PDCD5-Regulated Cell Fate Decision after Ultraviolet-Irradiation-Induced DNA Damage
|
|
|
- Garry Daniels
- 5 years ago
- Views:
Transcription
1 2582 Biophysical Journal Volume December PDCD5-Regulated Cell Fate Decision after Ultraviolet-Irradiation-Induced DNA Damage Changjing Zhuge, Ying Chang, Yanjun Li, Yingyu Chen, and Jinzhi Lei * Zhou Pei-Yuan Center for Applied Mathematics, Tsinghua University, Beijing, China; and Laboratory of Medical Immunology, School of Basic Medical Science, Peking University, Beijing, China ABSTRACT Programmed cell death 5 (PDCD5) is a human apoptosis-related molecule that is involved in both the cytoplasmic caspase-3 activity pathway (by regulating Bax translocation from cytoplasm to mitochondria) and the nuclear pathway (by interacting with Tip6). In this study, we developed a mathematical model of the PDCD5-regulated switching of the cell response from DNA repair to apoptosis after ultraviolet irradiation-induced DNA damage. We established the model by combining several hypotheses with experimental observations. Our simulations indicate that the ultimate cell response to DNA damage is dependent on a signal threshold mechanism, and the PDCD5 promotion of Bax translocation plays an essential role in PDCD5-regulated cell apoptosis. Furthermore, the model simulations revealed that PDCD5 nuclear translocation can attenuate cell apoptosis, and PDCD5 interactions with Tip6 can accelerate DNA damage-induced apoptosis, but the final cell fate decision is insensitive to the PDCD5-Tip6 interaction. These results are consistent with experimental observations. The effect of recombinant human PDCD5 was also investigated and shown to sensitize cells to DNA damage by promoting caspase-3 activity. INTRODUCTION DNA damage is a high-frequency event that occurs in living cells. A mammalian genome undergoes ~, modifications per day, each of which results in a finite probability of residual damage (). These damaging events can corrupt the genetic information and threaten the health of an organism. Therefore, a complex network of DNA damage responses (DDRs) has evolved to sense and respond to DNA damage (2). In multicellular organisms, activation of DDR results in two primary responses: DNA repair and genomic restoration or, if the damaged DNA cannot be sufficiently repaired, execution of a cell death program such as apoptosis (3 6). The molecular mechanism by which a cell makes the decision to promote survival (i.e., DNA repair) or death (i.e., by triggering the apoptosis program) is not yet clear and remains a challenging issue in cell biology (2). Ultraviolet (UV) irradiation is commonly used in the laboratory to induce DNA damage that can be detected by the ataxia telangiectasia Rad3 related (ATR) protein, a member of the ataxia telangiectasia mutated (ATM) family of DNA damage sensors (7). ATM proteins bind directly to free DNA ends and catalyze phosphorylation cascades to transmit damage signals to cell cycle checkpoints and repair proteins (8). Pathways downstream of the damage signals interact with each other to regulate cell responses. Among the downstream effects, the sumoylation of Tip6 and upregulation of programmed cell death 5 (PDCD5) are crucial for cell fate decisions, and both are associated with p53 signaling (9,). p53 is a core regulator of the decision between apoptosis and other fates in response to DNA damage (,2). ATM Submitted June 6, 2, and accepted for publication October 28, 2. *Correspondence: jzlei@mail.tsinghua.edu.cn Editor: Leah Edelstein-Keshet. proteins and other checkpoint activations lead to phosphorylation of p53, which greatly increases its stability (2). Several amino acids of p53 proteins are consistently phosphorylated and acetylated in response to different signals, which then induce different downstream pathways, including DNA repair, cell cycle arrest, and apoptosis (3). Acetylation of p53 at lysine 2 (K2) can increase the expression of the proapoptotic Bcl-2 family member protein Bax. Phosphorylation of p53 at serine 2 (S2) allows p2 protein synthesis and subsequent G checkpoint activation. p53 proteins can induce Mdm2, which in turn negatively affects p53 levels, resulting in a negative feedback loop (4). This feedback loop generates oscillations in both p53 and Mdm2 levels that are important in the mediation of the DDR. As mentioned above, Bax is a proapoptotic member of the Bcl-2 family that is induced by p53 acetylation at K2. After synthesis, Bax proteins are translocated to the outer mitochondrial membrane, where they form channels or even large holes for the release of cytochrome c into the cytosol from the mitochondria (5). Once released, cytochrome c associates with Apaf- (Apaf) and pro-caspase-9 to form apoptosomes. The apoptosome generates active caspase-9, which then cleaves pro-caspase-3 to active caspase-3 (5). Caspase-3 inactivates PARP and DNA-PK, two key enzymes involved in the homeostatic maintenance of genomic integrity, to disable the normal DNA repair process and thereby induce apoptosis (6). Tip6 is a histone acetyltransferase (HAT) that is involved in the cellular response to DNA damage (9). In response to DNA double-stranded breaks, Tip6 is recruited to DNA lesions to participate in both the initial and final stages of repair (7). Tip6 is also required for acetylation of the endogenous p53 protein at K2, and therefore the Ó 2 by the Biophysical Society //2/2582/ $2. doi:.6/j.bpj.2..44
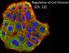
2 PDCD5-Regulated Cell Fate Decision 2583 p53-dependent induction of proapoptotic target genes, such as Bax, in response to DNA damage (8). Upon UV-irradiation-induced DNA damage, the initial step of this pathway involves site-specific sumoylation of Tip6 at lysines 43 (K43) and 45 (K45) via Ubc9. This sumoylation is thought to be essential for DDR via a p53-dependent pathway, because the inhibition of Tip6 sumoylation is known to abrogate the p53-dependent DDR (9). PDCD5 (formerly referred to as the TF- cell apoptosisrelated gene 9 (TFAR9)) is a novel gene that is known to be related to the regulation of apoptosis (9). Decreased expression of PDCD5 has been detected in various human tumors, including astrocytic glioma (2), chondrosarcoma (2), gastric cancer (22), lung cancer (23), and myelogenous leukemia (24). A genome-wide study ranked the PDCD5 gene as the 47th most significant gene among a pool of expressed genes in tumors (25), and overexpression of PDCD5 in tumor cells has been found to enhance apoptosis (9). The PDCD5 protein is apparently upregulated in cells upon apoptotic stimulation (9), enhances caspase-3 activity by modulating Bax translocation from the cytosol to the mitochondrial membrane (26), and then translocates rapidly from the cytoplasm to the nucleus by a yet unknown mechanism (27). The latter event of PDCD5 nuclear translocation is suggested to be an early marker for apoptosis, preceding the externalization of phosphatidylserine and fragmentation of chromosome DNA, and occurs in parallel with the loss of mitochondrial membrane potential (27). This phenomenon is independent of both cell type and apoptosis-inducing stimuli (27). There are at least two known pathways through which PDCD5 regulates cellular apoptosis. In the cytoplasm, PDCD5 exerts its effects through the mitochondrial pathway by modulating Bax translocation, cytochrome c release, and caspase-3 activation, either directly or indirectly (26). Cells with decreased expression of PDCD5 by short interfering RNA against PDCD5 (sipdcd5) display reduced sensitivity to an apoptotic stimulus by Bax overexpression. Knockdown of PDCD5 expression in cells is known to result in the attenuation of the cytochrome c released from the mitochondria to the cytosol, and the inhibition of both caspase-3 activity and pro-caspase-3 cleavage (26). Another pathway of cellular apoptosis occurs in the nucleus, where PDCD5 interacts with Tip6 and functions as a cooperator in promoting Tip6 HAT activity and DNA damage-induced apoptosis (). PDCD5 can bind to Tip6 and enhance the stability of Tip6 proteins. The level of binding is significantly increased after UV irradiation, which enhances the HAT activity of Tip6 and Tip6-dependent histone acetylation. Furthermore, PDCD5 increases Tip6- dependent K2 acetylation of p53 and participates in the p53-dependent expression of apoptosis-related genes (). Because the decreased expression of PDCD5 has been characterized for some human tumors, recent studies have focused on the potential clinical management of tumors by recombinant human PDCD5 (rhpdcd5) (28 33). RhPDCD5 was shown to enter a variety of cells by clathrinindependent endocytosis (3), significantly facilitating tumor cell apoptosis triggered by chemotherapy (3,32). To understand the effect of rhpdcd5 therapy, we applied our computational model in this study to quantitatively assess the influence of PDCD5 pathways on the DDR. Although the PDCD5 protein has been studied for more than years, and many interactions are known to be involved in the DDR pathway, questions remain about the importance of these interactions and how they cross talk to regulate the cell fate decision. Our goal in this study was to answer these questions with the use of a computational model. Numerous studies have described computational models of apoptosis pathways and DDRs (6,34 44). However, most of these studies focused on either the cytoplasmic pathway or p53 dynamics in the cellular response to DNA damage, and little work has been performed for other parts in the nuclear pathway. To the best of our knowledge, no mathematical model has been presented for the PDCD5 regulatory pathways in this regard. In this work, we attempted to establish such a model by integrating known experimentally supported functions of PDCD5, and to assess the importance of each PDCD5 interaction in the DDR pathways, as well as the connections between these interactions. To that end, we developed a mathematical model for PDCD5-regulated cell fate decisions in response to UV-irradiation-induced DNA damage. In establishing the mathematical model, we formed hypotheses in accordance with experimental observations of the regulation of PDCD5 nuclear translocation, PDCD5 interactions with Tip6, and PDCD5-mediated regulation of Bax translocation. We tested these hypotheses by comparing our model simulations with experimental evidence. Using the developed model, we discuss mechanisms by which PDCD5 regulates DNA damage-induced apoptosis. This work will provide the basis for further investigations of PDCD5 regulation in the DDR. MODEL AND METHODS Model description A schematic representation of the PDCD5 signaling network in response to UV-irradiation-induced DNA damage is shown in Fig.. Our goal in this study was to establish a model of the PDCD5 signaling pathway to understand how PDCD5 regulates cell fate. Therefore, we omitted other signaling pathways that may also be relevant for the DDR, but whose interconnections with PDCD5 regulation are not yet clear. For example, we omitted the p53-mdm2 regulation and simply assumed a time-dependent p53 dynamics after DNA damage (see the Supporting Material and Fig. S for discussions). In our model, we considered cellular responses to UV-irradiationinduced DNA damage, which is represented by ATR activity through the following mechanism (45,46): The multisubunit replication protein (RPA) Biophysical Journal ()
3 2584 Zhuge et al. FIGURE Schematic representation of the PDCD5 signaling network in response to UV-irradiation-induced DNA damage. State transitions and expressions are represented by arrowheaded solid lines, and the promotions of state transition or cell response are denoted by circle-headed lines. Inhibitions of protein activity or cell response are represented by bar-headed lines. Translocations are denoted by open arrowheaded dashed lines. Effects represented by the black lines are based on experimental data and known facts from literatures. The blue lines indicate assumptions in the current study (see text for details). complex has a high affinity for single-stranded DNA (ssdna), which is a common intermediate in the processing of many types of damaged DNA, including double-strand breaks (DSBs), UV-induced thymidine dimers, and mismatches in basepairing. Recruited by ssdna-rpa complexes at damage sites, ATR phosphorylates signal sensors/transducers (such as Rad7 or a complex of Rad9p, Husp, and Radp), leading to downstream responses of cell cycle arrest and DNA repair. For this reason, relocalization of ATR to sites of damage has been the proposed mode of ATR activation (45). ATR activation transmits DNA damage signals and triggers multiple damage-response pathways (2). In response to these signals, the expression of Tip6 and PDCD5 is rapidly upregulated, and p53 levels also increase within minutes after damage. The initial steps of the Tip6 regulation pathways involve the association of Tip6 with SUMO-conjugation enzymes and site-specific sumoylation of Tip6 at K43 and K45 (9). Tip6 sumoylation promotes its HAT activity, which in turn induces DNA repair. Furthermore, Tip6 sumoylation recruits p53 to the promyelocytic leukemia (PML) nuclear bodies to promote the acetylation of p53 protein at K2 and phosphorylation at S2, leading to expression of the proapoptotic protein Bax and the G arrest pathway, respectively (9,8). The G arrest pathway was omitted from this study. PDCD5 protein is weakly expressed and nearly evenly distributed in both the cytoplasm and nucleus (27). After the onset of an apoptotic stimulus, the cytoplasmic PDCD5 protein level first increases rapidly and then forms an inward gradient from the cytoplasm to the nucleus (,9,27). This pattern is maintained for a few hours (with the duration dependent on the type of cells and apoptosis stimulus) until the cytoplasmic PDCD5 proteins are triggered to translocate and accumulate in the nucleus (27). As the PDCD5 level in the nucleus becomes significantly higher than that in the cytoplasm, the concentration gradient of PDCD5 inverts. The following two observations are supported experimentally (27) (Fig. S2): ), the nucleus/cytoplasm ratio of the PDCD5 level is evidently < at the early stage and > later on; and 2), the cytoplasmic PDCD5 level is obviously decreased after nuclear translocation. We assume that the regulation of PDCD5 nuclear translocation is incorporated in the form of a hypothetical intermediary that is activated by a Tip6 downstream signal, and PDCD5 interactions with Tip6 can enhance the translocation activity (Fig. S3). Our model includes two PDCD5 signaling pathways. In the cytoplasmic pathway, PDCD5 increases the cleavage of pro-caspase-3 via enhancement of Bax translocation from the cytosol to the mitochondrial outer membrane (26). In the nucleus, PDCD5 proteins colocalize with Tip6, enhance the stability of Tip6 (Fig. S5), and promote both Tip6 HAT activity and K2 acetylation of p53 (Fig. S4) (). Accordingly, we assume in our model that the PDCD5 protein binds to Tip6, which increases its stability and HAT activity, as well as the Tip6-induced K2 acetylation of p53. Therefore, the PDCD5 protein is involved in both DNA repair and apoptotic pathways. Two feedback loops are shown in the model given in Fig.. When DNA repair is triggered, the cells begin the repair processes, which decrease the ATR activity and damage signals, resulting in a negative feedback loop. The other feedback loop comes from caspase-3, which is activated through increased Bax, cytochrome c release, and caspase-9 activation. The active caspase-3 disables normal DNA repair processes by inactivating PARP and DNA-PK (6). These regulatory processes form a positive feedback of DNA damage after UV irradiation. To summarize the above explanations, Biophysical Journal ()
4 PDCD5-Regulated Cell Fate Decision 2585 we argue that there are essentially two signaling pathways involving PDCD5 proteins that lead to a cell fate decision after DNA damage: the nuclear pathway that is controlled by the interaction of PDCD5 with Tip6, and the cytoplasmic pathway that is controlled by PDCD5-regulated Bax translocation. Moreover, the nuclear pathway is involved in both feedback loops, whereas the cytoplasmic pathway only contributes to the positive feedback. Formulations The model in Fig. can be formulated as a set of differential equations for the dynamics of molecular concentrations (or activities). There are five variables in our differential equation model: the concentrations of PDCD5 proteins in the nucleus [P] nu and cytoplasm [P] cyt, sumoylated Tip6 [T], the Tip6-PDCD5 complex (in nucleus) [TP], and ATR activity [A]. The dynamic processes of these quantities are modeled by a set of five ordinary differential equations. In addition, the total p53 protein concentration is assumed to be an increase function of the ATR activity. Other quantities in the model, including the HAT activity [H], p53 acetylation level [p53], concentration of Bax protein [Bax], and caspase-3 activity [C3], are assumed to depend on [P] nu, [P] cyt, [T], [TP], and [A] instantaneously by applying quasi-equilibrium assumptions. In the differential equations, the following biological processes are modeled: increased expression of Tip6 and PDCD5 after DNA damage, increased p53 level in response to DNA damage, nuclear translocation of PDCD5, cross nuclear membrane diffusion of PDCD5, binding and dissociation of Tip6 and PDCD5 in the nucleus, degradation of the molecules, and regulation of ATR activity through the DNA repair and apoptosis pathways. The complete model equations are given in the Supporting Material, with the following key assumptions taken into consideration: The production rates of Tip6 and PDCD5 are increase functions of the ATR activity [T]. The ATR activation rate is proportional to the degree of DNA damage, and decreases with respect to the Tip6 HAT activity and increases with caspase-3 activity. PDCD5 binds to sumoylated Tip6 to promote its HAT activity. PDCD5 binds to sumoylated Tip6 to promote K2 acetylation of p53. Furthermore, the total p53 level increases with ATR activity. In the cytoplasm pathway, PDCD5 promotes caspase-3 activity by regulating Bax translocation from the cytoplasm to the mitochondrial outer membrane, and Bax expression is upregulated by the K2 acetylated of p53. PDCD5 nuclear translocation is regulated by a Tip6 downstream signal, and PDCD5 interactions with Tip6 can enhance this translocation. Parameter estimation We estimated the model parameters according to available experimental results for UV-irradiation-induced apoptosis in U2OS cells (Fig. S4 and Fig. S5), such that our model yields a bistable response, i.e., a cell response to DNA damage with either success or failure in repairing the DNA sequence depending on stimulus strengths (Fig. S6). Details are provided in the Supporting Material. METHODS Numerical simulation In numerical simulations, we solved the model equations with the NDSolve function on the Mathematica 6. platform (47). Initial conditions were set to the steady state before the stimulus was applied. In each case, we solved the equations to t ¼ 4 h until the solutions reached a steady state, to mimic a UV-irradiation treatment of 4 h. Multicell simulation To simulate various responses of a group of cells, which are essentially different from each other, we modeled each individual cell by varying each model parameter randomly and independently within a range of 52% of its default value (as given by Table S). We kept the Hill coefficients unchanged because they usually relate to the qualitative properties of the interactions, which are the same for different cells of the same type. Thus, for a group of cells, we had an ensemble of sets of different parameter values. For each set of parameters, we solved the model equation with the above numerical method to obtain the response of a particular cell. The apoptosis probability could therefore be obtained by statistical calculations based on the simulation results of this group of cells. This procedure is described as multicell simulation. When we studied the dependence of the apoptosis probability on a particular parameter, we applied a multicell simulation in which the value of the given parameter was fixed while other parameters were randomly changed. RESULTS Threshold triggering of a cellular response to DNA damage To determine how cells respond to DNA damage with stimuli of different strengths, we numerically solved the model equations with stimulus strength v A varied from 4 to 2 2. The resulting time courses of ATR and caspase-3 activities are shown in Fig. 2, a and b. The results show a significant threshold mechanism of DDR. When the stimulus was low, ATR and caspase-3 activities were low at 4 h, indicating successful DNA repair. There was an obvious threshold (at approximately v A ¼.25) of the stimulus strength, such that when the stimulus was beyond the threshold, ATR and caspase-3 activities were high at 4 h, indicating a failure of DNA repair and initiation of apoptosis. Fig. 2 a shows obvious two-phase dynamics in ATR activity when the stimulus strength was high. The first phase occurred in the early stage, when ATR activities increased due to the appearance of damage signals, followed by a decrease in response to DNA repair. When the stimulus strength was low, DNA repair signals dominated the cellular responses, such that ATR activity decreased to a steady lowlevel state. However, when the stimulus strength was high, a second phase of increased ATR activity appeared in parallel with the increase of caspase-3 activity (Fig. 2 b). These results suggest that the second phase of ATR induction was due to the increase in caspase-3 activity, which in turn inhibited the DNA repair. A two-phase dynamic profile was also seen under strong stimulation in a recent study of a p53 network model of DDR (43). It is not clear whether there are any connections between these two-phase dynamics, and this issue requires further studies and experimental validation. Caspase-3 activation is usually considered as a marker of apoptosis induction, and therefore its activity can be used to characterize the probability of apoptosis. Fig. 2 c shows caspase-3 activity at 4 h as a function of stimulus strength. Biophysical Journal ()
5 ccaspase 3 activity 2586 Zhuge et al. a ATR activity v A Time (h) control sipdcd Stimulus strength (v ) A Stimulus threshold (v A ) b Caspase 3 activity d v A Time (h).2. v 2 k on α k 2 3 Ratio to the default value FIGURE 2 Simulation of cellular response to DNA damage with a threshold mechanism. (a) Time courses of ATR activities with different stimulus strengths (v A ). The arrow shows the direction of increasing v A from 4 to 2 with a step of 4 4 (same as in b). (b) Time courses of caspase-3 activities with different stimulus strengths. (c) Caspase-3 activity at 4 h after stimulation as a function of stimulus strength for control cells (v 2 ¼.39 3 ) and sipdcd5-transfected cells (v 2 ¼ ) are superimposed on the activity at steady state (lines). Other parameters are the same as described in Table S. (d) Dependence on stimuli thresholds (v A ) for induction of apoptosis with varying PDCD5 expression rate (v 2 ), binding rate of PDCD5 to Tip6 (k on ), efficiency of regulating Bax translocation by PDCD5 (a), and nuclear translocation rate (k ). The horizontal axis shows the ratio of each value to the corresponding default value. In each case, other parameters were taken as their default values. The threshold is defined as the value of stimulus strength v A such that caspase-3 activity is.5 at 4 h of stimulus. The result shows a clear transition from low to high activity at approximately v A ¼.25. We also examined the activity of sipdcd5-transfected cells in which the PDCD5 expression rate was reduced to one-third as in wild-type cells. A transition from low to high was also seen in sipdcd5 cells, but the threshold (approximately v A ¼.8) was much larger than that for wild-type cells. This result indicates that decreased expression of PDCD5 can inhibit apoptosis, in agreement with previously reported experiments (,9,26). Furthermore, we examined how the stimulus threshold for apoptosis induction depends on PDCD5 regulations. To that end, we individually changed the PDCD5 expression rate (v 2 ), binding rate of PDCD5 to Tip6 (k on ), efficiency of regulating Bax translocation by PDCD5 (a), and PDCD5 nuclear translocation rate (k ) (of course, keeping other parameters unchanged) to find the corresponding threshold (Fig. 2 d). The results show that the threshold decreased significantly when either v 2 or a increased, and increased with k, but was insensitive to changes in k on. These results suggest that the well-tuned PDCD5 expression rate v 2 and regulation efficiency a are important for cell response. Note that in our model, the PDCD5 cytoplasmic pathway is described by the parameter a, and the PDCD5 nuclear pathway is described by k on. The above results indicate that the apoptotic threshold is more sensitive to the PDCD5 cytoplasmic pathway than to the PDCD5 nuclear pathway. In Fig. 2 c, we note that for wild-type cells, the threshold for inducing apoptosis (v A ¼.25) was consistent with the bifurcation point of the high-to-low switch obtained from steady-state analysis (point A in Fig. S6, v A ¼.25) rather than the bifurcation point of the low-tohigh switch (point B in Fig. S6, v A ¼.97), as we expected. For sipdcd5-transfected cells, the threshold was between the two bifurcation points from the steady-state analysis (Fig. 2 c). These observations differ from what is known from bistability analysis, in which the threshold should be consistent with the bifurcation at a low-to-high switch (34,35,38). We argue that this is due to the appearance of the second phase of increased ATR activity, as discussed above, which could switch the cell response from DNA repair (low ATR and caspase-3 activities) to apoptosis (high ATR and caspase-3 activities), although the state of DNA repair was stable. Such a switch can be seen in time courses of molecular concentrations/activities (Fig. S7) in which the rapid increase in caspase-3 activity at ~2 h of simulation was evident, triggering apoptosis and stopping the DNA repair process. Experimental observations of such a rapid pulse-like increase in caspase-3 activity in p53-induced apoptosis were previously reported (48,49). PDCD5 enhancement of caspase-3 activity is required for DNA-damage-induced apoptosis PDCD5 is known to promote apoptosis by modulating Bax translocation, cytochrome c release, and caspase-3 activation (26). Here, using our model, we investigated how PDCD5 enhancement of caspase-3 activity would affect the cell response after DNA damage. Biophysical Journal ()
6 PDCD5-Regulated Cell Fate Decision 2587 In our model, we formulated the caspase-3 activity as a Hill-type function of Bax concentration, with the halfmaximal effective concentration (EC5) dependent on cytoplasmic PDCD5 concentration (see Supporting Material): ½BaxŠ n ½C3Š ¼c max K n þ½baxš n ; K m K ¼ m : () þ a½pš cyt Here the coefficient a represents the efficiency of cytoplasmic PDCD5 proteins, with a larger a-value indicating higher efficiency. In previous simulations, the stimulus threshold to induce apoptosis sensitively depended on the efficiency a, with larger a-values yielding lower thresholds (Fig. 2 d). To further investigate the role of PDCD5 regulation, we changed the efficiency a from zero to double the default value, and solved the equations. Fig. 3 a shows the simulation results for ATR and caspase-3 activity dynamics with different efficiencies. In contrast to the two-phase dynamics of cell response to DNA damage described above, here, when the efficiency a was small, ATR activity increased only once at the early stage (Fig. 3 a). When the efficiency was large, however, the ATR activity displayed the second phase of increase, in parallel with the increase of caspase- 3 activity, and both remained at a high-level steady state. These results suggest that the first induction phase of ATR activity was independent of the efficiency a, whereas the second phase was promoted by the increased efficiency. Fig. 3 b shows the simulation result of the timing of the second phase of ATR activity when it first crossed 3., as a function of the efficiency a. There was a minimum efficiency (a ¼.72), such that the second phase would not appear if a was smaller than this value; otherwise, the second phase would occur, the timing of which decreased with increasing a. As noted above, the second increase phase was required to switch the cell response from DNA repair to apoptosis. These results suggest that PDCD5 a 5 ATR activity 2 Caspase-3 activity α α Time (h) α FIGURE 3 Simulated cell responses with different efficiencies of PDCD5 regulating Bax translocation. (a) Time course of ATR activity (upper panel) and caspase-3 activity (lower panel). Arrows show directions of increasing a from to 4 with a step of.4. The dashed line shows ATR activity of 3. (an indicator of the second phase of ATR induction). (b) Dependence of the timing of the second phase activity on the efficiency a (see text for details). Stimulus strength v A ¼.26. b Second phase increasing time (h) modulation of caspase-3 activation in the cytoplasmic pathway is essential for DNA-damage-induced cell apoptosis. PDCD5 nuclear translocation can attenuate DNA-damage-induced cell apoptosis Previous experimental findings suggested that PDCD5 nuclear translocation is an early event for apoptosis (27). However, the role of this event in the cellular response to DNA damage (in particular, whether it is an apoptosis trigger or an early consequent signal of apoptosis) remains unclear. In previous simulations, we showed that increasing the rate of PDCD5 nuclear translocation yielded a higher stimulus threshold for apoptosis induction (Fig. 2 d). To further investigate the reason for this dependence, we studied the dynamic behaviors of cytoplasmic PDCD5 concentrations and caspase-3 activity while varying the PDCD5 nuclear translocation rate (Fig. 4 a). A direct consequence of PDCD5 nuclear translocation was a decrease in the cytoplasmic PDCD5 concentration, which in turn attenuated caspase-3 activity. When the translocation was off (k ¼ ), the cytoplasmic PDCD5 level increased with time in response to DNA damage, thereby sustaining the caspase-3 activity at a high level. When k was increased, the cytoplasmic PDCD5 level decreased after the activation of its nuclear translocation, causing the caspase-3 activity to be reduced to a low level (Fig. 4 a). To further validate the above observation, we calculated the apoptosis probability using the multicell simulation method. Fig. 4 b shows the obtained apoptosis probability as a function of k, which decreased with increasing k. These results suggest that PDCD5 nuclear translocation may not be necessary for DNA-damage-induced apoptosis, and increasing the a Cytoplasmic PDCD5 2 Caspase-3 activity k k Time (h) b Apoptosis probability k (ratio to the default value) FIGURE 4 Simulation results for cellular responses with different values of PDCD5 nuclear translocation rate. (a) Time courses of cytoplasmic PDCD5 concentration (upper panel) and caspase-3 activity (lower panel), with k ¼ 4 4 (down triangles), k ¼ 2 4 (up triangles), and k ¼ (circles). Arrows show the directions of increasing k.(b) Simulation result of apoptosis probability as a function of k. The probability was obtained by the multicell simulation method (see Materials and Methods for details). A cell was identified as apoptotic if its caspase-3 activity was >.5 at 4 h of simulation. Stimulus strength v A ¼.26. Biophysical Journal ()
7 acaspase-3 activity 2588 Zhuge et al. translocation rate can instead attenuate apoptosis by decreasing the cytoplasmic PDCD5 concentration. We speculate that PDCD5 nuclear translocation is not a triggering signal for apoptosis, and is more likely to be an early subsequent event of apoptosis. DNA-damage-induced apoptosis is unlikely sensitive to PDCD5 interactions with Tip6 Previous experimental results showed that PDCD5 interacts with Tip6 and functions as a cooperator in acetyltransferase activity and DNA-damage-induced apoptosis (). To examine the effects of PDCD5 interactions with Tip6, we changed the association rate k on of PDCD5 binding to Tip6 and solved the model equations. Fig. 5 a shows the simulated caspase-3 activity with k on increased from zero to double the default value. These results revealed that turning off the PDCD5 interaction could attenuate apoptosis, and increasing the binding rate could accelerate the increasing phase of caspase-3 activity, thereby accelerating DNA-damage-induced apoptosis, in agreement with experimental observations (). We also noted that the final activity was insensitive to the binding rate when k on was away from zero (for example, larger than one-fifth of the default value). Furthermore, Fig. 5 b shows the apoptosis probability obtained by the multicell simulation method, which displayed a weak dependence on the binding rate k on. These results suggest that DNA-damage-induced apoptosis is insensitive to the PDCD5 interactions with Tip6. It would be of interest to further clarify the significance of this interaction in vivo. rhpdcd5 sensitizes cells to DNA damage by promoting caspase-3 activity The simulations described above showed that our model qualitatively agrees well with experimental observations. Next, we applied the model to study how rhpdcd5 can affect DNAdamage-induced cell apoptosis. To that end, we replaced the cytoplasmic PDCD5 production rate v P ([A]) by v rh þ v P ([A]), where v rh is a constant representing the increased rate of cytoplasmic PDCD5 proteins due to the injection of rhpdcd5. Furthermore, we assumed that rhpdcd5 proteins share same functions as endogenous PDCD5. Fig. S8 shows the simulation results of apoptosis probabilities for wild-type and sipdcd5 treated cells, with and without rhpdcd5, as functions of stimulus strength. The results show that administration of rhpdcd5 increased the apoptosis probability for both wild-type and sipdcd5 cells, but this effect was more significant for sipdcd5 cells than for wild-type cells. As noted above, the modulation of cytoplasmic PDCD5 proteins to caspase-3 activity was important for PDCD5-regulated cell apoptosis. Thus, the effect of rhpdcd5 was mainly due to the increase of cytoplasm PDCD5 proteins, which enhanced caspase-3 activity. Finally, in Fig. S8, we note that in the case of a weak stimulus and after rhpdcd5 administration, the apoptosis probabilities showed an even decreased dependence on the stimulus strength v A. This is a result of decreased cytoplasmic PDCD5 protein concentration due to PDCD5 nuclear translocation. DISCUSSION PDCD5 is a novel gene that is involved in the determination of cell fate after DNA damage (,9), and decreased expression of PDCD5 has been detected in various human tumors (2 22,24). In cells undergoing apoptosis, the PDCD5 level in the cytoplasm is rapidly upregulated. At least two pathways may be involved in PDCD5-mediated enhancement of apoptosis. In the nucleus, PDCD5 interacts with Tip6 to promote both Tip6 HAT activity and Tip6- dependent K2 acetylation of p53, the latter of which increases the proapoptotic Bcl-2 family genes and induces apoptosis (). In the cytoplasm, decreased expression of PDCD5 was found to attenuate cell apoptosis and caspase- 3 activity induced by Bax expression, which suggests that PDCD5 may modulate caspase-3 activity through a cytoplasmic pathway (26). In recent studies, rhpdcd5 was found to facilitate tumor cell apoptosis triggered by chemotherapy (3 33). Therefore, overexpression of PDCD5 can enhance cell apoptosis, and administration of rhpdcd5 may have an important role in cancer chemotherapy. Nevertheless, to improve the safety of its use in clinical practice, k on Time (h) b Apoptosis probability k on (ratio to the default value) FIGURE 5 Simulation of cellular response to different binding rates of PDCD5 interactions with Tip6. (a) Dynamics of caspase-3 activity with k on increasing from zero to double the default value. The arrow shows the direction of increasing k on.(b) Apoptosis probability as a function of k on. The probability was obtained by the multicell simulation method (see Materials and Methods for details). A cell was identified as apoptotic if its caspase-3 activity was >.5 at 4 h after stimulation. Stimulus strength v A ¼.26. Biophysical Journal ()
8 PDCD5-Regulated Cell Fate Decision 2589 there is a pressing need to uncover the molecular mechanism by which PDCD5 accelerates cellular apoptosis, and to quantitatively investigate its role in the cell fate decision after chemotherapy. In this study, we attempted to quantitatively study the PDCD5-regulated cell fate decision after UV-irradiationinduced DNA damage. We developed a mathematical model for the regulatory pathways of PDCD5 in response to DNA damage. In this model, the PDCD5 protein level in the cytoplasm is upregulated after DNA damage, and cytoplasmic PDCD5 proteins can enhance caspase-3 activity by modulating Bax translocation. The two PDCD5 signaling pathways included in our model are based on experimental observations. In the cytoplasmic pathway, the PDCD5 proteins promote caspase-3 activity by facilitating the translocation of Bax from the cytosol to the outer membrane of the mitochondria. In the nuclear pathway, the PDCD5 protein binds to Tip6, thereby stabilizing it and promoting its HAT activity and Tip6-dependent p53 acetylation at K2. The two pathways are connected by PDCD5 nuclear translocation. Our model includes two feedback loops for the DNA damage signal that is represented by ATR activity: ), a negative feedback loop through the DNA repair pathway that tends to reduce the damage signal; and 2), a positive feedback through the apoptosis pathway that tends to increase the damage signal. Thus, the cell fate decision depends on the competition between these two pathways. The PDCD5 nuclear pathway is contained in both feedback loops, whereas the cytoplasmic pathway is involved only in the positive feedback loop. We propose several crucial hypotheses based on experimental observations (,26,27): ) PDCD5 nuclear translocation is regulated by a Tip6 downstream signal, and this signal can be enhanced by PDCD5. 2) PDCD5 binds to Tip6 to form a complex, and this complex can stabilize Tip6, promote its HAT activity, and enhance K2 acetylation of p53. 3) Cytoplasmic PDCD5 promotes caspase-3 activity by regulating Bax translocation from the cytoplasm to the outer membrane of the mitochondria. We estimated the model parameters in accordance with experimental results from apoptotic U2OS cells induced by UV irradiation. The model equations were solved numerically on the Mathematica 6. platform. To minimize the effect of specific parameter values, we applied the method of multicell simulation in which parameters are taken randomly within a range 52% of their default values. Using the multicell simulations, we obtained apoptosis probabilities of a group of cells whose parameters are random over the given range. Our simulations showed that cells respond to DNA damage with apoptosis triggered at a certain threshold. The threshold was shown to be sensitive to the PDCD5 expression rate and the efficiency of PDCD5-promoted caspase-3 activation, but insensitive to the PDCD5 nuclear translocation rate and the binding rate of PDCD5 to Tip6. In particular, PDCD5-enhancing caspase-3 activity is required for DNA-damage-induced apoptosis. On the basis of our simulations, we propose the following simplified model of PDCD5-regulated apoptosis after DNA damage (Fig. S9):. After DNA damage occurs, expressions of Tip6 and PDCD5 are upregulated, and the p53 level increases rapidly. PDCD5 proteins are mostly distributed in the cytoplasm. Meanwhile, the Tip6 HAT activity is activated and initiates the DNA repair process. 2. Tip6 induces K2 acetylation of p53, which in turn upregulates the expression of Bax. PDCD5 interactions with Tip6 can enhance the K2 acetylation of p Bax proteins translocate from the cytosol to mitochondria outer membranes and promote the release of cytochrome c, which induces caspase-3 activation. Cytoplasmic PDCD5 proteins can enhance caspase-3 activation by modulating Bax translocation. 4. Caspase-3 inhibits the DNA repair process and activates apoptosis. PDCD5 nuclear translocation is an early signal of apoptosis, associated with an accumulation of PDCD5 proteins in the nucleus. Finally, we applied our model to study how rhpdcd5 enhances cell apoptosis after DNA damage, which is often an effect of cancer chemotherapy. The results suggest that administration of rhpdcd5 sensitized cells to DNA damage by promoting caspase-3 activity through the cytoplasmic pathway. Many previous studies involving computational models of damage responses (6,4 44) concentrated on the cell fate decision mediated by p53, which is considered to be a key regulator in the genetic network of DDR. However, in this study we focused on a less-studied protein, PDCD5, which was not considered in previous models. This protein is involved in the DDR network through several interactions. Thus, the proposed model of PDCD5 interactions can be considered an extension of the known computational models of DDR. This extension provides a more comprehensive understanding of cell-fate decisions after DNA damage, and is important for investigations of the development of rhpdcd5 in cancer therapy. To our knowledge, this study is the first attempt to establish a computational model for PDCD5-regulated determination of the cell fate decision after DNA damage. We chose to focus on the PDCD5-related molecular regulatory networks, and omitted other signaling pathways that are involved in cell fate decisions but do not have a clear relation to PDCD5. This simplification was meant to help us elucidate the specific molecular mechanism of PDCD5 signaling. Nevertheless, a more complete signaling network is certainly necessary to enhance our understanding of cell fate decisions in response to DNA damage. Additional work in this direction is required before our model can be applied within a larger context. Biophysical Journal ()
9 259 Zhuge et al. SUPPORTING MATERIAL Mathematical modeling, parameter estimations, nine figures, a table, and references are available at S6-3495()36-3. This work was supported by the grants from the National Key Project for Basic Research of China (973, 2CB93) and the National Natural Science Foundation of China ( and 973). REFERENCES. Friedberg, E., G. Walker, and W. Sider DNA Repair and Mutagenesis. ASM Press, Washington, DC. 2. Rich, T., R. L. Allen, and A. H. Wyllie. 2. Defying death after DNA damage. Nature. 47: d Adda di Fagagna, F. 28. Living on a break: cellular senescence as a DNA-damage response. Nat. Rev. Cancer. 8: MacCallum, D. E., T. R. Hupp,., P. A. Hall The p53 response to ionising radiation in adult and developing murine tissues. Oncogene. 3: Vousden, K. H., and D. P. Lane. 27. p53 in health and disease. Nat. Rev. Mol. Cell Biol. 8: Zhang, X.-P., F. Liu,., W. Wang. 29. Cell fate decision mediated by p53 pulses. Proc. Natl. Acad. Sci. USA. 6: Hoekstra, M. F Responses to DNA damage and regulation of cell cycle checkpoints by the ATM protein kinase family. Curr. Opin. Genet. Dev. 7: Smith, G. C., R. B. Cary,., S. P. Jackson Purification and DNA binding properties of the ataxia-telangiectasia gene product ATM. Proc. Natl. Acad. Sci. USA. 96: Cheng, Z., Y. Ke,., X. Yao. 28. Functional characterization of TIP6 sumoylation in UV-irradiated DNA damage response. Oncogene. 27: Erratum in Oncogene 28;27:82.. Xu, L., Y. Chen,., D. Ma. 29. PDCD5 interacts with Tip6 and functions as a cooperator in acetyltransferase activity and DNA damage-induced apoptosis. Neoplasia. : Sionov, R. V., and Y. Haupt The cellular response to p53: the decision between life and death. Oncogene. 8: Meek, D. W. 29. Tumour suppression by p53: a role for the DNA damage response? Nat. Rev. Cancer. 9: Meek, D. W Mechanisms of switching on p53: a role for covalent modification? Oncogene. 8: Lev Bar-Or, R., R. Maya,., M. Oren. 2. Generation of oscillations by the p53-mdm2 feedback loop: a theoretical and experimental study. Proc. Natl. Acad. Sci. USA. 97: Hengartner, M. O. 2. The biochemistry of apoptosis. Nature. 47: Robertson, J. D., S. Orrenius, and B. Zhivotovsky. 2. Review: nuclear events in apoptosis. J. Struct. Biol. 29: Murr, R., J. I. Loizou,., Z. Herceg. 26. Histone acetylation by Trrap-Tip6 modulates loading of repair proteins and repair of DNA double-strand breaks. Nat. Cell Biol. 8: Sykes, S. M., H. S. Mellert,., S. B. McMahon. 26. Acetylation of the p53 DNA-binding domain regulates apoptosis induction. Mol. Cell. 24: Liu, H., Y. Wang,., D. Ma TFAR9, a novel apoptosis-related gene cloned from human leukemia cell line TF-, could enhance apoptosis of some tumor cells induced by growth factor withdrawal. Biochem. Biophys. Res. Commun. 254: Li, H., Q. Wang,., L. Zhang. 28. Reduced expression of PDCD5 is associated with high-grade astrocytic gliomas. Oncol. Rep. 2: Chen, C., H. Zhou,., Q. Ma. 2. Prognostic significance of downregulated expression of programmed cell death 5 in chondrosarcoma. J. Surg. Oncol. 2: Yang, Y. H., M. Zhao,., Y. Y. Lu. 26. Expression of programmed cell death 5 gene involves in regulation of apoptosis in gastric tumor cells. Apoptosis. : Spinola, M., P. Meyer,., A. Braun. 26. Association of the PDCD5 locus with lung cancer risk and prognosis in smokers. J. Clin. Oncol. 24: Ruan, G. R., Y. Z. Qin,., Y. R. Liu. 26. Abnormal expression of the programmed cell death 5 gene in acute and chronic myeloid leukemia. Leuk. Res. 3: Storey, J. D., and R. Tibshirani. 23. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA. : Chen, L. N., Y. Wang,., Y. Y. Chen. 26. Short interfering RNA against the PDCD5 attenuates cell apoptosis and caspase-3 activity induced by Bax overexpression. Apoptosis. :. 27. Chen, Y., R. Sun,., D. Ma. 2. Nuclear translocation of PDCD5 (TFAR9): an early signal for apoptosis? FEBS Lett. 59: Ruan, G. R., H. S. Zhao,., X. J. Huang. 28. Adenovirus-mediated PDCD5 gene transfer sensitizes K562 cells to apoptosis induced by idarubicin in vitro and in vivo. Apoptosis. 3: Xie, M., J. H. Niu,., G. R. Ruan. 29. A novel triple-regulated oncolytic adenovirus carrying PDCD5 gene exerts potent antitumor efficacy on common human leukemic cell lines. Apoptosis. 4: Wang, Y., D. Li,., D. Ma. 26. Cellular uptake of exogenous human PDCD5 protein. J. Biol. Chem. 28: Wang, Y., L. Shi,., X. Ke. 29. Recombinant human PDCD5 protein enhances chemo-sensitivities of hematologic maliganacies. Chin. Sci. Bull. 54: Chen, C., H. Zhou,., Y. Chen. 2. Recombinant human PDCD5 sensitizes chondrosaromas to cisplatin chemotherapy in vitro and in vivo. Apoptosis. 5: Shi, L., Q. Song,., Y. Wang. 2. Potent antitumor activities of recombinant human PDCD5 protein in combination with chemotherapy drugs in K562 cells. Biochem. Biophys. Res. Commun. 396: Eissing, T., H. Conzelmann,., P. Scheurich. 24. Bistability analyses of a caspase activation model for receptor-induced apoptosis. J. Biol. Chem. 279: Bagci, E. Z., Y. Vodovotz,., I. Bahar. 26. Bistability in apoptosis: roles of bax, bcl-2, and mitochondrial permeability transition pores. Biophys. J. 9: Bentele, M., I. Lavrik,., R. Eils. 24. Mathematical modeling reveals threshold mechanism in CD95-induced apoptosis. J. Cell Biol. 66: Hua, F., M. G. Cornejo,., D. A. Lauffenburger. 25. Effects of Bcl-2 levels on Fas signaling-induced caspase-3 activation: molecular genetic tests of computational model predictions. J. Immunol. 75: Legewie, S., N. Blüthgen, and H. Herzel. 26. Mathematical modeling identifies inhibitors of apoptsis as mediators of positive feedback and bistability. PLOS Comput. Biol. 2: Gu, C., J. Zhang,., J. Lei. 2. A trigger model of apoptosis induced by tumor necrosis factor signaling. BMC Syst. Biol. 5(Suppl ):S3. 4. Li, Z., M. Ni,., C. Tang. 2. Decision making of the p53 network: death by integration. J. Theor. Biol. 27: Sun, T., C. Chen,., P. Shen. 29. Modeling the role of p53 pulses in DNA damage-induced cell death decision. BMC Bioinformatics. : Zhang, T., P. Brazhnik, and J. J. Tyson. 27. Exploring mechanisms of the DNA-damage response: p53 pulses and their possible relevance to apoptosis. Cell Cycle. 6: Zhang, X.-P., F. Liu, and W. Wang. 2. Two-phase dynamics of p53 in the DNA damage response. Proc. Natl. Acad. Sci. USA. 8: Biophysical Journal ()
10 PDCD5-Regulated Cell Fate Decision Tyson, J. J., W. T. Baumann,., R. Clarke. 2. Dynamic modelling of oestrogen signalling and cell fate in breast cancer cells. Nat. Rev. Cancer. : Su, T. T. 26. Cellular responses to DNA damage: one signal, multiple choices. Annu. Rev. Genet. 4: Ulrich, H. D., and H. Walden. 2. Ubiquitin signalling in DNA replication and repair. Nat. Rev. Mol. Cell Biol. : Wolfram Research, Inc Mathematica, version 6.. Wolfram Research, Inc., Champaign, IL. 48. Geske, F. J., A. C. Nelson,., L. E. Gerschenson. 2. DNA repair is activated in early stages of p53-induced apoptosis. Cell Death Differ. 7: Geske, F. J., R. Lieberman,., L. E. Gerschenson. 2. Early stages of p53-induced apoptosis are reversible. Cell Death Differ. 8:82 9. Biophysical Journal ()
Supplementary Information
 Supplementary Information Significance of p53 dynamics in regulating apoptosis in response to ionizing radiation, and polypharmacological strategies Bing Liu 1,, Divesh Bhatt 1,, Zoltan N. Oltvai 2, Joel
Supplementary Information Significance of p53 dynamics in regulating apoptosis in response to ionizing radiation, and polypharmacological strategies Bing Liu 1,, Divesh Bhatt 1,, Zoltan N. Oltvai 2, Joel
Molecular biology :- Cancer genetics lecture 11
 Molecular biology :- Cancer genetics lecture 11 -We have talked about 2 group of genes that is involved in cellular transformation : proto-oncogenes and tumour suppressor genes, and it isn t enough to
Molecular biology :- Cancer genetics lecture 11 -We have talked about 2 group of genes that is involved in cellular transformation : proto-oncogenes and tumour suppressor genes, and it isn t enough to
609G: Concepts of Cancer Genetics and Treatments (3 credits)
 Master of Chemical and Life Sciences Program College of Computer, Mathematical, and Natural Sciences 609G: Concepts of Cancer Genetics and Treatments (3 credits) Text books: Principles of Cancer Genetics,
Master of Chemical and Life Sciences Program College of Computer, Mathematical, and Natural Sciences 609G: Concepts of Cancer Genetics and Treatments (3 credits) Text books: Principles of Cancer Genetics,
C-Phycocyanin (C-PC) is a n«sjfc&c- waefc-jduble phycobiliprotein. pigment isolated from Spirulina platensis. This water- soluble protein pigment is
 ' ^Summary C-Phycocyanin (C-PC) is a n«sjfc&c- waefc-jduble phycobiliprotein pigment isolated from Spirulina platensis. This water- soluble protein pigment is of greater importance because of its various
' ^Summary C-Phycocyanin (C-PC) is a n«sjfc&c- waefc-jduble phycobiliprotein pigment isolated from Spirulina platensis. This water- soluble protein pigment is of greater importance because of its various
Identification of influential proteins in the classical retinoic acid signaling pathway
 Ghaffari and Petzold Theoretical Biology and Medical Modelling (2018) 15:16 https://doi.org/10.1186/s12976-018-0088-7 RESEARCH Open Access Identification of influential proteins in the classical retinoic
Ghaffari and Petzold Theoretical Biology and Medical Modelling (2018) 15:16 https://doi.org/10.1186/s12976-018-0088-7 RESEARCH Open Access Identification of influential proteins in the classical retinoic
Supporting information (SI) Cell fate decision mediated by p53 pulses Xiao-Peng Zhang, Feng Liu, Zhang Cheng, and Wei Wang
 Supporting information (SI) Cell fate decision mediated by p53 pulses Xiao-Peng Zhang, Feng Liu, Zhang Cheng, and Wei Wang In this work, we aim at the p53 signaling network in untransformed mammalian cells,
Supporting information (SI) Cell fate decision mediated by p53 pulses Xiao-Peng Zhang, Feng Liu, Zhang Cheng, and Wei Wang In this work, we aim at the p53 signaling network in untransformed mammalian cells,
Stochastic Simulation of P53, MDM2 Interaction
 Human Journals Research Article May 2017 Vol.:6, Issue:3 All rights are reserved by B.Odgerel et al. Stochastic Simulation of P53, MDM2 Interaction Keywords: DNA damage, P53, MDM2 interactions, propensity,
Human Journals Research Article May 2017 Vol.:6, Issue:3 All rights are reserved by B.Odgerel et al. Stochastic Simulation of P53, MDM2 Interaction Keywords: DNA damage, P53, MDM2 interactions, propensity,
Silibinin i activates p53-caspase-2 pathway and causes caspase-mediated cleavage of Cip1/p21 in apoptosis
 Silibinin i activates p53-caspase-2 pathway and causes caspase-mediated cleavage of Cip1/p21 in apoptosis induction in bladder transitional-cell papilloma RT4 cells: Evidence for a regulatory loop between
Silibinin i activates p53-caspase-2 pathway and causes caspase-mediated cleavage of Cip1/p21 in apoptosis induction in bladder transitional-cell papilloma RT4 cells: Evidence for a regulatory loop between
Apoptosis Oncogenes. Srbová Martina
 Apoptosis Oncogenes Srbová Martina Cell Cycle Control point Cyclin B Cdk1 Cyclin D Cdk4 Cdk6 Cyclin A Cdk2 Cyclin E Cdk2 Cyclin-dependent kinase (Cdk) have to bind a cyclin to become active Regulation
Apoptosis Oncogenes Srbová Martina Cell Cycle Control point Cyclin B Cdk1 Cyclin D Cdk4 Cdk6 Cyclin A Cdk2 Cyclin E Cdk2 Cyclin-dependent kinase (Cdk) have to bind a cyclin to become active Regulation
SUPPLEMENTARY INFORMATION
 Supplementary Discussion The cell cycle machinery and the DNA damage response network are highly interconnected and co-regulated in assuring faithful duplication and partition of genetic materials into
Supplementary Discussion The cell cycle machinery and the DNA damage response network are highly interconnected and co-regulated in assuring faithful duplication and partition of genetic materials into
BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney
 BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney Page 2: Immune Mechanisms & Molecular Biology of Host Defence (Prof Campbell) Page 45: Infection and Implications for Cell
BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney Page 2: Immune Mechanisms & Molecular Biology of Host Defence (Prof Campbell) Page 45: Infection and Implications for Cell
Introduction to pathology lecture 5/ Cell injury apoptosis. Dr H Awad 2017/18
 Introduction to pathology lecture 5/ Cell injury apoptosis Dr H Awad 2017/18 Apoptosis = programmed cell death = cell suicide= individual cell death Apoptosis cell death induced by a tightly regulated
Introduction to pathology lecture 5/ Cell injury apoptosis Dr H Awad 2017/18 Apoptosis = programmed cell death = cell suicide= individual cell death Apoptosis cell death induced by a tightly regulated
Removal of Shelterin Reveals the Telomere End-Protection Problem
 Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Cancer Biology How a cell responds to DNA Damage
 1 Cancer Biology How a cell responds to DNA Damage Jann Sarkaria Department of Oncology Mayo Clinic 2 EDUCATIONAL GOALS How proteins can transmit signals to each other. The definition of a tumor suppressor
1 Cancer Biology How a cell responds to DNA Damage Jann Sarkaria Department of Oncology Mayo Clinic 2 EDUCATIONAL GOALS How proteins can transmit signals to each other. The definition of a tumor suppressor
p53 and Apoptosis: Master Guardian and Executioner Part 2
 p53 and Apoptosis: Master Guardian and Executioner Part 2 p14arf in human cells is a antagonist of Mdm2. The expression of ARF causes a rapid increase in p53 levels, so what would you suggest?.. The enemy
p53 and Apoptosis: Master Guardian and Executioner Part 2 p14arf in human cells is a antagonist of Mdm2. The expression of ARF causes a rapid increase in p53 levels, so what would you suggest?.. The enemy
Basics of Radiation Biology
 Basics of Radiation Biology Sally A. Amundson Columbia University Center for Radiological Research http://www.cmcr.columbia.edu/ Overview Radiation damage to cells DNA Effects of radiation damage on cells
Basics of Radiation Biology Sally A. Amundson Columbia University Center for Radiological Research http://www.cmcr.columbia.edu/ Overview Radiation damage to cells DNA Effects of radiation damage on cells
Basics of Radiation Biology
 Basics of Radiation Biology Sally A. Amundson Columbia University Center for Radiological Research http://www.cmcr.columbia.edu/ Overview Radiation damage to cells DNA Effects of radiation damage on cells
Basics of Radiation Biology Sally A. Amundson Columbia University Center for Radiological Research http://www.cmcr.columbia.edu/ Overview Radiation damage to cells DNA Effects of radiation damage on cells
Cell cycle and Apoptosis. Chalermchai Mitrpant
 Cell cycle and Apoptosis 2556 Chalermchai Mitrpant Overview of the cell cycle Outline Regulatory mechanisms controlling cell cycle Progression of the cell cycle Checkpoint of the cell cycle Phases of the
Cell cycle and Apoptosis 2556 Chalermchai Mitrpant Overview of the cell cycle Outline Regulatory mechanisms controlling cell cycle Progression of the cell cycle Checkpoint of the cell cycle Phases of the
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Research Communication
 IUBMB Life, 58(11): 659 663, November 2006 Research Communication Sequestration Shapes the Response of Signal Transduction Cascades Nils Blu thgen Molecular Neurobiology, Free University Berlin, and Institute
IUBMB Life, 58(11): 659 663, November 2006 Research Communication Sequestration Shapes the Response of Signal Transduction Cascades Nils Blu thgen Molecular Neurobiology, Free University Berlin, and Institute
Introduction. Cancer Biology. Tumor-suppressor genes. Proto-oncogenes. DNA stability genes. Mechanisms of carcinogenesis.
 Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Lecture 14 - The cell cycle and cell death
 02.17.10 Lecture 14 - The cell cycle and cell death The cell cycle: cells duplicate their contents and divide The cell cycle may be divided into 4 phases The cell cycle triggers essential processes (DNA
02.17.10 Lecture 14 - The cell cycle and cell death The cell cycle: cells duplicate their contents and divide The cell cycle may be divided into 4 phases The cell cycle triggers essential processes (DNA
Chapt 15: Molecular Genetics of Cell Cycle and Cancer
 Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
Regulation of Cell Division (Ch. 12)
 Regulation of Cell Division (Ch. 12) Coordination of cell division A multicellular organism needs to coordinate cell division across different tissues & organs critical for normal growth, development &
Regulation of Cell Division (Ch. 12) Coordination of cell division A multicellular organism needs to coordinate cell division across different tissues & organs critical for normal growth, development &
Cell Cycle, Mitosis, and Microtubules. LS1A Final Exam Review Friday 1/12/07. Processes occurring during cell cycle
 Cell Cycle, Mitosis, and Microtubules LS1A Final Exam Review Friday 1/12/07 Processes occurring during cell cycle Replicate chromosomes Segregate chromosomes Cell divides Cell grows Cell Growth 1 The standard
Cell Cycle, Mitosis, and Microtubules LS1A Final Exam Review Friday 1/12/07 Processes occurring during cell cycle Replicate chromosomes Segregate chromosomes Cell divides Cell grows Cell Growth 1 The standard
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Mitosis and the Cell Cycle
 Mitosis and the Cell Cycle Chapter 12 The Cell Cycle: Cell Growth & Cell Division Where it all began You started as a cell smaller than a period at the end of a sentence Getting from there to here Cell
Mitosis and the Cell Cycle Chapter 12 The Cell Cycle: Cell Growth & Cell Division Where it all began You started as a cell smaller than a period at the end of a sentence Getting from there to here Cell
The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein
 THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
THESIS BOOK The functional investigation of the interaction between TATA-associated factor 3 (TAF3) and p53 protein Orsolya Buzás-Bereczki Supervisors: Dr. Éva Bálint Dr. Imre Miklós Boros University of
Cell Quality Control. Peter Takizawa Department of Cell Biology
 Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Oncolytic virus strategy
 Oncolytic viruses Oncolytic virus strategy normal tumor NO replication replication survival lysis Oncolytic virus strategy Mechanisms of tumor selectivity of several, some of them naturally, oncolytic
Oncolytic viruses Oncolytic virus strategy normal tumor NO replication replication survival lysis Oncolytic virus strategy Mechanisms of tumor selectivity of several, some of them naturally, oncolytic
Crosstalk between Adiponectin and IGF-IR in breast cancer. Prof. Young Jin Suh Department of Surgery The Catholic University of Korea
 Crosstalk between Adiponectin and IGF-IR in breast cancer Prof. Young Jin Suh Department of Surgery The Catholic University of Korea Obesity Chronic, multifactorial disorder Hypertrophy and hyperplasia
Crosstalk between Adiponectin and IGF-IR in breast cancer Prof. Young Jin Suh Department of Surgery The Catholic University of Korea Obesity Chronic, multifactorial disorder Hypertrophy and hyperplasia
Tumor cell reassortment within the cell cycle (including checkpoints and cell-cycle arrest)
 Tumor cell reassortment within the cell cycle (including checkpoints and cell-cycle arrest) Carsten Herskind Dept. of Radiation Oncology. Universitätsmedizin Mannheim Medical Faculty Mannheim, Heidelberg
Tumor cell reassortment within the cell cycle (including checkpoints and cell-cycle arrest) Carsten Herskind Dept. of Radiation Oncology. Universitätsmedizin Mannheim Medical Faculty Mannheim, Heidelberg
Dynamics of p53 and NF-κB regulation in response to DNA damage and identification of target proteins suitable for therapeutic intervention
 Poltz and Naumann BMC Systems Biology 2012, 6:125 RESEARCH ARTICLE Open Access Dynamics of p53 and NF-κB regulation in response to DNA damage and identification of target proteins suitable for therapeutic
Poltz and Naumann BMC Systems Biology 2012, 6:125 RESEARCH ARTICLE Open Access Dynamics of p53 and NF-κB regulation in response to DNA damage and identification of target proteins suitable for therapeutic
Multistep nature of cancer development. Cancer genes
 Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Biochemistry 673 Your Name: Regulation of Metabolism Assoc. Prof. Jason Kahn Final Exam (150 points total) May 17, 2005
 Biochemistry 673 Your Name: Regulation of Metabolism Assoc. Prof. Jason Kahn Final Exam (150 points total) May 17, 2005 You have 120 minutes for this exam. Explanations should be concise and clear. Generous
Biochemistry 673 Your Name: Regulation of Metabolism Assoc. Prof. Jason Kahn Final Exam (150 points total) May 17, 2005 You have 120 minutes for this exam. Explanations should be concise and clear. Generous
BioNSi: A Discrete Biological Network Simulator Tool
 BioNSi: A Discrete Biological Network Simulator Tool Amir Rubinstein 1, Noga Bracha 1, Liat Rudner 1, Noga Zucker 1, Hadas E. Sloin 2, and Benny Chor 1 1 Blavatnik School of Computer Science, Tel Aviv
BioNSi: A Discrete Biological Network Simulator Tool Amir Rubinstein 1, Noga Bracha 1, Liat Rudner 1, Noga Zucker 1, Hadas E. Sloin 2, and Benny Chor 1 1 Blavatnik School of Computer Science, Tel Aviv
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition
 Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Tumour growth environment modulates Chk1 signalling pathways and sensitivity to Chk1 inhibition Andrew J Massey Supplementary Information Supplementary Figure S1. Related to Fig. 1. (a) HT29 or U2OS cells
Apoptosis Chapter 9. Neelu Yadav PhD
 Apoptosis Chapter 9 Neelu Yadav PhD Neelu.Yadav@Roswellpark.org 1 Apoptosis: Lecture outline Apoptosis a programmed cell death pathway in normal homeostasis Core Apoptosis cascade is conserved Compare
Apoptosis Chapter 9 Neelu Yadav PhD Neelu.Yadav@Roswellpark.org 1 Apoptosis: Lecture outline Apoptosis a programmed cell death pathway in normal homeostasis Core Apoptosis cascade is conserved Compare
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Lecture 10. G1/S Regulation and Cell Cycle Checkpoints. G1/S regulation and growth control G2 repair checkpoint Spindle assembly or mitotic checkpoint
 Lecture 10 G1/S Regulation and Cell Cycle Checkpoints Outline: G1/S regulation and growth control G2 repair checkpoint Spindle assembly or mitotic checkpoint Paper: The roles of Fzy/Cdc20 and Fzr/Cdh1
Lecture 10 G1/S Regulation and Cell Cycle Checkpoints Outline: G1/S regulation and growth control G2 repair checkpoint Spindle assembly or mitotic checkpoint Paper: The roles of Fzy/Cdc20 and Fzr/Cdh1
Chapter 12. Regulation of Cell Division. AP Biology
 Chapter 12. Regulation of Cell Division Coordination of cell division! Multicellular organism " need to coordinate across different parts of organism! timing of cell division! rates of cell division "
Chapter 12. Regulation of Cell Division Coordination of cell division! Multicellular organism " need to coordinate across different parts of organism! timing of cell division! rates of cell division "
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
The Need for a PARP in vivo Pharmacodynamic Assay
 The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D. Chief Scientific Officer Trevigen, Inc. Gaithersburg, MD Poly(ADP-ribose) polymerases are promising therapeutic targets. In response
The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D. Chief Scientific Officer Trevigen, Inc. Gaithersburg, MD Poly(ADP-ribose) polymerases are promising therapeutic targets. In response
Modeling of Cancer Signaling Pathways
 Modeling of Cancer Signaling Pathways by Remziye Karabekmez A thesis presented to the University of Waterloo in fulfillment of the thesis requirement for the degree of Master of Mathematics in Applied
Modeling of Cancer Signaling Pathways by Remziye Karabekmez A thesis presented to the University of Waterloo in fulfillment of the thesis requirement for the degree of Master of Mathematics in Applied
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Programmed Cell Death (apoptosis)
 Programmed Cell Death (apoptosis) Stereotypic death process includes: membrane blebbing nuclear fragmentation chromatin condensation and DNA framentation loss of mitochondrial integrity and release of
Programmed Cell Death (apoptosis) Stereotypic death process includes: membrane blebbing nuclear fragmentation chromatin condensation and DNA framentation loss of mitochondrial integrity and release of
Tala Saleh. Ahmad Attari. Mamoun Ahram
 23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
Part-4. Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death
 Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
Cell cycle and apoptosis
 Cell cycle and apoptosis Cell cycle Definition Stages and steps Cell cycle Interphase (G1/G0, S, and G2) Mitosis (prophase, metaphase, anaphase, telophase, karyokinesis, cytokinesis) Control checkpoints
Cell cycle and apoptosis Cell cycle Definition Stages and steps Cell cycle Interphase (G1/G0, S, and G2) Mitosis (prophase, metaphase, anaphase, telophase, karyokinesis, cytokinesis) Control checkpoints
shehab Moh Tarek ... ManarHajeer
 3 shehab Moh Tarek... ManarHajeer In the previous lecture we discussed the accumulation of oxygen- derived free radicals as a mechanism of cell injury, we covered their production and their pathologic
3 shehab Moh Tarek... ManarHajeer In the previous lecture we discussed the accumulation of oxygen- derived free radicals as a mechanism of cell injury, we covered their production and their pathologic
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
Transformation of Normal HMECs (Human Mammary Epithelial Cells) into Metastatic Breast Cancer Cells: Introduction - The Broad Picture:
 Transformation of Normal HMECs (Human Mammary Epithelial Cells) into Metastatic Breast Cancer Cells: Introduction - The Broad Picture: Spandana Baruah December, 2016 Cancer is defined as: «A disease caused
Transformation of Normal HMECs (Human Mammary Epithelial Cells) into Metastatic Breast Cancer Cells: Introduction - The Broad Picture: Spandana Baruah December, 2016 Cancer is defined as: «A disease caused
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy se
 A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
Regulation of Cell Division. AP Biology
 Regulation of Cell Division 2006-2007 Coordination of cell division A multicellular organism needs to coordinate cell division across different tissues & organs critical for normal growth, development
Regulation of Cell Division 2006-2007 Coordination of cell division A multicellular organism needs to coordinate cell division across different tissues & organs critical for normal growth, development
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Regulators of Cell Cycle Progression
 Regulators of Cell Cycle Progression Studies of Cdk s and cyclins in genetically modified mice reveal a high level of plasticity, allowing different cyclins and Cdk s to compensate for the loss of one
Regulators of Cell Cycle Progression Studies of Cdk s and cyclins in genetically modified mice reveal a high level of plasticity, allowing different cyclins and Cdk s to compensate for the loss of one
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
The Biochemistry of apoptosis
 The Biochemistry of apoptosis 1 1 The apoptosis is composed of multiple biochemical events 2 2 Biochemical, cellular, and molecular events in Apoptosis 1. Membrane blebbing; phosphatidyl serine exposure
The Biochemistry of apoptosis 1 1 The apoptosis is composed of multiple biochemical events 2 2 Biochemical, cellular, and molecular events in Apoptosis 1. Membrane blebbing; phosphatidyl serine exposure
What is the Warburg Effect
 What is the Warburg Effect Roles nutrients play in the biochemistry of a cell Thus, proliferating cells must acquire more nutrients, convert them into biosynthetic building blocks, and coordinate the reactions
What is the Warburg Effect Roles nutrients play in the biochemistry of a cell Thus, proliferating cells must acquire more nutrients, convert them into biosynthetic building blocks, and coordinate the reactions
Division Ave. High School AP Biology
 Regulation of Cell Division 2008-2009 Coordination of cell division A multicellular organism needs to coordinate cell division across different tissues & organs u critical for normal growth, development
Regulation of Cell Division 2008-2009 Coordination of cell division A multicellular organism needs to coordinate cell division across different tissues & organs u critical for normal growth, development
Brian T Burgess, DO, PhD, GYN Oncology Fellow Rachel W. Miller, MD, GYN Oncology
 Brian T Burgess, DO, PhD, GYN Oncology Fellow Rachel W. Miller, MD, GYN Oncology Epithelial Ovarian Cancer - Standard Current Treatment: Surgery with De-bulking + Platinum-Taxane based Chemotherapy - No
Brian T Burgess, DO, PhD, GYN Oncology Fellow Rachel W. Miller, MD, GYN Oncology Epithelial Ovarian Cancer - Standard Current Treatment: Surgery with De-bulking + Platinum-Taxane based Chemotherapy - No
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/305/ra106/dc1 Supplementary Materials for Controlling Long-Term Signaling: Receptor Dynamics Determine Attenuation and Refractory Behavior of the TGF-β Pathway
www.sciencesignaling.org/cgi/content/full/6/305/ra106/dc1 Supplementary Materials for Controlling Long-Term Signaling: Receptor Dynamics Determine Attenuation and Refractory Behavior of the TGF-β Pathway
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: W81XWH-05-1-0494 TITLE: Tumor Suppression by BRCA-1: A Critical Role at DNA Replication Forks PRINCIPAL INVESTIGATOR: Jean Gautier, Ph.D. CONTRACTING ORGANIZATION: Columbia University
AD Award Number: W81XWH-05-1-0494 TITLE: Tumor Suppression by BRCA-1: A Critical Role at DNA Replication Forks PRINCIPAL INVESTIGATOR: Jean Gautier, Ph.D. CONTRACTING ORGANIZATION: Columbia University
Regulation of Cell Division
 Regulation of Cell Division Two HeLa cancer cells are just completing cytokinesis. Explain how the cell division of cancer cells like these is misregulated. Identify genetic and other changes that might
Regulation of Cell Division Two HeLa cancer cells are just completing cytokinesis. Explain how the cell division of cancer cells like these is misregulated. Identify genetic and other changes that might
Signaling. Dr. Sujata Persad Katz Group Centre for Pharmacy & Health research
 Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Molecular Cell Biology (Bio 5068) Cell Cycle I. Ron Bose, MD PhD November 14, 2017
 Molecular Cell Biology (Bio 5068) Cell Cycle I Ron Bose, MD PhD November 14, 2017 CELL DIVISION CYCLE M G2 S G1 DISCOVERY AND NAMING OF CYCLINS A protein (called cyclin ) was observed to increase as cells
Molecular Cell Biology (Bio 5068) Cell Cycle I Ron Bose, MD PhD November 14, 2017 CELL DIVISION CYCLE M G2 S G1 DISCOVERY AND NAMING OF CYCLINS A protein (called cyclin ) was observed to increase as cells
Biology is the only subject in which multiplication is the same thing as division
 The Cell Cycle Biology is the only subject in which multiplication is the same thing as division Why do cells divide? For reproduction asexual reproduction For growth one-celled organisms from fertilized
The Cell Cycle Biology is the only subject in which multiplication is the same thing as division Why do cells divide? For reproduction asexual reproduction For growth one-celled organisms from fertilized
9 Metabolic trigger: control of methionine metabolism
 9 Metabolic trigger: control of methionine metabolism M.V. Martinov 1,V.M.Vitvitsky 1,E.V.Mosharov 2,R.Banerjee 2,F.I.Ataullakhanov 1 1 National Research Center for Hematology, Moscow, Russia 125167 2
9 Metabolic trigger: control of methionine metabolism M.V. Martinov 1,V.M.Vitvitsky 1,E.V.Mosharov 2,R.Banerjee 2,F.I.Ataullakhanov 1 1 National Research Center for Hematology, Moscow, Russia 125167 2
Review. Ageing 2: Cancer! Review: Mutations. Mutations 2/14/11. The Raw Material for Evolution. The Double Edged Sword
 Ageing 2: Cancer! Review: The force of natural selection declines with ageing due to increase in extrinsic mortality (= weakening of natural selection) and reduction in reproduction with age (selection
Ageing 2: Cancer! Review: The force of natural selection declines with ageing due to increase in extrinsic mortality (= weakening of natural selection) and reduction in reproduction with age (selection
Introduction to Cancer Biology
 Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Supplementary data for:
 Supplementary data for: Prediction of treatment efficacy for prostate cancer using a mathematical model Huiming Peng 1, Weiling Zhao 1, Hua Tan 1, Zhiwei Ji 1, Jingsong Li 2, King Li 1, Xiaobo Zhou 1,*
Supplementary data for: Prediction of treatment efficacy for prostate cancer using a mathematical model Huiming Peng 1, Weiling Zhao 1, Hua Tan 1, Zhiwei Ji 1, Jingsong Li 2, King Li 1, Xiaobo Zhou 1,*
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Title cooperative gene expression.
 Title Rigor of cell fate decision cooperative gene expression by vari by p53 Author(s) Murakami, Yohei; Takada, Shoji Citation BIOPHYSICS (2012), 8: 41-50 Issue Date 2012 URL http://hdl.handle.net/2433/154582
Title Rigor of cell fate decision cooperative gene expression by vari by p53 Author(s) Murakami, Yohei; Takada, Shoji Citation BIOPHYSICS (2012), 8: 41-50 Issue Date 2012 URL http://hdl.handle.net/2433/154582
microrna as a Potential Vector for the Propagation of Robustness in Protein Expression and Oscillatory Dynamics within a cerna Network
 microrna as a Potential Vector for the Propagation of Robustness in Protein Expression and Oscillatory Dynamics within a cerna Network Claude Gérard*,Béla Novák Oxford Centre for Integrative Systems Biology,
microrna as a Potential Vector for the Propagation of Robustness in Protein Expression and Oscillatory Dynamics within a cerna Network Claude Gérard*,Béla Novák Oxford Centre for Integrative Systems Biology,
Early cell death (FGF) B No RunX transcription factor produced Yes No differentiation
 Solution Key - Practice Questions Question 1 a) A recent publication has shown that the fat stem cells (FSC) can act as bone stem cells to repair cavities in the skull, when transplanted into immuno-compromised
Solution Key - Practice Questions Question 1 a) A recent publication has shown that the fat stem cells (FSC) can act as bone stem cells to repair cavities in the skull, when transplanted into immuno-compromised
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001
 BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
BIOL 4374/BCHS 4313 Cell Biology Exam #1 February 13, 2001 SS# Name This exam is worth a total of 100 points. The number of points each question is worth is shown in parentheses. Good luck! 1. (2) The
Pancreatic Cancer Research and HMGB1 Signaling Pathway
 Pancreatic Cancer Research and HMGB1 Signaling Pathway Haijun Gong*, Paolo Zuliani*, Anvesh Komuravelli*, James R. Faeder #, Edmund M. Clarke* * # The Hallmarks of Cancer D. Hanahan and R. A. Weinberg
Pancreatic Cancer Research and HMGB1 Signaling Pathway Haijun Gong*, Paolo Zuliani*, Anvesh Komuravelli*, James R. Faeder #, Edmund M. Clarke* * # The Hallmarks of Cancer D. Hanahan and R. A. Weinberg
KEY CONCEPT QUESTIONS IN SIGNAL TRANSDUCTION
 Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signaling Apoptosis. Scott André Oakes, M.D. Dept. of Pathology Univ. of Calif-San Francisco. Cyt c Release BAX/BAK. Apoptosome Formation
 Signaling Apoptosis Cyt c Release BAX/BAK Apoptosome Formation Caspase Activation Scott André Oakes, M.D. Dept. of Pathology Univ. of Calif-San Francisco Why do we need cell death? Sculpt Organs Control
Signaling Apoptosis Cyt c Release BAX/BAK Apoptosome Formation Caspase Activation Scott André Oakes, M.D. Dept. of Pathology Univ. of Calif-San Francisco Why do we need cell death? Sculpt Organs Control
Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Balancing speed and accuracy of polyclonal T cell activation: A role for extracellular feedback
 Balancing speed and accuracy of polyclonal T cell activation: A role for extracellular feedback The Harvard community has made this article openly available. Please share how this access benefits you.
Balancing speed and accuracy of polyclonal T cell activation: A role for extracellular feedback The Harvard community has made this article openly available. Please share how this access benefits you.
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 Name: Key 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function
BIO360 Fall 2013 Quiz 1 Name: Key 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function
Removal of Shelterin Reveals the Telomere End-Protection Problem
 Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Chapter 9 Signal Transduction and Cell Growth
 Part II Principles of Individual Cell Function Chapter 9 One characteristic of organisms is that they exhibit various behaviors in response to changes in their environment (i.e., the outside world). Cells
Part II Principles of Individual Cell Function Chapter 9 One characteristic of organisms is that they exhibit various behaviors in response to changes in their environment (i.e., the outside world). Cells
Cancer. Throughout the life of an individual, but particularly during development, every cell constantly faces decisions.
 Cancer Throughout the life of an individual, but particularly during development, every cell constantly faces decisions. Should it divide? Yes No--> Should it differentiate? Yes No-->Should it die? Yes-->Apoptosis
Cancer Throughout the life of an individual, but particularly during development, every cell constantly faces decisions. Should it divide? Yes No--> Should it differentiate? Yes No-->Should it die? Yes-->Apoptosis
Protocol Abstract and Schema
 Protocol Abstract and Schema A Phase I Trial of p28 (NSC745104), a Non-HDM2 mediated peptide inhibitor of p53 ubiquitination in pediatric patients with recurrent or progressive CNS tumors Description and
Protocol Abstract and Schema A Phase I Trial of p28 (NSC745104), a Non-HDM2 mediated peptide inhibitor of p53 ubiquitination in pediatric patients with recurrent or progressive CNS tumors Description and
Regulation of the Cell Cycle
 Regulation of the Cell Cycle 21 I. OVERVIEW Quiescent differentiated cell / can be induced to re-enter the active cell cycle. urvival Cell division Apoptosis 1 Daughter cells Apoptic cell enescent cell
Regulation of the Cell Cycle 21 I. OVERVIEW Quiescent differentiated cell / can be induced to re-enter the active cell cycle. urvival Cell division Apoptosis 1 Daughter cells Apoptic cell enescent cell
DSB. Double-Strand Breaks causate da radiazioni stress ossidativo farmaci
 DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to the detection and repair of DNA damage DSBs induce a local decrease
DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to the detection and repair of DNA damage DSBs induce a local decrease
Cancer and Gene Alterations - 1
 Cancer and Gene Alterations - 1 Cancer and Gene Alteration As we know, cancer is a disease of unregulated cell growth. Although we looked at some of the features of cancer when we discussed mitosis checkpoints,
Cancer and Gene Alterations - 1 Cancer and Gene Alteration As we know, cancer is a disease of unregulated cell growth. Although we looked at some of the features of cancer when we discussed mitosis checkpoints,
Kinase Inhibitor p21 WAF1/CIP1 in Apoptosis and Autophagy
 Pivotal Role of the Cyclin-dependent Kinase Inhibitor p21 WAF1/CIP1 in Apoptosis and Autophagy Keishi Fujiwara, Shigeru Daido, Akitsugu Yamamoto, Ryuji Kobayash, Tomohisa Yokoyama, Hiroshi Aok, Eiji Iwado,
Pivotal Role of the Cyclin-dependent Kinase Inhibitor p21 WAF1/CIP1 in Apoptosis and Autophagy Keishi Fujiwara, Shigeru Daido, Akitsugu Yamamoto, Ryuji Kobayash, Tomohisa Yokoyama, Hiroshi Aok, Eiji Iwado,
Lecture 15. Signal Transduction Pathways - Introduction
 Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis
 Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis MUDr. Jiří Vachtenheim, CSc. CELL CYCLE - SUMMARY Basic terminology: Cyclins conserved proteins with homologous regions; their cellular
Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis MUDr. Jiří Vachtenheim, CSc. CELL CYCLE - SUMMARY Basic terminology: Cyclins conserved proteins with homologous regions; their cellular
Early Embryonic Development
 Early Embryonic Development Maternal effect gene products set the stage by controlling the expression of the first embryonic genes. 1. Transcription factors 2. Receptors 3. Regulatory proteins Maternal
Early Embryonic Development Maternal effect gene products set the stage by controlling the expression of the first embryonic genes. 1. Transcription factors 2. Receptors 3. Regulatory proteins Maternal
Chemistry 107 Exam 4 Study Guide
 Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
Phospho-AKT Sampler Kit
 Phospho-AKT Sampler Kit E 0 5 1 0 0 3 Kits Includes Cat. Quantity Application Reactivity Source Akt (Ab-473) Antibody E021054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit Akt (Phospho-Ser473) Antibody
Phospho-AKT Sampler Kit E 0 5 1 0 0 3 Kits Includes Cat. Quantity Application Reactivity Source Akt (Ab-473) Antibody E021054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit Akt (Phospho-Ser473) Antibody
Part I. Boolean modelling exercises
 Part I. Boolean modelling exercises. Glucose repression of Icl in yeast In yeast Saccharomyces cerevisiae, expression of enzyme Icl (isocitrate lyase-, involved in the gluconeogenesis pathway) is important
Part I. Boolean modelling exercises. Glucose repression of Icl in yeast In yeast Saccharomyces cerevisiae, expression of enzyme Icl (isocitrate lyase-, involved in the gluconeogenesis pathway) is important
AperTO - Archivio Istituzionale Open Access dell'università di Torino
 AperTO - Archivio Istituzionale Open Access dell'università di Torino From the nucleus to the mitochondria and backthe odyssey of a multitask STAT3 This is the author's manuscript Original Citation: From
AperTO - Archivio Istituzionale Open Access dell'università di Torino From the nucleus to the mitochondria and backthe odyssey of a multitask STAT3 This is the author's manuscript Original Citation: From
Protein kinases are enzymes that add a phosphate group to proteins according to the. ATP + protein OH > Protein OPO 3 + ADP
 Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
General Principles of Endocrine Physiology
 General Principles of Endocrine Physiology By Dr. Isabel S.S. Hwang Department of Physiology Faculty of Medicine University of Hong Kong The major human endocrine glands Endocrine glands and hormones
General Principles of Endocrine Physiology By Dr. Isabel S.S. Hwang Department of Physiology Faculty of Medicine University of Hong Kong The major human endocrine glands Endocrine glands and hormones
