Chromatin Structure & Gene activity part 2
|
|
|
- Roy Taylor
- 5 years ago
- Views:
Transcription
1 Chromatin Structure & Gene activity part 2
2 Figure Make sure that you understand it. It is good practice for identifying the purpose for various controls.
3 Chromatin remodeling Acetylation helps to make DNA more accessible but by itself it is not enough. Also need additional chromatin remodeling. What is it? Don t really know all of the types that fall under this heading. 1. movement of nucleosomes away 2. other ways to loosen their grip on the DNA. 3. changing the nucleosome so that other proteins can move it or loosen it. 4. changing the nucleosome so that other proteins will RECOGNIZE it. pg 410
4 Chromatin remodeling List of protein complex families SWI/SNF - SwitchSniff ISWI - limitation switch NuRD INO80 Produce nucleosome free regions around enhancers & promoters pg 410
5 human interferon-beta gene (IFN-ß) Dimitris Thanos human interferon-beta gene (IFN-ß) viral infection activates it. TF activators bind nucleosome free areas near promoter generating an enhanceosome.
6 Remodel IFN-ß overview Enhanceosome HATs N N N HATs acetylate N N N N ac ac ac ac N N N N represents enhanceosome Acetylated histones help attract CBP-RNAPII Holoenzyme. N CBP RNAPII Gg g TFIID N N N N ac ac ac ac pg 411 CBP has bromodomains and uses them to recognize acetylated histones.
7 Remodel IFN-ß overview SWI/SNF complex remodels nucleosome over the promoter, loosening it. CBP RNAPII Gg g TFIID ac ac g Gg NN N ac ac ac N ac ac ac TFIID binds to the TATA box and bends it, nucleosome moves 36 bp downstream. Transcript will now start. II AP RN IID TF CBP N N N ac ac ac ac ac N ac ac ac ac
8 Way too simple K8 acetylation of H4 recruits SWI/SNF complex K9 & K14 acetylation of H3 causes recruitment of TFIID
9 Histone code hypothesis/model Kinase binds enhanceosome & phosphoryaltes H3 S10. NOW GCN 5 can acetylate H3 K14. enhanceosome enhanceosome recruits GCN5 The GCN5 HAT acetylates H4 K8 & K3 K9 Histone code is complete & now proteins use it. Acetylated H4K8 is recognized by SWI/SNF. SWI/SNF remodels nucleosome. TFIID is attracted by acetylated H3 K9 and H3 K14. Nucleosome can now lets TFIID have access to the TATA box. which it binds. Then it bends the DNA and the nucleosome moves. Could figure this out by changing K s to A s and doing ChromIP.
10 How? The ChIP assay chromatin immunoprecipitation assay Allows one to determine which proteins and molecules are present on a transcriptional control region. Allows one to detect changes in abundance of proteins and molecules on a transcriptional control region.
11 *
12 Agalioti et al 2002 Laboratory of Dimitris Thanos Fig th ed. Pattern is not random It has been postulated that phosphorylation of S10 is required for K14 acetylation. Time course supports this. TFIID binds after K14 is acetylated Time course of Sendai virus infection of HeLa cells. From the lab of Dimitris Thanos
13 Histone code hypothesis/model Kinase binds enhanceosome & phosphoryaltes H3 S10. NOW GCN 5 can acetylate H3 K14. enhanceosome enhanceosome recruits GCN5 The GCN5 HAT acetylates H4 K8 & K3 K9 Histone code is complete & now proteins use it. Acetylated H4K8 is recognized by SWI/SNF. SWI/SNF remodels nucleosome. TFIID is attracted by acetylated H3 K9 and H3 K14. Nucleosome can now lets TFIID have access to the TATA box. which it binds. Then it bends the DNA and the nucleosome moves. Could figure this out by changing K s to A s and doing ChromIP.
14 Histone code hypothesis/model Proteins associated with the enhancer modify the histones so that they are recognized by the General Transcription factor machinery.
15 Silencing Deacetylases can cause the condensation of chromatin. Condensation is the compression or packing that comes from many interactions between adjacent nucleosomes. Huge stretches are found in telomeres and centromeres. pg 416
16 c HAT M H3 M HDM M K4 Epigenesis M M Acetylation (activating) Deacetylation Deation Methylation (activating) M K9 A S10 P HDAC HMT K14 A K18 K23 A PK HDM A M K27 Phosphorylation (activating) S28 P M Deation Methylation (repressing) Dephosphorylation PP M K36 M HMT M K79 Histone tail basal transcript transcription; o heterochromat silenced (botto continuum of s (top right); repr Enrichment of and atio binding of tran or repressors (R transcriptional suggests that in subject to reac remains uncert modifications o ation an residues. H3 ph phosphorylatio is catalysed by reversed by his ation (w is catalysed by reversed by his phosphorylatio reversed by pro been identified residue. Panels Rev. Neurosci. R Tsankova, N, Renthal, W, Kumar, A, Nestler, EJ (2007) Epigenetic regulation in psychiatric disorders. Nat Rev Neurosci, 8: MAY 2007 VOLUME 8
17 Histone ation Drosophila Repressed lys4 lys9 H3 lys20 H4 Repressors HP1 & polycomb can bind. Stimulated lys4 lys9 lys20 H3 H4 Now can be bound by the activator Brahma but not by the repressors HP1 and polycomb. Interpreted to mean that the proteins are interpreting a code. Methylated K9 has a different effect when it is with ated K4 and ated K20. pg 418.
18 Histone ation Drosophila pg 392 4th ed. Repressed lys4 lys9 H3 lys20 H4 Repressors HP1 & polycomb can bind. Stimulated lys4 lys9 lys20 H3 H4 Now can be bound by the activator Brahma but not by the repressors HP1 and polycomb. Interpreted to mean that the proteins are interpreting a code. Methylated K9 has a different effect when it is with ated K4 and ated K20. Interpreted to mean that the proteins are interpreting a code. Methylated K9 has a different effect when it is with ated K4 and ated K20. pg 418.
19 Modifications in one histone can affect (allow or deny) a modification in an another histone.
20 Histone ation Drosophila Repressed lys4 lys9 H3 lys20 H4 Repressors HP1 & polycomb can bind. Stimulated lys4 lys9 lys20 H3 H4 Now can be bound by the activator Brahma but not by the repressors HP1 and polycomb. Interpreted to mean that the proteins are interpreting a code. Methylated K9 has a different effect when it is with ated K4 and ated K20. Interpreted to mean that the proteins are interpreting a code. Methylated K9 has a different effect when it is with ated K4 and ated K20. pg 418.
21 How many characters are there in this code? More than 100 modifications have been detected by NMR. Many scientists find this to be disquieting because you can make a huge number of unique patterns (code words) with a 100 character alphabet.
22
23 Figure Once transcription begins do all of the nucleosomes have to be bumped off?
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Lecture 10. Eukaryotic gene regulation: chromatin remodelling
 Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Lecture 10 Eukaryotic gene regulation: chromatin remodelling Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation sequences Transcriptional
Gene Expression DNA RNA. Protein. Metabolites, stress, environment
 Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Transcription and chromatin. General Transcription Factors + Promoter-specific factors + Co-activators
 Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
Transcription and chromatin General Transcription Factors + Promoter-specific factors + Co-activators Cofactor or Coactivator 1. work with DNA specific transcription factors to make them more effective
Lecture 8. Eukaryotic gene regulation: post translational modifications of histones
 Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Lecture 8 Eukaryotic gene regulation: post translational modifications of histones Recap.. Eukaryotic RNA polymerases Core promoter elements General transcription factors Enhancers and upstream activation
Chromatin Remodeling: A Novel Mechanism. of Psychotropic Drug Action
 Molecular Pharmacology This article has Fast not been Forward. copyedited Published and formatted. The on final May version 25, 2006 may differ as doi:10.1124/mol.106.027078 from this version. Chromatin
Molecular Pharmacology This article has Fast not been Forward. copyedited Published and formatted. The on final May version 25, 2006 may differ as doi:10.1124/mol.106.027078 from this version. Chromatin
Epigenetics. Lyle Armstrong. UJ Taylor & Francis Group. f'ci Garland Science NEW YORK AND LONDON
 ... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
... Epigenetics Lyle Armstrong f'ci Garland Science UJ Taylor & Francis Group NEW YORK AND LONDON Contents CHAPTER 1 INTRODUCTION TO 3.2 CHROMATIN ARCHITECTURE 21 THE STUDY OF EPIGENETICS 1.1 THE CORE
Host cell activation
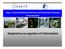 Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
Dept. of Internal Medicine/Infectious and Respiratory Diseases Stefan Hippenstiel Epigenetics as regulator of inflammation Host cell activation LPS TLR NOD2 MDP TRAF IKK NF-κB IL-x, TNFα,... Chromatin
Transcriptional and Epigenetic Mechanisms of Addiction
 Transcriptional and Epigenetic Mechanisms of Addiction Eric J. Nestler Mount Sinai School of Medicine New York, NY Dr. Ray Fuller There is every reason to be optimistic that in the future we will find
Transcriptional and Epigenetic Mechanisms of Addiction Eric J. Nestler Mount Sinai School of Medicine New York, NY Dr. Ray Fuller There is every reason to be optimistic that in the future we will find
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Epigenetic Mechanisms
 RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
RCPA Lecture Epigenetic chanisms Jeff Craig Early Life Epigenetics Group, MCRI Dept. of Paediatrics Overview What is epigenetics? Chromatin The epigenetic code What is epigenetics? the interactions of
Epigenetics: The Future of Psychology & Neuroscience. Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1
 Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Epigenetics: The Future of Psychology & Neuroscience Richard E. Brown Psychology Department Dalhousie University Halifax, NS, B3H 4J1 Nature versus Nurture Despite the belief that the Nature vs. Nurture
Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling
 Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling Resources: Latchman Lodish chapter 10, 20 Helmreich, chapter 11 http://www.nursa.org,
Biochemistry 673 Lecture 2 Jason Kahn, UMCP Introduction to steroid hormone receptor (nuclear receptor) signalling Resources: Latchman Lodish chapter 10, 20 Helmreich, chapter 11 http://www.nursa.org,
On the way of revealing coactivator complexes cross talk during transcriptional activation
 DOI 10.1186/s13578-016-0081-y Cell & Bioscience REVIEW Open Access On the way of revealing coactivator complexes cross talk during transcriptional activation Aleksey N. Krasnov *, Marina Yu. Mazina, Julia
DOI 10.1186/s13578-016-0081-y Cell & Bioscience REVIEW Open Access On the way of revealing coactivator complexes cross talk during transcriptional activation Aleksey N. Krasnov *, Marina Yu. Mazina, Julia
Epigenetics Armstrong_Prelims.indd 1 04/11/2013 3:28 pm
 Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Epigenetics Epigenetics Lyle Armstrong vi Online resources Accessible from www.garlandscience.com, the Student and Instructor Resource Websites provide learning and teaching tools created for Epigenetics.
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS
 R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
R. Piazza (MD, PhD), Dept. of Medicine and Surgery, University of Milano-Bicocca EPIGENETICS EPIGENETICS THE STUDY OF CHANGES IN GENE EXPRESSION THAT ARE POTENTIALLY HERITABLE AND THAT DO NOT ENTAIL A
Structure and Function of Fusion Gene Products in. Childhood Acute Leukemia
 Structure and Function of Fusion Gene Products in Childhood Acute Leukemia Chromosomal Translocations Chr. 12 Chr. 21 der(12) der(21) A.T. Look, Science 278 (1997) Distribution Childhood ALL TEL-AML1 t(12;21)
Structure and Function of Fusion Gene Products in Childhood Acute Leukemia Chromosomal Translocations Chr. 12 Chr. 21 der(12) der(21) A.T. Look, Science 278 (1997) Distribution Childhood ALL TEL-AML1 t(12;21)
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc
 Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) FS 2014 P Matthias & RG Clerc P. Matthias, 16 April 2014 DNA methylation basics Acetylation Acetyltransferases/Deacetylases
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Transcription and RNA processing Lecture 7 Biology 3310/4310 Virology Spring 2018 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at the
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD,
 Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
DNA and Histone Methylation in Learning and Memory
 EXPERIMENTAL BIOLOGY ANNUAL MEETING April 2010 DNA and Histone Methylation in Learning and Memory J. David Sweatt Dept of Neurobiology McKnight Brain Institute UAB School of Medicine The Molecular Basis
EXPERIMENTAL BIOLOGY ANNUAL MEETING April 2010 DNA and Histone Methylation in Learning and Memory J. David Sweatt Dept of Neurobiology McKnight Brain Institute UAB School of Medicine The Molecular Basis
Einführung in die Genetik
 Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
Einführung in die Genetik Prof. Dr. Kay Schneitz (EBio Pflanzen) http://plantdev.bio.wzw.tum.de schneitz@wzw.tum.de Prof. Dr. Claus Schwechheimer (PlaSysBiol) http://wzw.tum.de/sysbiol claus.schwechheimer@wzw.tum.de
INTERACTION DRUG BODY
 INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
Stem Cell Epigenetics
 Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Stem Cell Epigenetics Philippe Collas University of Oslo Institute of Basic Medical Sciences Norwegian Center for Stem Cell Research www.collaslab.com Source of stem cells in the body Somatic ( adult )
Deciphering the Transcriptional Histone Acetylation Code for a Human Gene
 Cell, Vol. 111, 381 392, November 1, 2002, Copyright 2002 by Cell Press Deciphering the Transcriptional Histone Acetylation Code for a Human Gene Theodora Agalioti, 1 Guoying Chen, 1 and Dimitris Thanos
Cell, Vol. 111, 381 392, November 1, 2002, Copyright 2002 by Cell Press Deciphering the Transcriptional Histone Acetylation Code for a Human Gene Theodora Agalioti, 1 Guoying Chen, 1 and Dimitris Thanos
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure
 Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Biochemical Determinants Governing Redox Regulated Changes in Gene Expression and Chromatin Structure Frederick E. Domann, Ph.D. Associate Professor of Radiation Oncology The University of Iowa Iowa City,
Are you the way you are because of the
 EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
EPIGENETICS Are you the way you are because of the It s my fault!! Nurture Genes you inherited from your parents? Nature Experiences during your life? Similar DNA Asthma, Autism, TWINS Bipolar Disorders
Where Splicing Joins Chromatin And Transcription. 9/11/2012 Dario Balestra
 Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
Where Splicing Joins Chromatin And Transcription 9/11/2012 Dario Balestra Splicing process overview Splicing process overview Sequence context RNA secondary structure Tissue-specific Proteins Development
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Atsuko Masumi. 1. Introduction. 2. Histone Acetyltransferases (HATs)
 Journal of Biomedicine and Biotechnology Volume 2011, Article ID 640610, 6 pages doi:10.1155/2011/640610 Review Article Histone Acetyltransferases as Regulators of Nonhistone Proteins: The Role of Interferon
Journal of Biomedicine and Biotechnology Volume 2011, Article ID 640610, 6 pages doi:10.1155/2011/640610 Review Article Histone Acetyltransferases as Regulators of Nonhistone Proteins: The Role of Interferon
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3
 Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE
 Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE If looking for the book Systems Analysis of Chromatin-Related Protein Complexes in Cancer in pdf format, then you have come
Systems Analysis Of Chromatin-Related Protein Complexes In Cancer READ ONLINE If looking for the book Systems Analysis of Chromatin-Related Protein Complexes in Cancer in pdf format, then you have come
The Role of Protein Domains in Cell Signaling
 The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
Expert Intelligence for Better Decisions Epigenetics:
 Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Control of nuclear receptor function by local chromatin structure
 MINIREVIEW Control of nuclear receptor function by local chromatin structure Malgorzata Wiench, Tina B. Miranda and Gordon L. Hager Laboratory of Receptor Biology and Gene Expression, National Cancer Institute,
MINIREVIEW Control of nuclear receptor function by local chromatin structure Malgorzata Wiench, Tina B. Miranda and Gordon L. Hager Laboratory of Receptor Biology and Gene Expression, National Cancer Institute,
6. Heterochromatin allows for silencing of large domains of the genome and appears to utilize a combination of mechanisms to do so.
 Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 18, 2014 Chromatin Structure and Its Regulation 2 Key Points 1. There are at least two ways in which arrays of chromatin
Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 18, 2014 Chromatin Structure and Its Regulation 2 Key Points 1. There are at least two ways in which arrays of chromatin
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
Overview: Conducting the Genetic Orchestra Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development
EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
 EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The EpiQuik Circulating Acetyl Histone H3K18 ELISA Kit (Colorimetric)
Tat inhibition by didehydro Cortistatin A promotes heterochromatin formation at the HIV 1 long terminal repeat
 https://doi.org/10.1186/s13072-019-0267-8 Epigenetics & Chromatin RESEARCH Open Access Tat inhibition by didehydro Cortistatin A promotes heterochromatin formation at the HIV 1 long terminal repeat Chuan
https://doi.org/10.1186/s13072-019-0267-8 Epigenetics & Chromatin RESEARCH Open Access Tat inhibition by didehydro Cortistatin A promotes heterochromatin formation at the HIV 1 long terminal repeat Chuan
5. Heterochromatin allows for silencing of large domains of the genome and utilizes a combination of mechanisms to do so.
 Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 1, 2016 Chromatin Structure and Its Regulation 2 Key Points 1. Histones provide a versatile regulatory platform through
Biochemistry 201 Biological Regulatory Mechanisms Lecturer: Geeta Narlikar February 1, 2016 Chromatin Structure and Its Regulation 2 Key Points 1. Histones provide a versatile regulatory platform through
9 th TRR81 PhD Minisymposium Kinases as regulators of chromatin structure and transcription
 9 th TRR81 PhD Minisymposium Kinases as regulators of chromatin structure and transcription Friday, 20 th of November 2015, 13:00h Institute of Molecular Biology and Tumor Research (IMT) Philipps-University
9 th TRR81 PhD Minisymposium Kinases as regulators of chromatin structure and transcription Friday, 20 th of November 2015, 13:00h Institute of Molecular Biology and Tumor Research (IMT) Philipps-University
Removal of Shelterin Reveals the Telomere End-Protection Problem
 Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
HDAC1 Inhibitor Screening Assay Kit
 HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Repression of IP-10 by Interactions between Histone Deacetylation and Hypermethylation in Idiopathic Pulmonary Fibrosis
 MOLECULAR AND CELLULAR BIOLOGY, June 2010, p. 2874 2886 Vol. 30, No. 12 0270-7306/10/$12.00 doi:10.1128/mcb.01527-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Repression of
MOLECULAR AND CELLULAR BIOLOGY, June 2010, p. 2874 2886 Vol. 30, No. 12 0270-7306/10/$12.00 doi:10.1128/mcb.01527-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Repression of
Constitutive heterochromatin formation and transcription in mammals
 Saksouk et al. Epigenetics & Chromatin 2015, 8:3 REVIEW Constitutive heterochromatin formation and transcription in mammals Nehmé Saksouk, Elisabeth Simboeck and Jérôme Déjardin * Open Access Abstract
Saksouk et al. Epigenetics & Chromatin 2015, 8:3 REVIEW Constitutive heterochromatin formation and transcription in mammals Nehmé Saksouk, Elisabeth Simboeck and Jérôme Déjardin * Open Access Abstract
Histone octamer assembly
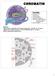 CHROMATIN Chromatin Histone structure Nucleosome Chromatin fiber Histone acetylation Chromatin transcription Regulation of gene expression: -Cell cycle control (+/-) -Cell differentiation Genome integrity
CHROMATIN Chromatin Histone structure Nucleosome Chromatin fiber Histone acetylation Chromatin transcription Regulation of gene expression: -Cell cycle control (+/-) -Cell differentiation Genome integrity
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
doi:10.1038/nature23267 Discussion Our findings reveal unique roles for the methylation states of histone H3K9 in RNAi-dependent and - independent heterochromatin formation. Clr4 is the sole S. pombe enzyme
TRANSCRIPTION FACTOR EFFECTS
 Department of Cell- and Molecular Biology Karolinska Institutet TRANSCRIPTION FACTOR EFFECTS ON CHROMATIN ORGANISATION AND GENE REGULATION Per-Henrik Holmqvist Stockholm 2005 Per-Henrik Holmqvist 2005
Department of Cell- and Molecular Biology Karolinska Institutet TRANSCRIPTION FACTOR EFFECTS ON CHROMATIN ORGANISATION AND GENE REGULATION Per-Henrik Holmqvist Stockholm 2005 Per-Henrik Holmqvist 2005
Protein methylation CH 3
 Protein methylation CH 3 methionine S-adenosylmethionine (SAM or adomet) Methyl group of the methionine is activated by the + charge of the adjacent sulfur atom. SAM S-adenosylhomocysteine homocysteine
Protein methylation CH 3 methionine S-adenosylmethionine (SAM or adomet) Methyl group of the methionine is activated by the + charge of the adjacent sulfur atom. SAM S-adenosylhomocysteine homocysteine
Epigenetics DNA methylation. Biosciences 741: Genomics Fall, 2013 Week 13. DNA Methylation
 Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Epigenetics DNA methylation Biosciences 741: Genomics Fall, 2013 Week 13 DNA Methylation Most methylated cytosines are found in the dinucleotide sequence CG, denoted mcpg. The restriction enzyme HpaII
Epigenetic reprogramming of tumor and stem cell genomes by exogenous delivery of Transcription Factors
 Pilar Blancafort Associate Professor School of Anatomy, Physiology and Human Biology UWA! Epigenetic reprogramming of tumor and stem cell genomes by exogenous delivery of Transcription Factors The University
Pilar Blancafort Associate Professor School of Anatomy, Physiology and Human Biology UWA! Epigenetic reprogramming of tumor and stem cell genomes by exogenous delivery of Transcription Factors The University
Post-translational modifications of proteins in gene regulation under hypoxic conditions
 203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
Epigenetics and diseases. Genetical analysis Irma takács
 Epigenetics and diseases Genetical analysis Irma takács 30.11.2010. Epigenetic diseases Introduction Mechanism Diseases Miracle: Identical DNA different cells EPIGENETICS Epigenetics C.H. Waddington in
Epigenetics and diseases Genetical analysis Irma takács 30.11.2010. Epigenetic diseases Introduction Mechanism Diseases Miracle: Identical DNA different cells EPIGENETICS Epigenetics C.H. Waddington in
Biochimica et Biophysica Acta
 BBAGRM-00286; No. of pages: 19; 4C: 8, 12 Biochimica et Biophysica Acta xxx (2010) xxx xxx Contents lists available at ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbagrm
BBAGRM-00286; No. of pages: 19; 4C: 8, 12 Biochimica et Biophysica Acta xxx (2010) xxx xxx Contents lists available at ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbagrm
Chromatin-Based Regulation of Gene Expression
 Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Chromatin-Based Regulation of Gene Expression.George J. Quellhorst, Jr., PhD.Associate Director, R&D.Biological Content Development Topics to be Discussed Importance of Chromatin-Based Regulation Mechanism
Prokaryotes and eukaryotes alter gene expression in response to their changing environment
 Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
Chapter 18 Prokaryotes and eukaryotes alter gene expression in response to their changing environment In multicellular eukaryotes, gene expression regulates development and is responsible for differences
REVIEW. Andrew J Bannister 1, Tony Kouzarides 1. Introduction. Histone acetylation. npg. such as repair, replication and recombination.
 REVIEW Cell Research (2011) 21:381-395. 2011 IBCB, SIBS, CAS All rights reserved 1001-0602/11 $ 32.00 www.nature.com/cr npg Regulation of chromatin by histone modifications Andrew J Bannister 1, Tony Kouzarides
REVIEW Cell Research (2011) 21:381-395. 2011 IBCB, SIBS, CAS All rights reserved 1001-0602/11 $ 32.00 www.nature.com/cr npg Regulation of chromatin by histone modifications Andrew J Bannister 1, Tony Kouzarides
Stacie Bulfer Proteins Research Proposal April 27, Background
 Background Stacie Bulfer Proteins Research Proposal April 27, 2003 DNA Packaging in Eukaryotes Eukaryotes have five histones: H1, H2A, H2B, H3 and H4 which form repeating units that compact DNA. Histones
Background Stacie Bulfer Proteins Research Proposal April 27, 2003 DNA Packaging in Eukaryotes Eukaryotes have five histones: H1, H2A, H2B, H3 and H4 which form repeating units that compact DNA. Histones
Epigenetics q&more 01.11
 Laurie. Knight, istockphoto.com Epigenetics 6 Bookmarks About the reading of genes in the Book of Life Prof. Dr. Manfred Jung, Julia M. Wagner, Institute for Pharmaceutical Sciences, Albert-Ludwig-University
Laurie. Knight, istockphoto.com Epigenetics 6 Bookmarks About the reading of genes in the Book of Life Prof. Dr. Manfred Jung, Julia M. Wagner, Institute for Pharmaceutical Sciences, Albert-Ludwig-University
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Oncology 520 Epigene0cs and Cancer. February 16, 2011
 Oncology 520 Epigene0cs and Cancer February 16, 2011 Outline What is epigene0cs? What are examples of epigene0c processes? What is the epigenome and how is it decoded? Are epigene0c mechanisms important
Oncology 520 Epigene0cs and Cancer February 16, 2011 Outline What is epigene0cs? What are examples of epigene0c processes? What is the epigenome and how is it decoded? Are epigene0c mechanisms important
Structure and function of active chromatin and DNase I hypersensitive sites
 MINIREVIEW Structure and function of active chromatin and DNase I hypersensitive sites Peter N. Cockerill Experimental Haematology, Leeds Institute of Molecular Medicine, University of Leeds, UK Keywords
MINIREVIEW Structure and function of active chromatin and DNase I hypersensitive sites Peter N. Cockerill Experimental Haematology, Leeds Institute of Molecular Medicine, University of Leeds, UK Keywords
Removal of Shelterin Reveals the Telomere End-Protection Problem
 Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
Removal of Shelterin Reveals the Telomere End-Protection Problem DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to
DSB. Double-Strand Breaks causate da radiazioni stress ossidativo farmaci
 DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to the detection and repair of DNA damage DSBs induce a local decrease
DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci DSB e CROMATINA Higher-order chromatin packaging is a barrier to the detection and repair of DNA damage DSBs induce a local decrease
DSB. Double-Strand Breaks causate da radiazioni stress ossidativo farmaci
 DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci METODI DDR foci formation in irradiated (2 Gy) cells fixed 2 h later IRIF IRradiation Induced Focus Laser micro-irradiation DDR
DSB Double-Strand Breaks causate da radiazioni stress ossidativo farmaci METODI DDR foci formation in irradiated (2 Gy) cells fixed 2 h later IRIF IRradiation Induced Focus Laser micro-irradiation DDR
Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells
 Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells Immunofluorescence staining of H3K9me3 and 53BP1 in PH and HGADFN003 (HG003) cells at 24 h after γ-irradiation. Scale bar,
Supplementary Figure S1: Defective heterochromatin repair in HGPS progeroid cells Immunofluorescence staining of H3K9me3 and 53BP1 in PH and HGADFN003 (HG003) cells at 24 h after γ-irradiation. Scale bar,
Hox genes. Discovered in Drosophila in 1923 by Bridges and Morgan Antennapaedia complex ANT-C Bithorax complex BX-C
 Hox genes Establish body plan during development Specify head to tail axis of animal embryos Head Hox genes, abdomen hox genes. Mutations can cause one body part to transform to another 39 transcription
Hox genes Establish body plan during development Specify head to tail axis of animal embryos Head Hox genes, abdomen hox genes. Mutations can cause one body part to transform to another 39 transcription
Transcription and RNA processing
 Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
Transcription and RNA processing Lecture 7 Biology W3310/4310 Virology Spring 2016 It is possible that Nature invented DNA for the purpose of achieving regulation at the transcriptional rather than at
Eukaryotic Gene Regulation
 Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Eukaryotic Gene Regulation Chapter 19: Control of Eukaryotic Genome The BIG Questions How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different,
Cancer and Gene Regulation
 OpenStax-CNX module: m44548 1 Cancer and Gene Regulation OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44548 1 Cancer and Gene Regulation OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder Labs August 30, 2012
 Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
PhD THESIS Epigenetic mechanisms involved in stem cell differentiation
 Romanian Academy Institute of Cellular Biology and Pathology "Nicolae Simionescu" PhD THESIS Epigenetic mechanisms involved in stem cell differentiation Coordinator: Acad. Maya Simionescu PhD Student:
Romanian Academy Institute of Cellular Biology and Pathology "Nicolae Simionescu" PhD THESIS Epigenetic mechanisms involved in stem cell differentiation Coordinator: Acad. Maya Simionescu PhD Student:
BIO360 Fall 2013 Quiz 1
 BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
BIO360 Fall 2013 Quiz 1 1. Examine the diagram below. There are two homologous copies of chromosome one and the allele of YFG carried on the light gray chromosome has undergone a loss-of-function mutation.
The role of epigenetics in breast. cancer progression
 The role of epigenetics in breast cancer progression Aslı Dilber Yıldırım January 2013 Master thesis, Cancer Genomics & Developmental Biology Under daily supervision of Rick van Nuland, MSc Examiners:
The role of epigenetics in breast cancer progression Aslı Dilber Yıldırım January 2013 Master thesis, Cancer Genomics & Developmental Biology Under daily supervision of Rick van Nuland, MSc Examiners:
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN
 Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
Epigenetic Principles and Mechanisms Underlying Nervous System Function in Health and Disease Mark F. Mehler MD, FAAN Institute for Brain Disorders and Neural Regeneration F.M. Kirby Program in Neural
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
H19 Chromatin Insulator in Development and DiseaseEpigenetic Regulation of the
 Comprehensive Summaries of Uppsala Dissertations from the Faculty of Science and Technology 825 H19 Chromatin Insulator in Development and DiseaseEpigenetic Regulation of the BY CLAES HOLMGREN ACTA UNIVERSITATIS
Comprehensive Summaries of Uppsala Dissertations from the Faculty of Science and Technology 825 H19 Chromatin Insulator in Development and DiseaseEpigenetic Regulation of the BY CLAES HOLMGREN ACTA UNIVERSITATIS
Gene Regulation. Bacteria. Chapter 18: Regulation of Gene Expression
 Chapter 18: Regulation of Gene Expression A Biology 2013 1 Gene Regulation rokaryotes and eukaryotes alter their gene expression in response to their changing environment In multicellular eukaryotes, gene
Chapter 18: Regulation of Gene Expression A Biology 2013 1 Gene Regulation rokaryotes and eukaryotes alter their gene expression in response to their changing environment In multicellular eukaryotes, gene
Identifying the Combinatorial Effects of Histone Modifications by Association Rule Mining in Yeast
 Evolutionary Bioinformatics Original Research Open Access Full open access to this and thousands of other papers at http://www.la-press.com. Identifying the Combinatorial Effects of Histone Modifications
Evolutionary Bioinformatics Original Research Open Access Full open access to this and thousands of other papers at http://www.la-press.com. Identifying the Combinatorial Effects of Histone Modifications
DNA Methylation and Demethylation as Targets for Anticancer Therapy
 Biochemistry (Moscow), Vol. 70, No. 5, 2005, pp. 533-549. Translated from Biokhimiya, Vol. 70, No. 5, 2005, pp. 651-669. Original Russian Text Copyright 2005 by Szyf. DNA Methylation and Demethylation
Biochemistry (Moscow), Vol. 70, No. 5, 2005, pp. 533-549. Translated from Biokhimiya, Vol. 70, No. 5, 2005, pp. 651-669. Original Russian Text Copyright 2005 by Szyf. DNA Methylation and Demethylation
The molecular mechanisms of latency and reactivation in CD4 + T cells during HIV infection Suzanna Huppelschoten
 The molecular mechanisms of latency and reactivation in CD4 + T cells during HIV infection Suzanna Huppelschoten March, 2011 Masterthesis Infection and Immunity Written under supervision of L.M van den
The molecular mechanisms of latency and reactivation in CD4 + T cells during HIV infection Suzanna Huppelschoten March, 2011 Masterthesis Infection and Immunity Written under supervision of L.M van den
Genetics and Genomics in Medicine Chapter 6 Questions
 Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Genetics and Genomics in Medicine Chapter 6 Questions Multiple Choice Questions Question 6.1 With respect to the interconversion between open and condensed chromatin shown below: Which of the directions
Epigenetics, brain evolution and behaviour
 Available online at www.sciencedirect.com Frontiers in Neuroendocrinology xxx (2008) xxx xxx Review Epigenetics, brain evolution and behaviour Eric B. Keverne a, *, James P. Curley b a Sub-Department of
Available online at www.sciencedirect.com Frontiers in Neuroendocrinology xxx (2008) xxx xxx Review Epigenetics, brain evolution and behaviour Eric B. Keverne a, *, James P. Curley b a Sub-Department of
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 18 Regulation of Gene Expression PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 2. Aims & Objectives
 2.1. Statement of the problem: Earlier reports have shown ambiguous alteration of histone marks in response to DNA damage in asynchronized population of cells. These histone marks not only undergo dynamic
2.1. Statement of the problem: Earlier reports have shown ambiguous alteration of histone marks in response to DNA damage in asynchronized population of cells. These histone marks not only undergo dynamic
Dominic J Smiraglia, PhD Department of Cancer Genetics. DNA methylation in prostate cancer
 Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Chromatin Regulation and the Histone Code in HIV Latency
 YALE JOURNAL OF BIOLOGY AND MEDICINE 90 (2017), pp.229-243. Review Chromatin Regulation and the Histone Code in HIV Latency Anne-Marie W. Turner a,b and David M. Margolis a,b,c,d,* a UNC HIV Cure Center,
YALE JOURNAL OF BIOLOGY AND MEDICINE 90 (2017), pp.229-243. Review Chromatin Regulation and the Histone Code in HIV Latency Anne-Marie W. Turner a,b and David M. Margolis a,b,c,d,* a UNC HIV Cure Center,
Transcriptional and post-transcriptional regulation of telomerase reverse transcriptase (htert) expression. Zheng Ge 葛峥
 Division of Hematology, Department of Medicine Karolinska Institutet, Stockholm, Sweden Transcriptional and post-transcriptional regulation of telomerase reverse transcriptase (htert) expression The role
Division of Hematology, Department of Medicine Karolinska Institutet, Stockholm, Sweden Transcriptional and post-transcriptional regulation of telomerase reverse transcriptase (htert) expression The role
Brd4 Coactivates Transcriptional Activation of NF- B via Specific Binding to Acetylated RelA
 MOLECULAR AND CELLULAR BIOLOGY, Mar. 2009, p. 1375 1387 Vol. 29, No. 5 0270-7306/09/$08.00 0 doi:10.1128/mcb.01365-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Brd4 Coactivates
MOLECULAR AND CELLULAR BIOLOGY, Mar. 2009, p. 1375 1387 Vol. 29, No. 5 0270-7306/09/$08.00 0 doi:10.1128/mcb.01365-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Brd4 Coactivates
Transcription Through Chromatin - Dynamic Organization of Genes
 GENERAL ARTCLE Transcription Through Chromatin - Dynamic Organization of Genes Hari Kishore A and Tapas K Kundu Tapas K Kundu is Assistant Professor at the Jawaharlal Nehru Centre for Advanced Scientific
GENERAL ARTCLE Transcription Through Chromatin - Dynamic Organization of Genes Hari Kishore A and Tapas K Kundu Tapas K Kundu is Assistant Professor at the Jawaharlal Nehru Centre for Advanced Scientific
Supplemental Figure 1: Asymmetric chromatin maturation leads to epigenetic asymmetries on sister chromatids.
 Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
Supplemental Material: Annu. Rev. Cell Dev. Biol. 2017. 33:291 318 https://doi.org/10.1146/annurev-cellbio-100616-060447 The Inherent Asymmetry of DNA Replication Snedeker, Wooten, and Chen Supplemental
Thesis for doctoral degree (Ph.D.) 2009 Characterization of Gcn5 Histone acetyltransferase in Schizosaccharomyces pombe Anna Johnsson
 Thesis for doctoral degree (Ph.D.) 2009 Thesis for doctoral degree (Ph.D.) 2009 Characterization of Gcn5 Histone acetyltransferase in Schizosaccharomyces pombe Anna Johnsson Characterization of Gcn5 Histone
Thesis for doctoral degree (Ph.D.) 2009 Thesis for doctoral degree (Ph.D.) 2009 Characterization of Gcn5 Histone acetyltransferase in Schizosaccharomyces pombe Anna Johnsson Characterization of Gcn5 Histone
Molecular Cell Biology. Prof. D. Karunagaran. Department of Biotechnology. Indian Institute of Technology Madras
 Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module-9 Molecular Basis of Cancer, Oncogenes and Tumor Suppressor Genes Lecture 6 Epigenetics
Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module-9 Molecular Basis of Cancer, Oncogenes and Tumor Suppressor Genes Lecture 6 Epigenetics
Epigenomics. Ivana de la Serna Block Health Science
 Epigenomics Ivana de la Serna Block Health Science 388 383-4111 ivana.delaserna@utoledo.edu Outline 1. Epigenetics-definition and overview 2. DNA methylation/hydroxymethylation 3. Histone modifications
Epigenomics Ivana de la Serna Block Health Science 388 383-4111 ivana.delaserna@utoledo.edu Outline 1. Epigenetics-definition and overview 2. DNA methylation/hydroxymethylation 3. Histone modifications
Transcriptional and epigenetic regulation of T-helper lineage specification
 Subhash K. Tripathi Riitta Lahesmaa Transcriptional and epigenetic regulation of T-helper lineage specification Authors addresses Subhash K. Tripathi 1,2,3, Riitta Lahesmaa 1 1 Turku Centre for Biotechnology,
Subhash K. Tripathi Riitta Lahesmaa Transcriptional and epigenetic regulation of T-helper lineage specification Authors addresses Subhash K. Tripathi 1,2,3, Riitta Lahesmaa 1 1 Turku Centre for Biotechnology,
