Nucleocytoplasmic Shuttling of Influenza A Virus Proteins
|
|
|
- Theresa Alexander
- 5 years ago
- Views:
Transcription
1 Viruses 2015, 7, ; doi: /v Review OPEN ACCESS viruses ISSN Nucleocytoplasmic Shuttling of Influenza A Virus Proteins Jing Li 1,, Meng Yu 1,2,, Weinan Zheng 1 and Wenjun Liu 1,2, * 1 Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing , China; s: lj418@163.com (J.L.); zhengdayumeng@163.com (M.Y.); zwn@foxmail.com (W.Z.) 2 University of Chinese Academy of Sciences, Beijing , China These authors contributed equally to this work. * Author to whom correspondence should be addressed; liuwj@im.ac.cn; Tel.: ; Fax: Academic Editor: Alexander Ploss Received: 24 December 2014 / Accepted: 20 May 2015 / Published: 22 May 2015 Abstract: Influenza viruses transcribe and replicate their genomes in the nuclei of infected host cells. The viral ribonucleoprotein (vrnp) complex of influenza virus is the essential genetic unit of the virus. The viral proteins play important roles in multiple processes, including virus structural maintenance, mediating nucleocytoplasmic shuttling of the vrnp complex, virus particle assembly, and budding. Nucleocytoplasmic shuttling of viral proteins occurs throughout the entire virus life cycle. This review mainly focuses on matrix protein (M1), nucleoprotein (NP), nonstructural protein (NS1), and nuclear export protein (NEP), summarizing the mechanisms of their nucleocytoplasmic shuttling and the regulation of virus replication through their phosphorylation to further understand the regulation of nucleocytoplasmic shuttling in host adaptation of the viruses. Keywords: influenza A virus; nucleocytoplasmic shuttling; virus replication 1. Introduction Influenza A virus (IAV), which contains an eight-segment, single-stranded, negative-sense RNA genome, is a member of the Orthomyxovirus family. Further subtyping is based on the antigenicity of the hemagglutinin (HA) and neuramindase (NA) surface glycoproteins. Wild waterfowl are the natural
2 Viruses 2015, reservoir of IAV, which can infect many kinds of mammals, including humans [1]. Currently, two major subtype (H1N1 and H3N2) viruses are circulating among humans. Although still an area of active research, the prevention and control of IAV infection mainly rely on vaccines, and the trivalent inactivated vaccine, which is recommended by the World Health Organization (WHO), is widely used. 2. Influenza A Virus Proteins Influenza virions are roughly spherical, made up of a viral envelope, a matrix layer, and a central core. The envelope consists of HA and NA, usually in a 4:1 ratio [2], and the M2 protein acts as a proton channel. The oligomerization of M1 forms the matrix layer to stabilize the structure of the virion [3,4]. The trimeric polymerase complex (composed of PB1, PB2, and PA), nucleoprotein (NP), and viral RNA constitute the basic unit of the IAV genome for transcription and replication, which is called the viral RNA ribonucleoprotein (vrnp) [5]. In addition, IAV also encodes several other proteins, such as nuclear export protein (NEP), which participates in export of newly made vrnps; nonstructural protein 1 (NS1), which mainly inhibits the type I interferon (IFN-I) system of host cells; and PB1-F2, PB1-N40, and PA-X, PA-N155, PA-N182, M42, and NS3 [6 11], which all play various roles in the virus life cycle. Unlike most RNA viruses, IAV replicates in the host nucleus [12]. In the early stage of infection, vrnps must enter the nucleus for replication and transcription. Then, the newly synthesized proteins translocate back into the nucleus to help in the assembly and nuclear export of progeny vrnps. Finally, the formation of mature virions and budding occurs. Thus, nucleocytoplasmic shuttling is important throughout the entire IAV life cycle. The transport of proteins, as well as cellular and viral RNPs, from the cytoplasm to the nucleus (and vice versa) is an active, energy-dependent, signal-mediated process. Taking into account that virus RNAs transport as RNPs rather than naked RNAs, RNA-binding proteins play a key role in RNA transport. Most of these proteins both bind RNA and contain sequences that allow them to be exported/imported between the cytoplasm and nucleus. In this review, we discuss key findings that have advanced our understanding of the nucleocytoplasmic shuttling of influenza virus proteins (especially for the M1, NP, and NS protein) and their influence on virus replication. 3. Nucleocytoplasmic Shuttling Mechanism of IAV Proteins In eukaryotic cells, the transport of proteins and RNAs into and out of the nucleus through nuclear pore complexes (NPCs)-large macromolecular structures embedded in the double membrane of the nuclear envelope, requires the help of transporter molecules and is an active process. There are two types of transporter molecules, named importin and exportin based on their functions. The importins, α and β, both belong to the importin β super-family. The former is able to recognize and bind the nuclear location signals (NLSs) in the target protein, and the complex is recognized and bound by importin β, which subsequently interacts with the NPC to accomplish the translocation of target proteins. Like the importins, exportins are responsible for ferrying cargo out of the nucleus via the same working principle. Chromosome region maintenance protein 1 (CRM1) is an exportin that recognizes and binds to target proteins carrying a nuclear export signal (NES) and mediates their export from the nucleus [13]. The classic NLSs recognized by importin α consist of two classes. The first is represented by the NLS of the Simian virus 40 (SV40) Large T-antigen (LT) protein and is composed of several contiguous basic amino acids ( 126 PKKKRKV 132, the basic amino acids are shown in bold). The second category is
3 Viruses 2015, represented by the NLS of nucleoplasmin, which is characterized by continuous basic amino acid sequences at both ends separated by >10 random amino acid residues ( 156 KRPAATKKAGQAKKKK 171, the basic amino acids are shown in bold) [14 16]. There are many subtypes in the importin α family, with different cargo specificities for different NLSs. Additionally, the conformation and phosphorylation of target proteins NLSs are also associated with importin α [17]. The NES recognized by CRM1 is composed of hydrophobic amino acids with larger side chains that appear at regular intervals. Its characteristic sequence is Φ-X(2-3)-Φ-X(2-3)-Φ-X-Φ, where Φ stands for Leu, Val, Ile, Met, or Phe, and X can be any amino acid [18]. After the first NLS of SV40 LT was identified in 1984, functional NLSs and NESs were subsequently found in influenza virus proteins, including PB1, PB2, PA, NP, M1, NS1, and NEP (Table 1). The proteins with NLSs can be recognized and bound by importin α for nuclear entry. Another non-classical nuclear import factor, Ran binding protein 5 (RanBP5), interacts with PB1-PA dimer to transport into the nucleus [19,20]. After the completion of replication, vrnps require the help of two viral proteins, M1 and NEP, which contain the NESs for recognition by CRM1 to move out of the nucleus for assembly and budding of progeny viruses [21 24]. The following section describes the nucleocytoplasmic shuttling mechanism of four IAV proteins (M1, NP, NS1, and NEP) in detail Nucleocytoplasmic Shuttling of M1 M1 is the most abundant protein in the virion and forms the matrix layer inside the virus envelope. The exterior matrix layer combines with HA, NA, and M2 located on the envelope, and the interior matrix layer combines with vrnp. Both layers form the virion structure. Initially, it was thought that M1 interacted with HA and NA inside the cytoplasm. Subsequently, the M1 protein was found to enter the nucleus and be indispensable for vrnp nuclear export [25]. Aside from the well-characterized basic amino acid-rich NLS [26], there is another basic amino acid stretch at position that is highly conserved among influenza A and B virus M1 proteins and plays a critical role in virus replication. Indeed, there is evidence showing that R77A or R78A substitutions in M1 result in aberrant M1 intracellular localization and defects in virus assembly and budding [27]. The viral polymerase proteins (PA, PB1, and PB2) are imported into the nucleus to assemble vrnps, together with NP and viral RNA. It is well established that vrnp nuclear export is mediated by the nuclear presence of M1 and NEP. In the absence of a functional NES, M1 is not able to interact with CRM1 itself. NEP likely bridges the M1 and CRM1 interaction by binding to M1 with its C-terminus, while the two N-terminal NESs interact with CRM1 and allow access to the CRM1-dependent export pathway (Figure 1). Our pervious experiments show that intracellular distribution of M1 is not sensitive to the CRM1 inhibitor leptomycin B (LMB), further confirming the indispensable role of NEP for vrnp nuclear export. Furthermore, the M1 individual nuclear export was specially dependent on its NES and critical for influenza A virus replication [24].
4 Viruses 2015, Table 1. List of influenza A virus protein nuclear localization signals (NLSs) and nuclear export signals (NESs). Protein NLS NES References NLS1: 34 RLRR 38, highly conserved [28] NS1 138 FDRLETLILL 147 NLS2: 216 PKQKRK 221, two NLSs act independently [29] PB1 187 RKRRVRDNMTKKMVTQRTIGKRKQR 211 [30], bipartite NLS [31] PB2 PA M1 NP NEP NLS1: 449 GIESIDNVMGMIGILPDMTPSTEMSMRGVRISKMGVDETSSAEKIV 495, required for efficient import NLS2: 736 KRKR 739, bipartite NLS K736 required for efficient import NLS1: 124 RREVHIYYLEKANKIK 139, bipartite NLS NLS2: E154 required for efficient import 101 RKLKR 105 NLS1: 3 TKGTKRSYEQM 13, unconventional NLS, 3Ser crucial for N-terminal phosphorylation NLS2: 198 KGINDRNFWRGENGRRTR 216, bipartite NLS Passive diffusion, no need of NLS 59 ILGFVFTLTV 68 L66A, V68A mutation impairs M1 nuclear export NES1: 24 EIRASVGKMIDGIGRFYIQMCTELKL 49 NES2: 183 VKGVGTMVMELIRMI 197 NES3: 248 PGNAEFEDLIFLARSALILRGSVAHKS 274 NES1: 12 ILMRMSKMQL 21 NES2: 31 IITQFESLKI 40 [32] [33] [34] [26] [24] [35] [36] [37] [38] [39] [40] [41] [42] [21] [43] [44]
5 Viruses 2015, Figure 1. Model of the nucleocytoplasmic shuttling of influenza virus proteins and vrnps. The importin/exportin molecules recognize and bind the NLSs/NESs in the target proteins and then are bound by the importin/exportin molecule, which subsequently interacts with the NPC to accomplish the nucleocytoplasmic shuttling. The importins α-β and RanBP5 pathway separately involved into the vrnps import. CRM1 is a commonly utilized exportin for influenza virus vrnps. The export of IAV RNPs seems to depend on a RanGTP-CRM1-NEP-M1-RNP model for transport out of the nucleus as a complex Nucleocytoplasmic Shuttling of NP There are at least two NLSs in NP, including the unconventional NLS1 (3 13), which interacts with importin α [36], and NLS2 ( ) [39], which together with NLS1 was originally hypothesized to be a traditional bipartite NLS. However, the crystal structure of NP shows that the two clusters are too close to be recognized by importins as a bipartite NLS [45]. Although it is hard to judge which of the two NLSs contributes most to the import of vrnp, import is almost completely inhibited when both NLSs are blocked using antibodies [46,47]. In addition, in vitro experiments and a reverse genetics approach reveal that NLS1 plays a pivotal role in mediating vrnp nuclear import [48,49]. Three NESs have been identified in NP that have the familiar sequence characteristics of common export signals. Two of them do not rely on CRM1 for export, though the third does. Further, all three
6 Viruses 2015, must be simultaneously mutated to inhibit NP nuclear export, proving their functional independence [42]. The three NESs are closely related to the nuclear export of vrnp, and mutation of the NP NESs inhibits the replication of the virus in the post-transcriptional stage. Through function comparison via mutation of all seven NESs in IAV proteins, it was found that NP NES3 played a key role in the replication of the virus, but the mechanism for this phenomenon has not been elucidated [50] Nucleocytoplasmic Shuttling of NS1 NS1 is mainly located inside the nucleus and transports to the cytoplasm in the late stage of infection. Two NLSs and one NES have been identified in NS1 [28]. S221 in NLS2 is very important for the nuclear localization of NS1. Further, the nuclear export of NS1 does not (or only partially) rely on CRM1 [51]. By comparison among different strains, it was found that the NS1 NES is highly conserved, and mutations of conserved residues, such as L141A and A149V, lead to the retention of NS1 inside the nucleus during the late stage of infection [52] Nucleocytoplasmic Shuttling of NEP NEP is known as a nuclear export protein. It may not need a NLS as it is only 14 kda in size, and at present, no NLS has been found in NEP. Through the use of an inhibitor of NLS-dependent active nuclear transport and an energy depletion assay, it was demonstrated that NEP enters the nucleus by passive diffusion [44]. NEP is involved in the nuclear export of vrnps [53]. NEP is able to interact with CRM1 in mammalian two-hybrid systems, suggesting that the nuclear export of NEP depends on binding to CRM1 [54]. Further, it has been demonstrated that the NES1 and NES2 of NEP [43] are CRM1 dependent and jointly participate in the nuclear export of NEP and vrnp. Interestingly, the NEPs of different strains result in distinct cellular distribution patterns, forming unique nuclear aggregates, with the weaker binding capacity to CRM1 due to one amino acid change in the NES, perhaps facilitating the export of the vrnp complex [44]. 4. Biological Significance of Nucleocytoplasmic Shuttling on IAV Replication 4.1. Interactions among Members of the vrnp Nucleocytoplasmic Shuttling Complex The vrnp is the basic unit for the replication and transcription of IAV. The interaction between members of the vrnp complex is of great significance for the structural stability and function of the complex. Several functional domains of related proteins involved in the interaction with the members of the polymerase complex [55,56]. For example, several binding sites on NP have been identified, including two NP-NP interaction regions (aa and ), two RNA interaction regions (aa 1 77 and ), two PB2 interaction regions (aa and ), and three polymerase interaction regions (R204, W207, and R208) [57,58]. As with the relevant interaction regions in the trimeric polymerase, these regions support the formation of the vrnp and are required for its function. It has been found that disturbing the self-polymerization of NP and destroying the interaction between NP and PB2 or NP and PB1 regulate the replication of influenza virus in a negative way [59]. The
7 Viruses 2015, C-terminus of NEP also interacts with PB2 and PB1 in the viral polymerase complex to regulate the activity of the polymerase, resulting in alteration of the synthesis of vrna [60]. In the process of infection, IAV enters the host cytoplasm, and at the same time, the M2 ion channel opens, internally acidifying the virus. Subsequently, the matrix layer depolymerizes, and as a result, M1 and vrnp dissociate from each other [55]. This conformational change results in the exposure of the NLS, which leads to the entry of the vrnp into the nucleus for replication and transcription. Each polymerase subunit and NP contain their own NLSs, and it is possible that the monomers are transported into the nucleus (for NP and PB2) or that PB1-PA dimers or trimers form in the cytoplasm before nuclear import, perhaps as part of a mechanism regulating different phases of the infection [61]. It has been intriguing trying to identify which NLSs play a decisive role in transport into the nucleus. Indeed, structure-function experiments demonstrate that the role of NLS1 is very important in mediating vrnp nuclear localization [62]. For vrnp nuclear export, a CRM1-NEP-M1-vRNP complex forms, where CRM1 recognizes and binds to the NES on NEP, accompanied by GTP hydrolysis (Figure 1). Thus, the nuclear export of vrnp depends on CRM1 [63]. Although NP contains an NES and can interact with CRM1, the NP/CRM1 interaction itself fails to induce the hydrolysis of RanGTP, suggesting that the NESs on NP cannot directly mediate vrnp nuclear export [23]. Indeed, M1 and NEP are necessary for this process [21,25]. M1 is exported from the nucleus via a CRM1-independent pathway, while all of the NESs of NEP are CRM1 dependent, suggesting that the NES on NEP directly mediates the nuclear export of vrnp. However, NES mutation on M1 leads to the nuclear retention of M1 and the accumulation of NEP, in addition to significantly reducing the virus titer, indicating that M1 plays an important role in vrnp nuclear export [24]. When the vrnp translocates out of the nucleus, the C-terminus of NEP binds to the N-terminus of M1, covering the NLS on M1 and inhibiting vrnp re-entry into the nucleus; mutation of NEP W78 destroys the interaction between NEP and M1 [23]. Experiments also suggest that the ability of NEP to support the nuclear export of vrnps by stabilizing the interaction between M1 and vrnps requires the NES-containing N-terminus, as well as the last three amino acids of NEP, and is functionally linked to the polymerase activity-enhancing function of NEP during viral genome replication [64] Phosphorylation of IAV Proteins to Regulate Nucleocytoplasmic Shuttling The phosphorylation of IAV proteins is an important regulatory mechanism that influences viral replication by affecting the nucleocytoplasmic shuttling of both the individual viral proteins themselves and the vrnp complex. Nuclear transport of M1 depends on phosphorylation mediated by protein kinase C (PKC), as PKC inhibition effectively decreases virus replication [65]. Mutation of the M1 Y132 phosphorylation site in A/WSN/1933 (H1N1) eliminates the interaction between M1 and importin α1, inhibiting M1 nuclear import, and thus having a negative influence on virus replication. JAK kinase inhibition can block this phosphorylation and impair WSN M1 entry into the nucleus but not H9N2 M1, suggesting that differential regulation of the interaction between M1 and importins exists in different subtypes of IAVs [66] (Figure 2). There are additional potential phosphorylation sites in the IAV M1 protein, such as T108, that are close to the NLS but whose functions have not been determined [67].
8 Viruses 2015, Figure 2. Model of Y132 phosphorylation participating in nuclear import of IAV M1. The conserved tyrosine 132 of M1 is a phosphorylation site. The NLS-neighboring phosphorylated Y132 residue is crucial for the cytoplasmic-nuclear translocation of M1 by increasing the binding capacity of M1 and nuclear import factor importin-α1, subsequently affecting the replication of IAV. NP is rich in serine residues, which is crucial for its phosphorylation, especially on the N-terminus of NLS1 [37]. Inhibiting phosphorylation of infected cells stabilizes the localization of NP in the nucleus and inhibits its nuclear export [38,68]. Several highly conserved phosphorylation sites have been identified in NP, such as S9/Y10 and S165. Mutation of the former significantly decreases viral titers [67], and the S165D mutation (which mimics phosphorylation) may maintain NP in a monomeric state, reducing its affinity for RNA [69]. In our lab, we demonstrated that the phosphorylation of S9, Y10, and Y296 was essential for virus growth in cell culture and modulation of activity of the viral polymerase in a mouse model and the nuclear-cytoplasmic shuttling of NP. The phosphorylation and de-phosphorylation of S9 and Y10 controlled nuclear import of NP by affecting the binding affinity between NP and importin-α. In addition, the phosphorylation of Y296 caused nuclear retention of NP by reducing the interaction between NP and CRM1 [70].
9 Viruses 2015, The phosphorylation of T215 in NS1 is mediated by the CDK/ERK pathway, and the T215A mutation attenuates virus replication [71]. The functional analysis of several phosphorylation sites, including T215, S48, and S42, shows that the S42D mutation eliminates dsrna binding by NS1, also attenuating virus replication [72]. 5. Conclusions To adapt to new host species, influenza viruses must overcome significant barriers to cross-species transmission and replication. The changes in the receptor-binding properties of the viral HA to adapt to new host species are well characterized, but the nuclear import machinery to determine the host tropism requires a better understanding. Previous studies show that the preference of PB2 for different importin-α isoforms (α1, α3, α4, α5, α6, and α7) differs between hosts. For efficient virus replication in different hosts, all viruses depend on importin-α1. Meanwhile, the polymerase subunit PB2 of avian viruses requires importin-α3, PB2 of mammalian virus shows importin-α7 specificity, and H1N1v replication depends on both, suggesting ongoing adaptation of this virus [73]. PB2 specificity for given importin-α isoform is mediated by both NLS and protein context. Researchers have found out PB2 627-NLS domain can induce selective importin binding in host-specific manner and areas where more research is required, including other transport associated protein in IAV. The specific interaction of the polymerase subunit PB2 and the NP of mammalian viruses with importin-α3/7 regulates the efficacy of viral replication and determine host range underlining the importance of the nuclear envelope in interspecies transmission [73 75]. Work in our laboratory also demonstrates differences in the nucleocytoplasmic shuttling of NPs between human-origin WSN and avian-origin IAVs, which impact the replication, adaptation, and pathogenicity of the viruses (unpublished). In addition, we have also found that the CA04 strain displays different vrnp distribution patterns, forming unique nuclear aggregates, with respect to WSN by comparing different strains of H1N1 NEP for vrnp nuclear translocation. In the further study, we confirmed that CA04 NEP interacts less efficiently with CRM1 and site T48 is responsible for the nuclear aggregation [44]. In view of NEP playing an important role in viral polymerase activity and vrnp nuclear transport efficiency, these differences may be one reason for divergence of the pathogenicity and host adaptation range between these virus strains. Future work will dissect the finely tuned interactions induced by natural/pressure selection to acquire cross-species activity. Phosphorylation is one of the most common and efficient post-translational modification methods in organisms, playing an important role in regulating many metabolic pathways. Nucleocytoplasmic shuttling of proteins and phosphorylation are closely related. Phosphorylation and dephosphorylation can strengthen or weaken the nucleocytoplasmic shuttling of a protein through a variety of mechanisms. For instance, phosphorylation of key NLS or NES residues on viral proteins can strengthen the affinity between the proteins and their nuclear transport receptors [76]. Alternatively, phosphorylation can strengthen the ability of a protein to separate itself from the NPC. In addition, phosphorylation can lead to a conformational change in the protein, selectively exposing its NLS or NES. In contrast, phosphorylation of the NPC or virus proteins can also inhibit nucleocytoplasmic shuttling. Regardless, research into the mechanism of how phosphorylation regulates nucleocytoplasmic shuttling is still at an early stage. Development of better quantitative methods to determine the influence of phosphorylation on the dynamic processes of protein transport will lead to a more comprehensive understanding of the
10 Viruses 2015, role of phosphorylation in the IAV life cycle, which is beneficial for influenza prevention and control. In addition, sumoylation is critical for protein-protein interactions in many biological systems. For example, the sumoylation of matrix protein M1 modulate the assembly and morphogenesis of influenza A virus [77,78], or sumoylation of NP is indispensable for virus growth and intracellular trafficking [79]. Further research found out that the sumoylation of M1 promotes M1-mediated vrnp nuclear export to increase viral replication [80]. Thereby, the modification of virus/cellular protein to further regulate the virus life cycle needs to be given more attention for further research. Acknowledgments This work was supported by grants from the National Natural Science Foundation of China (NSFC) (Grant No ), the National Key Technologies Research and Development Program of China (2013ZX ), China Ministry of Science and Technology (MOST) Project 973 (Grant Nos. 2012CB and 2011CB504705), the Key Research Program of the Chinese Academy of Sciences (KSZD-EW-Z ), and IDRC-APEIR Project Funding. W.J.L. is a principal investigator of the NSFC Innovative Research Group (Grant No ). Author Contributions J.L., M.Y., W.N.Z., and W.J.L. contributed to the content and editing of this article. Conflicts of Interest The authors declare no conflict of interest. References 1. Nelson, M.I.; Holmes, E.C. The evolution of epidemic influenza. Nat. Rev. Genet. 2007, 8, Kilbourne, E.D.; Johansson, B.E.; Grajower, B. Independent and disparate evolution in nature of influenza A virus hemagglutinin and neuraminidase glycoproteins. Proc. Natl. Acad. Sci. USA 1990, 87, Nayak, D.P.; Balogun, R.A.; Yamada, H.; Zhou, Z.H.; Barman, S. Influenza virus morphogenesis and budding. Virus Res. 2009, 143, Coloma, R.; Valpuesta, J.M.; Arranz, R.; Carrascosa, J.L.; Ortin, J.; Benito, J.M. The structure of a biologically active influenza virus ribonucleoprotein complex. PLoS Pathog. 2009, 5, Klumpp, K.; Ruigrok, R.W.H.; Baudin, F. Roles of the influenza virus polymerase and nucleoprotein in forming a functional RNP structure. EMBO J. 1997, 16, Selman, M.; Dankar, S.K.; Forbes, N.E.; Jia, J.-J.; Brown, E.G. Adaptive mutation in influenza A virus non-structural gene is linked to host switching and induces a novel protein by alternative splicing. Emerg. Microbes Infect. 2012, 1, e42.
11 Viruses 2015, Wise, H.M.; Hutchinson, E.C.; Jagger, B.W.; Stuart, A.D.; Kang, Z.H.; Robb, N.; Schwartzman, L.M.; Kash, J.C.; Fodor, E.; Firth, A.E.; et al. Identification of a novel splice variant form of the influenza A virus M2 ion channel with an antigenically distinct ectodomain. PLoS Pathog. 2012, 8, e Jagger, B.W.; Wise, H.M.; Kash, J.C.; Walters, K.A.; Wills, N.M.; Xiao, Y.L.; Dunfee, R.L.; Schwartzman, L.M.; Ozinsky, A.; Bell, G.L.; et al. An overlapping protein-coding region in influenza A virus segment 3 modulates the host response. Science 2012, 337, Wise, H.M.; Foeglein, A.; Sun, J.; Dalton, R.M.; Patel, S.; Howard, W.; Anderson, E.C.; Barclay, W.S.; Digard, P. A complicated message: Identification of a novel PB1-related protein translated from influenza A virus segment 2 mrna. J. Virol. 2009, 83, Muramoto, Y.; Noda, T.; Kawakami, E.; Akkina, R.; Kawaoka, Y. Identification of novel influenza A virus proteins translated from PA mrna. J. Virol. 2013, 87, Chen, W.; Calvo, P.A.; Amalide, D.; Gibbs, J.; Schubert, U.; Bacik, I.; Basta, S.; O Neill, R.; Schickli, J.; Palese, P.; et al. A novel influenza A virus mitochondrial protein that induces cell death. Nat. Med. 2001, 7, Stegmann, T.; M.Whitel, J.; Helenius, A. Intermediates in influenza induced membrane fusion. EMBO J. 1990, 9, Pemberton, L.F.; Blobelt, G.; Rosenblum, J.S. Transport routes through the nuclear pore complex. Curr. Opin. Cell Biol. 1998, 10, Makkerh, J.P.S.; Dingwall, C.; Laskey, R.A. Comparative mutagenesis of nuclear localization signals reveals the importance of neutral and acidic amino acids. Curr. Biol. 1996, 6, Dingwall, C.; Laskey, R.A. Nuclear import: A tale of two sites. Curr. Biol. 1998, 8, Wang, Y.E.; Park, A.; Lake, M.; Pentecost, M.; Torres, B.; Yun, T.E.; Wolf, M.C.; Holbrook, M.R.; Freiberg, A.N.; Lee, B. Ubiquitin-regulated nuclear-cytoplasmic trafficking of the nipah virus matrix protein is important for viral budding. PLoS Pathog. 2010, 6, e Jans, D.A.; Hubner, S. Regulation of protein transport to the nucleus: Central role of phosphorylation. Physiol. Rev. 1996, 76, Henderson, B.R.; Eleftheriou, A. A comparison of the activity, sequence specificity, and CRM1-dependence of different nuclear export signals. Exp. Cell Res. 2000, 256, Fodor, E.; Smith, M. The PA subunit is required for efficient nuclear accumulation of the PB1 subunit of the influenza A virus RNA polymerase complex. J. Virol. 2004, 78, Deng, T.; Engelhardt, O.G.; Thomas, B.; Akoulitchev, A.V.; Brownlee, G.G.; Fodor, E. Role of ran binding protein 5 in nuclear import and assembly of the influenza virus RNA polymerase complex. J. Virol. 2006, 80, E.O Neill, R.; Talon, J.; Palese, P. The influenza virus NEP (NS2 protein) mediates the nuclear export of viral ribonucleoproteins. EMBO J. 1998, 17, Bancroft, C.T.; Parslow, T.G. Evidence for segment-nonspecific packaging of the influenza A virus genome. J. Virol. 2002, 76, Akarsu, H.; Burmeister, W.P.; Petosa, C.; Petit, I. Crystal structure of the M1 protein-binding domain of the influenza A virus nuclear export protein (NEP/NS2). EMBO J. 2003, 22,
12 Viruses 2015, Cao, S.; Liu, X.; Yu, M.; Li, J.; Jia, X.; Bi, Y.; Sun, L.; Gao, G.F.; Liu, W. A nuclear export signal in the matrix protein of influenza A virus is required for efficient virus replication. J. Virol. 2012, 86, Sakaguchi, A.; Hirayama, E.; Hiraki, A.; Ishida, Y.-I.; Kim, J. Nuclear export of influenza viral ribonucleoprotein is temperature-dependently inhibited by dissociation of viral matrix protein. Virology 2003, 306, Ye, Z.; Robinson, D.; Wagner, R.R. Nucleus-targeting domain of the matrix protein (M1) of influenza virus. J. Virol. 1995, 69, Das, S.C.; Watanabe, S.; Hatta, M.; Noda, T.; Neumann, G.; Ozawa, M.; Kawaoka, Y. The highly conserved arginine residues at positions 76 through 78 of influenza A virus matrix protein M1 play an important role in viral replication by affecting the intracellular localization of M1. J. Virol. 2012, 86, Greenspan, D.; Palese, P.; Krystal, M. Two nuclear location signals in the influenza virus NS1 nonstructural protein. J. Virol. 1988, 62, Li, Y.; Yamakita, Y.; Krug, R.M. Regulation of a nuclear export signal by an adjacent inhibitory sequence: The effector domain of the influenza virus NS1 protein. Proc. Natl. Acad. Sci. USA 1998, 95, Nath, S.T.; Nayak, D.P. Function of two discrete regions is required for nuclear localization of polymerase basic protein 1 of A/WSN/33 influenza virus (H1N1). Mol. Cell. Biol. 1990, 10, Hutchinson, E.C.; Orr, O.E.; Man Liu, S.; Engelhardt, O.G.; Fodor, E. Characterization of the interaction between the influenza A virus polymerase subunit PB1 and the host nuclear import factor Ran-binding protein 5. J. Gen. Virol. 2011, 92, Mukaigawa, J.; Nayak, D.P. Two signals mediate nuclear localization of influenza virus (A/WSN/33) polymerase basic protein 2. J. Virol. 1991, 65, Tarendeau, F.; Boudet, J.; Guilligay, D.; Mas, P.J.; Bougault, C.M.; Boulo, S.; Baudin, F.; Ruigrok, R.W.; Daigle, N.; Ellenberg, J.; et al. Structure and nuclear import function of the C-terminal domain of influenza virus polymerase PB2 subunit. Nat. Struct. Mol. Biol. 2007, 14, Nieto, A.; Luna, S.D.L.; Bfircena, J.; Portela, A.; Ortin, J. Complex structure of the nuclear translocation signal of influenza virus polymerase PA subunit. J. Gen. Virol. 1994, 75, Fau, D.J.; Hurtley, S.M.; Warren, G. Reconstitution of an endocytic fusion event in a cell-free system. Cell 1985, 43, Wang, P.; Palese, P.; O Neill, R.E. The NPI-1/NPI-3 (karyopherin alpha) binding site on the influenza A virus nucleoprotein NP is a nonconventional nuclear localization signal. J. Virol. 1997, 71, Arrese, M.; Portela, A. Serine 3 is critical for phosphorylation at the N-terminal end of the nucleoprotein of influenza virus A/Victoria/3/75. J. Virol. 1996, 70, Neumann, G.; Castrucci, M.R.; Kawaoka, Y. Nuclear import and export of influenza virus nucleoprotein. J. Virol. 1997, 71, Weber, F.; Kochs, G.; Gruber, S.; Haller, O. A classical bipartite nuclear localization signal on Thogoto and influenza A virus nucleoproteins. Virology 1998, 250, 9 18.
13 Viruses 2015, Bullido, R.; Gomez-Puertas, P.; Albo, C.; Portela, A. Several protein regions contribute to determine the nuclear and cytoplasmic localization of the influenza A virus nucleoprotein. J. Gen. Virol. 2000, 81, Wu, W.W.; Pante, N. The directionality of the nuclear transport of the influenza A genome is driven by selective exposure of nuclear localization sequences on nucleoprotein. Virol. J. 2009, 6, e Yu, M.; Liu, X.; Cao, S.; Zhao, Z.; Zhang, K.; Xie, Q.; Chen, C.; Gao, S.; Bi, Y.; Sun, L.; et al. Identification and characterization of three novel nuclear export signals in the influenza A virus nucleoprotein. J. Virol. 2012, 86, Huang, S.; Chen, J.; Chen, Q.; Wang, H.; Yao, Y.; Chen, J.; Chena, Z. A second CRM1-dependent nuclear export signal in the influenza A virus NS2 protein contributes to the nuclear export of viral ribonucleoproteins. J. Virol. 2013, 87, Gao, S.; Wang, S.; Cao, S.; Sun, L.; Li, J.; Bi, Y.; Gao, G.F.; Liu, W. Characteristics of nucleocytoplasmic transport of H1N1 influenza A virus nuclear export protein. J. Virol. 2014, 88, Ye, Q.; Krug, R.M.; Tao, Y.J. The mechanism by which influenza A virus nucleoprotein forms oligomers and binds RNA. Nature 2006, 444, Zhang, H.; Hale, B.G.; Xu, K.; Sun, B. Viral and host factors required for avian H5N1 influenza A virus replication in mammalian cells. Viruses 2013, 5, Wu, W.W.; Sun, Y.H.; Pante, N. Nuclear import of influenza a viral ribonucleoprotein complexes is mediated by two nuclear localization sequences on viral nucleoprotein. Virol. J. 2007, 4, e Cros, J.F.; Garcia-Sastre, A.; Palese, P. An unconventional NLS is critical for the nuclear import of the influenza A virus nucleoprotein and ribonucleoprotein. Traffic 2005, 6, Ozawa, M.; Fujii, K.; Muramoto, Y.; Yamada, S.; Yamayoshi, S.; Takada, A.; Goto, H.; Horimoto, T.; Kawaoka, Y. Contributions of two nuclear localization signals of influenza A virus nucleoprotein to viral replication. J. Virol. 2007, 81, Chutiwitoonchai, N.; Kakisaka, M.; Yamada, K.; Aida, Y. Comparative analysis of seven viral nuclear export signals (NESs) reveals the crucial role of nuclear export mediated by the third NES consensus sequence of nucleoprotein (NP) in influenza A virus replication. PLoS ONE 2014, 9, e Han, H.; Cui, Z.Q.; Wang, W.; Zhang, Z.P.; Wei, H.P.; Zhou, Y.F.; Zhang, X.E. New regulatory mechanisms for the intracellular localization and trafficking of influenza A virus NS1 protein revealed by comparative analysis of A/PR/8/34 and A/Sydney/5/97. J. Gen. Virol. 2010, 91, Li, Z.; Jiang, Y.; Jiao, P.; Wang, A.; Zhao, F.; Tian, G.; Wang, X.; Yu, K.; Bu, Z.; Chen, H. The NS1 gene contributes to the virulence of H5N1 avian influenza viruses. J. Virol. 2006, 80, Neumann, G.; T.Hughes, M.; Kawaoka, Y. Influenza A virus NS2 protein mediates vrnp nuclear export through NES-independent interaction with hcrm1. EMBO J. 2000, 19, Iwatsuki Horimoto, K.; Horimoto, T.; Fujii, Y.; Kawaoka, Y. Generation of influenza A virus NS2 (NEP) mutants with an altered nuclear export signal sequence. J. Virol. 2004, 78,
14 Viruses 2015, Eisfeld, A.J.; Neumann, G.; Kawaoka, Y. At the centre: Influenza A virus ribonucleoproteins. Nat. Rev. Microbiol. 2015, 13, Hutchinson, E.C.; Fodor, E. Nuclear import of the influenza A virus transcriptional machinery. Vaccine 2012, 30, Elton, D.; Medcalf, L.; Bishop, K.; Harrison, D.; Digard, P. Identification of amino acid residues of influenza virus nucleoprotein essential for RNA binding. J. Virol. 1999, 73, Marklund, J.K.; Ye, Q.; Dong, J.; Tao, Y.J.; Krug, R.M. Sequence in the influenza A virus nucleoprotein required for viral polymerase binding and RNA synthesis. J. Virol. 2012, 86, Wang, Z.; Liu, X.; Zhao, Z.; Xu, C.; Zhang, K.; Chen, C.; Sun, L.; Gao, G.F.; Ye, X.; Liu, W. Cyclophilin E functions as a negative regulator to influenza virus replication by impairing the formation of the viral ribonucleoprotein complex. PLoS ONE 2011, 6, e Robb, N.C.; Smith, M.; Vreede, F.T.; Fodor, E. NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome. J. Gen. Virol. 2009, 90, Deng, T.; Sharps, J.; Fodor, E.; Brownlee, G.G. In vitro assembly of PB2 with a PB1-PA dimer supports a new model of assembly of influenza A virus polymerase subunits into a functional trimeric complex. J. Virol. 2005, 79, Wu, W.W.; Weaver, L.L.; Pante, N. Ultrastructural analysis of the nuclear localization sequences on influenza A ribonucleoprotein complexes. J. Mol. Biol. 2007, 374, Elton, D.; Simpson-Holley, M.; Archer, K.; Medcalf, L.; Hallam, R.; McCauley, J.; Digard, P. Interaction of the influenza virus nucleoprotein with the cellular CRM1-mediated nuclear export pathway. J. Virol. 2001, 75, Brunotte, L.; Flies, J.; Bolte, H.; Reuther, P.; Vreede, F.; Schwemmle, M. The nuclear export protein of H5N1 influenza A viruses recruits matrix 1 (M1) protein to the viral ribonucleoprotein to mediate nuclear export. J. Biol. Chem. 2014, 289, Halder, U.C.; Bhowmick, R.; Roy Mukherjee, T.; Nayak, M.K.; Chawla-Sarkar, M. Phosphorylation drives an apoptotic protein to activate antiapoptotic genes: Paradigm of influenza a matrix 1 protein function. J. Biol. Chem. 2013, 288, Wang, S.; Zhao, Z.; Bi, Y.; Sun, L.; Liu, X.; Liu, W. Tyrosine 132 phosphorylation of influenza A virus M1 protein is crucial for virus replication by controlling the nuclear import of M1. J. Virol. 2013, 87, Hutchinson, E.C.; Denham, E.M.; Thomas, B.; Trudgian, D.C.; Hester, S.S.; Ridlova, G.; York, A.; Turrell, L.; Fodor, E. Mapping the phosphoproteome of influenza A and B viruses by mass spectrometry. PLoS Pathog. 2012, 8, e Bui, M.; Myers, J.E.; Whittaker, G.R. Nucleo-cytoplasmic localization of influenza virus nucleoprotein depends on cell density and phosphorylation. Virus Res. 2003, 84, Chenavas, S.; Estrozi, L.F.; Slama-Schwok, A.; Delmas, B.; di Primo, C.; Baudin, F.; Li, X.; Crepin, T.; Ruigrok, R.W. Monomeric nucleoprotein of influenza A virus. PLoS Pathog. 2013, 9, Zheng, W.; Li, J.; Wang, S.; Cao, S.; Jiang, J.; Chen, C.; Ding, C.; Qin, C.; Ye, X.; Gao, G.F.; Liu, W. Phosphorylation controls the nuclear-cytoplasmic shuttling of influenza A virus nucleoprotein. J. Virol. 2015, doi: /jvi
15 Viruses 2015, Hale, B.G.; Knebel, A.; Botting, C.H.; Galloway, C.S.; Precious, B.L.; Jackson, D.; Elliott, R.M.; Randall, R.E. CDK/ERK-mediated phosphorylation of the human influenza A virus NS1 protein at threonine-215. Virology 2009, 383, Hsiang, T.Y.; Zhou, L.; Krug, R.M. Roles of the phosphorylation of specific serines and threonines in the NS1 protein of human influenza A viruses. J. Virol. 2012, 86, Gabriel, G.; Klingel, K.; Otte, A.; Thiele, S.; Hudjetz, B.; Arman-Kalcek, G.; Sauter, M.; Shmidt, T.; Rother, F.; Baumgarte, S.; et al. Differential use of importin-alpha isoforms governs cell tropism and host adaptation of influenza virus. Nat. Commun. 2011, 2, e Boivin, S.; Hart, D.J. Interaction of the influenza A virus polymerase PB2 C-terminal region with importin alpha isoforms provides insights into host adaptation and polymerase assembly. J. Biol. Chem. 2011, 286, Hudjetz, B.; Gabriel, G. Human-like PB2 627K influenza virus polymerase activity is regulated by importin-alpha1 and-alpha7. PLoS Pathog. 2012, 8, e Nardozzi, J.D.; Lott, K.; Cingolani, G. Phosphorylation meets nuclear import: A review. Cell Commun. Signal. 2010, 8, e Pal, S.; Santos, A.; Rosas, J.M.; Ortiz-Guzman, J.; Rosas-Acosta, G. Influenza A virus interacts extensively with the cellular SUMOylation system during infection. Virus Res. 2011, 158, Wu, C.Y.; Jeng, K.S.; Lai, M.M. The SUMOylation of matrix protein M1 modulates the assembly and morphogenesis of influenza A virus. J. Virol. 2011, 85, Han, Q.L.; Chang, C.; Li, L.; Klenk, C.; Cheng, J.K.; Chen, Y.X.; Xia, N.S.; Shu, Y.L.; Chen, Z.; Gabriel, G.; et al. Sumoylation of Influenza A Virus Nucleoprotein Is Essential for Intracellular Trafficking and Virus Growth. J. Virol. 2014, 88, Gao, S.; Wu, J.; Liu, R.Y.; Li, J.; Song, L.; Teng, Y.; Sheng, C.; Liu, D.; Yao, C.; Chen, H.; et al. Interaction of NS2 with AIMP2 facilitates the switch from ubiquitination to SUMOylation of M1 in influenza A virus-infected cells. J. Virol. 2015, 89, by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (
MOLECULAR CELL BIOLOGY
 1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
 http://nmhm.washingtondc.museum/collections/archives/agalleries/1918flu/ncp1603.jpg 1 https://assets-production-webvanta-com.s3-us-west- 2 2.amazonaws.com/000000/47/62/original/images/img_109_influenza/Spanish_flu_death_chart.jpg
http://nmhm.washingtondc.museum/collections/archives/agalleries/1918flu/ncp1603.jpg 1 https://assets-production-webvanta-com.s3-us-west- 2 2.amazonaws.com/000000/47/62/original/images/img_109_influenza/Spanish_flu_death_chart.jpg
The accumulation of influenza A virus segment 7 spliced mrnas is regulated by the NS1 protein
 Journal of General Virology (2012), 93, 113 118 DOI 10.1099/vir.0.035485-0 Short Communication Correspondence Ervin Fodor ervin.fodor@path.ox.ac.uk Received 24 June 2011 Accepted 12 September 2011 The
Journal of General Virology (2012), 93, 113 118 DOI 10.1099/vir.0.035485-0 Short Communication Correspondence Ervin Fodor ervin.fodor@path.ox.ac.uk Received 24 June 2011 Accepted 12 September 2011 The
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome
 Journal of General Virology (2009), 90, 1398 1407 DOI 10.1099/vir.0.009639-0 NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome Nicole C. Robb,3 Matt Smith,34 Frank
Journal of General Virology (2009), 90, 1398 1407 DOI 10.1099/vir.0.009639-0 NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome Nicole C. Robb,3 Matt Smith,34 Frank
Influenza viruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
 This article was published in an Elsevier journal. The attached copy is furnished to the author for non-commercial research and education use, including for instruction at the author s institution, sharing
This article was published in an Elsevier journal. The attached copy is furnished to the author for non-commercial research and education use, including for instruction at the author s institution, sharing
Plasmid-Driven Formation of Influenza Virus-Like Particles
 JOURNAL OF VIROLOGY, Jan. 2000, p. 547 551 Vol. 74, No. 1 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Plasmid-Driven Formation of Influenza Virus-Like
JOURNAL OF VIROLOGY, Jan. 2000, p. 547 551 Vol. 74, No. 1 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Plasmid-Driven Formation of Influenza Virus-Like
FurinDB: A Database of 20-Residue Furin Cleavage Site Motifs, Substrates and Their Associated Drugs
 Int. J. Mol. Sci. 2011, 12, 1060-1065; doi:10.3390/ijms12021060 OPEN ACCESS Technical Note International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms FurinDB: A Database of 20-Residue
Int. J. Mol. Sci. 2011, 12, 1060-1065; doi:10.3390/ijms12021060 OPEN ACCESS Technical Note International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms FurinDB: A Database of 20-Residue
Avian Influenza Virus H7N9. Dr. Di Liu Network Information Center Institute of Microbiology Chinese Academy of Sciences
 Avian Influenza Virus H7N9 Dr. Di Liu Network Information Center Institute of Microbiology Chinese Academy of Sciences Avian Influenza Virus RNA virus, Orthomyxoviruses Influenza A virus Eight Gene segments
Avian Influenza Virus H7N9 Dr. Di Liu Network Information Center Institute of Microbiology Chinese Academy of Sciences Avian Influenza Virus RNA virus, Orthomyxoviruses Influenza A virus Eight Gene segments
Modeling the Antigenic Evolution of Influenza Viruses from Sequences
 Modeling the Antigenic Evolution of Influenza Viruses from Sequences Taijiao Jiang Center of Systems Medicine, Chinese Academy of Medical Sciences Suzhou Institute of Systems Medicine October 8-10, 2015.
Modeling the Antigenic Evolution of Influenza Viruses from Sequences Taijiao Jiang Center of Systems Medicine, Chinese Academy of Medical Sciences Suzhou Institute of Systems Medicine October 8-10, 2015.
Transport of the Influenza Virus Genome from Nucleus to Nucleus
 Viruses 2013, 5, 2424-2446; doi:10.3390/v5102424 Review OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Transport of the Influenza Virus Genome from Nucleus to Nucleus Edward C. Hutchinson
Viruses 2013, 5, 2424-2446; doi:10.3390/v5102424 Review OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Transport of the Influenza Virus Genome from Nucleus to Nucleus Edward C. Hutchinson
CHARACTERIZING THE ROLE OF N TERMINUS OF INFLUENZA A NUCLEOPROTEIN FOR LOCATION AND VIRAL RNP ACTIVITY
 California State University, San Bernardino CSUSB ScholarWorks Electronic Theses, Projects, and Dissertations Office of Graduate Studies 6-2018 CHARACTERIZING THE ROLE OF N TERMINUS OF INFLUENZA A NUCLEOPROTEIN
California State University, San Bernardino CSUSB ScholarWorks Electronic Theses, Projects, and Dissertations Office of Graduate Studies 6-2018 CHARACTERIZING THE ROLE OF N TERMINUS OF INFLUENZA A NUCLEOPROTEIN
The RNA polymerase of influenza A virus: mechanisms of viral transcription and replication
 Acta virologica 57: 113 122, 2013 doi:10.4149/av_2013_02_113 The RNA polymerase of influenza A virus: mechanisms of viral transcription and replication E. Fodor Sir William Dunn School of Pathology, University
Acta virologica 57: 113 122, 2013 doi:10.4149/av_2013_02_113 The RNA polymerase of influenza A virus: mechanisms of viral transcription and replication E. Fodor Sir William Dunn School of Pathology, University
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Patricia Fitzgerald-Bocarsly
 FLU Patricia Fitzgerald-Bocarsly October 23, 2008 Orthomyxoviruses Orthomyxo virus (ortho = true or correct ) Negative-sense RNA virus (complementary to mrna) Five different genera Influenza A, B, C Thogotovirus
FLU Patricia Fitzgerald-Bocarsly October 23, 2008 Orthomyxoviruses Orthomyxo virus (ortho = true or correct ) Negative-sense RNA virus (complementary to mrna) Five different genera Influenza A, B, C Thogotovirus
number Done by Corrected by Doctor Ashraf
 number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
Dr. Ahmed K. Ali Attachment and entry of viruses into cells
 Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
TITLE: Influenza A (H7N9) virus evolution: Which genetic mutations are antigenically important?
 TITLE: Influenza A (H7N9) virus evolution: Which genetic mutations are antigenically important? AUTHORS: Joshua G. Petrie 1, Adam S. Lauring 2,3 AFFILIATIONS: 1 Department of Epidemiology, University of
TITLE: Influenza A (H7N9) virus evolution: Which genetic mutations are antigenically important? AUTHORS: Joshua G. Petrie 1, Adam S. Lauring 2,3 AFFILIATIONS: 1 Department of Epidemiology, University of
Correlated mutations in the four influenza proteins essential for viral RNA synthesis, host adaptation, and virulence: NP, PA, PB1, and PB2
 Vol.2, No.10, 1138-1147 (2010) http://dx.doi.org/10.4236/ns.2010.210141 Natural Science Correlated mutations in the four influenza proteins essential for viral RNA synthesis, host adaptation, and virulence:
Vol.2, No.10, 1138-1147 (2010) http://dx.doi.org/10.4236/ns.2010.210141 Natural Science Correlated mutations in the four influenza proteins essential for viral RNA synthesis, host adaptation, and virulence:
University of Groningen. Computational studies of influenza hemagglutinin Boonstra, Sander
 University of Groningen Computational studies of influenza hemagglutinin Boonstra, Sander IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
University of Groningen Computational studies of influenza hemagglutinin Boonstra, Sander IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
Structure and assembly of the influenza A virus ribonucleoprotein complex
 FEBS Letters 587 (2013) 1206 1214 journal homepage: www.febsletters.org Review Structure and assembly of the influenza A virus ribonucleoprotein complex Wenjie Zheng, Yizhi Jane Tao Department of Biochemistry
FEBS Letters 587 (2013) 1206 1214 journal homepage: www.febsletters.org Review Structure and assembly of the influenza A virus ribonucleoprotein complex Wenjie Zheng, Yizhi Jane Tao Department of Biochemistry
Fayth K. Yoshimura, Ph.D. September 7, of 7 HIV - BASIC PROPERTIES
 1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
Regulation of cell signaling cascades by influenza A virus
 The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 389 394 Regulation of cell signaling cascades
The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 389 394 Regulation of cell signaling cascades
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
The influenza virus nucleoprotein: a multifunctional RNA-binding protein pivotal to virus replication
 Journal of General Virology (2002), 83, 723 734. Printed in Great Britain... The influenza virus nucleoprotein: a multifunctional RNA-binding protein pivotal to virus replication Agustı n Portela 1 and
Journal of General Virology (2002), 83, 723 734. Printed in Great Britain... The influenza virus nucleoprotein: a multifunctional RNA-binding protein pivotal to virus replication Agustı n Portela 1 and
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Interaction of the Influenza Virus Nucleoprotein with the Cellular CRM1-Mediated Nuclear Export Pathway
 JOURNAL OF VIROLOGY, Jan. 2001, p. 408 419 Vol. 75, No. 1 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.1.408 419.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Interaction of
JOURNAL OF VIROLOGY, Jan. 2001, p. 408 419 Vol. 75, No. 1 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.1.408 419.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Interaction of
Cristina Cassetti, Ph.D.
 NIAID Extramural Research Update: Recombinant Influenza Viruses and Biosafety Cristina Cassetti, Ph.D. Influenza Program Officer Division of Microbiology and Infectious Diseases NIAID Influenza virus DMID
NIAID Extramural Research Update: Recombinant Influenza Viruses and Biosafety Cristina Cassetti, Ph.D. Influenza Program Officer Division of Microbiology and Infectious Diseases NIAID Influenza virus DMID
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Technology Overview. Summary
 Live Attenuated Influenza Vaccines with Altered NS1 Technology Overview Summary Transformative Technology: Live attenuated influenza vaccines (LAIVs) with precise, genetically stable truncations of the
Live Attenuated Influenza Vaccines with Altered NS1 Technology Overview Summary Transformative Technology: Live attenuated influenza vaccines (LAIVs) with precise, genetically stable truncations of the
Last time we talked about the few steps in viral replication cycle and the un-coating stage:
 Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Viral structure م.م رنا مشعل
 Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Protein Trafficking in the Secretory and Endocytic Pathways
 Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
Protein Trafficking in the Secretory and Endocytic Pathways The compartmentalization of eukaryotic cells has considerable functional advantages for the cell, but requires elaborate mechanisms to ensure
Influenza A Virus Multiplication and the Cellular SUMOylation System
 Chapter 1 Influenza A Virus Multiplication and the Cellular SUMOylation System Andrés Santos, Jason Chacón and Germán Rosas-Acosta Additional information is available at the end of the chapter http://dx.doi.org/10.5772/54351
Chapter 1 Influenza A Virus Multiplication and the Cellular SUMOylation System Andrés Santos, Jason Chacón and Germán Rosas-Acosta Additional information is available at the end of the chapter http://dx.doi.org/10.5772/54351
An Unconventional NLS is Critical for the Nuclear Import of the Influenza A Virus Nucleoprotein and Ribonucleoprotein
 Traffic 2005; 6: 205 213 Blackwell Munksgaard Copyright # Blackwell Munksgaard 2005 doi: 10.1111/j.1600-0854.2004.00263.x An Unconventional NLS is Critical for the Nuclear Import of the Influenza A Virus
Traffic 2005; 6: 205 213 Blackwell Munksgaard Copyright # Blackwell Munksgaard 2005 doi: 10.1111/j.1600-0854.2004.00263.x An Unconventional NLS is Critical for the Nuclear Import of the Influenza A Virus
Role of Ran Binding Protein 5 in Nuclear Import and Assembly of the Influenza Virus RNA Polymerase Complex
 JOURNAL OF VIROLOGY, Dec. 2006, p. 11911 11919 Vol. 80, No. 24 0022-538X/06/$08.00 0 doi:10.1128/jvi.01565-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Role of Ran Binding
JOURNAL OF VIROLOGY, Dec. 2006, p. 11911 11919 Vol. 80, No. 24 0022-538X/06/$08.00 0 doi:10.1128/jvi.01565-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Role of Ran Binding
Influenzavirus. Influenzavirus 10/27/15. Lord Randolphs report from Queen Marys trip to Scotland in 1562
 A disease under the influence of the stars or the moon. Lord Randolphs report from Queen Marys trip to Scotland in 1562 She fell acquaintance with a new disease which passed through her whole court neither
A disease under the influence of the stars or the moon. Lord Randolphs report from Queen Marys trip to Scotland in 1562 She fell acquaintance with a new disease which passed through her whole court neither
Reoviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Overview of virus life cycle
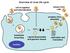 Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Reverse Genetics of RNA Viruses
 Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
LESSON 4.4 WORKBOOK. How viruses make us sick: Viral Replication
 DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
Coronaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Impact of Amino Acid Mutations in PB2, PB1-F2, and NS1 on the Replication and Pathogenicity of Pandemic (H1N1) 2009 Influenza Viruses
 JOURNAL OF VIROLOGY, May 2011, p. 4596 4601 Vol. 85, No. 9 0022-538X/11/$12.00 doi:10.1128/jvi.00029-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Impact of Amino Acid Mutations
JOURNAL OF VIROLOGY, May 2011, p. 4596 4601 Vol. 85, No. 9 0022-538X/11/$12.00 doi:10.1128/jvi.00029-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Impact of Amino Acid Mutations
REGULATORY MECHANISMS OF TRANSCRIPTION FACTOR FUNCTION
 Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) RG Clerc 04.04.2012 REGULATORY MECHANISMS OF TRANSCRIPTION FACTOR FUNCTION Protein synthesized Protein phosphorylated
Transcription Regulation And Gene Expression in Eukaryotes Cycle G2 (lecture 13709) RG Clerc 04.04.2012 REGULATORY MECHANISMS OF TRANSCRIPTION FACTOR FUNCTION Protein synthesized Protein phosphorylated
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Interaction of influenza virus proteins with nucleosomes
 Virology 332 (2005) 329 336 www.elsevier.com/locate/yviro Interaction of influenza virus proteins with nucleosomes Inmaculada Garcia-Robles a, Hatice Akarsu a,b, Christoph W. Mqller a, Rob W.H. Ruigrok
Virology 332 (2005) 329 336 www.elsevier.com/locate/yviro Interaction of influenza virus proteins with nucleosomes Inmaculada Garcia-Robles a, Hatice Akarsu a,b, Christoph W. Mqller a, Rob W.H. Ruigrok
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
numbe r Done by Corrected by Doctor
 numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
Palindromes drive the re-assortment in Influenza A
 www.bioinformation.net Hypothesis Volume 7(3) Palindromes drive the re-assortment in Influenza A Abdullah Zubaer 1, 2 * & Simrika Thapa 1, 2 1Swapnojaatra Bioresearch Laboratory, DataSoft Systems, Dhaka-15,
www.bioinformation.net Hypothesis Volume 7(3) Palindromes drive the re-assortment in Influenza A Abdullah Zubaer 1, 2 * & Simrika Thapa 1, 2 1Swapnojaatra Bioresearch Laboratory, DataSoft Systems, Dhaka-15,
MCB Chapter 11. Topic E. Splicing mechanism Nuclear Transport Alternative control modes. Reading :
 MCB Chapter 11 Topic E Splicing mechanism Nuclear Transport Alternative control modes Reading : 419-449 Topic E Michal Linial 14 Jan 2004 1 Self-splicing group I introns were the first examples of catalytic
MCB Chapter 11 Topic E Splicing mechanism Nuclear Transport Alternative control modes Reading : 419-449 Topic E Michal Linial 14 Jan 2004 1 Self-splicing group I introns were the first examples of catalytic
Protein Modeling Event
 Protein Modeling Event School Name: School Number: Team Member 1: Team Member 2: : Pre-Build Score: On-Site Build Score: Test Score: Tie Breaker: Total: Final Rank: Part I: Pre-Build (40% of total score)
Protein Modeling Event School Name: School Number: Team Member 1: Team Member 2: : Pre-Build Score: On-Site Build Score: Test Score: Tie Breaker: Total: Final Rank: Part I: Pre-Build (40% of total score)
Conserved methionine 165 of matrix protein contributes to the nuclear import and is essential for influenza A virus replication
 Švančarová and Betáková Virology Journal (2018) 15:187 https://doi.org/10.1186/s12985-018-1056-x RESEARCH Open Access Conserved methionine 165 of matrix protein contributes to the nuclear import and is
Švančarová and Betáková Virology Journal (2018) 15:187 https://doi.org/10.1186/s12985-018-1056-x RESEARCH Open Access Conserved methionine 165 of matrix protein contributes to the nuclear import and is
The Role of the NS Segment of Influenza A Virus in Setting Host Range and Pathogenicity. Matthew Luke Turnbull
 This thesis has been submitted in fulfilment of the requirements for a postgraduate degree (e.g. PhD, MPhil, DClinPsychol) at the University of Edinburgh. Please note the following terms and conditions
This thesis has been submitted in fulfilment of the requirements for a postgraduate degree (e.g. PhD, MPhil, DClinPsychol) at the University of Edinburgh. Please note the following terms and conditions
Nuclear Pore Tutorial
 Michael Brenner, Harvard January 7, 2010 Nuclear Pore Tutorial Outline (1) What is the nuclear pore/what does it look like? What are the proteins that make it up? (2) How does the nuclear pore work? The
Michael Brenner, Harvard January 7, 2010 Nuclear Pore Tutorial Outline (1) What is the nuclear pore/what does it look like? What are the proteins that make it up? (2) How does the nuclear pore work? The
Lipid raft-a gateway for passing through the cell membrane for pathogens
 16 3 2004 6 Chinese Bulletin of Life Sciences Vol. 16, No. 3 Jun., 2004 10040374(2004)03014404 200031 / GPI (GPI) Q241 Q257 R37 A Lipid rafta gateway for passing through the cell membrane for pathogens
16 3 2004 6 Chinese Bulletin of Life Sciences Vol. 16, No. 3 Jun., 2004 10040374(2004)03014404 200031 / GPI (GPI) Q241 Q257 R37 A Lipid rafta gateway for passing through the cell membrane for pathogens
7.012 Problem Set 6 Solutions
 Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
Virus Entry/Uncoating
 Virus Entry/Uncoating Delivery of genome to inside of a cell Genome must be available for first step of replication The Problem--barriers to infection Virion Barriers: Non-enveloped viruses capsid Enveloped
Virus Entry/Uncoating Delivery of genome to inside of a cell Genome must be available for first step of replication The Problem--barriers to infection Virion Barriers: Non-enveloped viruses capsid Enveloped
Virus-host interactions
 Virus-host interactions - Strategies viruses use to replicate their genomes in susceptible host cells replication - Strategies viruses use to move their genomes throughout susceptible host plants cell-to-cell
Virus-host interactions - Strategies viruses use to replicate their genomes in susceptible host cells replication - Strategies viruses use to move their genomes throughout susceptible host plants cell-to-cell
Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Pandemic Influenza influenza epidemic: realization of a worst-case scenario
 Pandemic Influenza October 9, 2006 1918 influenza epidemic: realization of a worst-case scenario First case: Albert Mitchell, Camp Funston, KS, March 11, 1918 Up to 20% of all humans infected 20-50 million
Pandemic Influenza October 9, 2006 1918 influenza epidemic: realization of a worst-case scenario First case: Albert Mitchell, Camp Funston, KS, March 11, 1918 Up to 20% of all humans infected 20-50 million
Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus from eight plasmids
 Journal of General Virology (2000), 81, 2843 2847. Printed in Great Britain... SHORT COMMUNICATION Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus
Journal of General Virology (2000), 81, 2843 2847. Printed in Great Britain... SHORT COMMUNICATION Unidirectional RNA polymerase I polymerase II transcription system for the generation of influenza A virus
Gag. Gag-Gag. TSG101 Nedd4. Two-Hybrid HIV Gag. Two-Hybrid. Two-Hybrid
 CytoTrap HIV Gag AnnexinA2 AUP1 AUP1 HIV AUP1 N Gag CytoTrap Gag-Gag TSG101 Nedd4 Two-Hybrid Two-Hybrid HIVGag Gag Two-Hybrid HIV Gag Gag HIV Two-Hybrid CytoTrap cdna Gag Gag CytoTrap Gag CytoTrap Gag-Gag
CytoTrap HIV Gag AnnexinA2 AUP1 AUP1 HIV AUP1 N Gag CytoTrap Gag-Gag TSG101 Nedd4 Two-Hybrid Two-Hybrid HIVGag Gag Two-Hybrid HIV Gag Gag HIV Two-Hybrid CytoTrap cdna Gag Gag CytoTrap Gag CytoTrap Gag-Gag
Overview: Chapter 19 Viruses: A Borrowed Life
 Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
Viral reproductive cycle
 Lecture 29: Viruses Lecture outline 11/11/05 Types of viruses Bacteriophage Lytic and lysogenic life cycles viruses viruses Influenza Prions Mad cow disease 0.5 µm Figure 18.4 Viral structure of capsid
Lecture 29: Viruses Lecture outline 11/11/05 Types of viruses Bacteriophage Lytic and lysogenic life cycles viruses viruses Influenza Prions Mad cow disease 0.5 µm Figure 18.4 Viral structure of capsid
Cellular cap-binding proteins associate with influenza virus mrnas
 Journal of General Virology (2011), 92, 1627 1634 DOI 10.1099/vir.0.029231-0 Cellular cap-binding proteins associate with influenza virus mrnas Katja Bier, Ashley York and Ervin Fodor Correspondence Ervin
Journal of General Virology (2011), 92, 1627 1634 DOI 10.1099/vir.0.029231-0 Cellular cap-binding proteins associate with influenza virus mrnas Katja Bier, Ashley York and Ervin Fodor Correspondence Ervin
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION
 Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Yutaka Sasaki 1, Kyoji Hagiwara 1, Michinori Kakisaka 1, Kazunori Yamada 1,2, Tomoyuki Murakami 1,2, Yoko Aida 1,2 * Abstract.
 Importin a3/qip1 Is Involved in Multiplication of Mutant Influenza Virus with Alanine Mutation at Amino Acid 9 Independently of Nuclear Transport Function Yutaka Sasaki 1, Kyoji Hagiwara 1, Michinori Kakisaka
Importin a3/qip1 Is Involved in Multiplication of Mutant Influenza Virus with Alanine Mutation at Amino Acid 9 Independently of Nuclear Transport Function Yutaka Sasaki 1, Kyoji Hagiwara 1, Michinori Kakisaka
علم األحياء الدقيقة Microbiology Introduction to Virology & Immunology
 علم األحياء الدقيقة Microbiology Introduction to Virology & Immunology What is a virus? Viruses may be defined as acellular organisms whose genomes consist of nucleic acid (DNA or RNA), and which obligatory
علم األحياء الدقيقة Microbiology Introduction to Virology & Immunology What is a virus? Viruses may be defined as acellular organisms whose genomes consist of nucleic acid (DNA or RNA), and which obligatory
Lecture 15. Signal Transduction Pathways - Introduction
 Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Bioinformation Volume 5
 Identification of sequence mutations affecting hemagglutinin specificity to sialic acid receptor in influenza A virus subtypes Usman Sumo Friend Tambunan*, Ramdhan Department of Chemistry, Faculty of Mathematics
Identification of sequence mutations affecting hemagglutinin specificity to sialic acid receptor in influenza A virus subtypes Usman Sumo Friend Tambunan*, Ramdhan Department of Chemistry, Faculty of Mathematics
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Basis and Clinical Applications of Interferon
 Interferon Therapy Basis and Clinical Applications of Interferon JMAJ 47(1): 7 12, 2004 Jiro IMANISHI Professor, Kyoto Prefectural University of Medicine Abstract: Interferon (IFN) is an antiviral substance
Interferon Therapy Basis and Clinical Applications of Interferon JMAJ 47(1): 7 12, 2004 Jiro IMANISHI Professor, Kyoto Prefectural University of Medicine Abstract: Interferon (IFN) is an antiviral substance
Final Report. Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD. Wendy Fong ( 方美珍 ) CNIC; Beijing, China
 Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
PB1-F2 Amino Acids Regulate Influenza A Viral Polymerase Activity
 Journal of Basic & Applied Sciences, 2014, 10, 1-6 1 PB1-F2 Amino Acids Regulate Influenza A Viral Polymerase Activity Yumi Ueda, Motoko Tanaka, Yukihiro Kyan, Mitsutaka Yoshida, Kenji Sasahara and Kyoko
Journal of Basic & Applied Sciences, 2014, 10, 1-6 1 PB1-F2 Amino Acids Regulate Influenza A Viral Polymerase Activity Yumi Ueda, Motoko Tanaka, Yukihiro Kyan, Mitsutaka Yoshida, Kenji Sasahara and Kyoko
Lahore University of Management Sciences. BIO314 Virology and Microbiology (Spring 2015)
 BIO314 Virology and Microbiology (Spring 2015) Instructor Room. Office Hours Email Telephone Secretary/TA TA Office Hours Course URL (if any) Shaper Mirza and Sadia Hamera Shaper.Mirza@uth.tmc.edu Course
BIO314 Virology and Microbiology (Spring 2015) Instructor Room. Office Hours Email Telephone Secretary/TA TA Office Hours Course URL (if any) Shaper Mirza and Sadia Hamera Shaper.Mirza@uth.tmc.edu Course
Mapping the Phosphoproteome of Influenza A and B Viruses by Mass Spectrometry
 Mapping the Phosphoproteome of Influenza A and B Viruses by Mass Spectrometry Edward C. Hutchinson, Eleanor M. Denham, Benjamin Thomas, David C. Trudgian a, Svenja S. Hester, Gabriela Ridlova b, Ashley
Mapping the Phosphoproteome of Influenza A and B Viruses by Mass Spectrometry Edward C. Hutchinson, Eleanor M. Denham, Benjamin Thomas, David C. Trudgian a, Svenja S. Hester, Gabriela Ridlova b, Ashley
Evolution of influenza
 Evolution of influenza Today: 1. Global health impact of flu - why should we care? 2. - what are the components of the virus and how do they change? 3. Where does influenza come from? - are there animal
Evolution of influenza Today: 1. Global health impact of flu - why should we care? 2. - what are the components of the virus and how do they change? 3. Where does influenza come from? - are there animal
RECAP FROM MONDAY AND TUESDAY LECTURES
 RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
Chapter 6- An Introduction to Viruses*
 Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide gel electrophoresis/genetics)
 Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
The humoral immune responses to IBV proteins.
 The humoral immune responses to IBV proteins. E. Dan Heller and Rosa Meir The Hebrew University of Jerusalem, Israel COST FA1207 meeting WG2 + WG3, Budapest, Jan. 2015 1 IBV encodes four major structural
The humoral immune responses to IBV proteins. E. Dan Heller and Rosa Meir The Hebrew University of Jerusalem, Israel COST FA1207 meeting WG2 + WG3, Budapest, Jan. 2015 1 IBV encodes four major structural
Human leukocyte antigen-b27 alleles in Xinjiang Uygur patients with ankylosing spondylitis
 Human leukocyte antigen-b27 alleles in Xinjiang Uygur patients with ankylosing spondylitis H.-Y. Zou, W.-Z. Yu, Z. Wang, J. He and M. Jiao Institute of Clinical Medicine, Urumqi General Hospital, Lanzhou
Human leukocyte antigen-b27 alleles in Xinjiang Uygur patients with ankylosing spondylitis H.-Y. Zou, W.-Z. Yu, Z. Wang, J. He and M. Jiao Institute of Clinical Medicine, Urumqi General Hospital, Lanzhou
brought to you by and REFERENCES
 brought to you by www.thebacteriophages.org and www.phage.org REFERENCES 1. Butcher, S. J., T. Dokland, P. M. Ojala, D. H. Bamford, and S. D. Fuller. 1997. Intermediates in the assembly pathway of the
brought to you by www.thebacteriophages.org and www.phage.org REFERENCES 1. Butcher, S. J., T. Dokland, P. M. Ojala, D. H. Bamford, and S. D. Fuller. 1997. Intermediates in the assembly pathway of the
Hierarchy among Viral RNA (vrna) Segments in Their Role in vrna Incorporation into Influenza A Virions
 JOURNAL OF VIROLOGY, Mar. 2006, p. 2318 2325 Vol. 80, No. 5 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.5.2318 2325.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Hierarchy among
JOURNAL OF VIROLOGY, Mar. 2006, p. 2318 2325 Vol. 80, No. 5 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.5.2318 2325.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Hierarchy among
CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION
 CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION What is Cytokine? Secreted popypeptide (protein) involved in cell-to-cell signaling. Acts in paracrine or autocrine fashion through specific cellular receptors.
CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION What is Cytokine? Secreted popypeptide (protein) involved in cell-to-cell signaling. Acts in paracrine or autocrine fashion through specific cellular receptors.
Size nm m m
 1 Viral size and organization Size 20-250nm 0.000000002m-0.000000025m Virion structure Capsid Core Acellular obligate intracellular parasites Lack organelles, metabolic activities, and reproduction Replicated
1 Viral size and organization Size 20-250nm 0.000000002m-0.000000025m Virion structure Capsid Core Acellular obligate intracellular parasites Lack organelles, metabolic activities, and reproduction Replicated
Section 6. Junaid Malek, M.D.
 Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Name Section Problem Set 6
 Name Section 7.012 Problem Set 6 Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed of lipids
Name Section 7.012 Problem Set 6 Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed of lipids
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Bio 401 Sp2014. Multiple Choice (Circle the letter corresponding to the correct answer)
 MIDTERM EXAM KEY (60 pts) You should have 5 pages. Please put your name on every page. You will have 70 minutes to complete the exam. You may begin working as soon as you receive the exam. NOTE: the RED
MIDTERM EXAM KEY (60 pts) You should have 5 pages. Please put your name on every page. You will have 70 minutes to complete the exam. You may begin working as soon as you receive the exam. NOTE: the RED
Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting
 Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting Question No. 1 of 10 Question 1. Which of the following statements about the nucleus is correct? Question #01 A. The
Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting Question No. 1 of 10 Question 1. Which of the following statements about the nucleus is correct? Question #01 A. The
Seroepidemiological Evidence of Avian Influenza A Virus Transmission to Pigs in Southern China
 JCM Accepts, published online ahead of print on 21 November 2012 J. Clin. Microbiol. doi:10.1128/jcm.02625-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 Seroepidemiological
JCM Accepts, published online ahead of print on 21 November 2012 J. Clin. Microbiol. doi:10.1128/jcm.02625-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 Seroepidemiological
Medical Virology. Herpesviruses, Orthomyxoviruses, and Retro virus. - Herpesviruses Structure & Composition: Herpesviruses
 Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
Coronaviruses cause acute, mild upper respiratory infection (common cold).
 Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
Supplementary Fig. S1. Schematic diagram of minigenome segments.
 open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
SEQUENCE FEATURE VARIANT TYPES
 SEQUENCE FEATURE VARIANT TYPES DEFINITION OF SFVT: The Sequence Feature Variant Type (SFVT) component in IRD (http://www.fludb.org) is a relatively novel approach that delineates specific regions, called
SEQUENCE FEATURE VARIANT TYPES DEFINITION OF SFVT: The Sequence Feature Variant Type (SFVT) component in IRD (http://www.fludb.org) is a relatively novel approach that delineates specific regions, called
Supplementary Discussion 1: The mechanistic model of NIK1-mediated antiviral
 doi:10.1038/nature14171 SUPPLEMENTARY DISCUSSION Supplementary Discussion 1: The mechanistic model of NIK1-mediated antiviral signaling 1. Stress-induced oligomerization of the extracellular domain of
doi:10.1038/nature14171 SUPPLEMENTARY DISCUSSION Supplementary Discussion 1: The mechanistic model of NIK1-mediated antiviral signaling 1. Stress-induced oligomerization of the extracellular domain of
Interaction of Polymerase Subunit PB2 and NP with Importin a1 Is a Determinant of Host Range of Influenza A Virus
 Interaction of Polymerase Subunit PB2 and NP with Importin a1 Is a Determinant of Host Range of Influenza A Virus Gülsah Gabriel, Astrid Herwig, Hans-Dieter Klenk * Institute of Virology, Philipps University
Interaction of Polymerase Subunit PB2 and NP with Importin a1 Is a Determinant of Host Range of Influenza A Virus Gülsah Gabriel, Astrid Herwig, Hans-Dieter Klenk * Institute of Virology, Philipps University
Virology 396 (2010) Contents lists available at ScienceDirect. Virology. journal homepage:
 Virology 396 (2010) 125 134 Contents lists available at ScienceDirect Virology journal homepage: www.elsevier.com/locate/yviro Mechanisms and functional implications of the degradation of host RNA polymerase
Virology 396 (2010) 125 134 Contents lists available at ScienceDirect Virology journal homepage: www.elsevier.com/locate/yviro Mechanisms and functional implications of the degradation of host RNA polymerase
