TRANSCRIPTION CAPPING
|
|
|
- Joel August Howard
- 5 years ago
- Views:
Transcription
1 Messenger RNA
2
3 Eucaryotic gene expression transcription START site exons upstream elements promoter elements TRANSCRIPTION CAPPING introns gene m 7 G pre-mrna m 7 G SPLICING POLYADENYLATION AAAAAAAAAn polyadenylated pre-mrna m 7 G TRANSPORT AAAAAAAAAn mature mrna TRANSLATION
4 mrna structures AUG UAA cap AAAAAAAAAAAAA 5 UTR 3 UTR ORF
5 Eucaryotic gene expression transcription START site exons upstream elements promoter elements TRANSCRIPTION CAPPING introns gene m 7 G pre-mrna m 7 G SPLICING POLYADENYLATION AAAAAAAAAn polyadenylated pre-mrna m 7 G TRANSPORT AAAAAAAAAn mature mrna TRANSLATION
6 Why the Cap structure is important? 1) RNA stability 2) Favours the mrna transport to the cytoplasm 3) Increases transla?on (it binds to eif4e that belongs to transla?on ini?a?on complex)
7 7-metilguanosina This involves the addi?on of a modified guanine base to the 50 end of the RNA. Specifically, it is a methylated guanine, and it is joined to the RNA transcript by an unusual 5 5 linkage involving three phosphates CAP is added at very early stage of transcrip?on ini?a?on estremità 5 della RNA catena The 5-5 phosphodiester bond makes the molecule resistant to the exonuclease ac?vity. I vitro synthe?zed RNA without CAP are rapidly degraded
8 pre-mrna capping The 5 cap is created in three enzyma?c steps: 1. a phosphate group is removed from the 5 end of the transcript. RNA triphosphatase 2. GMP moiety is added. RNA guanylyltransferase 3. GMP nucleo?de is modified by the addi?on of a methyl group. RNA(G-7-) methyltransferase
9 5 CAP favours the mrna transport to the cytoplasm CBC: CAP binding complex
10 Eucaryotic gene expression transcription START site exons upstream elements promoter elements TRANSCRIPTION CAPPING introns gene m 7 G pre-mrna m 7 G SPLICING POLYADENYLATION AAAAAAAAAn polyadenylated pre-mrna m 7 G TRANSPORT AAAAAAAAAn mature mrna TRANSLATION
11 How is the func4on of the polya tail? 1) RNA stability 2) Favours the mrna transport to the cytoplasm 3) Increases transla?on efficiency by favou?ng the loading of ribosomal 40S subunit 4) mrna 3 end forma?on allows efficient transcrip?on termina?on.
12 Looking for consenus sequence SEQUENCE ALLINEAMENT OF cdnas STARTING FROM THE POLYA TAIL 5 UTR 3 UTR nt globin AUG UAA GGUUAUCCAUCAAUAAA.GCUAUACGCAAAA n Ig AUG UAA CCACUGGGCCAAUAAA.GCUAUACGCAAAA n Ovoalbumin AUG UAA GCAACCUCGAAUAAA.GCUAUACGCAAAA n Polymerase AUG UAA AUCUGGAGGAAUAAA.GCUAUACGCAAAA n Myc AUG UAA UAUAGAUCCAAUAAA.GCUAUACGCAAAA n AAUAAA consensus
13 Efficiency consensus Polyadenyla?on efficiency Influence of consensus sequence AAUAAA!
14 Site- specific muta4on Recombinant DNA preparation * mutazione transfection mut1 mut2 mut3 WT polya Northern blot RNA extraction After 24 h HeLa cells
15 3 end forma?on in mammalian cells Cleavage and polyadenyla?on site 5 AAUAAA GU rich nt 5 AAUAAA OH P GU rich 3 5 AAUAAA A 250 degrada?on
16 Cleavage and Polyadenylation Specific Factor" CPSF! 5 AAUAAA CA SPECIFIC COMPLEXES ARE INVOLVED IN mrna 3 END PROCESSING sito di taglio Cleavage Stimulation Factor" CstF! " PAP 3 Poly- A polymerase GU- rich CFII! CFI! Curr Opin Cell Biol Jun;16(3):272-8" HeLa NE AAUAAA AAGAAA complex EMSA assay for tes?ng the interac?on between CPSF 160 and consensus sequence AAUAAA sito di taglio free RNA
17 3 - End Forma4on: RNA Processing Cleavage and Polyadenyla?on Specific Factor Cleavage Factors Cleavage S?mula?on Factor
18 18
19 19
20 degrada4on 20
21 21
22 22
23 CAP and polya tail influence efficient translation
24 Polyadenylation is linked to termination mrna Torpedo Model AAAAAAAAA pre- mrna CPSF pa Xrn2 CstF RNAPII RNAPII
25 Alterna?ve polyadenyla?on of the immunoglobulin μ heavy chain gene sub obtimal pa obtimal pa Secreted terminus Membrane bound terminus Secreted terminus Membrane bound terminus
26 CSTF- 64 levels control the alterna?ve processing of mrna. In not ac?vated B cells the limi?ng concentra?on of CSTF allows the recogni?on of the stronger polyadenyla?on signal. The immuglobulin produced will then contain a por?on for binding to the membrane. Aler the ac?va?on of B cells CSTF levels are increased and this allows the use of the weaker polyadenyla?on site that will be preferen?ally used because it will be the first to be transcribed. CstF-64 The polyadenylation factor CstF-64 regulates alternative processing of IgM heavy chain premrna during B cell differentiation. Takagaki Y, Seipelt RL, Peterson ML, Manley JL. Cell 1996 Nov 29;87(5):941-52
27 Eucaryotic gene expression transcription START site exons upstream elements promoter elements TRANSCRIPTION CAPPING introns gene m 7 G pre-mrna m 7 G SPLICING POLYADENYLATION AAAAAAAAAn polyadenylated pre-mrna m 7 G TRANSPORT AAAAAAAAAn mature mrna TRANSLATION
28 Eukaryotic genes contain introns! Fig. 14.2
29 Identification of introns by R-looping Gene dell ovalbumina di pollo ibridato con il suo mrna e visualizzato al microscopio elemronico Exon:intron organization of the ovalbumin gene Exon Intron
30 Specific sequences inside the pre- mrna indicate where the spicing event has to take place. These sequences indicate the boundary between introns and exons.
31 Consensus Sequences Surrounding the 5 and 3 Splice Sites 31
32 The intron is removed in a Form Called Lariat and the Flanking Exons are joined Two trans- esterifica4ons: Step 1: The OH of the conserved A at the branch site amacks the phosphoryl group of the conserved G in the 5 splice site. As result, the 5 exon is released and the 5 - end of the intron forms a three- way junc?on structure. Step 2: The OH of the 5 exon amacks the phosphoryl group at the 3 splice site. As a consequence, the 5 and 3 exons are joined and the intron is released in a lariat shape
33 Nuclear pre-mrna splicing proceeds through two trans-esterification reactions! Fig. 14.4
34 Splicing occurs in a spliceosome an RNA-protein complex spliceosome (~100 proteins + 5 small RNAs) U1, U2, U4/U5/U6 pre- mrna spliced mrna Splicing works similarly in different organisms, for example in yeast, flies, worms, plants and animals.
35 Five snrnas are involved in pre-mrna splicing"
36 U4/U5/U6 tri-snrnps U1 and U2 snrnps
37 Ⅰ Assembly, rearrangement, and catalysis within the spliceosome: the splicing pathway Assembly step 1 1. U1 recognizes 5 splice site. 2. One subunit of U2AF binds to Py tract and the other to the 3 splice site. The former subunits interacts with BBP and helps it to bind to the branch point. 3. Early (E) complex is formed U1 BBP U2AF pg A 3 E complex
38 Assembly step 2 1. U2 binds to the branch site, and then A complex is formed. 2. The base- pairing between the U2 and the branch site is such that the branch site A is extruded. This A residue is available to react with the 5 splice site. U1 U2AF pg A 3 U2 A complex
39 Assembly step 3 1. U4, U5 and U6 form the tri- snrnp Par?cle. 2. With the entry of the tri- snrnp, the A complex is converted into the B complex. U1 U2AF pg A 3 U6 U5 U4 U2 U2AF65 35 A complex U5 U1 U2 U6 U4 A B complex
40 U1 U5 U2 B complex U6 U4 A U1 U6 U5 U4 U2 A A C complex in which the catalysis has not occurred yet U4 U6 U5 U2 A
41 Catalysis Step 1 Forma?on of the C complex produces the ac4ve site, with U2 and U6 RNAs being brought together. Forma?on of the ac?ve site juxtaposes the 5 splice site of the pre- mrna and the branch site, allowing the branched A residue to a]ack the 5 splice site to accomplish the first transesterfica?on reac?on. U4 U6 U5 U2 A
42 Catalysis Step 2 U5 snrnp helps to bring the two exons together, and aids the second transesterifica?on reac?on, in which the 3 - OH of the 5 exon amacks the 3 splice site. Final Step Release of the mrna product and the snrnps. U5 U5 U6 U2 A U6 U2 A
43 Spliceosome & ATP RNA- RNA Rearrangements - I 43
44 Spliceosome & ATP RNA- RNA Rearrangements - II 44
45 Pre- mrna Splicing is Accelerated by RNA- RNA Basepairing: snrnps 45
46 How to study splicing in vitro procedure -in vitro transcribed RNA -RNA plus nuclear extract -RNA isolation ot different time points -elettrophoresis for the analysis of the reaction products RNA analysis RNA isolation nuclear extract
47 Recognition of canonical splice sites?? BBP GU A AG GU YYYYYY 10-15% of single nucleo?de muta?ons causing disease affect splice sites From: H. Stark, P. Dube, R. Luhrmann & B. Kastner, Nature 409, (2001)
48 RNA splicing gets the most out of genes In animals, complexity depends less upon the number of genes than upon the number of different ways they can splice the RNA
49 Genome sequence analysis! Introns are abundant! 94% human, 85% fruitfly, 95% nemtode, 95% plant genes! Human genome (26,000-35,000 genes)! Only 2-5 times as many genes as in fruitflies (13,600)! or in nematodes (19,000)! Alternative splicing is abundant!! ~75% of human gene transcripts are alternatively spliced!
50 Alternative splicing The presence of mul?ple introns in many eukaryo?c genes permits expression of mul?ple, related proteins from a single gene by means of alterna?ve slicing, an important mechanism for the produc?on of different forms of proteins, called isoforms, by different types of cells. Nearly 75% of all human genes are expressed as alterna?vely spliced mrnas, leading to an expansion of the coding capacity of our genome. intron 1 intron 2 intron 3 intron Usually all introns must be removed before the mrna can be translated to produce protein
51 Alternative Splicing! 5 splice site switching! 3 splice site switching! Exon Skipping! Mutually exclusive exons! Intron retention!
52
53
54
55 Splicing modulators
56 Multiple introns may be spliced differently in different circumstances, for example in different tissues. Heart muscle mrna pre- mrna Uterine muscle mrna hus one gene can encode more than one protein. The proteins are similar but not identical and may have distinct properties. This is important in complex organisms
57 Detec?on of alterna?ve splicing by Northern blorng Total RNA Transfer solution RNA Probe against exon
Messenger RNA maturation. Dr.ssa Mariangela Morlando
 Messenger RNA maturation Dr.ssa Mariangela Morlando Features of RNA molecule Synthe(sed from a DNA template (transcrip(on) Single strand molecule Contains ribose in place of deoxyribose Contains Uracil
Messenger RNA maturation Dr.ssa Mariangela Morlando Features of RNA molecule Synthe(sed from a DNA template (transcrip(on) Single strand molecule Contains ribose in place of deoxyribose Contains Uracil
Eukaryotic mrna is covalently processed in three ways prior to export from the nucleus:
 RNA Processing Eukaryotic mrna is covalently processed in three ways prior to export from the nucleus: Transcripts are capped at their 5 end with a methylated guanosine nucleotide. Introns are removed
RNA Processing Eukaryotic mrna is covalently processed in three ways prior to export from the nucleus: Transcripts are capped at their 5 end with a methylated guanosine nucleotide. Introns are removed
Genetics. Instructor: Dr. Jihad Abdallah Transcription of DNA
 Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Genetics Instructor: Dr. Jihad Abdallah Transcription of DNA 1 3.4 A 2 Expression of Genetic information DNA Double stranded In the nucleus Transcription mrna Single stranded Translation In the cytoplasm
Mechanism of splicing
 Outline of Splicing Mechanism of splicing Em visualization of precursor-spliced mrna in an R loop Kinetics of in vitro splicing Analysis of the splice lariat Lariat Branch Site Splice site sequence requirements
Outline of Splicing Mechanism of splicing Em visualization of precursor-spliced mrna in an R loop Kinetics of in vitro splicing Analysis of the splice lariat Lariat Branch Site Splice site sequence requirements
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics
 1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
1. Identify and characterize interesting phenomena! 2. Characterization should stimulate some questions/models! 3. Combine biochemistry and genetics to gain mechanistic insight! 4. Return to step 2, as
Processing of RNA II Biochemistry 302. February 13, 2006
 Processing of RNA II Biochemistry 302 February 13, 2006 Precursor mrna: introns and exons Intron: Transcribed RNA sequence removed from precursor RNA during the process of maturation (for class II genes:
Processing of RNA II Biochemistry 302 February 13, 2006 Precursor mrna: introns and exons Intron: Transcribed RNA sequence removed from precursor RNA during the process of maturation (for class II genes:
Processing of RNA II Biochemistry 302. February 14, 2005 Bob Kelm
 Processing of RNA II Biochemistry 302 February 14, 2005 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation (non proteincoding sequence) Discovered by P. Sharp
Processing of RNA II Biochemistry 302 February 14, 2005 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation (non proteincoding sequence) Discovered by P. Sharp
Chapter 10 - Post-transcriptional Gene Control
 Chapter 10 - Post-transcriptional Gene Control Chapter 10 - Post-transcriptional Gene Control 10.1 Processing of Eukaryotic Pre-mRNA 10.2 Regulation of Pre-mRNA Processing 10.3 Transport of mrna Across
Chapter 10 - Post-transcriptional Gene Control Chapter 10 - Post-transcriptional Gene Control 10.1 Processing of Eukaryotic Pre-mRNA 10.2 Regulation of Pre-mRNA Processing 10.3 Transport of mrna Across
Processing of RNA II Biochemistry 302. February 18, 2004 Bob Kelm
 Processing of RNA II Biochemistry 302 February 18, 2004 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation Discovered by P. Sharp and R. Roberts in late 1970s
Processing of RNA II Biochemistry 302 February 18, 2004 Bob Kelm What s an intron? Transcribed sequence removed during the process of mrna maturation Discovered by P. Sharp and R. Roberts in late 1970s
Figure mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic).
 Splicing Figure 14.3 mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic). mrna Splicing rrna and trna are also sometimes spliced;
Splicing Figure 14.3 mouse globin mrna PRECURSOR RNA hybridized to cloned gene (genomic). mouse globin MATURE mrna hybridized to cloned gene (genomic). mrna Splicing rrna and trna are also sometimes spliced;
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004
 Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 May 3, 2004 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes
 Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes Question No. 1 of 10 1. Which of the following statements about gene expression control in eukaryotes is correct? Question #1 (A)
Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes Question No. 1 of 10 1. Which of the following statements about gene expression control in eukaryotes is correct? Question #1 (A)
1. Investigate the structure of the trna Synthase in complex with a trna molecule. (pdb ID 1ASY).
 Problem Set 11 (Due Nov 25 th ) 1. Investigate the structure of the trna Synthase in complex with a trna molecule. (pdb ID 1ASY). a. Why don t trna molecules contain a 5 triphosphate like other RNA molecules
Problem Set 11 (Due Nov 25 th ) 1. Investigate the structure of the trna Synthase in complex with a trna molecule. (pdb ID 1ASY). a. Why don t trna molecules contain a 5 triphosphate like other RNA molecules
RNA Processing in Eukaryotes *
 OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
Molecular Biology (BIOL 4320) Exam #2 April 22, 2002
 Molecular Biology (BIOL 4320) Exam #2 April 22, 2002 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Molecular Biology (BIOL 4320) Exam #2 April 22, 2002 Name SS# This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses after the question number. Good
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
MCB Chapter 11. Topic E. Splicing mechanism Nuclear Transport Alternative control modes. Reading :
 MCB Chapter 11 Topic E Splicing mechanism Nuclear Transport Alternative control modes Reading : 419-449 Topic E Michal Linial 14 Jan 2004 1 Self-splicing group I introns were the first examples of catalytic
MCB Chapter 11 Topic E Splicing mechanism Nuclear Transport Alternative control modes Reading : 419-449 Topic E Michal Linial 14 Jan 2004 1 Self-splicing group I introns were the first examples of catalytic
MODULE 3: TRANSCRIPTION PART II
 MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
RECAP (1)! In eukaryotes, large primary transcripts are processed to smaller, mature mrnas.! What was first evidence for this precursorproduct
 RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
RECAP (1) In eukaryotes, large primary transcripts are processed to smaller, mature mrnas. What was first evidence for this precursorproduct relationship? DNA Observation: Nuclear RNA pool consists of
Pre-mRNA has introns The splicing complex recognizes semiconserved sequences
 Adding a 5 cap Lecture 4 mrna splicing and protein synthesis Another day in the life of a gene. Pre-mRNA has introns The splicing complex recognizes semiconserved sequences Introns are removed by a process
Adding a 5 cap Lecture 4 mrna splicing and protein synthesis Another day in the life of a gene. Pre-mRNA has introns The splicing complex recognizes semiconserved sequences Introns are removed by a process
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
Protein Synthesis
 Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
TRANSCRIPTION. DNA à mrna
 TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
TRANSCRIPTION DNA à mrna Central Dogma Animation DNA: The Secret of Life (from PBS) http://www.youtube.com/watch? v=41_ne5ms2ls&list=pl2b2bd56e908da696&index=3 Transcription http://highered.mcgraw-hill.com/sites/0072507470/student_view0/
Bio 111 Study Guide Chapter 17 From Gene to Protein
 Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins.
 Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Novel RNAs along the Pathway of Gene Expression. (or, The Expanding Universe of Small RNAs)
 Novel RNAs along the Pathway of Gene Expression (or, The Expanding Universe of Small RNAs) Central Dogma DNA RNA Protein replication transcription translation Central Dogma DNA RNA Spliced RNA Protein
Novel RNAs along the Pathway of Gene Expression (or, The Expanding Universe of Small RNAs) Central Dogma DNA RNA Protein replication transcription translation Central Dogma DNA RNA Spliced RNA Protein
MECHANISMS OF ALTERNATIVE PRE-MESSENGER RNA SPLICING
 Annu. Rev. Biochem. 2003. 72:291 336 doi: 10.1146/annurev.biochem.72.121801.161720 Copyright 2003 by Annual Reviews. All rights reserved First published online as a Review in Advance on February 27, 2003
Annu. Rev. Biochem. 2003. 72:291 336 doi: 10.1146/annurev.biochem.72.121801.161720 Copyright 2003 by Annual Reviews. All rights reserved First published online as a Review in Advance on February 27, 2003
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Where Splicing Joins Chromatin And Transcription. 9/11/2012 Dario Balestra
 Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
REGULATED AND NONCANONICAL SPLICING
 REGULATED AND NONCANONICAL SPLICING RNA Processing Lecture 3, Biological Regulatory Mechanisms, Hiten Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice site consensus sequences do have
REGULATED AND NONCANONICAL SPLICING RNA Processing Lecture 3, Biological Regulatory Mechanisms, Hiten Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice site consensus sequences do have
Translation. Host Cell Shutoff 1) Initiation of eukaryotic translation involves many initiation factors
 Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
genomics for systems biology / ISB2020 RNA sequencing (RNA-seq)
 RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER
 REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
Pol II - mrna synthesis
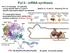 Pol II - mrna synthesis Pol II, S. cerevisiae (12 subunits) - core by specific Rpb1-3, 9 and 11 - Rpb5-6, 8, 10 and 12 - shared by Pol I-III - specific subcomplex Rpb4/7 not essential - TF: transcription
Pol II - mrna synthesis Pol II, S. cerevisiae (12 subunits) - core by specific Rpb1-3, 9 and 11 - Rpb5-6, 8, 10 and 12 - shared by Pol I-III - specific subcomplex Rpb4/7 not essential - TF: transcription
Antibodies and T Cell Receptor Genetics Generation of Antigen Receptor Diversity
 Antibodies and T Cell Receptor Genetics 2008 Peter Burrows 4-6529 peterb@uab.edu Generation of Antigen Receptor Diversity Survival requires B and T cell receptor diversity to respond to the diversity of
Antibodies and T Cell Receptor Genetics 2008 Peter Burrows 4-6529 peterb@uab.edu Generation of Antigen Receptor Diversity Survival requires B and T cell receptor diversity to respond to the diversity of
V23 Regular vs. alternative splicing
 Regular splicing V23 Regular vs. alternative splicing mechanistic steps recognition of splice sites Alternative splicing different mechanisms how frequent is alternative splicing? Effect of alternative
Regular splicing V23 Regular vs. alternative splicing mechanistic steps recognition of splice sites Alternative splicing different mechanisms how frequent is alternative splicing? Effect of alternative
Messenger RNA (mrna) is never alone
 Messenger RNA (mrna) is never alone mrna is coated and compacted by RNA-binding proteins (RBPs), forming large messenger ribonucleoprotein particles (mrnps). RBPs assemble on nascent and mature mrnas.
Messenger RNA (mrna) is never alone mrna is coated and compacted by RNA-binding proteins (RBPs), forming large messenger ribonucleoprotein particles (mrnps). RBPs assemble on nascent and mature mrnas.
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
Biochemistry 2000 Sample Question Transcription, Translation and Lipids. (1) Give brief definitions or unique descriptions of the following terms:
 (1) Give brief definitions or unique descriptions of the following terms: (a) exon (b) holoenzyme (c) anticodon (d) trans fatty acid (e) poly A tail (f) open complex (g) Fluid Mosaic Model (h) embedded
(1) Give brief definitions or unique descriptions of the following terms: (a) exon (b) holoenzyme (c) anticodon (d) trans fatty acid (e) poly A tail (f) open complex (g) Fluid Mosaic Model (h) embedded
This thesis is based on the following papers, referred to in the text by their roman numerals:
 List of papers This thesis is based on the following papers, referred to in the text by their roman numerals: I II Margaret Rush, Xiaomin Zhao, Stefan Schwartz. (2005) A splicing enhancer in the E4 coding
List of papers This thesis is based on the following papers, referred to in the text by their roman numerals: I II Margaret Rush, Xiaomin Zhao, Stefan Schwartz. (2005) A splicing enhancer in the E4 coding
Prediction and Statistical Analysis of Alternatively Spliced Exons
 Prediction and Statistical Analysis of Alternatively Spliced Exons T.A. Thanaraj 1 and S. Stamm 2 The completion of large genomic sequencing projects revealed that metazoan organisms abundantly use alternative
Prediction and Statistical Analysis of Alternatively Spliced Exons T.A. Thanaraj 1 and S. Stamm 2 The completion of large genomic sequencing projects revealed that metazoan organisms abundantly use alternative
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 NOVEMBER 2, 2006
Generation of antibody diversity October 18, Ram Savan
 Generation of antibody diversity October 18, 2016 Ram Savan savanram@uw.edu 441 Lecture #10 Slide 1 of 30 Three lectures on antigen receptors Part 1 : Structural features of the BCR and TCR Janeway Chapter
Generation of antibody diversity October 18, 2016 Ram Savan savanram@uw.edu 441 Lecture #10 Slide 1 of 30 Three lectures on antigen receptors Part 1 : Structural features of the BCR and TCR Janeway Chapter
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Viral Control of SR Protein Activity
 Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 1078 Viral Control of SR Protein Activity BY CAMILLA ESTMER NILSSON ACTA UNIVERSITATIS UPSALIENSIS UPPSALA 2001 Dissertation
Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 1078 Viral Control of SR Protein Activity BY CAMILLA ESTMER NILSSON ACTA UNIVERSITATIS UPSALIENSIS UPPSALA 2001 Dissertation
Computational Biology I LSM5191
 Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression II TRANSLATION RIBOSOMES: protein synthesizing machines Translation takes place on defined
Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture Notes: Transcriptome: Molecular Biology of Gene Expression II TRANSLATION RIBOSOMES: protein synthesizing machines Translation takes place on defined
Post-Transcriptional Regulation: Splicing
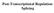 Post-Transcriptional Regulation: Splicing Pre-RNA terminal modifications Splicing Editing chemical modifications cutting capping polya base change X X pre-rrna 45S (poli) intron removal new chemical groups
Post-Transcriptional Regulation: Splicing Pre-RNA terminal modifications Splicing Editing chemical modifications cutting capping polya base change X X pre-rrna 45S (poli) intron removal new chemical groups
RNA (Ribonucleic acid)
 RNA (Ribonucleic acid) Structure: Similar to that of DNA except: 1- it is single stranded polunucleotide chain. 2- Sugar is ribose 3- Uracil is instead of thymine There are 3 types of RNA: 1- Ribosomal
RNA (Ribonucleic acid) Structure: Similar to that of DNA except: 1- it is single stranded polunucleotide chain. 2- Sugar is ribose 3- Uracil is instead of thymine There are 3 types of RNA: 1- Ribosomal
Insulin mrna to Protein Kit
 Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Objectives: Prof.Dr. H.D.El-Yassin
 Protein Synthesis and drugs that inhibit protein synthesis Objectives: 1. To understand the steps involved in the translation process that leads to protein synthesis 2. To understand and know about all
Protein Synthesis and drugs that inhibit protein synthesis Objectives: 1. To understand the steps involved in the translation process that leads to protein synthesis 2. To understand and know about all
DNA codes for RNA, which guides protein synthesis.
 Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
V24 Regular vs. alternative splicing
 Regular splicing V24 Regular vs. alternative splicing mechanistic steps recognition of splice sites Alternative splicing different mechanisms how frequent is alternative splicing? Effect of alternative
Regular splicing V24 Regular vs. alternative splicing mechanistic steps recognition of splice sites Alternative splicing different mechanisms how frequent is alternative splicing? Effect of alternative
Alternative splicing control 2. The most significant 4 slides from last lecture:
 Alternative splicing control 2 The most significant 4 slides from last lecture: 1 2 Regulatory sequences are found primarily close to the 5 -ss and 3 -ss i.e. around exons. 3 Figure 1 Elementary alternative
Alternative splicing control 2 The most significant 4 slides from last lecture: 1 2 Regulatory sequences are found primarily close to the 5 -ss and 3 -ss i.e. around exons. 3 Figure 1 Elementary alternative
Alternative Splicing and Genomic Stability
 Alternative Splicing and Genomic Stability Kevin Cahill cahill@unm.edu http://dna.phys.unm.edu/ Abstract In a cell that uses alternative splicing, the total length of all the exons is far less than in
Alternative Splicing and Genomic Stability Kevin Cahill cahill@unm.edu http://dna.phys.unm.edu/ Abstract In a cell that uses alternative splicing, the total length of all the exons is far less than in
The Multifunctional Nature of the Adenovirus L4-22K Protein
 Digital Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 1185 The Multifunctional Nature of the Adenovirus L4-22K Protein SUSAN LAN ACTA UNIVERSITATIS UPSALIENSIS UPPSALA 2016
Digital Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 1185 The Multifunctional Nature of the Adenovirus L4-22K Protein SUSAN LAN ACTA UNIVERSITATIS UPSALIENSIS UPPSALA 2016
Role of small nuclear RNAs in eukaryotic gene expression
 The Authors Journal compilation 2013 Biochemical Society Essays Biochem. (2013) 54, 79 90: doi: 10.1042/BSE0540079 Role of small nuclear RNAs in eukaryotic gene expression 6 Saba Valadkhan 1 and Lalith
The Authors Journal compilation 2013 Biochemical Society Essays Biochem. (2013) 54, 79 90: doi: 10.1042/BSE0540079 Role of small nuclear RNAs in eukaryotic gene expression 6 Saba Valadkhan 1 and Lalith
MCB 102 Third Exam Spring 2015
 MCB 102 Third Exam Spring 2015 Problem 1 Problem 2 Problem 3 Problem 4 Problem 5 Problem 6 Problem 7 Problem 8 Problem 9 Problem 10 (14 points) (9 points) (10 points) (9 points) (5 points) (6 points) (7
MCB 102 Third Exam Spring 2015 Problem 1 Problem 2 Problem 3 Problem 4 Problem 5 Problem 6 Problem 7 Problem 8 Problem 9 Problem 10 (14 points) (9 points) (10 points) (9 points) (5 points) (6 points) (7
UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME
 UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME RNA Splicing Lecture 2, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Eight essential ATPases
UNDERSTANDING THE ROLE OF ATP HYDROLYSIS IN THE SPLICEOSOME RNA Splicing Lecture 2, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Eight essential ATPases
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
Islamic University Faculty of Medicine
 Islamic University Faculty of Medicine 2012 2013 2 RNA is a modular structure built from a combination of secondary and tertiary structural motifs. RNA chains fold into unique 3 D structures, which act
Islamic University Faculty of Medicine 2012 2013 2 RNA is a modular structure built from a combination of secondary and tertiary structural motifs. RNA chains fold into unique 3 D structures, which act
Pyrimidine Tracts between the 5 Splice Site and Branch Point Facilitate Splicing and Recognition of a Small Drosophila Intron
 MOLECULAR AND CELLULAR BIOLOGY, May 1997, p. 2774 2780 Vol. 17, No. 5 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology Pyrimidine Tracts between the 5 Splice Site and Branch Point
MOLECULAR AND CELLULAR BIOLOGY, May 1997, p. 2774 2780 Vol. 17, No. 5 0270-7306/97/$04.00 0 Copyright 1997, American Society for Microbiology Pyrimidine Tracts between the 5 Splice Site and Branch Point
Received 26 January 1996/Returned for modification 28 February 1996/Accepted 15 March 1996
 MOLECULAR AND CELLULAR BIOLOGY, June 1996, p. 3012 3022 Vol. 16, No. 6 0270-7306/96/$04.00 0 Copyright 1996, American Society for Microbiology Base Pairing at the 5 Splice Site with U1 Small Nuclear RNA
MOLECULAR AND CELLULAR BIOLOGY, June 1996, p. 3012 3022 Vol. 16, No. 6 0270-7306/96/$04.00 0 Copyright 1996, American Society for Microbiology Base Pairing at the 5 Splice Site with U1 Small Nuclear RNA
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Frank Rigo and Harold G. Martinson*
 MOLECULAR AND CELLULAR BIOLOGY, Jan. 2008, p. 849 862 Vol. 28, No. 2 0270-7306/08/$08.00 0 doi:10.1128/mcb.01410-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Functional Coupling
MOLECULAR AND CELLULAR BIOLOGY, Jan. 2008, p. 849 862 Vol. 28, No. 2 0270-7306/08/$08.00 0 doi:10.1128/mcb.01410-07 Copyright 2008, American Society for Microbiology. All Rights Reserved. Functional Coupling
RNA regulons in Hox 5 UTRs confer ribosome specificity to gene regula=on
 RNA regulons in Hox 5 UTRs confer ribosome specificity to gene regula=on Shifeng Xue, Siqi Tian, Kotaro Fujii, Wipapat Kladwang, Rhiju Das & Maria Barna Nature, Volume 157, Pages 33-38 (1 st of January
RNA regulons in Hox 5 UTRs confer ribosome specificity to gene regula=on Shifeng Xue, Siqi Tian, Kotaro Fujii, Wipapat Kladwang, Rhiju Das & Maria Barna Nature, Volume 157, Pages 33-38 (1 st of January
Splicing-coupled 3 0 end formation requires a terminal splice acceptor site, but not intron excision
 Published online 8 May Nucleic Acids Research,, Vol. 4, No. 4 7 74 doi:.9/nar/gkt446 Splicing-coupled end formation requires a terminal splice acceptor site, but not intron excision Lee Davidson and Steven
Published online 8 May Nucleic Acids Research,, Vol. 4, No. 4 7 74 doi:.9/nar/gkt446 Splicing-coupled end formation requires a terminal splice acceptor site, but not intron excision Lee Davidson and Steven
Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy
 7.4 - Translation 7.4.1 - Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy Each amino acid has a specific trna-activating
7.4 - Translation 7.4.1 - Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy Each amino acid has a specific trna-activating
Chapter 5. Generation of lymphocyte antigen receptors
 Chapter 5 Generation of lymphocyte antigen receptors Structural variation in Ig constant regions Isotype: different class of Ig Heavy-chain C regions are encoded in separate genes Initially, only two of
Chapter 5 Generation of lymphocyte antigen receptors Structural variation in Ig constant regions Isotype: different class of Ig Heavy-chain C regions are encoded in separate genes Initially, only two of
SR Proteins. The SR Protein Family. Advanced article. Brenton R Graveley, University of Connecticut Health Center, Farmington, Connecticut, USA
 s s Brenton R Graveley, University of Connecticut Health Center, Farmington, Connecticut, USA Klemens J Hertel, University of California Irvine, Irvine, California, USA Members of the serine/arginine-rich
s s Brenton R Graveley, University of Connecticut Health Center, Farmington, Connecticut, USA Klemens J Hertel, University of California Irvine, Irvine, California, USA Members of the serine/arginine-rich
Bi 8 Lecture 17. interference. Ellen Rothenberg 1 March 2016
 Bi 8 Lecture 17 REGulation by RNA interference Ellen Rothenberg 1 March 2016 Protein is not the only regulatory molecule affecting gene expression: RNA itself can be negative regulator RNA does not need
Bi 8 Lecture 17 REGulation by RNA interference Ellen Rothenberg 1 March 2016 Protein is not the only regulatory molecule affecting gene expression: RNA itself can be negative regulator RNA does not need
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Abstract. a11111 RESEARCH ARTICLE
 RESEARCH ARTICLE Systematic Profiling of Poly(A)+ Transcripts Modulated by Core 3 End Processing and Splicing Factors Reveals Regulatory Rules of Alternative Cleavage and Polyadenylation Wencheng Li 1,2,
RESEARCH ARTICLE Systematic Profiling of Poly(A)+ Transcripts Modulated by Core 3 End Processing and Splicing Factors Reveals Regulatory Rules of Alternative Cleavage and Polyadenylation Wencheng Li 1,2,
Point total. Page # Exam Total (out of 90) The number next to each intermediate represents the total # of C-C and C-H bonds in that molecule.
 This exam is worth 90 points. Pages 2- have questions. Page 1 is for your reference only. Honor Code Agreement - Signature: Date: (You agree to not accept or provide assistance to anyone else during this
This exam is worth 90 points. Pages 2- have questions. Page 1 is for your reference only. Honor Code Agreement - Signature: Date: (You agree to not accept or provide assistance to anyone else during this
PROTEIN SYNTHESIS. It is known today that GENES direct the production of the proteins that determine the phonotypical characteristics of organisms.
 PROTEIN SYNTHESIS It is known today that GENES direct the production of the proteins that determine the phonotypical characteristics of organisms.» GENES = a sequence of nucleotides in DNA that performs
PROTEIN SYNTHESIS It is known today that GENES direct the production of the proteins that determine the phonotypical characteristics of organisms.» GENES = a sequence of nucleotides in DNA that performs
DEVELOPING BIOINFORMATICS TOOLS FOR THE STUDY OF ALTERNATIVE SPLICING IN EUKARYOTIC GENES LIM YUN PING
 DEVELOPING BIOINFORMATICS TOOLS FOR THE STUDY OF ALTERNATIVE SPLICING IN EUKARYOTIC GENES LIM YUN PING SCHOOL OF MECHANICAL & PRODUCTION ENGINEERING 2005 TOOLS FOR STUDY OF ALTERNATIVE SPLICING LIM YUN
DEVELOPING BIOINFORMATICS TOOLS FOR THE STUDY OF ALTERNATIVE SPLICING IN EUKARYOTIC GENES LIM YUN PING SCHOOL OF MECHANICAL & PRODUCTION ENGINEERING 2005 TOOLS FOR STUDY OF ALTERNATIVE SPLICING LIM YUN
Elongation and pre-mrna processing RNAPII
 Elongation and pre-mrna processing RNAPII Introduction Themes Stage 1 - first steps - RNA polymerase II is released Stage 2 - promoter proximal events - pausing, checkpoint control Stage 3 - productive
Elongation and pre-mrna processing RNAPII Introduction Themes Stage 1 - first steps - RNA polymerase II is released Stage 2 - promoter proximal events - pausing, checkpoint control Stage 3 - productive
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
GENOME-WIDE COMPUTATIONAL ANALYSIS OF SMALL NUCLEAR RNA GENES OF ORYZA SATIVA (INDICA AND JAPONICA)
 GENOME-WIDE COMPUTATIONAL ANALYSIS OF SMALL NUCLEAR RNA GENES OF ORYZA SATIVA (INDICA AND JAPONICA) M.SHASHIKANTH, A.SNEHALATHARANI, SK. MUBARAK AND K.ULAGANATHAN Center for Plant Molecular Biology, Osmania
GENOME-WIDE COMPUTATIONAL ANALYSIS OF SMALL NUCLEAR RNA GENES OF ORYZA SATIVA (INDICA AND JAPONICA) M.SHASHIKANTH, A.SNEHALATHARANI, SK. MUBARAK AND K.ULAGANATHAN Center for Plant Molecular Biology, Osmania
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
BIO 5099: Molecular Biology for Computer Scientists (et al)
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
RECAP FROM MONDAY AND TUESDAY LECTURES
 RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Cytopathogenesis and Inhibition of Host Gene Expression by RNA Viruses
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Dec. 2000, p. 709 724 Vol. 64, No. 4 1092-2172/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Cytopathogenesis and Inhibition
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Dec. 2000, p. 709 724 Vol. 64, No. 4 1092-2172/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Cytopathogenesis and Inhibition
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Spliceosome Pathway. is required for a stable interaction between U2 snrnp and
 MOLECULAR AND CELLULAR BIOLOGY, Sept. 1988, p. 3755-3760 Vol. 8, No. 9 0270-7306/88/093755-06$02.00/0 Copyright 1988, American Society for Microbiology Early Commitment of Yeast Pre-mRNA to the Spliceosome
MOLECULAR AND CELLULAR BIOLOGY, Sept. 1988, p. 3755-3760 Vol. 8, No. 9 0270-7306/88/093755-06$02.00/0 Copyright 1988, American Society for Microbiology Early Commitment of Yeast Pre-mRNA to the Spliceosome
Interaktionen von RNAs und Proteinen
 Sonja Prohaska Computational EvoDevo Universitaet Leipzig May 5, 2017 RNA-protein binding vs. DNA-protein binding Is binding of proteins to RNA the same as binding of proteins to DNA? Will the same rules
Sonja Prohaska Computational EvoDevo Universitaet Leipzig May 5, 2017 RNA-protein binding vs. DNA-protein binding Is binding of proteins to RNA the same as binding of proteins to DNA? Will the same rules
Identification of Novel Compounds That Inhibit HIV-1 Gene Expression by Targeting Viral RNA Processing
 Identification of Novel Compounds That Inhibit HIV-1 Gene Expression by Targeting Viral RNA Processing by Ahalya Balachandran A thesis submitted in conformity with the requirements for the degree of Master
Identification of Novel Compounds That Inhibit HIV-1 Gene Expression by Targeting Viral RNA Processing by Ahalya Balachandran A thesis submitted in conformity with the requirements for the degree of Master
Investigating the mechanism of alternative splicing regulation of the RNA-binding proteins T-STAR and Sam68
 Investigating the mechanism of alternative splicing regulation of the RNA-binding proteins T-STAR and Sam68 Marina Danilenko Institute of Genetic Medicine A thesis submitted for the degree of Doctor of
Investigating the mechanism of alternative splicing regulation of the RNA-binding proteins T-STAR and Sam68 Marina Danilenko Institute of Genetic Medicine A thesis submitted for the degree of Doctor of
MODULE 4: SPLICING. Removal of introns from messenger RNA by splicing
 Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
Central Dogma. Central Dogma. Translation (mrna -> protein)
 Central Dogma Central Dogma Translation (mrna -> protein) mrna code for amino acids 1. Codons as Triplet code 2. Redundancy 3. Open reading frames 4. Start and stop codons 5. Mistakes in translation 6.
Central Dogma Central Dogma Translation (mrna -> protein) mrna code for amino acids 1. Codons as Triplet code 2. Redundancy 3. Open reading frames 4. Start and stop codons 5. Mistakes in translation 6.
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being A Eukaryote. Eukaryotic Cells. Basic eukaryotes have:
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 15: Being a Eukaryote: From DNA to Protein, A Tour of the Eukaryotic Cell. Christiaan van Woudenberg Being A Eukaryote Basic eukaryotes
Complete Student Notes for BIOL2202
 Complete Student Notes for BIOL2202 Revisiting Translation & the Genetic Code Overview How trna molecules interpret a degenerate genetic code and select the correct amino acid trna structure: modified
Complete Student Notes for BIOL2202 Revisiting Translation & the Genetic Code Overview How trna molecules interpret a degenerate genetic code and select the correct amino acid trna structure: modified
U2 toggles iteratively between the stem IIa and stem IIc conformations to promote pre-mrna splicing
 U2 toggles iteratively between the stem IIa and stem IIc conformations to promote pre-mrna splicing Angela K. Hilliker, 1 Melissa A. Mefford, 2 and Jonathan P. Staley 1,3 1 Department of Molecular Genetics
U2 toggles iteratively between the stem IIa and stem IIc conformations to promote pre-mrna splicing Angela K. Hilliker, 1 Melissa A. Mefford, 2 and Jonathan P. Staley 1,3 1 Department of Molecular Genetics
Sec$on 8. Gene$c Muta$ons, Ribosome Structure and the Tetracyclines
 Sec$on 8 Gene$c Muta$ons, Ribosome Structure and the Tetracyclines Explain the mechanism of ac$on of the transcrip$onal and transla$onal inhibitors Requires knowledge of: Sec$on 6 Sec$on 7 Sec$on 8 How
Sec$on 8 Gene$c Muta$ons, Ribosome Structure and the Tetracyclines Explain the mechanism of ac$on of the transcrip$onal and transla$onal inhibitors Requires knowledge of: Sec$on 6 Sec$on 7 Sec$on 8 How
MCB Test 1 Mueckler Review
 MCB Test 1 Mueckler Review 9-30-17 Func%ons of Cellular Membranes 1. Plasma membrane acts as a selec%vely permeable barrier to the environment Uptake of nutrients Waste disposal Maintains intracellular
MCB Test 1 Mueckler Review 9-30-17 Func%ons of Cellular Membranes 1. Plasma membrane acts as a selec%vely permeable barrier to the environment Uptake of nutrients Waste disposal Maintains intracellular
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
The U1 snrnp Base Pairs with the 5 Splice Site within a Penta-snRNP Complex
 MOLECULAR AND CELLULAR BIOLOGY, May 2003, p. 3442 3455 Vol. 23, No. 10 0270-7306/03/$08.00 0 DOI: 10.1128/MCB.23.10.3442 3455.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
MOLECULAR AND CELLULAR BIOLOGY, May 2003, p. 3442 3455 Vol. 23, No. 10 0270-7306/03/$08.00 0 DOI: 10.1128/MCB.23.10.3442 3455.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved.
