CHAPTER 3. SILAC mouse-based screen to identify differentially expressed proteins in liver of induced Dicer knockout mice
|
|
|
- Elwin Blair
- 5 years ago
- Views:
Transcription
1 CHAPTER 3 SILAC mouse-based screen to identify differentially expressed proteins in liver of induced Dicer knockout mice 3.1 Introduction As mentioned earlier, few studies have been carried out to investigate proteome alteration associated with induced Dicer knockout. We have reported altered proteome of small intestine upon ablation of Dicer [49]. We utilized inducible Cre-loxP knockout system for deletion of Dicer. For the quantitative profiling of the liver proteome, we spiked SILAC labeled mouse liver. Previously, various in vitro labeling methods including DIGE, 18 O and itraq labeling have been adopted for quantitative proteomic profiling of mouse systems [50-55]. Gelbased quantitative proteomics approaches suffer from low reproducibility and hampered separation due to the limited pi range. Back exchange of 18 O isotope, compression of itraq ratios and probability of introduction of manual errors in labeling are other major limitations of the in vitro quantitation approaches. Label free quantitation has been used as an alternative quantitative proteomic approach. However, it requires highly reproducible LC-MS conditions, which are difficult to achieve especially across multiple sample runs [56]. Therefore, in vivo labeling strategies such as 15 N labeling and SILAC are the preferred approaches for quantitative proteomics [57, 58]. Challenges in achieving complete labeling and complexity of the quantitation data are some of the major limitations of the 15 N labeling. As a result, SILAC has been the method of choice for in vivo labeling. However, until recently, use of SILAC was limited to cell lines. With the development of 13 C 6 -Lysine enriched mice food, mice can be labeled in vivo, extending the application of SILAC to animal systems [49, 58, 22
2 59]. This quantitation approach is free of manual errors in labeling as proteins are completely labeled in vivo. We developed SILAC mice-based quantitative proteomics assay to identify the differentially expressed proteins upon depletion of Dicer in liver. We carried out high resolution mass spectrometry analysis and identified 2,137 proteins. We calculated ratio-of-ratios to identify relative quantitation of proteins. Amongst the 257 proteins upregulated in liver of induced Dicer knockout, we observed enrichment of proteins involved in fatty acid metabolism as the regulated subproteome by Dicer. We further carried out MRM assays to validate the candidate proteins which include peroxisomal bifunctional enzyme, phosphoenolpyruvate carboxykinase 1, Cyp3A13, Cyp3A41 and myristoylated alanine-rich protein kinase C substrate. Our findings highlight crucial roles of Dicer in regulation of proteins involved in lipid transport and metabolism in mouse liver. We propose that the SILAC mice-based proteomic screening coupled with MRM assays provide a robust pipeline for a systematic in vivo quantitative proteomic analysis. 23
3 3.2 Materials and Methods Generation of inducible Dicer knockout mouse We selected Cre-loxP system to generate inducible knockout mice. ROSA26- CreERT2 mice and mice with floxed Dicer exon 21 and 22 were crossed. The progeny was responsive to tamoxifen, resulting in deletion of floxed Dicer exon 21 and 22. ROSA26-CreERT2 mice were used as control. Mice were monitored daily for any obvious pathology. On the day 8 post induction, mice were starved for 3 hr. prior to sacrifice. Livers were harvested and washed with PBS to get rid of blood Generation of SILAC mice As described previously, SILAC mice were generated by feeding the stable isotope-labeled mouse food obtained from Cambridge Isotope Laboratories (Mouse Feed Labeling Kit, catalog number: MLK-LYS-C) [49, 60]. Briefly, twenty-eight days old female mouse (F0) was fed diet containing 13 C 6 -Lysine and mated 14 days later. Heavy diet was maintained until F1 pups were weaned. One F1 female mouse was kept on heavy diet and propagated to F2. Littermate female mice (F0) were fed light diet to generate unlabeled control mice. Labeling efficiency was monitored from blood, liver and lung specimens collected at 10 weeks of age from F0 and 4 weeks of age from the F1 generation Sample preparation and basic RPLC Liver tissues were excised from five uninduced and induced Dicer knockout mice. Tissues were lysed in 9 M urea and protein concentrations were measured using BCA assays. Uninduced and induced Dicer knockout liver lysates were 24
4 pooled and spiked with liver lysate from SILAC mouse at 2:1 (w/w) ratio. The spiked lysates were digested in solution using Lys-C protease. Lys-C was added at 1:50 (w/w) (Lysyl Endopeptidase Mass Spectrometry grade, Wako Chemical USA, Richmond, VA) to the lysates and incubated at 37 C for 4 hr. Additional Lys-C was added to the pre-digested lysate at 1:50 (w/w) ratio and incubated for 12 hr. Basic RPLC (brplc) was carried out as described previously [49]. Briefly, Lys-C digests were reconstituted in brplc solvent A (10 mm triethylammonium bicarbonate, ph 9.5) and were separated on XBridge BEH C18 Column (Waters, UK) with a linear increase in gradient from 5 to 100% of 10 mm TEABC with 90% acetonitrile (ph 9.5) over 30 min. and persisting for 10 minutes. For each condition, 24 fractions were collected and dried before LC- MS/MS analysis LC-MS/MS analysis LC-MS/MS analysis of 48 brplc fractions was carried out using Eksigent nano LC interfaced with the LTQ-Orbitrap XL ETD mass spectrometer (Thermo Scientific, San Jose, CA). The peptides were loaded on a trap column (75 µm 2 cm) packed with C18 material (5 μm Magic C18 AQ) at a flow rate of 5 μl/min of 97% solvent A (3% acetonitrile and 0.1% formic acid) and separated on an analytical column (75 µm 12 cm) packed with the same material using linear gradient of solvent B (0.1% formic acid in 90% acetonitrile) from 10 % to 60% solvent B for 60 min, to 97% solvent B from 74 to 90 min. All the MS spectra were acquired on an Orbitrap analyzer at the resolving power of 60,000 at 400 m/z while the data dependent MS/MS spectra were acquired using an LTQ analyzer. Ten most intense precursor ions from a survey scan within m/z 25
5 range from 350 to 1,800 above 1,000 of intensity were isolated with a 2 Da window and fragmented by CID with 30% normalized collision energy. The precursors were excluded, after fragmentation, for 30 seconds with a 7 ppm window. Maximum ion injection times were set to 500 msec for MS and 200 msec for MS/MS. The automatic gain control targets were set to for MS in the Orbitrap, for MS n in the LTQ Data analysis Mass spectrometry data analysis was carried out using Proteome Discoverer 1.2 suite (Thermo Fisher Scientific, Bremen, Germany). Precursor mass range of 300 to 5,000 Da and signal to noise ratio of 1.5 were used as selection criteria for generation of peak lists. NCBI RefSeq 42 containing mouse proteins with known contaminants (29,109 entries) was used as a reference database. SEQUEST and Mascot algorithms were used to carry out database searches. The parameters used for database searches include trypsin as a protease with allowed one missed cleavage, carbamidomethyl cysteine as a fixed modification and 13 C 6 -Lysine, oxidation of methionine as variable modifications. MS error window of 10 ppm and MS/MS error window of 0.8 Da were allowed. As described earlier, LC-MS/MS data was searched against a reversed database to calculate 1% false discovery rate score cut-off [61]. Ratio-of-ratio was calculated to determine differentially expressed proteins in induced Dicer knockout liver [59] MRM assays MRM assays were designed to detect and validate the protein level changes in induced Dicer knockout mice. Five differentially expressed proteins were selected as candidates for MRM analysis. Skyline v1.2 was used to create a 26
6 transition list of proteotypic peptides from the selected proteins [62]. Preference was given to proteotypic peptides with precursor charge of +2 that did not contain cysteine and methionine. In solution digestion of equal amounts of four uninduced and induced Dicer knockout mice liver samples was carried out and the peptides were acidified and stored at -80 C. All samples were analyzed in triplicate on TSQ Quantum Ultra (Thermo, San Jose, CA) interfaced with Agilent s 1100 series LC. Peptides were enriched on a trap column (5 μm, 75 μm 2 cm.) and separated using analytical column (3 μm, 75 μm 10 cm) with a linear gradient of 90% ACN 0.1% formic acid for 80 min at a constant flow rate of 350 nl/min. Both columns were packed in-house using Magic C18 AQ (Michrom Bioresources). Spray voltage of 2.5 kv was applied and ion transfer tube was maintained at 275 C. The collision energy for each transition was optimized based on the formula C.E. = (m/z) Data was acquired with Q1 and Q3 set at 0.4 and 0.7 unit mass resolutions, respectively Bioinformatics analysis for enriching candidate mirnas Proteins upregulated in the liver of induced Dicer knockout were taken as queries to search for the potential regulating mirnas. From TargetScan (Release 6.2), we downloaded the dataset, Summary Counts to identify seed sites in 3 UTR of mrnas and the representative target mirnas [63]. RefSeq was use as the reference database for mouse mrnas. Gene symbols were used to derive the intersection of the proteins and the predicted regulating mirnas. Only mirnas belonging to the species Mus musculus (mmu-mir) were used for analysis. 27
7 3.3 Results and Discussion Dicer is essential for survival of adult mice To assess the role of Dicer in adult mice and to circumvent embryonic lethality, we adopted an inducible Cre-loxP knockout system. The ubiquitous expression of Cre recombinase (Cre-ERT2) was driven by ROSA26 loci. The Cre-ERT2 mice were crossed to mice with floxed exons 21 and 22 of Dicer gene which encode the RNase III domain. Upon administration of tamoxifen, the Cre recombinase translocates to the nucleus deleting floxed exons 21 and 22. RT- PCR targeting the junctional region of exons 20 and 21 demonstrated that 80% of the mice had disrupted Dicer1. Mice were monitored daily for any obvious abnormalities. We observed that 5/10 mice developed diarrhea. The Dicer ablation proved to be lethal in most of the cases. As described earlier, by day 10 after inducing ablation of Dicer, 80% of the mice died [49]. The major histopathological changes observed by day 8 post induction of Dicer knockout included inflamed small intestine and bone marrow depletion with the reduced number of myeloid lineage cells. Dysregulation of lipid metabolism and proteomic changes associated with ablation of Dicer in small intestine were reported by our group earlier [49]. Liver performs crucial functions including metabolism of lipids, glucose and detoxification. Upon sacrifice of the Dicer knockout mice, we did not observe any gross abnormality of liver. The histological studies of the same specimens did not reveal any evident pathology. 28
8 Figure 3. SILAC mouse based quantitative proteomic workflow to identify differentially expressed proteins in liver of induced Dicer knockout mice. Liver lysates of five uninduced and five induced Dicer knockout mice were spiked at 2:1 ratio with liver lysates of two uninduced SILAC mice. Proteolysis was carried out using Lys-C protease. Digests were separated on brplc to obtain 24 fractions each for uninduced and induced Dicer knockout spiked liver lysates. LC-MS/MS analysis of the brplc fractions was carried out on an Orbitrap-XL ETD mass spectrometer. Database searches and quantitation was carried out using Proteome Discoverer (v 1.2) and ratio of ratio was calculated to identify differentially regulated proteins in the liver of induced Dicer knockout mice. Candidate upregulated proteins in induced Dicer knockout were validated using MRM assays. 29
9 3.3.2 Development of quantitative proteomic screen to identify the target Dicer proteome in mouse liver The aim of designing quantitative proteomic assay was to develop a suitable labeled internal standard that will allow systematic quantitation of proteins. SILAC mice have been previously used to explore proteomic changes associated with aging [59]. We developed SILAC mice-based screen as depicted in Figure 3. In order to minimize the inter-individual variation, we pooled liver lysates from five mice. As described in the methods, the SILAC mice were metabolically labeled by feeding 13 C 6 -Lysine labeled diet. As a consequence, all proteins were labeled in vivo with 13 C 6 -Lysine, making them amenable labeled controls. Uninduced and induced Dicer knockout liver lysates were spiked with liver lysate of SILAC mice at the 2:1 ratio. As generation and maintenance of SILAC mice is expensive, spiking of the SILAC mouse tissues provides as an economical alternative for in vivo quantitative proteomics. Proteolysis was carried out in-solution using Lys-C protease which specifically hydrolyses C-terminal of Lysine to acquire quantitation for the maximum number of peptides with paired 13 C 6 -Lysine containing peptides from SILAC mice. Reversed phase liquid chromatography operating at the macroflow rate in the high ph condition (brplc) was used to obtain 24 fractions per condition. The LC-MS/MS analysis was carried out on nanoflow reversed phase liquid chromatography coupled to LTQ-Orbitrap XL ETD mass spectrometer. MS was acquired at a high resolution (60,000 at 400 m/z) to obtain high accuracy quantitation information. In total, we analyzed 48 LC-MS/MS fractions. At better than 1% FDR, we identified 58,684 peptides corresponding to 2,134 proteins. 30
10 A Peroxisomal bifunctional enzyme (Ehhadh) B LGILDVVVK Phosphoenolpyruvate carboxykinase 1 (Pck1) GLGGVNVEELFGISK Relative Abundance Uninduced Induced knockout Light Heavy Light Relative Abundance m/z Heavy m/z y Relative Abundance Uninduced Light Heavy m/z 100 Relative Abundance Induced knockout Light y Heavy m/z y Relative Abundance b y y y7 y5 y b a4 b b m/z 800 y y Relative Abundance y b a y b7-nh y y b6-nh b m/z b y b b b b Figure 4. Representative MS and MS/MS spectra of upregulated proteins in liver induced Dicer knockout mice. A) MS and MS/MS of LGILDVVVK from upregulated peroxisomal bifunctional enzyme in liver of induced Dicer knockout mice. B) MS and MS/MS of GLGGVNVEELFGISK from upregulated phosphoenolpyruvate carboxykinase 1 in liver of induced Dicer knockout mice Differentially regulated proteins in induced Dicer knockout liver In uninduced mice liver, 1,412 proteins and in induced Dicer knockout mice liver, 1,993 proteins were profiled. We selected 1,217 proteins quantified in common and calculated ratio of SILAC ratios to obtain relative quantitation of proteins. We set an arbitrary threshold of 2-fold to select the differentially regulated proteins in induced Dicer knockout mouse liver. Out of the 319 differentially regulated proteins, 257 proteins were upregulated as listed in appendix I. 31
11 The upregulated proteins include aldehyde oxidase (4.5-fold), myristoylated alanine-rich protein kinase C substrate (Marcks) (4.3-fold), sorbin, SH3 domaincontaining protein 1 (3.4-fold), filamin A (3-fold) and phosphoenolpyruvate carboxykinase 1 (Pck1) (2.7-fold). Representative MS and MS/MS of upregulated proteins, peroxisomal bifunctional enzyme and phosphoenolpyruvate carboxykinase are illustrated in Figure 4. The downregulated proteins include 15-hydroxyprostaglandin dehydrogenase (6- fold), cytochrome P450 2C29 (5.5-fold) and alcohol dehydrogenase 4 (4.5-fold). Our dataset represents enrichment of cytochrome P450 family members which were upregulated in Dicer knockout liver such as cytochrome P450 2d22 (15.4- fold), cytochrome P450 3a41b (9.4-fold), cytochrome P450 4v3 (5.8-fold), cytochrome P450 3a13 (4.3-fold), cytochrome P450 2c70 (2.8-fold) and cytochrome P450 2a12 (2-fold). Cytochrome P450 is a family of heme proteins that function as oxygenases. These perform diverse functions including xenobiotic metabolism, fatty acid metabolism and steroid hormone synthesis. Upregulation of cytochrome P450 family of proteins can indicate elevation in the ROS. Association of the polymorphisms in cytochrome P450 family members and hepatocellular carcinoma has been established in previous studies [64, 65] PPARα targets are upregulated upon depletion of Dicer in liver To identify enriched classes of molecules, we analyzed the differentially regulated proteins using DAVID (v 6.7) [66]. Our dataset reflected enrichment of proteins involved in PPAR signaling. PPARs are steroid hormone nuclear receptors. There are multiple isoforms of PPAR with varying tissue expression. PPARα is the predominant isoform expressed in liver [67]. It is known to target 32
12 genes involved in fatty acid catabolism, gluconeogenesis and ketone body synthesis and liporotein assembly [68, 69]. Figure 5. Microsomal and peroxisomal lipid oxidation pathways Upregulated enzymes involved in ω - oxidation of fatty acids and peroxisomal β - oxidation of fatty acids in liver of induced Dicer knockout mice, most of which have been reported to be PPARα targets. Key enzymes involved in the steps of peroxisomal β-fatty acid oxidation included peroxisomal acyl-coenzyme A oxidase 2 (Acox2), peroxisomal multifunctional enzyme type 2 (Hsd17b4), peroxisomal bifunctional enzyme (Ehhadh) and peroxisomal 3- ketoacyl-coa thiolase A (Acaa1a). PPARα targets involved in lipid transport proteins were also upregulated in the liver of induced Dicer knockout mice. 33
13 One of the key pathways regulated by PPARα is fatty acid catabolism. As depicted in Figure 5, some of the major enzymatic steps involved in peroxisomal β -oxidation include oxidation, hydration, dehydrogenation and thiolytic cleavage of fatty acids. Acox2 acyl-coa oxidase was upregulated (2-fold) in induced Dicer knockout liver which acts upon bile acid intermediates and branched chain fatty acids. The second and third steps of β oxidation involve hydratase and 3-hydroxyacyl-CoaA dehydrogenase, for which peroxisome harbors hydroxysteroid (17-beta) dehydrogenase 4 or peroxisomal bifunctional enzyme type 2 and peroxisomal bifunctional enzyme which show overlapping functions. Hydroxysteroid (17-beta) dehydrogenase 4 was 1.4-fold upregulated while peroxisomal bifunctional enzyme was 3.7-fold upregulated in induced Dicer knockout liver. Peroxisomal bifunctional enzyme is involved in production of medium chain dicarboxylic acids. Acaa1a is a peroxisomal 3- ketoacyl-coa thiolase A which performs the last step of β oxidation. It was 4.1- fold upregulated in induced Dicer knockout liver. The substrates for peroxisomal β oxidation are end products of ω-oxidation of fatty acids which is initiated by microsomal Cyp4 family of cytochromes. Cyp4V3 of mouse is a homolog of human Cyp4V2 which functions as fatty acid ω-hydroxylase. Cyp4V3 was 5.8-fold upregulated in the Dicer knockout liver. Though Cyp4V3 is not reported to be a direct target of PPARα, upregulation of Cyp4V3 along with the enzymes involved in peroxisomal β oxidation, indicate over activation of oxidation of fatty acids. These events might have led to the observed loss of weight in the induced Dicer knockout mice. We also identified several other PPARα targets which were upregulated in. Other upregulated PPARα targets in induced Dicer knockout liver include proteins involved in lipid transport such as ATP-binding cassette sub-family D member 3 (11-fold), 34
14 apolipoprotein A-IV (3.9-fold), perilipin 2 (3.4-fold) and solute carrier family 27 (fatty acid transporter), member 2 (2.3-fold). Representative proteins involved in lipid metabolism or transport which were upregulated in liver of induced Dicer knockout mice are listed in Table1. As described earlier, down-regulation of Dicer is reported to be associated with hepatocellular carcinoma [45, 46]. PPARα pathway is also observed to be associated with hepatocellular carcinoma [70]. In our study, PPARα targets were upregulated upon depletion of Dicer. We also observed over-expression of cytochrome P450 family of proteins which can be attributed to elevation of ROS stress due to over activation of microsomal and peroxisomal fatty acid oxidation. The chronic effect of Dicer depletion on the PPAR signaling and development of hepatocellular carcinoma needs to be further explored. Protein Gene symbol KO/Ctrl 1. Sodium/bile acid cotransporter Slc10a ATP-binding cassette sub-family D member 3 Abcd Cytochrome P450 4V3 Cyp4v Peroxisomal 3-ketoacyl-CoA thiolase A Acaa1a Peroxisomal bifunctional enzyme Ehhadh Perilipin-2 Plin Peroxisomal acyl-coenzyme A oxidase 2 Acox2 2.0 Table 1. A partial list of proteins involved in lipid metabolism List of proteins, gene symbols and fold-change upregulation in liver of induced Dicer knockout mice (KO) compared to uninduced Dicer knockout mice (Ctrl) Validation of candidate targets of Dicer using MRM assays To validate upregulated proteins in liver of induced Dicer knockout mice using a complementary mass spectrometry-based method, we designed MRM assays for peroxisomal bifunctional enzyme (Ehhadh), phosphoenolpyruvate carboxykinase (Pck1), Cyp3a41, Cyp3a13 and myristoylated alanine-rich C- 35
15 kinase substrate (Marcks). We selected 4 uninduced and 4 induced Dicer knockout mice. At least four transitions were monitored for a minimum of one unique peptide for each protein. For normalization of data, housekeeping protein, isocitrate dehydrogenase (Idh2) was selected as an unchanged internal standard from LC-MS/MS data. We monitored LGILDVVVK (z = +2, m/z = 478.3) and GWYQYDKPLGR (z = +2, m/z = 691.8) for validation of Ehhadh, a PPARα target which is involved in peroxisomal β-oxidation of fatty acids. As depicted in Figure 6, LGILDVVVK was 5.2-fold upregulated in liver of induced Dicer knockout mice corroborating our findings. Similarly as listed in Table 2, we also monitored LTPIGYIPK (z = +2, m/z = 501.3), LQEEIDETLPNK (z = +2, m/z 714.8), LQDEIDAALPNK (z = +2, m/z = 663.8), VNGDASPAAAEPGAK (z = +2, m/z: 677.8) corresponding to Pck1, Cyp3a41, Cyp3a13 and Marcks, respectively. Quantitation obtained using MRM-based assays was in agreement with the quantitation obtained from the LC-MS/MS data. We observed that MRM provided as a robust and antibody-independent approach for validation of differentially expressed proteins in our study. Protein 1 Peroxisomal bifunctional enzyme 2 Phosphoenolpyruvate carboxykinase 1 3 Cytochrome P450 3A41 4 Cytochrome P450 3A13 5 Myristoylated alaninerich C-kinase substrate Gene symbol Peptide Induced Dicer knockout / uninduced (MRM) Induced Dicer knockout / uninduced (SILAC spike) Ehhadh LGILDVVVK GWYQYDKPLGR 8.7 Pck1 LTPIGYIPK Cyp3a41 LQEEIDETLPNK b Cyp3a13 LQDEIDAALPNK Marcks VNGDASPAAAEPGA K
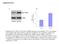
16 Table 2. List of validated proteins upregulated in induced Dicer knockout using MRM assays List of proteins, gene symbols, MRM-based quantitation and relative fold-change upregulation data from spiked SILAC mice LC-MS/MS analysis Figure 6. MRM-based validation of upregulated proteins in liver of induced Dicer knockout mice. A) Four transitions were monitored for LGILDVVVK from peroxisomal bifunctional enzyme in uninduced and induced Dicer knockout mice. Box and whiskers plot depict relative abundance in the uninduced and induced liver knockout mice. B) Four transitions were monitored for LQEEIDETLPNK of phosphoenolpyruvate carboxykinase 1 in liver of induced Dicer knockout mice. Box and whiskers plot show relative abundance in the uninduced and induced liver knockout mice Bioinformatics analysis of the enriched upstream mirnas. 37
17 Dicer-dependent mirna generation is the predominant pathway in maturation of mirnas. We hypothesized that upon depletion of Dicer; mirna maturation is severely affected, resulting in dysregulation of downstream mirna targets. Thus, we carried out a bioinformatics analysis using TargetScan algorithm to identify the upstream mirnas of the upregulated proteins in liver of induced Dicer knockout mice. We identified 598 unique mouse mirnas (mmu-mirs) predicted to be regulators of the upregulated proteins in induced Dicer knockout liver. Some of the predicted upstream mirnas targeting multiple upregulated proteins included mmu-mir-124 and mmu-mir-200b. mir-124 has been described as a tumor suppressor gene in recent studies carried out on hepatocellular carcinoma [71, 72]. Vimentin (2-fold upregulated), an EMT marker has been demonstrated previously as direct target of mir-124 [72]. Apart from vimentin, multiple upregulated proteins associated with actin binding and membrane trafficking are predicted targets of mir-124. mir-143 was identified as upstream mirna of multiple proteins involved in lipid metabolism or transport which included peroxisomal bifunctional enzyme, Abcd1 and perilipin 2. Marcks was also predicted as a target of mir-143. mir-143 is known to be involved in regulation of lipid metabolism [73, 74]. It induces adipocyte differentiation with accumulation of triglycerides. We hypothesize that upon downregulation of mir-143, the target genes involved in fatty acid oxidation are upregulated. Further studies are required to investigate functions of these candidate mirnas and their targets in liver. Also, studies need to be carried out to identify the correlation of downregulation of Dicer, mirnas, PPARα downstream targets possibly with carcinogenesis. 38
Quantification with Proteome Discoverer. Bernard Delanghe
 Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
The distribution of log 2 ratio (H/L) for quantified peptides. cleavage sites in each bin of log 2 ratio of quantified. peptides
 Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Quantitative chromatin proteomics reveals a dynamic histone. post-translational modification landscape that defines asexual
 Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes
 Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev
 Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS
 Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS Jenny Chen, Hongxia Wang, Zhiqi Hao, Patrick Bennett, and Greg Kilby Thermo Fisher
Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS Jenny Chen, Hongxia Wang, Zhiqi Hao, Patrick Bennett, and Greg Kilby Thermo Fisher
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Mass Spectrometry. Mass spectrometer MALDI-TOF ESI/MS/MS. Basic components. Ionization source Mass analyzer Detector
 Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry
 Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry
 Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
NIH Public Access Author Manuscript J Proteome Res. Author manuscript; available in PMC 2014 July 05.
 NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
Unsupervised Identification of Isotope-Labeled Peptides
 Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Supporting information
 Supporting information A novel lipidomics workflow for improved human plasma identification and quantification using RPLC-MSn methods and isotope dilution strategies Evelyn Rampler 1,2,3, Angela Criscuolo
Supporting information A novel lipidomics workflow for improved human plasma identification and quantification using RPLC-MSn methods and isotope dilution strategies Evelyn Rampler 1,2,3, Angela Criscuolo
Supporting information
 Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Shotgun Proteomics MS/MS. Protein Mixture. proteolysis. Peptide Mixture. Time. Abundance. Abundance. m/z. Abundance. m/z 2. Abundance.
 Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Sequence Coverage (%) Profilin-1 P UD 2
 Protein Name Accession Number (Swissprot) Sequence Coverage (%) No. of MS/MS Queries Mascot Score 1 Reference Cytoskeletal proteins Beta-actin P60709 37 14 298 Alpha-actin P68032 33 10 141 20 Beta-actin-like
Protein Name Accession Number (Swissprot) Sequence Coverage (%) No. of MS/MS Queries Mascot Score 1 Reference Cytoskeletal proteins Beta-actin P60709 37 14 298 Alpha-actin P68032 33 10 141 20 Beta-actin-like
O O H. Robert S. Plumb and Paul D. Rainville Waters Corporation, Milford, MA, U.S. INTRODUCTION EXPERIMENTAL. LC /MS conditions
 Simplifying Qual/Quan Analysis in Discovery DMPK using UPLC and Xevo TQ MS Robert S. Plumb and Paul D. Rainville Waters Corporation, Milford, MA, U.S. INTRODUCTION The determination of the drug metabolism
Simplifying Qual/Quan Analysis in Discovery DMPK using UPLC and Xevo TQ MS Robert S. Plumb and Paul D. Rainville Waters Corporation, Milford, MA, U.S. INTRODUCTION The determination of the drug metabolism
Quantification of PtdInsP 3 molecular species in cells and tissues by mass spectrometry
 Nature Methods Quantification of PtdInsP 3 molecular species in cells and tissues by mass spectrometry Jonathan Clark, Karen E Anderson, Veronique Juvin, Trevor S Smith, Fredrik Karpe, Michael J Wakelam,
Nature Methods Quantification of PtdInsP 3 molecular species in cells and tissues by mass spectrometry Jonathan Clark, Karen E Anderson, Veronique Juvin, Trevor S Smith, Fredrik Karpe, Michael J Wakelam,
Extended Mass Range Triple Quadrupole for Routine Analysis of High Mass-to-charge Peptide Ions
 Extended Mass Range Triple Quadrupole for Routine Analysis of High Mass-to-charge Peptide Ions Application Note Targeted Proteomics Authors Linfeng Wu, Christine A. Miller, Jordy Hsiao, Te-wei Chu, Behrooz
Extended Mass Range Triple Quadrupole for Routine Analysis of High Mass-to-charge Peptide Ions Application Note Targeted Proteomics Authors Linfeng Wu, Christine A. Miller, Jordy Hsiao, Te-wei Chu, Behrooz
LC/MS/MS SOLUTIONS FOR LIPIDOMICS. Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS
 LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast
 Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Characterization of an Unknown Compound Using the LTQ Orbitrap
 Characterization of an Unknown Compound Using the LTQ rbitrap Donald Daley, Russell Scammell, Argenta Discovery Limited, 8/9 Spire Green Centre, Flex Meadow, Harlow, Essex, CM19 5TR, UK bjectives unknown
Characterization of an Unknown Compound Using the LTQ rbitrap Donald Daley, Russell Scammell, Argenta Discovery Limited, 8/9 Spire Green Centre, Flex Meadow, Harlow, Essex, CM19 5TR, UK bjectives unknown
MS1 and MS2 crosstalk in label free quantitation of mass spectrometry data independent acquisitions
 MS1 and MS2 crosstalk in label free quantitation of mass spectrometry data independent acquisitions MS1 528.18 +++ m/z 568.98 ++ m/z 678.34 ++ m/z MS2/SWATH June 9th, 2013 Matthew J. Rardin SIRT3 regulated
MS1 and MS2 crosstalk in label free quantitation of mass spectrometry data independent acquisitions MS1 528.18 +++ m/z 568.98 ++ m/z 678.34 ++ m/z MS2/SWATH June 9th, 2013 Matthew J. Rardin SIRT3 regulated
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan
 PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
for the Identification of Phosphorylated Peptides
 Application of a Data Dependent Neutral-Loss Experiment on the Finnigan LTQ for the Identification of Phosphorylated Peptides Gargi Choudhary Diane Cho Thermo Electron, San Jose, CA Abstracted from posters
Application of a Data Dependent Neutral-Loss Experiment on the Finnigan LTQ for the Identification of Phosphorylated Peptides Gargi Choudhary Diane Cho Thermo Electron, San Jose, CA Abstracted from posters
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring
 Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Quantitative Analysis of Vit D Metabolites in Human Plasma using Exactive System
 Quantitative Analysis of Vit D Metabolites in Human Plasma using Exactive System Marta Kozak Clinical Research Applications Group Thermo Fisher Scientific San Jose CA Clinical Research use only, Not for
Quantitative Analysis of Vit D Metabolites in Human Plasma using Exactive System Marta Kozak Clinical Research Applications Group Thermo Fisher Scientific San Jose CA Clinical Research use only, Not for
Phospholipid characterization by a TQ-MS data based identification scheme
 P-CN1716E Phospholipid characterization by a TQ-MS data based identification scheme ASMS 2017 MP-406 Tsuyoshi Nakanishi 1, Masaki Yamada 1, Ningombam Sanjib Meitei 2, 3 1 Shimadzu Corporation, Kyoto, Japan,
P-CN1716E Phospholipid characterization by a TQ-MS data based identification scheme ASMS 2017 MP-406 Tsuyoshi Nakanishi 1, Masaki Yamada 1, Ningombam Sanjib Meitei 2, 3 1 Shimadzu Corporation, Kyoto, Japan,
REDOX PROTEOMICS. Roman Zubarev.
 REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
SIEVE 2.1 Proteomics Example
 SIEVE 2.1 Proteomics Example Software Overview What is SIEVE? SIEVE is Thermo Scientific s differential software solution. SIEVE will continue to enhance our current product for label-free differential
SIEVE 2.1 Proteomics Example Software Overview What is SIEVE? SIEVE is Thermo Scientific s differential software solution. SIEVE will continue to enhance our current product for label-free differential
Learning Objectives. Overview of topics to be discussed 10/25/2013 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS
 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
4.2 RESULTS AND DISCUSSION
 phosphorylated proteins on treatment with Erlotinib.This chapter describes the finding of the SILAC experiment. 4.2 RESULTS AND DISCUSSION This is the first global report of its kind using dual strategies
phosphorylated proteins on treatment with Erlotinib.This chapter describes the finding of the SILAC experiment. 4.2 RESULTS AND DISCUSSION This is the first global report of its kind using dual strategies
Dr. Erin E. Chambers Waters Corporation. Presented by Dr. Diego Rodriguez Cabaleiro Waters Europe Waters Corporation 1
 Development of an SPE-LC/MS/MS Assay for the Simultaneous Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid in Support of Alzheimer s Research Dr. Erin E. Chambers Waters Corporation Presented
Development of an SPE-LC/MS/MS Assay for the Simultaneous Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid in Support of Alzheimer s Research Dr. Erin E. Chambers Waters Corporation Presented
Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform
 Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform Abstract Targeted proteomics for biomarker verification/validation
Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform Abstract Targeted proteomics for biomarker verification/validation
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Supporting Information
 Supporting Information Development of a High Coverage Pseudotargeted Lipidomics Method Based on Ultra-High Performance Liquid Chromatography-Mass Spectrometry Qiuhui Xuan 1,2#, Chunxiu Hu 1#, Di Yu 1,2,
Supporting Information Development of a High Coverage Pseudotargeted Lipidomics Method Based on Ultra-High Performance Liquid Chromatography-Mass Spectrometry Qiuhui Xuan 1,2#, Chunxiu Hu 1#, Di Yu 1,2,
4-Plex itraq Based Quantitative Proteomic Analysis Using an Agilent Accurate -Mass Q-TOF
 4-Plex itraq Based Quantitative Proteomic Analysis Using an Agilent Accurate -Mass Q-TOF Application Note Authors H. C. Harsha, G. S. S. Kumar, and A. Pandey Institute of Bioinformatics Bangalore India
4-Plex itraq Based Quantitative Proteomic Analysis Using an Agilent Accurate -Mass Q-TOF Application Note Authors H. C. Harsha, G. S. S. Kumar, and A. Pandey Institute of Bioinformatics Bangalore India
LECTURE-15. itraq Clinical Applications HANDOUT. Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS
 LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature13418 Supplementary Results: USP30 opposes autophagic flux In HEK-293 cells, USP30 overexpression increased basal LC3-II levels, dependent on enzymatic activity,
SUPPLEMENTARY INFORMATION doi:10.1038/nature13418 Supplementary Results: USP30 opposes autophagic flux In HEK-293 cells, USP30 overexpression increased basal LC3-II levels, dependent on enzymatic activity,
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow
 Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
New Solvent Grade Targeted for Trace Analysis by UHPLC-MS
 New Solvent Grade Targeted for Trace Analysis by UHPLC-MS Subhra Bhattacharya, Deva H. Puranam, and Stephen C. Roemer Thermo Fisher Scientific Fisher Chemical, One Reagent Lane, Fair Lawn, NJ Material
New Solvent Grade Targeted for Trace Analysis by UHPLC-MS Subhra Bhattacharya, Deva H. Puranam, and Stephen C. Roemer Thermo Fisher Scientific Fisher Chemical, One Reagent Lane, Fair Lawn, NJ Material
Edgar Naegele. Abstract
 Simultaneous determination of metabolic stability and identification of buspirone metabolites using multiple column fast LC/TOF mass spectrometry Application ote Edgar aegele Abstract A recent trend in
Simultaneous determination of metabolic stability and identification of buspirone metabolites using multiple column fast LC/TOF mass spectrometry Application ote Edgar aegele Abstract A recent trend in
Screening and Speciation of Raw and Processed Meat Products
 vmethod Application for Food Testing Screening and Speciation of Raw and Processed Meat Products A Selective and Robust LC-MS/MS Method for Multiple Meat Speciation and Authentication on the QTRAP 4500
vmethod Application for Food Testing Screening and Speciation of Raw and Processed Meat Products A Selective and Robust LC-MS/MS Method for Multiple Meat Speciation and Authentication on the QTRAP 4500
Using Multiple Mass Defect Filters and Higher Energy Collisional Dissociation on an LTQ Orbitrap XL for Fast, Sensitive and Accurate Metabolite ID
 Application ote: 417 Using Multiple Mass Defect Filters and Higher Energy Collisional Dissociation on an LTQ rbitrap XL for Fast, Sensitive and Accurate Metabolite ID Yingying Huang 1, Shirley Liu 2, Shichang
Application ote: 417 Using Multiple Mass Defect Filters and Higher Energy Collisional Dissociation on an LTQ rbitrap XL for Fast, Sensitive and Accurate Metabolite ID Yingying Huang 1, Shirley Liu 2, Shichang
Lehninger 5 th ed. Chapter 17
 Lehninger 5 th ed. Chapter 17 December 26, 2010 Prof. Shimon Schuldiner Email: Shimon.Schuldiner@huji.ac.il Phone: 6585992 CHAPTER 17 Fatty Acid Catabolism Key topics: How fats are digested in animals
Lehninger 5 th ed. Chapter 17 December 26, 2010 Prof. Shimon Schuldiner Email: Shimon.Schuldiner@huji.ac.il Phone: 6585992 CHAPTER 17 Fatty Acid Catabolism Key topics: How fats are digested in animals
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
MS/MS as an LC Detector for the Screening of Drugs and Their Metabolites in Race Horse Urine
 Application Note: 346 MS/MS as an LC Detector for the Screening of Drugs and Their Metabolites in Race Horse Urine Gargi Choudhary and Diane Cho, Thermo Fisher Scientific, San Jose, CA Wayne Skinner and
Application Note: 346 MS/MS as an LC Detector for the Screening of Drugs and Their Metabolites in Race Horse Urine Gargi Choudhary and Diane Cho, Thermo Fisher Scientific, San Jose, CA Wayne Skinner and
Figure S6. A-J) Annotated UVPD mass spectra for top ten peptides found among the peptides identified by Byonic but not SEQUEST + Percolator.
 Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Quantitative LC-MS/MS Analysis of Glucagon. Veniamin Lapko, Ph.D June 21, 2011
 Quantitative LC-MS/MS Analysis of Glucagon Veniamin Lapko, Ph.D June 21, 2011 Contents Comparison with small molecule LC-MS/MS LC-MS/MS sensitivity of peptides detection Stability: neat vs. matrix solutions
Quantitative LC-MS/MS Analysis of Glucagon Veniamin Lapko, Ph.D June 21, 2011 Contents Comparison with small molecule LC-MS/MS LC-MS/MS sensitivity of peptides detection Stability: neat vs. matrix solutions
New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics
 New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics July.3.13 Ken Miller Vice President of Marketing, Life Sciences Mass Spectrometry 1 The world leader in serving science Omics & the
New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics July.3.13 Ken Miller Vice President of Marketing, Life Sciences Mass Spectrometry 1 The world leader in serving science Omics & the
Comparison of mass spectrometers performances
 Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Shotgun metaproteomics of the human distal gut microbiota. Present by Lei Chen
 Shotgun metaproteomics of the human distal gut microbiota Present by Lei Chen (lc6@indana.edu) Outline Background What are the goals? Materials and Methods Results Discussion Background The human gastrointestinal
Shotgun metaproteomics of the human distal gut microbiota Present by Lei Chen (lc6@indana.edu) Outline Background What are the goals? Materials and Methods Results Discussion Background The human gastrointestinal
Michal Godula Thermo Fisher Scientific. The world leader in serving science
 Resolving the food authenticity challenges using advanced isotopic ratio and Thermo Scientific Orbitrap high resolution mass spectrometry tools in practice Michal Godula Thermo Fisher Scientific The world
Resolving the food authenticity challenges using advanced isotopic ratio and Thermo Scientific Orbitrap high resolution mass spectrometry tools in practice Michal Godula Thermo Fisher Scientific The world
Increased Identification Coverage and Throughput for Complex Lipidomes
 Increased Identification Coverage and Throughput for Complex Lipidomes Reiko Kiyonami, David Peake, Yingying Huang, Thermo Fisher Scientific, San Jose, CA USA Application Note 607 Key Words Q Exactive
Increased Identification Coverage and Throughput for Complex Lipidomes Reiko Kiyonami, David Peake, Yingying Huang, Thermo Fisher Scientific, San Jose, CA USA Application Note 607 Key Words Q Exactive
Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry
 Bruker Daltonics Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry Abstract Analysis of the N-glycosylation pattern on
Bruker Daltonics Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry Abstract Analysis of the N-glycosylation pattern on
Rapid, Simple Impurity Characterization with the Xevo TQ Mass Spectrometer
 Robert Plumb, Michael D. Jones, and Marian Twohig Waters Corporation, Milford, MA, USA INTRODUCTION The detection and characterization of impurities and degradation products of an active pharmaceutical
Robert Plumb, Michael D. Jones, and Marian Twohig Waters Corporation, Milford, MA, USA INTRODUCTION The detection and characterization of impurities and degradation products of an active pharmaceutical
New Instruments and Services
 New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance
 Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
MASS SPECTROMETRY BASED METABOLOMICS. Pavel Aronov. ABRF2010 Metabolomics Research Group March 21, 2010
 MASS SPECTROMETRY BASED METABOLOMICS Pavel Aronov ABRF2010 Metabolomics Research Group March 21, 2010 Types of Experiments in Metabolomics targeted non targeted Number of analyzed metabolites is limited
MASS SPECTROMETRY BASED METABOLOMICS Pavel Aronov ABRF2010 Metabolomics Research Group March 21, 2010 Types of Experiments in Metabolomics targeted non targeted Number of analyzed metabolites is limited
Profiling the Distribution of N-Glycosylation in Therapeutic Antibodies using the QTRAP 6500 System
 Profiling the Distribution of N-Glycosylation in Therapeutic Antibodies using the QTRAP 6500 System Scheduled MRM Pro Algorithm for Increased Efficiency of Targeted Detection Jenny Albanese 1, Christie
Profiling the Distribution of N-Glycosylation in Therapeutic Antibodies using the QTRAP 6500 System Scheduled MRM Pro Algorithm for Increased Efficiency of Targeted Detection Jenny Albanese 1, Christie
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid
 Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
Biologic Oxidation BIOMEDICAL IMPORTAN
 Biologic Oxidation BIOMEDICAL IMPORTAN Chemically, oxidation is defined as the removal of electrons and reduction as the gain of electrons. Thus, oxidation is always accompanied by reduction of an electron
Biologic Oxidation BIOMEDICAL IMPORTAN Chemically, oxidation is defined as the removal of electrons and reduction as the gain of electrons. Thus, oxidation is always accompanied by reduction of an electron
Robust extraction, separation, and quantitation of structural isomer steroids from human plasma by SPE-UHPLC-MS/MS
 TECHNICAL NOTE 21882 Robust extraction, separation, and quantitation of structural isomer steroids human plasma by SPE-UHPLC-MS/MS Authors Jon Bardsley 1, Kean Woodmansey 1, and Stacy Tremintin 2 1 Thermo
TECHNICAL NOTE 21882 Robust extraction, separation, and quantitation of structural isomer steroids human plasma by SPE-UHPLC-MS/MS Authors Jon Bardsley 1, Kean Woodmansey 1, and Stacy Tremintin 2 1 Thermo
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/407/ra127/dc1 Supplementary Materials for Loss of FTO in adipose tissue decreases Angptl4 translation and alters triglyceride metabolism Chao-Yung Wang,* Shian-Sen
www.sciencesignaling.org/cgi/content/full/8/407/ra127/dc1 Supplementary Materials for Loss of FTO in adipose tissue decreases Angptl4 translation and alters triglyceride metabolism Chao-Yung Wang,* Shian-Sen
SUPPLEMENTAL DATA AGING, April 2013, Vol.5 No.4
 SUPPLEMENTAL DATA Figure S1. Under CR conditions, the atg32δ mutation elevates the extent of oxidative damage to proteins. WT and atg32δ strains were cultured in the nutrient rich YP medium initially containing
SUPPLEMENTAL DATA Figure S1. Under CR conditions, the atg32δ mutation elevates the extent of oxidative damage to proteins. WT and atg32δ strains were cultured in the nutrient rich YP medium initially containing
* Skyline LC SRM. 2 Skyline LC-MS/MS. Skyline University of Washington Windows.
 Proteome Letters 2016 1 89-94 * *E-mail: fmatsuda@ist.osaka-u.ac.jp 565-0871 1-5 2016 4 20 2016 5 26 2016 5 27 LC SRM LC SRM 1 LC-MS/MS LC 1 3 SRM SRM LC-MS Skyline SRM LC-MS LC SRM 2 Skyline Skyline University
Proteome Letters 2016 1 89-94 * *E-mail: fmatsuda@ist.osaka-u.ac.jp 565-0871 1-5 2016 4 20 2016 5 26 2016 5 27 LC SRM LC SRM 1 LC-MS/MS LC 1 3 SRM SRM LC-MS Skyline SRM LC-MS LC SRM 2 Skyline Skyline University
Supporting Information: Protein Corona Analysis of Silver Nanoparticles Exposed to Fish Plasma
 Supporting Information: Protein Corona Analysis of Silver Nanoparticles Exposed to Fish Plasma Jiejun Gao 1, Lu Lin 2, Alexander Wei 2,*, and Maria S. Sepúlveda 1,* 1 Department of Forestry and Natural
Supporting Information: Protein Corona Analysis of Silver Nanoparticles Exposed to Fish Plasma Jiejun Gao 1, Lu Lin 2, Alexander Wei 2,*, and Maria S. Sepúlveda 1,* 1 Department of Forestry and Natural
More structural information with MS n
 PRODUCT SPECIFICATIONS The LTQ XL linear ion trap mass spectrometer More structural information with MS n The LTQ XL linear ion trap mass spectrometer delivers more structural information faster and with
PRODUCT SPECIFICATIONS The LTQ XL linear ion trap mass spectrometer More structural information with MS n The LTQ XL linear ion trap mass spectrometer delivers more structural information faster and with
2.5. AMPK activity
 Supplement Fig. A 3 B phos-ampk 2.5 * Control AICAR AMPK AMPK activity (Absorbance at 45 nm) 2.5.5 Control AICAR Supplement Fig. Effects of AICAR on AMPK activation in macrophages. J774. macrophages were
Supplement Fig. A 3 B phos-ampk 2.5 * Control AICAR AMPK AMPK activity (Absorbance at 45 nm) 2.5.5 Control AICAR Supplement Fig. Effects of AICAR on AMPK activation in macrophages. J774. macrophages were
Multiple Reaction Monitoring for Direct Quantitation of Intact Proteins using a Triple Quadrupole Mass Spectrometer
 ELECTRONIC SUPPLEMENTARY INFORMATION Multiple Reaction Monitoring for Direct Quantitation of Intact Proteins using a Triple Quadrupole Mass Spectrometer Evelyn H. Wang 1, Peter C. Combe 2, and Kevin A.
ELECTRONIC SUPPLEMENTARY INFORMATION Multiple Reaction Monitoring for Direct Quantitation of Intact Proteins using a Triple Quadrupole Mass Spectrometer Evelyn H. Wang 1, Peter C. Combe 2, and Kevin A.
Neosolaniol. [Methods listed in the Feed Analysis Standards]
![Neosolaniol. [Methods listed in the Feed Analysis Standards] Neosolaniol. [Methods listed in the Feed Analysis Standards]](/thumbs/86/94003482.jpg) Neosolaniol [Methods listed in the Feed Analysis Standards] 1 Simultaneous analysis of mycotoxins by liquid chromatography/ tandem mass spectrometry [Feed Analysis Standards, Chapter 5, Section 1 9.1 ]
Neosolaniol [Methods listed in the Feed Analysis Standards] 1 Simultaneous analysis of mycotoxins by liquid chromatography/ tandem mass spectrometry [Feed Analysis Standards, Chapter 5, Section 1 9.1 ]
Conflict of Interest Statement
 Utilizing an Accurate Mass and Retention Time Library to Facilitate Biomarker Discovery Nichole Reisdorph, PhD and Cole Michel, BS Mass Spectrometry Facility Skaggs School of Pharmacy and Pharmaceutical
Utilizing an Accurate Mass and Retention Time Library to Facilitate Biomarker Discovery Nichole Reisdorph, PhD and Cole Michel, BS Mass Spectrometry Facility Skaggs School of Pharmacy and Pharmaceutical
Impurity Identification using a Quadrupole - Time of Flight Mass Spectrometer QTOF
 Impurity Identification using a Quadrupole - Time of Flight Mass Spectrometer QTOF PUSHER TOF DETECTOR ZSPRAY TM Ion Source SAMPLING CONE SKIMMER RF HEXAPOLE RF HEXAPOLE QUADRUPOLE IN NARROW BANDPASS MODE
Impurity Identification using a Quadrupole - Time of Flight Mass Spectrometer QTOF PUSHER TOF DETECTOR ZSPRAY TM Ion Source SAMPLING CONE SKIMMER RF HEXAPOLE RF HEXAPOLE QUADRUPOLE IN NARROW BANDPASS MODE
TECHNICAL NOTE. Accurate and fast proteomics analysis of human plasma with PlasmaDive and SpectroDive
 TECHNICAL NOTE Accurate and fast proteomics analysis of human plasma with PlasmaDive and SpectroDive In this technical note you will learn about: Step-by-step set-up of parallel reaction monitoring (PRM)
TECHNICAL NOTE Accurate and fast proteomics analysis of human plasma with PlasmaDive and SpectroDive In this technical note you will learn about: Step-by-step set-up of parallel reaction monitoring (PRM)
APPLICATION NOTE. Highly reproducible and Comprehensive Proteome Profiling of Formalin-Fixed Paraffin-Embedded (FFPE) Tissues Slices
 APPLICATION NOTE Highly reproducible and Comprehensive Proteome Profiling of Formalin-Fixed Paraffin-Embedded (FFPE) Tissues Slices INTRODUCTION Preservation of tissue biopsies is a critical step to This
APPLICATION NOTE Highly reproducible and Comprehensive Proteome Profiling of Formalin-Fixed Paraffin-Embedded (FFPE) Tissues Slices INTRODUCTION Preservation of tissue biopsies is a critical step to This
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging
 Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Advances in Hybrid Mass Spectrometry
 The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
LC-MS/MS Method for the Determination of 21 Opiates and Opiate Derivatives in Urine
 LC-MS/MS Method for the Determination of 21 Opiates and Opiate Derivatives in Urine J. Jones, S. Westwood, T. Liddicoat, L. Pereira, T. Edge Thermo Fisher Scientific, Manor Park, Runcorn, UK Overview Purpose:
LC-MS/MS Method for the Determination of 21 Opiates and Opiate Derivatives in Urine J. Jones, S. Westwood, T. Liddicoat, L. Pereira, T. Edge Thermo Fisher Scientific, Manor Park, Runcorn, UK Overview Purpose:
INTRODUCTION CH 3 CH CH 3 3. C 37 H 48 N 6 O 5 S 2, molecular weight Figure 1. The Xevo QTof MS System.
 Fast and Sensitive in vitro Metabolism Study of Rate and Routes of Clearance for Ritonavir using UPLC CUPLED with the Xevo QTof MS System Jose Castro-Perez, Kate Yu, John Shockcor, Henry Shion, Emma Marsden-Edwards,
Fast and Sensitive in vitro Metabolism Study of Rate and Routes of Clearance for Ritonavir using UPLC CUPLED with the Xevo QTof MS System Jose Castro-Perez, Kate Yu, John Shockcor, Henry Shion, Emma Marsden-Edwards,
Mass Spectrometry Infrastructure
 Mass Spectrometry Infrastructure Todd Williams, Ph.D. Director KU Mass Spectrometry and Analytical Proteomics Laboratory Mass Spectrometry Lab B025 Malott Hall Mission The Mass Spectrometry and analytical
Mass Spectrometry Infrastructure Todd Williams, Ph.D. Director KU Mass Spectrometry and Analytical Proteomics Laboratory Mass Spectrometry Lab B025 Malott Hall Mission The Mass Spectrometry and analytical
Glycerolipid Analysis. LC/MS/MS Analytical Services
 Glycerolipid Analysis LC/MS/MS Analytical Services Molecular Characterization and Quantitation of Glycerophospholipids in Commercial Lecithins by High Performance Liquid Chromatography with Mass Spectrometric
Glycerolipid Analysis LC/MS/MS Analytical Services Molecular Characterization and Quantitation of Glycerophospholipids in Commercial Lecithins by High Performance Liquid Chromatography with Mass Spectrometric
Lipid Metabolism. Catabolism Overview
 Lipid Metabolism Pratt & Cornely, Chapter 17 Catabolism Overview Lipids as a fuel source from diet Beta oxidation Mechanism ATP production Ketone bodies as fuel 1 High energy More reduced Little water
Lipid Metabolism Pratt & Cornely, Chapter 17 Catabolism Overview Lipids as a fuel source from diet Beta oxidation Mechanism ATP production Ketone bodies as fuel 1 High energy More reduced Little water
MSSimulator. Simulation of Mass Spectrometry Data. Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany
 Chris Bielow Algorithmic Bioinformatics, Institute for Computer Science MSSimulator Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany Simulation of Mass Spectrometry Data Motivation
Chris Bielow Algorithmic Bioinformatics, Institute for Computer Science MSSimulator Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany Simulation of Mass Spectrometry Data Motivation
Integrated Targeted Quantitation Method for Insulin and its Therapeutic Analogs
 Integrated Targeted Quantitation Method for Insulin and its Therapeutic Analogs Eric Niederkofler, 1 Dobrin Nedelkov, 1 Urban Kiernan, 1 David Phillips, 1 Kemmons Tubbs, 1 Scott Peterman, 2 Bryan Krastins,
Integrated Targeted Quantitation Method for Insulin and its Therapeutic Analogs Eric Niederkofler, 1 Dobrin Nedelkov, 1 Urban Kiernan, 1 David Phillips, 1 Kemmons Tubbs, 1 Scott Peterman, 2 Bryan Krastins,
Tala Saleh. Razi Kittaneh ... Nayef Karadsheh
 Tala Saleh Razi Kittaneh... Nayef Karadsheh β-oxidation of Fatty Acids The oxidation of fatty acids occurs in 3 steps: Step 1: Activation of the Fatty acid FA + HS-CoA + ATP FA-CoA + AMP + PPi - The fatty
Tala Saleh Razi Kittaneh... Nayef Karadsheh β-oxidation of Fatty Acids The oxidation of fatty acids occurs in 3 steps: Step 1: Activation of the Fatty acid FA + HS-CoA + ATP FA-CoA + AMP + PPi - The fatty
Molecular Cell, Volume 46. Supplemental Information
 Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
A Robustness Study for the Agilent 6470 LC-MS/MS Mass Spectrometer
 A Robustness Study for the Agilent 7 LC-MS/MS Mass Spectrometer Application Note Clinical Research Authors Linda Côté, Siji Joseph, Sreelakshmy Menon, and Kevin McCann Agilent Technologies, Inc. Abstract
A Robustness Study for the Agilent 7 LC-MS/MS Mass Spectrometer Application Note Clinical Research Authors Linda Côté, Siji Joseph, Sreelakshmy Menon, and Kevin McCann Agilent Technologies, Inc. Abstract
Roles of Lipids. principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular
 Roles of Lipids principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular = Oxidation of fatty acids Central energy-yielding pathway in animals. O
Roles of Lipids principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular = Oxidation of fatty acids Central energy-yielding pathway in animals. O
Analysis of Testosterone, Androstenedione, and Dehydroepiandrosterone Sulfate in Serum for Clinical Research
 Analysis of Testosterone, Androstenedione, and Dehydroepiandrosterone Sulfate in Serum for Clinical Research Dominic Foley, Michelle Wills, and Lisa Calton Waters Corporation, Wilmslow, UK APPLICATION
Analysis of Testosterone, Androstenedione, and Dehydroepiandrosterone Sulfate in Serum for Clinical Research Dominic Foley, Michelle Wills, and Lisa Calton Waters Corporation, Wilmslow, UK APPLICATION
Rapid Analysis of Water-Soluble Vitamins in Infant Formula by Standard-Addition
 Rapid Analysis of Water-Soluble Vitamins in Infant Formula by Standard-Addition Evelyn Goh Waters Pacific, Singapore APPLICATION BENEFITS This method allows for the simultaneous analysis of 12 water-soluble
Rapid Analysis of Water-Soluble Vitamins in Infant Formula by Standard-Addition Evelyn Goh Waters Pacific, Singapore APPLICATION BENEFITS This method allows for the simultaneous analysis of 12 water-soluble
Relative Quantitation of Human Polymorphonuclear Leukocyte Cell Membrane GPEtn Lipids
 Relative Quantitation of Human Polymorphonuclear Leukocyte Cell Membrane GPEtn Lipids Using the QTRAP System with mtraq Reagents Karin A. Zemski-Berry 1, John M. Hevko 2, and Robert C. Murphy 1 1 Department
Relative Quantitation of Human Polymorphonuclear Leukocyte Cell Membrane GPEtn Lipids Using the QTRAP System with mtraq Reagents Karin A. Zemski-Berry 1, John M. Hevko 2, and Robert C. Murphy 1 1 Department
Loss of protein association causes cardiolipin degradation in Barth syndrome
 SUPPLEMENTARY INFORMATION Loss of protein association causes cardiolipin degradation in Barth syndrome Yang Xu 1, Colin K.L. Phoon 2, Bob Berno 5, Kenneth D Souza 6, Esthelle Hoedt 4, Guoan Zhang 4, Thomas
SUPPLEMENTARY INFORMATION Loss of protein association causes cardiolipin degradation in Barth syndrome Yang Xu 1, Colin K.L. Phoon 2, Bob Berno 5, Kenneth D Souza 6, Esthelle Hoedt 4, Guoan Zhang 4, Thomas
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group
 4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
Voet Biochemistry 3e John Wiley & Sons, Inc.
 * * Voet Biochemistry 3e Lipid Metabolism Part I: (Chap. 25, sec.1-3) Glucose C 6 H 12 O 6 + 6 O 2 6 CO 2 + 6 H 2 O G o = -2823 kj/mol Fats (palmitic acid) C 16 H 32 O 2 + 23 O 2 16 CO 2 + 16 H 2 O G o
* * Voet Biochemistry 3e Lipid Metabolism Part I: (Chap. 25, sec.1-3) Glucose C 6 H 12 O 6 + 6 O 2 6 CO 2 + 6 H 2 O G o = -2823 kj/mol Fats (palmitic acid) C 16 H 32 O 2 + 23 O 2 16 CO 2 + 16 H 2 O G o
A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids
 A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids Jeremy Netto, 1 Stephen Wong, 1 Federico Torta, 2 Pradeep Narayanaswamy, 2
A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids Jeremy Netto, 1 Stephen Wong, 1 Federico Torta, 2 Pradeep Narayanaswamy, 2
