Insights into the mechanism and inhibition of fatty acid amide hydrolase from quantum mechanics/molecular mechanics (QM/MM) modelling
|
|
|
- Edwin Holmes
- 6 years ago
- Views:
Transcription
1 Enzyme Mechanisms: Fast Reaction and Computational Approaches 363 Insights into the mechanism and inhibition of fatty acid amide hydrolase from quantum mechanics/molecular mechanics (QM/MM) modelling Alessio Lodola*, Marco Mor*, Jitnapa Sirirak and Adrian J. Mulholland 1 *Dipartimento Farmaceutico, Università degli Studi di Parma, viale G.P. Usberti 27/A Campus Universitario, I Parma, Italy, and Centre for Computational Chemistry, School of Chemistry, University of Bristol, Bristol BS8 1TS, U.K. Abstract FAAH (fatty acid amide hydrolase) is a promising target for the treatment of several central nervous system and peripheral disorders. Combined QM/MM (quantum mechanics/molecular mechanics) calculations have elucidated the role of its unusual catalytic triad in the hydrolysis of oleamide and oleoylmethyl ester substrates, and have identified the productive inhibitor-binding orientation for the carbamoylating compound URB524. These are potentially crucial insights for designing new covalent inhibitors of this drug target. Introduction Computational modelling [e.g. with combined QM/MM (quantum mechanics/molecular mechanics) methods] is increasingly making practical contributions to our understanding of enzyme-catalysed reaction mechanisms [1]. QM (quantum chemical electronic structure) methods allow the modelling of chemical reactions, and are potentially highly accurate for small molecules. However, electronic structure calculations require large computational resources, which significantly limits the size of the system that can be treated, making calculations on whole enzymes (composed of thousands of atoms) prohibitive. Calculations on enzymecatalysed reactions are possible with QM/MM methods: they link QM and MM representations of different parts of a system [2]. The combination of these approaches contains the fundamentals necessary to describe the PESs (potential energy surfaces) relevant to enzymatic chemistry, at least to a first approximation [3]. In the QM/MM framework, the reactive region of the enzyme active site is described by a QM technique (e.g. semi-empirical, density functional theory or ab initio molecular orbital methods); the QM region should be of sufficient size to include crucial electronic structure effects in the reaction [4,5]. Semi-empirical molecular orbital techniques such as AM1 and PM3, although not suitable for all systems, allow large QM regions to be treated but often overestimate reaction barriers [6]. Reliable calculations have increasingly been made possible by the development of QM/MM methods based on density functional theory and Key words: fatty acid amide hydrolase (FAAH), inhibitor design, quantum mechanics/molecular mechanics (QM/MM), reaction mechanism, tetrahedral intermediate, transition state. Abbreviations used: FAAH, fatty acid amide hydrolase; OA, oleamide; OME, oleoylmethyl ester; PES, potential energy surface; QM/MM, quantum mechanics/molecular mechanics; TI, tetrahedral intermediate; TS, transition state. 1 To whom correspondence should be addressed ( Adrian.Mulholland@bristol.ac.uk). on ab initio electron correlation approaches such as the MP2 perturbation method and coupled-cluster theory [7,8]. Many important drug targets are enzymes. Ligand design could significantly benefit from detailed, molecularlevel knowledge of their reaction mechanisms. QM/MM modelling has the potential to show in atomic detail how TSs (transition states) and intermediates are stabilized within enzymes. Increasingly, mechanistic simulations will provide important insights, of the type described here, for medicinal chemists and will thus help in the design of new inhibitors and the development of predictive models of drug metabolism [9]. FAAH (fatty acid amide hydrolase): a case study for QM/MM reaction modelling FAAH is a key enzyme in endocannabinoid metabolism [10], and is a promising target for treatment of nervous system disorders such as anxiety, pain and depression [11]. FAAH is a member of the amidase signature family endowed with alys 142 Ser 217 Ser 241 catalytic triad [12]. FAAH shows a remarkable ability to hydrolyse amides and esters at similar rates, by a mechanism in which acylation is rate limiting [13]. Considerable experimental effort [14] has been invested into elucidating the detailed mechanism of hydrolysis by FAAH. Lys 142 appears to serve as a key acid and base in distinct steps of the catalytic cycle. As a base, Lys 142 activates the Ser 241 nucleophile for attack on the substrate amide carbonyl. As an acid, Lys 142 readily protonates the substrate-leaving group leading to its expulsion (Figure 1). The impact of Lys 142 on Ser 241 nucleophile strength and leaving group protonation occurs indirectly, via the bridging Ser 217 of the triad, which acts as a proton shuttle [15]. The three-dimensional structure of FAAH has been determined by X-ray crystallography [16], prompting computational investigations on the interactions of FAAH Biochem. Soc. Trans. (2009) 37, ; doi: /bst
2 364 Biochemical Society Transactions (2009) Volume 37, part 2 Figure 1 Proposed acylation mechanism for FAAH [17] YisNH 2 for OA (the label N am is used for its leaving group nitrogen in the text) and CH 3 OforOME(O es is used as a label for its leaving group oxygen in the text). Figure 2 PM3-CHARMM22 QM/MM potential energy profiles for the hydrolysis of OA (black) [17,21] and OME (grey) in FAAH A G are relevant configurations along the calculated pathways: A is the substrate complex; C is the TI; E is the protonated TI; G is the product; B, D and F are the (approximate) TS structures for the different steps. Y is NH 2 for OA and CH 3 OforOME. with substrates and with inhibitors. We have modelled the mechanism of the first step of the acylation reaction of the substrate OA (oleamide) [17]. An adiabatic mapping approach was applied to scan the PES for the reaction at the PM3-CHARMM22 QM/MM level. Density functional theory calculations (with the B3LYP functional) were also performed, for a better description of the reaction energetics [18]. In our model, the first reaction step of the acylation consists of three events (Figure 1). A proton is abstracted from Ser 241 and, via the bridging residue cis-ser 217, is transferred to the general base Lys 142. The activated Ser 241 attacks the carbonyl group of the OA substrate, leading to the formation of the TI (tetrahedral intermediate). To model these events, two reaction co-ordinates were applied, defined in terms of interatomic distances: R x defined as [d(o 1, H 1 ) d(o 2,H 1 ) d(o 1, C)] including proton abstraction from Ser 241 by Ser 217 and the nucleophilic attack by Ser 241 ; R y, defined as [(do 2,H 2 ] d[(n, H 2 )], describes the proton transfer between Ser 217 and Lys 142. R x and R y were increased respectively in steps of 0.2 and 0.1 Å (1 Å = 0.1 nm) with harmonic restraints of 5000 kcal mol 1 Å 2 (1 kcal = kj) as described in [17]. The PM3-CHARMM22 PES indicated that the deprotonation of the Ser 241 nucleophile is the rate-limiting step of the reaction, with a barrier of 36 kcal mol 1 at this level of theory (Figure 2). As the subsequent nucleophilic attack takes place with no barrier, TI formation and Ser 241 deprotonation are concerted events. The TI (configuration C, Figure 2) was calculated to be less stable than the Michaelis complex (substrate complex, A) by 26 kcal mol 1, indicating its transient character. The TI is significantly stabilized by the enzyme; however, comparison of the PM3-CHARMM22 PES and the in vacuo PM3 PES showed that the highest level of enzyme stabilization along the pathway was reached at TI due to crucial hydrogen bonds involving the charged oxygen of the OA and NH groups of the oxyanion hole residues.
3 Enzyme Mechanisms: Fast Reaction and Computational Approaches 365 Figure 3 Structures of important points along the QM/MM reaction pathway for the acylation reaction with OA in FAAH ([17], see also Figure 2) Left-hand panel: TI (C in Figure 2). Middle panel: protonated TI (E in Figure 2). Right-hand panel: acylenzyme (G in Figure 2). Energy correction at the higher B3LYP/6-31+G(d) level gave a barrier of 18 kcal mol 1, close to the experimentally deduced activation barrier for the overall reaction of OA of 16 kcal mol 1 [13]. The effect of conformational fluctuations on catalysis is a key issue of debate in enzymology [19]. QM/MM calculations based on energy minimization alone, e.g. adiabatic mapping along a reaction co-ordinate (as opposed to simulations incorporating the effects of thermal fluctuations, e.g. molecular dynamics or Monte Carlo simulations [20]), may hide potential pitfalls due to conformational transitions at the protein active site. We have investigated the relation between conformational fluctuations and reactivity for the first step of the acylation reaction between FAAH and OA. PESs were calculated at the PM3-CHARMM22 and B3LYP/6-31+G(d)//PM3-CHARMM22 levels by using several conformations (four) of the enzyme substrate complex. The results showed that geometrical fluctuations of the active site can significantly affect the overall energetic barrier and that important conformational changes occur in the active site in the reaction [21]. Reaction of OA apparently proceeds via a minor conformation of the enzyme, in which the TS is better stabilized. We have also modelled the second step of the FAAH OA acylation reaction at the PM3-CHARMM22 QM/MM level, employing the same type of adiabatic mapping approach. Formation of the acylenzyme from the TI requires several proton transfer processes involving both Lys 142 and Ser 217. Once the leaving group nitrogen is protonated, the breakdown of the amide bond can be completed. Various reaction co-ordinates were tested in QM/MM calculations of PESs for this reaction step, and only one was found to give a smooth, satisfactory shape. This surface was calculated starting from the TI, splitting the modelling of the reaction into two events: (i) protonation of the leaving group and (ii) breaking of C N bond. For step (i), two reaction coordinates were employed, namely R r defined as the difference between atomic distances [d(o 2, H 1 ) d(n am, H 1 )] moving the proton H 1 from Ser 217 to N am of OA, and R s defined as [d(n, H 2 ) d(o 2,H 2 )] moving proton H 2 from Lys 142 back to Ser 217. Both reaction co-ordinates were restrained to a sequence of values with harmonic restraints of 5000 kcal mol 1 Å 2, and increased in steps of 0.1 Å; geometry optimization was performed at each point to achieve an energy gradient of 0.01 kcal mol 1 Å 1. The PM3-CHAR- MM22 PES showed a mechanism where the protonation of the basic nitrogen of OA (N am )isconcertedwiththe transfer of proton H 2 from Lys 142 to Ser 217. However, visual inspection of the structures along the modelled pathway showed a rearrangement of the TI due to the pyramidal inversion of the N am. This event is crucial, because the lone pair on N am needs to be correctly oriented towards the oxygen of Ser 217 to accept the H 1 proton (Figure 3). All these events lead to configuration E (Figure 2) overcoming a barrier of 40 kcal mol 1 (relative to the substrate complex), 3 kcal mol 1 higher than that calculated for the deprotonation of Ser 241. The protonated TI is calculated to be more stable than the TI itself: the energy of E is 12 kcal mol 1 with respect to the substrate complex. Finally, the expulsion of the leaving group (e.g. neutral ammonia in the reaction of OA) was modelled by increasing (in steps of 0.1 Å) the reaction co-ordinate R z,[d(c, N am )]. This reaction led to the formation of the acylenzyme G overcoming a barrier of 16 kcal mol 1 (relative to the substrate complex). The full energy profile for the acylation reaction indicates that formation of the TI is not the rate-limiting step, because the highest overall barrier was found for the protonation of the leaving group. However, the small difference between the calculated barriers (36 compared with 40 kcal mol 1 ) suggests that the acylation depends on both reaction steps, with no step being clearly rate limiting. The general picture of the mechanism of OA hydrolysis in FAAH is similar to that found in similar computational work by Tubert-Brohman et al. [22], who performed free energy perturbation simulations for the reaction at the PDDG/PM3 QM/MM level. It is encouraging that independent simulations by different groups with different QM/MM techniques arrive at the same qualitative mechanistic conclusions, and strengthen these conclusions. A small difference between the results is that Tubert-Brohman et al. [22] found the highest
4 366 Biochemical Society Transactions (2009) Volume 37, part 2 barrier for TI formation (rather then collapse). This difference may be related to the different calculated relative energies of the TI, as the energy cost for the protonation and expulsion of the leaving group (E-F-G pathway in Figure 2) is comparable with both methods ( 15 kcal mol 1 relative to the energy of the TI). In our calculations, the TI is less stable than the substrate complex by 26 kcal mol 1, whereas with the QM/MM/FEP (free energy perturbation) method the TI is predicted to be more stable than the substrate complex by kcal mol 1 [22]. This may be due to the more extensive conformational sampling performed by Tubert-Brohman et al. As noted above, potential energy barriers (as reported here) are dependent on the conformation of the enzyme used for modelling; another important consideration is that the PM3 method overestimates barriers for the reaction. The potential energy profiles described here (e.g. Figure 2) should not be directly compared with experimental values for these reasons. They are, however, useful, for comparison of alternative substrates and conformations. To investigate the experimental observation that FAAH cleaves amides somewhat faster than esters [13], we have also modelled the acylation mechanism for OME (oleoylmethyl ester). The FAAH OME system was prepared by applying the same procedure described for OA [17]. The same PM3- CHARMM22 adiabatic mapping approach was used to explore PESs of the reaction. The PES for the first step of the acylation was explored by applying the same reaction co-ordinates (R x and R y ) and the same harmonic restraints as for OA. Deprotonation of Ser 241 was found to have a barrier of 44 kcal mol 1, relative to the substrate complex A (Figure 2). This event is in concert with the nucleophilic attack and leads to the TI. The subsequent breakdown of the TI was divided into two events, similar to the modelling of the OA reaction: (i) protonation of the alcohol-leaving group and (ii) breaking of the C O bond. For step (i), two reaction co-ordinates were employed, namely R r defined as [d(o 2,H 1 ) d(o es H 1 )] moving the proton H 1 from Ser 217 to O es of OME, and R s defined as [d(n, H 2 ) d(o 2,H 2 )] moving proton H 2 from Lys 142 back to Ser 217. R r and R s were increased in steps of 0.1 Å with harmonic restraints of 5000 kcal mol 1 Å 2. For step (ii) (expulsion of methanol), the reaction co-ordinate R z {the distance [(C, O es )]} was increased in steps of 0.1 Å. The highest energy point along the path was the TS for protonation of the leaving group oxygen (O es ). This process led to E (Figure 2) overcoming a barrier of 47 kcal mol 1 relative to the substrate complex A. The subsequent expulsion of methanol happened spontaneously, as no effective barrier divides E from the acylenzyme. For OME, as for OA, the reaction rate is predicted to depend to a similar extent on TI formation and collapse. Tubert- Brohman et al. [22] have also compared the reactions of these two substrates, but only the first step of acylation for OME. Figure 2 summarizes the energetics (at the PM3- CHARMM22 level) for acylation of OA and OME. The overall calculated barrier (relative to A) for the amide cleavage is 40 kcal mol 1, 7 kcal mol 1 less than the barrier for the ester. These results are in qualitative agreement with kinetic investigations on OA and OME substrates: the experimental barrier for the hydrolysis of OA is lower by 0.6 kcal mol 1 than that for OME [13]. The calculations suggest that this unusual preference for the amide depends both on a more efficient stabilization of the TS for the deprotonation of Ser 241 and on higher basicity of the nitrogen N am compared with that of oxygen O es in the TI. We have also used this QM/MM modelling approach to investigate the reaction between FAAH and the carbamic inhibitor URB524 [23]. Carbamates and urea compounds inactivate FAAH by carbamoylating the Ser 241 nucleophile [24,25]. URB524 and its derivatives can be docked in two possible orientations within the FAAH catalytic site (called orientations I and II), both of which place the carbamic group close to Ser 241. Traditional computational tools employed in drug discovery (such as docking and scoring) failed to discriminate clearly between these two binding orientations [26,27]. We have studied the carbamoylation reaction of this inhibitor in FAAH, employing the PM3-CHARMM22 potential, with B3LYP/6-31G+(d) energy corrections. PESs were calculated for each binding orientation, and TS structures and intermediates were identified along the reaction profiles. The calculations clearly showed that reaction was significantly energetically preferred in orientation II, identifying II as the productive binding mode [23]. These findings are consistent with the recent development of potent new FAAH inhibitors, and with linear interaction energy calculations [28,29] for a series of 22 new FAAH carbamoylating compounds [30]. Altogether, our results indicate that QM/MM modelling of reaction mechanisms of covalent inhibitors in enzyme targets has the potential to provide detailed information for drug design in cases where docking approaches alone may be inadequate [31]. Acknowledgements We thank our co-authors and collaborators in relevant work reviewed here. Funding A.J.M. is supported by an Engineering and Physical Sciences Research Council Leadership Fellowship [grant number EP/G007705/1]. References 1 Cavalli, A., Carloni, P. and Recanatini, M. (2006) Target-related applications of first principles quantum chemical methods in drug design. Chem. Rev. 106, Field, M.J., Bash, P.A. and Karplus, M. (1990) A combined quantum mechanical and molecular mechanical potential for molecular dynamics simulations. J. Comput. Chem. 11, Warshel, A. (2003) Computer simulations of enzyme catalysis: methods, progress, and insights. Annu. Rev. Biophys. Biomol. Struct. 32, Friesner, R.A. and Guallar, V. (2005) Ab initio quantum chemical and mixed quantum mechanics/molecular mechanics (QM/MM) methods for studying enzymatic catalysis. Annu. Rev. Phys. Chem. 56, Senn, H.S. and Thiel, W. (2007) QM/MM studies of enzymes. Curr. Opin. Chem. Biol. 11,
5 Enzyme Mechanisms: Fast Reaction and Computational Approaches Ridder, L. and Mulholland, A.J. (2003) Modeling biotransformation reactions by combined quantum mechanical/molecular mechanical approaches: from structure to activity. Curr. Top. Med. Chem. 3, Claeyssens, F., Harvey, J.N., Manby, F.R., Mata, R.A., Mulholland, A.J., Ranaghan, K.E., Schütz, M., Thiel, S., Thiel, W. and Werner, H.-J. (2006) High accuracy computation of reaction barriers in enzymes. Angew. Chem. Int. Ed. Engl. 45, Mulholland, A.J. (2007) Chemical accuracy in QM/MM calculations on enzyme-catalysed reactions. Chem. Cent. J. 1, Mulholland, A.J. (2005) Modelling enzyme reaction mechanisms, specificity and catalysis. Drug Discov. Today 10, Piomelli, D. (2003) The molecular logic of endocannabinoid signalling. Nat. Rev. Neurosci. 4, Piomelli, D., Tarzia, G., Duranti, A., Tontini, A., Mor, M., Compton, T.R., Dasse, O., Monaghan, E.P., Parrott, J.A. and Putman, D. (2006) Pharmacological profile of the selective FAAH inhibitor KDS-4103 (URB597). CNS Drug Rev. 12, Labar, G. and Michaux, C. (2007) Fatty acid amide hydrolase: from characterization to therapeutics. Chem. Biodiversity 4, McKinney, M.K. and Cravatt, B.F. (2003) Evidence for distinct roles in catalysis for residues of the serine-serine-lysine catalytic triad of fatty acid amide hydrolase. J. Biol. Chem. 278, McKinney, M.K. and Cravatt, B.F. (2005) Structure and function of fatty acid amide hydrolase. Annu. Rev. Biochem. 74, Ahn, K., McKinney, M.K. and Cravatt, B.F. (2008) Enzymatic pathways that regulate endocannabinoid signaling in the nervous system. Chem. Rev. 108, Bracey, M.H., Hanson, M.A., Masuda, K.R., Stevens, R.C. and Cravatt, B. F. (2002) Structural adaptation in a membrane enzyme that terminates endocannabinoid signaling. Science 298, Lodola, A., Mor, M., Hermann, J.C., Tarzia, G., Piomelli, D. and Mulholland, A.J. (2005) QM/MM modelling of oleamide hydrolysis in fatty acid amide hydrolase (FAAH) reveals a new mechanism of nucleophile activation. Chem. Commun., Mulholland, A.J. (2008) Computational enzymology: modelling the mechanisms of biological catalysts. Biochem. Soc. Trans. 36, Klähn, M., Braun-Sand, S., Rosta, E. and Warshel, A. (2005) On possible pitfalls in ab initio quantum mechanics/molecular mechanics minimization approaches for studies of enzymatic reactions. J. Phys. Chem. B 109, van der Kamp, M.W., Shaw, K.E., Woods, C.J. and Mulholland, A.J. (2008) Biomolecular simulation and modelling: status, progress and prospects. J. R. Soc. Interface 5, S173 S Lodola, A., Mor, M., Zurek, J., Tarzia, G., Piomelli, D., Harvey, J.N. and Mulholland, A.J. (2007) Conformational effects in enzyme catalysis: reaction via a high energy conformation in fatty acid amide Hydrolase. Biophys. J. 92, L20 L22 22 Tubert-Brohman, I., Acevedo, O. and Jorgensen, W.L. (2006) Elucidation of hydrolysis mechanisms for fatty acid amide hydrolase and its Lys142Ala variant via QM/MM simulations. J. Am. Chem. Soc. 128, Lodola, A., Mor, M., Rivara, S., Christov, C., Tarzia, G., Piomelli, D. and Mulholland, A.J. (2008) Identification of productive inhibitor binding orientation in fatty acid amide hydrolase (FAAH) by QM/MM mechanistic modelling. Chem. Commun., Alexander, J.P. and Cravatt, B.F. (2005) Mechanism of carbamate inactivation of FAAH: implications for the design of covalent inhibitors and in vivo functional probes for enzymes. Chem. Biol. 12, Ahn, K., Johnson, D.S., Fitzgerald, L.R., Liimatta, M., Arendse, A., Stevenson, T., Lund, E.T., Nugent, R.A., Nomanbhoy, T.K., Alexander, J.P. and Cravatt, B.F. (2007) Novel mechanistic class of fatty acid amide hydrolase inhibitors with remarkable selectivity. Biochemistry 46, Mor, M., Rivara, S., Lodola, A., Plazzi, P.V., Tarzia, G., Duranti, A., Tontini, A., Piersanti, G., Kathuria, S. and Piomelli, D. (2008) Cyclohexylcarbamic acid 3 -or4 -substituted biphenyl-3-yl esters as fatty acid amide hydrolase inhibitors: synthesis, quantitative structure-activity relationships, and molecular modeling studies. J. Med. Chem. 47, Tarzia, G., Duranti, A., Gatti, G., Piersanti, G., Tontini, A., Rivara, S., Lodola, A., Plazzi, P.V., Mor, M., Kathuria, S. and Piomelli, D. (2006) Synthesis and structure activity relationships of FAAH inhibitors: cyclohexylcarbamic acid biphenyl esters with chemical modulation at the proximal phenyl ring. ChemMedChem 1, Carlson, H.A. and Jorgensen, W.L. (1995) An extended linear response method for determining free energies of hydration. J. Phys. Chem. 99, Zhou, R., Friesner, R.A., Ghosh, A., Rizzo, R.C., Jorgensen, W.L. and Levy, R.M. (2001) New linear interaction method for binding affinity calculations using a continuum solvent model. J. Phys. Chem. B 105, Mor, M., Lodola, A., Rivara, S., Vacondio, F., Duranti, A., Tontini, A., Sanchini, S., Piersanti, G., Clapper, J.R., King, A.R. et al. (2008) Synthesis and quantitative structure activity relationship of fatty acid amide hydrolase inhibitors: modulation at the n-portion of biphenyl-3-yl alkylcarbamates. J. Med. Chem. 51, Lodola, A., Woods, C.J. and Mulholland, A.J. (2008) Applications and advances of QM/MM methods in computational enzymology. Annu. Rep. Comput. Chem. 4, Received 17 November 2008 doi: /bst
Previous Class. Today. Term test I discussions. Detection of enzymatic intermediates: chymotrypsin mechanism
 Term test I discussions Previous Class Today Detection of enzymatic intermediates: chymotrypsin mechanism Mechanistic Understanding of Enzymemediated Reactions Ultimate goals: Identification of the intermediates,
Term test I discussions Previous Class Today Detection of enzymatic intermediates: chymotrypsin mechanism Mechanistic Understanding of Enzymemediated Reactions Ultimate goals: Identification of the intermediates,
PAPER No. : 16, Bioorganic and biophysical chemistry MODULE No. : 22, Mechanism of enzyme catalyst reaction (I) Chymotrypsin
 Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Bio 100 Serine Proteases 9/26/11
 Assigned Reading: 4th ed. 6.4.1 The Chymotrypsin Mechanism Involves Acylation And Deacylation Of A Ser Residue p. 213 BOX 20-1 Penicillin and β-lactamase p. 779 6.5.7 Some Enzymes Are Regulated By Proteolytic
Assigned Reading: 4th ed. 6.4.1 The Chymotrypsin Mechanism Involves Acylation And Deacylation Of A Ser Residue p. 213 BOX 20-1 Penicillin and β-lactamase p. 779 6.5.7 Some Enzymes Are Regulated By Proteolytic
Chymotrypsin Lecture. Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin
 Chymotrypsin Lecture Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin What s so great about enzymes? They accomplish large rate accelerations (10 10-10
Chymotrypsin Lecture Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin What s so great about enzymes? They accomplish large rate accelerations (10 10-10
6. The catalytic mechanism of arylsulfatase A and its theoretical investigation
 6. The catalytic mechanism of arylsulfatase A and its theoretical investigation When the crystal structure of arylsulfatase A was solved, a remarkable structural analogy to another hydrolytic enzyme, the
6. The catalytic mechanism of arylsulfatase A and its theoretical investigation When the crystal structure of arylsulfatase A was solved, a remarkable structural analogy to another hydrolytic enzyme, the
MITOCW watch?v=xms9dyhqhi0
 MITOCW watch?v=xms9dyhqhi0 The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high-quality, educational resources for free.
MITOCW watch?v=xms9dyhqhi0 The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high-quality, educational resources for free.
Lecture 18 (10/27/17) Lecture 18 (10/27/17)
 Reading: Ch6; 225-232 Lecture 18 (10/27/17) Problems: Ch5 (text); 2 Ch6 (study guide-facts); 5, 6, 7, 14 NEXT Reading: Ch5; 164, 166-169 Problems: none Remember Monday at 6:30 in PHO-206 is the first MB
Reading: Ch6; 225-232 Lecture 18 (10/27/17) Problems: Ch5 (text); 2 Ch6 (study guide-facts); 5, 6, 7, 14 NEXT Reading: Ch5; 164, 166-169 Problems: none Remember Monday at 6:30 in PHO-206 is the first MB
Chemical Mechanism of Enzymes
 Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
All chemicals were obtained from Aldrich, Acros, Fisher, or Fluka and were used without
 Supplemental Data Alexander et al. Experimental Procedures General Methods for Inhibitor Synthesis All chemicals were obtained from Aldrich, Acros, Fisher, or Fluka and were used without further purification,
Supplemental Data Alexander et al. Experimental Procedures General Methods for Inhibitor Synthesis All chemicals were obtained from Aldrich, Acros, Fisher, or Fluka and were used without further purification,
Enzyme Catalysis-Serine Proteases
 Enzyme Catalysis-Serine Proteases Concepts to be learned Activation Energy Transition State Example: Proteases Requirements for proteolysis Families of proteases Protein Folds used by proteases for catalysis
Enzyme Catalysis-Serine Proteases Concepts to be learned Activation Energy Transition State Example: Proteases Requirements for proteolysis Families of proteases Protein Folds used by proteases for catalysis
CHAPTER 9: CATALYTIC STRATEGIES. Chess vs Enzymes King vs Substrate
 CHAPTER 9: CATALYTIC STRATEGIES Chess vs Enzymes King vs Substrate INTRODUCTION CHAPTER 9 What are the sources of the catalytic power and specificity of enzymes? Problems in reactions in cells Neutral
CHAPTER 9: CATALYTIC STRATEGIES Chess vs Enzymes King vs Substrate INTRODUCTION CHAPTER 9 What are the sources of the catalytic power and specificity of enzymes? Problems in reactions in cells Neutral
Glycosidic bond cleavage
 Glycosidases and Glycosyltransferases Introduction to Inverting/Retaining Mechanisms Inhibitor design Chemical Reaction Proposed catalytic mechanisms Multiple slides courtesy of Harry Gilbert with Wells
Glycosidases and Glycosyltransferases Introduction to Inverting/Retaining Mechanisms Inhibitor design Chemical Reaction Proposed catalytic mechanisms Multiple slides courtesy of Harry Gilbert with Wells
A. B. C. D. E. F. G. H. I. J. K. Ser/Thr. Ser/Thr. Ser/Thr. Ser/Thr. Ser/Thr. Asn. Asn. Asn. Asn. Asn. Asn
 A. B. C. D. E. F. "3 "3!4!3 Ser/Thr "3!4!3!4 Asn Asn Ser/Thr Asn!3!6 Ser/Thr G. H. I. J. K.!3 Ser/Thr Ser/Thr 4 4 2 2 6 3 6 Asn Asn Asn Glycosidases and Glycosyltransferases Introduction to Inverting/Retaining
A. B. C. D. E. F. "3 "3!4!3 Ser/Thr "3!4!3!4 Asn Asn Ser/Thr Asn!3!6 Ser/Thr G. H. I. J. K.!3 Ser/Thr Ser/Thr 4 4 2 2 6 3 6 Asn Asn Asn Glycosidases and Glycosyltransferases Introduction to Inverting/Retaining
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Figure 1. A ribbon diagram of the aldolase (A) and a close up of the active site (B) including the bound substrate.
 Problem Set 4 (C-C bond formation, phosphoryl transfer reactions and the role of ATP) 1. Chemists can use the same strategies as nature to make new carbon-carbon bonds stereospecifically using enzymes
Problem Set 4 (C-C bond formation, phosphoryl transfer reactions and the role of ATP) 1. Chemists can use the same strategies as nature to make new carbon-carbon bonds stereospecifically using enzymes
Carboxylic Acid Derivatives Reading Study Problems Key Concepts and Skills Lecture Topics: Structures and reactivity of carboxylic acid derivatives
 Carboxylic Acid Derivatives Reading: Wade chapter 21, sections 21-1- 21-16 Study Problems: 21-45, 21-46, 21-48, 21-49, 21-50, 21-53, 21-56, 21-58, 21-63 Key Concepts and Skills: Interpret the spectra of
Carboxylic Acid Derivatives Reading: Wade chapter 21, sections 21-1- 21-16 Study Problems: 21-45, 21-46, 21-48, 21-49, 21-50, 21-53, 21-56, 21-58, 21-63 Key Concepts and Skills: Interpret the spectra of
From Structure to Function (II): Enzyme Structure & Catalysis
 BCHS 6229 Protein Structure and Function Lecture 5 (Oct 25, 2011) From Structure to Function (II): Enzyme Structure & Catalysis 1 Outline Catalysis: Overview Active site geometry Proximity and ground-state
BCHS 6229 Protein Structure and Function Lecture 5 (Oct 25, 2011) From Structure to Function (II): Enzyme Structure & Catalysis 1 Outline Catalysis: Overview Active site geometry Proximity and ground-state
7.05 Spring 2004 March 12, Recitation #4
 7.05 Spring 2004 March 12, 2004 Recitation #4 ontact Information TA: Victor Sai Recitation: Friday, 34pm, 2132 Email: sai@mit.edu ffice ours: Friday, 45pm, 2132 Unit 2 Schedule Recitation/Exam Date Lectures
7.05 Spring 2004 March 12, 2004 Recitation #4 ontact Information TA: Victor Sai Recitation: Friday, 34pm, 2132 Email: sai@mit.edu ffice ours: Friday, 45pm, 2132 Unit 2 Schedule Recitation/Exam Date Lectures
Peptide hydrolysis uncatalyzed half-life = ~450 years HIV protease-catalyzed half-life = ~3 seconds
 Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Chapter 11: Enzyme Catalysis
 Chapter 11: Enzyme Catalysis Matching A) high B) deprotonated C) protonated D) least resistance E) motion F) rate-determining G) leaving group H) short peptides I) amino acid J) low K) coenzymes L) concerted
Chapter 11: Enzyme Catalysis Matching A) high B) deprotonated C) protonated D) least resistance E) motion F) rate-determining G) leaving group H) short peptides I) amino acid J) low K) coenzymes L) concerted
UNIVERSITY OF GUELPH CHEM 4540 ENZYMOLOGY Winter 2005 Quiz #2: March 24, 2005, 11:30 12:50 Instructor: Prof R. Merrill ANSWERS
 UNIVERSITY F GUELPH CHEM 4540 ENZYMLGY Winter 2005 Quiz #2: March 24, 2005, 11:30 12:50 Instructor: Prof R. Merrill ANSWERS Instructions: Time allowed = 80 minutes. Total marks = 30. This quiz represents
UNIVERSITY F GUELPH CHEM 4540 ENZYMLGY Winter 2005 Quiz #2: March 24, 2005, 11:30 12:50 Instructor: Prof R. Merrill ANSWERS Instructions: Time allowed = 80 minutes. Total marks = 30. This quiz represents
Mechanisms of Enzymes
 Mechanisms of Enzymes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy How enzymes work * Chemical reactions have an energy
Mechanisms of Enzymes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy How enzymes work * Chemical reactions have an energy
1. Measurement of the rate constants for simple enzymatic reaction obeying Michaelis- Menten kinetics gave the following results: =3x10-5 = 30μM
 1. Measurement of the rate constants for simple enzymatic reaction obeying Michaelis- Menten kinetics gave the following results: k 1 = 2 x 10 8 M -1 s -1, k 2 = 1 x 10 3 s -1, k 3 = 5 x 10 3 s -1 a) What
1. Measurement of the rate constants for simple enzymatic reaction obeying Michaelis- Menten kinetics gave the following results: k 1 = 2 x 10 8 M -1 s -1, k 2 = 1 x 10 3 s -1, k 3 = 5 x 10 3 s -1 a) What
Statin inhibition of HMG-CoA reductase: a 3-dimensional view
 Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Molecular Graphics Perspective of Protein Structure and Function
 Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Catalysis & specificity: Proteins at work
 Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Final Exam Chemistry 391 Structural Biochemistry Fall Do not open the exam until ready to begin! Rules of the Game:
 Name Practice for 2018 Final Exam Chemistry 391 Structural Biochemistry Fall 2016 Do not open the exam until ready to begin! ules of the Game: This is a take-home Exam. The exam is due on Thursday, December
Name Practice for 2018 Final Exam Chemistry 391 Structural Biochemistry Fall 2016 Do not open the exam until ready to begin! ules of the Game: This is a take-home Exam. The exam is due on Thursday, December
Enzymes: The Catalysts of Life
 Chapter 6 Enzymes: The Catalysts of Life Lectures by Kathleen Fitzpatrick Simon Fraser University Activation Energy and the Metastable State Many thermodynamically feasible reactions in a cell that could
Chapter 6 Enzymes: The Catalysts of Life Lectures by Kathleen Fitzpatrick Simon Fraser University Activation Energy and the Metastable State Many thermodynamically feasible reactions in a cell that could
Enzyme Catalytic Mechanisms. Dr. Kevin Ahern
 Enzyme Catalytic Mechanisms Dr. Kevin Ahern Cleave Peptide Bonds Specificity of Cutting Common Active Site Composition/Structure Mechanistically Well Studied Chymotrypsin Chymotrypsin Catalysis H2O Chymotrypsin
Enzyme Catalytic Mechanisms Dr. Kevin Ahern Cleave Peptide Bonds Specificity of Cutting Common Active Site Composition/Structure Mechanistically Well Studied Chymotrypsin Chymotrypsin Catalysis H2O Chymotrypsin
Adenosine triphosphate (ATP)
 Adenosine triphosphate (ATP) 1 High energy bonds ATP adenosine triphosphate N NH 2 N -O O P O O P O- O- O O P O- O CH 2 H O H N N adenine phosphoanhydride bonds (~) H OH ribose H OH Phosphoanhydride bonds
Adenosine triphosphate (ATP) 1 High energy bonds ATP adenosine triphosphate N NH 2 N -O O P O O P O- O- O O P O- O CH 2 H O H N N adenine phosphoanhydride bonds (~) H OH ribose H OH Phosphoanhydride bonds
NIH Public Access Author Manuscript J Am Chem Soc. Author manuscript; available in PMC 2008 September 29.
 NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
Chapter 18. Carboxylic Acids and Their Derivatives. Nucleophilic Addition-Elimination at the Acyl Carbon
 Chapter 18 Carboxylic Acids and Their Derivatives. Nucleophilic Addition-Elimination at the Acyl Carbon Carboxylic Acids Organic compounds characterized by their acidity Contains COOH group (must be at
Chapter 18 Carboxylic Acids and Their Derivatives. Nucleophilic Addition-Elimination at the Acyl Carbon Carboxylic Acids Organic compounds characterized by their acidity Contains COOH group (must be at
Esters of Carboxylic Acids These are derivatives of carboxylic acids where the hydroxyl group is replaced by an alkoxy group.
 Carboxylic acid Derivatives Carboxylic acid derivatives are described as compounds that can be converted to carboxylic acids via simple acidic or basic hydrolysis. The most important acid derivatives are
Carboxylic acid Derivatives Carboxylic acid derivatives are described as compounds that can be converted to carboxylic acids via simple acidic or basic hydrolysis. The most important acid derivatives are
Chapter 18 Carboxylic Acids and Their Derivatives. Nucleophilic Addition- Elimination at the Acyl Carbon
 Chapter 18 Carboxylic Acids and Their Derivatives. Nucleophilic Addition- Elimination at the Acyl Carbon Introduction The carboxyl group (-CO 2 H) is the parent group of a family of compounds called acyl
Chapter 18 Carboxylic Acids and Their Derivatives. Nucleophilic Addition- Elimination at the Acyl Carbon Introduction The carboxyl group (-CO 2 H) is the parent group of a family of compounds called acyl
MCB 102 Discussion, Spring 2012
 MB Discussion, Spring 2012 Practice Problems 1. Effect of enzymes on reactions Which of the listed effects would be brought about by any enzyme catalyzing the following simple reaction? k 1 S P where K
MB Discussion, Spring 2012 Practice Problems 1. Effect of enzymes on reactions Which of the listed effects would be brought about by any enzyme catalyzing the following simple reaction? k 1 S P where K
CHM 341 C: Biochemistry I. Test 2: October 24, 2014
 CHM 341 C: Biochemistry I Test 2: ctober 24, 2014 This test consists of 14 questions worth points. Make sure that you read the entire question and answer each question clearly and completely. To receive
CHM 341 C: Biochemistry I Test 2: ctober 24, 2014 This test consists of 14 questions worth points. Make sure that you read the entire question and answer each question clearly and completely. To receive
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
Structure of Alkenes In ethene (ethylene) each carbon is bonded to 3 other atoms, with zero nonbonding electrons => sp 2 hybridization.
 Structure and Synthesis of Alkenes Alkenes (olefins) are hydrocarbons which have carbon carbon double bonds. A double bond is a bond and a bond. Double bond B.D.E. bond B.D.E. = 146 kcal/mol = 83 kcal/mol
Structure and Synthesis of Alkenes Alkenes (olefins) are hydrocarbons which have carbon carbon double bonds. A double bond is a bond and a bond. Double bond B.D.E. bond B.D.E. = 146 kcal/mol = 83 kcal/mol
Lecture 19. Nucleophilic Acyl Substitution Y - + X - Y X R C X. April 2, Chemistry 328N
 Lecture 19 Nucleophilic Acyl Substitution X Y - - Y X X - Y April 2, 2019 hemistry 328N Acid-catalyzed Esterification (also called Fischer esterification) H H 3 H H H 2 H 3 Please study the mechanism hemistry
Lecture 19 Nucleophilic Acyl Substitution X Y - - Y X X - Y April 2, 2019 hemistry 328N Acid-catalyzed Esterification (also called Fischer esterification) H H 3 H H H 2 H 3 Please study the mechanism hemistry
Functional Derivatives of Carboxylic Acids
 Functional Derivatives of Carboxylic Acids Derivatives of Carboxylic Acids are compounds in which the OH of a carboxyl group has been replaced by CI, OOCR, NH2, or OR'to convert acid chlorides,anhydrides,
Functional Derivatives of Carboxylic Acids Derivatives of Carboxylic Acids are compounds in which the OH of a carboxyl group has been replaced by CI, OOCR, NH2, or OR'to convert acid chlorides,anhydrides,
Carboxylic Acid Derivatives: Nucleophilic Acyl Substitution
 Carboxylic Acid Derivatives: Nucleophilic Acyl Substitution Carboxylic Acid Derivatives Carboxylic acid derivatives. Acyl chloride Acid anhydride Ester Amide Nucleophilic acyl substitution 19.1 Nomenclature
Carboxylic Acid Derivatives: Nucleophilic Acyl Substitution Carboxylic Acid Derivatives Carboxylic acid derivatives. Acyl chloride Acid anhydride Ester Amide Nucleophilic acyl substitution 19.1 Nomenclature
Chapter 20: Carboxylic Acid Derivatives: Nucleophilic Acyl Substitution
 hapter 20: arboxylic Acid Derivatives: ucleophilic Acyl Substitution 20.1: omenclature of arboxylic Acid Derivatives (please read) carboxylic acid -oic acid ' ester -oate ' lactone cyclic ester l acid
hapter 20: arboxylic Acid Derivatives: ucleophilic Acyl Substitution 20.1: omenclature of arboxylic Acid Derivatives (please read) carboxylic acid -oic acid ' ester -oate ' lactone cyclic ester l acid
molecules ISSN
 Molecules 2000, 5, 1399-1407 molecules ISSN 1420-3049 http://www.mdpi.org Substitution Effects on Reactivity of N-Acyl-2-amino-2-desoxyglucopyranoses. Quantum Chemical Study Aušra Vektariene 1, *, Arvydas
Molecules 2000, 5, 1399-1407 molecules ISSN 1420-3049 http://www.mdpi.org Substitution Effects on Reactivity of N-Acyl-2-amino-2-desoxyglucopyranoses. Quantum Chemical Study Aušra Vektariene 1, *, Arvydas
Fundamentals of Organic Chemistry CHEM 109 For Students of Health Colleges Credit hrs.: (2+1)
 Fundamentals of Organic Chemistry CHEM 109 For Students of Health Colleges Credit hrs.: (2+1) King Saud University College of Science, Chemistry Department CHEM 109 CHAPTER 7. CARBOXYLIC ACIDS AND THEIR
Fundamentals of Organic Chemistry CHEM 109 For Students of Health Colleges Credit hrs.: (2+1) King Saud University College of Science, Chemistry Department CHEM 109 CHAPTER 7. CARBOXYLIC ACIDS AND THEIR
Chapter 10. Carboxylic Acids and Derivatives. Naming Carboxylic Acids and Derivatives. Carboxylic Acids: RCOOH (RCO 2 H)
 Chapter 10 Carboxylic Acids and Derivatives Naming Carboxylic Acids and Derivatives Carboxylic Acids: RCH (RC 2 H) The functional group of a carboxylic acid is a carboxyl group (carbonyl & hydroxyl group)
Chapter 10 Carboxylic Acids and Derivatives Naming Carboxylic Acids and Derivatives Carboxylic Acids: RCH (RC 2 H) The functional group of a carboxylic acid is a carboxyl group (carbonyl & hydroxyl group)
The MOLECULES of LIFE
 The MOLECULES of LIFE Physical and Chemical Principles Solutions Manual Prepared by James Fraser and Samuel Leachman Chapter 16 Principles of Enzyme Catalysis Problems True/False and Multiple Choice 1.
The MOLECULES of LIFE Physical and Chemical Principles Solutions Manual Prepared by James Fraser and Samuel Leachman Chapter 16 Principles of Enzyme Catalysis Problems True/False and Multiple Choice 1.
Carboxylic Acids. The Importance of Carboxylic Acids (RCO 2 H)
 Carboxylic Acids The Importance of Carboxylic Acids (RCO 2 H) Starting materials for acyl derivatives (esters, amides, and acid chlorides) Abundant in nature from oxidation of aldehydes and alcohols in
Carboxylic Acids The Importance of Carboxylic Acids (RCO 2 H) Starting materials for acyl derivatives (esters, amides, and acid chlorides) Abundant in nature from oxidation of aldehydes and alcohols in
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached
 BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
CARBOXYLIC ACIDS AND THEIR DERIVATIVES: NUCLEOPHILIC ADDITION-ELIMINATION AT THE ACYL CARBON
 CARBOXYLIC ACIDS AND THEIR DERIVATIVES: NUCLEOPHILIC ADDITION-ELIMINATION AT THE ACYL CARBON RED ANT WAS SOURCE OF FORMIC ACID (RCOOH) Lecture 8 ORGANIC CHEMISTRY 2 Introduction The carboxyl group (-CO
CARBOXYLIC ACIDS AND THEIR DERIVATIVES: NUCLEOPHILIC ADDITION-ELIMINATION AT THE ACYL CARBON RED ANT WAS SOURCE OF FORMIC ACID (RCOOH) Lecture 8 ORGANIC CHEMISTRY 2 Introduction The carboxyl group (-CO
Lecture 20. Herman Emil Fischer Nobel Prize 1902 Sugars, Esters and Purines. April 4, Chemistry 328N
 Lecture 20 April 4, 2019 Herman Emil Fischer 1852-1919 Nobel Prize 1902 Sugars, Esters and Purines Acid-catalyzed Esterification (also called Fischer esterification) CH CH 3 H H H 2 CCH 3 Please study
Lecture 20 April 4, 2019 Herman Emil Fischer 1852-1919 Nobel Prize 1902 Sugars, Esters and Purines Acid-catalyzed Esterification (also called Fischer esterification) CH CH 3 H H H 2 CCH 3 Please study
1. For the following reaction, at equilibrium [S] = 5 mm, [P] = 0.5 mm, and k f = 10 s -1. k f
![1. For the following reaction, at equilibrium [S] = 5 mm, [P] = 0.5 mm, and k f = 10 s -1. k f 1. For the following reaction, at equilibrium [S] = 5 mm, [P] = 0.5 mm, and k f = 10 s -1. k f](/thumbs/77/75734355.jpg) 1. For the following reaction, at equilibrium [S] = 5 mm, [P] = 0.5 mm, and k f = 10 s -1. S k f k r P a) Calculate K eq (the equilibrium constant) and k r. b) A catalyst increases k f by a factor of 10
1. For the following reaction, at equilibrium [S] = 5 mm, [P] = 0.5 mm, and k f = 10 s -1. S k f k r P a) Calculate K eq (the equilibrium constant) and k r. b) A catalyst increases k f by a factor of 10
X-ray Crystallographic Analysis of r-ketoheterocycle Inhibitors Bound to a Humanized Variant of Fatty Acid Amide Hydrolase
 J. Med. Chem. XXXX, XXX, 000 000 A DOI: 10.1021/jm9012196 X-ray Crystallographic Analysis of r-ketoheterocycle Inhibitors Bound to a Humanized Variant of Fatty Acid Amide Hydrolase Mauro Mileni, Joie Garfunkle,
J. Med. Chem. XXXX, XXX, 000 000 A DOI: 10.1021/jm9012196 X-ray Crystallographic Analysis of r-ketoheterocycle Inhibitors Bound to a Humanized Variant of Fatty Acid Amide Hydrolase Mauro Mileni, Joie Garfunkle,
Enzymes. Enzymes accelerate chemical reactions as the engine accelerates this drag race.
 Chapter 30 Enzymes Enzymes accelerate chemical reactions as the engine accelerates this drag race. Introduction to General, Organic, and Biochemistry, 10e John Wiley & Sons, Inc Morris Hein, Scott Pattison,
Chapter 30 Enzymes Enzymes accelerate chemical reactions as the engine accelerates this drag race. Introduction to General, Organic, and Biochemistry, 10e John Wiley & Sons, Inc Morris Hein, Scott Pattison,
Chemistry 135, First Exam. September 23, Chem 135, Exam 1 SID:
 Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Break Desired Compound into 3 Smaller Ones
 Break Desired Compound into 3 Smaller nes A B C Indole ydrazine Phenol When creating a covalent link between model compounds move the charge on the deleted into the carbon to maintain integer charge (i.e.
Break Desired Compound into 3 Smaller nes A B C Indole ydrazine Phenol When creating a covalent link between model compounds move the charge on the deleted into the carbon to maintain integer charge (i.e.
Finding the Sweet Spot- Mechanism Guided Design of Glycosidase Inhibitors. Jahnabi Roy CHEM 575 Seminar 11/01/12
 Finding the Sweet Spot- Mechanism Guided Design of Glycosidase Inhibitors Jahnabi Roy CHEM 575 Seminar 11/01/12 Glycans and Glycosyl Hydrolases http://cellbiology.med.unsw.edu.au/units/science/lecture0803.htm
Finding the Sweet Spot- Mechanism Guided Design of Glycosidase Inhibitors Jahnabi Roy CHEM 575 Seminar 11/01/12 Glycans and Glycosyl Hydrolases http://cellbiology.med.unsw.edu.au/units/science/lecture0803.htm
Chapter 19: Carboxylic Acid Derivatives: Nucleophilic Acyl Substitution 19.1: Nomenclature of Carboxylic Acid Derivatives (please read)
 problem 18.33b - = 128.7 123.9 179.7 146.8 147.4 45.3 18.0 161 hapter 19: arboxylic Acid Derivatives: ucleophilic Acyl Substitution 19.1: omenclature of arboxylic Acid Derivatives (please read) carboxylic
problem 18.33b - = 128.7 123.9 179.7 146.8 147.4 45.3 18.0 161 hapter 19: arboxylic Acid Derivatives: ucleophilic Acyl Substitution 19.1: omenclature of arboxylic Acid Derivatives (please read) carboxylic
Student Biochemistry I Homework III Due 10/13/04 64 points total (48 points based on text; 16 points for Swiss-PDB viewer exercise)
 Biochemistry I Homework III Due 10/13/04 64 points total (48 points based on text; 16 points for Swiss-PDB viewer exercise) 1). 20 points total T or F; if false, provide a brief rationale as to why. Only
Biochemistry I Homework III Due 10/13/04 64 points total (48 points based on text; 16 points for Swiss-PDB viewer exercise) 1). 20 points total T or F; if false, provide a brief rationale as to why. Only
Computational Insights into the Structural Properties and Catalytic Functions of Selenoprotein Glutathione Peroxidase (GPx)
 1 1 Computational Insights into the Structural Properties and Catalytic Functions of Selenoprotein Glutathione Peroxidase (GPx) Rajeev Prabhakar, Keiji Morokuma, and Djamaladdin G. Musaev 1.1 Introduction
1 1 Computational Insights into the Structural Properties and Catalytic Functions of Selenoprotein Glutathione Peroxidase (GPx) Rajeev Prabhakar, Keiji Morokuma, and Djamaladdin G. Musaev 1.1 Introduction
Physical properties: C L = L. Cl, NH 2, OCH 3, OH, OCR O O O NH 2 CH 3 N(CH 3 ) 2. Sol. in H 2 O
 Lecture Notes hem 51 S. King hapter 22 arboxylic Acids and their Derivatives: Nucleophilic Acyl Substitution I. Structure and Physical Properties: Type 2 carbonyl compounds (carboxylic acids and derivatives)
Lecture Notes hem 51 S. King hapter 22 arboxylic Acids and their Derivatives: Nucleophilic Acyl Substitution I. Structure and Physical Properties: Type 2 carbonyl compounds (carboxylic acids and derivatives)
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids
 Amino acids 1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids 5-To understand amino acids synthesis Amino
Amino acids 1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids 5-To understand amino acids synthesis Amino
Concept 8.3: ATP powers cellular work by coupling exergonic reactions to endergonic reactions
 Concept 8.3: ATP powers cellular work by coupling exergonic reactions to endergonic reactions A cell does three main kinds of work: Chemical Transport Mechanical To do work, cells manage energy resources
Concept 8.3: ATP powers cellular work by coupling exergonic reactions to endergonic reactions A cell does three main kinds of work: Chemical Transport Mechanical To do work, cells manage energy resources
Alkenes are very useful in syntheses -they allow us to convert into many of the other types of functional groups.
 Chapter 7: Alkenes: reactions and synthesis Alkenes are very useful in syntheses -they allow us to convert into many of the other types of functional groups. 7.1 Preparation of alkenes: preview Addition
Chapter 7: Alkenes: reactions and synthesis Alkenes are very useful in syntheses -they allow us to convert into many of the other types of functional groups. 7.1 Preparation of alkenes: preview Addition
ESTERS AND RELATED CARBOXYLIC ACID DERIVATIVES. Jack DeRuiter
 ESTES AD ELATED ABYLI AID DEIVATIVES I. Structure and Preparation Jack Deuiter Esters are derivatives of carboxylic acids that arise via replacement of the hydroxyl () portion of the acid function with
ESTES AD ELATED ABYLI AID DEIVATIVES I. Structure and Preparation Jack Deuiter Esters are derivatives of carboxylic acids that arise via replacement of the hydroxyl () portion of the acid function with
Hydrolytic transformations involving amide-, ester bonds are the easiest to perform
 Hydrolytic reactions Hydrolytic transformations involving amide-, ester bonds are the easiest to perform Proteins used for these reactions are proteases, esterases or lipases. A favourite class of enzymes
Hydrolytic reactions Hydrolytic transformations involving amide-, ester bonds are the easiest to perform Proteins used for these reactions are proteases, esterases or lipases. A favourite class of enzymes
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Probing the catalytic mechanism of an antifibrotic copper metallodrug
 Probing the catalytic mechanism of an antifibrotic copper metallodrug 4 December 2017 Lizelle Lubbe Supervisor: Prof Ed Sturrock Transition metals offer benefit of designing compounds with complex architectures,
Probing the catalytic mechanism of an antifibrotic copper metallodrug 4 December 2017 Lizelle Lubbe Supervisor: Prof Ed Sturrock Transition metals offer benefit of designing compounds with complex architectures,
Biochemistry: Macromolecules
 1 Biology: Macromolecules 2 Carbohydrates Carbohydrate organic compound containing carbon, hydrogen, & oxygen in a 1:2:1 ratio Meaning: hydrated carbon ratio of h:0 is 2:1 (same as in water) Source: plants
1 Biology: Macromolecules 2 Carbohydrates Carbohydrate organic compound containing carbon, hydrogen, & oxygen in a 1:2:1 ratio Meaning: hydrated carbon ratio of h:0 is 2:1 (same as in water) Source: plants
Chapter Three (Biochemistry)
 Chapter Three (Biochemistry) 1 SECTION ONE: CARBON COMPOUNDS CARBON BONDING All compounds can be classified in two broad categories: organic compounds and inorganic compounds. Organic compounds are made
Chapter Three (Biochemistry) 1 SECTION ONE: CARBON COMPOUNDS CARBON BONDING All compounds can be classified in two broad categories: organic compounds and inorganic compounds. Organic compounds are made
Evidence for Distinct Roles in Catalysis for Residues of the Serine- Serine-Lysine Catalytic Triad of Fatty Acid Amide Hydrolase* S
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 278, No. 39, Issue of September 26, pp. 37393 37399, 2003 2003 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Evidence for
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 278, No. 39, Issue of September 26, pp. 37393 37399, 2003 2003 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Evidence for
Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex
 John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
PROTEOMICS August 27 31, 2007 Peter D'Eustachio - MSB
 PROTEOMICS August 27 31, 2007 Peter D'Eustachio - MSB 328 e-mail: deustp01@med.nyu.edu GOALS OF THIS SEGMENT OF THE COURSE Recognize the structures of the 20 amino acids; understand their properties as
PROTEOMICS August 27 31, 2007 Peter D'Eustachio - MSB 328 e-mail: deustp01@med.nyu.edu GOALS OF THIS SEGMENT OF THE COURSE Recognize the structures of the 20 amino acids; understand their properties as
1/3/2011. Chapter 17 Carboxylic Acids and Their Derivatives. Nucleophilic Addition- Elimination at the Acyl Carbon
 Introduction The carboxyl group (-CO 2 H) is the parent group of a family of compounds called acyl compounds or carboxylic acid derivatives Chapter 17 Carboxylic Acids and Their Derivatives. Nucleophilic
Introduction The carboxyl group (-CO 2 H) is the parent group of a family of compounds called acyl compounds or carboxylic acid derivatives Chapter 17 Carboxylic Acids and Their Derivatives. Nucleophilic
The effects of ph on Type VII-NA Bovine Intestinal Mucosal Alkaline Phosphatase Activity
 The effects of ph on Type VII-NA Bovine Intestinal Mucosal Alkaline Phosphatase Activity ANDREW FLYNN, DYLAN JONES, ERIC MAN, STEPHEN SHIPMAN, AND SHERMAN TUNG Department of Microbiology and Immunology,
The effects of ph on Type VII-NA Bovine Intestinal Mucosal Alkaline Phosphatase Activity ANDREW FLYNN, DYLAN JONES, ERIC MAN, STEPHEN SHIPMAN, AND SHERMAN TUNG Department of Microbiology and Immunology,
Stereoselective C,C-bond formation. Cyclizations of biradicals*
 Pure Appl. Chem., Vol. 72, No. 9, pp. 1623 1629, 2000. 2000 IUPAC Stereoselective C,C-bond formation. Cyclizations of biradicals* Bernd Giese, Frédérique Barbosa, Christian Stähelin, Stefan Sauer, Philipp
Pure Appl. Chem., Vol. 72, No. 9, pp. 1623 1629, 2000. 2000 IUPAC Stereoselective C,C-bond formation. Cyclizations of biradicals* Bernd Giese, Frédérique Barbosa, Christian Stähelin, Stefan Sauer, Philipp
Molecular Dynamics of Sialic Acid Analogues and their Interaction with Influenza Hemagglutinin
 Research Papers www.ijpsonline.com Molecular Dynamics of Sialic Acid Analogues and their Interaction with Influenza Hemagglutinin T. FEMLIN BLESSIA, V. S. RAPHEAL 1 AND D. J. S. SHARMILA* Department of
Research Papers www.ijpsonline.com Molecular Dynamics of Sialic Acid Analogues and their Interaction with Influenza Hemagglutinin T. FEMLIN BLESSIA, V. S. RAPHEAL 1 AND D. J. S. SHARMILA* Department of
Six Types of Enzyme Catalysts
 Six Types of Enzyme Catalysts Although a huge number of reactions occur in living systems, these reactions fall into only half a dozen types. The reactions are: 1. Oxidation and reduction. Enzymes that
Six Types of Enzyme Catalysts Although a huge number of reactions occur in living systems, these reactions fall into only half a dozen types. The reactions are: 1. Oxidation and reduction. Enzymes that
PHARMACOPHORE MODELING
 CHAPTER 9 PHARMACOPHORE MODELING 9.1 PHARMACOPHORES AS VIRTUAL SCREENING TOOLS The pharmacophore concept has been widely used over the past decades for rational drug design approaches and has now been
CHAPTER 9 PHARMACOPHORE MODELING 9.1 PHARMACOPHORES AS VIRTUAL SCREENING TOOLS The pharmacophore concept has been widely used over the past decades for rational drug design approaches and has now been
Chapter 20: Carboxylic Acids and Nitriles شیمی آلی 2
 Chapter 20: Carboxylic Acids and Nitriles شیمی آلی 2 Dr M. Mehrdad University of Guilan, Department of Chemistry, Rasht, Iran m-mehrdad@guilan.ac.ir Based on McMurry s Organic Chemistry, 7 th edition The
Chapter 20: Carboxylic Acids and Nitriles شیمی آلی 2 Dr M. Mehrdad University of Guilan, Department of Chemistry, Rasht, Iran m-mehrdad@guilan.ac.ir Based on McMurry s Organic Chemistry, 7 th edition The
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/ MCQs. Location : 102, 105, 106, 301, 302
 FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
CELLULAR METABOLISM. Metabolic pathways can be linear, branched, cyclic or spiral
 CHM333 LECTURE 24 & 25: 3/27 29/13 SPRING 2013 Professor Christine Hrycyna CELLULAR METABOLISM What is metabolism? - How cells acquire, transform, store and use energy - Study reactions in a cell and how
CHM333 LECTURE 24 & 25: 3/27 29/13 SPRING 2013 Professor Christine Hrycyna CELLULAR METABOLISM What is metabolism? - How cells acquire, transform, store and use energy - Study reactions in a cell and how
Organic Chemistry - Problem Drill 23: Amino Acids, Peptides, and Proteins
 rganic Chemistry - Problem Drill 23: Amino Acids, Peptides, and Proteins No. 1 of 10 1. Which amino acid does not contain a chiral center? (A) Serine (B) Proline (C) Alanine (D) Phenylalanine (E) Glycine
rganic Chemistry - Problem Drill 23: Amino Acids, Peptides, and Proteins No. 1 of 10 1. Which amino acid does not contain a chiral center? (A) Serine (B) Proline (C) Alanine (D) Phenylalanine (E) Glycine
Objectives. Carbon Bonding. Carbon Bonding, continued. Carbon Bonding
 Biochemistry Table of Contents Objectives Distinguish between organic and inorganic compounds. Explain the importance of carbon bonding in biological molecules. Identify functional groups in biological
Biochemistry Table of Contents Objectives Distinguish between organic and inorganic compounds. Explain the importance of carbon bonding in biological molecules. Identify functional groups in biological
Carboxylic Acid Derivatives
 arboxylic Acid Derivatives The most important derivatives of carboxylic acids are l " ' ' acid halide acid anhydride an ester an amide Although not direct derivatives, nitriles, -, are related to carboxylic
arboxylic Acid Derivatives The most important derivatives of carboxylic acids are l " ' ' acid halide acid anhydride an ester an amide Although not direct derivatives, nitriles, -, are related to carboxylic
Final Exam Chemistry 391 Structural Biochemistry Fall Do not open the exam until ready to begin! Rules of the Game:
 ame Practice Exam Solutions 2018 Final Exam Chemistry 391 Structural Biochemistry Fall 2016 Do not open the exam until ready to begin! ules of the Game: This is a take-home Exam. The exam is due on Thursday,
ame Practice Exam Solutions 2018 Final Exam Chemistry 391 Structural Biochemistry Fall 2016 Do not open the exam until ready to begin! ules of the Game: This is a take-home Exam. The exam is due on Thursday,
Carboxylic Acids and their Derivatives I
 2302272 Org Chem II Part I Lecture 5 Carboxylic Acids and their Derivatives I Instructor: Dr. Tanatorn Khotavivattana E-mail: tanatorn.k@chula.ac.th Recommended Textbook: Chapter 20 in Organic Chemistry,
2302272 Org Chem II Part I Lecture 5 Carboxylic Acids and their Derivatives I Instructor: Dr. Tanatorn Khotavivattana E-mail: tanatorn.k@chula.ac.th Recommended Textbook: Chapter 20 in Organic Chemistry,
Docking Analysis of Darunavir as HIV Protease Inhibitors
 Journal of Computational Methods in Molecular Design, 2012, 2 (1):39-43 Scholars Research Library (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA Docking Analysis
Journal of Computational Methods in Molecular Design, 2012, 2 (1):39-43 Scholars Research Library (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA Docking Analysis
PAPER No. : 16 Bioorganic and biophysical chemistry MODULE No. : 25 Coenzyme-I Coenzyme A, TPP, B12 and biotin
 Subject Paper No and Title Module No and Title Module Tag 16, Bio organic and Bio physical chemistry 25, Coenzyme-I : Coenzyme A, TPP, B12 and CHE_P16_M25 TABLE OF CONTENTS 1. Learning Outcomes 2. Introduction
Subject Paper No and Title Module No and Title Module Tag 16, Bio organic and Bio physical chemistry 25, Coenzyme-I : Coenzyme A, TPP, B12 and CHE_P16_M25 TABLE OF CONTENTS 1. Learning Outcomes 2. Introduction
Fatty acid breakdown
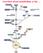 Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Chem 239 C: Chapter 19 Carboxylic Acid Derivatives Goals: Description, Nomenclature Reactivity of Carboxylic Acid Derivatives Nucleophylic Acyl
 Chem 239 C: Chapter 19 Carboxylic Acid Derivatives Goals: Description, Nomenclature Reactivity of Carboxylic Acid Derivatives Nucleophylic Acyl Substitution Acyl Chlorides Carboxylic Acid Anhydrides, Esters
Chem 239 C: Chapter 19 Carboxylic Acid Derivatives Goals: Description, Nomenclature Reactivity of Carboxylic Acid Derivatives Nucleophylic Acyl Substitution Acyl Chlorides Carboxylic Acid Anhydrides, Esters
Previous Class. Today. Spectrophotometry Spectrofluorimetry Radioactive procedures. ph dependence of Enzyme Catalysis (focus on pgs.
 Spectrophotometry Spectrofluorimetry Radioactive procedures Previous Class Today ph dependence of Enzyme Catalysis (focus on pgs. 169-176) ph Effects on Enzyme Activity Structural Considerations: Extreme
Spectrophotometry Spectrofluorimetry Radioactive procedures Previous Class Today ph dependence of Enzyme Catalysis (focus on pgs. 169-176) ph Effects on Enzyme Activity Structural Considerations: Extreme
1. Draw a standard line bond structure for compounds of the following molecular formulas:
 TF ame: LS1a Fall 06 Problem Set #1 Due Friday 9/29 at noon in your TF s drop box on the 2 nd floor of Science Center all questions including extra ones should be turned in 1. Draw a standard line bond
TF ame: LS1a Fall 06 Problem Set #1 Due Friday 9/29 at noon in your TF s drop box on the 2 nd floor of Science Center all questions including extra ones should be turned in 1. Draw a standard line bond
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
We will usually use the common name for an enzyme, such as carboxypeptidase, or chymotrypsin.
 Chapter 11 - Enzymatic Catalysis Introduction: In the late 1800's the Buchner brothers discovered that yeast extracts were capable of alcholic fermentation, thus refuting Pasteur s contention that intact
Chapter 11 - Enzymatic Catalysis Introduction: In the late 1800's the Buchner brothers discovered that yeast extracts were capable of alcholic fermentation, thus refuting Pasteur s contention that intact
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Protein Primer, Lumry I, Chapter 10, Enzyme structure, middle
 Protein Primer, Lumry I, Chapter 10, Enzyme structure, 5-15-03 10-1 Chapter 10. Enzyme structure A major feature of the trypsin family of proteases is the palindromic pattern taken by the B factors of
Protein Primer, Lumry I, Chapter 10, Enzyme structure, 5-15-03 10-1 Chapter 10. Enzyme structure A major feature of the trypsin family of proteases is the palindromic pattern taken by the B factors of
MITOCW watch?v=lcih8faydgk
 MITOCW watch?v=lcih8faydgk The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high-quality educational resources for free. To
MITOCW watch?v=lcih8faydgk The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high-quality educational resources for free. To
Chemical Biology - Chem 370 (3 credits)
 Chemical Biology - Chem 370 (3 credits) Spring Semester 2015 Instructors: Dr. Jeff Jones, Fulmer 406/408, 335-5983, jpj@wsu.edu Dr. ChulHee Kang, Fulmer 264, 509-335-1409, chkang@wsu.edu Class Meeting:
Chemical Biology - Chem 370 (3 credits) Spring Semester 2015 Instructors: Dr. Jeff Jones, Fulmer 406/408, 335-5983, jpj@wsu.edu Dr. ChulHee Kang, Fulmer 264, 509-335-1409, chkang@wsu.edu Class Meeting:
