Genotypes of patients with factor XI deficiency Mutation Location Domain Type Genotype Origin Activity U/dL
|
|
|
- Andrew Blankenship
- 5 years ago
- Views:
Transcription
1 Genotypes of patients with factor XI deficiency Mutation Location Domain Type Genotype Origin Activity U/dL Antigen U/dL Expression comments Reference nt-1296g>a Promoter Promoter Regulatory Comp het Nigeria 3 1 nt 823 del C Exon 8 Apple 3 Frameshift Met-18Ile* Cys128stop* Exon 2 Exon 5 Signal Apple 2 Missense Comp het <1 <3 2 3 Met-18Ile* Exon 2 Signal Missense Het 43,50 Yes 2 3 Ser-4Leu Exon 2 Signal Missense Hom Morocco Gly-1Arg Exon 2 Signal Missense Comp het China Gln263stop Exon 8 Apple3 Nonsense IVS2+6 T>G + Intron 2 Splicing Hom Czechoslovakia ins AG Glu1stop Exon 2 Apple 1 Nonsense Het 43 2 nt 73 del 14 bp Exon 3 Apple 1 Frameshift Comp het Non-Ashkenazi <1 <1 7 Glu117stop Exon 5 Apple 2 Nonsense Jew nt 78 del GGA Exon 3 Apple 1 Frameshift Het Italy Thr10Ala Exon 3 Apple 1 Missense Comp het 22 9 Cys527Arg* Exon 14 Catalytic Missense Asp16His* Exon 3 Apple 1 Missense Het Yes 10,11 Val20Ala Exon 3 Apple 1 Missense Het France 32 Yes 7 Pro23Leu Exon 3 Apple 1 Missense Het Africa Pro23Gln Exon 3 Apple 1 Missense Het 56 2 Ser24Arg Exon 3 Apple 1 Missense Het Ala25Thr Exon 3 Apple 1 Missense Comp het Glu602Gln Exon 15 Catalytic Missense Cys28Phe Exon 3 Apple 1 Missense Het Gln29His Exon 3 Apple 1 Missense Het 2 Thr33Pro* Exon 3 Apple 1 Missense Het Iran Thr33Pro* Exon 3 Apple 1 Missense Het Turkey Yes 16 Thr33Ile Exon 3 Apple 1 Missense Het Iran Cys38Arg* Exon 3 Apple 1 Missense Hom French Basque <1 Yes Founder mutation 17
2 Cys38Arg* Exon 3 Apple 1 Missense Het Cys38Arg* Exon 3 Apple 1 Missense Comp het Arg308His Exon 9 Apple 4 Missense Cys38Arg* Exon 3 Apple 1 Missense Comp het Portugal <1 12 Pro382Leu* Exon 11 Catalytic Missense Cys38Arg* Exon 3 Apple 1 Missense Comp het French Basque 1-2 Yes 17 Tyr493His Exon 13 Catalytic Missense Yes Cys38stop Exon 3 Apple 1 Nonsense Het Ireland Cys38Trp Exon 3 Apple 1 Missense Het Yes 8 nt 192 ins G Exon 3 Apple 1 Frameshift Comp het West France <1 <1 19 Gln88stop* Exon 4 Apple 1 Nonsense Arg54Pro Exon 3 Apple 1 Missense Het Czechoslovakia Yes 8 IVS3+2 T>A Intron 3 - Splicing Comp het Czechoslovakia Lys252Ile* Exon 8 Apple 3 Missense nt 226 del T Exon 4 Apple 1 Frameshift Hom France 1 20 Cys58Arg Exon 4 Apple 1 Missense Het 42 2 Cys58Tyr Exon 4 Apple 1 Missense Het Yemenite Jew 25 Yes 7 Cys58Phe Exon 4 Apple 1 Missense Hom <1 2 Ser78Phe* Exon 4 Apple 1 Missense Het Gly79Ala Exon 4 Apple 1 Missense Comp het Czechoslovakia Glu117stop* Exon 5 Apple 2 Nonsense Ser81Tyr Exon 4 Apple 1 Missense Het Lys83Arg Exon 4 Apple 1 Missense Het nt 308 ins 7bp Exon 4 Apple 1 Frameshift Het Portugal Gln88stop* Exon 4 Apple 1 Nonsense Hom West France <1 <1 19 Gln88stop* Exon 4 Apple 1 Nonsense Comp het West France Thr575Met* Exon 15 Catalytic Missense nt 325 G>A* Exon 4 Apple 1 Splicing Het Italy 52 Yes 22 nt 325 G>A* Exon 4 Apple 1 Splicing Het Australia Reported as Ala91Thr nt 325 G>A* Exon 4 Apple 1 Splicing Comp het Australia 2 23 Gly104Asp* Exon 5 Apple 2 Missense nt 325 G>A* Phe283Leu* Exon 4 Exon 9 Apple 1 Apple 4 Splicing Missense Comp het Italy
3 nt 325 G>A* Reported as Ala91Thr Glu323Lys* Exon 4 Exon 9 Apple 1 Apple 4 Splicing Missense Comp het 9 2 IVS4+1 G>A* Intron 4 Splicing Het 45 2 IVS4+1 G>A* Intron 4 - Splicing Comp het 2 2 Cys128stop* Exon 5 Apple 2 Nonsense Cys92Gly Exon 5 Apple 2 Missense Het Vietnam Met102Thr Exon 5 Apple 2 Missense Het 40,45 Yes 2,3 Gly104Asp* Exon 5 Apple 2 Missense Het 33 2,11 Gln116stop* Exon 5 Apple 2 Nonsense Het Austria Gln116stop* Exon 5 Apple 2 Nonsense Comp het Italy < Thr123Met Exon 5 Apple 2 Missense Glu117stop (type II)* Exon 5 Apple 2 Nonsense Hom Ashkenazi and non Ashkenazi Jews, Arabs Glu117stop* Exon 5 Apple 2 Nonsense Hom Portugal (non Jewish) Glu117stop* Exon 5 Apple 2 Nonsense Hom Italy (non Jewish) <1 1 Yes Common in Ashkenazi and Iraqi Jews, Arabs. Founder mutation 1 2 unrelated families with Jewish haplotype 1 Several unrelated patients with the Jewish haplotype Glu117stop* Exon 5 Apple 2 Nonsense Comp het Italy <1 34 Cys118Arg Exon 5 Apple 2 Missense Glu117stop* Exon 5 Apple 2 Nonsense Comp het Italy < Cys122Tyr* Exon 5 Apple 2 Missense Glu117stop* Exon 5 Apple 2 Nonsense Comp het France <1 20 Cys212Ser Exon 7 Apple 3 Missense Glu117stop* Exon 5 Apple 2 Nonsense Comp het <1 <3 2 26,27,28,29, 30, ,33
4 Gln233stop* Exon 7 Apple 3 Nonsense Glu117stop* Cys237Tyr and Cys321Phe Exon 6 Exon 8 Exon 9 Apple 1 Apple 3 Apple 4 Nonsense Missense Polymorphism Comp het Mixed Basque/ Jewish <1 Yes Yes Yes 17 Glu117stop* (type II) Phe283Leu* (type III) Glu117stop* Cys309stop* Glu117stop* Gly350Arg Glu117stop* Tyr427Cys Exon 5 Exon 9 Apple 2 Nonsense Missense Comp het Ashkenazi Jewish 2-8 Common in Ashkenazi Jews Exon 5 Exon 9 Apple 2 Apple 4 Nonsense Nonsense Comp het Italy < Exon 5 Apple 2 Nonsense Comp het France <1 <1 20 Exon 10 Apple 4 Missense Exon 5 Apple 2 Nonsense Comp het Ashkenazi Jew <1 <1-7 Exon 12 Catalytic Missense Yes Cys118stop* Exon 5 Apple 2 Nonsense Hom Italy <1 32 Cys118stop* Exon 5 Apple 2 Nonsense Comp het <1 2 Pro382Leu* Exon 11 Catalytic Missense Cys122Tyr* Exon 5 Apple 2 Missense Hom Italy 4 3 Yes 8 Cys122Tyr* Exon 5 Apple 2 Missense Het 25 4 Cys128stop* Exon 5 Apple 2 Nonsense Het UK Cys128stop* Exon 5 Apple 2 Nonsense Hom UK <1 36,37 Cys128stop* Exon 5 Apple 2 Nonsense Comp het UK <1 <1 36 Undefined Cys128stop* Exon 5 Apple 2 Nonsense Comp het <2 36 nt? ins 1 bp Exon 4 Apple 1 Frameshift Cys128stop* Exon 5 Apple 2 Nonsense Comp het Australia <2 23 Gene deletion* Deletion Thr132Met Exon 5 Apple 2 Missense Het 23 2 Tyr133Cys* Exon 5 Apple 2 Missense Het 2,14 Tyr133Cys* Exon 5 Apple 2 Missense Comp het <1 2 Cys128stop* Exon 5 Apple 2 Nonsense Tyr133Ser* Exon 5 Apple 2 Missense Het 50 Yes 2,3 Ala134Pro Exon 5 Apple 2 Missense Hom <1 2 IVS5+5 G>C Intron 5 Splicing 10 26,27,28
5 IVS5-2 A>G* Intron 5 Splicing Hom 3 11 IVS5-2 A>G* Intron 5 Splicing Het Italy ,4 Gly155Glu Exon 6 Apple 2 Missense Het IVS6+3 A>G* Intron 6 Splicing Het Basque 40 2,17,20,25 France IVS6+3 A>G* Intron 6 Splicing Hom Mexico <1 7 IVS6+3 A>G* Intron 6 - Splicing Comp het Italy Arg184Gly Exon 7 Apple 3 Missense Yes Ala181Val Exon 7 Apple 3 Missense Comp het France Ala412Thr* Exon 11 Catalytic Missense Cys182Tyr* Exon 7 Apple 3 Missense Het 34 2 Cys182Tyr* Exon 7 Apple 3 Missense Het Australia Pro188Ser* Exon 7 Apple 3 Missense Het France 30 4 Pro188Ser* Cys527Arg* Exon 7 Exon 14 Apple 3 Catalytic Missense Missense Comp het 3 9 nt 644 del 6 bp* Exon 7 Apple 3 In frame del Het Austria nt 644 del 6 bp* Exon 7 Apple 3 In frame del Het Iran nt 644 del 6 bp* Exon 7 Apple 3 In frame del Comp het Italy <1 5 Yes 32 Nonsense Glu117stop* Exon 5 Apple 2 Arg210stop Exon 7 Apple 3 Nonsense Comp het Lebanon <1 <3 19 Gly336Arg Exon 10 Apple 4 Missense Cys212Arg Exon 7 Apple 3 Missense Hom <1 2 Gly217Ser Exon 7 Apple 3 Missense Comp het Turkey Yes 16 Trp501stop* Exon 13 catalytic Nonsense Phe221Ser Exon 7 Apple 3 Missense Hom Japan <3 Yes 39 nt 717 ins T Exon 7 Apple 3 Frameshift Comp het Australia Arg250His Exon 8 Apple 3 Missense Ser225Phe Exon 7 Apple 3 Missense Het Yes 40 Gln226stop* Exon 7 Apple 3 Nonsense Hom Japan <3 39 Gln226stop* Exon 7 Apple 3 Nonsense Comp het Japan <3 39 Gly400Val* Exon 11 Catalytic Missense Gln226Arg* Ser248Asn Exon 7 Exon 8 Apple 3 Apple 3 Polymorphism Missense Het African American Yes Yes Polymorphism 41,42,43
6 Trp228stop Exon 7 Apple 3 Nonsense Comp het China 44 Trp383stop Exon 11 Catalytic Nonsense Trp228Cys Exon 7 Apple 3 Missense Hom Italy 1.6 <5 45 Arg234Ile Exon 7 Apple 3 Missense Het 38 2 Arg234Lys Exon 7 Apple 3 Missense Het Australia Arg234Ser* Exon 8 Apple 3 Missense Het 42 2 Arg234Ser* Exon 8 Apple 3 Missense Comp het Undefined Glu243Asp Exon 8 Apple 3 Missense Comp het France Pro520Leu* Exon 14 Catalytic Missense Gly245Glu Exon 8 Apple 3 Missense Het Arg250Cys* Exon 8 Apple 3 Missense Hom 8 2 Arg250Cys* Exon 8 Apple 3 Missense Het 53 2 Lys252Ile* Exon 8 Apple 3 Missense Comp het Caucasian 4 Yes 2,46 Cys128stop* Exon 5 Apple 2 Nonsense Lys252Ile* Exon 8 Apple 3 Missense Comp het Australia Arg308Cys* Exon 9 Apple 4 Missense Gln263stop* Exon 8 Apple 3 Nonsense Hom Japan <1 <1 47 Gln263stop* Exon 8 Apple 3 Nonsense Comp het Japan <1 <1 48 Undefined Gln263stop* Exon 8 Apple 3 Nonsense Comp het China 1 49 Tyr351stop Exon 10 Catalytic Nonsense Val271Leu Exon 8 Apple 3 Splicing? Comp het India Gly460Arg* Exon 12 Catalytic Missense Phe283Leu* (type III) Exon 9 Apple 4 Missense Hom Ashkenazi Jewish Phe283Leu* (type III) 3-14 Yes Common in Ashkenazi Jews, founder mutation Exon 9 Apple 4 Missense Het Italy 4 subjects with the Jewish haplotype Phe283Leu* Exon 9 Apple 4 Missense Comp het Ashkenazi Jew Glu323Lys* Exon 9 Apple 4 Missense Phe283Leu* Exon 9 Apple 4 Missense Comp het Czechoslovakia Ala412Thr* Exon 11 catalytic Missense Phe283Leu* Exon 9 Apple 4 Missense Comp het France 1 < ,27,28,29, 30,51 33
7 Arg479stop* Exon 13 catalytic Nonsense nt 908 del G Exon 9 Apple 4 Frameshift Het nt 919 del G Exon 9 Apple 4 Frameshift Het 14 Glu297Lys* Exon 9 Apple 4 Missense Het France 40, ,12 Glu297Lys* Exon 9 Apple 4 Missense Het Israeli Arab Glu297Lys* Exon 9 Apple 4 Missense Het Glu297Lys* Exon 9 Apple 4 Missense Comp het Belgium <2 <5 7 Cys527Tyr Exon 14 Catalytic Missense Yes nt 951 ins 19 bp Exon 9 Apple 4 Frameshift Het Leu302Pro Exon 9 Apple 4 Missense Yes 10 nt961 del TG* Exon 9 Apple 4 Frameshift Hom France <1 <1 4,7 nt961 del TG* Exon 9 Apple 4 Frameshift Het 57 2 Thr304Ile* Exon 9 Apple 4 Missense Het 51 Yes 2,10 Thr304Ile* Exon 9 Apple 4 Missense Hom France Val307Phe* Exon 9 Apple 4 Missense Het Val307Phe* Exon 9 Apple 4 Missense Het Arg308Cys* Exon 9 Apple 4 Missense Het Caucasian 41, ,52 Cys309stop* Exon 9 Apple 4 Nonsense Comp het Italy <1 <2 24 Gly578Cys Exon 15 Catalytic Missense Thr313Ile Exon 9 Apple 4 Missense Hom <1 15 Glu323Lys* Exon 9 Apple 4 Missense Yes 10 nt1026 G>T* Exon 9 Apple 4 Splicing Comp het Portugal 2 7 Cys398Tyr* Exon 11 Catalytic Missense nt1026 G>T* Exon 9 Apple 4 Splicing Comp het Portugal Lys518Asn Exon 14 Catalytic Missense nt 1027 ins G Exon 9 Apple 4 Frameshift Comp het Caucasian 4 Unknown 53 Phe283Leu* Missense Jewish ancestry IVS9+5 G>T Intron 9 Splicing Het IVS9-2 A>G Intron 9 Splicing 10 nt1075 del A* Exon 10 Apple 4 Frameshift Hom Portugal 1 21 nt1075 del A* Exon 10 Apple 4 Frameshift Hom Morocco 2 <5 7 nt1075 del A* Exon 10 Apple 4 Frameshift Comp het Portugal 2 7
8 Cys398Tyr* Exon 11 Catalytic Missense Ile341Met Exon 10 Apple 4 Missense Het 2 Leu342Pro Exon 10 Apple 4 Missense Hom Iran <1 15 Gly344Arg Exon 10 Apple 4 Missense Het 38 2 Gly350Ala Exon 10 Apple 4 Missense Comp het West France Cys581stop* Exon 15 Catalytic Nonsense Gly350Glu Exon 10 Apple 4 Missense Japan Yes 54 Tyr351Ser Exon 10 Apple 4 Missense Hom India <1 50 Leu355Ser Exon 10 Apple 4 Missense Het 67 2 Cys356Arg Exon 10 Apple 4 Missense Het 40 2 IVS10+1 G>A Intron 10 Splicing Het Japan IVS10+5 G>A* Intron 10 Splicing Het 73 2 IVS10+5 G>A* Intron 10 Splicing Het IVS10-4 del Intron 10 Splicing Hom China <10 <10 55 gttg Val371Ile Exon 11 Catalytic Missense Het Italy Yes 56 Gly372Ala Exon 11 Catalytic Missense Comp het Iran 4 57 Glu547Lys* Exon 14 Catalytic Missense Ala375Val Exon 11 Catalytic Missense Het Arg378Cys Exon 11 Catalytic Missense Het Yes 2,3 Trp381Leu Exon 11 Catalytic Missense Het France Thr386Asn Exon 11 Catalytic Missense Hom Arab 2 58 Leu387Gln Exon 11 Catalytic Missense Het His388Pro Exon 11 Catalytic Missense Het Thr389Pro Exon 11 Catalytic Missense Het 35 2 Cys398Tyr* Exon 11 Catalytic Missense Hom <1 Yes 2,40 Cys398Tyr* Exon 11 Catalytic Missense Het 25,39 2,40 Gly400Val* Exon 11 Catalytic Missense Het Italy/ 15 <20 Yes 31 Czechoslovakia Gly400Val* Exon 11 Catalytic Missense Japan <1 31,59 (Nagoya II) Gly400Val* Exon 11 Catalytic Missense Hom China 2 2,31 Gln406stop Exon 11 Catalytic Nonsense Het 51 2 Trp407Cys Exon 11 Catalytic Missense Het Africa Thr410Ile Exon 11 Catalytic Missense Het 38 13
9 Ala412Ser Exon 11 Catalytic Missense Het Ala412Thr* Exon 11 Catalytic Missense Hom Australia 2 23 Ala412Val* Exon 11 Catalytic Missense Het Ala412Val* Exon 11 Catalytic Missense Comp het <1 14 Gene deletion* IVS11+12 G>A Intron 11 Splicing Het IVS11-10 T>A Intron 11 Splicing Hom Portugal 1 21 Arg425Cys Exon 12 Catalytic Missense Het 46 2 Gln433Glu Exon 12 Catalytic Missense Comp het Japan <1 60 nt 1560 ins G* Exon 13 Catalytic Frameshift Phe442Val Exon 12 Catalytic Missense Het Glu447stop Exon 12 Catalytic Nonsense Comp het Japan <1 61 nt1560 ins G* Exon 13 Frameshift Gly460Arg* Exon 12 Catalytic Missense Hom <1 2 Gly460Arg* Exon 12 Catalytic Missense Het Gly460Arg* Exon 12 Catalytic Missense Hom India <1 Another patient 12,50 from Sri Lanka Thr475Ile* Exon 12 Catalytic Missense Het Yes 2,23,62 Thr475Ile* Exon 12 Catalytic Missense Het France Arg479stop* Exon 13 Catalytic Nonsense Het 55 2 Cys482Arg Exon 13 Catalytic Missense Comp het 9 13 Undefined Cys482Trp Exon 13 Catalytic Missense Het 37 2 Ser485Pro Exon 13 Catalytic Missense Hom Iran <1 15 Trp497Gly Exon 13 Catalytic Missense Het Italy Trp497Cys Exon 13 Catalytic Missense Het Sri Lanka Val498Met Exon 13 Catalytic Missense Comp het Korea 1 63 nt 1560 ins G* Exon 13 Catalytic Frameshift Trp501stop Exon 13 Catalytic Nonsense Hom Japan <5 <5 64 Trp501Cys* Exon 13 Catalytic Missense Hom Lebanon 1.6 <1 65,66 IVS13+2 T>G Exon 13 Splicing Het Ashkenazi Jew Pro520Leu* Exon 14 Catalytic Missense Het Yes 2,14,67 Pro520Leu* Exon 14 Catalytic Missense Het France Pro520Leu* Exon 14 Catalytic Missense Het Combined with heterozygote 68
10 FVII deficiency Gly544Ser Exon 14 catalytic Missense Het Glu547Lys* Exon 14 Catalytic Missense Het ,14 Glu547Lys* Exon 14 Catalytic Missense Hom nt1714 del 3+ IVS14 del 11* (type IV) Glu117stop* IVS14+1 G>A* (type I) Glu117stop* IVS14+1 G>A* (type I) Exon14 Exon 5 Catalytic Apple 2 Splicing Nonsense Comp het Ashkenazi Jewish Intron 14 Catalytic Splicing Comp het Ashkenazi Jewish Exon 5 Apple 2 Nonsense Intron 14 Catalytic Splicing Hom Ashkenazi Jewish <1 69 <1 26 <1 Prevalent in Ashkenazi Jews. Founder mutation IVS14+1 G>A* Intron 14 Catalytic Splicing Het 71 2 IVS14-2 A>G Intron 14 Catalytic Splicing Het Iran Gly555Glu* Exon 15 Catalytic Missense Hom Bucharan Jew <1 100 Yes 71 Gly555Glu* Exon 15 Catalytic Missense Het 51 2 Cys563Phe Exon 15 Catalytic Missense Het 45 2 Trp569Ser Exon 15 Catalytic Missense Het Germany <20 Yes 31 Thr575Met* Exon 15 Catalytic Missense Hom Lebanon Thr575Met* Exon 15 Catalytic Missense Het Yes 2,3 Ser576Arg Exon 15 Catalytic Missense Het Caucasian ,52 Glu579Lys Exon 15 Catalytic Missense Het 15 4 Cys581stop* Exon 15 Catalytic Nonsense Hom West France Tyr590His Exon 15 Catalytic Missense Hom Iran <1 15 Tyr590stop* Exon 15 Catalytic Nonsense Het 30 2,14 Trp599Arg Exon 15 Catalytic Missense Hom Japan <1 <1 72 Ile600Ser* Exon 15 Catalytic Missense Hom <2 2 Ile600Ser* Exon 15 Catalytic Missense Het ,14 Leu601Pro Exon 15 Catalytic Missense Hom Italy <1 <1 Yes 8 Exons Catalytic Big deletion Hom Tunis <1 7 deletion Gene deletion* Whole gene Het 32 2,14,73 70
11 deletion I Gene deletion* whole gene deletion II Het 2 Nucleotide numbers are based on the Genebank file M13142 using the A (nucleotide 44) of the ATG initiator methionine as +1. *A mutation that was identified in more than 1 family.
12 Mutations causing Factor XI deficiency according to their types Promoter Missense Nonsense Splice Deletion/Insertion nt G>A M-18I* S81Y A184G T304I* P382L* W497G G1X IVS2+6 T>G + 10 ins AG nt 73 del 14 bp nt -54 G>A S-4L K83R P188S* V307F* T386N W497C C38X nt 325 G>A* nt 78 del GGA G-1R C92G C212R R308C* L387Q V498M Q88X* IVS3+2 T>A nt 192 ins G T10A M102T C212S R308H H388P W501C* Q116X* IVS4+1 G>A* nt 226 del T D16H* G104D* G217S T313I T389P K518N E117X* IVS5+5 G>C nt 308 ins 7 bp V20A C118R F221S C321F C398Y* P520L* C118x* IVS5-2 A>G* nt 643 del 6 bp* P23L C122Y* S225F E323K* G400V* C527Y C128X* IVS6+3 A>G* nt 717 ins T P23Q T123M W228C G336R W407S C527R* R210X IVS9 +5 G>T nt 823 del C S24R T132M R234I I341M T410I G544S Q226X IVS9-2A>G nt 908 del G A25T Y133S* R234K L342P A412T* E547K* W228X IVS10+1 G>A nt 919 del G C28F Y133C* R234S* G344R A412V* G555E* Q233X IVS10+5 G>A* nt dup ins 19 bp Q29H A134P C237Y G350R A412S C563F Q263X* IVS10-4 del GTTG nt 961 del TG* T33P* G155E E243D G350A R425C W569S C309X* IVS11+12 G>A (nt1304)t nt 1026 G>T* T33I L172P G245E G350E Y427C T575M* Y351X IVS11-10 T>A nt 1027 ins G C38R* A181V S248N Y351S Q433E S576R W383X IVS13+2 T>G nt 1075 del A* C38W C182Y* R250C L355S F442V G578C Q406X nt 1714 del 3+ IVS14 del 14 nt 1560 ins G R54P A184G R250H C356R G460R* E579K E447X IVS14+1G>A* C58R P188S* K252I* V371I T475I* Y590H R479X* IVS14-2 A>G C58Y G155E V271Lsplice? G372A C482W W599R W501X* C58F L172P F283L* A375V C482R I600S* C581X* S78F* A181V E297K* R378C S485P L601P Y590X* G79A C182Y* L302P W381L Y493H E602Q L601P * Mutation was identified in more than 1one family
13 References 1. Vasileiadis I, El-Ali M, Nanas S, et al: First diagnosis of factor XI deficiency in a patient with subarachnoid haemorrhage. Blood Coagul Fibrinolysis 20:309, Mitchell M, Mountford r, Butler R, et al: Spectrum of factor XI (F11) mutations in the UK population 116 index cases and 140 mutations. Hum Mutat 27:829, Mitchell MJ, Dai L, Clarke JB, et al: Characterisation of five factor XI mutations. Thromb Haemost 97:884, Quelin F, Mathonnet F, Potentini-Esnault C, et al: Identification of five novel mutations in the factor XI gene (F11) of patients with factor XI deficiency. Blood Coagul Fibrinolysis 17:69, Wang J, Wang X, Dai J, et al: A case of factor XI deficiency caused by compound heterozygous F11 gene mutation. Haemophilia 15:603, Castaman G, Giacomelli SH, Habart D, et al: Factor XI gene mutations in factor XI deficient patients of the Czech Republic. Am J Hematol 83:916, Zucker M, Zivelin A, Landau M, et al: Characterization of seven novel mutations causing factor XI deficiency. Haematologica 92:1375, Spena S, Asselta R, Caccia S, et al: Analysis of the structural effects of four novel and a previously known mutations causing factor XI deficiency. Thromb Haemost 102:603, Rugeri L, Quelin F, Chatard B, et al: Thrombin generation in patients with factor XI deficiency and clinical bleeding risk. Haemophilia 16:771, Pugh RE, McVey JH, Tuddenham EGD, Hancock JF: Six point mutations that cause factor XI deficiency. Blood 85:1509, Mitchell M, Harrington P, Cutler J, et al: Eighteen unrelated patients with factor XI deficiency, four novel mutations and a 100% detection rate by denaturing highperformance liquid chromatography. Br J Haematol 121:500, Quelin F, Francois D, d'oiron R: Factor XI deficiency: Identification of six novel missense mutations (P23L, P69T, C92G, E243D, W497C and E547K). Haematologica 90:1149, Saunders RE, Shiltagh N, Gomez K, et al: Structural analysis of eight novel and 112 previously reported missense mutations in the interactive FXI mutation database reveals new insight on FXI deficiency. Thromb Haemost 102:287, Hill M, McLeod F, Franks H, et al: Genetic analysis in FXI deficiency: six novel mutations and the use of a polymerase chain reaction-based test to define a whole gene deletion. Br J Haematol 129:825, 2005.
14 15. Fard-Esfahani P, Lari GR, Ravanbod S, et al: Seven novel point mutations in the F11 gene in Iranian FXI-deficient patients. Haemophilia 14:91, Berber E, Rimoldi V, Usluer S, et al: Characterization of the genetic basis of FXI deficiency in two Turkish patients. Haemophilia 16:564, Zivelin A, Bauduer F, Ducout L, et al: Factor XI deficiency in French Basques is caused predominantly by an ancestral Cys38Arg mutation in the factor XI gene. Blood 99:2448, Ramadan KM, McNulty O, Anderson JA, Jones FG, Winter PC: A novel factor XI gene mutation associated with mild factor XI deficiency in a symptomatic patient. Blood Coagul Fibrinolysis 17:499, Quelin F, Trossaert M, Sigaud M et al: Molecular basis of severe factor XI deficiency in seven families from the west of France. Seven novel mutations, including an ancient Q88X mutation. J Thromb Haemost 2:71, Quelin F, Frere C, Pouymayou C, et al: Prospective analysis of factor XI deficiencies in the Marseilles area identified four novel mutations among 12 consecutive unrelated families. Blood Coagul fibrinolysis 20:84, Ventura C, Santos AIM, Tavares A, et al: Molecular genetic analysis of factor XI deficiency: Identification of five novel gene alterations and the origin of type II mutation in Portuguese families. Thromb Haemost 84:833, Guella I, Solda G, Spena S, et al: Molecular characterization of two novel mutations causing factor XI deficiency: A splicing defect and a missense mutation responsible for a CRM+ defect. Thromb Haemost 99:523, Duncan EM, Casey GJ, Fenech MP, et al: Partial and severe factor XI deficiency in South Australia and the usefulness of factor XI mutation analysis for diagnosis. Pathology 40:401, Castaman G, Giacomelli SH, Dragani A, et al: Severe factor XI deficiency in the Abruzzo region of Italy is associated to different FXI gene mutations. Haematologica 93:957, Dossenbach-Glaninger A, Hopmeir P: Coagulation factor XI gene analysis in three factor XI deficient Austrian patients. Eur J Haematol 76:317, Asakai R, Chung DW, Batnoff OD, Davie EW: Factor XI (plasma thromboplastin antecedent) deficiency in Ashkenazi Jews is a bleeding disorder that can result from three types of point mutations. Proc Natl Acad Sci USA 86:7667, Asakai R, Chung DW, Davie EW, Seligsohn U: Factor XI deficiency in Ashkenazi Jews in Israel. New Engl J Med 325:153, Hancock JF, Wieland K, Pugh RE, et al: A molecular genetic study of factor XI deficiency. Blood 77:1942, 1991.
15 29. Shpilberg O, Peretz H, Zivelin A, et al: One of the two common mutations causing factor XI deficiency in Ashkenazi Jews (type II) is also prevalent in Iraqi Jews, who represent the ancient gene pool of Jews. Blood 85:429, Peretz H, Mulai A, Usher S, et al: The two common mutations causing factor XI deficiency in Jews stem from distinct founders: One of ancient Middle Eastern origin and another of more recent European origin. Blood 90:2654, Kravtsov DV, Wu W, Meijers JC, et al: Dominant factor XI deficiency caused by mutations in the factor XI catalytic domain. Blood 104:128, Zadra G, Asselta R, Malcovati M, et al: Molecular genetic analysis of severe coagulation factor XI deficiency in six Italian patients. Haematologica 89:1332, Zadra G, Asselta R, Tenchini ML et al: Simultaneous genotyping of coagulation factor XI type II and type III mutations by multiplex real-time polymerase chain reaction to determine their prevalence in healthy and factor XI-deficient Italians. Haematologica 93:715, Tomaiuolo M, Favuzzi G, Cappucci F, et al: Factor XI deficiency: Two novel mutations in asymptomatic Italian patients. Haemophilia 16:767, Imanaka Y, Lal K, Nishimura T, et al: Identification of two novel mutations in non- Jewish factor XI deficiency. Br J Haematol 90:916, Bolton-Maggs PHB, Peretz H, Butler R, et al: A common ancestral mutation (C128X) occurring in 11 non-jewish families from the UK with factor XI deficiency. J Thromb Haemost 2:918, Zacharski LR, French EE: Factor XI (PTA) deficiency in an English-American kindred. Thromb Haemost 39:215, De Raucourt E, De Mazancourt P, Quelin F: Four novel FXI gene mutations in three factor XI-deficient patients. Blood Coagul Fibrionolysis 19:240, Okumura K, Kyotani M, Kawai R, et al: Recurrent mutations of factor XI gene in Japanese. Int J Hematol 83:462, Kravtsov DV, Monahan PE, Gailani D: A classification system for cross-reactive material negative factor XI deficiency. Blood 105:4671, Martincic D, Zimmerman SA, Ware RE, et al: Identification of mutations and polymorphisms in the factor XI genes of an African American family by Dideoxyfingerprinting. Blood 92:3309, Cargill M, Altshuler D, Ireland J, et al: Characterization of single-nucleotide polymorphisms in coding regions of human genes. Nat Genet 22:231, 1999.
16 43. Sun MF, Baglia FA, Ho D, et al: Defective binding of factor XI-N248 to activated human platelets. Blood 98:125, Wu WM, Want HL, Wang XF, et al: Identification of two novel factor XI non-sense mutation Trp228stop and Trp383stop in a Chinese pedigree of congenital factor XI deficiency. Zhonghua Xue Ye Xue Za Zhi 24:126, Alhaq A, Mitchell M, Sethi M, et al: Identification of a novel mutation in a non-jewish factor XI deficient kindred. Br J Haematol 104:44, Dai L, Mitchell M, Carson P, et al: Severe factor XI deficiency caused by compound heterozygosity. Br J Haematol 125:817, Sato E, Kawamata N, Kato A, Oshimi K: A novel mutation that leads to a congenital factor XI deficiency in a Japanese family. Am J Hematol 63:165, Kawaguchi T, Koga S, Hongo H, et al: A novel type of factor XI deficiency showing compound genetic abnormalities: a nonsense mutation and an impaired transcription. Int J Hematol 71:84, Au WY, Cheung JW, Lam CCK, Kwong YL: Two factor XI mutations in a Chinese family with factor XI deficiency. Am J Hematol 74:136, Jayandharan G, Shaji RV, Nair SC, et al: Novel missense mutations in two patients with factor XI deficiency (Val271Leu and Tyr351Ser) and one patient with combined factor XI and factor IX deficiency (Phe349Val). J Thromb Haemost 3:808, Meijers JC, Davie EW, Chung DW: Expression of human blood coagulation factor XI: Characterization of the defect in factor XI type III deficiency. Blood 79:1435, Mitchell M, Cutler J, Thompson S, et al: Heterozygous factor XI deficiency association with three novel mutations. Br J Haematol 107:763, Dossenbach-Glaninger A, Krugluger W, Schrattbauer K, et al: Severe factor XI deficiency caused by compound heterozygosity for the type III mutation and a novel insertion in exon 9 (codons 324/325+G). Br J Haematol 114:875, Meijers JC, Mulvihill ER, Davie EW, Chung DW: Apple four in human blood coagulation factor XI mediates dimmer formation. Biochemistry 31:4680, Xie S, Wang HL, Wang XF, et al: The inherited coagulation factor XI deficiency caused by intronic mutation IVS J-4delgttg. Zhonghua Xue Ye Xue Za Zhi 26:144, Bozzao C, Rimoldi V, Asselta R, et al: A novel factor XI missense mutation (Val371Ile) in the activation loop is responsible for a case of mild type II factor XI deficiency. FEBS J 274:6128, Karimi M, Jafari H, Lahsaeizadeh S, et al: Factor XI deficiency in Southern Iran: Identification of a novel missense mutation. Ann Hemato 88:359, 2009
17 58. Wistinghausen B, Reischer A, Oddoux C, et al: Severe factor XI deficiency in an Arab family associated with a novel mutation in exon 11. Br J Haematol 99:575, Sato T, Iyama S, Araki N, et al: Factor XI deficiency caused by a mutation of Gly400Val. Rinsho Ketsueki 48:148, Ishikawa N, Okada S, Sato T, et al: A novel mutation (Gln433Glu) in exon 12 combined with the G insertion in exon 13 causes severe factor XI deficiency in Japanese patients. Blood Coagul Fibrinolysis 18:519, Tsukahara A, Yamada T, Takagi A, et al: Compound heterozygosity for two novel mutations in a severe factor XI deficiency. Am J Hematol 73:279, McVey JH, Lal K, Imanaka Y, et al: Characterisation of blood coagulation factor XI T475I. Thromb Haemost 93:1082, Kwon MJ, Kim HJ, Bang SH, Kim SH: Severe factor XI deficiency in a Korean woman with a novel missense mutation (Val498Met) and duplication G mutation in exon 13 of the F11 gene. Blood Coagul Fibrinolysis 19:679, Iijima K, Udagawa A, Kawasaki H, et al: A factor XI deficiency associated with a nonsense mutation (Trp501stop) in the catalytic domain. Br J Haematol 111:556, de Moerloose P, Germanos-Haddad M, Boehlen F, Neerman-Arbez M: Severe factor XI deficiency in a Lebanese family: Identification of a novel missense mutation (Trp501Cys) in the catalytic domain. Blood Coagul Fibrinolysis 15:269, Germanos-Haddad M, de Moerloose P, Boehlen F, et al: Homozygosity for a Thr575Met missense mutation in the catalytic domain associated with factor XI deficiency. Haematologica 90:418, Gailani D, Schmidt A, Sun MF, Bolton-Maggs PH, Bajaj SP: A cross-reactive material positive variant of coagulation factor XI (FXI) with a catalytic defect. J Thromb Haemost, 5:781, Quelin F, De Raucourt E, Mathonnet F, et al: Characterization of combined factor VII and factor XI deficiencies. Haemophilia 14:639, Peretz H, Zivelin A, Usher S, Seligsohn U: A 14-bp deletion (Codon 554 del AAGgtaacagagtg) at exon 14/Intron N junction of the coagulation factor XI gene disrupts splicing and causes severe factor XI deficiency. Hum Mutat 8:77, Peretz H, Salomon O, Seligsohn U et al: Unpublished 71. Zivelin A, Ogawa T, Bulvik S, et al: Severe factor XI deficiency caused by a Gly 555 to Glu mutation (factor XI-Glu 555 ): A cross-reactiv emaqterial positive variant defective in factor IX activation. J Thromb Haemost 2:1782, Takamiya O, Machida S, Yamamoto M: Factor XI deficiency with a novel homozygous mutation Trp599Arg near the C-terminal region. Haematologica 90:999, 2005.
18 73. Mitchell M, Dai L, Savidge G, Alhaq A: An Alu-mediated 31.5-kb deletion as the cause of factor XI deficiency in 2 unrelated patients. Blood 104:2394, 2004.
Supplemental Table 1. Country of origin of the samples collected for the studies included in this systematic review. Authors PubMed ID.
 Supplemental Note In order to obtain all articles relevant to this study, key search terms were used in various combinations with the AND operator. Search terms included: pyrazinamide, resistance, Mycobacterium
Supplemental Note In order to obtain all articles relevant to this study, key search terms were used in various combinations with the AND operator. Search terms included: pyrazinamide, resistance, Mycobacterium
Genotypes of patients with factor XIII deficiency Gene Mutation Location Domain Type Genotype Origin Activit y U/dL F13A
 Genotypes of patients with factor XIII deficiency Gene Mutation Location Domain Type Genotype Origin Activit y U/dL F13A F13A F13A nt -7 del 20 bp and Ala394Val IVS4+2 ins T (Nagoya 1) IVS1-7 ins TT Arg326Gln*
Genotypes of patients with factor XIII deficiency Gene Mutation Location Domain Type Genotype Origin Activit y U/dL F13A F13A F13A nt -7 del 20 bp and Ala394Val IVS4+2 ins T (Nagoya 1) IVS1-7 ins TT Arg326Gln*
Nomenclature What is in a name? My name Joseˊ Jimenez = Bill Dana John L.C. Savony = Frank Fontaine
 Nomenclature What is in a name? My name Joseˊ Jimenez = Bill Dana John L.C. Savony = Frank Fontaine Three categories of Amyloid Terms 1. Amyloidologists, Amyloid Centers, Researchers official nomenclature
Nomenclature What is in a name? My name Joseˊ Jimenez = Bill Dana John L.C. Savony = Frank Fontaine Three categories of Amyloid Terms 1. Amyloidologists, Amyloid Centers, Researchers official nomenclature
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples
 A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
Citation for published version (APA): Verschueren, S. (2017). Systemic causes of heavy menstrual bleeding [Groningen]: Rijksuniversiteit Groningen
![Citation for published version (APA): Verschueren, S. (2017). Systemic causes of heavy menstrual bleeding [Groningen]: Rijksuniversiteit Groningen Citation for published version (APA): Verschueren, S. (2017). Systemic causes of heavy menstrual bleeding [Groningen]: Rijksuniversiteit Groningen](/thumbs/83/87655756.jpg) University of Groningen Systemic causes of heavy menstrual bleeding Verschueren, Sophie IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
University of Groningen Systemic causes of heavy menstrual bleeding Verschueren, Sophie IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
Genotypes of patients with factor X deficiency
 Mutation Met-0Val (Nice I) Pro0Ser (Nice II) nt del GC Phe1Ser* Val-Phe* Undefined Val-Phe* Gln16stop Leu-2Pro Ser-0Arg (Shanghai) Ala-26Asp* Gly-Arg* (Santo Domingo) Gly-Arg* Gly-Arg* Gly-Arg* (Santo
Mutation Met-0Val (Nice I) Pro0Ser (Nice II) nt del GC Phe1Ser* Val-Phe* Undefined Val-Phe* Gln16stop Leu-2Pro Ser-0Arg (Shanghai) Ala-26Asp* Gly-Arg* (Santo Domingo) Gly-Arg* Gly-Arg* Gly-Arg* (Santo
SUPPLEMENTARY INFORMATION. Rare independent mutations in renal salt handling genes contribute to blood pressure variation
 SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
modified dye uptake assay including formazan test EC 90 not tested plaque reduction assay
 Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Deliverable 2.1 List of relevant genetic variants for pre-emptive PGx testing
 GA N 668353 H2020 Research and Innovation Deliverable 2.1 List of relevant genetic variants for pre-emptive PGx testing WP N and Title: WP2 - Towards shared European Guidelines for PGx Lead beneficiary:
GA N 668353 H2020 Research and Innovation Deliverable 2.1 List of relevant genetic variants for pre-emptive PGx testing WP N and Title: WP2 - Towards shared European Guidelines for PGx Lead beneficiary:
Fabry Disease X-linked genetic, multi-organ disorder. Fabry disease screening program in Hypertrophic p Cardiomyopathy: preliminary results.
 Fabry Disease X-linked genetic, multi-organ disorder Fabry disease screening program in Hypertrophic p Cardiomyopathy: y preliminary results. Globotriaosylceramide, GL3 Brain -galactosidase A Eyes Lactosylceramide
Fabry Disease X-linked genetic, multi-organ disorder Fabry disease screening program in Hypertrophic p Cardiomyopathy: y preliminary results. Globotriaosylceramide, GL3 Brain -galactosidase A Eyes Lactosylceramide
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E,
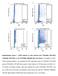 Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
The role of E148Q in FMF. Elon Pras Institute of Human Genetics Sheba Medical Center
 The role of E148Q in FMF Elon Pras Institute of Human Genetics Sheba Medical Center Familial Mediterranean Fever (FMF) Acute attacks of fever accompanied by: Peritonitis Pleuritis Arthritis Erysipelas
The role of E148Q in FMF Elon Pras Institute of Human Genetics Sheba Medical Center Familial Mediterranean Fever (FMF) Acute attacks of fever accompanied by: Peritonitis Pleuritis Arthritis Erysipelas
Epidemiology of Mutations for Cystic Fibrosis
 Appendixes Appendix A Epidemiology of Mutations for Cystic Fibrosis The differential distribution of mutations causing cystic fibrosis (CF) has clear implications for carrier screening. Besides DF508,
Appendixes Appendix A Epidemiology of Mutations for Cystic Fibrosis The differential distribution of mutations causing cystic fibrosis (CF) has clear implications for carrier screening. Besides DF508,
Neuropathy. Nerves before and after TTR. Márcia Waddington Cruz. CEPARM HUCFF-UFRJ.
 Neuropathy. Nerves before and after TTR. Márcia Waddington Cruz. CEPARM HUCFF-UFRJ. Amyloidosis Amyloid deposit. Precursor proteins. Fibrilar ptn. Lesion to tissues. Mechanism? 2 A TTR. SSA or Wild Type
Neuropathy. Nerves before and after TTR. Márcia Waddington Cruz. CEPARM HUCFF-UFRJ. Amyloidosis Amyloid deposit. Precursor proteins. Fibrilar ptn. Lesion to tissues. Mechanism? 2 A TTR. SSA or Wild Type
Exon I M1V 1A>G Kolker et al, delG Spector & Sharer, unpublished /90 delc Mushimoto et al, 2011
 Exon I M1V 1A>G Kolker et al, 2007 --- 11delG Spector & Sharer, unpublished 2006 --- 89/90 delc Mushimoto et al, 2011 Intron 1 IVS1+5G>T --- Greenberg et al, 1995 Exon II --- 109-110delCA Tsai et al, 2017
Exon I M1V 1A>G Kolker et al, 2007 --- 11delG Spector & Sharer, unpublished 2006 --- 89/90 delc Mushimoto et al, 2011 Intron 1 IVS1+5G>T --- Greenberg et al, 1995 Exon II --- 109-110delCA Tsai et al, 2017
Endocrine Surgery. Characteristics of the Germline MEN1 Mutations in Korea: A Literature Review ORIGINAL ARTICLE. The Korean Journal of INTRODUCTION
 ORIGINAL ARTICLE ISSN 1598-1703 (Print) ISSN 2287-6782 (Online) Korean J Endocrine Surg 2014;14:7-11 The Korean Journal of Endocrine Surgery Characteristics of the Germline MEN1 Mutations in Korea: A Literature
ORIGINAL ARTICLE ISSN 1598-1703 (Print) ISSN 2287-6782 (Online) Korean J Endocrine Surg 2014;14:7-11 The Korean Journal of Endocrine Surgery Characteristics of the Germline MEN1 Mutations in Korea: A Literature
Transgenic Mice and Genetargeting
 Transgenic Mice and Genetargeting mice In Biomedical Science Techniques of transgenic and gene-targeting mice are indispensable for analyses of in vivo functions of particular genes and roles of their
Transgenic Mice and Genetargeting mice In Biomedical Science Techniques of transgenic and gene-targeting mice are indispensable for analyses of in vivo functions of particular genes and roles of their
Supplementary Figure 1: Classification scheme for non-synonymous and nonsense germline MC1R variants. The common variants with previously established
 Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Structure and function of factor XI
 Review article Structure and function of factor XI Jonas Emsley, 1 Paul A. McEwan, 1 and David Gailani 2 1 School of Pharmacy, Centre for Biomolecular Sciences, University of Nottingham, Nottingham, United
Review article Structure and function of factor XI Jonas Emsley, 1 Paul A. McEwan, 1 and David Gailani 2 1 School of Pharmacy, Centre for Biomolecular Sciences, University of Nottingham, Nottingham, United
Dr Rosline Hassan Haematology Department, School of Medical Sciences, Universiti Sains Malaysia, Kelantan
 Dr Rosline Hassan Haematology Department, School of Medical Sciences, Universiti Sains Malaysia, Kelantan THE FIRST ASEAN FEDERATION OF HAEMATOLOGY AND THE VIIITH MALAYSIAN NATIONAL HAEMATOLOGY SCIENTIFIC
Dr Rosline Hassan Haematology Department, School of Medical Sciences, Universiti Sains Malaysia, Kelantan THE FIRST ASEAN FEDERATION OF HAEMATOLOGY AND THE VIIITH MALAYSIAN NATIONAL HAEMATOLOGY SCIENTIFIC
LE IPERCHILOMICRONEMIE FAMILIARI
 LE IPERCHILOMICRONEMIE FAMILIARI Livia Pisciotta Di.M.I.-Università di Genova HYPERCHYLOMICRONEMIA SYNDROME Ø Hypertriglyceridemia (>10 mmol/l, >877 mg/dl) Ø Fasting chylomicronemia Ø Recurrent abdominal
LE IPERCHILOMICRONEMIE FAMILIARI Livia Pisciotta Di.M.I.-Università di Genova HYPERCHYLOMICRONEMIA SYNDROME Ø Hypertriglyceridemia (>10 mmol/l, >877 mg/dl) Ø Fasting chylomicronemia Ø Recurrent abdominal
Intron 1 IVS1+5G>T --- Greenberg et al, Exon II del2 Tsai et al, 2017 K42R 125A>G Spector et al, unpublished
 Exon I M1V 1A>G Kolker et al, 2007 --- 11delG Spector & Sharer, unpublished S25L 74C>T Spector et al, unpublished --- 89/90 delc Mushimoto et al, 2011 88-91del4+IVS1 gtca Spector et al, unpublished Intron
Exon I M1V 1A>G Kolker et al, 2007 --- 11delG Spector & Sharer, unpublished S25L 74C>T Spector et al, unpublished --- 89/90 delc Mushimoto et al, 2011 88-91del4+IVS1 gtca Spector et al, unpublished Intron
LAL: significato clinico e biologico delle mutazioni di Bcr-Abl
 LAL: significato clinico e biologico delle mutazioni di Bcr-Abl Simona Soverini Dipartimento di Ematologia e Scienze Oncologiche L. e A. Seràgnoli Università di Bologna The vast majority of Ph+ ALL patients
LAL: significato clinico e biologico delle mutazioni di Bcr-Abl Simona Soverini Dipartimento di Ematologia e Scienze Oncologiche L. e A. Seràgnoli Università di Bologna The vast majority of Ph+ ALL patients
Beta Thalassemia Case Study Introduction to Bioinformatics
 Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Molecular genetics of maple syrup urine disease in the Turkish population
 The Turkish Journal of Pediatrics 2009; 51: 97-102 Original Molecular genetics of maple syrup urine disease in the Turkish population Kerstin Gorzelany 1, Ali Dursun 2, Turgay Coşkun 2, Serap H. Kalkanoğlu-Sivri
The Turkish Journal of Pediatrics 2009; 51: 97-102 Original Molecular genetics of maple syrup urine disease in the Turkish population Kerstin Gorzelany 1, Ali Dursun 2, Turgay Coşkun 2, Serap H. Kalkanoğlu-Sivri
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication
 Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
The many faces of MDR 3 deficiency
 The many faces of MDR 3 deficiency Relevance of canalicular membrane transporting proteins for human liver disease Peter LM Jansen and Marleen de Vree Academic Medical Center Amsterdam, The Netherlands
The many faces of MDR 3 deficiency Relevance of canalicular membrane transporting proteins for human liver disease Peter LM Jansen and Marleen de Vree Academic Medical Center Amsterdam, The Netherlands
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP
 Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
LIPOPROTEINE ATEROGENE E ANTI-ATEROGENE ATEROGENE
 LIPOPROTEINE ATEROGENE E ANTI-ATEROGENE ATEROGENE Sebastiano Calandra Dipartimento di Scienze Biomediche Università di Modena e Reggio Emilia Incidence Rate/1000 200-150 - 100-50 - Women 0 Men
LIPOPROTEINE ATEROGENE E ANTI-ATEROGENE ATEROGENE Sebastiano Calandra Dipartimento di Scienze Biomediche Università di Modena e Reggio Emilia Incidence Rate/1000 200-150 - 100-50 - Women 0 Men
Genetic Tests for the Better Outcome of VTE? 서울대학교병원혈액종양내과윤성수
 Genetic Tests for the Better Outcome of VTE? 서울대학교병원혈액종양내과윤성수 Thrombophilia A hereditary or acquired disorder predisposing to thrombosis Questions Why should we test? Who should we test For what disorders?
Genetic Tests for the Better Outcome of VTE? 서울대학교병원혈액종양내과윤성수 Thrombophilia A hereditary or acquired disorder predisposing to thrombosis Questions Why should we test? Who should we test For what disorders?
EU Market Situation for Poultry. Committee for the Common Organisation of the Agricultural Markets 20 April 2017
 EU Market Situation for Poultry Committee for the Common Organisation of the Agricultural Markets 2 April 217 -9.% -11.% -5.% -.1% -.7% -2.2% 3.% 1.7% 1.2%.8%.6%.%.%.% 7.9% 7.% 6.4% 6.2% 6.% 5.5% 5.3%
EU Market Situation for Poultry Committee for the Common Organisation of the Agricultural Markets 2 April 217 -9.% -11.% -5.% -.1% -.7% -2.2% 3.% 1.7% 1.2%.8%.6%.%.%.% 7.9% 7.% 6.4% 6.2% 6.% 5.5% 5.3%
Complement mediated renal disease. Tim Goodship Institute of Genetic Medicine Newcastle University
 Complement mediated renal disease Tim Goodship Institute of Genetic Medicine Newcastle University Evidence of complement activation is IgA and HSP Membranous Lupus nephritis ANCA associated Anti-GBM MPGN,
Complement mediated renal disease Tim Goodship Institute of Genetic Medicine Newcastle University Evidence of complement activation is IgA and HSP Membranous Lupus nephritis ANCA associated Anti-GBM MPGN,
Current genotyping in the Czech Republic
 The European Perspective: Current genotyping in the Czech Republic MUDr. Martin Písačka Reference laboratory for immunohematology ÚHKT Prag 37.Jahreskongress DGTI, 24.September 2004, Mannheim Pre-genotyping
The European Perspective: Current genotyping in the Czech Republic MUDr. Martin Písačka Reference laboratory for immunohematology ÚHKT Prag 37.Jahreskongress DGTI, 24.September 2004, Mannheim Pre-genotyping
A Quick Guide to the. I507del. Mutation CFTR SCIENCE
 A Quick Guide to the I507del Mutation CFTR SCIENCE 2016 Vertex Pharmaceuticals Incorporated VXR-HQ-02-00045a(1) 03/2016 Loss of CFTR activity is the underlying cause of cystic fibrosis (CF) 1 Spectrum
A Quick Guide to the I507del Mutation CFTR SCIENCE 2016 Vertex Pharmaceuticals Incorporated VXR-HQ-02-00045a(1) 03/2016 Loss of CFTR activity is the underlying cause of cystic fibrosis (CF) 1 Spectrum
TTR-FAP: Diagnosis and treatment Zürich June 19,2014. Ole B Suhr Umeå University and University Hospital Department of Medicine Umeå Sweden
 TTR-FAP: Diagnosis and treatment Zürich June 19,2014 Ole B Suhr Umeå University and University Hospital Department of Medicine Umeå Sweden Diagnosis of ATTR amyloidosis Clinical symptoms of ATTR- amyloidosis
TTR-FAP: Diagnosis and treatment Zürich June 19,2014 Ole B Suhr Umeå University and University Hospital Department of Medicine Umeå Sweden Diagnosis of ATTR amyloidosis Clinical symptoms of ATTR- amyloidosis
Prevalence and clinical implications of BRCA1/2 germline mutations in Chinese women with breast cancer Yuntao Xie M.D., Ph.D.
 Prevalence and clinical implications of BRCA1/2 germline mutations in Chinese women with breast cancer Yuntao Xie M.D., Ph.D. Hereditary Cancer Center, Peking University Cancer Hospital 1 Breast cancer
Prevalence and clinical implications of BRCA1/2 germline mutations in Chinese women with breast cancer Yuntao Xie M.D., Ph.D. Hereditary Cancer Center, Peking University Cancer Hospital 1 Breast cancer
Venous Thrombosis in Asia
 Venous Thrombosis in Asia Pantep Angchaisuksiri, M.D. Professor of Medicine, Mahidol University, Thailand Adjunct Associate Professor, University of North Carolina, Chapel Hill, USA Venous Thromboembolism
Venous Thrombosis in Asia Pantep Angchaisuksiri, M.D. Professor of Medicine, Mahidol University, Thailand Adjunct Associate Professor, University of North Carolina, Chapel Hill, USA Venous Thromboembolism
Deep-Sequencing of HIV-1
 Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Clinical and Molecular Genetic Spectrum of Slovenian Patients with CGD
 Clinical and Molecular Genetic Spectrum of Slovenian Patients with CGD Avčin T, Debeljak M, Markelj G, Anderluh G*, Glavnik V, Kuhar M University Children s Hospital Ljubljana and *Biotechnical Faculty,
Clinical and Molecular Genetic Spectrum of Slovenian Patients with CGD Avčin T, Debeljak M, Markelj G, Anderluh G*, Glavnik V, Kuhar M University Children s Hospital Ljubljana and *Biotechnical Faculty,
CANCER GENETICS PROVIDER SURVEY
 Dear Participant, Previously you agreed to participate in an evaluation of an education program we developed for primary care providers on the topic of cancer genetics. This is an IRB-approved, CDCfunded
Dear Participant, Previously you agreed to participate in an evaluation of an education program we developed for primary care providers on the topic of cancer genetics. This is an IRB-approved, CDCfunded
Dr Simon McRae Department of Haematology SA Pathology
 Dr Simon McRae Department of Haematology SA Pathology Normally defined as factor VIII levels >5-40 IU/dL Proportion of patients with mild HA varies between centres - 32% patients with mild HA in a large
Dr Simon McRae Department of Haematology SA Pathology Normally defined as factor VIII levels >5-40 IU/dL Proportion of patients with mild HA varies between centres - 32% patients with mild HA in a large
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL SUPPLEMENTAL MATERIAL Supplemental Methods Phenotype Prediction Analyses In order to assess the phylogenetic properties of nssnvs, sequence conservation analysis was conducted using
SUPPLEMENTAL MATERIAL SUPPLEMENTAL MATERIAL Supplemental Methods Phenotype Prediction Analyses In order to assess the phylogenetic properties of nssnvs, sequence conservation analysis was conducted using
Ethnic differences in GRHPR mutations in patients with primary hyperoxaluria type 2
 Clin Genet 2014: 86: 342 348 Printed in Singapore. All rights reserved Short Report 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12292 Ethnic differences
Clin Genet 2014: 86: 342 348 Printed in Singapore. All rights reserved Short Report 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12292 Ethnic differences
A Quick Guide to the G A. Mutation CFTR SCIENCE
 A Quick Guide to the 1717-1G A Mutation FR SIENE 2016 Vertex Pharmaceuticals Incorporated VXR-HQ-02-00045a(1) 03/2016 Loss of FR activity is the underlying cause of cystic fibrosis (F) 1 Spectrum of Phenotypes
A Quick Guide to the 1717-1G A Mutation FR SIENE 2016 Vertex Pharmaceuticals Incorporated VXR-HQ-02-00045a(1) 03/2016 Loss of FR activity is the underlying cause of cystic fibrosis (F) 1 Spectrum of Phenotypes
Von Willebrand Disease. Alison Street Malaysia April 2010
 Von Willebrand Disease Alison Street Malaysia April 2010 Physiology of VWF OUTLINE Clinical presentation of VWD Classification of VWD with emphases on Type 1, 2B and 2N disease Testing for VWD Treatment
Von Willebrand Disease Alison Street Malaysia April 2010 Physiology of VWF OUTLINE Clinical presentation of VWD Classification of VWD with emphases on Type 1, 2B and 2N disease Testing for VWD Treatment
Clinical Spectrum and Genetic Mechanism of GLUT1-DS. Yasushi ITO (Tokyo Women s Medical University, Japan)
 Clinical Spectrum and Genetic Mechanism of GLUT1-DS Yasushi ITO (Tokyo Women s Medical University, Japan) Glucose transporter type 1 (GLUT1) deficiency syndrome Mutation in the SLC2A1 / GLUT1 gene Deficiency
Clinical Spectrum and Genetic Mechanism of GLUT1-DS Yasushi ITO (Tokyo Women s Medical University, Japan) Glucose transporter type 1 (GLUT1) deficiency syndrome Mutation in the SLC2A1 / GLUT1 gene Deficiency
THE ROLE OF CFTR MUTATIONS IN CAUSING CYSTIC FIBROSIS (CF)
 THE ROLE OF CFTR MUTATIONS IN CAUSING CYSTIC FIBROSIS (CF) Vertex Pharmaceuticals Incorporated, 50 Northern Avenue, Boston, MA 02210. Vertex and the Vertex triangle logo are registered trademarks for Vertex
THE ROLE OF CFTR MUTATIONS IN CAUSING CYSTIC FIBROSIS (CF) Vertex Pharmaceuticals Incorporated, 50 Northern Avenue, Boston, MA 02210. Vertex and the Vertex triangle logo are registered trademarks for Vertex
Iranian Journal of Basic Medical Sciences
 Iranian Journal of Basic Medical Sciences ijbms.mums.ac.ir Nonsense-mediated mrna decay among coagulation factor genes Shirin Shahbazi 1 * 1 Department of Medical Genetics, Faculty of Medical Sciences,
Iranian Journal of Basic Medical Sciences ijbms.mums.ac.ir Nonsense-mediated mrna decay among coagulation factor genes Shirin Shahbazi 1 * 1 Department of Medical Genetics, Faculty of Medical Sciences,
GENETIC VARIATIONS AND POLYMORPHISMS OF GPROTEIN-COUPLED RECEPTORS: Functional and Therapeutic Implications
 Annu. Rev. Pharmacol. Toxicol. 2001. 41:593 624 Copyright c 2001 by Annual Reviews. All rights reserved GENETIC VARIATIONS AND POLYMORPHISMS OF GPROTEIN-COUPLED RECEPTORS: Functional and Therapeutic Implications
Annu. Rev. Pharmacol. Toxicol. 2001. 41:593 624 Copyright c 2001 by Annual Reviews. All rights reserved GENETIC VARIATIONS AND POLYMORPHISMS OF GPROTEIN-COUPLED RECEPTORS: Functional and Therapeutic Implications
Mutation analysis of CHCHD10 in neurodegenerative diseases, including Parkinson s disease Ekaterina Rogaeva
 Mutation analysis of CHCHD10 in neurodegenerative diseases, including Parkinson s disease Ekaterina Rogaeva Tanz Centre for Research in Neurodegenerative Diseases, University of Toronto Faculty of Medicine,
Mutation analysis of CHCHD10 in neurodegenerative diseases, including Parkinson s disease Ekaterina Rogaeva Tanz Centre for Research in Neurodegenerative Diseases, University of Toronto Faculty of Medicine,
Analysis of myocilin mutations in 1703 glaucoma patients from five different populations
 1999 Oxford University Press Human Molecular Genetics, 1999, Vol. 8, No. 5 899 905 Analysis of myocilin mutations in 1703 glaucoma patients from five different populations John H. Fingert 1, Elise Héon
1999 Oxford University Press Human Molecular Genetics, 1999, Vol. 8, No. 5 899 905 Analysis of myocilin mutations in 1703 glaucoma patients from five different populations John H. Fingert 1, Elise Héon
CPTR title slide. A Standardized System for Grading Mutations in Mycobacterium tuberculosis for Association with Drug Resistance
 CPTR title slide A Standardized System for Grading Mutations in Mycobacterium tuberculosis for Association with Drug Resistance PAOLO MIOTTO CPTR 2017 Workshop, March 20 23 The Need A lack of user-friendly
CPTR title slide A Standardized System for Grading Mutations in Mycobacterium tuberculosis for Association with Drug Resistance PAOLO MIOTTO CPTR 2017 Workshop, March 20 23 The Need A lack of user-friendly
Susanne Schnittger. Workflow of molecular investigations in JAK2-negative MPNs - the Munich experience
 Susanne Schnittger Workflow of molecular investigations in JAK2negative MPNs the Munich experience Cohort single centre experience to apply new markers in a daily diagnostic work flow total: 20,547 cases
Susanne Schnittger Workflow of molecular investigations in JAK2negative MPNs the Munich experience Cohort single centre experience to apply new markers in a daily diagnostic work flow total: 20,547 cases
Kalydeco. Kalydeco (ivacaftor) Description
 Federal Employee Program 1310 G Street, N.W. Washington, D.C. 20005 202.942.1000 Fax 202.942.1125 5.45.03 Subject: Kalydeco Page: 1 of 6 Last Review Date: November 30, 2018 Kalydeco Description Kalydeco
Federal Employee Program 1310 G Street, N.W. Washington, D.C. 20005 202.942.1000 Fax 202.942.1125 5.45.03 Subject: Kalydeco Page: 1 of 6 Last Review Date: November 30, 2018 Kalydeco Description Kalydeco
Dr. Ayman Mohsen Mashi, MBBS Consultant Hematology & Blood Transfusion Department Head, Laboratory & Blood Bank King Fahad Central Hospital, Gazan,
 Dr. Ayman Mohsen Mashi, MBBS Consultant Hematology & Blood Transfusion Department Head, Laboratory & Blood Bank King Fahad Central Hospital, Gazan, KSA amashi@moh.gov.sa 24/02/2018 β-thalassemia syndromes
Dr. Ayman Mohsen Mashi, MBBS Consultant Hematology & Blood Transfusion Department Head, Laboratory & Blood Bank King Fahad Central Hospital, Gazan, KSA amashi@moh.gov.sa 24/02/2018 β-thalassemia syndromes
Medium-Chain Acyl-CoA Dehydrogenase (MCAD) splicing mutations identified in newborns with an abnormal MS/MS profile
 EURASNET Workshop on RNA splicing and genetic diagnosis, London, UK Medium-Chain Acyl-CoA Dehydrogenase (MCAD) splicing mutations identified in newborns with an abnormal MS/MS profile Brage Storstein Andresen
EURASNET Workshop on RNA splicing and genetic diagnosis, London, UK Medium-Chain Acyl-CoA Dehydrogenase (MCAD) splicing mutations identified in newborns with an abnormal MS/MS profile Brage Storstein Andresen
Identification of a novel duplication mutation in the VHL gene in a large Chinese family with Von Hippel-Lindau (VHL) syndrome
 Identification of a novel duplication mutation in the VHL gene in a large Chinese family with Von Hippel-Lindau (VHL) syndrome L.H. Cao 1, B.H. Kuang 2, C. Chen 1, C. Hu 2, Z. Sun 1, H. Chen 2, S.S. Wang
Identification of a novel duplication mutation in the VHL gene in a large Chinese family with Von Hippel-Lindau (VHL) syndrome L.H. Cao 1, B.H. Kuang 2, C. Chen 1, C. Hu 2, Z. Sun 1, H. Chen 2, S.S. Wang
HCV NS3 Protease Drug Resistance
 test code: cpt code: 10000 87902 category: Infectious Disease HCV genotype 1a CODON Glecaprevir 1a Grazoprevir 1a Paritaprevir 1a Simeprevir 1a Voxilaprevir 1a V36A S 13 R 10 Comment V36C V36G V36I V36L
test code: cpt code: 10000 87902 category: Infectious Disease HCV genotype 1a CODON Glecaprevir 1a Grazoprevir 1a Paritaprevir 1a Simeprevir 1a Voxilaprevir 1a V36A S 13 R 10 Comment V36C V36G V36I V36L
IVACAFTOR THE ISRAELI EXPERIENCE ADI DAGAN MD THE ISRAELI CF CENTER SHEBA MEDICAL CENTER, TEL-HASHOMER
 IVACAFTOR THE ISRAELI EXPERIENCE ADI DAGAN MD THE ISRAELI CF CENTER SHEBA MEDICAL CENTER, TEL-HASHOMER February 21, 2014 U.S. Food and Drug Administration Approves KALYDECO (ivacaftor) for Use in Eight
IVACAFTOR THE ISRAELI EXPERIENCE ADI DAGAN MD THE ISRAELI CF CENTER SHEBA MEDICAL CENTER, TEL-HASHOMER February 21, 2014 U.S. Food and Drug Administration Approves KALYDECO (ivacaftor) for Use in Eight
Tobacco: World Markets and Trade
 United States Department of Agriculture Foreign Agricultural Service Circular Series FT -09-05 Sep. 2005 List of Tables Tobacco: World Markets and Trade Table 2 U.S. Tobacco Trade: 2004-2005 Table 3 Unmanufactured
United States Department of Agriculture Foreign Agricultural Service Circular Series FT -09-05 Sep. 2005 List of Tables Tobacco: World Markets and Trade Table 2 U.S. Tobacco Trade: 2004-2005 Table 3 Unmanufactured
Personalized Healthcare Update
 Dr. Kai - Oliver Wesche Market Development Manager, Personalized Healthcare QIAGEN Personalized Healthcare Update Pioneering Personalized Medicine through Partnering TOMTOVOK BKM120 Zelboraf QIAGEN partners:
Dr. Kai - Oliver Wesche Market Development Manager, Personalized Healthcare QIAGEN Personalized Healthcare Update Pioneering Personalized Medicine through Partnering TOMTOVOK BKM120 Zelboraf QIAGEN partners:
Mutation analysis of BRCA1 and BRCA2 from 793 Korean patients with sporadic breast cancer
 Clin Genet 2006: 70: 496 501 Printed in Singapore. All rights reserved Short Report # 2006 The Authors Journal compilation # 2006 Blackwell Munksgaard CLINICAL GENETICS doi: 10.1111/j.1399-0004.2006.00717.x
Clin Genet 2006: 70: 496 501 Printed in Singapore. All rights reserved Short Report # 2006 The Authors Journal compilation # 2006 Blackwell Munksgaard CLINICAL GENETICS doi: 10.1111/j.1399-0004.2006.00717.x
Spherical Bearings Heavy Duty Equipments
 Spherical Bearings Heavy Duty Equipments Highlights Quality Service Price Wbf Replacement Parts adaptableto > Caterpillar >Komatsu >Volvo 1 WBF SPHERICAL BEARINGS adaptable to Caterpillar Part No. Description
Spherical Bearings Heavy Duty Equipments Highlights Quality Service Price Wbf Replacement Parts adaptableto > Caterpillar >Komatsu >Volvo 1 WBF SPHERICAL BEARINGS adaptable to Caterpillar Part No. Description
Supplementary Figure 1 Weight and body temperature of ferrets inoculated with
 Supplementary Figure 1 Weight and body temperature of ferrets inoculated with A/Anhui/1/2013 (H7N9) influenza virus. (a) Body temperature and (b) weight change of ferrets after intranasal inoculation with
Supplementary Figure 1 Weight and body temperature of ferrets inoculated with A/Anhui/1/2013 (H7N9) influenza virus. (a) Body temperature and (b) weight change of ferrets after intranasal inoculation with
Von Willebrand Disease: Mutations, Von Willebrand Factor Variance and Genetic Drift
 Von Willebrand Disease: Mutations, Von Willebrand Factor Variance and Genetic Drift Student: Cecilia Carlsson Pecharromán Supervisor: Torbjörn Säll Institution: Department of Biology, Lund University E-mail:
Von Willebrand Disease: Mutations, Von Willebrand Factor Variance and Genetic Drift Student: Cecilia Carlsson Pecharromán Supervisor: Torbjörn Säll Institution: Department of Biology, Lund University E-mail:
POTENTIAL LINK BETWEEN GAUCHER DISEASE PATHWAYS
 POTENTIAL LINK BETWEEN GAUCHER DISEASE PATHWAYS AND THOSE OF PARKINSON DISEASE Hanna Rosenbaum, MD Hematology and Bone Marrow Transplantation Rambam Medical Center and Bruce Rappaport Faculty of Medicine
POTENTIAL LINK BETWEEN GAUCHER DISEASE PATHWAYS AND THOSE OF PARKINSON DISEASE Hanna Rosenbaum, MD Hematology and Bone Marrow Transplantation Rambam Medical Center and Bruce Rappaport Faculty of Medicine
Frequencies and geographic distributions of genetic mutations in transthyretinand non-transthyretin-related familial amyloidosis
 Clin Genet 2015: 88: 396 400 Printed in Singapore. All rights reserved Short Report Frequencies and geographic distributions of genetic mutations in transthyretinand non-transthyretin-related familial
Clin Genet 2015: 88: 396 400 Printed in Singapore. All rights reserved Short Report Frequencies and geographic distributions of genetic mutations in transthyretinand non-transthyretin-related familial
PRIMARY OPEN-ANGLE GLAUCOMA IS ONE OF THE
 Age-dependent Prevalence of Mutations at the GLC1A Locus in Primary Open-angle Glaucoma SATOKO SHIMIZU, MD, PAUL R. LICHTER, MD, A. TIM JOHNSON, MD, PHD, ZHAOHUI ZHOU, PHD, MISAO HIGASHI, BS, MARIA GOTTFREDSDOTTIR,
Age-dependent Prevalence of Mutations at the GLC1A Locus in Primary Open-angle Glaucoma SATOKO SHIMIZU, MD, PAUL R. LICHTER, MD, A. TIM JOHNSON, MD, PHD, ZHAOHUI ZHOU, PHD, MISAO HIGASHI, BS, MARIA GOTTFREDSDOTTIR,
J. C.-H. Yang 1, L.V. Sequist 2, S. L. Geater 3, C.-M. Tsai 4, T. Mok 5, M. H. Schuler 6, N. Yamamoto 7, D. Massey 8, V. Zazulina 8, Yi-Long Wu 9
 Activity of afatinib in uncommon epidermal growth factor receptor (EGFR) mutations: Findings from three prospective trials of afatinib in EGFR mutation-positive lung cancer J. C.-H. Yang 1, L.V. Sequist
Activity of afatinib in uncommon epidermal growth factor receptor (EGFR) mutations: Findings from three prospective trials of afatinib in EGFR mutation-positive lung cancer J. C.-H. Yang 1, L.V. Sequist
Genetic epidemiology of MODY in the Czech republic: new mutations in the MODY genes HNF-4α, GCK and HNF-1α
 Diabetologia (2003) 46:291 295 DOI 10.1007/s00125-002-1010-7 Genetic epidemiology of MODY in the Czech republic: new mutations in the MODY genes HNF-4α, GCK and HNF-1α S. Pruhova 1, J. Ek 2, J. Lebl 1,
Diabetologia (2003) 46:291 295 DOI 10.1007/s00125-002-1010-7 Genetic epidemiology of MODY in the Czech republic: new mutations in the MODY genes HNF-4α, GCK and HNF-1α S. Pruhova 1, J. Ek 2, J. Lebl 1,
Vertical Magnetic Separation of Circulating Tumor Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients
 Vertical Magnetic Separation of Circulating Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients Chang Eun Yoo 1,2#, Jong-Myeon Park 3#, Hui-Sung Moon 1,2, Je-Gun Joung 2, Dae-Soon Son
Vertical Magnetic Separation of Circulating Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients Chang Eun Yoo 1,2#, Jong-Myeon Park 3#, Hui-Sung Moon 1,2, Je-Gun Joung 2, Dae-Soon Son
Genetic Diagnosis of Liver Diseases
 The Hong Kong Association for the Study of Liver Diseases 27 th Annual Scientific Meeting and International Symposium on Hepatology 16 November 2014 Genetic Diagnosis of Liver Diseases Dr Chloe Mak MD,
The Hong Kong Association for the Study of Liver Diseases 27 th Annual Scientific Meeting and International Symposium on Hepatology 16 November 2014 Genetic Diagnosis of Liver Diseases Dr Chloe Mak MD,
Original Article Characterization of phenylalanine hydroxylase gene mutations in phenylketonuria in Xinjiang of China
 Int J Clin Exp Med 2014;7(11):4406-4412 www.ijcem.com /ISSN:1940-5901/IJCEM0002101 Original Article Characterization of phenylalanine hydroxylase gene mutations in phenylketonuria in Xinjiang of China
Int J Clin Exp Med 2014;7(11):4406-4412 www.ijcem.com /ISSN:1940-5901/IJCEM0002101 Original Article Characterization of phenylalanine hydroxylase gene mutations in phenylketonuria in Xinjiang of China
Variant interpretation exercise. ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18
 Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Pharmacogenomics in Rare Diseases: Development Strategy for Ivacaftor as a Therapy for Cystic Fibrosis
 Pharmacogenomics in Rare Diseases: Development Strategy for Ivacaftor as a Therapy for Cystic Fibrosis Federico Goodsaid Vice President Strategic Regulatory Intelligence Vertex Pharmaceuticals Is there
Pharmacogenomics in Rare Diseases: Development Strategy for Ivacaftor as a Therapy for Cystic Fibrosis Federico Goodsaid Vice President Strategic Regulatory Intelligence Vertex Pharmaceuticals Is there
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/20497 holds various files of this Leiden University dissertation. Author: Siegerink, Bob Title: Prothrombotic factors and the risk of myocardial infarction
Cover Page The handle http://hdl.handle.net/1887/20497 holds various files of this Leiden University dissertation. Author: Siegerink, Bob Title: Prothrombotic factors and the risk of myocardial infarction
HCV NS3 Protease Drug Resistance
 test code: cpt code: 10000 87902 category: Infectious Disease HCV genotypes 1a CODON Simeprevir 1a Boceprevir 1a Telaprevir 1a Paritaprevir 1a Grazoprevir 1a V36A Comment R 3 R 10, 11 R 16 S 24 V36C V36G
test code: cpt code: 10000 87902 category: Infectious Disease HCV genotypes 1a CODON Simeprevir 1a Boceprevir 1a Telaprevir 1a Paritaprevir 1a Grazoprevir 1a V36A Comment R 3 R 10, 11 R 16 S 24 V36C V36G
Smoking, human papillomavirus infection, and p53 mutation as risk factors in oropharyngeal cancer: a case-control study
 RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Smoking, human papillomavirus infection, and p53 as risk factors in oropharyngeal cancer: a case-control study PKS Chan *, JSY Chor, AC Vlantis, TL
RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Smoking, human papillomavirus infection, and p53 as risk factors in oropharyngeal cancer: a case-control study PKS Chan *, JSY Chor, AC Vlantis, TL
Characterization of Hepatitis B Virus (HBV) Among Liver Patients in Kenya
 Characterization of Hepatitis B Virus (HBV) Among Liver Patients in Kenya By MISSIANI OCHWOTO (Medical Research officer, KEMRI) Julius Oyugi 2, Dufton Mwaengo 2, James Kimotho 1 Carla Osiowy 3 and Elijah
Characterization of Hepatitis B Virus (HBV) Among Liver Patients in Kenya By MISSIANI OCHWOTO (Medical Research officer, KEMRI) Julius Oyugi 2, Dufton Mwaengo 2, James Kimotho 1 Carla Osiowy 3 and Elijah
Global Measles and Rubella Update November 2018
 Global Measles and Rubella Update November 218 Measles Number of Reported Measles by WHO Regions 218 Region Member States* Suspected cases Measles cases Clin Epi Lab Jan Feb Mar Apr May Jun Jul Aug Sep
Global Measles and Rubella Update November 218 Measles Number of Reported Measles by WHO Regions 218 Region Member States* Suspected cases Measles cases Clin Epi Lab Jan Feb Mar Apr May Jun Jul Aug Sep
review July/August 2003 Vol. 5 No. 4
 review July/August 2003 Vol. 5 No. 4 Can mucopolysaccharidosis type I disease severity be predicted based on a patient s genotype? A comprehensive review of the literature Nancy J. Terlato, PharmD 1, and
review July/August 2003 Vol. 5 No. 4 Can mucopolysaccharidosis type I disease severity be predicted based on a patient s genotype? A comprehensive review of the literature Nancy J. Terlato, PharmD 1, and
SALSA MLPA KIT P060-B2 SMA
 SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
Determination of APTT factor sensitivity the misguiding guideline
 International Journal of Laboratory Hematology ORIGINAL ARTICLE The Official journal of the International Society for Laboratory Hematology INTERNATIONAL JOURNAL OF LABORATORY HEMATOLOGY Determination
International Journal of Laboratory Hematology ORIGINAL ARTICLE The Official journal of the International Society for Laboratory Hematology INTERNATIONAL JOURNAL OF LABORATORY HEMATOLOGY Determination
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13898 Supplementary Information Table 1 Kras mutation status of carcinogen-induced mouse lung adenomas Tumour Treatment Strain Grade Genotype Kras status (WES)* Kras status (Sanger) 32T1
doi:10.1038/nature13898 Supplementary Information Table 1 Kras mutation status of carcinogen-induced mouse lung adenomas Tumour Treatment Strain Grade Genotype Kras status (WES)* Kras status (Sanger) 32T1
Large observational study to UNderstand the Global impact of Severe Acute respiratory FailurE (LUNG-SAFE)
 Large observational study to UNderstand the Global impact of Severe Acute respiratory FailurE (LUNG-SAFE) John Laffey, Giacomo Bellani, Tai Pham, Eddy Fan, Antonio Pesenti on behalf of the LUNG SAFE Investigators
Large observational study to UNderstand the Global impact of Severe Acute respiratory FailurE (LUNG-SAFE) John Laffey, Giacomo Bellani, Tai Pham, Eddy Fan, Antonio Pesenti on behalf of the LUNG SAFE Investigators
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each
 Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
Country-wise and Item-wise Exports of Animal By Products Value Rs. Lakh Quantity in '000 Unit: Kgs Source: MoC Export Import Data Bank
 Country-wise and Item-wise Exports of Animal By Products Value Rs. Lakh Quantity in '000 Unit: Kgs Source: MoC Export Import Data Bank Country CAPEXIL Description HS Code Value 2010-2011 Quantity 2010-2011
Country-wise and Item-wise Exports of Animal By Products Value Rs. Lakh Quantity in '000 Unit: Kgs Source: MoC Export Import Data Bank Country CAPEXIL Description HS Code Value 2010-2011 Quantity 2010-2011
screening procedures Disease resistant to full-dose imatinib ( 600 mg/day) or intolerant to any dose of imatinib
 Table S1. Study inclusion and exclusion criteria Inclusion criteria Aged 18 years Signed and dated informed consent form prior to protocol-specific screening procedures Cytogenetic- or PCR-based diagnosis
Table S1. Study inclusion and exclusion criteria Inclusion criteria Aged 18 years Signed and dated informed consent form prior to protocol-specific screening procedures Cytogenetic- or PCR-based diagnosis
Concurrent Practical Session ACMG Classification
 Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
A Case of Factor XII Deficiency Which was Found in Recurrent Spontaneous Abortion. Y. S. Nam, I. H. Kim, T. K. Yoon, C. N. Lee and K. Y.
 12 1 A Case of Factor XII Deficiency Which was Found in Recurrent Spontaneous Abortion Y S Nam, I H Kim, T K Yoon, C N Lee and K Y Cha Department of Obstetrics and Gynecology, College of Medicine, Pocheon
12 1 A Case of Factor XII Deficiency Which was Found in Recurrent Spontaneous Abortion Y S Nam, I H Kim, T K Yoon, C N Lee and K Y Cha Department of Obstetrics and Gynecology, College of Medicine, Pocheon
Investigating the prospect of. develop optimal transfusion
 Investigating the prospect of applying RHD genotyping to develop optimal transfusion strategies for weak D patients in Ireland Paula Holton 1, Diarmaid O Donghaile 2, Sorcha Ni Loingsigh 2, Mark Lambert
Investigating the prospect of applying RHD genotyping to develop optimal transfusion strategies for weak D patients in Ireland Paula Holton 1, Diarmaid O Donghaile 2, Sorcha Ni Loingsigh 2, Mark Lambert
Supplementary Online Content
 Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
MRC-Holland MLPA. Description version 19;
 SALSA MLPA probemix P6-B2 SMA Lot B2-712, B2-312, B2-111, B2-511: As compared to the previous version B1 (lot B1-11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). SPINAL
SALSA MLPA probemix P6-B2 SMA Lot B2-712, B2-312, B2-111, B2-511: As compared to the previous version B1 (lot B1-11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). SPINAL
The molecular basis of phenylketonuria in Koreans
 J Hum Genet (2004) 49:617 621 DOI 10.1007/s10038-004-0197-5 ORIGINAL ARTICLE Dong Hwan Lee Æ Soo Kyung Koo Æ Kwang-Soo Lee Young-Joo Yeon Æ Hyun-Jeong Oh Æ Sang-Wun Kim Sook-Jin Lee Æ Sung-Soo Kim Æ Jong-Eun
J Hum Genet (2004) 49:617 621 DOI 10.1007/s10038-004-0197-5 ORIGINAL ARTICLE Dong Hwan Lee Æ Soo Kyung Koo Æ Kwang-Soo Lee Young-Joo Yeon Æ Hyun-Jeong Oh Æ Sang-Wun Kim Sook-Jin Lee Æ Sung-Soo Kim Æ Jong-Eun
Letter to the Editor. Dear Editors
 Atherosclerosis 44 (999) 443 447 Letter to the Editor Detection of a new compound heterozygote (del G 96 /G4A) for lipoprotein lipase deficiency and a comparative haplotype analysis of the mutant lipoprotein
Atherosclerosis 44 (999) 443 447 Letter to the Editor Detection of a new compound heterozygote (del G 96 /G4A) for lipoprotein lipase deficiency and a comparative haplotype analysis of the mutant lipoprotein
Hyperammonemia in Pediatric Clinics: A review of ornithine transcarbamylase deficiency (OTCD) based on our case studies
 JMA Medical Awards Hyperammonemia in Pediatric Clinics: A review of ornithine transcarbamylase deficiency (OTCD) based on our case studies JMAJ 47(4): 160 165, 2004 Ichiro MATSUDA Professor Emeritus, Kumamoto
JMA Medical Awards Hyperammonemia in Pediatric Clinics: A review of ornithine transcarbamylase deficiency (OTCD) based on our case studies JMAJ 47(4): 160 165, 2004 Ichiro MATSUDA Professor Emeritus, Kumamoto
E148Q as A Familial Mediterranean Fever-Causing Mutation: A Clinical-Based Study R A Jarjour ABSTRACT
 E148Q as A Familial Mediterranean Fever-Causing Mutation: A Clinical-Based Study R A Jarjour ABSTRACT Objective: to evaluate the clinical implications of E148Q mutation in familial Mediterranean fever
E148Q as A Familial Mediterranean Fever-Causing Mutation: A Clinical-Based Study R A Jarjour ABSTRACT Objective: to evaluate the clinical implications of E148Q mutation in familial Mediterranean fever
