Cancer results from the sequential accumulation of mutations
|
|
|
- Sharlene Francis
- 6 years ago
- Views:
Transcription
1 Prevalence of somatic alterations in the colorectal cancer cell genome Tian-Li Wang*, Carlo Rago*, Natalie Silliman*, Janine Ptak*, Sanford Markowitz, James K. V. Willson, Giovanni Parmigiani*, Kenneth W. Kinzler*, Bert Vogelstein*, and Victor E. Velculescu* *Howard Hughes Medical Institute and The Sidney Kimmel Comprehensive Cancer Center, Johns Hopkins Medical Institutions, Baltimore, MD 21231; and Howard Hughes Medical Institute, Cancer Center and Department of Medicine, Case Western Reserve University, Ireland Cancer Center, Department of Medicine, and Research Institute, University Hospitals of Cleveland, Cleveland, OH Contributed by Bert Vogelstein, December 31, 2001 Although a small fraction of human cancers have increased rates of somatic mutation because of known deficiencies in DNA repair, little is known about the prevalence of somatic alterations in the vast majority of human cancers. To systematically assess nonsynonymous somatic alterations in colorectal neoplasia, we used DNA sequencing to analyze 3.2 Mb of coding tumor DNA comprising 1,811 exons from 470 genes. In total, we identified only three distinct somatic mutations, comprising two missense changes and one 14-bp deletion, each in a different gene. The accumulation of approximately one nonsynonymous somatic change per Mb of tumor DNA is consistent with a rate of mutation in tumor cells that is similar to that of normal cells. These data suggest that most sporadic colorectal cancers do not display a mutator phenotype at the nucleotide level. They also have significant implications for the interpretation of somatic mutations in candidate tumor-suppressor genes. Cancer results from the sequential accumulation of mutations in specific genes. As new mutations provide a growth advantage to a cell within the tumor, the selected clone outgrows the precursor lesion, and successive waves of mutation, clonal expansion, and growth lead to cancer progression. Throughout this process of mutation and selection, the acquisition of new mutations is thought to be the rate-limiting step, setting the timeline of tumor development. Although an increase in the rate of mutation should accelerate this process, there has been debate as to whether genetic instability or a mutator phenotype (1) in addition to selection is actually required for neoplasia. Some have argued that an increased mutation rate at the nucleotide level is necessary to account for all of the mutations present in cancers (1), while others have suggested that the normal somatic mutation rate, coupled with selection, is sufficient to explain tumor formation in most cases (2). Also, it has been suggested that a different form of genetic instability, involving gross changes in chromosomes rather than subtle changes in nucleotide sequence, drives most human cancers (1, 3, 4). Recent work has demonstrated that a small fraction of cancers clearly display genetic instability at the nucleotide level because of defects in DNA repair processes. The first demonstration of this principle was made through the discovery that patients with xeroderma pigmentosum (XP) and related disorders have hereditary defects in genes encoding nucleotide excision repair enzymes (5). These patients develop skin tumors as a result of their inability to repair mutations caused by UV irradiation. XP and related disorders are inherited in an autosomal recessive fashion and are accordingly rare. Another form of repair defect, involving mismatch repair, has been shown to be implicated in the tumors of all patients with hereditary nonpolyposis colon cancer (HNPCC) syndrome (6), as well as 12% of patients with nonfamilial forms of colorectal cancer (7). HNPCC is inherited in an autosomal dominant fashion and accounts for 3 5% of the total colorectal cancer cases in the Western world (6, 8). The mismatch repair defects are predominantly due to inactivation of the hmsh2 or hmlh1 genes, although defects in other mismatch repair genes play a role in a subset of HNPCC cases. Although mismatch repair defects have been shown to lead to a widespread nucleotide instability, microsatellite sequences are particularly susceptible to this form of repair defect (9, 10). Accordingly, microsatellite instability (MIN), defined by insertion and deletion errors in repetitive elements, has been shown to occur in all colorectal tumors harboring mismatch repair defects. MIN also has been found in a small fraction of other cancers, including those of the stomach, endometrium, and ovary. Unlike the aforementioned examples, the vast majority of human cancers seem to possess intact mismatch repair and base excision repair capabilities, and it is unclear whether they possess a mutator phenotype at the nucleotide level. Some studies have suggested an increase in the number of point mutations in tumors, for example, by analysis of published data on multiple mutations in the p53 gene (11), but such measurements have been limited to individual genes and may be compromised by a number of experimental variables. Other studies have suggested an elevated rate of somatic mutation in colorectal tumors on the basis of the analysis of sequences adjacent to dinucleotide repeats (12). To obtain an unbiased measurement of the prevalence of nucleotide alterations, we surveyed a large number of coding regions in the genomes of colorectal cancers. Our results, described below, have significant implications for theories relating instability to naturally occurring human tumorigenesis and the interpretation of somatic mutations in specific genes in such cancers. Material and Methods Collection of Gene Sequences. Gene sequences were obtained from the Celera Discovery System (http: cds.celera.com) and Gen- Bank (http: databases. Determination of coding sequence within each transcript was based on annotation present in the databases. In each case, we used computational approaches to confirm start and stop codons and ORFs. Primer Design and Oligonucleotide Synthesis. Exon sequences and adjacent intronic sequences for each gene were extracted from Celera Discovery System or from GenBank databases. Primers for PCR amplification and DNA sequencing were designed by using the PRIMER 3 program (13) (http: www-genome.wi.mit. edu cgi-bin primer primer3 Forward primers were localized to regions no closer than 100 bp from the exon boundaries; forward sequencing primers were designed so that they would not overlap forward PCR primers and to extend no Abbreviations: XP, xeroderma pigmentosum; MIN, microsatellite instability; SNP, single nucleotide polymorphism. To whom reprint requests should be addressed at: The Johns Hopkins Oncology Center, 1650 Orleans Street, Room 589, Baltimore, MD velculescu@jhmi.edu. The publication costs of this article were defrayed in part by page charge payment. This article must therefore be hereby marked advertisement in accordance with 18 U.S.C solely to indicate this fact PNAS March 5, 2002 vol. 99 no. 5 cgi doi pnas
2 closer than 50 bp to the 5 exon boundary. Reverse amplification primers were localized to regions no closer than 10 bp to the 3 exon boundary. Oligonucleotides for PCR amplification and sequencing were synthesized by Genelink, NY. Isolation of Tumor DNA. Because primary tumors always contain nonneoplastic cells, presumptive mutations can be masked or difficult to interpret. To circumvent this difficulty, we analyzed early passage cell lines passaged in vitro or in nude mice, in which sequence changes were unambiguous. It has been shown previously that the genetic alterations in such cell lines usually are indistinguishable from those found in the primary tumors (14 16). Surgically removed colorectal tumors were disaggregated and implanted into nude mice or into in vitro culture conditions, as described (14). DNA was purified by standard SDS-proteinase K digestion and phenol-chloroform extraction or by DNAEasy Tissue kit (Qiagen, Chatsworth, CA). Determination of MIN Status in Tumor Samples. A large number of dinucleotide or tetranucleotide microsatellite sequences ( 20) and the BAT26 mononucleotide marker (17) were evaluated in each of the tumors used in this cohort. PCR products were separated and analyzed on either manual sequencing gels or by using an SCE well capillary electrophoresis system (SpectruMedix, State College, PA). PCR Amplification. Each PCR was performed in 10 l containing 0.25 mm dntps, 1 M forward and reverse primers, 6% DMSO, 1 PCR buffer (Invitrogen), 5 ng DNA, and 0.5 units platinum Taq (Invitrogen). PCR was carried out in 384-well format (Applied Biosystems) with a touchdown PCR program: 94 C(2 min); 3 cycles of 94 C (30 sec), 64 C (30 sec), 70 C (30 sec); 3 cycles of 94 C (30 sec), 61 C (30 sec), 70 C (30 sec); 3 cycles of 94 C (30 sec), 58 C (30 sec), 70 C (30 sec); 35 cycles of 94 C for (30 sec), 57 C (30 sec), 70 C (30 sec); 1 cycle of 70 C (5 min). After PCR, samples were purified by isopropanol precipitation and resuspended in TE (10 mm Tris, ph mm EDTA). DNA Sequencing. Purified PCR products were sequenced by using Big Dye Terminator v.3.0 Kit (Applied Biosystems). Cycle sequencing reactions were carried out in 384-well format at 35 cycles of 96 C for 10 s, 50 C for 5 s, and 60 C for 4 min. Sequencing reactions were precipitated by mixing with addition of a 20- l mixture containing 87.5% ethanol and M sodium acetate. The precipitated products were resuspended in sample loading buffer (Hi-Di Formamide, Applied Biosystems) and were analyzed with a SCE well capillary electrophoresis system (SpectruMedix). Sequencing Data Analysis. Sequencing data were analyzed by using BASECALL v.3.39 (SpectruMedix). SEQMAN (DNAstar, Madison, WI) and SEQUENCHER (Gene Codes, Ann Arbor, MI) were used to assemble sequence chromatograms and consensus reference sequences. Assemblies were visually inspected to identify potential alterations. Coding sequences containing nonsynonymous alterations were searched against Celera Discovery System RefSNP database to determine if the changes corresponded to previously identified polymorphisms. Statistical Analysis. The interval estimate for somatic mutations per Mb tumor DNA represents a highest posterior density region based on an exact a posteriori analysis, assuming a Poisson model for the distribution of mutations on the genome, and a uniform a priori distribution on the unknown mutation rate (18). It is preferable to standard asymptotic confidence intervals because there are few mutations and the likelihood function is skewed. Additional details on these analyses and statistical tools for evaluating whether observed mutation frequencies are Table 1. Summary of sequence analysis and observed alterations Selected genes 504 Exons extracted 2,206 Genes analyzed 470 Exons analyzed 1,811 Average exon size 152 bp Average number of tumors analyzed per exon 11.4 Total sequence data obtained 6.5 Mb Total coding DNA analyzed 3.2 Mb Nonsynonymous changes observed 320 Previously identified SNPs 90 Novel coding SNPs 227 Somatic mutations 3 above the background mutation frequency are available at http: astor.som.jhmi.edu gp trab. Results and Discussion To explore the frequency of somatic alterations in coding regions of tumor DNA, we selected a total of 504 genes for analysis. By using a combination of the public (19) and private (Celera; ref. 20) genome databases, we extracted available genomic data for a total of 2,206 exons contained in these genes and designed primers for PCR amplification and sequencing. The sequences of these genes were determined in a panel of 12 colorectal cancer cell lines. Cell lines rather than primary tumors were used to ensure that nonneoplastic cells within the tumors did not complicate the sequence analysis. These cell lines were generally of early passage, and each was extensively examined with several different kinds of microsatellites to ensure that no MIN was present, thus excluding mismatch repair deficiency as a source of potential somatic alterations in these cases. The cell lines also were chosen to contain regions of loss of heterozygosity (LOH) in areas containing genes of interest to increase the sensitivity of detecting somatic mutations by DNA sequencing, as any alteration observed would appear as a homozygous change. The exons described above were amplified from tumor Fig. 1. Approach to somatic mutation discovery. After identifying genes of interest, the Celera and public genome databases were used to extract exon sequences and intronic regions to design primers for PCR amplification and sequencing. Successfully amplified and sequenced exons were assembled and compared with exon reference sequences to identify nonsynonymous alterations. Any potential alterations were then compared with polymorphism databases to exclude known SNPs. For remaining alterations, PCR products from matched normal and tumor DNA were sequenced to identify novel SNPs and somatic mutations. MEDICAL SCIENCES Wang et al. PNAS March 5, 2002 vol. 99 no
3 Table 2. Genes identified with somatic alterations Gene name Gene symbol Celera transcript ID GenBank accession Nucleotide change* Amino acid change Eukaryotic translation initiation factor 3, EIF3S1 hct30575 NM G T 267 D Y subunit 1 Neurofilament 3 (150 kd medium) NEF3 hct7649 NM C T 16 R W N A N A hct AA bp deletion Frameshift and stop at aa 59 *Nucleotide positions are relative to the transcript sequences present in the Celera Discovery System. Represents an unspliced expressed sequence tag that is homologous to one exon of the Celera predicted transcript. DNA samples using PCR and were subjected to dideoxy sequencing. A total of 1,811 exons were successfully amplified from 470 genes and sequenced from an average of 11 different tumor DNA samples (Table 1). The average size of examined exons was 150 bp and ranged up to 699 bp in length. Both intronic splice acceptor and donor regions were analyzed, with the exception of cases where the exon size was greater than the length of readable sequencing product. A total of 6.5 Mb of sequence data was obtained in this way (Table 1). Sequencing results were analyzed by assembling data of related exons from different tumors and comparing them in silico to reference sequences (Fig. 1). We confined our analysis to 3.2 Mb that encoded exons and focused on protein altering (nonsynonymous) changes, as these provided the best indicators of alterations most likely to have an effect on tumorigenesis. A total of 320 alterations resulting in nonsynonymous changes were identified in the 3.2-Mb assembly. Observed changes first were compared with polymorphism databases to determine whether they corresponded to previously recognized single nucleotide polymorphisms (SNPs). A total of 90 SNPs were identified and excluded in this manner. The remaining changes, representing potential somatic alterations, then were analyzed by sequencing of DNA from normal tissues of the corresponding patients. Two hundred and twenty seven of these changes were present in the corresponding patient s normal tissues. These changes thus represented previously unidentified SNPs within exons. After analysis of all potential changes by sequencing of both tumor and matched normal DNA samples, only three somatic alterations were identified, each in a different gene (Table 2). Two of these corresponded to missense changes in well characterized genes: agtottransversion at nucleotide position 799 of the eukaryotic translation initiation factor 3 subunit 1 gene, resulting in an aspartate to tyrosine change at amino acid position 267 (Fig. 2), and a C to T transition at nucleotide position 46 of the neurofilament 3 gene, resulting in an arginine to tryptophan change at amino acid position 16 (Fig. 3). The third alteration corresponded to a 14-bp deletion in a gene of unknown function (Celera hct ) at nucleotide position 154, resulting in a frameshift and introduction of premature translational termination codon at amino acid position 54 (Fig. 4). All three changes occurred in different tumor samples, and no mutations in the remaining exons of these genes were detected in the 10 additional tumors examined. The identification of three somatic alterations from 3.2 Mb of analyzed DNA corresponds to a somatic mutation frequency of approximately one alteration ( alterations, 95% probability interval) per Mb of tumor DNA, or approximately 3,000 alterations ( alterations, 95% probability interval) per haploid genome. However, this represents an underestimate of the number of somatic changes in a tumor genome, as we examined only nonsynonymous changes. On the basis of genome-wide polymorphism studies (21, 22), it has been shown that nonsynonymous and synonymous changes have similar frequencies in coding DNA, and that the frequency of coding and noncoding changes is approximately the same. Therefore, one would expect that our somatic mutation frequency would be increased approximately 2-fold. This probably represents the higher end of such an estimate, because approximately two-thirds of random coding mutations alter an amino acid, and the selection pressure on these nonsynonymous changes is likely to be substantially less at the single-cell level than at the organismal level. Accordingly, the majority of cancers would be expected to carry less than 6,000 alterations per genome. The total somatic alterations in a cancer represent those that accumulated during normal somatic cell division as well as in the tumor during successive waves of clonal expansion and growth. Each of the bottlenecks that the tumor cell passes through (including initiation plus each wave of clonal expansion) leads to fixation of all mutations that have previously occurred in the tumor cell s Fig. 2. Sequence chromatograms showing somatic alteration in eukaryotic translation initiation factor 3, subunit 1. Arrow indicates the location of nucleotide change in the tumor DNA sequence. Fig. 3. Sequence chromatograms showing somatic alteration in neurofilament 3 (150 kd medium). Arrow indicates the location of nucleotide change in the tumor DNA sequence cgi doi pnas Wang et al.
4 Fig. 4. Sequence chromatograms showing somatic alteration in Celera predicted transcript hct Arrow indicates the region of the 14-bp deletion in the tumor DNA sequence. progenitors. Colorectal cancers are thought to develop from epithelial stem cells residing in colonic crypts (23 25). As the entire intestinal epithelium replaces itself every few days, a typical colon epithelial stem cell is thought to undergo 100 cell divisions per year and 4,000 divisions before the onset of most adenomas at age 40. Additionally, as cell turnover far exceeds net tumor growth (2, 26, 27), during the 20-yr progression of a colorectal carcinoma, at least another 2,000 cell divisions would occur. By using the estimated mutation rate in normal somatic human cells of nucleotides per cell per division (1, 28, 29), one would expect that a 20-year old tumor removed from a 60-year old individual would have accumulated 1,800 18,000 alterations per haploid genome. This number of mutations is similar to the prevalence of alterations we observed in sporadic cancers and suggests that normal mutation rates are sufficient to explain tumor progression in most cancers. Like all large-scale studies, our approach has several limitations. First, the majority of genes analyzed, including the three that contained somatic alterations, were present in regions that had undergone LOH in the tumor DNA. Although there is no current evidence to suggest a different rate of point mutations in regions of LOH, it is possible that our conclusions would be altered by analyses of additional genes from heterozygous regions. Second, our analysis was performed on cell lines rather than primary tumors, which may allow for accumulation of mutations during in vitro passaging. However, all of the mutations observed were clonal in the tumors analyzed, making it unlikely that they developed during the few in vitro passages of the cell lines. This possibility, even if true, would only decrease the actual number of mutations calculated to be present in the tumors and, therefore, would not impact on the major conclusions of this study. Third, if we assume a hierarchical model of epithelial cell generation in colonic crypts rather than the conventional single stem cell model (30), only 50 instead of 4,000 cell divisions would occur in an average tumor progenitor cell, and the number of expected mutations per tumor genome would be reduced from a range of 1,800 18,000 to 600 6,000. Although the current experimental evidence does not support such a model (25), the reduced number of mutations remains within several-fold of our observed mutation rate within tumors and would suggest at best a weak increase in mutation rates in cancer cells. Despite these potential limitations, it is clear that these results have several significant implications. One implication is that they do not disprove the argument that genetic instability is an inherent feature of tumorigenesis. Instead, they emphasize that, although a subtle instability at the nucleotide level does not play a role in most colorectal cancers, a different form of genetic instability generally occurs in these tumors. In fact, extensive studies of polymorphic markers in our panel of tumors showed that they each had numerous allelic losses, consistent with widespread chromosomal changes (31). Additionally, the results highlight the importance of specific mutations in the genes that drive neoplasia. Because the studies described above show that passenger mutations (i.e., random mutations, with no functional significance) are found so rarely in these tumors, great confidence can be placed in the significance of somatic mutations in genes like APC and p53, which occur in nearly all of the colorectal cancers represented in our panel. Lastly, the results are important for interpreting the significance of mutations in candidate tumor-suppressor genes. The search for other tumorsuppressor genes involved in common cancers continues to be active, and the best evidence to validate such candidates comes from mutational data (32). On the one hand, our data show that mutations in any given gene are likely to be uncommon unless selected for during tumorigenesis. On the other hand, even random genes like those studied here are occasionally mutated in these cancers as a result of normal mutational processes and bottlenecks during the neoplastic process. A prudent conclusion from these results, therefore, would be to invoke the requirement for a significantly higher mutation frequency for example, at least 2 independent examples of functionally altering mutations from a panel of no more than 20 tumors to implicate a gene as a candidate tumor suppressor. To help investigators rigorously determine whether the number of observed mutations in a gene of interest is significantly above the expected background mutation frequency, we have developed simple statistical tools, freely available at http: astor.som.jhmi.edu gp trab. We thank members of the Molecular Genetics Laboratory for helpful discussions. This work was supported by the Maryland Cigarette Restitution Fund, the Clayton Fund, and National Institutes of Health Grants CA62924, CA57345, and CA MEDICAL SCIENCES Wang et al. PNAS March 5, 2002 vol. 99 no
5 1. Loeb, L. A. (2001) Cancer Res. 61, Tomlinson, I. & Bodmer, W. (1999) Nat. Med. 5, Boland, C. R. & Ricciardiello, L. (1999) Proc. Natl. Acad. Sci. USA 96, Lengauer, C., Kinzler, K. W. & Vogelstein, B. (1998) Nature (London) 396, Bootsma, D., Kraemer, K. H., Cleaver, J. E. & Hoeijmakers, J. H. J. (1998) in The Genetic Basis of Human Cancer, Vol. 1, eds. Vogelstein, B. & Kinzler, K. W. (McGraw Hill, New York), pp Lynch, H. T. & de La Chapelle, A. (1999) J. Med. Genet. 36, Perucho, M. (1996) Biol. Chem. 377, Boland, R. C. (1998) in The Genetic Basis of Human Cancer, Vol. 1, eds. Vogelstein, B. & Kinzler, K. W. (McGraw Hill, New York), pp Sia, E. A., Jinks-Robertson, S. & Petes, T. D. (1997) Mutat. Res. 383, Thomas, D. C., Umar, A. & Kunkel, T. A. (1996) Mutat. Res. 350, Strauss, B. S. (1997) Carcinogenesis 18, Stoler, D. L., Chen, N., Basik, M., Kahlenberg, M. S., Rodriguez-Bigas, M. A., Petrelli, N. J. & Anderson, G. R. (1999) Proc. Natl. Acad. Sci. USA 96, Rozen, S. & Skaletsky, H. (2000) Methods Mol. Biol. 132, Thiagalingam, S., Lengauer, C., Leach, F. S., Schutte, M., Hahn, S. A., Overhauser, J., Willson, J. K., Markowitz, S., Hamilton, S. R., Kern, S. E., et al. (1996) Nat. Genet. 13, Riggins, G. J., Thiagalingam, S., Rozenblum, E., Weinstein, C. L., Kern, S. E., Hamilton, S. R., Willson, J. K., Markowitz, S. D., Kinzler, K. W. & Vogelstein, B. (1996) Nat. Genet. 13, Cahill, D. P., Lengauer, C., Yu, J., Riggins, G. J., Willson, J. K., Markowitz, S. D., Kinzler, K. W. & Vogelstein, B. (1998) Nature (London) 392, Parsons, R., Myeroff, L. L., Liu, B., Willson, J. K., Markowitz, S. D., Kinzler, K. W. & Vogelstein, B. (1995) Cancer Res. 55, Schervish, M. (1995) Statistical Theory (Springer, New York). 19. Lander, E. S., Linton, L. M., Birren, B., Nusbaum, C., Zody, M. C., Baldwin, J., Devon, K., Dewar, K., Doyle, M., FitzHugh, W., et al. (2001) Nature (London) 409, Venter, J. C., Adams, M. D., Myers, E. W., Li, P. W., Mural, R. J., Sutton, G. G., Smith, H. O., Yandell, M., Evans, C. A., Holt, R. A., et al. (2001) Science 291, Cargill, M., Altshuler, D., Ireland, J., Sklar, P., Ardlie, K., Patil, N., Shaw, N., Lane, C. R., Lim, E. P., Kalyanaraman, N., et al. (1999) Nat. Genet. 22, Halushka, M. K., Fan, J. B., Bentley, K., Hsie, L., Shen, N., Weder, A., Cooper, R., Lipshutz, R. & Chakravarti, A. (1999) Nat. Genet. 22, Lipkin, M., Bell, B. & Shelrock, P. (1963) J. Clin. Invest. 42, Shorter, R. G., Moertel, C. G. & Titus, J. L. (1964) Am. J. Dig. Dis. 9, Potten, C. S. & Loeffler, M. (1990) Development (Cambridge, U.K.) 110, Matsui, T., Yao, T. & Iwashita, A. (2000) World J. Surg. 24, Shimomatsuya, T., Tanigawa, N. & Muraoka, R. (1991) Jpn. J. Cancer Res. 82, Drake, J. W. (1969) Nature (London) 221, Elmore, E., Kakunaga, T. & Barrett, J. C. (1983) Cancer Res. 43, Morris, J. A. (1999) J. Theor. Biol. 199, Thiagalingam, S., Laken, S., Willson, J. K., Markowitz, S. D., Kinzler, K. W., Vogelstein, B. & Lengauer, C. (2001) Proc. Natl. Acad. Sci. USA 98, Haber, D. & Harlow, E. (1997) Nat. Genet. 16, cgi doi pnas Wang et al.
Multistep nature of cancer development. Cancer genes
 Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Multistep nature of cancer development Phenotypic progression loss of control over cell growth/death (neoplasm) invasiveness (carcinoma) distal spread (metastatic tumor) Genetic progression multiple genetic
Tumor suppressor genes D R. S H O S S E I N I - A S L
 Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
Cancer develops after somatic mutations overcome the multiple
 Genetic variation in cancer predisposition: Mutational decay of a robust genetic control network Steven A. Frank* Department of Ecology and Evolutionary Biology, University of California, Irvine, CA 92697-2525
Genetic variation in cancer predisposition: Mutational decay of a robust genetic control network Steven A. Frank* Department of Ecology and Evolutionary Biology, University of California, Irvine, CA 92697-2525
Chapter 5. Gazzoli I, Loda M, Garber J, Syngal S and Kolodner RD. Cancer Research 2002 Jul 15; 62(14):
 Chapter 5 A hereditary nonpolyposis colorectal carcinoma case associated with hypermethylation of the MLH1 gene in normal tissue and loss of heterozygosity of the unmethylated allele in the resulting microsatellite
Chapter 5 A hereditary nonpolyposis colorectal carcinoma case associated with hypermethylation of the MLH1 gene in normal tissue and loss of heterozygosity of the unmethylated allele in the resulting microsatellite
Accumulated Clonal Genetic Alterations in Familial and Sporadic Colorectal Carcinomas with Widespread Instability in Microsatellite Sequences
 American Journal of Pathology, Vol. 153, No. 4, October 1998 Copyright American Society for Investigative Pathology Accumulated Clonal Genetic Alterations in Familial and Sporadic Colorectal Carcinomas
American Journal of Pathology, Vol. 153, No. 4, October 1998 Copyright American Society for Investigative Pathology Accumulated Clonal Genetic Alterations in Familial and Sporadic Colorectal Carcinomas
MRC-Holland MLPA. Description version 29;
 SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
Development of Carcinoma Pathways
 The Construction of Genetic Pathway to Colorectal Cancer Moriah Wright, MD Clinical Fellow in Colorectal Surgery Creighton University School of Medicine Management of Colon and Diseases February 23, 2019
The Construction of Genetic Pathway to Colorectal Cancer Moriah Wright, MD Clinical Fellow in Colorectal Surgery Creighton University School of Medicine Management of Colon and Diseases February 23, 2019
CANCER. Inherited Cancer Syndromes. Affects 25% of US population. Kills 19% of US population (2nd largest killer after heart disease)
 CANCER Affects 25% of US population Kills 19% of US population (2nd largest killer after heart disease) NOT one disease but 200-300 different defects Etiologic Factors In Cancer: Relative contributions
CANCER Affects 25% of US population Kills 19% of US population (2nd largest killer after heart disease) NOT one disease but 200-300 different defects Etiologic Factors In Cancer: Relative contributions
LESSON 3.2 WORKBOOK. How do normal cells become cancer cells? Workbook Lesson 3.2
 For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
2003 Landes Bioscience. Not for distribution.
 [Cancer Biology & Therapy 1:6, 685-692, November/December 2002]; 2002 Landes Bioscience Research Paper Dynamics of Genetic Instability in Sporadic and Familial Colorectal Cancer Natalia L. Komarova 1,2
[Cancer Biology & Therapy 1:6, 685-692, November/December 2002]; 2002 Landes Bioscience Research Paper Dynamics of Genetic Instability in Sporadic and Familial Colorectal Cancer Natalia L. Komarova 1,2
Asingle inherited mutant gene may be enough to
 396 Cancer Inheritance STEVEN A. FRANK Asingle inherited mutant gene may be enough to cause a very high cancer risk. Single-mutation cases have provided much insight into the genetic basis of carcinogenesis,
396 Cancer Inheritance STEVEN A. FRANK Asingle inherited mutant gene may be enough to cause a very high cancer risk. Single-mutation cases have provided much insight into the genetic basis of carcinogenesis,
Cancer and sornat ic evolution
 Chapter 1 Cancer and sornat ic evolution 1.1 What is cancer? The development and healthy life of a human being requires the cooperation of more than ten million cells for the good of the organism. This
Chapter 1 Cancer and sornat ic evolution 1.1 What is cancer? The development and healthy life of a human being requires the cooperation of more than ten million cells for the good of the organism. This
CANCER GENETICS PROVIDER SURVEY
 Dear Participant, Previously you agreed to participate in an evaluation of an education program we developed for primary care providers on the topic of cancer genetics. This is an IRB-approved, CDCfunded
Dear Participant, Previously you agreed to participate in an evaluation of an education program we developed for primary care providers on the topic of cancer genetics. This is an IRB-approved, CDCfunded
SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407
 SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407 The Mismatch Repair (MMR) system is critical for the maintenance of genomic stability. MMR increases the fidelity of DNA
SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407 The Mismatch Repair (MMR) system is critical for the maintenance of genomic stability. MMR increases the fidelity of DNA
Landes Bioscience Not for distribution.
 [Cell Cycle 3:3, 358-362; March 2004]; 2004 Landes Bioscience Report Linear Model of Colon Cancer Initiation Franziska Michor 1 Yoh Iwasa 2 Harith Rajagopalan 3 Christoph Lengauer 3 Martin A. Nowak 1,
[Cell Cycle 3:3, 358-362; March 2004]; 2004 Landes Bioscience Report Linear Model of Colon Cancer Initiation Franziska Michor 1 Yoh Iwasa 2 Harith Rajagopalan 3 Christoph Lengauer 3 Martin A. Nowak 1,
HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007
 MIT OpenCourseWare http://ocw.mit.edu HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
The concept that tumors develop through the accumulation of
 The role of chromosomal instability in tumor initiation Martin A. Nowak, Natalia L. Komarova, Anirvan Sengupta, Prasad V. Jallepalli, Ie-Ming Shih**, Bert Vogelstein**, and Christoph Lengauer Institute
The role of chromosomal instability in tumor initiation Martin A. Nowak, Natalia L. Komarova, Anirvan Sengupta, Prasad V. Jallepalli, Ie-Ming Shih**, Bert Vogelstein**, and Christoph Lengauer Institute
Colon Cancer and Hereditary Cancer Syndromes
 Colon Cancer and Hereditary Cancer Syndromes Gisela Keller Institute of Pathology Technische Universität München gisela.keller@lrz.tum.de Colon Cancer and Hereditary Cancer Syndromes epidemiology models
Colon Cancer and Hereditary Cancer Syndromes Gisela Keller Institute of Pathology Technische Universität München gisela.keller@lrz.tum.de Colon Cancer and Hereditary Cancer Syndromes epidemiology models
Can chromosomal instability initiate tumorigenesis?
 Seminars in Cancer Biology 15 (2005) 43 49 Review Can chromosomal instability initiate tumorigenesis? Franziska Michor a, Yoh Iwasa b, Bert Vogelstein c, Christoph Lengauer c, Martin A. Nowak a, a Program
Seminars in Cancer Biology 15 (2005) 43 49 Review Can chromosomal instability initiate tumorigenesis? Franziska Michor a, Yoh Iwasa b, Bert Vogelstein c, Christoph Lengauer c, Martin A. Nowak a, a Program
The Human Major Histocompatibility Complex
 The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
International Journal of Science, Environment and Technology, Vol. 6, No 5, 2017,
 International Journal of Science, Environment and Technology, Vol. 6, No 5, 2017, 2792 2796 ISSN 2278-3687 (O) 2277-663X (P) POLYMORPHISM IN EXON 9 OF HEAT SHOCK TRANSCRIPTION FACTOR 4 (HSF4) GENE IN EXOTIC
International Journal of Science, Environment and Technology, Vol. 6, No 5, 2017, 2792 2796 ISSN 2278-3687 (O) 2277-663X (P) POLYMORPHISM IN EXON 9 OF HEAT SHOCK TRANSCRIPTION FACTOR 4 (HSF4) GENE IN EXOTIC
B Base excision repair, in MUTYH-associated polyposis and colorectal cancer, BRAF testing, for hereditary colorectal cancer, 696
 Index Note: Page numbers of article titles are in boldface type. A Adenomatous polyposis, familial. See Familial adenomatous polyposis. Anal anastomosis, ileal-pouch, proctocolectomy with, in FAP, 591
Index Note: Page numbers of article titles are in boldface type. A Adenomatous polyposis, familial. See Familial adenomatous polyposis. Anal anastomosis, ileal-pouch, proctocolectomy with, in FAP, 591
Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3.
 Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3. MN1 (Accession No. NM_002430) MN1-1514F 5 -GGCTGTCATGCCCTATTGAT Exon 1 MN1-1882R 5 -CTGGTGGGGATGATGACTTC Exon
Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3. MN1 (Accession No. NM_002430) MN1-1514F 5 -GGCTGTCATGCCCTATTGAT Exon 1 MN1-1882R 5 -CTGGTGGGGATGATGACTTC Exon
Anatomic Molecular Pathology: An Emerging Field
 Anatomic Molecular Pathology: An Emerging Field Antonia R. Sepulveda M.D., Ph.D. University of Pennsylvania asepu@mail.med.upenn.edu 2008 ASIP Annual Meeting Anatomic pathology (U.S.) is a medical specialty
Anatomic Molecular Pathology: An Emerging Field Antonia R. Sepulveda M.D., Ph.D. University of Pennsylvania asepu@mail.med.upenn.edu 2008 ASIP Annual Meeting Anatomic pathology (U.S.) is a medical specialty
Role of Paired Box9 (PAX9) (rs ) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis
 EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
Clinicopathologic Characteristics of Left-Sided Colon Cancers with High Microsatellite Instability
 The Korean Journal of Pathology 29; 43: 428-34 DOI: 1.4132/KoreanJPathol.29.43.5.428 Clinicopathologic Characteristics of Left-Sided Colon Cancers with High Microsatellite Instability Sang Kyum Kim Junjeong
The Korean Journal of Pathology 29; 43: 428-34 DOI: 1.4132/KoreanJPathol.29.43.5.428 Clinicopathologic Characteristics of Left-Sided Colon Cancers with High Microsatellite Instability Sang Kyum Kim Junjeong
Molecular mechanisms of human carcinogenesis
 Cancer: Cell Structures, Carcinogens and Genomic Instability Edited by Leon P. Bignold 2006 Birkhäuser Verlag/Switzerland 321 Molecular mechanisms of human carcinogenesis William B. Coleman 1 and Gregory
Cancer: Cell Structures, Carcinogens and Genomic Instability Edited by Leon P. Bignold 2006 Birkhäuser Verlag/Switzerland 321 Molecular mechanisms of human carcinogenesis William B. Coleman 1 and Gregory
New: P077 BRCA2. This new probemix can be used to confirm results obtained with P045 BRCA2 probemix.
 SALSA MLPA KIT P045-B2 BRCA2/CHEK2 Lot 0410, 0609. As compared to version B1, four reference probes have been replaced and extra control fragments at 100 and 105 nt (X/Y specific) have been included. New:
SALSA MLPA KIT P045-B2 BRCA2/CHEK2 Lot 0410, 0609. As compared to version B1, four reference probes have been replaced and extra control fragments at 100 and 105 nt (X/Y specific) have been included. New:
Introduction. Chapter 1
 Introduction Chapter 1 1.1 Colorectal cancer Transformation from normal cell to cancer cell is thought to be a multi-step process involving the accumulation of genetic alterations in oncogenes, tumor
Introduction Chapter 1 1.1 Colorectal cancer Transformation from normal cell to cancer cell is thought to be a multi-step process involving the accumulation of genetic alterations in oncogenes, tumor
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Computational Systems Biology: Biology X
 Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
Introduction to Cancer Biology
 Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Célia DeLozier-Blanchet
 The Genetics Consultation in OB-GYN : Hereditary cancers Célia DeLozier-Blanchet Division of Medical Genetics, Geneva University Hospital It is probable that all cancers are genetic! genetic vs. hereditary
The Genetics Consultation in OB-GYN : Hereditary cancers Célia DeLozier-Blanchet Division of Medical Genetics, Geneva University Hospital It is probable that all cancers are genetic! genetic vs. hereditary
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Hereditary non polyposis colorectal cancer in a random sample of colorectal cancer patients
 Basic Science Hereditary non polyposis colorectal cancer in a random sample of colorectal cancer patients Nada Pavlović-Čalić 1, Izet Eminović 2, Vesna Hadžiabdić 3, Kasim Muminhodžić 1, Radovan Komel
Basic Science Hereditary non polyposis colorectal cancer in a random sample of colorectal cancer patients Nada Pavlović-Čalić 1, Izet Eminović 2, Vesna Hadžiabdić 3, Kasim Muminhodžić 1, Radovan Komel
Comments on Significance of candidate cancer genes as assessed by the CaMP score by Parmigiani et al.
 Comments on Significance of candidate cancer genes as assessed by the CaMP score by Parmigiani et al. Holger Höfling Gad Getz Robert Tibshirani June 26, 2007 1 Introduction Identifying genes that are involved
Comments on Significance of candidate cancer genes as assessed by the CaMP score by Parmigiani et al. Holger Höfling Gad Getz Robert Tibshirani June 26, 2007 1 Introduction Identifying genes that are involved
The first attempts to formulate a quantitative description of
 Multistage carcinogenesis and the incidence of colorectal cancer E. Georg Luebeck* and Suresh H. Moolgavkar Fred Hutchinson Cancer Research Center, 1100 Fairview Avenue North, P.O. Box 19024, Seattle,
Multistage carcinogenesis and the incidence of colorectal cancer E. Georg Luebeck* and Suresh H. Moolgavkar Fred Hutchinson Cancer Research Center, 1100 Fairview Avenue North, P.O. Box 19024, Seattle,
MRC-Holland MLPA. Description version 29; 31 July 2015
 SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0114. As compared to the previous B2 version (lot 0813 and 0912), 11 target probes are replaced or added, and 10 new reference probes are included. P082
SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0114. As compared to the previous B2 version (lot 0813 and 0912), 11 target probes are replaced or added, and 10 new reference probes are included. P082
DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK
 CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
MODULE NO.14: Y-Chromosome Testing
 SUBJECT Paper No. and Title Module No. and Title Module Tag FORENSIC SIENCE PAPER No.13: DNA Forensics MODULE No.21: Y-Chromosome Testing FSC_P13_M21 TABLE OF CONTENTS 1. Learning Outcome 2. Introduction:
SUBJECT Paper No. and Title Module No. and Title Module Tag FORENSIC SIENCE PAPER No.13: DNA Forensics MODULE No.21: Y-Chromosome Testing FSC_P13_M21 TABLE OF CONTENTS 1. Learning Outcome 2. Introduction:
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/22278 holds various files of this Leiden University dissertation. Author: Cunha Carvalho de Miranda, Noel Filipe da Title: Mismatch repair and MUTYH deficient
Cover Page The handle http://hdl.handle.net/1887/22278 holds various files of this Leiden University dissertation. Author: Cunha Carvalho de Miranda, Noel Filipe da Title: Mismatch repair and MUTYH deficient
Genome 371, Autumn 2018 Quiz Section 9: Genetics of Cancer Worksheet
 Genome 371, Autumn 2018 Quiz Section 9: Genetics of Cancer Worksheet All cancer is due to genetic mutations. However, in cancer that clusters in families (familial cancer) at least one of these mutations
Genome 371, Autumn 2018 Quiz Section 9: Genetics of Cancer Worksheet All cancer is due to genetic mutations. However, in cancer that clusters in families (familial cancer) at least one of these mutations
Genetic progression and the waiting time to cancer
 Genetic progression and the waiting time to cancer Niko Beerenwinkel *, Tibor Antal, David Dingli, Arne Traulsen, Kenneth W. Kinzler, Victor E. Velculescu, Bert Vogelstein, Martin A. Nowak 0 Program for
Genetic progression and the waiting time to cancer Niko Beerenwinkel *, Tibor Antal, David Dingli, Arne Traulsen, Kenneth W. Kinzler, Victor E. Velculescu, Bert Vogelstein, Martin A. Nowak 0 Program for
MRC-Holland MLPA. Description version 06; 23 December 2016
 SALSA MLPA probemix P417-B2 BAP1 Lot B2-1216. As compared to version B1 (lot B1-0215), two reference probes have been added and two target probes have a minor change in length. The BAP1 (BRCA1 associated
SALSA MLPA probemix P417-B2 BAP1 Lot B2-1216. As compared to version B1 (lot B1-0215), two reference probes have been added and two target probes have a minor change in length. The BAP1 (BRCA1 associated
oncogenes-and- tumour-suppressor-genes)
 Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
MRC-Holland MLPA. Description version 30; 06 June 2017
 SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0517, C1-0114. As compared to the previous B2 version (lot B2-0813, B2-0912), 11 target probes are replaced or added, and 10 new reference probes are
SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0517, C1-0114. As compared to the previous B2 version (lot B2-0813, B2-0912), 11 target probes are replaced or added, and 10 new reference probes are
Chapt 15: Molecular Genetics of Cell Cycle and Cancer
 Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
Ovarian cancer: 2012 Update Srini Prasad MD Univ Texas MD Anderson Cancer Center
 Ovarian cancer: 2012 Update Srini Prasad MD Univ Texas MD Anderson Cancer Center Ovarian cancer is not a single disease Ovarian Epithelial Tumors: Histological Spectrum* Type Frequency Histology High-Grade
Ovarian cancer: 2012 Update Srini Prasad MD Univ Texas MD Anderson Cancer Center Ovarian cancer is not a single disease Ovarian Epithelial Tumors: Histological Spectrum* Type Frequency Histology High-Grade
Keywords: microsatellite instability; multiple primary cancer; hereditary non-polyposis colorectal cancer
 790 Gut 2000;46:790 794 First Department of Internal Medicine, Sapporo Medical University, Sapporo, K Yamashita Y Arimura S Kurokawa F Itoh T Endo K Imai First Department of Surgery, Sapporo Medical University,
790 Gut 2000;46:790 794 First Department of Internal Medicine, Sapporo Medical University, Sapporo, K Yamashita Y Arimura S Kurokawa F Itoh T Endo K Imai First Department of Surgery, Sapporo Medical University,
MUTATIONS, MUTAGENESIS, AND CARCINOGENESIS. (Start your clickers)
 MUTATIONS, MUTAGENESIS, AND CARCINOGENESIS (Start your clickers) How do mutations arise? And how do they affect a cell and its organism? Mutations: heritable changes in genes Mutations occur in DNA But
MUTATIONS, MUTAGENESIS, AND CARCINOGENESIS (Start your clickers) How do mutations arise? And how do they affect a cell and its organism? Mutations: heritable changes in genes Mutations occur in DNA But
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Clonal evolution of human cancers
 Clonal evolution of human cancers -Pathology-based microdissection and genetic analysis precisely demonstrates molecular evolution of neoplastic clones- Hiroaki Fujii, MD Ageo Medical Laboratories, Yashio
Clonal evolution of human cancers -Pathology-based microdissection and genetic analysis precisely demonstrates molecular evolution of neoplastic clones- Hiroaki Fujii, MD Ageo Medical Laboratories, Yashio
SALSA MLPA probemix P169-C2 HIRSCHSPRUNG-1 Lot C As compared to version C1 (lot C1-0612), the length of one probe has been adjusted.
 mix P169-C2 HIRSCHSPRUNG-1 Lot C2-0915. As compared to version C1 (lot C1-0612), the length of one has been adjusted. Hirschsprung disease (HSCR), or aganglionic megacolon, is a congenital disorder characterised
mix P169-C2 HIRSCHSPRUNG-1 Lot C2-0915. As compared to version C1 (lot C1-0612), the length of one has been adjusted. Hirschsprung disease (HSCR), or aganglionic megacolon, is a congenital disorder characterised
Microsatellite instability is a rare phenomenon in transition from chronic to blastic phase chronic myeloid leukemia
 Turkish Journal of Cancer Vol.31/ No. 2/2001 Microsatellite instability is a rare phenomenon in transition from chronic to blastic phase chronic myeloid leukemia H. OGÜN SERCAN, ZEYNEP Y. SERCAN, SEFA
Turkish Journal of Cancer Vol.31/ No. 2/2001 Microsatellite instability is a rare phenomenon in transition from chronic to blastic phase chronic myeloid leukemia H. OGÜN SERCAN, ZEYNEP Y. SERCAN, SEFA
Introduction to Genetics
 Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
Cancer genetics
 Cancer genetics General information about tumorogenesis. Cancer induced by viruses. The role of somatic mutations in cancer production. Oncogenes and Tumor Suppressor Genes (TSG). Hereditary cancer. 1
Cancer genetics General information about tumorogenesis. Cancer induced by viruses. The role of somatic mutations in cancer production. Oncogenes and Tumor Suppressor Genes (TSG). Hereditary cancer. 1
Karyotype analysis reveals transloction of chromosome 22 to 9 in CML chronic myelogenous leukemia has fusion protein Bcr-Abl
 Chapt. 18 Cancer Molecular Biology of Cancer Student Learning Outcomes: Describe cancer diseases in which cells no longer respond Describe how cancers come from genomic mutations (inherited or somatic)
Chapt. 18 Cancer Molecular Biology of Cancer Student Learning Outcomes: Describe cancer diseases in which cells no longer respond Describe how cancers come from genomic mutations (inherited or somatic)
MEDICAL GENOMICS LABORATORY. Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG)
 Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
American Society of Cytopathology Core Curriculum in Molecular Biology
 American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 1 Molecular Basis of Cancer Molecular Oncology Keisha
American Society of Cytopathology Core Curriculum in Molecular Biology American Society of Cytopathology Core Curriculum in Molecular Biology Chapter 1 Molecular Basis of Cancer Molecular Oncology Keisha
Section D: The Molecular Biology of Cancer
 CHAPTER 19 THE ORGANIZATION AND CONTROL OF EUKARYOTIC GENOMES Section D: The Molecular Biology of Cancer 1. Cancer results from genetic changes that affect the cell cycle 2. Oncogene proteins and faulty
CHAPTER 19 THE ORGANIZATION AND CONTROL OF EUKARYOTIC GENOMES Section D: The Molecular Biology of Cancer 1. Cancer results from genetic changes that affect the cell cycle 2. Oncogene proteins and faulty
Risk of Colorectal Cancer (CRC) Hereditary Syndromes in GI Cancer GENETIC MALPRACTICE
 Identifying the Patient at Risk for an Inherited Syndrome Sapna Syngal, MD, MPH, FACG Director, Gastroenterology Director, Familial GI Program Dana-Farber/Brigham and Women s Cancer Center Associate Professor
Identifying the Patient at Risk for an Inherited Syndrome Sapna Syngal, MD, MPH, FACG Director, Gastroenterology Director, Familial GI Program Dana-Farber/Brigham and Women s Cancer Center Associate Professor
Chapter 9. Cells Grow and Reproduce
 Chapter 9 Cells Grow and Reproduce DNA Replication DNA polymerase Addition of a nucleotide to the 3 end of a growing strand Use dntps as substrate Release of pyrophosphate Initiation of Replication Replication
Chapter 9 Cells Grow and Reproduce DNA Replication DNA polymerase Addition of a nucleotide to the 3 end of a growing strand Use dntps as substrate Release of pyrophosphate Initiation of Replication Replication
Lynch Syndrome Screening for Endometrial Cancer: Basic Concepts 1/16/2017
 1 Hi, my name is Sarah Kerr. I m a pathologist at Mayo Clinic, where I participate in our high volume Lynch syndrome tumor testing practice. Today I hope to cover some of the basics needed to understand
1 Hi, my name is Sarah Kerr. I m a pathologist at Mayo Clinic, where I participate in our high volume Lynch syndrome tumor testing practice. Today I hope to cover some of the basics needed to understand
The molecular genetics of endometrial cancer
 The molecular genetics of endometrial cancer Lora Hedrick Ellenson, M.D. Department of Pathology and Laboratory Medicine Weill Medical College of Cornell University Introduction Classification of endometrial
The molecular genetics of endometrial cancer Lora Hedrick Ellenson, M.D. Department of Pathology and Laboratory Medicine Weill Medical College of Cornell University Introduction Classification of endometrial
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Policy Specific Section: Medical Necessity and Investigational / Experimental. October 14, 1998 March 28, 2014
 Medical Policy Genetic Testing for Colorectal Cancer Type: Medical Necessity and Investigational / Experimental Policy Specific Section: Laboratory/Pathology Original Policy Date: Effective Date: October
Medical Policy Genetic Testing for Colorectal Cancer Type: Medical Necessity and Investigational / Experimental Policy Specific Section: Laboratory/Pathology Original Policy Date: Effective Date: October
Genome - Wide Linkage Mapping
 Biological Sciences Initiative HHMI Genome - Wide Linkage Mapping Introduction This activity is based on the work of Dr. Christine Seidman et al that was published in Circulation, 1998, vol 97, pgs 2043-2048.
Biological Sciences Initiative HHMI Genome - Wide Linkage Mapping Introduction This activity is based on the work of Dr. Christine Seidman et al that was published in Circulation, 1998, vol 97, pgs 2043-2048.
Microsatellite Instability and Mismatch Repair Protein (hmlh1, hmsh2) Expression in Intrahepatic Cholangiocarcinoma
 The Korean Journal of Pathology 2005; 39: 9-14 Microsatellite Instability and Mismatch Repair Protein (hmlh1, hmsh2) Expression in Intrahepatic Cholangiocarcinoma Yun Kyung Kang Woo Ho Kim 1 Department
The Korean Journal of Pathology 2005; 39: 9-14 Microsatellite Instability and Mismatch Repair Protein (hmlh1, hmsh2) Expression in Intrahepatic Cholangiocarcinoma Yun Kyung Kang Woo Ho Kim 1 Department
Familial and Hereditary Colon Cancer
 Familial and Hereditary Colon Cancer Aasma Shaukat, MD, MPH, FACG, FASGE, FACP GI Section Chief, Minneapolis VAMC Associate Professor, Division of Gastroenterology, Department of Medicine, University of
Familial and Hereditary Colon Cancer Aasma Shaukat, MD, MPH, FACG, FASGE, FACP GI Section Chief, Minneapolis VAMC Associate Professor, Division of Gastroenterology, Department of Medicine, University of
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Lynch Syndrome. Angie Strang, PGY2
 Lynch Syndrome Angie Strang, PGY2 Background Previously hereditary nonpolyposis colorectal cancer Autosomal dominant inherited cancer susceptibility syndrome Caused by defects in the mismatch repair system
Lynch Syndrome Angie Strang, PGY2 Background Previously hereditary nonpolyposis colorectal cancer Autosomal dominant inherited cancer susceptibility syndrome Caused by defects in the mismatch repair system
T.S. IEFIMENKO, M. Z. ANTONYUK, V.S. MARTYNENKO
 - -, -. 1. Panayotov I., Tsujimoto H., Fertility restoration and NOR suppression caused by Aegilops mutica chromosomes in alloplasmic hybrids and lines // Euphytica. 1997. Vol. 94, 2. P. 145 149. 2. Eser
- -, -. 1. Panayotov I., Tsujimoto H., Fertility restoration and NOR suppression caused by Aegilops mutica chromosomes in alloplasmic hybrids and lines // Euphytica. 1997. Vol. 94, 2. P. 145 149. 2. Eser
Molecular biology of colorectal cancer
 Molecular biology of colorectal cancer Phil Quirke Yorkshire Cancer Research Centenary Professor of Pathology University of Leeds, UK Rapid pace of molecular change Sequencing changes 2012 1,000 genomes
Molecular biology of colorectal cancer Phil Quirke Yorkshire Cancer Research Centenary Professor of Pathology University of Leeds, UK Rapid pace of molecular change Sequencing changes 2012 1,000 genomes
Colonic polyps and colon cancer. Andrew Macpherson Director of Gastroentology University of Bern
 Colonic polyps and colon cancer Andrew Macpherson Director of Gastroentology University of Bern Improtance of the problem of colon cancers - Epidemiology Lifetime risk 5% Incidence/10 5 /annum (US Detroit
Colonic polyps and colon cancer Andrew Macpherson Director of Gastroentology University of Bern Improtance of the problem of colon cancers - Epidemiology Lifetime risk 5% Incidence/10 5 /annum (US Detroit
Genetics and Genomics in Endocrinology
 Genetics and Genomics in Endocrinology Dr. Peter Igaz MD MSc PhD 2 nd Department of Medicine Faculty of Medicine Semmelweis University Genetics-based endocrine diseases I. Monogenic diseases: Multiple
Genetics and Genomics in Endocrinology Dr. Peter Igaz MD MSc PhD 2 nd Department of Medicine Faculty of Medicine Semmelweis University Genetics-based endocrine diseases I. Monogenic diseases: Multiple
NEXT GENERATION SEQUENCING OPENS NEW VIEWS ON VIRUS EVOLUTION AND EPIDEMIOLOGY. 16th International WAVLD symposium, 10th OIE Seminar
 NEXT GENERATION SEQUENCING OPENS NEW VIEWS ON VIRUS EVOLUTION AND EPIDEMIOLOGY S. Van Borm, I. Monne, D. King and T. Rosseel 16th International WAVLD symposium, 10th OIE Seminar 07.06.2013 Viral livestock
NEXT GENERATION SEQUENCING OPENS NEW VIEWS ON VIRUS EVOLUTION AND EPIDEMIOLOGY S. Van Borm, I. Monne, D. King and T. Rosseel 16th International WAVLD symposium, 10th OIE Seminar 07.06.2013 Viral livestock
colorectal cancer Colorectal cancer hereditary sporadic Familial 1/12/2018
 colorectal cancer Adenocarcinoma of the colon and rectum is the third most common site of new cancer cases and deaths in men (following prostate and lung or bronchus cancer) and women (following breast
colorectal cancer Adenocarcinoma of the colon and rectum is the third most common site of new cancer cases and deaths in men (following prostate and lung or bronchus cancer) and women (following breast
BASIC CONCEPTS OF CANCER: GENOMIC DETERMINATION
 BASIC CONCEPTS OF CANCER: GENOMIC DETERMINATION Edith Oláh Corresponding author s address: Prof. Edith Olah, Ph.D., D.Sc. Department of Molecular Genetics, National Institute of Oncology, H-1525 Budapest,
BASIC CONCEPTS OF CANCER: GENOMIC DETERMINATION Edith Oláh Corresponding author s address: Prof. Edith Olah, Ph.D., D.Sc. Department of Molecular Genetics, National Institute of Oncology, H-1525 Budapest,
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
6.3 DNA Mutations. SBI4U Ms. Ho-Lau
 6.3 DNA Mutations SBI4U Ms. Ho-Lau DNA Mutations Gene expression can be affected by errors that occur during DNA replication. Some errors are repaired, but others can become mutations (changes in the nucleotide
6.3 DNA Mutations SBI4U Ms. Ho-Lau DNA Mutations Gene expression can be affected by errors that occur during DNA replication. Some errors are repaired, but others can become mutations (changes in the nucleotide
MRC-Holland MLPA. Description version 08; 18 November 2016
 SALSA MLPA probemix P122-D1 NF1 AREA Lot D1-1016. As compared to lot C2-0312, four probes in the NF1 area and one reference probe have been removed, four reference probes have been replaced and several
SALSA MLPA probemix P122-D1 NF1 AREA Lot D1-1016. As compared to lot C2-0312, four probes in the NF1 area and one reference probe have been removed, four reference probes have been replaced and several
Is ALS a multistep process?
 Is ALS a multistep process? Neil Pearce, London School of Hygiene and Tropical Medicine Ammar Al-Chalabi, Institute of Psychiatry, King s College London Zoe Rutter-Locher, King s College London Is ALS
Is ALS a multistep process? Neil Pearce, London School of Hygiene and Tropical Medicine Ammar Al-Chalabi, Institute of Psychiatry, King s College London Zoe Rutter-Locher, King s College London Is ALS
Colorectal adenocarcinoma leading cancer in developed countries In US, annual deaths due to colorectal adenocarcinoma 57,000.
 Colonic Neoplasia Remotti Colorectal adenocarcinoma leading cancer in developed countries In US, annual incidence of colorectal adenocarcinoma 150,000. In US, annual deaths due to colorectal adenocarcinoma
Colonic Neoplasia Remotti Colorectal adenocarcinoma leading cancer in developed countries In US, annual incidence of colorectal adenocarcinoma 150,000. In US, annual deaths due to colorectal adenocarcinoma
Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell
 Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell Wenxin Li Department of Biological Sciences Fordham University Abstract MEFV is a human gene that codes for an
Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell Wenxin Li Department of Biological Sciences Fordham University Abstract MEFV is a human gene that codes for an
Familial and Hereditary Colon Cancer
 Familial and Hereditary Colon Cancer Aasma Shaukat, MD, MPH, FACG, FASGE, FACP GI Section Chief, Minneapolis VAMC Associate Professor, Division of Gastroenterology, Department of Medicine, University of
Familial and Hereditary Colon Cancer Aasma Shaukat, MD, MPH, FACG, FASGE, FACP GI Section Chief, Minneapolis VAMC Associate Professor, Division of Gastroenterology, Department of Medicine, University of
MRC-Holland MLPA. Description version 12; 13 January 2017
 SALSA MLPA probemix P219-B3 PAX6 Lot B3-0915: Compared to version B2 (lot B2-1111) two reference probes have been replaced and one additional reference probe has been added. In addition, one flanking probe
SALSA MLPA probemix P219-B3 PAX6 Lot B3-0915: Compared to version B2 (lot B2-1111) two reference probes have been replaced and one additional reference probe has been added. In addition, one flanking probe
SALSA MLPA probemix P241-D2 MODY mix 1 Lot D As compared to version D1 (lot D1-0911), one reference probe has been replaced.
 mix P241-D2 MODY mix 1 Lot D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent diabetes
mix P241-D2 MODY mix 1 Lot D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent diabetes
GENETICS OF COLORECTAL CANCER: HEREDITARY ASPECTS By. Magnitude of the Problem. Magnitude of the Problem. Cardinal Features of Lynch Syndrome
 GENETICS OF COLORECTAL CANCER: HEREDITARY ASPECTS By HENRY T. LYNCH, M.D. 1 Could this be hereditary Colon Cancer 4 Creighton University School of Medicine Omaha, Nebraska Magnitude of the Problem Annual
GENETICS OF COLORECTAL CANCER: HEREDITARY ASPECTS By HENRY T. LYNCH, M.D. 1 Could this be hereditary Colon Cancer 4 Creighton University School of Medicine Omaha, Nebraska Magnitude of the Problem Annual
Disorders of Cell Growth & Neoplasia. Lecture 4 Molecular basis of cancer
 General Pathology VPM 152 Disorders of Cell Growth & Neoplasia Lecture 4 Molecular basis of cancer Enrique Aburto Apr 2010 Skin tumor in a 10-year-old Rottweiler. Considering the external appearance and
General Pathology VPM 152 Disorders of Cell Growth & Neoplasia Lecture 4 Molecular basis of cancer Enrique Aburto Apr 2010 Skin tumor in a 10-year-old Rottweiler. Considering the external appearance and
The Biology and Genetics of Cells and Organisms The Biology of Cancer
 The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
The Biology and Genetics of Cells and Organisms The Biology of Cancer Mendel and Genetics How many distinct genes are present in the genomes of mammals? - 21,000 for human. - Genetic information is carried
Advances in genetic diagnosis of neurological disorders
 Acta Neurol Scand 2014: 129 (Suppl. 198): 20 25 DOI: 10.1111/ane.12232 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd ACTA NEUROLOGICA SCANDINAVICA Review Article Advances in genetic diagnosis
Acta Neurol Scand 2014: 129 (Suppl. 198): 20 25 DOI: 10.1111/ane.12232 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd ACTA NEUROLOGICA SCANDINAVICA Review Article Advances in genetic diagnosis
The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation
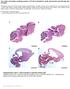 The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation Anita Becker-Heck#, Irene Zohn#, Noriko Okabe#, Andrew Pollock#, Kari Baker Lenhart,
The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation Anita Becker-Heck#, Irene Zohn#, Noriko Okabe#, Andrew Pollock#, Kari Baker Lenhart,
Gene finding. kuobin/
 Gene finding KUO-BIN LI, PH.D. http://www.bii.a-star.edu.sg/ kuobin/ Bioinformatics Institute 30 Medical Drive, Level 1, IMCB Building Singapore 117609 Republic of Singapore Gene finding (LSM5191) p.1
Gene finding KUO-BIN LI, PH.D. http://www.bii.a-star.edu.sg/ kuobin/ Bioinformatics Institute 30 Medical Drive, Level 1, IMCB Building Singapore 117609 Republic of Singapore Gene finding (LSM5191) p.1
SALSA MLPA KIT P060-B2 SMA
 SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
It is widely believed that cancer cells are derived from normal
 Top-down morphogenesis of colorectal tumors Ie-Ming Shih*, Tian-Li Wang*, Giovanni Traverso*, Kathy Romans*, Stanley R. Hamilton, Shmuel Ben-Sasson, Kenneth W. Kinzler*, and Bert Vogelstein* *The Howard
Top-down morphogenesis of colorectal tumors Ie-Ming Shih*, Tian-Li Wang*, Giovanni Traverso*, Kathy Romans*, Stanley R. Hamilton, Shmuel Ben-Sasson, Kenneth W. Kinzler*, and Bert Vogelstein* *The Howard
Serrated Polyps and a Classification of Colorectal Cancer
 Serrated Polyps and a Classification of Colorectal Cancer Ian Chandler June 2011 Structure Serrated polyps and cancer Molecular biology The Jass classification The familiar but oversimplified Vogelsteingram
Serrated Polyps and a Classification of Colorectal Cancer Ian Chandler June 2011 Structure Serrated polyps and cancer Molecular biology The Jass classification The familiar but oversimplified Vogelsteingram
The Whys OAP Annual Meeting CCO Symposium September 20. Immunohistochemical Assessment Dr. Terence Colgan Mount Sinai Hospital, Toronto
 Immunohistochemical Assessment of Mismatch Repair Proteins in Endometrial Cancer: The Whys and How Terence J. Colgan, MD Head of Gynaecological Pathology, Mount Sinai Hospital, University of Toronto, Toronto.
Immunohistochemical Assessment of Mismatch Repair Proteins in Endometrial Cancer: The Whys and How Terence J. Colgan, MD Head of Gynaecological Pathology, Mount Sinai Hospital, University of Toronto, Toronto.
Biallelic inactivation of hmlh1 by epigenetic gene silencing, a novel mechanism causing human MSI cancers
 Proc. Natl. Acad. Sci. USA Vol. 95, pp. 8698 8702, July 1998 Genetics Biallelic inactivation of hmlh1 by epigenetic gene silencing, a novel mechanism causing human MSI cancers MARTINA L. VEIGL*, LAKSHMI
Proc. Natl. Acad. Sci. USA Vol. 95, pp. 8698 8702, July 1998 Genetics Biallelic inactivation of hmlh1 by epigenetic gene silencing, a novel mechanism causing human MSI cancers MARTINA L. VEIGL*, LAKSHMI
