 Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
DETECTING HIGHLY DIFFERENTIATED COPY-NUMBER VARIANTS FROM POOLED POPULATION SEQUENCING
 DETECTING HIGHLY DIFFERENTIATED COPY-NUMBER VARIANTS FROM POOLED POPULATION SEQUENCING DANIEL R. SCHRIDER * Department of Biology and School of Informatics and Computing, Indiana University, 1001 E Third
DETECTING HIGHLY DIFFERENTIATED COPY-NUMBER VARIANTS FROM POOLED POPULATION SEQUENCING DANIEL R. SCHRIDER * Department of Biology and School of Informatics and Computing, Indiana University, 1001 E Third
NGS panels in clinical diagnostics: Utrecht experience. Van Gijn ME PhD Genome Diagnostics UMCUtrecht
 NGS panels in clinical diagnostics: Utrecht experience Van Gijn ME PhD Genome Diagnostics UMCUtrecht 93 Gene panels UMC Utrecht Cardiovascular disease (CAR) (5 panels) Epilepsy (EPI) (11 panels) Hereditary
NGS panels in clinical diagnostics: Utrecht experience Van Gijn ME PhD Genome Diagnostics UMCUtrecht 93 Gene panels UMC Utrecht Cardiovascular disease (CAR) (5 panels) Epilepsy (EPI) (11 panels) Hereditary
Updating penetrance estimates for syndromes with variable phenotypic manifestation. Adele Corrigan June 27th
 Updating penetrance estimates for syndromes with variable phenotypic manifestation Adele Corrigan June 27th Background Array CGH has led to increased identification of copy number variants (CNVs) Our understanding
Updating penetrance estimates for syndromes with variable phenotypic manifestation Adele Corrigan June 27th Background Array CGH has led to increased identification of copy number variants (CNVs) Our understanding
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.
 Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols.
 Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols. A-tailed DNA was ligated to T-tailed dutp adapters, circularized
Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols. A-tailed DNA was ligated to T-tailed dutp adapters, circularized
Mendel s Methods: Monohybrid Cross
 Mendel s Methods: Monohybrid Cross Mendel investigated whether the white-flowered form disappeared entirely by breeding the F1 purple flowers with each other. Crossing two purple F1 monohybrid plants is
Mendel s Methods: Monohybrid Cross Mendel investigated whether the white-flowered form disappeared entirely by breeding the F1 purple flowers with each other. Crossing two purple F1 monohybrid plants is
The Chromosomal Basis of Inheritance
 The Chromosomal Basis of Inheritance Factors and Genes Mendel s model of inheritance was based on the idea of factors that were independently assorted and segregated into gametes We now know that these
The Chromosomal Basis of Inheritance Factors and Genes Mendel s model of inheritance was based on the idea of factors that were independently assorted and segregated into gametes We now know that these
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD
 Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) cancer DNA
 www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
SUPPLEMENTARY FIGURES: Supplementary Figure 1
 SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016
 Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Multiple samples from five patients (P4, P8, P14, P15 and P17) with Barrett s esophagus and adjacent EAC show that the poor overlap is not a result of sampling bias. Bar graphs showing
Supplementary Figure 1 Multiple samples from five patients (P4, P8, P14, P15 and P17) with Barrett s esophagus and adjacent EAC show that the poor overlap is not a result of sampling bias. Bar graphs showing
CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE. Dr. Bahar Naghavi
 2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
Unit 1 Exploring and Understanding Data
 Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Figure 1: Transmission of Wing Shape & Body Color Alleles: F0 Mating. Figure 1.1: Transmission of Wing Shape & Body Color Alleles: Expected F1 Outcome
 I. Chromosomal Theory of Inheritance As early cytologists worked out the mechanism of cell division in the late 1800 s, they began to notice similarities in the behavior of BOTH chromosomes & Mendel s
I. Chromosomal Theory of Inheritance As early cytologists worked out the mechanism of cell division in the late 1800 s, they began to notice similarities in the behavior of BOTH chromosomes & Mendel s
Tutorial on Genome-Wide Association Studies
 Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
John Nguyen, Nozomi Nishimura, Robert Fetcho, Costantino Iadecola, Chris B. Schaffer
 Supplemental figures and text for Occlusion of cortical ascending venules causes blood flow decreases, reversals in flow direction, and vessel dilation in upstream capillaries John Nguyen, Nozomi Nishimura,
Supplemental figures and text for Occlusion of cortical ascending venules causes blood flow decreases, reversals in flow direction, and vessel dilation in upstream capillaries John Nguyen, Nozomi Nishimura,
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Lecture 20. Disease Genetics
 Lecture 20. Disease Genetics Michael Schatz April 12 2018 JHU 600.749: Applied Comparative Genomics Part 1: Pre-genome Era Sickle Cell Anaemia Sickle-cell anaemia (SCA) is an abnormality in the oxygen-carrying
Lecture 20. Disease Genetics Michael Schatz April 12 2018 JHU 600.749: Applied Comparative Genomics Part 1: Pre-genome Era Sickle Cell Anaemia Sickle-cell anaemia (SCA) is an abnormality in the oxygen-carrying
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
Chapter 15 Notes 15.1: Mendelian inheritance chromosome theory of inheritance wild type 15.2: Sex-linked genes
 Chapter 15 Notes The Chromosomal Basis of Inheritance Mendel s hereditary factors were genes, though this wasn t known at the time Now we know that genes are located on The location of a particular gene
Chapter 15 Notes The Chromosomal Basis of Inheritance Mendel s hereditary factors were genes, though this wasn t known at the time Now we know that genes are located on The location of a particular gene
Introduction to Genetics and Genomics
 2016 Introduction to enetics and enomics 3. ssociation Studies ggibson.gt@gmail.com http://www.cig.gatech.edu Outline eneral overview of association studies Sample results hree steps to WS: primary scan,
2016 Introduction to enetics and enomics 3. ssociation Studies ggibson.gt@gmail.com http://www.cig.gatech.edu Outline eneral overview of association studies Sample results hree steps to WS: primary scan,
Supplemental Figure 1. Small RNA size distribution from different soybean tissues.
 Supplemental Figure 1. Small RNA size distribution from different soybean tissues. The size of small RNAs was plotted versus frequency (percentage) among total sequences (A, C, E and G) or distinct sequences
Supplemental Figure 1. Small RNA size distribution from different soybean tissues. The size of small RNAs was plotted versus frequency (percentage) among total sequences (A, C, E and G) or distinct sequences
An Introduction to Quantitative Genetics I. Heather A Lawson Advanced Genetics Spring2018
 An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
Calling DNA variants SNVs, CNVs, and SVs. Steve Laurie Variant Effect Predictor Training Course Prague, 6 th November 2017
 1 Calling DNA variants SNVs, CNVs, and SVs Steve Laurie Variant Effect Predictor Training Course Prague, 6 th November 2017 Calling DNA variants SNVs, CNVs, SVs 2 1. What is a variant? 2. Paired End read
1 Calling DNA variants SNVs, CNVs, and SVs Steve Laurie Variant Effect Predictor Training Course Prague, 6 th November 2017 Calling DNA variants SNVs, CNVs, SVs 2 1. What is a variant? 2. Paired End read
Nature Genetics: doi: /ng Supplementary Figure 1. Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms.
 Supplementary Figure 1 Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms. IF images of wild-type (wt) and met-2 set-25 worms showing the loss of H3K9me2/me3 at the indicated developmental
Supplementary Figure 1 Immunofluorescence (IF) confirms absence of H3K9me in met-2 set-25 worms. IF images of wild-type (wt) and met-2 set-25 worms showing the loss of H3K9me2/me3 at the indicated developmental
Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University
 Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Supplemental Figures. Figure S1: 2-component Gaussian mixture model of Bourdeau et al. s fold-change distribution
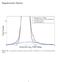 Supplemental Figures Figure S1: 2-component Gaussian mixture model of Bourdeau et al. s fold-change distribution 1 All CTCF Unreponsive RefSeq Frequency 0 2000 4000 6000 0 500 1000 1500 2000 Block Size
Supplemental Figures Figure S1: 2-component Gaussian mixture model of Bourdeau et al. s fold-change distribution 1 All CTCF Unreponsive RefSeq Frequency 0 2000 4000 6000 0 500 1000 1500 2000 Block Size
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
SUPPLEMENTARY INFORMATION
 Supplementary text Collectively, we were able to detect ~14,000 expressed genes with RPKM (reads per kilobase per million) > 1 or ~16,000 with RPKM > 0.1 in at least one cell type from oocyte to the morula
Supplementary text Collectively, we were able to detect ~14,000 expressed genes with RPKM (reads per kilobase per million) > 1 or ~16,000 with RPKM > 0.1 in at least one cell type from oocyte to the morula
Detection of copy number variations in PCR-enriched targeted sequencing data
 Detection of copy number variations in PCR-enriched targeted sequencing data German Demidov Parseq Lab, Saint-Petersburg University of Russian Academy of Sciences, current: Center for Genomic Regulation
Detection of copy number variations in PCR-enriched targeted sequencing data German Demidov Parseq Lab, Saint-Petersburg University of Russian Academy of Sciences, current: Center for Genomic Regulation
