Characterization of Lovastatin biosynthetic cluster proteins in Aspergillus terreus strain ATCC 20542
|
|
|
- Cordelia Wilkerson
- 6 years ago
- Views:
Transcription
1 Hypothesis Volume 6(7) Characterization of Lovastatin biosynthetic cluster proteins in Aspergillus terreus strain ATCC Thankaswamy Kosalai Subazini* & Gopal Ramesh Kumar Bioinformatics Lab, AU-KBC Research Centre, M.I.T campus of Anna University Chennai, Chromepet, Chennai , India; Thankaswamy Kosalai Subazini - subazinitk@au-kbc.org; Phone: , ; Fax: ; *Corresponding author Received December 21, 2010; Accepted February 22, 2011; Published June 23, 2011 Abstract: Aspergillus terreus is a filamentous ascomycota, which is prominent for its production of lovastatin, an antihypercholesterolemic drug. The commercial importance of lovastatin with annual sales of billions of dollars made us to focus on lovastatin biosynthetic cluster proteins. The analysis of these lovastatin biosynthetic cluster proteins with different perspectives such as physicochemical property, structure based analysis and functional studies were done to find out the role and function of every protein involved in the lovastatin biosynthesis pathway. Several computational tools are used to predict the physicochemical properties, secondary structural features, topology, patterns, domains and cellular location. There are 8 unidentified proteins in lovastatin biosynthetic cluster, in which 6 proteins have homologous partners, and annotation transfer is done based on the closely related homologous genes, and their structures are also modeled. The two other proteins that do not have homologous partners are predicted as PQ loop repeat protein that may be involved in glycosylation machinery and as thiolase-acyl activity by the integrated functional analysis approach. Keywords: Aspergillus terreus, Functional annotation, HMG CoA reductase inhibitor, Lovastatin, Sequence analysis, Structure analysis. Background: Aspergillus terreus, an important fungus is a major source for lovastatin, which treats cardiovascular disease by effectively inhibiting HMG CoA reductase activity, the rate limiting step in cholesterol biosynthesis. In addition, lovastatin is also involved in anti-inflammatory antioxidant activities and has the ability to inhibit proliferation and induce apoptosis in a variety of tumor cell lines [1]. Experimental evidences from literature reveal the presence of 18 proteins involved in lovastatin biosynthesis of A. terreus, ATCC Out of the 18 lovastatin biosynthetic cluster proteins it is found that 2 proteins are involved in regulatory mechanisms, 3 in transportation, 9 enzymes, 2 proteins of thiolases acyl-enzyme intermediate signature and PQ loop repeat, and 2 megas, [2] Lovastatin Nonaketide Synthase (LNKS) and Lovastatin Diketide Synthase (LDKS). The insilico analysis of physicochemical properties along with their topology and the functional analyses of these proteins are done by integrating standard functional annotation tools available online based on different methodologies. The final verification of functions of proteins is done by structure based analysis. Materials and Methodology: The protein sequences involved in lovastatin biosynthesis in ATCC strains are retrieved from NCBI ( database. Sequence analysis: Primary sequence analysis is done using sequence manipulation suite [3]. By using the suite the physicochemical properties like grand average of hydropathicity (GRAVY), ph, acidicity, basicity, Instability index, aliphatic index, extinction coefficient and molecular weight are calculated. Secondary structure prediction: The secondary structure of protein features such as helical content, beta sheet formations and turns, loops, and coil regions are predicted by GOR [4], available within Antherprot [5]. A number of trans-membrane helices are also identified from the sequences. Functional annotation: The functional annotation is done using different standard functional annotation tools available online such as KOGnitor [6] which incorporates orthologous information for eukaryotic sequence, [7], a tool that is based on protein families, ScanProsite [8] for identifying biologically significant sites, ProDom [9] for domain based analysis, and BLAST [10] for annotating function based on homology. SignalP [11] is used for determining whether the protein is secretory or non-secretory and ProtFun [12] for characterizing function by integrating post-translational and localization aspects of the protein. Tertiary structure prediction: The 3D structures of the proteins are predicted by using MODELLER9V8 [13]. The templates for proteins, which do not have structural homologs in Protein Data Bank (PDB) [14] (through blastp analysis) is determined using mgenthreader [15]. The model of the proteins was subjected to Swiss-PDB Viewer [16] for refinement (of side chains, problematic loops, removal of amino acid clashes and energy minimization). Finally, the refined models were subjected for validation. Backbone conformations are evaluated by Psi/Phi Ramachandran plot obtained from SAVS [17] and the Z-Score is obtained from Prosa web server [18]. Bioinformation 6(7): (2011) Biomedical Informatics
2 Results and Discussion: Primary sequence analysis: The primary sequence analysis revealed that the GRAVY indices of 13 lovastatin biosynthetic proteins are ranging from 0.03 to -0.5, which indicates they are hydrophilic and mostly predicted to be localized in extracellular and mitochondrial regions, the other 5 proteins (PQ loop repeat containing protein, HMG-CoA reductase, the 2 Permeases of the major facilitator superfamily, tricarboxylate transport protein) are hydrophobic (GRAVY values range from 0.05 to 0.6) and are localized in plasma membrane and basic in nature with ph value greater than 7. There are 2 more basic proteins, cytochrome monooxygenase and protein containing thiolase activity region. The rest of proteins are acidic in nature. The aliphatic index value of lovastatin cluster proteins ranges between 60 C and 110 C, indicating that the proteins are highly thermostable for wide temperature range. 11 proteins have instability index greater than 40 (Figure 1) and are unstable. is refined using Swiss PDB Viewer. The refined structures are validated and the scores obtained are shown in Table 2 (see Supplementary material). More than 80% of residues in modeled structures obey Ramachandran plot, and Prosa Z- scores are negative, which indicates the predicted structure correlated with the structural features of PDB. Homology modeling for the protein gi failed, because of the lack of homologous structures. The 3- D structure of modeled protein gi (Figure 3) with the template Beta- N-acetylhexosaminidase enzyme confirms that gi belongs to Glycosyl hydrolase family. Figure 2: Prediction of glycosylation sites for protein sequence with accession number gi using NetNGlyc Server. The vertical lines indicate the presence of 4 sequons crossing the threshold (horizontal line at 0.5) and 2 sequons crossing additional thresholds are predicted to be glycosylated. Figure 1: Physicochemical analysis on lovastatin biosynthetic cluster proteins. The graph shows the correlation observed in aliphatic, instability index and molecular weight of lovastatin biosynthetic cluster proteins. Secondary structure: The secondary structures of lovastatin biosynthetic cluster proteins revealed that all proteins in cluster have random coils dominated among secondary structure elements alpha helix, extended strand and beta turns. Topology predictions: In lovastatin biosynthetic cluster, there are 4 protein sequences with transmembrane helices, in which 2 protein sequences (efflux pump (gi ) and Major facilitator superfamily (gi )) are involved in membrane transport and has 12 transmembrane helices. The other two sequences with 6 transmembrane helices are enzymes predicted to be PQ loop repeat membrane protein (gi ) and HMG-CoA reductase (gi ). Functional analysis: The ScanProsite results showed that presence of protein sequence with accession number gi has the thiolases acyl-enzyme intermediate signature with the pattern [LIVM]-[NST]-{T}-x-C-[SAGLI]-[ST]-[SAG]- [LIVMFYNS]-x-[STAG]-[LIVM]-x(6)-[LIVM]. The BLAST results revealed that even though gi is highly homologous with immunoglobulin I-set domain protein it is also homologous with Glycosyl hydrolase family 67 N- terminus with 98% query coverage and E-value 9e-86. The presence of sequons NXS/T also confirms that gi codes for Glycosyl family protein. The sequons are verified by using Net-N-Glyc server [19] (Figure 2). Gi is distantly homologous to uroporphyrinogen decarboxylase with query coverage of 33% and E-value of 0.11 is predicted as a PQ-loop repeat family protein. The other proteins are having close homology and their functional prediction is done by annotation transfer from homologous sequences. The patterns and signature of lovastatin biosynthetic proteins along with their predicted functional roles of ATCC strain are shown in Table 1 (see Supplementary material). 3-D structure prediction and analysis: Homology based tertiary structure prediction for proteins are done using Modeller so as to verify the function predicted. Totally 5 models are generated for each of the individual protein and the best model of each protein Figure 3: Modelled structure for protein sequence with accession number gi The protein sequence of accession number gi is modeled using Modeller and the function is predicted as beta-nacetylhexosaminidases based on structural similarity. The N-glycosylation sites are represented by shaded regions in sequence and as stick in the 3-D structure. Conclusion: The insilico analysis of lovastatin biosynthetic cluster protein to understand their physicochemical, structural and functional properties are performed. Functional analysis of eight proteins performed by different function analysis tools based on domains, motifs and profiles improved the function annotation capability. The annotation revealed the presence of cis-aconitic acid decarboxylase of itaconic acid biosynthesis in lovastatin biosynthetic cluster, which shows that the two metabolites itaconic acid and lovastatin are related. Gi has thiolase activity domain and regulation of it may affect the energy level of the cells in metabolic state. From this analysis it is also found that the sequences with accession number gi and gi may be involved in glycosylation machinery as tailoring enzyme and can be used in preparing analogues of Lovastatin. The incorporation of these enzymes with cytochrome s into heterologous hosts may help in large scale production of lovastatin in microbial fermentations. Acknowledgment: Subazini TK is thankful to CSIR, Govt of India, New Delhi for the award of Senior Research fellowship (SRF). Bioinformation 6(7): (2011) Biomedical Informatics
3 References: [1] Martirosyan A et al. BMC Cancer : 103 [PMID: ] [2] Xie X et al. Chem Biol (11): 1161 [PMID: ] [3] Stothard P. Biotechniques : 1102 [PMID: ] [4] Garnier J et al. J Mol Biol (1): 97 [PMID: ] [5] Deleage G et al. Comput Biol Med (4): 259 [PMID: ] [6] Tatusov RL et al. BMC Bioinformatics : 41 [PMID: ] [7] Bateman A et al. Nucleic Acids Res : D138 [PMID: ] [8] Gasteiger E et al. Nucleic Acids Res : 3784 [PMID: ] [9] Servant F et al. Brief Bioinform : 246 [PMID: ] [10] Altschul SF et al. J Mol Biol : 403 [PMID: ] [11] Bendtsen JD et al. J Mol Biol : 783 [PMID: ] [12] Jensen LJ et al. J Mol Biol : 1257 [PMID: ] [13] Sali A & Blundell TL. J Mol Biol : 779 [PMID: ] [14] Berman HM et al. Acta Crystallogr D Biol Crystallogr : 899 [PMID: ] [15] Jones DT. J Mol Biol (4): 797 [PMID: ] [16] Kaplan W & Littlejohn TG. Brief Bioinform (2): 195 [PMID: ] [17] Lovell SC et al. Proteins (3): 437 [PMID: ] [18] Colovos C & Yeates TO. Protein Sci (9): 1511 [PMID: ] [19] Edited by P Kangueane Citation: Subazini & Kumar. Bioinformation 6(7): (2011) License statement: This is an open-access article, which permits unrestricted use, distribution, and reproduction in any medium, for non-commercial purposes, provided the original author and source are credited. Bioinformation 6(7): (2011) Biomedical Informatics
4 Supplementary material: Table 1: Functional analysis on lovastatin biosynthetic cluster. The analysis of lovastatin biosynthetic cluster proteins and their predicted domains and functional role are shown in this table. Accession number Domains Predicted from tool esterase Previous annotation Current Annotation Functional role gi gb AAD esterase gi gb AAD gi gb AAD cytochrome monooxygenase [Aspergillus terreus]. gi gb AAD34553 gi gb AAD34554 enoyl reductase GDSL-like Lipase/Acylhydrolase esterase Esterase Acts on ester bonds, carboxylesterase activity PQ loop repeat PQ-loop repeat containing protein Cytochrome, Blast cytochrome Cytochrome Membrane bound protein involved in Glycosylation machinary Monacolin J are likely to be catalyzed by cytochrome enzymes phospholipase/carboxyhydrolase KOGnitor oxidoreductase Involves in the hydrolysis of hydrophobic phosphonate ester compounds. Alcohol dehydrogenase GroESlike domain,zinc-binding dehydrogenase Zinc-binding oxidoreductase enoyl reductase Lovastatin Polyketide Enoyl Reductase (LovC) LovC with LNKS synthesizes dihydomonacolin J with successive ketoreduction and dehydration activity. gi gb AAD34555 transesterase gi gb AAD34556 hydroxymethylglutaryl-coenzyme A reductase Beta-lactamase class C and other penicillin binding proteins Beta-lactamase Sterol-sensing domain (SSD) profile, Hydroxymethylglutarylcoenzyme A reductases family profile, Hydroxymethylglutarylcoenzyme A reductases signature 1, 2,3 KOGnitor transesterase Transesterase (LovD) ScanProsite 3-hydroxy-3- methylglutarylcoenzyme A reductase 3-hydroxy-3- methylglutarylcoenzyme A reductase LovD catalyze last step of biosynthesis by transacylating acl group from LDKS to monacolin J to yield l t ti Involved in fatty acid metabolism gi gb AAD34557 regulatory protein Fungal Zn(2)-Cys(6) binuclear cluster domain,fungal specific transcription factor domain, Regulatory protein Fungal-type DNAbinding transcription factor Regulates a variety of cellular and metabolic processes. Zn(2)-C6 fungal-type DNAbinding domain profile, Zn(2)-C6 fungal-type DNA-binding domain signature, gi gb AAD34558 Major Facilitator Superfamily, Permeases of the major facilitator superfamily Possible roles for efflux pumps include removal of end products of metabolism and buffering the cytoplasm. located in the cytoplasmic region gi gb AAD34559 polyketide [Aspergillus terreus] Beta-ketoacyl, N- terminal domain,beta-ketoacyl, C-terminal domain,acyl transferase domain, polyketide Lovastatin diketide LDKS produce 2-methyl butyrate side chain. Acyl carrier protein phosphopantetheine domain profile, Beta-ketoacyl s active site, Sugar transport proteins signature 1, Bioinformation 6(7): (2011) Biomedical Informatics
5 gi gb AAD34560 Thiolases acyl-enzyme intermediate signature, ScanProsite Unknown Enzyme involved in Thiolase activity Enzyme involved in Thiolase activity gi gb AAD34561 regulatory protein gi gb AAD34562 Thiolase Fungal Zn(2)-Cys(6) binuclear cluster domain,fungal specific transcription factor domain Zn(2)-C6 fungal-type DNAbinding domain profile, Zn(2)-C6 fungal-type DNA-binding domain ProDom, regulatory protein Fungal-type DNAbinding transcription factor i t Mitochondrial carrier protein tricarboxylate transport protein, Solute carrier (Solcar) repeat profile, tricarboxylate transport protein, mitochondrial precursor Blast mitochondrial precursor Regulates a variety of cellular and metabolic processes. Involved in citrate-h+ /malate exchange. Provides a carbon source for fatty acid and sterol biosyntheses, and NAD+ for the glycolytic pathway. gi gb AAD34563 MmgE/PrpD family cis-aconitic acid decarboxylase Involved in itaconic acid biosynthesis gi gb AAD34564 gi gb AAD34565 cytochrome monooxygenase [Aspergillus terreus] Major Facilitator Superfamily, cytochrome monooxygenase, cytochrome monooxygenase Permeases of the major facilitator superfamily PHENYLACETATE HYDROXYLASE Transportation cytochrome monooxygenase gi gb AAD34566 Glycosidases family 67 N terminus Alpha-Glucuronidases are family 67 glycosidases that cleave the Alpha-1,2-glycosidic bond between 4-Omethyl-D-glucuronic acid and xylose units as part of an array of hemicellulosehydrolyzing enzymes gi gb AAD AF151722_1 lovastatin nonaketide Beta-ketoacyl, N- terminal domain,thiolase, N- terminal domain,beta-ketoacyl, C-terminal domain,acyl transferase domain hypothetical protein similar to polyketide lovastatin nonaketide LNKS is in iterative form and along with lovc; it synthesizes dihydomonacolin J Acyl carrier protein phosphopantetheine domain profile, Beta-ketoacyl s ti it Table 2: Structure validation report. Structure validation of different models for 6 proteins obtained from MODELLER Accession number RAMACHANDRAN PLOT TEMPLATE PROSA (Z-Score) Core (%) Allowed (%) General (%) Disallowed (%) Template Function gi YCD Yhr049w/FSH1 gi PW4 Glycerol-3-Phosphate Transporter gi OKC Mitochondrial ADP/ATP carrier gi HP0 Aminopeptidase N gi GFP multidrug transporter EmrD gi GH5 Beta-N-acetylhexosaminidase Bioinformation 6(7): (2011) Biomedical Informatics
LIPOPREDICT: Bacterial lipoprotein prediction server
 www.bioinformation.net Server Volume 8(8) LIPOPREDICT: Bacterial lipoprotein prediction server S Ramya Kumari, Kiran Kadam, Ritesh Badwaik & Valadi K Jayaraman* Centre for Development of Advanced Computing
www.bioinformation.net Server Volume 8(8) LIPOPREDICT: Bacterial lipoprotein prediction server S Ramya Kumari, Kiran Kadam, Ritesh Badwaik & Valadi K Jayaraman* Centre for Development of Advanced Computing
Lecture: 26 OXIDATION OF FATTY ACIDS
 Lecture: 26 OXIDATION OF FATTY ACIDS Fatty acids obtained by hydrolysis of fats undergo different oxidative pathways designated as alpha ( ), beta ( ) and omega ( ) pathways. -oxidation -Oxidation of fatty
Lecture: 26 OXIDATION OF FATTY ACIDS Fatty acids obtained by hydrolysis of fats undergo different oxidative pathways designated as alpha ( ), beta ( ) and omega ( ) pathways. -oxidation -Oxidation of fatty
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function
 Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
In Silico Sequence Analysis, Structure Prediction and Evaluation of Fatty Acid Synthase Protein (FAS) of Gallus gallus (Chicken) by Homology Modelling
 In Silico Sequence Analysis, Structure Prediction and Evaluation of Fatty Acid Synthase Protein (FAS) of Gallus gallus (Chicken) by Homology Modelling Aman Singh Bhalla 1, Roma Lal 2 1 Research Assistant,
In Silico Sequence Analysis, Structure Prediction and Evaluation of Fatty Acid Synthase Protein (FAS) of Gallus gallus (Chicken) by Homology Modelling Aman Singh Bhalla 1, Roma Lal 2 1 Research Assistant,
MITOCW watch?v=ddt1kusdoog
 MITOCW watch?v=ddt1kusdoog The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high-quality educational resources for free. To
MITOCW watch?v=ddt1kusdoog The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high-quality educational resources for free. To
Electron Transport Chain and Oxidative phosphorylation
 Electron Transport Chain and Oxidative phosphorylation So far we have discussed the catabolism involving oxidation of 6 carbons of glucose to CO 2 via glycolysis and CAC without any oxygen molecule directly
Electron Transport Chain and Oxidative phosphorylation So far we have discussed the catabolism involving oxidation of 6 carbons of glucose to CO 2 via glycolysis and CAC without any oxygen molecule directly
number Done by Corrected by Doctor Faisal Al-Khatibe
 number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
Metabolism Energy Pathways Biosynthesis. Catabolism Anabolism Enzymes
 Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes
 Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes The major site of acetoacetate and 3-hydorxybutyrate production is in the liver. 3-hydorxybutyrate is the
Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes The major site of acetoacetate and 3-hydorxybutyrate production is in the liver. 3-hydorxybutyrate is the
Tala Saleh. Razi Kittaneh ... Nayef Karadsheh
 Tala Saleh Razi Kittaneh... Nayef Karadsheh β-oxidation of Fatty Acids The oxidation of fatty acids occurs in 3 steps: Step 1: Activation of the Fatty acid FA + HS-CoA + ATP FA-CoA + AMP + PPi - The fatty
Tala Saleh Razi Kittaneh... Nayef Karadsheh β-oxidation of Fatty Acids The oxidation of fatty acids occurs in 3 steps: Step 1: Activation of the Fatty acid FA + HS-CoA + ATP FA-CoA + AMP + PPi - The fatty
Transcriptome wide identification and characterization of Starch Synthase enzyme in finger millet
 www.bioinformation.net Volume 14(7) Hypothesis Transcriptome wide identification and characterization of Starch Synthase enzyme in finger millet Rajhans Tyagi 1*, Apoorv Tiwari 2,3, Alok Kumar Gupta 4,
www.bioinformation.net Volume 14(7) Hypothesis Transcriptome wide identification and characterization of Starch Synthase enzyme in finger millet Rajhans Tyagi 1*, Apoorv Tiwari 2,3, Alok Kumar Gupta 4,
Ras and Cell Signaling Exercise
 Ras and Cell Signaling Exercise Learning Objectives In this exercise, you will use, a protein 3D- viewer, to explore: the structure of the Ras protein the active and inactive state of Ras and the amino
Ras and Cell Signaling Exercise Learning Objectives In this exercise, you will use, a protein 3D- viewer, to explore: the structure of the Ras protein the active and inactive state of Ras and the amino
biochem480 [Autumn2014] Enzyme bio-informatics project
![biochem480 [Autumn2014] Enzyme bio-informatics project biochem480 [Autumn2014] Enzyme bio-informatics project](/thumbs/91/104514013.jpg) biochem480 [Autumn2014] Enzyme bio-informatics project Edit out any stuff in italics for the final version Student Name: IUBSystematic Name: 3-hydroxy-3-methylglutaryl-CoA reductase Other protein Names:
biochem480 [Autumn2014] Enzyme bio-informatics project Edit out any stuff in italics for the final version Student Name: IUBSystematic Name: 3-hydroxy-3-methylglutaryl-CoA reductase Other protein Names:
Points 1. Following is the overall reaction catalyzed by the Calvin-Benson cycle:
 BCH 4054 February 22, 2002 HOUR TEST 2 NAME_ Points 1. Following is the overall reaction catalyzed by the Calvin-Benson cycle: CO 2 + 3ATP + 2NADPH 1/3 glyceraldehyde-3-p + 3ADP + 2NADP + Give the structures
BCH 4054 February 22, 2002 HOUR TEST 2 NAME_ Points 1. Following is the overall reaction catalyzed by the Calvin-Benson cycle: CO 2 + 3ATP + 2NADPH 1/3 glyceraldehyde-3-p + 3ADP + 2NADP + Give the structures
BIOL 158: BIOLOGICAL CHEMISTRY II
 BIOL 158: BIOLOGICAL CHEMISTRY II Lecture 5: Vitamins and Coenzymes Lecturer: Christopher Larbie, PhD Introduction Cofactors bind to the active site and assist in the reaction mechanism Apoenzyme is an
BIOL 158: BIOLOGICAL CHEMISTRY II Lecture 5: Vitamins and Coenzymes Lecturer: Christopher Larbie, PhD Introduction Cofactors bind to the active site and assist in the reaction mechanism Apoenzyme is an
BY: RASAQ NURUDEEN OLAJIDE
 BY: RASAQ NURUDEEN OLAJIDE LECTURE CONTENT INTRODUCTION CITRIC ACID CYCLE (T.C.A) PRODUCTION OF ACETYL CoA REACTIONS OF THE CITIRC ACID CYCLE THE AMPHIBOLIC NATURE OF THE T.C.A CYCLE THE GLYOXYLATE CYCLE
BY: RASAQ NURUDEEN OLAJIDE LECTURE CONTENT INTRODUCTION CITRIC ACID CYCLE (T.C.A) PRODUCTION OF ACETYL CoA REACTIONS OF THE CITIRC ACID CYCLE THE AMPHIBOLIC NATURE OF THE T.C.A CYCLE THE GLYOXYLATE CYCLE
Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids
 www.bioinformation.net Volume 12(12) Hypothesis Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids Ayub Shaik 1, Vishnu Thumma 2, Aruna Kumari Kotha 2, Sandhya
www.bioinformation.net Volume 12(12) Hypothesis Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids Ayub Shaik 1, Vishnu Thumma 2, Aruna Kumari Kotha 2, Sandhya
Structure modeling and antidiabetic activity of a seed protein of Momordica charantia in non-obese diabetic (NOD) mice
 www.bioinformation.net Hypothesis Volume 9(15) Structure modeling and antidiabetic activity of a seed protein of Momordica charantia in non-obese diabetic (NOD) mice Gagan Chhabra & Aparna Dixit* Gene
www.bioinformation.net Hypothesis Volume 9(15) Structure modeling and antidiabetic activity of a seed protein of Momordica charantia in non-obese diabetic (NOD) mice Gagan Chhabra & Aparna Dixit* Gene
CHAPTER 6 METABOLIC PATHWAY RECONSTRUCTION
 66 CHAPTER 6 METABOLIC PATHWAY RECONSTRUCTION 6.1 GENERAL A metabolic pathway is a series of biochemical reactions that occur within a cell and these chemical reactions are catalyzed by certain enzymes.
66 CHAPTER 6 METABOLIC PATHWAY RECONSTRUCTION 6.1 GENERAL A metabolic pathway is a series of biochemical reactions that occur within a cell and these chemical reactions are catalyzed by certain enzymes.
Insight into the mechanism of lipids binding and uptake by CD36 receptor
 www.bioinformation.net Hypothesis Volume 11(6) Insight into the mechanism of lipids binding and uptake by CD36 receptor Zineb Tarhda*, Azeddine Ibrahimi Biotechnology lab (MedBiotech), Faculté de Médecine
www.bioinformation.net Hypothesis Volume 11(6) Insight into the mechanism of lipids binding and uptake by CD36 receptor Zineb Tarhda*, Azeddine Ibrahimi Biotechnology lab (MedBiotech), Faculté de Médecine
!"#$%&' (#%) /&'(2+"( /&3&4,, ! " #$% - &'()!% *-sheet -(!-Helix - &'(&') +,(-. - &'()&+) /&%.(0&+(! - &'(1&2%( Basic amino acids
 Basic amino acids pk ~ 10.5 pk ~ 12.5 pk ~ 6.0 Polar 25!"#$%&' (#%)! " #$% - &'()!% *-sheet -(!-Helix - &'(&') +,(-. - &'()&+) /&%.(0&+(! - &'(1&2%( /&'(2+"( /&3&4,, :++55 ('&.! 6($.(" 40 > 3&4,, ('&.!
Basic amino acids pk ~ 10.5 pk ~ 12.5 pk ~ 6.0 Polar 25!"#$%&' (#%)! " #$% - &'()!% *-sheet -(!-Helix - &'(&') +,(-. - &'()&+) /&%.(0&+(! - &'(1&2%( /&'(2+"( /&3&4,, :++55 ('&.! 6($.(" 40 > 3&4,, ('&.!
Metabolism. Chapter 8 Microbial Metabolism. Metabolic balancing act. Catabolism Anabolism Enzymes. Topics. Metabolism Energy Pathways Biosynthesis
 Chapter 8 Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis Catabolism Anabolism Enzymes Metabolism 1 2 Metabolic balancing act Catabolism and anabolism simple model Catabolism Enzymes
Chapter 8 Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis Catabolism Anabolism Enzymes Metabolism 1 2 Metabolic balancing act Catabolism and anabolism simple model Catabolism Enzymes
Metabolism. Topic 11&12 (ch8) Microbial Metabolism. Metabolic Balancing Act. Topics. Catabolism Anabolism Enzymes
 Topic 11&12 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic Balancing Act Catabolism Enzymes involved in breakdown of complex
Topic 11&12 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic Balancing Act Catabolism Enzymes involved in breakdown of complex
BIOSYNTHESIS OF FATTY ACIDS. doc. Ing. Zenóbia Chavková, CSc.
 BIOSYNTHESIS OF FATTY ACIDS doc. Ing. Zenóbia Chavková, CSc. The pathway for the of FAs is not the reversal of the oxidation pathway Both pathways are separated within different cellular compartments In
BIOSYNTHESIS OF FATTY ACIDS doc. Ing. Zenóbia Chavková, CSc. The pathway for the of FAs is not the reversal of the oxidation pathway Both pathways are separated within different cellular compartments In
Enzymes what are they?
 Topic 11 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic balancing act Catabolism Enzymes involved in breakdown of complex
Topic 11 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic balancing act Catabolism Enzymes involved in breakdown of complex
Structural characterization of a flavonoid glycosyltransferase from Withania somnifera
 www.bioinformation.net Hypothesis Volume 8(19) Structural characterization of a flavonoid glycosyltransferase from Withania somnifera Santosh Kumar Ramachandra Jadhav 1, Krunal Arvind Patel 1, Bhushan
www.bioinformation.net Hypothesis Volume 8(19) Structural characterization of a flavonoid glycosyltransferase from Withania somnifera Santosh Kumar Ramachandra Jadhav 1, Krunal Arvind Patel 1, Bhushan
Fatty acid breakdown
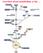 Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Bioinformatics for molecular biology
 Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
A look at macromolecules (Text pages 38-54) What is the typical chemical composition of a cell? (Source of figures to right: Madigan et al.
 A look at macromolecules (Text pages 38-54) What is the typical chemical composition of a cell? (Source of figures to right: Madigan et al. 2002 Chemical Bonds Ionic Electron-negativity differences cause
A look at macromolecules (Text pages 38-54) What is the typical chemical composition of a cell? (Source of figures to right: Madigan et al. 2002 Chemical Bonds Ionic Electron-negativity differences cause
Mrub_2765 is the version of E. coli FabZ in Meiothermus ruber, while Mrub_0266 is the version of E. coli FabI in Meiothermus ruber
 Augustana College Augustana Digital Commons Meiothermus ruber Genome Analysis Project Biology 2-2016 Mrub_2765 is the version of E. coli FabZ in Meiothermus ruber, while Mrub_0266 is the version of E.
Augustana College Augustana Digital Commons Meiothermus ruber Genome Analysis Project Biology 2-2016 Mrub_2765 is the version of E. coli FabZ in Meiothermus ruber, while Mrub_0266 is the version of E.
23.1 Lipid Metabolism in Animals. Chapter 23. Micelles Lipid Metabolism in. Animals. Overview of Digestion Lipid Metabolism in
 Denniston Topping Caret Copyright! The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Chapter 23 Fatty Acid Metabolism Triglycerides (Tgl) are emulsified into fat droplets
Denniston Topping Caret Copyright! The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Chapter 23 Fatty Acid Metabolism Triglycerides (Tgl) are emulsified into fat droplets
2-more complex molecules (fatty acyl esters) as triacylglycerols.
 ** Fatty acids exist in two forms:- 1-free fatty acids (unesterified) 2-more complex molecules (fatty acyl esters) as triacylglycerols. ** most tissues might use fatty acids as source of energy during
** Fatty acids exist in two forms:- 1-free fatty acids (unesterified) 2-more complex molecules (fatty acyl esters) as triacylglycerols. ** most tissues might use fatty acids as source of energy during
Carbohydrate Metabolism I
 Carbohydrate Metabolism I Outline Glycolysis Stages of glycolysis Regulation of Glycolysis Carbohydrate Metabolism Overview Enzyme Classification Dehydrogenase - oxidizes substrate using cofactors as
Carbohydrate Metabolism I Outline Glycolysis Stages of glycolysis Regulation of Glycolysis Carbohydrate Metabolism Overview Enzyme Classification Dehydrogenase - oxidizes substrate using cofactors as
Supplementary Information
 Supplementary Information Assembly and Clustering of Natural Antibiotics Guides Target Identification Chad W. Johnston 1,2, Michael A. Skinnider 1,2, Chris A. Dejong 1,2, Philip N. Rees 1,2, Gregory M.
Supplementary Information Assembly and Clustering of Natural Antibiotics Guides Target Identification Chad W. Johnston 1,2, Michael A. Skinnider 1,2, Chris A. Dejong 1,2, Philip N. Rees 1,2, Gregory M.
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
The three important structural features of proteins:
 The three important structural features of proteins: a. Primary (1 o ) The amino acid sequence (coded by genes) b. Secondary (2 o ) The interaction of amino acids that are close together or far apart in
The three important structural features of proteins: a. Primary (1 o ) The amino acid sequence (coded by genes) b. Secondary (2 o ) The interaction of amino acids that are close together or far apart in
Three-dimensional (3D) structure prediction of the American and African oil-palms β-ketoacyl-[acp] synthase-ii protein by comparative modelling
![Three-dimensional (3D) structure prediction of the American and African oil-palms β-ketoacyl-[acp] synthase-ii protein by comparative modelling Three-dimensional (3D) structure prediction of the American and African oil-palms β-ketoacyl-[acp] synthase-ii protein by comparative modelling](/thumbs/87/96045358.jpg) www.bioinformation.net Hypothesis Volume 10(3) Three-dimensional (3D) structure prediction of the American and African oil-palms β-ketoacyl-[acp] synthase-ii protein by comparative modelling Edina Wang,
www.bioinformation.net Hypothesis Volume 10(3) Three-dimensional (3D) structure prediction of the American and African oil-palms β-ketoacyl-[acp] synthase-ii protein by comparative modelling Edina Wang,
Secondary Structure North 72nd Street, Wauwatosa, WI Phone: (414) Fax: (414) dmoleculardesigns.com
 Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
Structure prediction and functional characterization of secondary metabolite proteins of Ocimum
 www.bioinformation.net Hypothesis Volume 6(8) Structure prediction and functional characterization of secondary metabolite proteins of Ocimum Sudeep Roy*, Nidhi Maheshwari, Rashi Chauhan, Naresh Kumar
www.bioinformation.net Hypothesis Volume 6(8) Structure prediction and functional characterization of secondary metabolite proteins of Ocimum Sudeep Roy*, Nidhi Maheshwari, Rashi Chauhan, Naresh Kumar
Section 1 Lecture 1- Origins of Life Life probably started by Hydrothermal Vents.
 Section 1 Lecture 1- Origins of Life Life probably started by Hydrothermal Vents. Photosynthesis originated around 3GA, as cells figured out how to fix CO2 and release O2. Eukaryotes originates 1.5-2.5
Section 1 Lecture 1- Origins of Life Life probably started by Hydrothermal Vents. Photosynthesis originated around 3GA, as cells figured out how to fix CO2 and release O2. Eukaryotes originates 1.5-2.5
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Ahmad Ulnar. Faisal Nimri ... Dr.Faisal
 24 Ahmad Ulnar Faisal Nimri... Dr.Faisal Fatty Acid Synthesis - Occurs mainly in the Liver (to store excess carbohydrates as triacylglycerols(fat)) and in lactating mammary glands (for the production of
24 Ahmad Ulnar Faisal Nimri... Dr.Faisal Fatty Acid Synthesis - Occurs mainly in the Liver (to store excess carbohydrates as triacylglycerols(fat)) and in lactating mammary glands (for the production of
Chapter 8. Metabolism. Topics in lectures 15 and 16. Chemical foundations Catabolism Biosynthesis
 Chapter 8 Topics in lectures 15 and 16 Metabolism Chemical foundations Catabolism Biosynthesis 1 Metabolism Chemical Foundations Enzymes REDOX Catabolism Pathways Anabolism Principles and pathways 2 Enzymes
Chapter 8 Topics in lectures 15 and 16 Metabolism Chemical foundations Catabolism Biosynthesis 1 Metabolism Chemical Foundations Enzymes REDOX Catabolism Pathways Anabolism Principles and pathways 2 Enzymes
P450 CYCLE. All P450s follow the same catalytic cycle of;
 P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
Cellular Respiration Harvesting Chemical Energy ATP
 Cellular Respiration Harvesting Chemical Energy ATP 2006-2007 What s the point? The point is to make ATP! ATP 2006-2007 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats,
Cellular Respiration Harvesting Chemical Energy ATP 2006-2007 What s the point? The point is to make ATP! ATP 2006-2007 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats,
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site.
 Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Find this material useful? You can help our team to keep this site up and bring you even more content consider donating via the link on our site. Still having trouble understanding the material? Check
Secondary Structure. by hydrogen bonds
 Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
FATTY ACID SYNTHESIS
 FATTY ACID SYNTHESIS Malonyl- CoA inhibits Carni1ne Palmitoyl Transferase I. Malonyl- CoA is a precursor for fa=y acid synthesis. Malonyl- CoA is produced from acetyl- CoA by the enzyme Acetyl- CoA Carboxylase.
FATTY ACID SYNTHESIS Malonyl- CoA inhibits Carni1ne Palmitoyl Transferase I. Malonyl- CoA is a precursor for fa=y acid synthesis. Malonyl- CoA is produced from acetyl- CoA by the enzyme Acetyl- CoA Carboxylase.
Cellular Respiration Stage 2 & 3. Glycolysis is only the start. Cellular respiration. Oxidation of Pyruvate Krebs Cycle.
 Cellular Respiration Stage 2 & 3 Oxidation of Pyruvate Krebs Cycle AP 2006-2007 Biology Glycolysis is only the start Glycolysis glucose pyruvate 6C 2x 3C Pyruvate has more energy to yield 3 more C to strip
Cellular Respiration Stage 2 & 3 Oxidation of Pyruvate Krebs Cycle AP 2006-2007 Biology Glycolysis is only the start Glycolysis glucose pyruvate 6C 2x 3C Pyruvate has more energy to yield 3 more C to strip
Syllabus for BASIC METABOLIC PRINCIPLES
 Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
3.0 Fatty Acids and Polyketides : Biosynthesis and Engineering
 3.0 Fatty Acids and Polyketides : Biosynthesis and Engineering Chemical tructure Chemical Formula: C 16 32 2 Palmitic Acid = LAD aponification Na Na = AP Present in animal and plant fats as triacyl glycerols
3.0 Fatty Acids and Polyketides : Biosynthesis and Engineering Chemical tructure Chemical Formula: C 16 32 2 Palmitic Acid = LAD aponification Na Na = AP Present in animal and plant fats as triacyl glycerols
Oxidation of Long Chain Fatty Acids
 Oxidation of Long Chain Fatty Acids Dr NC Bird Oxidation of long chain fatty acids is the primary source of energy supply in man and animals. Hibernating animals utilise fat stores to maintain body heat,
Oxidation of Long Chain Fatty Acids Dr NC Bird Oxidation of long chain fatty acids is the primary source of energy supply in man and animals. Hibernating animals utilise fat stores to maintain body heat,
Ch. 9 Cellular Respiration Stage 2 & 3: Oxidation of Pyruvate Krebs Cycle
 Ch. 9 Cellular Respiration Stage 2 & 3: Oxidation of Pyruvate Krebs Cycle 2006-2007 Glycolysis is only the start Glycolysis glucose pyruvate 6C Pyruvate has more energy to yield 3 more C to strip off (to
Ch. 9 Cellular Respiration Stage 2 & 3: Oxidation of Pyruvate Krebs Cycle 2006-2007 Glycolysis is only the start Glycolysis glucose pyruvate 6C Pyruvate has more energy to yield 3 more C to strip off (to
LIPID METABOLISM
 LIPID METABOLISM LIPOGENESIS LIPOGENESIS LIPOGENESIS FATTY ACID SYNTHESIS DE NOVO FFA in the blood come from :- (a) Dietary fat (b) Dietary carbohydrate/protein in excess of need FA TAG Site of synthesis:-
LIPID METABOLISM LIPOGENESIS LIPOGENESIS LIPOGENESIS FATTY ACID SYNTHESIS DE NOVO FFA in the blood come from :- (a) Dietary fat (b) Dietary carbohydrate/protein in excess of need FA TAG Site of synthesis:-
Module No. # 01 Lecture No. # 19 TCA Cycle
 Biochemical Engineering Prof. Dr. Rintu Banerjee Department of Agricultural and Food Engineering Asst. Prof. Dr. Saikat Chakraborty Department of Chemical Engineering Indian Institute of Technology, Kharagpur
Biochemical Engineering Prof. Dr. Rintu Banerjee Department of Agricultural and Food Engineering Asst. Prof. Dr. Saikat Chakraborty Department of Chemical Engineering Indian Institute of Technology, Kharagpur
Citrate Cycle. Lecture 28. Key Concepts. The Citrate Cycle captures energy using redox reactions
 Citrate Cycle Lecture 28 Key Concepts The Citrate Cycle captures energy using redox reactions Eight reactions of the Citrate Cycle Key control points in the Citrate Cycle regulate metabolic flux What role
Citrate Cycle Lecture 28 Key Concepts The Citrate Cycle captures energy using redox reactions Eight reactions of the Citrate Cycle Key control points in the Citrate Cycle regulate metabolic flux What role
Lipid Metabolism. Catabolism Overview
 Lipid Metabolism Pratt & Cornely, Chapter 17 Catabolism Overview Lipids as a fuel source from diet Beta oxidation Mechanism ATP production Ketone bodies as fuel 1 High energy More reduced Little water
Lipid Metabolism Pratt & Cornely, Chapter 17 Catabolism Overview Lipids as a fuel source from diet Beta oxidation Mechanism ATP production Ketone bodies as fuel 1 High energy More reduced Little water
Binding of activated isoniazid with acetyl-coa carboxylase from Mycobacterium tuberculosis
 www.bioinformation.net Hypothesis Volume 7(3) Binding of activated isoniazid with acetyl-coa carboxylase from Mycobacterium tuberculosis Ameeruddin Nusrath Unissa 1 *, Subramanian Sudha 2, Nagamiah Selvakumar
www.bioinformation.net Hypothesis Volume 7(3) Binding of activated isoniazid with acetyl-coa carboxylase from Mycobacterium tuberculosis Ameeruddin Nusrath Unissa 1 *, Subramanian Sudha 2, Nagamiah Selvakumar
BIOLOGY 103 Spring 2001 MIDTERM LAB SECTION
 BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
BIOLOGY 103 Spring 2001 MIDTERM NAME KEY LAB SECTION ID# (last four digits of SS#) STUDENT PLEASE READ. Do not put yourself at a disadvantage by revealing the content of this exam to your classmates. Your
A HLA-DRB supertype chart with potential overlapping peptide binding function
 A HLA-DRB supertype chart with potential overlapping peptide binding function Arumugam Mohanapriya 1,2, Satish Nandagond 1, Paul Shapshak 3, Uma Kangueane 1, Pandjassarame Kangueane 1, 2 * 1 Biomedical
A HLA-DRB supertype chart with potential overlapping peptide binding function Arumugam Mohanapriya 1,2, Satish Nandagond 1, Paul Shapshak 3, Uma Kangueane 1, Pandjassarame Kangueane 1, 2 * 1 Biomedical
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A
 Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biochemistry: A Short Course
 Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 28 Fatty Acid Synthesis 2013 W. H. Freeman and Company Chapter 28 Outline 1. The first stage of fatty acid synthesis is transfer
Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 28 Fatty Acid Synthesis 2013 W. H. Freeman and Company Chapter 28 Outline 1. The first stage of fatty acid synthesis is transfer
Summary of fatty acid synthesis
 Lipid Metabolism, part 2 1 Summary of fatty acid synthesis 8 acetyl CoA + 14 NADPH + 14 H+ + 7 ATP palmitic acid (16:0) + 8 CoA + 14 NADP + + 7 ADP + 7 Pi + 7 H20 1. The major suppliers of NADPH for fatty
Lipid Metabolism, part 2 1 Summary of fatty acid synthesis 8 acetyl CoA + 14 NADPH + 14 H+ + 7 ATP palmitic acid (16:0) + 8 CoA + 14 NADP + + 7 ADP + 7 Pi + 7 H20 1. The major suppliers of NADPH for fatty
BCH 4054 Spring 2001 Chapter 24 Lecture Notes
 BCH 4054 Spring 2001 Chapter 24 Lecture Notes 1 Chapter 24 Fatty Acid Catabolism 2 Fatty Acids as Energy Source Triglycerides yield 37 kj/g dry weight Protein 17 kj/g Glycogen 16 kj/g (even less wet weight)
BCH 4054 Spring 2001 Chapter 24 Lecture Notes 1 Chapter 24 Fatty Acid Catabolism 2 Fatty Acids as Energy Source Triglycerides yield 37 kj/g dry weight Protein 17 kj/g Glycogen 16 kj/g (even less wet weight)
number Done by Corrected by Doctor F. Al-Khateeb
 number 23 Done by A. Rawajbeh Corrected by Doctor F. Al-Khateeb Ketone bodies Ketone bodies are used by the peripheral tissues like the skeletal and cardiac muscles, where they are the preferred source
number 23 Done by A. Rawajbeh Corrected by Doctor F. Al-Khateeb Ketone bodies Ketone bodies are used by the peripheral tissues like the skeletal and cardiac muscles, where they are the preferred source
BCM 221 LECTURES OJEMEKELE O.
 BCM 221 LECTURES BY OJEMEKELE O. OUTLINE INTRODUCTION TO LIPID CHEMISTRY STORAGE OF ENERGY IN ADIPOCYTES MOBILIZATION OF ENERGY STORES IN ADIPOCYTES KETONE BODIES AND KETOSIS PYRUVATE DEHYDROGENASE COMPLEX
BCM 221 LECTURES BY OJEMEKELE O. OUTLINE INTRODUCTION TO LIPID CHEMISTRY STORAGE OF ENERGY IN ADIPOCYTES MOBILIZATION OF ENERGY STORES IN ADIPOCYTES KETONE BODIES AND KETOSIS PYRUVATE DEHYDROGENASE COMPLEX
These are example problems, which are similar to those you may see on the final exam.
 MCB102 / Metabolism Problem Set #3 Spring 2008 These are example problems, which are similar to those you may see on the final exam. QUESTION 1: /. Circle the correct answer, but if the answer is provide
MCB102 / Metabolism Problem Set #3 Spring 2008 These are example problems, which are similar to those you may see on the final exam. QUESTION 1: /. Circle the correct answer, but if the answer is provide
Figure 1 Original Advantages of biological reactions being catalyzed by enzymes:
 Enzyme basic concepts, Enzyme Regulation I III Carmen Sato Bigbee, Ph.D. Objectives: 1) To understand the bases of enzyme catalysis and the mechanisms of enzyme regulation. 2) To understand the role of
Enzyme basic concepts, Enzyme Regulation I III Carmen Sato Bigbee, Ph.D. Objectives: 1) To understand the bases of enzyme catalysis and the mechanisms of enzyme regulation. 2) To understand the role of
Chapter 10. 이화작용 : 에너지방출과보존 (Catabolism: Energy Release and Conservation)
 Chapter 10 이화작용 : 에너지방출과보존 (Catabolism: Energy Release and Conservation) 1 Fueling Processes Respiration 1 Most respiration involves use of an electron transport chain As electrons pass through the electron
Chapter 10 이화작용 : 에너지방출과보존 (Catabolism: Energy Release and Conservation) 1 Fueling Processes Respiration 1 Most respiration involves use of an electron transport chain As electrons pass through the electron
Palindromes drive the re-assortment in Influenza A
 www.bioinformation.net Hypothesis Volume 7(3) Palindromes drive the re-assortment in Influenza A Abdullah Zubaer 1, 2 * & Simrika Thapa 1, 2 1Swapnojaatra Bioresearch Laboratory, DataSoft Systems, Dhaka-15,
www.bioinformation.net Hypothesis Volume 7(3) Palindromes drive the re-assortment in Influenza A Abdullah Zubaer 1, 2 * & Simrika Thapa 1, 2 1Swapnojaatra Bioresearch Laboratory, DataSoft Systems, Dhaka-15,
Citrate Cycle Supplemental Reading
 Citrate Cycle Supplemental Reading Key Concepts - The Citrate Cycle captures energy using redox reactions - Eight enzymatic reactions of the Citrate Cycle - Key control points in the citrate cycle regulate
Citrate Cycle Supplemental Reading Key Concepts - The Citrate Cycle captures energy using redox reactions - Eight enzymatic reactions of the Citrate Cycle - Key control points in the citrate cycle regulate
In silico Analysis and 3D Modelling of SORD Protein in Diabetic Retinopathy
 Available online at www.scholarsresearchlibrary.com J. Comput. Method. Mol. Design, 2011, 1 (4):22-27 (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA In silico Analysis
Available online at www.scholarsresearchlibrary.com J. Comput. Method. Mol. Design, 2011, 1 (4):22-27 (http://scholarsresearchlibrary.com/archive.html) ISSN : 2231-3176 CODEN (USA): JCMMDA In silico Analysis
Respiration. Respiration. Respiration. How Cells Harvest Energy. Chapter 7
 How Cells Harvest Energy Chapter 7 Organisms can be classified based on how they obtain energy: autotrophs: are able to produce their own organic molecules through photosynthesis heterotrophs: live on
How Cells Harvest Energy Chapter 7 Organisms can be classified based on how they obtain energy: autotrophs: are able to produce their own organic molecules through photosynthesis heterotrophs: live on
Biochemistry Sheet 27 Fatty Acid Synthesis Dr. Faisal Khatib
 Page1 بسم رلاهللا On Thursday, we discussed the synthesis of fatty acids and its regulation. We also went on to talk about the synthesis of Triacylglycerol (TAG). Last time, we started talking about the
Page1 بسم رلاهللا On Thursday, we discussed the synthesis of fatty acids and its regulation. We also went on to talk about the synthesis of Triacylglycerol (TAG). Last time, we started talking about the
III. Metabolism The Citric Acid Cycle
 Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism The Citric Acid Cycle Slide 1 The Eight Steps of the Citric Acid Cycle Enzymes: 4 dehydrogenases (2 decarboxylation) 3
Department of Chemistry and Biochemistry University of Lethbridge III. Metabolism The Citric Acid Cycle Slide 1 The Eight Steps of the Citric Acid Cycle Enzymes: 4 dehydrogenases (2 decarboxylation) 3
Roles of Lipids. principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular
 Roles of Lipids principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular = Oxidation of fatty acids Central energy-yielding pathway in animals. O
Roles of Lipids principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular = Oxidation of fatty acids Central energy-yielding pathway in animals. O
Ch 07. Microbial Metabolism
 Ch 07 Microbial Metabolism SLOs Differentiate between metabolism, catabolism, and anabolism. Fully describe the structure and function of enzymes. Differentiate between constitutive and regulated enzymes.
Ch 07 Microbial Metabolism SLOs Differentiate between metabolism, catabolism, and anabolism. Fully describe the structure and function of enzymes. Differentiate between constitutive and regulated enzymes.
A cell has enough ATP to last for about three seconds.
 Energy Transformation: Cellular Respiration Outline 1. Energy and carbon sources in living cells 2. Sources of cellular ATP 3. Turning chemical energy of covalent bonds between C-C into energy for cellular
Energy Transformation: Cellular Respiration Outline 1. Energy and carbon sources in living cells 2. Sources of cellular ATP 3. Turning chemical energy of covalent bonds between C-C into energy for cellular
Fatty acids synthesis
 Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 30, 2014
 Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 30, 2014 PHRM 836 Exam I - 1 Name: Instructions 1. Check your exam to make certain that it has 10 pages including this cover
Examination I PHRM 836 Biochemistry for Pharmaceutical Sciences II September 30, 2014 PHRM 836 Exam I - 1 Name: Instructions 1. Check your exam to make certain that it has 10 pages including this cover
Details of Organic Chem! Date. Carbon & The Molecular Diversity of Life & The Structure & Function of Macromolecules
 Details of Organic Chem! Date Carbon & The Molecular Diversity of Life & The Structure & Function of Macromolecules Functional Groups, I Attachments that replace one or more of the hydrogens bonded to
Details of Organic Chem! Date Carbon & The Molecular Diversity of Life & The Structure & Function of Macromolecules Functional Groups, I Attachments that replace one or more of the hydrogens bonded to
a) The statement is true for X = 400, but false for X = 300; b) The statement is true for X = 300, but false for X = 200;
 1. Consider the following statement. To produce one molecule of each possible kind of polypeptide chain, X amino acids in length, would require more atoms than exist in the universe. Given the size of
1. Consider the following statement. To produce one molecule of each possible kind of polypeptide chain, X amino acids in length, would require more atoms than exist in the universe. Given the size of
AP Biology. Proteins. Proteins. Proteins. Amino acids H C OH H R. Effect of different R groups: Polar amino acids polar or charged & hydrophilic
 Most structurally & functionally diverse group : involved in almost everything enzymes (pepsin, DNA polymerase) structure (keratin, collagen) carriers & transport (, aquaporin) cell communication signals
Most structurally & functionally diverse group : involved in almost everything enzymes (pepsin, DNA polymerase) structure (keratin, collagen) carriers & transport (, aquaporin) cell communication signals
NBCE Mock Board Questions Biochemistry
 1. Fluid mosaic describes. A. Tertiary structure of proteins B. Ribosomal subunits C. DNA structure D. Plasma membrane structure NBCE Mock Board Questions Biochemistry 2. Where in the cell does beta oxidation
1. Fluid mosaic describes. A. Tertiary structure of proteins B. Ribosomal subunits C. DNA structure D. Plasma membrane structure NBCE Mock Board Questions Biochemistry 2. Where in the cell does beta oxidation
Oxidative Phosphorylation
 Electron Transport Chain (overview) The NADH and FADH 2, formed during glycolysis, β- oxidation and the TCA cycle, give up their electrons to reduce molecular O 2 to H 2 O. Electron transfer occurs through
Electron Transport Chain (overview) The NADH and FADH 2, formed during glycolysis, β- oxidation and the TCA cycle, give up their electrons to reduce molecular O 2 to H 2 O. Electron transfer occurs through
Chapter 26 Biochemistry 5th edition. phospholipids. Sphingolipids. Cholesterol. db=books&itool=toolbar
 http://www.ncbi.nlm.nih.gov/sites/entrez? db=books&itool=toolbar 1 The surface of a soap bubble is a bilayer formed by detergent molecules 2 Chapter 26 Biochemistry 5th edition phospholipids Sphingolipids
http://www.ncbi.nlm.nih.gov/sites/entrez? db=books&itool=toolbar 1 The surface of a soap bubble is a bilayer formed by detergent molecules 2 Chapter 26 Biochemistry 5th edition phospholipids Sphingolipids
TCA CYCLE (Citric Acid Cycle)
 TCA CYCLE (Citric Acid Cycle) TCA CYCLE The Citric Acid Cycle is also known as: Kreb s cycle Sir Hans Krebs Nobel prize, 1953 TCA (tricarboxylic acid) cycle The citric acid cycle requires aerobic conditions!!!!
TCA CYCLE (Citric Acid Cycle) TCA CYCLE The Citric Acid Cycle is also known as: Kreb s cycle Sir Hans Krebs Nobel prize, 1953 TCA (tricarboxylic acid) cycle The citric acid cycle requires aerobic conditions!!!!
Membranes: Membranes:
 Membranes: organize the chemical activities of cells by organizing different metabolic processes Control the flow of substances into or out of the cell The plasma membrane of the cell is selectively permeable
Membranes: organize the chemical activities of cells by organizing different metabolic processes Control the flow of substances into or out of the cell The plasma membrane of the cell is selectively permeable
Review Session 1. Control Systems and Homeostasis. Figure 1.8 A simple control system. Biol 219 Review Sessiono 1 Fall 2016
 Control Systems and Homeostasis Review Session 1 Regulated variables are kept within normal range by control mechanisms Keeps near set point, or optimum value Control systems local and reflex Input signal
Control Systems and Homeostasis Review Session 1 Regulated variables are kept within normal range by control mechanisms Keeps near set point, or optimum value Control systems local and reflex Input signal
f(x) = x R² = RPKM (M8.MXB) f(x) = x E-014 R² = 1 RPKM (M31.
 14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
Biosynthesis of Fatty Acids
 Biosynthesis of Fatty Acids Fatty acid biosynthesis takes place in the cytosol rather than the mitochondria and requires a different activation mechanism and different enzymes and coenzymes than fatty
Biosynthesis of Fatty Acids Fatty acid biosynthesis takes place in the cytosol rather than the mitochondria and requires a different activation mechanism and different enzymes and coenzymes than fatty
The further from the nucleus, the higher the electron s energy Valence shell electrons participate in biological reactions
 Chemistry of Life Revision: The further from the nucleus, the higher the electron s energy Valence shell electrons participate in biological reactions Atoms exchange electrons with other elements to form
Chemistry of Life Revision: The further from the nucleus, the higher the electron s energy Valence shell electrons participate in biological reactions Atoms exchange electrons with other elements to form
Mitochondria and ATP Synthesis
 Mitochondria and ATP Synthesis Mitochondria and ATP Synthesis 1. Mitochondria are sites of ATP synthesis in cells. 2. ATP is used to do work; i.e. ATP is an energy source. 3. ATP hydrolysis releases energy
Mitochondria and ATP Synthesis Mitochondria and ATP Synthesis 1. Mitochondria are sites of ATP synthesis in cells. 2. ATP is used to do work; i.e. ATP is an energy source. 3. ATP hydrolysis releases energy
Cellular Respiration Harvesting Chemical Energy ATP
 Cellular Respiration Harvesting Chemical Energy ATP 2006-2007 What s the point? The point is to make ATP! ATP 2006-2007 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats,
Cellular Respiration Harvesting Chemical Energy ATP 2006-2007 What s the point? The point is to make ATP! ATP 2006-2007 Harvesting stored energy Energy is stored in organic molecules carbohydrates, fats,
Lipids and Classification:
 Lipids and Classification: Lipids: Biological lipids are a chemically diverse group of organic compounds which are insoluble or only poorly soluble in water. They are readily soluble in non-polar solvents
Lipids and Classification: Lipids: Biological lipids are a chemically diverse group of organic compounds which are insoluble or only poorly soluble in water. They are readily soluble in non-polar solvents
CELL BIOLOGY - CLUTCH CH AEROBIC RESPIRATION.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF AEROBIC RESPIRATION Cellular respiration is a series of reactions involving electron transfers to breakdown molecules for (ATP) 1. Glycolytic pathway: Glycolysis
!! www.clutchprep.com CONCEPT: OVERVIEW OF AEROBIC RESPIRATION Cellular respiration is a series of reactions involving electron transfers to breakdown molecules for (ATP) 1. Glycolytic pathway: Glycolysis
What s the point? The point is to make ATP! ATP
 2006-2007 What s the point? The point is to make ATP! ATP Glycolysis 2 ATP Kreb s cycle 2 ATP Life takes a lot of energy to run, need to extract more energy than 4 ATP! There s got to be a better way!
2006-2007 What s the point? The point is to make ATP! ATP Glycolysis 2 ATP Kreb s cycle 2 ATP Life takes a lot of energy to run, need to extract more energy than 4 ATP! There s got to be a better way!
Biology 638 Biochemistry II Exam-3. (Note that you are not allowed to use any calculator)
 Biology 638 Biochemistry II Exam-3 (Note that you are not allowed to use any calculator) 1. In the non-cyclic pathway, electron pathway is. Select the most accurate one. a. PSII PC Cyt b 6 f PC PSI Fd-NADP
Biology 638 Biochemistry II Exam-3 (Note that you are not allowed to use any calculator) 1. In the non-cyclic pathway, electron pathway is. Select the most accurate one. a. PSII PC Cyt b 6 f PC PSI Fd-NADP
Figure S3 Differentially expressed fungal and plant genes organised by catalytic activity ontology Organisation of E. festucae (A) and L.
 sak WT Number of branches 3 WT sak Figure S1 Vasculature of plants infected with the E. festucae sak mutant. Light micrographs of perennial ryegrass blade tissue showing branching between the vasculature
sak WT Number of branches 3 WT sak Figure S1 Vasculature of plants infected with the E. festucae sak mutant. Light micrographs of perennial ryegrass blade tissue showing branching between the vasculature
