Supplementary Information
|
|
|
- April Parrish
- 6 years ago
- Views:
Transcription
1 Supplementary Information Assembly and Clustering of Natural Antibiotics Guides Target Identification Chad W. Johnston 1,2, Michael A. Skinnider 1,2, Chris A. Dejong 1,2, Philip N. Rees 1,2, Gregory M. Chen 1,2, Chelsea Walker 1,2, Shawn French 1, Eric D. Brown 1, Janos Berdy 3, Dennis Y. Liu 1,2 & Nathan A. Magarvey 1,2 * 1 Department of Biochemistry & Biomedical Sciences; M. G. DeGroote Institute for Infectious Disease Research; 2 Department of Chemistry & Chemical Biology; McMaster University, Hamilton, Canada L8S 4K1 3 Eötvös Loránd University, Budapest, Hungary * Corresponding author, address: magarv@mcmaster.ca
2 Supplementary Results Supplementary Figure 1. Antibioticome web application. The antibioticome search web application (accessible at provides a user-friendly mechanism to query the targeted antibioticome, in order to define the putative molecular target of a user-submitted molecule. A single molecular structure or database of molecular structures are submitted in SMILES format, and undergo in silico retrobiosynthesis followed by hierarchical clustering. The single highest-scoring targeted antibiotic match identified by computational retrobiosynthesis is reported for each submitted molecule, as well as the family of natural products to which that targeted antibiotic belongs, the mechanism of action of that family, and the confidence score assigned to the match.
3 Supplementary Figure 2. PRISM a web application for identifying biosynthetic gene clusters. Screenshot of the PRISM user interface. Supplementary Figure 3. PRISM results for the cephalosporin biosynthetic gene cluster. (a) Detection of the cephalosporin biosynthetic gene cluster. Red indicates nonribosomal peptide synthetase genes, brown indicates resistance genes, and green indicates tailoring enzymes. (b) Detection of an embedded resistance determinant related to the cephalosporin mode of action Class C beta lactamase.
4 Supplementary Figure 4. PRISM results for the daptomycin biosynthetic gene cluster. (a) Detection of the daptomycin biosynthetic gene cluster. Red indicates nonribosomal peptide synthetase genes, brown indicates resistance genes, green indicates tailoring enzymes, and gray represents other biosynthetic enzymes. (b) Detection of an embedded resistance determinant related to the daptomycin chemical scaffold, a daptomycin ABC transporter. Supplementary Figure 5. PRISM results for the erythromycin biosynthetic gene cluster. (a) Detection of the erythromycin biosynthetic gene cluster. Blue indicates polyketide synthase genes, brown indicates resistance genes, purple indicates sugar biosynthesis genes, green indicates tailoring enzymes, and gray represents other biosynthetic enzymes. (b) Detection of an embedded resistance determinant related to the erythromycin mode of action, the ribosome modifying rrna methyltransferase.
5 Supplementary Figure 6. PRISM results for the friulimicin biosynthetic gene cluster. (a) Detection of the friulimicin biosynthetic gene cluster. Red indicates nonribosomal peptide synthetase genes, brown indicates resistance genes, and gray represents other biosynthetic enzymes. (b) Detection of an embedded resistance determinant related to the friulimicin chemical scaffold, a friulimicin ABC transporter. Supplementary Figure 7. PRISM results for the mupirocin biosynthetic gene cluster. (a) Detection of the mupirocin biosynthetic gene cluster. Blue indicates polyketide synthase genes, brown indicates resistance genes, and gray represents other biosynthetic enzymes. (b) Detection of an embedded resistance determinant related to the mupirocin mode of action, a mupirocin resistant Ile-tRNA synthetase.
6 Supplementary Figure 8. PRISM results for the albomycin biosynthetic gene cluster. (a) Detection of the albomycin biosynthetic gene cluster. Red indicates nonribosomal peptide synthetase genes, brown indicates resistance genes, and green indicates tailoring enzymes. (b) Detection of an embedded resistance determinant related to the albomycin mode of action, an albomycin resistant Ser-tRNA synthetase. Supplementary Figure 9. PRISM results for the albicidin biosynthetic gene cluster. (a) Detection of the albicidin biosynthetic gene cluster. Red indicates nonribosomal peptide synthetase genes, blue represents polyketide synthetase genes, brown indicates resistance genes, green indicates tailoring enzymes, and gray represents other biosynthetic enzymes. (b) Detection of an embedded resistance determinant related to the albicidin chemical scaffold, an albicidin ABC transporter.
7 Supplementary Figure 10. PRISM results for the teicoplanin biosynthetic gene cluster. (a) Detection of the teicoplanin biosynthetic gene cluster. Red indicates nonribosomal peptide synthetase genes, brown indicates resistance genes, green indicates tailoring enzymes, and gray represents other biosynthetic enzymes. (b) Detection of an embedded resistance determinant related to the teicoplanin mode of action the D-ala-D-lac glycopeptide-resistant ligase. Supplementary Figure 11. PRISM results for the capuramycin biosynthetic gene cluster. (a) Detection of the capuramycin biosynthetic gene cluster. Red indicates nonribosomal peptide synthetase genes, brown indicates resistance genes, purple indicates sugar biosynthesis enzymes, and gray represents other biosynthetic enzymes. (b) Detection of an embedded resistance determinant specific to the capuramycin chemical scaffold A phosphotransferase.
8 Supplementary Figure 12. PRISM results for the telomycin biosynthetic gene cluster. (a) Detection of the telomycin biosynthetic gene cluster. Red indicates nonribosomal peptide synthetase genes, brown indicates resistance genes, green indicates tailoring enzymes, and gray represents other biosynthetic enzymes. (b) Detection of a generic ABC transporter as the sole known resistance determinant detected in the telomycin gene cluster. Supplementary Figure 13. Telomycin does not cause hemolysis. A solution of 0.25% sheep red blood cell suspension in phosphate-buffered saline was incubated for 1 h at 37 C with either a serial dilution of telomycin, 1% Triton X-100 (positive lysis control), DMSO alone (1%; negative lysis control), or without added content (negative lysis control). After 1 h RBCs were pelleted and lysis was assessed by measuring absorbance of the supernatant at 540 nm.
9 Supplementary Figure 14. Summary of observed spontaneous telomycin-resistance mutations. Sequenced genomes of telomycin-resistant S. aureus Newman and B. subtilis 168 were compared to sensitive parent strains, revealing mutations in the dominant, house-keeping cardiolipin synthase genes cls2 and clsa, respectively. Mutations occurred in the catalytic domains of cardiolipin synthase, and either resulted in truncations, or missense mutations near the active site HKD motifs. Supplementary Figure 15. Fluorescence microscopy of anionic-lipid dye NAO. (a) Structure of the cardiolipin dye 10-N-nonyl-acridine orange (NAO). (b) Fluorescence microscopy images of B. subtilis with the standard membrane dye FM4-64 and the standard cardiolipin dye NAO. Summed pixel intensities over the length of a selected bacteria are depicted below.
10 Supplementary Tables Gene Predicted Function Strand Amino Acids tlo1 Alpha-mannosidase tlo2 Hydrolase tlo3 ABC transporter permease tlo4 ABC transporter permease tlo5 Extracellular binding domain tlo6 Hypothetical protein tlo7 Putative glucokinase tlo8 LacI-family transcription regulator tlo9 Hypothetical protein tlo10 Hydrolase tlo11 Transcriptional regulator tlo12 Tryptophan dehydrogenase tlo13 Antitermination regulator tlo14 GntR-family transcription regulator tlo15 Anthranilate phosphoribosyltransferase tlo16 Phospholipase tlo17 HTH regulatory protein tlo18 Fatty acyl adenylate ligase tlo19 Acyl carrier protein + 90 tlo20 NRPS tlo21 NRPS tlo22 NRPS tlo23 Cytochrome P tlo24 MbtH protein + 75 tlo25 Acylase Transposase Transposase tlo26 / Hydrolase tlo27 Aminotransferase-methyltransferase tlo28 Ferritin-like tlo29 Cytochrome P tlo30 ABC transporter membrane protein tlo31 ABC transporter ATPase tlo32 Proline hydroxylase tlo33 Hypothetical protein tlo34 Hypothetical protein Supplementary Table 1. Telomycin gene cluster analysis.
11 Bacterial strain Telomycin MIC % Cardiolipin Staphylococcus aureus Newman 8 g/ml 100% S. aureus Newman ClsA Q148stop 128 g/ml 1.7% S. aureus Newman ClsA A388stop 128 g/ml 3.8% Bacillus subtilis g/ml 100% B. subtilis 168 ClsA P442L 16 g/ml 10.2% Supplementary Table 2. Cardiolipin levels present in spontaneously telomycin-resistant mutant strains. [NaCl] (M) S. aureus wild type S. aureus Telo R Supplementary Table 3. Increasing osmotic stress increases telomycin potency. MICs of sensitive and telomycin-resistant (cls2 A388stop) S. aureus cultured in CAMHB containing 0, 0.5, 1, or 1.5 M NaCl. MICs were determined after 16 h growth, and are shown in μg ml -1. B. subtilis 168 wildtype B. subtilis 168 Telo R S. aureus Newman wildtype S. aureus Newman Telo R (1) (2) (3) (4) (5) (6) (7) >128 (8) di-5-hydroxytryptophan telomycin (9) di-5-methoxytryptophan telomycin (10) >128 >128 >128 > >128 Supplementary Table 4. MICs of telomycin antibiotics (1-10). Sensitive and resistant strains were exposed to telomycins and MICs were measured by microdilution in CAMHB. MICs were determined after 16 h growth, and are shown in μg ml -1.
12 Supplementary Datasets Supplementary Dataset 1. Results from retrobiosynthetic analysis of microbial modular natural product antibacterials. A retrobiosynthetic algorithm was used to cluster specific antibacterial natural products by their chemical scaffolds. Clustering analysis yielded proposed classes based on conserved chemical scaffold, with the number of structures in each class is listed along with a representative chemical structure and statement on the mechanism of action or known cross resistance. Molecules which are not present in this dataset were not accepted as inputs by our retrobiosynthetic algorithm or were not specific antibacterial agents. Supplementary Dataset 2. Hidden Markov models used by PRISM to detect antibiotic resistance genes.
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function
 Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
Figure S3 Differentially expressed fungal and plant genes organised by catalytic activity ontology Organisation of E. festucae (A) and L.
 sak WT Number of branches 3 WT sak Figure S1 Vasculature of plants infected with the E. festucae sak mutant. Light micrographs of perennial ryegrass blade tissue showing branching between the vasculature
sak WT Number of branches 3 WT sak Figure S1 Vasculature of plants infected with the E. festucae sak mutant. Light micrographs of perennial ryegrass blade tissue showing branching between the vasculature
Supplementary Figure 1. The proposed biosynthetic pathways for the A-ring transformation of the
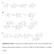 Supplementary Figure 1. The proposed biosynthetic pathways for the A-ring transformation of the kinamycin and lomaiviticin antibiotics. A, by Steven J. Gould; B, by Emily P. Balskus; C, by Bradley S. Moore.
Supplementary Figure 1. The proposed biosynthetic pathways for the A-ring transformation of the kinamycin and lomaiviticin antibiotics. A, by Steven J. Gould; B, by Emily P. Balskus; C, by Bradley S. Moore.
A look at macromolecules (Text pages 38-54) What is the typical chemical composition of a cell? (Source of figures to right: Madigan et al.
 A look at macromolecules (Text pages 38-54) What is the typical chemical composition of a cell? (Source of figures to right: Madigan et al. 2002 Chemical Bonds Ionic Electron-negativity differences cause
A look at macromolecules (Text pages 38-54) What is the typical chemical composition of a cell? (Source of figures to right: Madigan et al. 2002 Chemical Bonds Ionic Electron-negativity differences cause
Figure S1: Abundance of Fe related proteins in oceanic Synechococcus sp. strain WH8102 in. Ferredoxin. Ferredoxin. Fe ABC transporter
 0.004 SW nm 0.04 0.03 nm nm 0.16 0.1 nm nm 0.40 0.3 nm nm 0.80 0.5 nm 1.6 1 nm 16 10 nm 160 100 nm 0 nm 0.04 nm 0.16 nm 0.40 nm 0.80 nm 1.6 nm 16 nm 160 nm Spectral counts 0 nm 0.04 nm 0.16 nm 0.40 nm
0.004 SW nm 0.04 0.03 nm nm 0.16 0.1 nm nm 0.40 0.3 nm nm 0.80 0.5 nm 1.6 1 nm 16 10 nm 160 100 nm 0 nm 0.04 nm 0.16 nm 0.40 nm 0.80 nm 1.6 nm 16 nm 160 nm Spectral counts 0 nm 0.04 nm 0.16 nm 0.40 nm
Supplementary Information. Sonorensin: A new bacteriocin with potential of an anti-biofilm agent and a food
 Supplementary Information Sonorensin: A new bacteriocin with potential of an anti-biofilm agent and a food biopreservative Lipsy Chopra, Gurdeep Singh, Kautilya Kumar Jena and Debendra K. Sahoo* Biochemical
Supplementary Information Sonorensin: A new bacteriocin with potential of an anti-biofilm agent and a food biopreservative Lipsy Chopra, Gurdeep Singh, Kautilya Kumar Jena and Debendra K. Sahoo* Biochemical
f(x) = x R² = RPKM (M8.MXB) f(x) = x E-014 R² = 1 RPKM (M31.
 14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
Acetyl-CoA or BC acyl-coa. Int1-ACP. FabH. FabF. Anti-FabF Platensimycin ACP. Fak
 Initiation Acetyl-CoA AccBCDA FabZ Int-ACP Elongation [N+] FabG Int1-ACP Acetyl-CoA or BC acyl-coa FabH FabD CoA Malonyl-ACP Malonyl CoA ACP Anti-FabI Triclosan AFN-15 FabI acyl-acp PlsX PlsY Pls C Lipoic
Initiation Acetyl-CoA AccBCDA FabZ Int-ACP Elongation [N+] FabG Int1-ACP Acetyl-CoA or BC acyl-coa FabH FabD CoA Malonyl-ACP Malonyl CoA ACP Anti-FabI Triclosan AFN-15 FabI acyl-acp PlsX PlsY Pls C Lipoic
LIPOPREDICT: Bacterial lipoprotein prediction server
 www.bioinformation.net Server Volume 8(8) LIPOPREDICT: Bacterial lipoprotein prediction server S Ramya Kumari, Kiran Kadam, Ritesh Badwaik & Valadi K Jayaraman* Centre for Development of Advanced Computing
www.bioinformation.net Server Volume 8(8) LIPOPREDICT: Bacterial lipoprotein prediction server S Ramya Kumari, Kiran Kadam, Ritesh Badwaik & Valadi K Jayaraman* Centre for Development of Advanced Computing
Point total. Page # Exam Total (out of 90) The number next to each intermediate represents the total # of C-C and C-H bonds in that molecule.
 This exam is worth 90 points. Pages 2- have questions. Page 1 is for your reference only. Honor Code Agreement - Signature: Date: (You agree to not accept or provide assistance to anyone else during this
This exam is worth 90 points. Pages 2- have questions. Page 1 is for your reference only. Honor Code Agreement - Signature: Date: (You agree to not accept or provide assistance to anyone else during this
CHAPTER 6 METABOLIC PATHWAY RECONSTRUCTION
 66 CHAPTER 6 METABOLIC PATHWAY RECONSTRUCTION 6.1 GENERAL A metabolic pathway is a series of biochemical reactions that occur within a cell and these chemical reactions are catalyzed by certain enzymes.
66 CHAPTER 6 METABOLIC PATHWAY RECONSTRUCTION 6.1 GENERAL A metabolic pathway is a series of biochemical reactions that occur within a cell and these chemical reactions are catalyzed by certain enzymes.
Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples:
 Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Supplemental Information for: Localization of anionic phospholipids in Escherichia coli cells
 Supplemental Information for: Localization of anionic phospholipids in Escherichia coli cells Piercen M. Oliver, a John A. Crooks, a Mathias Leidl, b Earl J. Yoon, a Alan Saghatelian, b and Douglas B.
Supplemental Information for: Localization of anionic phospholipids in Escherichia coli cells Piercen M. Oliver, a John A. Crooks, a Mathias Leidl, b Earl J. Yoon, a Alan Saghatelian, b and Douglas B.
Macrolides, Clindamycin & Ketolides Polymyxins
 Macrolides, Clindamycin & Ketolides Polymyxins Kwan Soo Ko Macrolides - Erythromycin - Azithromycin - Clarithromycin Lincosamides - Lincomycin - Clindamycin Unrelated chemically But, many similar biological
Macrolides, Clindamycin & Ketolides Polymyxins Kwan Soo Ko Macrolides - Erythromycin - Azithromycin - Clarithromycin Lincosamides - Lincomycin - Clindamycin Unrelated chemically But, many similar biological
Genome-wide association studies for understanding pathogen evolution. Samuel K. Sheppard
 Genome-wide association studies for understanding pathogen evolution Samuel K. Sheppard Pathogenic bacteria Population genomics and evolution Sheppard et al. Science 2008; 11: 237-239. Sheppard et al.
Genome-wide association studies for understanding pathogen evolution Samuel K. Sheppard Pathogenic bacteria Population genomics and evolution Sheppard et al. Science 2008; 11: 237-239. Sheppard et al.
An Introduction to Carbohydrates
 An Introduction to Carbohydrates Carbohydrates are a large class of naturally occurring polyhydroxy aldehydes and ketones. Monosaccharides also known as simple sugars, are the simplest carbohydrates containing
An Introduction to Carbohydrates Carbohydrates are a large class of naturally occurring polyhydroxy aldehydes and ketones. Monosaccharides also known as simple sugars, are the simplest carbohydrates containing
Investigations of Pactamycin Biosynthesis Andrew Osborn
 Investigations of Pactamycin Biosynthesis Andrew Osborn Research Mentors: Dr. Taifo Mahmud, Dr. Kerry McPhail A Bioresource Research Thesis Seminar Presentation Streptomyces pactum produces the antitumor
Investigations of Pactamycin Biosynthesis Andrew Osborn Research Mentors: Dr. Taifo Mahmud, Dr. Kerry McPhail A Bioresource Research Thesis Seminar Presentation Streptomyces pactum produces the antitumor
Supplemental Information
 Supplemental Information Screening of strong constitutive promoters in the S. albus transcriptome via RNA-seq The total RNA of S. albus J1074 was isolated after 24 hrs and 72 hrs of cultivation at 30 C
Supplemental Information Screening of strong constitutive promoters in the S. albus transcriptome via RNA-seq The total RNA of S. albus J1074 was isolated after 24 hrs and 72 hrs of cultivation at 30 C
AP Biology Summer Assignment Chapter 3 Quiz
 AP Biology Summer Assignment Chapter 3 Quiz 2016-17 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. All of the following are found in a DNA nucleotide
AP Biology Summer Assignment Chapter 3 Quiz 2016-17 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. All of the following are found in a DNA nucleotide
Student name ID # 2. (4 pts) What is the terminal electron acceptor in respiration? In photosynthesis?
 1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? 2. (4 pts) What is the terminal electron acceptor
1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? 2. (4 pts) What is the terminal electron acceptor
Nature Methods: doi: /nmeth Supplementary Figure 1. Salipro lipid particles.
 Supplementary Figure 1 Salipro lipid particles. (a) Gel filtration analysis of Saposin A after incubation with the indicated detergent solubilised lipid solutions. The generation of Saposin A-lipid complexes
Supplementary Figure 1 Salipro lipid particles. (a) Gel filtration analysis of Saposin A after incubation with the indicated detergent solubilised lipid solutions. The generation of Saposin A-lipid complexes
Mechanisms of Daptomycin Resistance and the Seesaw Effect in Multi-Drug Resistant Enterococci. NAME OF STUDENT Candidacy Exam August 25, 2017
 Mechanisms of Daptomycin Resistance and the Seesaw Effect in Multi-Drug Resistant Enterococci NAME OF STUDENT Candidacy Exam August 25, 2017 Major nosocomial pathogen Endocarditis, bacteremia, UTIs, meningitis
Mechanisms of Daptomycin Resistance and the Seesaw Effect in Multi-Drug Resistant Enterococci NAME OF STUDENT Candidacy Exam August 25, 2017 Major nosocomial pathogen Endocarditis, bacteremia, UTIs, meningitis
Syllabus for BASIC METABOLIC PRINCIPLES
 Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
Resistance to new anti-grampositive. Roland Leclercq, Microbiology, CHU Cote de Nacre, Caen, France
 Resistance to new anti-grampositive agents Roland Leclercq, Microbiology, CHU Cote de Nacre, Caen, France Recently available antimicrobials against MDR Gram-positive infections Cyclic lipopeptide: daptomycin
Resistance to new anti-grampositive agents Roland Leclercq, Microbiology, CHU Cote de Nacre, Caen, France Recently available antimicrobials against MDR Gram-positive infections Cyclic lipopeptide: daptomycin
In vitro bactericidal assay Fig. S8 Gentamicin protection assay Phagocytosis assay
 In vitro bactericidal assay Mouse bone marrow was isolated from the femur and the tibia. Cells were suspended in phosphate buffered saline containing.5% BSA and 2 mm EDTA and filtered through a cell strainer.
In vitro bactericidal assay Mouse bone marrow was isolated from the femur and the tibia. Cells were suspended in phosphate buffered saline containing.5% BSA and 2 mm EDTA and filtered through a cell strainer.
Discussion of Prism modules and predicted interactions (Fig. 4)
 SUPPLEMENTARY NOTES Discussion of Prism modules and predicted interactions (Fig. 4) a. Interactions of the TCA-cycle, respiratory chain, and ATP synthetase with the amino acid biosynthesis modules. Given
SUPPLEMENTARY NOTES Discussion of Prism modules and predicted interactions (Fig. 4) a. Interactions of the TCA-cycle, respiratory chain, and ATP synthetase with the amino acid biosynthesis modules. Given
Spectrum of vancomycin and susceptibility testing
 Spectrum of vancomycin and susceptibility testing Olivier Denis Service de Microbiologie Laboratoire de bactériologie Service de Microbiologie Hôpital Erasme Glycopeptides Vancomycin 1958 < Amycolatopsis
Spectrum of vancomycin and susceptibility testing Olivier Denis Service de Microbiologie Laboratoire de bactériologie Service de Microbiologie Hôpital Erasme Glycopeptides Vancomycin 1958 < Amycolatopsis
Analysis of Sphingosine Kinase Activity in Single Natural Killer Cells from Peripheral Blood
 Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information: Analysis of Sphingosine Kinase Activity in Single
Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2015 Electronic Supplementary Information: Analysis of Sphingosine Kinase Activity in Single
1. endoplasmic reticulum This is the location where N-linked oligosaccharide is initially synthesized and attached to glycoproteins.
 Biology 4410 Name Spring 2006 Exam 2 A. Multiple Choice, 2 pt each Pick the best choice from the list of choices, and write it in the space provided. Some choices may be used more than once, and other
Biology 4410 Name Spring 2006 Exam 2 A. Multiple Choice, 2 pt each Pick the best choice from the list of choices, and write it in the space provided. Some choices may be used more than once, and other
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supporting Information
 Supporting Information Discovery of linear low-cationic peptides to target methicillin-resistant Staphylococcus aureus in vivo Yuan Liu, Meirong Song, Shuangyang Ding,, Kui Zhu*,,, Beijing Advanced Innovation
Supporting Information Discovery of linear low-cationic peptides to target methicillin-resistant Staphylococcus aureus in vivo Yuan Liu, Meirong Song, Shuangyang Ding,, Kui Zhu*,,, Beijing Advanced Innovation
Affinity of Doripenem and Comparators to Penicillin-Binding Proteins in Escherichia coli and ACCEPTED
 AAC Accepts, published online ahead of print on February 00 Antimicrob. Agents Chemother. doi:./aac.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
AAC Accepts, published online ahead of print on February 00 Antimicrob. Agents Chemother. doi:./aac.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Hong-qi Sun, Xue-mei Lu, Pei-ji Gao* State Key Laboratory of Microbial Technology, Shandong University, Jinan , China.
 Brazilian Journal of Microbiology (2011) 42: 410-414 ISSN 1517-8382 THE EXPLORATION OF THE ANTIBACTERIAL MECHANISM OF FE 3+ AGAINST BACTERIA Hong-qi Sun, Xue-mei Lu, Pei-ji Gao* State Key Laboratory of
Brazilian Journal of Microbiology (2011) 42: 410-414 ISSN 1517-8382 THE EXPLORATION OF THE ANTIBACTERIAL MECHANISM OF FE 3+ AGAINST BACTERIA Hong-qi Sun, Xue-mei Lu, Pei-ji Gao* State Key Laboratory of
A Novel Lantibiotic Acting on Bacterial Cell Wall Synthesis Produced by the Uncommon FLAVIA MARINELLI. DBSM, University of Insubria, Varese Italy
 A Novel Lantibiotic Acting on Bacterial Cell Wall Synthesis Produced by the Uncommon Actinomycete Planomonospora sp. FLAVIA MARINELLI DBSM, University of Insubria, Varese Italy Vicuron Pharmaceuticals,
A Novel Lantibiotic Acting on Bacterial Cell Wall Synthesis Produced by the Uncommon Actinomycete Planomonospora sp. FLAVIA MARINELLI DBSM, University of Insubria, Varese Italy Vicuron Pharmaceuticals,
Table removed due to copyright reasons.
 atural Product Biosynthesis Relevance of natural products in the pharmaceutical industry (see Table 2 in Butler J. at. Prod. 2004, 67, 2141-2153.)) Table removed due to copyright reasons. 2 * a phosphopantetheinyl
atural Product Biosynthesis Relevance of natural products in the pharmaceutical industry (see Table 2 in Butler J. at. Prod. 2004, 67, 2141-2153.)) Table removed due to copyright reasons. 2 * a phosphopantetheinyl
Loss of protein association causes cardiolipin degradation in Barth syndrome
 SUPPLEMENTARY INFORMATION Loss of protein association causes cardiolipin degradation in Barth syndrome Yang Xu 1, Colin K.L. Phoon 2, Bob Berno 5, Kenneth D Souza 6, Esthelle Hoedt 4, Guoan Zhang 4, Thomas
SUPPLEMENTARY INFORMATION Loss of protein association causes cardiolipin degradation in Barth syndrome Yang Xu 1, Colin K.L. Phoon 2, Bob Berno 5, Kenneth D Souza 6, Esthelle Hoedt 4, Guoan Zhang 4, Thomas
Biochemistry 2000 Sample Question Transcription, Translation and Lipids. (1) Give brief definitions or unique descriptions of the following terms:
 (1) Give brief definitions or unique descriptions of the following terms: (a) exon (b) holoenzyme (c) anticodon (d) trans fatty acid (e) poly A tail (f) open complex (g) Fluid Mosaic Model (h) embedded
(1) Give brief definitions or unique descriptions of the following terms: (a) exon (b) holoenzyme (c) anticodon (d) trans fatty acid (e) poly A tail (f) open complex (g) Fluid Mosaic Model (h) embedded
PFK Activity Assay Kit (Colorimetric)
 PFK Activity Assay Kit (Colorimetric) Catalog Number KA3761 100 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information...
PFK Activity Assay Kit (Colorimetric) Catalog Number KA3761 100 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information...
Screening of bacteria producing amylase and its immobilization: a selective approach By Debasish Mondal
 Screening of bacteria producing amylase and its immobilization: a selective approach By Debasish Mondal Article Summary (In short - What is your article about Just 2 or 3 lines) Category: Bacillus sp produce
Screening of bacteria producing amylase and its immobilization: a selective approach By Debasish Mondal Article Summary (In short - What is your article about Just 2 or 3 lines) Category: Bacillus sp produce
Aminoglycoside activity observed on single pre-translocation ribosome complexes
 correction notice Nat. Chem. Biol. 6, 54 62 (2010) Aminoglycoside activity observed on single pre-translocation ribosome complexes Michael B Feldman, Daniel S Terry, Roger B Altman & Scott C Blanchard
correction notice Nat. Chem. Biol. 6, 54 62 (2010) Aminoglycoside activity observed on single pre-translocation ribosome complexes Michael B Feldman, Daniel S Terry, Roger B Altman & Scott C Blanchard
Physiological Adaptation. Microbial Physiology Module 4
 Physiological Adaptation Microbial Physiology Module 4 Topics Coordination of Metabolic Reactions Regulation of Enzyme Activity Regulation of Gene Expression Global Control, Signal Transduction and Twocomponent
Physiological Adaptation Microbial Physiology Module 4 Topics Coordination of Metabolic Reactions Regulation of Enzyme Activity Regulation of Gene Expression Global Control, Signal Transduction and Twocomponent
Extracellular Enzymes Lab
 Biochemistry Extracellular Enzymes Lab All organisms convert small organic compounds, such as glucose, into monomers required for the production of macromolecules; e.g., Building Blocks Monomers Macromolecules
Biochemistry Extracellular Enzymes Lab All organisms convert small organic compounds, such as glucose, into monomers required for the production of macromolecules; e.g., Building Blocks Monomers Macromolecules
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Introduction 1/3. The protagonist of our story: the prokaryotic ribosome
 Introduction 1/3 The protagonist of our story: the prokaryotic ribosome Introduction 2/3 The core functional domains of the ribosome (the peptidyl trasferase center and the decoding center) are composed
Introduction 1/3 The protagonist of our story: the prokaryotic ribosome Introduction 2/3 The core functional domains of the ribosome (the peptidyl trasferase center and the decoding center) are composed
Genetic information flows from mrna to protein through the process of translation
 Genetic information flows from mrn to protein through the process of translation TYPES OF RN (RIBONUCLEIC CID) RN s job - protein synthesis (assembly of amino acids into proteins) Three main types: 1.
Genetic information flows from mrn to protein through the process of translation TYPES OF RN (RIBONUCLEIC CID) RN s job - protein synthesis (assembly of amino acids into proteins) Three main types: 1.
SUPPLEMENTARY INFORMATION
 ARTICLE NUMBER: 16198 DOI: 10.1038/NMICROBIOL.2016.198 Genome reduction in an abundant and ubiquitous soil bacterium, Candidatus Udaeobacter copiosus Tess E Brewer 1, 2, Kim M Handley 3, Paul Carini 1,
ARTICLE NUMBER: 16198 DOI: 10.1038/NMICROBIOL.2016.198 Genome reduction in an abundant and ubiquitous soil bacterium, Candidatus Udaeobacter copiosus Tess E Brewer 1, 2, Kim M Handley 3, Paul Carini 1,
Figure 1 Original Advantages of biological reactions being catalyzed by enzymes:
 Enzyme basic concepts, Enzyme Regulation I III Carmen Sato Bigbee, Ph.D. Objectives: 1) To understand the bases of enzyme catalysis and the mechanisms of enzyme regulation. 2) To understand the role of
Enzyme basic concepts, Enzyme Regulation I III Carmen Sato Bigbee, Ph.D. Objectives: 1) To understand the bases of enzyme catalysis and the mechanisms of enzyme regulation. 2) To understand the role of
New Insights into Peptidoglycan Biosynthesis. Louis B. Rice Warren Alpert Medical School of Brown University Providence, RI
 New Insights into Peptidoglycan Biosynthesis Louis B. Rice Warren Alpert Medical School of Brown University Providence, RI Outline of Presentation Review role of penicillin-binding proteins in cell wall
New Insights into Peptidoglycan Biosynthesis Louis B. Rice Warren Alpert Medical School of Brown University Providence, RI Outline of Presentation Review role of penicillin-binding proteins in cell wall
Cases in employees. Cases. Day of onset (July)
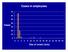 Cases in employees 70 60 50 Cases 40 30 20 10 0 0 2 4 6 8 10 12 14 16 18 20 22 24 26 28 30 Day of onset (July) 70 60 50 40 Cases 30 Cases in employees and visitors Employees Visitors 20 10 0 0 2 4 6 8
Cases in employees 70 60 50 Cases 40 30 20 10 0 0 2 4 6 8 10 12 14 16 18 20 22 24 26 28 30 Day of onset (July) 70 60 50 40 Cases 30 Cases in employees and visitors Employees Visitors 20 10 0 0 2 4 6 8
Biology. Lectures winter term st year of Pharmacy study
 Biology Lectures winter term 2008 1 st year of Pharmacy study 3 rd Lecture Chemical composition of living matter chemical basis of life. Atoms, molecules, organic compounds carbohydrates, lipids, proteins,
Biology Lectures winter term 2008 1 st year of Pharmacy study 3 rd Lecture Chemical composition of living matter chemical basis of life. Atoms, molecules, organic compounds carbohydrates, lipids, proteins,
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Selective protection of an ARF1-GTP signaling axis by a bacterial scaffold induces bidirectional trafficking arrest.
 Selective protection of an ARF1-GTP signaling axis by a bacterial scaffold induces bidirectional trafficking arrest. Andrey S. Selyunin, L. Evan Reddick, Bethany A. Weigele, and Neal M. Alto Supplemental
Selective protection of an ARF1-GTP signaling axis by a bacterial scaffold induces bidirectional trafficking arrest. Andrey S. Selyunin, L. Evan Reddick, Bethany A. Weigele, and Neal M. Alto Supplemental
Supplementary Information
 Supplementary Information The conifer biomarkers dehydroabietic and abietic acids are widespread in Cyanobacteria Maria Sofia Costa, Adriana Rego, Vitor Ramos, Tiago Afonso, Sara Freitas, Marco Preto,
Supplementary Information The conifer biomarkers dehydroabietic and abietic acids are widespread in Cyanobacteria Maria Sofia Costa, Adriana Rego, Vitor Ramos, Tiago Afonso, Sara Freitas, Marco Preto,
Lecture 13 (10/13/17)
 Lecture 13 (10/13/17) Reading: Ch6; 187-189, 204-205 Problems: Ch4 (text); 2, 3 NXT (after xam 2) Reading: Ch6; 190-191, 194-195, 197-198 Problems: Ch6 (text); 5, 6, 7, 24 OUTLIN NZYMS: Binding & Catalysis
Lecture 13 (10/13/17) Reading: Ch6; 187-189, 204-205 Problems: Ch4 (text); 2, 3 NXT (after xam 2) Reading: Ch6; 190-191, 194-195, 197-198 Problems: Ch6 (text); 5, 6, 7, 24 OUTLIN NZYMS: Binding & Catalysis
Biomolecules: amino acids
 Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Antibacterial activity of nitric oxide (NO) releasing silver nanoparticles (AgNPs)
 Antibacterial activity of nitric oxide (NO) releasing silver nanoparticles (AgNPs) Dr Amedea B. Seabra amedea.seabra@ufabc.edu.br Universidade Federal do ABC, Brazil 1 Silver nanoparticles (AgNPs) 2 Silver
Antibacterial activity of nitric oxide (NO) releasing silver nanoparticles (AgNPs) Dr Amedea B. Seabra amedea.seabra@ufabc.edu.br Universidade Federal do ABC, Brazil 1 Silver nanoparticles (AgNPs) 2 Silver
USA. Supplemental Information
 Cloning and sequencing of the kedarcidin biosynthetic gene cluster from Streptoalloteichus sp. ATCC 53650 revealing new insights into biosynthesis of the enediyne family of antitumor antibiotics Jeremy
Cloning and sequencing of the kedarcidin biosynthetic gene cluster from Streptoalloteichus sp. ATCC 53650 revealing new insights into biosynthesis of the enediyne family of antitumor antibiotics Jeremy
Department of Chemistry, Université de Montréal, C.P. 6128, Succursale centre-ville, Montréal, Québec, H3C 3J7, Canada.
 Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
2.28E E AKT/CAT P-3-4:
 Supplemental Table 1: IPA network analysis of the microarray data presented in Figure 6 showing the most significant molecular and cellular function classifications along with the respective number of
Supplemental Table 1: IPA network analysis of the microarray data presented in Figure 6 showing the most significant molecular and cellular function classifications along with the respective number of
Student Handout. green B 1. foam pieces. ) into a single unit to model the substrate in this reaction.
 Student Handout Introduction Enzymes are specialized proteins that catalyze or speed up chemical reactions within cells. The substance upon which an enzyme acts is called a substrate. Substrates are small
Student Handout Introduction Enzymes are specialized proteins that catalyze or speed up chemical reactions within cells. The substance upon which an enzyme acts is called a substrate. Substrates are small
Biological Pathways. Janick Mathys
 Biological Pathways Janick Mathys Biological Pathways Definition Biochemical compounds Biological interactions Energy Control interactions Levels of abstraction Types of biological pathways Integration
Biological Pathways Janick Mathys Biological Pathways Definition Biochemical compounds Biological interactions Energy Control interactions Levels of abstraction Types of biological pathways Integration
CHEM-643 Biochemistry Mid-term Examination 8:00 10:00, Friday, 2 November 2007
 EM-643 Biochemistry Mid-term Examination 8:00 10:00, Friday, 2 November 2007 Name Dr.. White - Instructor There are 10 pages to this examination including this page. Write your name on each new page. Read
EM-643 Biochemistry Mid-term Examination 8:00 10:00, Friday, 2 November 2007 Name Dr.. White - Instructor There are 10 pages to this examination including this page. Write your name on each new page. Read
Name: Date: Block: Biology 12
 Name: Date: Block: Biology 12 Provincial Exam Review: Cell Processes and Applications January 2003 Use the following diagram to answer questions 1 and 2. 1. Which labelled organelle produces most of the
Name: Date: Block: Biology 12 Provincial Exam Review: Cell Processes and Applications January 2003 Use the following diagram to answer questions 1 and 2. 1. Which labelled organelle produces most of the
Supplementary Table 1. Functional avidities of survivin-specific T-cell clones against LML-peptide
 Supplementary Table 1. Functional avidities of survivin-specific T-cell clones against LML-peptide pulsed T2 cells. clone avidity by 4-hour 51 Cr-release assay 50% lysis at E:T 10:1 [LML peptide, M] #24
Supplementary Table 1. Functional avidities of survivin-specific T-cell clones against LML-peptide pulsed T2 cells. clone avidity by 4-hour 51 Cr-release assay 50% lysis at E:T 10:1 [LML peptide, M] #24
Alan Weiner BIOCHEM 530 Friday, MEB 248 October 23, 2015 RNA structure, the ribosome, structure-based drug design
 Alan Weiner BIOCHEM 530 Friday, MEB 248 October 23, 2015 RNA structure, the ribosome, structure-based drug design Crick's "Central Dogma"? DNA makes RNA makes protein Crick's "Central Dogma"? DNA makes
Alan Weiner BIOCHEM 530 Friday, MEB 248 October 23, 2015 RNA structure, the ribosome, structure-based drug design Crick's "Central Dogma"? DNA makes RNA makes protein Crick's "Central Dogma"? DNA makes
LAB 5 - Enzymes BACKGROUND INFORMATION
 LAB 5 - Enzymes BACKGROUND INFORMATION Chemical Reactions The cells of organisms, from bacteria to plants to animals, carry out hundreds to thousands of chemical reactions that must be properly coordinated
LAB 5 - Enzymes BACKGROUND INFORMATION Chemical Reactions The cells of organisms, from bacteria to plants to animals, carry out hundreds to thousands of chemical reactions that must be properly coordinated
Supplementary Figure 1. Tamoxifen does not promote chemotaxis or chemokinesis in the absence of additional stimulation. Transwell chemotaxis assays
 Supplementary Figure 1. Tamoxifen does not promote chemotaxis or chemokinesis in the absence of additional stimulation. Transwell chemotaxis assays revealed that tamoxifen treatment alone did not stimulate
Supplementary Figure 1. Tamoxifen does not promote chemotaxis or chemokinesis in the absence of additional stimulation. Transwell chemotaxis assays revealed that tamoxifen treatment alone did not stimulate
Microbiology - Problem Drill 16: Antibiotics. Question No. 1 of 10. Question. Feedback. Question
 Microbiology - Problem Drill 16: Antibiotics No. 1 of 10 1. An effective chemotherapeutic drug should have. (A) Low therapeutic index (B) More toxicity (C) Selective toxicity (D) Mutation inducing properties
Microbiology - Problem Drill 16: Antibiotics No. 1 of 10 1. An effective chemotherapeutic drug should have. (A) Low therapeutic index (B) More toxicity (C) Selective toxicity (D) Mutation inducing properties
SUPPORTING INFORMATION. Studies on antimicrobial evaluation of some 1-((1-(1H-benzo[d]imidazol-2-
![SUPPORTING INFORMATION. Studies on antimicrobial evaluation of some 1-((1-(1H-benzo[d]imidazol-2- SUPPORTING INFORMATION. Studies on antimicrobial evaluation of some 1-((1-(1H-benzo[d]imidazol-2-](/thumbs/79/79646314.jpg) SUPPORTIG IFORMATIO Studies on antimicrobial evaluation of some 1-((1-(1H-benzo[d]imidazol-2- yl)ethylidene)amino)-6-((arylidene)amino)-2-oxo-4-phenyl-1,2-dihydropyridine-3,5- dicarbonitriles C Desai*,
SUPPORTIG IFORMATIO Studies on antimicrobial evaluation of some 1-((1-(1H-benzo[d]imidazol-2- yl)ethylidene)amino)-6-((arylidene)amino)-2-oxo-4-phenyl-1,2-dihydropyridine-3,5- dicarbonitriles C Desai*,
ab ATP Synthase Enzyme Activity Microplate Assay Kit
 ab109714 ATP Synthase Enzyme Activity Microplate Assay Kit Instructions for Use For the quantitative measurement of ATP Synthase activity in samples from Human, Rat and Cow This product is for research
ab109714 ATP Synthase Enzyme Activity Microplate Assay Kit Instructions for Use For the quantitative measurement of ATP Synthase activity in samples from Human, Rat and Cow This product is for research
The Cell Membrane (Ch. 7)
 The Cell Membrane (Ch. 7) Phospholipids Phosphate head hydrophilic Fatty acid tails hydrophobic Arranged as a bilayer Phosphate attracted to water Fatty acid repelled by water Aaaah, one of those structure
The Cell Membrane (Ch. 7) Phospholipids Phosphate head hydrophilic Fatty acid tails hydrophobic Arranged as a bilayer Phosphate attracted to water Fatty acid repelled by water Aaaah, one of those structure
Algal Biofuels Research: Using basic science to maximize fuel output. Jacob Dums, PhD candidate, Heike Sederoff Lab March 9, 2015
 Algal Biofuels Research: Using basic science to maximize fuel output Jacob Dums, PhD candidate, jtdums@ncsu.edu Heike Sederoff Lab March 9, 2015 Outline Research Approach Dunaliella Increase Oil Content
Algal Biofuels Research: Using basic science to maximize fuel output Jacob Dums, PhD candidate, jtdums@ncsu.edu Heike Sederoff Lab March 9, 2015 Outline Research Approach Dunaliella Increase Oil Content
ab CytoPainter Golgi/ER Staining Kit
 ab139485 CytoPainter Golgi/ER Staining Kit Instructions for Use Designed to detect Golgi bodies and endoplasmic reticulum by microscopy This product is for research use only and is not intended for diagnostic
ab139485 CytoPainter Golgi/ER Staining Kit Instructions for Use Designed to detect Golgi bodies and endoplasmic reticulum by microscopy This product is for research use only and is not intended for diagnostic
Calibration Protocols
 Calibration Protocols (1)OD 600 Reference point LUDOX Protocol Materials: 1ml LUDOX CL-X (provided in kit) ddh20 (provided by team) 96 well plate, black with clear flat bottom preferred (provided by team)
Calibration Protocols (1)OD 600 Reference point LUDOX Protocol Materials: 1ml LUDOX CL-X (provided in kit) ddh20 (provided by team) 96 well plate, black with clear flat bottom preferred (provided by team)
4/12/17. Cells. Cell Structure. Ch. 2 Cell Structure and Func.on. Range of Cell Sizes BIOL 100
 Ch. 2 Cell Structure and Func.on BIOL 100 Cells Fundamental units of life Cell theory All living things are composed of one or more cells. The cell is the most basic unit of life. All cells come from pre-existing
Ch. 2 Cell Structure and Func.on BIOL 100 Cells Fundamental units of life Cell theory All living things are composed of one or more cells. The cell is the most basic unit of life. All cells come from pre-existing
Bjoern Peters La Jolla Institute for Allergy and Immunology Buenos Aires, Oct 31, 2012
 www.iedb.org Bjoern Peters bpeters@liai.org La Jolla Institute for Allergy and Immunology Buenos Aires, Oct 31, 2012 Overview 1. Introduction to the IEDB 2. Application: 2009 Swine-origin influenza virus
www.iedb.org Bjoern Peters bpeters@liai.org La Jolla Institute for Allergy and Immunology Buenos Aires, Oct 31, 2012 Overview 1. Introduction to the IEDB 2. Application: 2009 Swine-origin influenza virus
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL
 Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Glycolysis - Plasmodium
 Apicomplexan Biochemistry Basics Toxoplasma Good cell biology model Genome sequencing not completed Virtual pathways Cryptosporidium The strange one Genome sequence completed Virtual pathways Plasmodium
Apicomplexan Biochemistry Basics Toxoplasma Good cell biology model Genome sequencing not completed Virtual pathways Cryptosporidium The strange one Genome sequence completed Virtual pathways Plasmodium
Extracellular Enzymes Lab
 Biochemistry Extracellular Enzymes Lab All organisms convert small organic compounds, such as glucose, into monomers required for the production of macromolecules; e.g., Building Blocs Monomers Macromolecules
Biochemistry Extracellular Enzymes Lab All organisms convert small organic compounds, such as glucose, into monomers required for the production of macromolecules; e.g., Building Blocs Monomers Macromolecules
Biochemistry 2 Recita0on Amino Acid Metabolism
 Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Biochemistry: A Short Course
 Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 30 Amino Acid Degradation and the Urea Cycle 2013 W. H. Freeman and Company Chapter 30 Outline Amino acids are obtained from the
Tymoczko Berg Stryer Biochemistry: A Short Course Second Edition CHAPTER 30 Amino Acid Degradation and the Urea Cycle 2013 W. H. Freeman and Company Chapter 30 Outline Amino acids are obtained from the
Activity: Biologically Important Molecules
 Activity: Biologically Important Molecules AP Biology Introduction We have already seen in our study of biochemistry that the molecules that comprise living things are carbon-based, and that they are thought
Activity: Biologically Important Molecules AP Biology Introduction We have already seen in our study of biochemistry that the molecules that comprise living things are carbon-based, and that they are thought
Medical Microbiology
 Lecture 5!!!!!!ƒš!!Œ!!! š!!œ!! Œ!!!! Dr. Ismail I. Daood Medical Microbiology!! Systematic Bacteriology Gram-Positive Cocci : GENUS : Staphylococcus : The general properties of Staphylococcus are Gram-
Lecture 5!!!!!!ƒš!!Œ!!! š!!œ!! Œ!!!! Dr. Ismail I. Daood Medical Microbiology!! Systematic Bacteriology Gram-Positive Cocci : GENUS : Staphylococcus : The general properties of Staphylococcus are Gram-
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2919 Direct observation of the influence of cardiolipin and antibiotics on lipid II binding to MurJ Jani
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants
 Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
Daptomycin-Resistant Enterococcus faecalis Diverts the Antibiotic Molecule from the Division Septum and Remodels Cell Membrane Phospholipids
 RESEARCH ARTICLE Daptomycin-Resistant Enterococcus faecalis Diverts the Antibiotic Molecule from the Division Septum and Remodels Cell Membrane Phospholipids Truc T. Tran, a,b Diana Panesso, a,c Nagendra
RESEARCH ARTICLE Daptomycin-Resistant Enterococcus faecalis Diverts the Antibiotic Molecule from the Division Septum and Remodels Cell Membrane Phospholipids Truc T. Tran, a,b Diana Panesso, a,c Nagendra
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
Supporting Information
 Supporting Information Dauvillée et al. 10.1073/pnas.0907424106 Fig. S1. Iodine screening of the C. cohnii mutant bank. Each single colony was grown on rich-medium agar plates then vaporized with iodine.
Supporting Information Dauvillée et al. 10.1073/pnas.0907424106 Fig. S1. Iodine screening of the C. cohnii mutant bank. Each single colony was grown on rich-medium agar plates then vaporized with iodine.
EFFECT OF SOME AMINO ACIDS ON THE GROWTH AND L-GLUTAMIC ACID FERMENTATION BY AN AUXOTROPHIC MUTANT Micrococcus glutamicus AB 100.
 S. Ganguly et. al. / International Journal on Pharmaceutical and Biomedical Research (IJPBR) Vol. 2(1), 2011, 21-25 EFFECT OF SOME AMINO ACIDS ON THE GROWTH AND L-GLUTAMIC ACID FERMENTATION BY AN AUXOTROPHIC
S. Ganguly et. al. / International Journal on Pharmaceutical and Biomedical Research (IJPBR) Vol. 2(1), 2011, 21-25 EFFECT OF SOME AMINO ACIDS ON THE GROWTH AND L-GLUTAMIC ACID FERMENTATION BY AN AUXOTROPHIC
Biology EOC Review. Saturday Session
 Biology EOC Review Saturday Session Cells DNA Ribosome Cytoplasm Cell Membrane Prokaryote Eukaryote Prokaryotic Bacteria Flagellum Cell Membrane (Plasma) Cell Wall Eukaryotic Animal Mitochondria Ribosome
Biology EOC Review Saturday Session Cells DNA Ribosome Cytoplasm Cell Membrane Prokaryote Eukaryote Prokaryotic Bacteria Flagellum Cell Membrane (Plasma) Cell Wall Eukaryotic Animal Mitochondria Ribosome
A Level. A Level Biology. Biological Molecules and Enzyme Questions. AQA, OCR, Edexcel. Name: Total Marks: Page 1
 AQA, OCR, Edexcel A Level A Level Biology Biological Molecules and Enzyme Questions Name: Total Marks: Page 1 Q1.The diagram represents part of the human digestive system. The organs are labelled A F.
AQA, OCR, Edexcel A Level A Level Biology Biological Molecules and Enzyme Questions Name: Total Marks: Page 1 Q1.The diagram represents part of the human digestive system. The organs are labelled A F.
Electron Transport Chain and Oxidative phosphorylation
 Electron Transport Chain and Oxidative phosphorylation So far we have discussed the catabolism involving oxidation of 6 carbons of glucose to CO 2 via glycolysis and CAC without any oxygen molecule directly
Electron Transport Chain and Oxidative phosphorylation So far we have discussed the catabolism involving oxidation of 6 carbons of glucose to CO 2 via glycolysis and CAC without any oxygen molecule directly
*For complete material(s) information, refer to
 Butler Community College Science, Technology, Engineering, and Math Division Robert Carlson New Fall 2017 Implemented Fall 2018 COURSE OUTLINE Biochemistry Course Description CH 275. Biochemistry. 4 hours
Butler Community College Science, Technology, Engineering, and Math Division Robert Carlson New Fall 2017 Implemented Fall 2018 COURSE OUTLINE Biochemistry Course Description CH 275. Biochemistry. 4 hours
Dental Research Institute, Faculty of Dentistry, University of Toronto, Toronto, Canada *For correspondence:
 Zymogram Assay for the Detection of Peptidoglycan Hydrolases in Streptococcus mutans Delphine Dufour and Céline M. Lévesque * Dental Research Institute, Faculty of Dentistry, University of Toronto, Toronto,
Zymogram Assay for the Detection of Peptidoglycan Hydrolases in Streptococcus mutans Delphine Dufour and Céline M. Lévesque * Dental Research Institute, Faculty of Dentistry, University of Toronto, Toronto,
Host Double Strand Break Repair Generates HIV-1 Strains Resistant to CRISPR/Cas9
 Host Double Strand Break Repair Generates HIV-1 Strains Resistant to CRISPR/Cas9 Kristine E. Yoder, a * and Ralf Bundschuh b a Department of Molecular Virology, Immunology and Medical Genetics, Center
Host Double Strand Break Repair Generates HIV-1 Strains Resistant to CRISPR/Cas9 Kristine E. Yoder, a * and Ralf Bundschuh b a Department of Molecular Virology, Immunology and Medical Genetics, Center
