SUPPLEMENTAL MATERIAL
|
|
|
- Charles Paul
- 5 years ago
- Views:
Transcription
1 SUPPLEMENTAL MATERIAL
2 SUPPLEMENTAL MATERIAL Supplemental Methods Phenotype Prediction Analyses In order to assess the phylogenetic properties of nssnvs, sequence conservation analysis was conducted using primary sequences from the University of California Santa Cruz Genome Browser ( 1 In order to calculate the degree of conservation across orthologs, the primary sequences from 44 species including primates, other placental mammals, monotremes, and non-mammalian vertebrates were aligned. Additionally, the 8 paralog sequences of the Na v 1 family were obtained from UniProt and aligned to the Na v 1.5 protein sequence using ClustalW. As sequences within this family have regions of low similarity, each individual region (i.e. N-terminal, DI-S1, DI-S1/2, DI-S2, DI-S2/3, DI-S3, DI-S3/4, DI-S4, DI- S4/5, DI-S5, DI-S5/6, DI-S6, DI/DII, etc.) were aligned individually and subsequently assembled into a continuous sequence relative to the Na v 1.5 protein sequence. The number of substitutions at each position (degree of nonidentity) was calculated by assessing the number of primary sequences at each position that harbored a residue that did not match the corresponding human residue. This number was separately for both orthologs and paralogs and genetic variants were then classified as occurring at a position with either no substitutions or > 1 substitution(s) in the orthologs or paralogs. In order to assess the physicochemical properties of nssnvs, Grantham chemical scores were calculated using the Grantham amino acid difference matrix as previously described. 2 Grantham values range from 15 (most conservative) to 215 (most radical), with values 150 considered radical, 100 to 149 considered moderately radical, 50 to 99 considered moderately
3 conservative, and < 50 considered conservative. For the purposes of this study, nssnvs were classified as radical (Grantham value 100) or conservative (Grantham value < 100). Mutation Assessor, version 2, provides predictions based on evolutionary conservation based on protein homologs and has been previously validated on disease associated (OMIM) and polymorphic variants. The assumptions and exact methodology employed by the current version of the SIFT algorithm have been described previously. 3 Mutation assessor classified each variant based on a calculated functional impact score into either neutral, low, medium, and high. In this study, neutral and low were combined as benign and medium and high were combined as damaging. SIFT, version 4.0.5, an additional conservation-based metric was used to analyze the Na v 1.5 protein sequence and provide phenotype predictions for each nssnv identified in cases and controls using the default settings. The assumptions and exact methodology employed by the current version of the SIFT algorithm have been described previously. 4 For the purposes of this study, nssnvs were classified as either Tolerated or Damaging based on the SIFT prediction. PolyPhen2, version 2.1.0, was used to analyze the effect of nssnvs on the secondary and tertiary protein structure of the Na v 1.5 channel using information derived from the Protein Databank (PDB) and Database of Secondary Structure Assignments (DSSP) using default settings. The assumptions and exact methodology of the PolyPhen2 algorithm have been described previously. 5 PolyPhen2 classified each variant as probably damaging, possibly damaging, or benign. For this study, those nssnvs labeled as probably damaging or possibly damaging were combined as damaging.
4 Condel, version 1.5, provides phenotypic classifications based on a weighted average of the normalized scores from SIFT, PolyPhen2, and Mutation Assessor. The assumptions and exact methodology of the Condel algorithm have been described previously. 6 Each variant was classified as either deleterious or neutral based on the Condel predictions. APPRAISE is a Bayesian logistic regression model that integrates multiple lines of evidence, including the domain containing the variant, variant class, SIFT/Polyphen/Grantham missense predictions, conservation and frequency in population databases, to evaluate the probability that a rare variant found in an individual with long QT syndrome is the cause of their disease. 7 For the purposes of this study, nssnvs were classified as pathogenic (posterior probability of pathogenicity 0.9) or conservative (posterior probability of pathogenicity value < 0.9). Calculation of Estimated Prediction Values In order to estimate the likelihood of disease causation, an estimated predictive value (EPV, defined as the probability of pathogenicity for a mutation identified in a case; EPV = (case frequency control frequency)/case frequency) was employed. 8 Briefly, these calculations rely on the simplifying assumptions that (1) the rate of background genetic variation is the same for the case and control populations and (2) all mutations found in controls are benign, background mutations, given the low prevalence of SCN5A-mediated BrS and LQTS. Applying these principles, we then calculated estimated predictive values (EPVs). The upper and lower bounds of the 95% confidence intervals (95% CI) were calculated for all EPVs using the formula: CI=1 1/(ê{ln (RR)±z*[SE(log RR)]}), where RR is the relative ratio (mutation frequency in cases divided by the mutation frequency in controls), z= for 1 α=95%, and SE[log(RR)] is the standard error around the log of RR. All EPVs calculated here
5 are specific to clinically definite cases as defined previously above and would be over-estimates if applied to a less definite case. Site-directed mutagenesis The SCN5A missense mutations were engineered by site-directed mutagenesis (QuikChange Site-Directed Mutagenesis Kit, Stratagene, La Jolla, CA) using PCR technique into two common alternatively spliced SCN5A transcripts, H558/Q1077del (Genbank accession no.ay148488) or R558/Q1077del and H558/Q1077 (Genbank accession no. AC ), of the human cardiac voltage-dependent Na+ channel α subunit SCN5A cdna (kindly provided by Dr. Jonathan C. Makielski, University of Wisconsin, Madison, WI) in the pcdna3 vector (Invitrogen, Carlsbad, CA). The integrity of the constructs was verified by DNA sequencing. Heterologous expression of mutant and wild-type sodium channels HEK-293 cells were cultured in minimum essential medium supplemented with 1% nonessential amino acid solution, 10% horse serum, 1% sodium pyruvate solution, and 1.4% penicillin/ streptomycin solution in a 5% CO 2 incubator at 37 C. 1 μg SCN5A wild type (WT) or mutant cdna were co-transfected with 1 μg or 0.25 μg Green Fluorescence Protein (GFP) cdna (kindly provided by Dr. Gianrico Farrugia, Mayo Clinic, Rochester, MN) with the use of 3μl Lipofectamine (Invitrogen, Carlsbad, CA). Transfected HEK-293 cells were cultured in OPTI- MEM (Gibco, Carlsbad, CA) and incubated for 24 hours. Cells exhibiting green fluorescence were selected for electrophysiological experiments. Electrophysiological measurements and data analysis Standard whole-cell patch clamp technique was used to measure SCN5A wild type and mutant sodium currents at room temperature (22-24 C) with the use of an Axopatch 200B amplifier, Digidata 1440A and pclamp 10.2 software (Axon Instruments, Sunnyvale, CA). The
6 extracellular (bath) solution contained (mmol/l): 140NaCl, 4 KCl, 1.8 CaCl 2, 0.75 MgCl 2, and 5 HEPES, ph adjusted to 7.4 with NaOH. The pipette solution contained (mmol/l): 120 CsF, 20 CsCl, 2 EGTA, and 5 HEPES, ph adjusted to 7.4 with CsOH. Microelectrodes were pulled on a P-97 puller (Sutter Instruments, Novato, CA) and fire polished to a final resistance of 2-3 MΏ. Series resistance was compensated by 80-85%. Currents were filtered at 5 khz and digitized at khz with an eight-pole Bessel filter. The voltage-dependence of activation, steady-state inactivation and late inward sodium current (INaL) were determined using voltage-clamp protocols described in the relevant figures (Figures 1-5) and figure legend. In all protocols, a holding potential of -100 mv or -120 mv and a start to start interval of 1-3 s were used. Data were analyzed using Clampfit (Axon Instruments, Sunnyvale, CA), Excel (Microsoft, Redmond, WA), and fitted with Origin 8 (OriginLab Corporation, Northampton, MA) software. The voltage-dependence of activation curve was fitted with a Boltzmann function: GNa/GNa, max= {1+exp [(V-V1/2)/k]}-1, where V1/2 and k are the half-maximal voltage of activation and the slope factor respectively, and GNa =INa / (V-Vrev), where Vrev is the reversal potential. The steady-state inactivation curve was fitted with a Boltzmann function: INa/INa, max= {1+exp [(V-V1/2)/k]}-1, where V1/2 and k are the half-maximal voltage of inactivation and the slope factor respectively. Late I Na was measured at the end of 700 ms long depolarization. Statistical analysis Results are expressed as mean ± SEM. Student t test was performed to determine statistical significance between two groups. A p<0.05 was considered to be significant. Supplemental References 1. Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D. The human genome browser at ucsc. Genome Res. 2002;12:
7 2. Grantham R. Amino acid difference formula to help explain protein evolution. Science. 1974;185: Reva B, Antipin Y, Sander C. Predicting the functional impact of protein mutations: Application to cancer genomics. Nucleic Acids Res. 2011;39:e Kumar P, Henikoff S, Ng PC. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc. 2009;4: Ramensky V, Bork P, Sunyaev S. Human non-synonymous snps: Server and survey. Nucleic Acids Res. 2002;30: Gonzalez-Perez A, Lopez-Bigas N. Improving the assessment of the outcome of nonsynonymous snvs with a consensus deleteriousness score, Condel. Am J Hum Genet. 2011;88: Ruklisa D, Ware JS, Walsh R, Balding DJ, Cook SA. Bayesian models for syndrome- and gene-specific probabilities of novel variant pathogenicity. Genome Med. 2015;7:5 8. Kapa S, Tester DJ, Salisbury BA, Harris-Kerr C, Pungliya MS, Alders M, Wilde AA, Ackerman MJ. Genetic testing for long-qt syndrome: Distinguishing pathogenic mutations from benign variants. Circulation. 2009;120:
8 Status Nucleotid e Change Variant Region BrS (2111) LQT (2888) Control (8975) Ortholog Paralog Grantham SIFT Condel MAss PP2 Composite case nssnv 3 G>A M1I N-term Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 9 C>G N3K N-term Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 44 G>C R15T N-term Benign Benign Benign Benign Benign Patho Patho 2 case nssnv 53 G>A R18Q N-term Benign Benign Benign Patho Patho Patho Patho 4 control nssnv 52 C>T R18W N-term Benign Benign Patho Patho Patho Patho Patho 4 control nssnv 59 C>T S20F N-term Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 80 G>A R27H N-term Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 89 A>G E30G N-term Benign Benign Benign Patho Patho Patho Patho 4 Polymorphism 100 C>T R34C N-term Benign Benign Patho Patho Patho Patho Patho 4 control nssnv 101 G>A R34H N-term Benign Benign Benign Patho Patho Patho Patho 4 control nssnv 103 G>A G35S N-term Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 128 G>A R43Q N-term Patho Benign Benign Benign Patho Benign Benign 2 case nssnv 142 G>A E48K N-term Benign Benign Benign Benign Patho Benign Patho 2 case nssnv 154 C>T P52S N-term Benign Patho Benign Patho Patho Patho Benign 4 case nssnv 158 G>A R53Q N-term Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 185 A>C K62T N-term Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 210 T>G N70K N-term Benign Benign Benign Patho Patho Benign Patho 3 control nssnv 209 A>G N70S N-term Benign Benign Benign Benign Benign Benign Patho 1 control nssnv 235 C>T P79S N-term Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 250 G>A D84N N-term Patho Patho Benign Patho Patho Patho Patho 6 Polymorphism 253 C>A P85T N-term Benign Patho Benign Patho Patho Patho Patho 5 case nssnv 278 T>C F93S N-term Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 281 T>G I94S N-term Benign Benign Patho Patho Patho Patho Patho 4 control nssnv 280 A>G I94V N-term Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 310 C>T R104W N-term Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 311 G>A R104Q N-term Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 310 C>G R104G N-term Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 327 C>A N109K N-term Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 343 A>G S115G N-term Benign Benign Benign Patho Patho Patho Patho 4 Polymorphism 355 C>T P119S N-term Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 361 C>T R121W N-term Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 362 G>A R121Q N-term Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 373 G>C V125L N-term Benign Benign Benign Patho Patho Benign Benign 2 case nssnv 376 A>G K126E N-term Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 407 T>C L136P DI-S Benign Benign Benign Patho Patho Patho Patho 4 control nssnv 414 G>A M138I DI-S Patho Benign Benign Benign Patho Patho Patho 4 case nssnv 436 G>A V146M DI-S Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 481 G>C E161Q DI-S Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 481 G>A E161K DI-S Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 496 G>A A166T DI-S Benign Benign Benign Patho Patho Benign Patho 3
9 control nssnv 508 T>A F170I DI-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 525 G>C K175N DI-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 533 C>G A178G DI-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 544 T>C C182R DI-S2/ Patho Benign Patho Patho Patho Patho Patho 5 case nssnv 554 C>T A185V DI-S2/ Benign Benign Benign Patho Benign Benign Patho 2 control nssnv 553 G>A A185T DI-S2/ Benign Benign Benign Benign Patho Benign Patho 2 control nssnv 601 A>G I201V DI-S Patho Patho Benign Patho Patho Benign Benign 4 case nssnv 611 C>T A204V DI-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 635 T>A L212Q DI-S3/ Patho Benign Patho Patho Patho Benign Patho 4 case nssnv 635 T>C L212P DI-S3/ Patho Benign Benign Patho Patho Patho Patho 5 Polymorphism 647 C>T S216L DI-S3/ Patho Benign Patho Patho Patho Patho Patho 5 Polymorphism 659 C>T T220I DI-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 665 G>A R222Q DI-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 667 G>C V223L DI-S Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 673 C>T R225W DI-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 677 C>T A226V DI-S Benign Patho Benign Patho Patho Patho Patho 5 control nssnv 694 G>A V232I DI-S Patho Patho Benign Benign Patho Patho Patho 5 case nssnv 718 G>A V240M DI-S4/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 739 G>C V247L DI-S4/ Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 799 A>G I267V DI-S Benign Benign Benign Benign Patho Benign Benign 1 case nssnv 808 C>A Q270K DI-S Benign Patho Benign Patho Patho Patho Patho 5 case nssnv 825 C>A N275K DI-S Benign Patho Benign Patho Patho Benign Patho 4 case nssnv 827 T>A L276Q DI-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 832 C>G H278D DI-S5/ Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 844 C>T R282C DI-S5/ Benign Benign Patho Patho Patho Patho Patho 4 Polymorphism 856 G>T A286S DI-S5/ Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 865 G>A G289S DI-S5/ Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 872 A>G N291S DI-S5/ Benign Benign Benign Benign Patho Benign Patho 2 Polymorphism 895 T>A L299M DI-S5/ Benign Benign Benign Benign Patho Benign Benign 1 case nssnv 898 G>A V300I DI-S5/ Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 944 T>C L315P DI-S5/ Benign Benign Benign Patho Patho Benign Patho 3 case nssnv 959 C>A T320N DI-S5/ Benign Benign Benign Benign Benign Benign Patho 1 case nssnv 974 T>G L325R DI-S5/ Patho Benign Patho Patho Patho Patho Benign 4 case nssnv 1007 C>T P336L DI-S5/ Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 1018 C>T R340W DI-S5/ Benign Benign Patho Patho Patho Benign Patho 3 control nssnv 1019 G>T R340L DI-S5/ Benign Benign Patho Benign Benign Benign Benign 0 control nssnv 1036 G>A E346K DI-S5/ Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 1042 C>G P348A DI-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 1052 G>T G351V DI-S5/ Benign Benign Patho Patho Patho Patho Patho 4 case nssnv 1052 G>A G351D DI-S5/ Benign Benign Benign Patho Patho Patho Patho 4
10 control nssnv 1051 G>A G351S DI-S5/ Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 1066 G>A D356N DI-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 1099 C>T R367C DI-S5/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 1100 G>T R367L DI-S5/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 1100 G>A R367H DI-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 1106 T>A M369K DI-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 1109 C>T T370M DI-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 1120 T>G W374G DI-S5/ Patho Patho Patho Patho Patho Patho Patho 6 control nssnv 1127 G>A R376H DI-S5/ Benign Benign Benign Patho Patho Patho Patho 4 control nssnv 1126 C>T R376C DI-S5/ Benign Benign Patho Patho Patho Patho Patho 4 case nssnv 1156 G>A G386R DI-S5/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 1157 G>A G386E DI-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 1186 G>C V396L DI-S Patho Benign Benign Patho Patho Benign Patho 4 case nssnv 1187 T>C V396A DI-S Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 1190 T>C I397T DI-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 1218 C>G N406K DI-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 1225 C>G L409V DI-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 1231 G>A V411M DI-S Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 1247 A>G Y416C DI/DII Patho Patho Patho Patho Patho Patho Patho 6 Polymorphism 1282 G>A E428K DI/DII Benign Benign Benign Benign Patho Benign Benign 1 case nssnv 1315 G>A E439K DI/DII Benign Benign Benign Patho Patho Patho Patho 4 control nssnv 1336 G>A E446K DI/DII Benign Patho Benign Patho Patho Patho Patho 5 Polymorphism 1340 C>G A447G DI/DII Patho Benign Benign Benign Patho Benign Patho 3 control nssnv 1345 A>G T449A DI/DII Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 1373 G>A R458H DI/DII Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 1381 T>G L461V DI/DII Benign Benign Benign Benign Patho Benign Benign 1 case nssnv 1385 A>C E462A DI/DII Benign Benign Patho Benign Patho Benign Patho 2 control nssnv 1384 G>A E462K DI/DII Benign Benign Benign Benign Patho Benign Patho 2 control nssnv 1405 G>A V469I DI/DII Benign Benign Benign Benign Benign Benign Patho 1 Polymorphism 1425 A>C R475S DI/DII Benign Benign Patho Benign Benign Benign Patho 1 Polymorphism 1441 C>T R481W DI/DII Benign Benign Patho Patho Patho Benign Benign 2 control nssnv 1442 G>A R481Q DI/DII Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 1502 A>G D501G DI/DII Patho Patho Benign Benign Patho Patho Patho 5 control nssnv 1537 C>T R513C DI/DII Benign Benign Patho Patho Benign Benign Patho 2 control nssnv 1561 A>G K521E DI/DII Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 1568 G>A R523H DI/DII Benign Benign Benign Benign Patho Benign Patho 2 Polymorphism 1571 C>A S524Y DI/DII Benign Benign Patho Patho Patho Patho Patho 4 control nssnv 1577 G>A R526H DI/DII Benign Benign Benign Benign Patho Benign Benign 1 case nssnv 1588 T>G F530V DI/DII Benign Benign Benign Benign Patho Patho Patho 3 case nssnv 1595 T>G F532C DI/DII Patho Benign Patho Patho Patho Patho Patho 5
11 Polymorphism 1598 G>A R533H DI/DII Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 1604 G>A R535Q DI/DII Patho Benign Benign Benign Patho Benign Patho 3 case nssnv 1629 T>A F543L DI/DII Patho Patho Benign Patho Patho Patho Patho 6 Polymorphism 1652 C>T A551V DI/DII Benign Benign Benign Benign Benign Benign Patho 1 case nssnv 1654 G>A G552R DI/DII Benign Benign Patho Patho Patho Patho Patho 4 control nssnv 1654 G>T G552W DI/DII Benign Benign Patho Patho Patho Patho Patho 4 Polymorphism 1673 A>G H558R DI/DII Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 1703 G>A R568H DI/DII Benign Benign Benign Patho Patho Patho Benign 3 case nssnv 1705 C>T R569W DI/DII Benign Benign Patho Patho Patho Patho Patho 4 case nssnv 1712 G>T S571I DI/DII Benign Benign Patho Patho Patho Patho Patho 4 control nssnv 1715 C>T A572V DI/DII Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 1714 G>T A572S DI/DII Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 1715 C>A A572D DI/DII Benign Benign Patho Benign Benign Benign Benign 0 control nssnv 1735 G>A G579R DI/DII Benign Benign Patho Benign Patho Benign Benign 1 control nssnv 1776 C>A N592K DI/DII Benign Benign Benign Benign Patho Patho Patho 3 control nssnv 1787 A>G D596G DI/DII Benign Benign Benign Benign Patho Benign Patho 2 control nssnv 1802 T>C V601A DI/DII Benign Benign Benign Benign Patho Benign Patho 2 control nssnv 1844 G>A G615E DI/DII Benign Patho Benign Benign Patho Benign Patho 3 Polymorphism 1852 C>T L618F DI/DII Benign Benign Benign Benign Patho Benign Patho 2 control nssnv 1855 C>T L619F DI/DII Patho Benign Benign Patho Patho Benign Patho 4 case nssnv 1858 C>T R620C DI/DII Benign Benign Patho Patho Patho Benign Patho 3 case nssnv 1895 C>T T632M DI/DII Benign Benign Benign Patho Patho Benign Patho 3 control nssnv 1913 G>A G638D DI/DII Benign Benign Benign Benign Benign Benign Patho 1 case nssnv 1915 G>A G639R DI/DII Benign Benign Patho Benign Benign Benign Benign 0 case nssnv 1918 C>G P640A DI/DII Benign Benign Benign Patho Patho Benign Benign 2 Polymorphism 1940 C>A A647D DI/DII Benign Benign Patho Benign Benign Benign Benign 0 case nssnv 1943 C>T P648L DI/DII Benign Benign Benign Benign Patho Benign Benign 1 case nssnv 1960 G>A E654K DI/DII Benign Benign Benign Patho Patho Patho Patho 4 Polymorphism 1967 C>T P656L DI/DII Benign Benign Benign Patho Patho Benign Patho 3 control nssnv 1976 G>A R659Q DI/DII Benign Benign Benign Patho Patho Benign Patho 3 control nssnv 1981 C>T R661W DI/DII Patho Benign Patho Patho Patho Patho Patho 5 Polymorphism 1984 G>T A662S DI/DII Benign Benign Benign Patho Patho Benign Patho 3 Polymorphism 2014 G>A A672T DI/DII Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 2018 T>C L673P DI/DII Benign Benign Benign Patho Patho Benign Patho 3 control nssnv 2038 C>T R680C DI/DII Benign Benign Patho Patho Benign Benign Benign 1 case nssnv 2047 T>G C683G DI/DII Benign Patho Patho Patho Patho Patho Benign 4 case nssnv 2065 C>T R689C DI/DII Benign Benign Patho Patho Patho Patho Patho 4 Polymorphism 2066 G>A R689H DI/DII Benign Benign Benign Patho Patho Patho Benign 3 Polymorphism 2074 C>A Q692K DI/DII Benign Benign Benign Patho Benign Benign Benign 1 control nssnv 2078 G>A R693H DI/DII Benign Benign Benign Benign Benign Benign Benign 0
12 control nssnv 2077 C>T R693C DI/DII Benign Benign Patho Benign Patho Benign Patho 2 control nssnv 2102 C>T P701L DI/DII Benign Benign Benign Patho Patho Patho Patho 4 control nssnv 2114 C>T S705F DI/DII Benign Benign Patho Patho Patho Benign Patho 3 control nssnv 2150 C>T P717L DII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 2192 C>T T731I DII-S Patho Patho Benign Patho Patho Patho Benign 5 case nssnv 2204 C>T A735V DII-S Patho Patho Benign Patho Patho Patho Benign 5 control nssnv 2236 G>A E746K DII-S1/ Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 2249 A>G Q750R DII-S Benign Benign Benign Patho Benign Benign Benign 1 case nssnv 2254 G>A G752R DII-S Benign Patho Patho Patho Patho Patho Patho 5 case nssnv 2273 G>A G758E DII-S Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 2291 T>G M764R DII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 2314 G>A D772N DII-S2/ Benign Patho Benign Patho Patho Patho Patho 5 case nssnv 2317 C>T P773S DII-S2/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 2365 G>A V789I DII-S Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 2423 G>C R808P DII-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 2447 T>A F816Y DII-S Benign Patho Benign Patho Patho Patho Patho 5 Polymorphism 2497 G>A G833R DII-S4/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 2516 T>C L839P DII-S4/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 2542 A>T I848F DII-S Benign Patho Benign Patho Patho Patho Patho 5 case nssnv 2553 C>A F851L DII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 2599 G>C E867Q DII-S5/ Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 2632 C>T R878C DII-S5/ Patho Benign Patho Patho Patho Patho Patho 5 case nssnv 2633 G>A R878H DII-S5/ Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 2657 A>C H886P DII-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 2677 C>T R893C DII-S5/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 2678 G>A R893H DII-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 2701 G>A E901K DII-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 2729 C>T S910L DII-S5/ Benign Benign Patho Patho Patho Benign Patho 3 case nssnv 2743 T>C C915R DII-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 2750 T>G L917R DII-S Benign Benign Patho Patho Patho Patho Patho 4 Polymorphism 2770 G>A V924I DII-S Benign Patho Benign Patho Benign Benign Benign 2 case nssnv 2780 A>G N927S DII-S Patho Benign Benign Patho Patho Patho Benign 4 case nssnv 2783 T>C L928P DII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 2804 T>C L935P DII-S Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 2878 C>A Q960K DII/DIII Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 2893 C>T R965C DII/DIII Benign Patho Patho Patho Patho Patho Patho 5 case nssnv 2894 G>A R965H DII/DIII Benign Patho Benign Patho Patho Patho Patho 5 case nssnv 2894 G>T R965L DII/DIII Benign Patho Patho Patho Patho Patho Patho 5 Polymorphism 2905 G>T G969C DII/DIII Patho Benign Patho Patho Patho Patho Patho 5 control nssnv 2923 C>T R975W DII/DIII Benign Benign Patho Patho Patho Patho Patho 4
13 case nssnv 2942 G>T C981F DII/DIII Benign Benign Patho Benign Benign Benign Patho 1 Polymorphism 2944 T>C C982R DII/DIII Benign Benign Patho Benign Benign Benign Patho 1 Polymorphism 2957 G>A R986Q DII/DIII Benign Patho Benign Benign Benign Benign Benign 1 case nssnv 2989 G>A A997T DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 2989 G>T A997S DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 3010 T>C C1004R DII/DIII Benign Benign Patho Benign Patho Patho Patho 3 control nssnv 3032 C>T P1011L DII/DIII Benign Benign Benign Benign Patho Benign Benign 1 control nssnv 3047 C>T T1016M DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 3080 G>A R1027Q DII/DIII Benign Benign Benign Benign Patho Patho Patho 3 control nssnv 3096 G>T E1032D DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 3094 G>A E1032K DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 3104 G>A G1035D DII/DIII Benign Benign Benign Patho Benign Patho Patho 3 control nssnv 3110 G>T G1037V DII/DIII Benign Benign Patho Benign Benign Benign Patho 1 Polymorphism 3118 G>A G1040R DII/DIII Benign Benign Patho Benign Patho Benign Benign 1 case nssnv 3157 G>A E1053K DII/DIII Patho Benign Benign Benign Patho Patho Patho 4 case nssnv 3164 A>G D1055G DII/DIII Patho Patho Benign Patho Patho Patho Benign 5 case nssnv 3206 C>T T1069M DII/DIII Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 3236 C>A S1079Y DII/DIII Benign Benign Patho Patho Patho Benign Patho 3 control nssnv 3235 T>A S1079T DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 3245 T>C V1082A DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 3262 G>A A1088T DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 3269 C>T P1090L DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 3292 G>T V1098L DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 3299 C>T A1100V DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 3308 C>A S1103Y DII/DIII Benign Benign Patho Patho Patho Benign Benign 2 control nssnv 3319 G>A E1107K DII/DIII Benign Benign Benign Benign Benign Benign Patho 1 case nssnv 3338 C>T A1113V DII/DIII Patho Patho Benign Benign Benign Benign Benign 2 case nssnv 3340 G>A D1114N DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 3346 C>T R1116W DII/DIII Benign Benign Patho Patho Patho Benign Benign 2 Polymorphism 3347 G>A R1116Q DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 3388 G>A E1130K DII/DIII Patho Benign Benign Benign Benign Benign Benign 1 control nssnv 3392 C>T T1131I DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 3391 A>T T1131S DII/DIII Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 3419 G>C S1140T DII/DIII Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 3496 G>A D1166N DII/DIII Benign Benign Benign Benign Patho Benign Benign 1 control nssnv 3520 C>T R1174W DII/DIII Benign Benign Patho Patho Patho Patho Benign 3 control nssnv 3524 G>A R1175H DII/DIII Benign Benign Benign Benign Patho Benign Patho 2 control nssnv 3539 C>T A1180V DII/DIII Benign Benign Benign Patho Benign Benign Benign 1 control nssnv 3542 T>C V1181A DII/DIII Benign Benign Benign Patho Patho Patho Benign 3 Polymorphism 3578 G>A R1193Q DII/DIII Benign Benign Benign Benign Benign Benign Benign 0
14 Polymorphism 3577 C>T R1193W DII/DIII Benign Benign Patho Patho Patho Patho Patho 4 case nssnv 3596 A>C Y1199S DII/DIII Benign Benign Patho Patho Patho Patho Patho 4 control nssnv 3604 G>A V1202M DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 3656 G>A S1219N DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 3673 G>A E1225K DIII-S1/ Benign Patho Benign Patho Patho Patho Patho 5 case nssnv 3682 T>C Y1228H DIII-S1/ Patho Benign Benign Benign Benign Patho Patho 3 case nssnv 3694 C>T R1232W DIII-S1/ Benign Benign Patho Patho Patho Patho Patho 4 case nssnv 3695 G>A R1232Q DIII-S1/ Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 3716 T>C L1239P DIII-S Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 3718 G>C E1240Q DIII-S Patho Benign Benign Patho Patho Patho Patho 5 control nssnv 3727 G>A D1243N DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 3746 T>A V1249D DIII-S Benign Benign Patho Patho Patho Patho Patho 4 Polymorphism 3751 G>A V1251M DIII-S Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 3758 A>G E1253G DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 3784 G>A G1262S DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 3813 G>C W1271C DIII-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 3823 G>A D1275N DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 3833T>A I1278N DIII-S Benign Patho Patho Patho Patho Patho Patho 5 control nssnv 3835 G>A V1279I DIII-S Benign Patho Benign Patho Patho Patho Patho 5 case nssnv 3847 C>A L1283M DIII-S Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 3863 C>G A1288G DIII-S Patho Patho Benign Patho Patho Patho Patho 6 Polymorphism 3878 T>C F1293S DIII-S3/ Benign Benign Patho Patho Patho Benign Patho 3 Polymorphism 3907 C>T R1303W DIII-S Patho Patho Patho Patho Patho Patho Patho 6 control nssnv 3911 C>T T1304M DIII-S Benign Patho Benign Patho Patho Patho Patho 5 Polymorphism 3922 C>T L1308F DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 3932 T>C L1311P DIII-S Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 3956 G>T G1319V DIII-S4/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 3968 T>G V1323G DIII-S4/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 3974 A>G N1325S DIII-S4/ Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 3976 G>T A1326S DIII-S4/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 3988G>A A1330T DIII-S4/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 3995 C>T P1332L DIII-S4/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4000 A>G I1334V DIII-S4/ Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 4003 A>G M1335V DIII-S4/ Patho Benign Benign Patho Patho Benign Patho 4 case nssnv 4012 C>G L1338V DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4018 G>A V1340I DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4030 T>C F1344L DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4036 C>A L1346I DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4037 T>C L1346P DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4052 T>G M1351R DIII-S Patho Benign Benign Patho Patho Patho Patho 5
15 case nssnv 4057 G>A V1353M DIII-S Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 4070 C>T A1357V DIII-S Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 4072 G>T G1358W DIII-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4077 G>T K1359N DIII-S Benign Patho Benign Patho Patho Patho Patho 5 case nssnv 4079 T>G F1360C DIII-S5/ Benign Benign Patho Patho Patho Patho Patho 4 case nssnv 4088 G>A C1363Y DIII-S5/ Patho Patho Patho Patho Patho Patho Patho 6 control nssnv 4090 A>G I1364V DIII-S5/ Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 4098 G>T Q1366H DIII-S5/ Benign Benign Benign Benign Patho Benign Patho 2 case nssnv 4145 G>T S1382I DIII-S5/ Benign Benign Patho Patho Patho Patho Patho 4 control nssnv 4161 G>C L1387F DIII-S5/ Benign Benign Benign Benign Patho Patho Benign 2 case nssnv 4213 G>A V1405M DIII-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4213 G>C V1405L DIII-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4216 G>C G1406R DIII-S5/ Patho Benign Patho Patho Patho Patho Patho 5 case nssnv 4217 G>A G1406E DIII-S5/ Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 4222 G>A G1408R DIII-S5/ Patho Benign Patho Patho Patho Patho Patho 5 case nssnv 4226 A>G Y1409C DIII-S5/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4234 C>T L1412F DIII-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4255 A>G K1419E DIII-S5/ Patho Patho Benign Patho Patho Benign Patho 5 case nssnv 4258 G>C G1420R DIII-S5/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4279 G>T A1427S DIII-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4283 C>T A1428V DIII-S5/ Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 4282 G>T A1428S DIII-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4294 A>G R1432G DIII-S5/ Patho Benign Patho Patho Patho Patho Patho 5 case nssnv 4296 G>C R1432S DIII-S5/ Patho Benign Patho Patho Patho Patho Patho 5 case nssnv 4298 G>T G1433V DIII-S5/ Benign Benign Patho Patho Patho Patho Patho 4 case nssnv 4313 C>T P1438L DIII-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4321 G>C E1441Q DIII-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4342 A>C I1448L DIII-S Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 4343 T>C I1448T DIII-S Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 4346 A>G Y1449C DIII-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4352 T>A V1451D DIII-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4373 C>A S1458Y DIII-S Benign Benign Patho Patho Patho Patho Patho 4 case nssnv 4387 A>T N1463Y DIII-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4402 G>T V1468F DIII-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4415 A>G N1472S DIII/DIV Benign Patho Benign Patho Patho Patho Benign 4 case nssnv 4418 T>G F1473C DIII/DIV Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4442 G>A G1481E DIII/DIV Benign Patho Benign Patho Patho Patho Patho 5 case nssnv 4459 A>C M1487L DIII/DIV Benign Patho Benign Patho Patho Patho Benign 4 case nssnv 4463 C>G T1488R DIII/DIV Patho Patho Benign Patho Patho Patho Benign 5 case nssnv 4467 G>T E1489D DIII/DIV Patho Patho Benign Patho Patho Patho Benign 5
16 case nssnv 4478 A>G K1493R DIII/DIV Patho Patho Benign Patho Patho Patho Benign 5 case nssnv 4484 A>C Y1495S DIII/DIV Benign Patho Patho Patho Patho Patho Patho 5 case nssnv 4492 A>G M1498V DIII/DIV Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4501 C>G L1501V DIII/DIV Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4515 G>T K1505N DIII/DIV Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 4534 C>T R1512W DIII/DIV Patho Patho Patho Patho Patho Patho Patho 6 Polymorphism 4535 G>A R1512Q DIII/DIV Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 4544 A>G N1515S DIII/DIV Patho Patho Benign Patho Benign Patho Patho 5 case nssnv 4562 T>A I1521K DIII/DIV Benign Benign Patho Patho Patho Patho Benign 3 control nssnv 4571 T>C I1524T DIV-S Benign Benign Benign Patho Patho Patho Patho 4 control nssnv 4573 G>A V1525M DIV-S Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 4594 G>A V1532I DIV-S Benign Benign Benign Benign Benign Benign Patho 1 control nssnv 4595 T>C V1532A DIV-S Benign Benign Benign Patho Patho Benign Patho 3 case nssnv 4642 G>A E1548K DIV-S1/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4680 G>C L1560F DIV-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4712 T>G F1571C DIV-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4720 G>A E1574K DIV-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4745 T>C L1582P DIV-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4747 C>T R1583C DIV-S2/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4748 G>A R1583H DIV-S2/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4779 C>G I1593M DIV-S Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 4781 T>C F1594S DIV-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4786 T>A F1596I DIV-S Patho Benign Benign Patho Patho Patho Benign 4 case nssnv 4810 G>A V1604M DIV-S Patho Benign Benign Patho Patho Benign Patho 4 case nssnv 4838 A>T Q1613L DIV-S3/ Benign Benign Patho Patho Patho Benign Benign 2 case nssnv 4859 C>T T1620M DIV-S3/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4868 G>A R1623Q DIV-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4868 G>T R1623L DIV-S Patho Patho Patho Patho Patho Patho Benign 5 case nssnv 4877 G>A R1626H DIV-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4886 G>A R1629Q DIV-S Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 4913 G>A R1638Q DIV-S Benign Benign Benign Patho Patho Patho Patho 4 control nssnv 4916 G>C G1639A DIV-S Benign Benign Benign Benign Patho Benign Patho 2 case nssnv 4925 G>A G1642E DIV-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4930 C>T R1644C DIV-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4931 G>A R1644H DIV-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4948 C>T L1650F DIV-S4/ Benign Patho Benign Patho Patho Patho Patho 5 case nssnv 4955 T>C M1652T DIV-S4/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4978 A>G I1660V DIV-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 4981 G>A G1661R DIV-S Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 4981 G>C G1661R DIV-S Patho Patho Patho Patho Patho Patho Patho 6
17 case nssnv 4999 G>A V1667I DIV-S Patho Benign Benign Patho Patho Benign Patho 4 case nssnv 5015 C>A S1672Y DIV-S Benign Benign Patho Patho Patho Patho Patho 4 case nssnv 5038 G>A A1680T DIV-S Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 5092 G>A A1698T DIV-S5/ Benign Benign Benign Patho Patho Benign Patho 3 case nssnv 5126 C>T T1709M DIV-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 5126 C>G T1709R DIV-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 5134 G>A G1712S DIV-S5/ Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 5141 A>G D1714G DIV-S5/ Benign Patho Benign Patho Patho Patho Patho 5 case nssnv 5164 A>G N1722D DIV-S5/ Benign Benign Benign Patho Patho Patho Patho 4 case nssnv 5168 C>A T1723N DIV-S5/ Benign Benign Benign Patho Patho Benign Patho 3 case nssnv 5182 T>C C1728R DIV-S5/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 5184 C>G C1728W DIV-S5/ Patho Patho Patho Patho Patho Patho Patho 6 control nssnv 5215 C>T R1739W DIV-S5/ Benign Benign Patho Patho Patho Patho Patho 4 case nssnv 5218 G>A G1740R DIV-S5/ Patho Patho Patho Patho Patho Patho Patho 6 case nssnv 5227 G>A G1743R DIV-S5/ Patho Benign Patho Patho Patho Patho Patho 5 case nssnv 5228 G>A G1743E DIV-S5/ Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 5282 T>A L1761H DIV-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 5281 C>T L1761F DIV-S Patho Patho Benign Patho Patho Benign Patho 5 case nssnv 5287 G>A V1763M DIV-S Patho Benign Benign Patho Patho Benign Patho 4 case nssnv 5290 G>T V1764F DIV-S Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 5296 A>C M1766L DIV-S Patho Patho Benign Patho Patho Benign Patho 5 case nssnv 5302 A>G I1768V DIV-S Patho Patho Benign Patho Patho Benign Patho 5 case nssnv 5329 G>A V1777M C-term Patho Benign Benign Patho Patho Patho Patho 5 control nssnv 5336 C>T T1779M C-term Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 5350 G>A E1784K C-term Patho Benign Benign Patho Patho Patho Patho 5 Polymorphism 5360 G>A S1787N C-term Patho Benign Benign Patho Patho Patho Patho 5 case nssnv 5384 A>G Y1795C C-term Patho Patho Patho Patho Patho Patho Patho 6 control nssnv 5389 A>G I1797V C-term Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 5435 C>T S1812L C-term Benign Benign Patho Benign Benign Benign Benign 0 control nssnv 5477 G>A R1826H C-term Patho Benign Benign Benign Patho Patho Patho 4 control nssnv 5476 C>T R1826C C-term Patho Benign Patho Patho Patho Patho Patho 5 control nssnv 5494 C>G Q1832E C-term Benign Benign Benign Patho Patho Benign Patho 3 Polymorphism 5507 T>C I1836T C-term Benign Benign Benign Patho Patho Patho Benign 3 case nssnv 5516 A>G D1839G C-term Patho Patho Benign Patho Patho Patho Patho 6 control nssnv 5524 A>T M1842L C-term Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 5581 G>A V1861I C-term Patho Patho Benign Patho Patho Patho Patho 6 case nssnv 5616 G>C K1872N C-term Patho Benign Benign Patho Patho Patho Patho 5 Polymorphism 5689 C>T R1897W C-term Benign Benign Patho Benign Patho Benign Patho 2 Polymorphism 5693 G>A R1898H C-term Patho Benign Benign Patho Patho Patho Patho 5 control nssnv 5692 C>T R1898C C-term Patho Benign Patho Patho Patho Patho Patho 5
18 case nssnv 5701 G>C E1901Q C-term Patho Patho Benign Patho Patho Benign Patho 5 Polymorphism 5701 G>A E1901K C-term Patho Patho Benign Patho Patho Benign Patho 5 Polymorphism 5711 C>T S1904L C-term Benign Benign Patho Patho Patho Benign Patho 3 case nssnv 5726 A>G Q1909R C-term Patho Patho Benign Patho Patho Benign Patho 5 control nssnv 5755 C>T R1919C C-term Patho Benign Patho Patho Patho Benign Patho 4 Polymorphism 5756 G>A R1919H C-term Patho Benign Benign Patho Patho Benign Patho 4 control nssnv 5770 G>A A1924T C-term Patho Benign Benign Benign Patho Benign Patho 3 control nssnv 5795 C>T A1932V C-term Benign Patho Benign Benign Patho Benign Benign 2 case nssnv 5803 G>A G1935S C-term Benign Benign Benign Benign Benign Benign Benign 0 case nssnv 5812 G>A E1938K C-term Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 5849 A>G Y1950C C-term Benign Benign Patho Patho Benign Benign Benign 1 Polymorphism 5851 G>T V1951L C-term Benign Benign Benign Benign Patho Benign Benign 1 Polymorphism 5873 G>A R1958Q C-term Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 5885 C>T P1962L C-term Benign Patho Benign Patho Patho Patho Benign 4 Polymorphism 5904 C>G I1968M C-term Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 5903 T>G I1968S C-term Benign Benign Patho Benign Benign Benign Benign 0 case nssnv 5929 T>A Y1977N C-term Benign Benign Patho Patho Patho Patho Patho 4 control nssnv 5961 C>A N1987K C-term Benign Benign Benign Benign Benign Benign Patho 1 Polymorphism 5963 T>G L1988R C-term Benign Benign Patho Benign Benign Benign Benign 0 Polymorphism 5972 G>A R1991Q C-term Benign Benign Benign Benign Benign Benign Benign 0 control nssnv 5971 C>T R1991W C-term Benign Benign Patho Patho Patho Benign Patho 3 control nssnv 6007 G>A D2003N C-term Benign Benign Benign Patho Patho Benign Patho 3 case nssnv 6010 T>G F2004V C-term Benign Benign Benign Benign Patho Benign Benign 1 Polymorphism 6010 T>C F2004L C-term Benign Benign Benign Benign Benign Benign Benign 0 Polymorphism 6016 C>G P2006A C-term Benign Benign Benign Benign Patho Benign Benign 1 case nssnv 6034 C>T R2012C C-term Benign Benign Patho Patho Patho Benign Patho 3 Abbreviations: Mass, Mutation Assesor; PP2, PolyPhen2; Patho, pathogenic
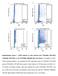
19 1 0.8 Sensitivity LQTS nsnvs BrS nsnvs All Case nsnvs specificity Supplemental Figure 1: Receiver operator curve used to identify the number of synergist tools required to achieve the highest specificity and sensitivity for the differentiation of case- and control- rare nssnvs.
20 Supplemental Table 2: Summary of Cellular EP parameters for SCN5A variants from our lab SCN5A Variants Current Density Activation Inactivation Late Current pa/pf (-20 mv) V 1/2 (mv) V 1/2 (mv) (%) WT-Q1077del ±71.7 (10) -35.7±0.8 (10) -79.4±0.9 (10) 0.12±0.03 (9) E30G-Q1077del ±85.9 (11) -35.9±0.9 (12) -76.8±0.5 (12)* 0.22±0.06 (11) WT-Q1077del ±45.6 (9) -40.0±0.8 (9) -87.2±0.8 (9) 0.17±0.03 (7) E48K-Q1077del ±32.5 (7) -36.1±1.6 (7)* -89.9±1.0 (7) 0.17±0.03 (6) WT-Q ±47.1 (8) -40.7±0.6 (8) -88.6±0.5 (7) 0.16±0.06 (7) E48K-Q ±48.7 (9) -36.9±1.7 (9) -88.2±0.9 (9) 0.22±0.05 (9) WT-Q1077del ±86.6 (12) -37.1±1.0 (12) -79.0±0.3 (6) 0.13±0.04 (7) Y87C-Q1077del ±68.1 (14) -36.5±0.8 (14) -80.5±0.9 (14) 0.15±0.05 (10) WT-Q1077del ±85.3 (13) -37.1±1.4 (13) -79.7±0.7 (13) 0.19±0.07 (10) R190Q-Q1077del ±102.1 (13) -37.0±1.5 (13) -78.9±0.7 (13) 0.23±0.07 (10) WT-Q1077del ±76.5 (15) -33.4±1.2 (15) -77.9±0.5 (15) 0.23±0.08 (7) S216L-Q1077del ±27.2 (15)* -35.4±1.4 (15) -77.6±0.8 (15) 0.19±0.06 (7) WT-Q1077del ±84.4 (13) -37.0±1.4 (13) -81.2±0.7 (13) 0.3±0.05 (9) Q245K-Q1077del ±103.2 (12) -40.7±1.5 (12) -80.5±1.0 (12) 0.3±0.1 (8) WT-Q1077del ±74.3 (11) -35.2±1.0 (11) -79.6±0.8 (10) 0.17±0.1 (8) I397F-Q1077del ±38.4 (13)* -42.1±0.7 (13)* -74.5±1.5 (13)* 0.82±0.1 (13)*
21 WT-Q ±84.1 (9) -38.3±0.3 (9) -79.9±1.2 (8) 0.16±0.06 (8) I397F-Q ±37.2 (12)* -37.9±1.4 (12) -70.8±0.5 (12)* 1.08±0.09 (8)* WT-Q1077del ±61.5 (14) -35.4±1.5 (14) -82.8±0.7 (14) 0.21±0.06 (11) E462A-Q1077del ±72.6 (13) -38.0±1.4 (11) -81.6±0.9 (13) 0.20±0.03 (13) WT-Q ±58.0 (11) -36.2±1.4 (11) -82.4±1.0 (11) 0.15±0.02 (10) E462A-Q ±63.3 (14) -38.4±1.1 (14) -81.5±0.7 (14) 0.20±0.03 (13) WT-Q1077del ±81.3 (15) -30.3±1.9 (15) -78.3±0.8 (13) 0.26±0.05 (11) E462K-Q1077del ±62.7 (15) -32.2±1.4 (15) -81.3±0.7 (15) 0.22±0.05 (7) WT-Q1077del/R ±74.4 (15) -33.6±1.7 (15) -79.4±0.7 (14) 0.16±0.04 (11) E462K-Q1077del/R ±100.7 (13) -36.8±0.8 (13) -80.2±0.4 (12) 0.19±0.06 (10) WT-Q1077del ±88.4 (13) -34.9±1.4 (13) -81.8±0.7 (13) 0.21±0.09 (12) R569G-Q1077del ±86.8 (13) -42.0±1.1 (13)* -81.1±1.0 (13) 0.19±0.05 (9) WT-Q1077del ±85.3 (15) -34.4±1.5 (15) -80.1±0.4 (14) 0.24±0.06 (10) R569W-Q1077del ±92.2 (9) -35.7±1.5 (9) -79.4±0.5 (9) 0.16±0.02 (7) WT-Q1077del ±84.4 (13) -37.0±1.4 (13) -81.2±0.7 (13) 0.30±0.05 (9) R620C-Q1077del ±124.7 (10) -36.5±0.9 (10) -78.9±0.7 (10)* 0.18±0.06 (8) WT-Q1077del ±91.2 (11) -35.1±2.0 (11) -80.5±0.5 (11) 0.22±0.06 (9) P627L-Q1077del ±93.4 (10) -36.2±1.2 (10) -79.5±0.4 (8) 0.18±0.07 (5) WT-Q1077del ±81.3 (15) -30.3±1.9 (15) -78.3±0.8 (13) 0.26±0.05 (11)
modified dye uptake assay including formazan test EC 90 not tested plaque reduction assay
 Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E,
 Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples
 A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Original Article. Enhanced Classification of Brugada Syndrome Associated and Long-QT Syndrome Associated Genetic Variants in the SCN5A-Encoded Na v
 Original Article Enhanced Classification of Brugada Syndrome Associated and Long-QT Syndrome Associated Genetic Variants in the SCN5A-Encoded Na v 1.5 Cardiac Sodium Channel Jamie D. Kapplinger, BA; John
Original Article Enhanced Classification of Brugada Syndrome Associated and Long-QT Syndrome Associated Genetic Variants in the SCN5A-Encoded Na v 1.5 Cardiac Sodium Channel Jamie D. Kapplinger, BA; John
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication
 Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
SUPPLEMENTARY INFORMATION. Rare independent mutations in renal salt handling genes contribute to blood pressure variation
 SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Supplementary Figure 1 Weight and body temperature of ferrets inoculated with
 Supplementary Figure 1 Weight and body temperature of ferrets inoculated with A/Anhui/1/2013 (H7N9) influenza virus. (a) Body temperature and (b) weight change of ferrets after intranasal inoculation with
Supplementary Figure 1 Weight and body temperature of ferrets inoculated with A/Anhui/1/2013 (H7N9) influenza virus. (a) Body temperature and (b) weight change of ferrets after intranasal inoculation with
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP
 Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
HCV NS3 Protease Drug Resistance
 test code: cpt code: 10000 87902 category: Infectious Disease HCV genotype 1a CODON Glecaprevir 1a Grazoprevir 1a Paritaprevir 1a Simeprevir 1a Voxilaprevir 1a V36A S 13 R 10 Comment V36C V36G V36I V36L
test code: cpt code: 10000 87902 category: Infectious Disease HCV genotype 1a CODON Glecaprevir 1a Grazoprevir 1a Paritaprevir 1a Simeprevir 1a Voxilaprevir 1a V36A S 13 R 10 Comment V36C V36G V36I V36L
Pharmacogenomics in Rare Diseases: Development Strategy for Ivacaftor as a Therapy for Cystic Fibrosis
 Pharmacogenomics in Rare Diseases: Development Strategy for Ivacaftor as a Therapy for Cystic Fibrosis Federico Goodsaid Vice President Strategic Regulatory Intelligence Vertex Pharmaceuticals Is there
Pharmacogenomics in Rare Diseases: Development Strategy for Ivacaftor as a Therapy for Cystic Fibrosis Federico Goodsaid Vice President Strategic Regulatory Intelligence Vertex Pharmaceuticals Is there
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13898 Supplementary Information Table 1 Kras mutation status of carcinogen-induced mouse lung adenomas Tumour Treatment Strain Grade Genotype Kras status (WES)* Kras status (Sanger) 32T1
doi:10.1038/nature13898 Supplementary Information Table 1 Kras mutation status of carcinogen-induced mouse lung adenomas Tumour Treatment Strain Grade Genotype Kras status (WES)* Kras status (Sanger) 32T1
Human TRPC6 Ion Channel Cell Line
 TECHNICAL DATA SHEET ValiScreen Ion Channel Cell Line Caution: For Laboratory Use. A research product for research purposes only Human TRPC6 Ion Channel Cell Line Product No.: AX-012-C Lot No.: 512-548-A
TECHNICAL DATA SHEET ValiScreen Ion Channel Cell Line Caution: For Laboratory Use. A research product for research purposes only Human TRPC6 Ion Channel Cell Line Product No.: AX-012-C Lot No.: 512-548-A
CPTR title slide. A Standardized System for Grading Mutations in Mycobacterium tuberculosis for Association with Drug Resistance
 CPTR title slide A Standardized System for Grading Mutations in Mycobacterium tuberculosis for Association with Drug Resistance PAOLO MIOTTO CPTR 2017 Workshop, March 20 23 The Need A lack of user-friendly
CPTR title slide A Standardized System for Grading Mutations in Mycobacterium tuberculosis for Association with Drug Resistance PAOLO MIOTTO CPTR 2017 Workshop, March 20 23 The Need A lack of user-friendly
Vertical Magnetic Separation of Circulating Tumor Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients
 Vertical Magnetic Separation of Circulating Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients Chang Eun Yoo 1,2#, Jong-Myeon Park 3#, Hui-Sung Moon 1,2, Je-Gun Joung 2, Dae-Soon Son
Vertical Magnetic Separation of Circulating Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients Chang Eun Yoo 1,2#, Jong-Myeon Park 3#, Hui-Sung Moon 1,2, Je-Gun Joung 2, Dae-Soon Son
Deep-Sequencing of HIV-1
 Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Deep-Sequencing of HIV-1 The quest for true variants Alexander Thielen, Martin Däumer 09.05.2015 Limitations of drug resistance testing by standard-sequencing Blood plasma RNA extraction RNA Reverse Transcription/
Sample Metrics. Allele Frequency (%) Read Depth Ploidy. Gene CDS Effect Protein Effect. LN Metastasis Tumor Purity Computational Pathology 80% 60%
 Supplemental Table 1: Estimated tumor purity, allele frequency, and independent read depth for all gene mutations classified as either potentially pathogenic or VUS in the metatastic and primary tumor
Supplemental Table 1: Estimated tumor purity, allele frequency, and independent read depth for all gene mutations classified as either potentially pathogenic or VUS in the metatastic and primary tumor
Relative activity (%) SC35M
 a 125 Bat (H17N) b 125 A/WSN (H1N1) Relative activity (%) 0 75 50 25 Relative activity (%) 0 75 50 25 0 Pos. Neg. PA PB1 Pos. Neg. NP PA PB1 PB2 0 Pos. Neg. NP PA PB1 PB2 SC35M Bat Supplementary Figure
a 125 Bat (H17N) b 125 A/WSN (H1N1) Relative activity (%) 0 75 50 25 Relative activity (%) 0 75 50 25 0 Pos. Neg. PA PB1 Pos. Neg. NP PA PB1 PB2 0 Pos. Neg. NP PA PB1 PB2 SC35M Bat Supplementary Figure
HCV NS3 Protease Drug Resistance
 test code: cpt code: 10000 87902 category: Infectious Disease HCV genotypes 1a CODON Simeprevir 1a Boceprevir 1a Telaprevir 1a Paritaprevir 1a Grazoprevir 1a V36A Comment R 3 R 10, 11 R 16 S 24 V36C V36G
test code: cpt code: 10000 87902 category: Infectious Disease HCV genotypes 1a CODON Simeprevir 1a Boceprevir 1a Telaprevir 1a Paritaprevir 1a Grazoprevir 1a V36A Comment R 3 R 10, 11 R 16 S 24 V36C V36G
Personalized Healthcare Update
 Dr. Kai - Oliver Wesche Market Development Manager, Personalized Healthcare QIAGEN Personalized Healthcare Update Pioneering Personalized Medicine through Partnering TOMTOVOK BKM120 Zelboraf QIAGEN partners:
Dr. Kai - Oliver Wesche Market Development Manager, Personalized Healthcare QIAGEN Personalized Healthcare Update Pioneering Personalized Medicine through Partnering TOMTOVOK BKM120 Zelboraf QIAGEN partners:
History (August 2010) Therapy for Experienced Patients. History (September 2010) History (November 2010) 12/2/11
 (August 2010) Therapy for Experienced Patients Hiroyu Hatano, MD, MHS Assistant Professor of Medicine University of California San Francisco Medical Management of AIDS December 2011 42M HIV (CD4=450, VL=6250,
(August 2010) Therapy for Experienced Patients Hiroyu Hatano, MD, MHS Assistant Professor of Medicine University of California San Francisco Medical Management of AIDS December 2011 42M HIV (CD4=450, VL=6250,
Genetic Analysis of Allosteric Signaling in RhaR from Escherichia coli and Characterization of the VirF Protein from Shigella flexneri
 Genetic Analysis of Allosteric Signaling in RhaR from Escherichia coli and Characterization of the VirF Protein from Shigella flexneri By Bria Collette Kettle Submitted to the graduate degree program in
Genetic Analysis of Allosteric Signaling in RhaR from Escherichia coli and Characterization of the VirF Protein from Shigella flexneri By Bria Collette Kettle Submitted to the graduate degree program in
Fabry Disease X-linked genetic, multi-organ disorder. Fabry disease screening program in Hypertrophic p Cardiomyopathy: preliminary results.
 Fabry Disease X-linked genetic, multi-organ disorder Fabry disease screening program in Hypertrophic p Cardiomyopathy: y preliminary results. Globotriaosylceramide, GL3 Brain -galactosidase A Eyes Lactosylceramide
Fabry Disease X-linked genetic, multi-organ disorder Fabry disease screening program in Hypertrophic p Cardiomyopathy: y preliminary results. Globotriaosylceramide, GL3 Brain -galactosidase A Eyes Lactosylceramide
LAL: significato clinico e biologico delle mutazioni di Bcr-Abl
 LAL: significato clinico e biologico delle mutazioni di Bcr-Abl Simona Soverini Dipartimento di Ematologia e Scienze Oncologiche L. e A. Seràgnoli Università di Bologna The vast majority of Ph+ ALL patients
LAL: significato clinico e biologico delle mutazioni di Bcr-Abl Simona Soverini Dipartimento di Ematologia e Scienze Oncologiche L. e A. Seràgnoli Università di Bologna The vast majority of Ph+ ALL patients
Using SuSPect to Predict the Phenotypic Effects of Missense Variants. Chris Yates UCL Cancer Institute
 Using SuSPect to Predict the Phenotypic Effects of Missense Variants Chris Yates UCL Cancer Institute c.yates@ucl.ac.uk Outline SAVs and Disease Development of SuSPect Features included Feature selection
Using SuSPect to Predict the Phenotypic Effects of Missense Variants Chris Yates UCL Cancer Institute c.yates@ucl.ac.uk Outline SAVs and Disease Development of SuSPect Features included Feature selection
Key determinants of pathogenicity
 Key determinants of pathogenicity Session 6: Determining pathogenicity and genotype-phenotype correlation J. Peter van Tintelen MD PhD Clinical geneticist Academic Medical Center Amsterdam, the Netherlands
Key determinants of pathogenicity Session 6: Determining pathogenicity and genotype-phenotype correlation J. Peter van Tintelen MD PhD Clinical geneticist Academic Medical Center Amsterdam, the Netherlands
Susanne Schnittger. Workflow of molecular investigations in JAK2-negative MPNs - the Munich experience
 Susanne Schnittger Workflow of molecular investigations in JAK2negative MPNs the Munich experience Cohort single centre experience to apply new markers in a daily diagnostic work flow total: 20,547 cases
Susanne Schnittger Workflow of molecular investigations in JAK2negative MPNs the Munich experience Cohort single centre experience to apply new markers in a daily diagnostic work flow total: 20,547 cases
Deliverable 2.1 List of relevant genetic variants for pre-emptive PGx testing
 GA N 668353 H2020 Research and Innovation Deliverable 2.1 List of relevant genetic variants for pre-emptive PGx testing WP N and Title: WP2 - Towards shared European Guidelines for PGx Lead beneficiary:
GA N 668353 H2020 Research and Innovation Deliverable 2.1 List of relevant genetic variants for pre-emptive PGx testing WP N and Title: WP2 - Towards shared European Guidelines for PGx Lead beneficiary:
Supplemental material to this article can be found at:
 Supplemental material to this article can be found at: http://molpharm.aspetjournals.org/content/suppl/2010/09/20/mol.110.066332.dc1 0026-895X/10/7806-1124 1134$20.00 MOLECULAR PHARMACOLOGY Vol. 78, No.
Supplemental material to this article can be found at: http://molpharm.aspetjournals.org/content/suppl/2010/09/20/mol.110.066332.dc1 0026-895X/10/7806-1124 1134$20.00 MOLECULAR PHARMACOLOGY Vol. 78, No.
Resistance to Integrase Strand Transfer Inhibitors
 NORTHWEST AIDS EDUCATION AND TRAINING CENTER Resistance to Integrase Strand Transfer Inhibitors David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases
NORTHWEST AIDS EDUCATION AND TRAINING CENTER Resistance to Integrase Strand Transfer Inhibitors David Spach, MD Clinical Director, Northwest AETC Professor of Medicine, Division of Infectious Diseases
CHAPTER 8 ONCOGENIC MARKER DETECTION FROM P53 MUTANT AMINO-ACID SUBSTITUTIONS
 134 CHAPTER 8 ONCOGENIC MARKER DETECTION FROM P53 MUTANT AMINO-ACID SUBSTITUTIONS The recent past has witnessed a rapid rise in the utilization of computational techniques to aid and accelerate biological
134 CHAPTER 8 ONCOGENIC MARKER DETECTION FROM P53 MUTANT AMINO-ACID SUBSTITUTIONS The recent past has witnessed a rapid rise in the utilization of computational techniques to aid and accelerate biological
About OMICS Group Conferences
 About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
Resistance Workshop. 3rd European HIV Drug
 3rd European HIV Drug Resistance Workshop March 30-April 1 st, 2005 Christine Hughes, PharmD Clinical Associate Professor Faculty of Pharmacy & Pharmaceutical Sciences University of Alberta Tenofovir resistance
3rd European HIV Drug Resistance Workshop March 30-April 1 st, 2005 Christine Hughes, PharmD Clinical Associate Professor Faculty of Pharmacy & Pharmaceutical Sciences University of Alberta Tenofovir resistance
Frequency(%) KRAS G12 KRAS G13 KRAS A146 KRAS Q61 KRAS K117N PIK3CA H1047 PIK3CA E545 PIK3CA E542K PIK3CA Q546. EGFR exon19 NFS-indel EGFR L858R
 Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
THE UMD TP53 MUTATION DATABASE UPDATES AND BENEFITS. Pr. Thierry Soussi
 THE UMD TP53 MUTATION DATABASE UPDATES AND BENEFITS Pr. Thierry Soussi thierry.soussi@ki.se thierry.soussi@upmc.fr TP53: 33 YEARS AND COUNTING STRUCTURE FUNCTION RELATIONSHIP OF WILD AND MUTANT TP53 1984
THE UMD TP53 MUTATION DATABASE UPDATES AND BENEFITS Pr. Thierry Soussi thierry.soussi@ki.se thierry.soussi@upmc.fr TP53: 33 YEARS AND COUNTING STRUCTURE FUNCTION RELATIONSHIP OF WILD AND MUTANT TP53 1984
Is action potential threshold lowest in the axon?
 Supplementary information to: Is action potential threshold lowest in the axon? Maarten H. P. Kole & Greg J. Stuart Supplementary Fig. 1 Analysis of action potential (AP) threshold criteria. (a) Example
Supplementary information to: Is action potential threshold lowest in the axon? Maarten H. P. Kole & Greg J. Stuart Supplementary Fig. 1 Analysis of action potential (AP) threshold criteria. (a) Example
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE
 CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
Update on Zoonotic Infections with Variant Influenza A Viruses in the USA
 Update on Zoonotic Infections with Variant Influenza A Viruses in the USA Todd Davis Zoonotic Virus Team Virology, Surveillance and Diagnosis Branch Influenza Division CDC, Atlanta OFFLU Swine Influenza
Update on Zoonotic Infections with Variant Influenza A Viruses in the USA Todd Davis Zoonotic Virus Team Virology, Surveillance and Diagnosis Branch Influenza Division CDC, Atlanta OFFLU Swine Influenza
Changing demographics of smoking and its effects during therapy
 Changing demographics of smoking and its effects during therapy Egbert F. Smit MD PhD. Dept. Pulmonary Diseases, Vrije Universiteit Medical Centre, Amsterdam, The Netherlands Smoking prevalence adults
Changing demographics of smoking and its effects during therapy Egbert F. Smit MD PhD. Dept. Pulmonary Diseases, Vrije Universiteit Medical Centre, Amsterdam, The Netherlands Smoking prevalence adults
Cellular/Molecular The Journal of Neuroscience, November 29, (48):
 12566 The Journal of Neuroscience, November 29, 2006 26(48):12566 12575 Cellular/Molecular Na v 1.7 Mutant A863P in Erythromelalgia: Effects of Altered Activation and Steady-State Inactivation on Excitability
12566 The Journal of Neuroscience, November 29, 2006 26(48):12566 12575 Cellular/Molecular Na v 1.7 Mutant A863P in Erythromelalgia: Effects of Altered Activation and Steady-State Inactivation on Excitability
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each
 Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
2/7/16. Neurons maintain a negative membrane potential. Membrane potential. Ion conductances determine the membrane potential
 Neurons maintain a negative membrane potential. V Ion channels are key regulators of membrane potential. Low Na + 2mM High K + 125mM Low Ca + (10-7 ) Low Cl - (5mM) Membrane potential. V ENa= RT/nF ln[na+]o/[na+]in
Neurons maintain a negative membrane potential. V Ion channels are key regulators of membrane potential. Low Na + 2mM High K + 125mM Low Ca + (10-7 ) Low Cl - (5mM) Membrane potential. V ENa= RT/nF ln[na+]o/[na+]in
Supplementary Figure 1: Classification scheme for non-synonymous and nonsense germline MC1R variants. The common variants with previously established
 Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Supplementary Figure 1: Classification scheme for nonsynonymous and nonsense germline MC1R variants. The common variants with previously established classifications 1 3 are shown. The effect of novel missense
Enhanced Classification of Brugada Syndrome- and Long QT Syndrome- Associated Genetic Variants in the SCN5A-Encoded Na v 1.5 Cardiac Sodium Channel
 DOI: 10.1161/CIRCGENETICS.114.000831 Enhanced Classification of Brugada Syndrome- and Long QT Syndrome- Associated Genetic Variants in the SCN5A-Encoded Na v 1.5 Cardiac Sodium Channel Running title: Kapplinger
DOI: 10.1161/CIRCGENETICS.114.000831 Enhanced Classification of Brugada Syndrome- and Long QT Syndrome- Associated Genetic Variants in the SCN5A-Encoded Na v 1.5 Cardiac Sodium Channel Running title: Kapplinger
What do we need to know about RAVs clinically?
 14 th European HIV & Hepatitis Workshop Rome, 25-27 May, 2016 What do we need to know about RAVs clinically? Stefan Zeuzem, MD University of Frankfurt Germany Background Resistance associated variants
14 th European HIV & Hepatitis Workshop Rome, 25-27 May, 2016 What do we need to know about RAVs clinically? Stefan Zeuzem, MD University of Frankfurt Germany Background Resistance associated variants
Congenital long QT syndrome of particularly malignant course connected with so far unknown mutation in the sodium channel SCN5A gene
 CASE REPORT Cardiology Journal 2013, Vol. 20, No. 1, pp. 78 82 10.5603/CJ.2013.0012 Copyright 2013 Via Medica ISSN 1897 5593 Congenital long QT syndrome of particularly malignant course connected with
CASE REPORT Cardiology Journal 2013, Vol. 20, No. 1, pp. 78 82 10.5603/CJ.2013.0012 Copyright 2013 Via Medica ISSN 1897 5593 Congenital long QT syndrome of particularly malignant course connected with
Mapping Evolutionary Pathways of HIV-1 Drug Resistance. Christopher Lee, UCLA Dept. of Chemistry & Biochemistry
 Mapping Evolutionary Pathways of HIV-1 Drug Resistance Christopher Lee, UCLA Dept. of Chemistry & Biochemistry Stalemate: We React to them, They React to Us E.g. a virus attacks us, so we develop a drug,
Mapping Evolutionary Pathways of HIV-1 Drug Resistance Christopher Lee, UCLA Dept. of Chemistry & Biochemistry Stalemate: We React to them, They React to Us E.g. a virus attacks us, so we develop a drug,
The Complexity of Simple Genetics
 The Complexity of Simple Genetics? The ciliopathies: a journey into variable penetrance and expressivity Bardet-Biedl Syndrome Allelism at a single locus is insufficient to explain phenotypic variability
The Complexity of Simple Genetics? The ciliopathies: a journey into variable penetrance and expressivity Bardet-Biedl Syndrome Allelism at a single locus is insufficient to explain phenotypic variability
A total of 2,822 Mexican dyslipidemic cases and controls were recruited at INCMNSZ in
 Supplemental Material The N342S MYLIP polymorphism is associated with high total cholesterol and increased LDL-receptor degradation in humans by Daphna Weissglas-Volkov et al. Supplementary Methods Mexican
Supplemental Material The N342S MYLIP polymorphism is associated with high total cholesterol and increased LDL-receptor degradation in humans by Daphna Weissglas-Volkov et al. Supplementary Methods Mexican
Supplementary Information
 Hyperpolarization-activated cation channels inhibit EPSPs by interactions with M-type K + channels Meena S. George, L.F. Abbott, Steven A. Siegelbaum Supplementary Information Part 1: Supplementary Figures
Hyperpolarization-activated cation channels inhibit EPSPs by interactions with M-type K + channels Meena S. George, L.F. Abbott, Steven A. Siegelbaum Supplementary Information Part 1: Supplementary Figures
Supplementary Online Content
 Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
ELECTRONIC SUPPLEMENTARY MATERIAL - Cellular and Molecular Life Sciences -
 ELECTRONIC SUPPLEMENTARY MATERIAL - Cellular and Molecular Life Sciences - Single nucleotide evolution quantifies the importance of each site along the structure of mitochondrial carriers Ciro Leonardo
ELECTRONIC SUPPLEMENTARY MATERIAL - Cellular and Molecular Life Sciences - Single nucleotide evolution quantifies the importance of each site along the structure of mitochondrial carriers Ciro Leonardo
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
Molecular Characterization of the NF2 Gene in Korean Patients with Neurofibromatosis Type 2: A Report of Four Novel Mutations
 Korean J Lab Med 2010;30:190-4 DOI 10.3343/kjlm.2010.30.2.190 Original Article Diagnostic Genetics Molecular Characterization of the NF2 Gene in Korean Patients with Neurofibromatosis Type 2: A Report
Korean J Lab Med 2010;30:190-4 DOI 10.3343/kjlm.2010.30.2.190 Original Article Diagnostic Genetics Molecular Characterization of the NF2 Gene in Korean Patients with Neurofibromatosis Type 2: A Report
NGS panels in clinical diagnostics: Utrecht experience. Van Gijn ME PhD Genome Diagnostics UMCUtrecht
 NGS panels in clinical diagnostics: Utrecht experience Van Gijn ME PhD Genome Diagnostics UMCUtrecht 93 Gene panels UMC Utrecht Cardiovascular disease (CAR) (5 panels) Epilepsy (EPI) (11 panels) Hereditary
NGS panels in clinical diagnostics: Utrecht experience Van Gijn ME PhD Genome Diagnostics UMCUtrecht 93 Gene panels UMC Utrecht Cardiovascular disease (CAR) (5 panels) Epilepsy (EPI) (11 panels) Hereditary
ERCC2mutations as predictors of response to cisplatinin bladder cancer
 ERCC2mutations as predictors of response to cisplatinin bladder cancer Eliezer (Eli) Van Allen, MD Instructor, Harvard Medical School Dana-Farber Cancer Institute Broad Institute of MIT and Harvard August
ERCC2mutations as predictors of response to cisplatinin bladder cancer Eliezer (Eli) Van Allen, MD Instructor, Harvard Medical School Dana-Farber Cancer Institute Broad Institute of MIT and Harvard August
nachr α 4 β 2 CHO Cell Line
 B SYS GmbH nachr α 4 β 2 CHO Cell Line Cell Culture Conditions B SYS GmbH B SYS GmbH nachr α 4 β 2 CHO Page 2 TABLE OF CONTENTS 1 BACKGROUND...3 1.1 Human Nicotinic Acetylcholine Receptors...3 1.2 B SYS
B SYS GmbH nachr α 4 β 2 CHO Cell Line Cell Culture Conditions B SYS GmbH B SYS GmbH nachr α 4 β 2 CHO Page 2 TABLE OF CONTENTS 1 BACKGROUND...3 1.1 Human Nicotinic Acetylcholine Receptors...3 1.2 B SYS
ULTRA-DEEP SEQUENCING OF THE BCR-ABL KINASE DOMAIN FOR BETTER THERAPEUTIC TAILORING OF PHILADELPHIA CHROMOSOME-POSITIVE LEUKEMIA PATIENTS
 ULTRA-DEEP SEQUENCING OF THE BCR-ABL KINASE DOMAIN FOR BETTER THERAPEUTIC TAILORING OF PHILADELPHIA CHROMOSOME-POSITIVE LEUKEMIA PATIENTS Simona Soverini, PhD Department of Experimental, Diagnostic and
ULTRA-DEEP SEQUENCING OF THE BCR-ABL KINASE DOMAIN FOR BETTER THERAPEUTIC TAILORING OF PHILADELPHIA CHROMOSOME-POSITIVE LEUKEMIA PATIENTS Simona Soverini, PhD Department of Experimental, Diagnostic and
Kidney. Heart. Lung. Sirt1. Gapdh. Mouse IgG DAPI. Rabbit IgG DAPI
 a e Na V 1.5 Ad-LacZ Ad- 110KD b Scn5a/ (relative to Ad-LacZ) f 150 100 50 0 p = 0.65 Ad-LacZ Ad- c Heart Lung Kidney Spleen 110KD d fl/fl c -/- DAPI 20 µm Na v 1.5 250KD fl/fl Rabbit IgG DAPI fl/fl Mouse
a e Na V 1.5 Ad-LacZ Ad- 110KD b Scn5a/ (relative to Ad-LacZ) f 150 100 50 0 p = 0.65 Ad-LacZ Ad- c Heart Lung Kidney Spleen 110KD d fl/fl c -/- DAPI 20 µm Na v 1.5 250KD fl/fl Rabbit IgG DAPI fl/fl Mouse
Mechanisms of cold sensitivity of paramyotonia congenita mutation R1448H and overlap syndrome mutation M1360V
 J Physiol (2003), 547.3, pp. 691 698 DOI: 10.1113/jphysiol.2002.033928 The Physiological Society 2003 www.jphysiol.org Mechanisms of cold sensitivity of paramyotonia congenita mutation R1448H and overlap
J Physiol (2003), 547.3, pp. 691 698 DOI: 10.1113/jphysiol.2002.033928 The Physiological Society 2003 www.jphysiol.org Mechanisms of cold sensitivity of paramyotonia congenita mutation R1448H and overlap
The role of E148Q in FMF. Elon Pras Institute of Human Genetics Sheba Medical Center
 The role of E148Q in FMF Elon Pras Institute of Human Genetics Sheba Medical Center Familial Mediterranean Fever (FMF) Acute attacks of fever accompanied by: Peritonitis Pleuritis Arthritis Erysipelas
The role of E148Q in FMF Elon Pras Institute of Human Genetics Sheba Medical Center Familial Mediterranean Fever (FMF) Acute attacks of fever accompanied by: Peritonitis Pleuritis Arthritis Erysipelas
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
6/12/2018. Disclosures. Clinical Genomics The CLIA Lab Perspective. Outline. COH HopeSeq Heme Panels
 Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Genetic Predictors of Radiosensitivity.
 Genetic Predictors of Radiosensitivity. Richard G. Stock, MD Professor, Director of Genito-Urinary Oncology Department of Radiation Oncology Ichan School of Medicine at Mount Sinai New York, NY DISCLOSURE
Genetic Predictors of Radiosensitivity. Richard G. Stock, MD Professor, Director of Genito-Urinary Oncology Department of Radiation Oncology Ichan School of Medicine at Mount Sinai New York, NY DISCLOSURE
Study Design - GT 1 Retreatment
 Retreatment of Patients Who Failed 8 or 12 Weeks of Ledipasvir/Sofosbuvir-Based Regimens With Ledipasvir/Sofosbuvir for 24 Weeks Eric Lawitz, Steven Flamm, Jenny C. Yang, Phillip S. Pang, Yanni Zhu, Evguenia
Retreatment of Patients Who Failed 8 or 12 Weeks of Ledipasvir/Sofosbuvir-Based Regimens With Ledipasvir/Sofosbuvir for 24 Weeks Eric Lawitz, Steven Flamm, Jenny C. Yang, Phillip S. Pang, Yanni Zhu, Evguenia
Paroxysmal extreme pain disorder mutations within the D3/S4 S5 linker of Nav1.7 cause moderate destabilization of fast inactivation
 J Physiol 586.17 (2008) pp 4137 4153 4137 Paroxysmal extreme pain disorder mutations within the D3/S4 S5 linker of Nav1.7 cause moderate destabilization of fast inactivation Brian W. Jarecki, Patrick L.
J Physiol 586.17 (2008) pp 4137 4153 4137 Paroxysmal extreme pain disorder mutations within the D3/S4 S5 linker of Nav1.7 cause moderate destabilization of fast inactivation Brian W. Jarecki, Patrick L.
Support. Overview. Auditory Dys-synchrony. Auditory Brainstem Response. Potential Causes
 Potential Role of Genetic Testing in Auditory Neuropathy/Dys-synchrony Christina Runge-Samuelson, Ph.D., CCC-A Associate Professor Co-Director, Koss Cochlear Implant Program Department of tolaryngology
Potential Role of Genetic Testing in Auditory Neuropathy/Dys-synchrony Christina Runge-Samuelson, Ph.D., CCC-A Associate Professor Co-Director, Koss Cochlear Implant Program Department of tolaryngology
(43) Publication date: 04 September 2014 ( ) (22) Filing Date: 27 February 2014 ( )
 (54) Title (EN): INCOHERENT TYPE-III MATERIALS FOR CHARGE CARRIERS CONTROL DEVICES (54) Title (FR): MATÉRIAUX DE TYPE III INCOHÉRENTS POUR DISPOSITIFS DE RÉGULATION DE PORTEURS DE CHARGES (72) Inventor(s):
(54) Title (EN): INCOHERENT TYPE-III MATERIALS FOR CHARGE CARRIERS CONTROL DEVICES (54) Title (FR): MATÉRIAUX DE TYPE III INCOHÉRENTS POUR DISPOSITIFS DE RÉGULATION DE PORTEURS DE CHARGES (72) Inventor(s):
Case presentations from diagnostic exome sequencing results in ion channels M. Koko, U. Hedrich, H. Lerche DASNE meeting 2017, Eisenach
 Case presentations from diagnostic exome sequencing results in ion channels M. Koko, U. Hedrich, H. Lerche DASNE meeting 2017, Eisenach CASE 1: Clinical picture Patient s presentation: 38 year old jewish
Case presentations from diagnostic exome sequencing results in ion channels M. Koko, U. Hedrich, H. Lerche DASNE meeting 2017, Eisenach CASE 1: Clinical picture Patient s presentation: 38 year old jewish
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #2
 Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #2 1. (PeiXi) You are performing research on a novel ion channel and want to learn some of its characteristics. a) When you conducted voltage clamp
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #2 1. (PeiXi) You are performing research on a novel ion channel and want to learn some of its characteristics. a) When you conducted voltage clamp
Clinical Genetics. Functional validation in a diagnostic context. Robert Hofstra. Leading the way in genetic issues
 Clinical Genetics Leading the way in genetic issues Functional validation in a diagnostic context Robert Hofstra Clinical Genetics Leading the way in genetic issues Future application of exome sequencing
Clinical Genetics Leading the way in genetic issues Functional validation in a diagnostic context Robert Hofstra Clinical Genetics Leading the way in genetic issues Future application of exome sequencing
An international compendium of mutations in the SCN5A-encoded cardiac sodium channel in patients referred for Brugada syndrome genetic testing
 An international compendium of mutations in the SCN5A-encoded cardiac sodium channel in patients referred for Brugada syndrome genetic testing Jamie D. Kapplinger, BA,* David J. Tester, BS,* Marielle Alders,
An international compendium of mutations in the SCN5A-encoded cardiac sodium channel in patients referred for Brugada syndrome genetic testing Jamie D. Kapplinger, BA,* David J. Tester, BS,* Marielle Alders,
Mutational and phenotypical spectrum of phenylalanine hydroxylase deficiency in Denmark
 Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Received 21 November 2005/Returned for modification 20 January 2006/Accepted 27 March 2006
 JOURNAL OF CLINICAL MICROBIOLOGY, June 2006, p. 1930 1943 Vol. 44, No. 6 0095-1137/06/$08.00 0 doi:10.1128/jcm.02415-05 Copyright 2006, American Society for Microbiology. All Rights Reserved. Evaluation
JOURNAL OF CLINICAL MICROBIOLOGY, June 2006, p. 1930 1943 Vol. 44, No. 6 0095-1137/06/$08.00 0 doi:10.1128/jcm.02415-05 Copyright 2006, American Society for Microbiology. All Rights Reserved. Evaluation
Cross-talk between Two Partners Regulates the Gating of the K ATP Channel Pore
 International Conference on Metabolic Syndromes Cross-talk between Two Partners Regulates the Gating of the K ATP Channel Pore Dr Hussein N Rubaiy h.rubaiy@leeds.ac.uk School of Medicine, University of
International Conference on Metabolic Syndromes Cross-talk between Two Partners Regulates the Gating of the K ATP Channel Pore Dr Hussein N Rubaiy h.rubaiy@leeds.ac.uk School of Medicine, University of
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation
 Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Construction of a hepatocellular carcinoma cell line that stably expresses stathmin with a Ser25 phosphorylation site mutation J. Du 1, Z.H. Tao 2, J. Li 2, Y.K. Liu 3 and L. Gan 2 1 Department of Chemistry,
Kalydeco. Kalydeco (ivacaftor) Description
 Federal Employee Program 1310 G Street, N.W. Washington, D.C. 20005 202.942.1000 Fax 202.942.1125 5.45.03 Subject: Kalydeco Page: 1 of 6 Last Review Date: November 30, 2018 Kalydeco Description Kalydeco
Federal Employee Program 1310 G Street, N.W. Washington, D.C. 20005 202.942.1000 Fax 202.942.1125 5.45.03 Subject: Kalydeco Page: 1 of 6 Last Review Date: November 30, 2018 Kalydeco Description Kalydeco
WO 2012/ A3. 15 November 2012 ( ) P O P C T
 (12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International
screening procedures Disease resistant to full-dose imatinib ( 600 mg/day) or intolerant to any dose of imatinib
 Table S1. Study inclusion and exclusion criteria Inclusion criteria Aged 18 years Signed and dated informed consent form prior to protocol-specific screening procedures Cytogenetic- or PCR-based diagnosis
Table S1. Study inclusion and exclusion criteria Inclusion criteria Aged 18 years Signed and dated informed consent form prior to protocol-specific screening procedures Cytogenetic- or PCR-based diagnosis
Medical Policy An independent licensee of the Blue Cross Blue Shield Association
 Cystic Fibrosis Transmembrane Page 1 of 11 Medical Policy An independent licensee of the Blue Cross Blue Shield Association Title: Cystic Fibrosis Transmembrane Conductance Regulator (CFTR) Prime Therapeutics
Cystic Fibrosis Transmembrane Page 1 of 11 Medical Policy An independent licensee of the Blue Cross Blue Shield Association Title: Cystic Fibrosis Transmembrane Conductance Regulator (CFTR) Prime Therapeutics
Monitoring for Drug Resistance by Genotyping. Urvi M Parikh, PhD MTN Virology Core Lab
 Monitoring for Drug Resistance by Genotyping Urvi M Parikh, PhD MTN Virology Core Lab Outline What is Drug Resistance? Genotyping Algorithm Standard vs Sensitive Resistance Testing Sequencing Protocols
Monitoring for Drug Resistance by Genotyping Urvi M Parikh, PhD MTN Virology Core Lab Outline What is Drug Resistance? Genotyping Algorithm Standard vs Sensitive Resistance Testing Sequencing Protocols
Spherical Bearings Heavy Duty Equipments
 Spherical Bearings Heavy Duty Equipments Highlights Quality Service Price Wbf Replacement Parts adaptableto > Caterpillar >Komatsu >Volvo 1 WBF SPHERICAL BEARINGS adaptable to Caterpillar Part No. Description
Spherical Bearings Heavy Duty Equipments Highlights Quality Service Price Wbf Replacement Parts adaptableto > Caterpillar >Komatsu >Volvo 1 WBF SPHERICAL BEARINGS adaptable to Caterpillar Part No. Description
SUPPLEMENTARY INFORMATION. Divergent TLR7/9 signaling and type I interferon production distinguish
 SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
Potassium Channelopathies: Consequences and Impact on Treatment December 4, 2010
 Potassium Channelopathies: Consequences and Impact on Treatment December 4, 2010 Karen S. Wilcox, Ph.D. Department of Pharmacology & Toxicology Anticonvulsant Drug Development Program University of Utah
Potassium Channelopathies: Consequences and Impact on Treatment December 4, 2010 Karen S. Wilcox, Ph.D. Department of Pharmacology & Toxicology Anticonvulsant Drug Development Program University of Utah
LIPOPROTEINE ATEROGENE E ANTI-ATEROGENE ATEROGENE
 LIPOPROTEINE ATEROGENE E ANTI-ATEROGENE ATEROGENE Sebastiano Calandra Dipartimento di Scienze Biomediche Università di Modena e Reggio Emilia Incidence Rate/1000 200-150 - 100-50 - Women 0 Men
LIPOPROTEINE ATEROGENE E ANTI-ATEROGENE ATEROGENE Sebastiano Calandra Dipartimento di Scienze Biomediche Università di Modena e Reggio Emilia Incidence Rate/1000 200-150 - 100-50 - Women 0 Men
supplementary information
 Figure S1 Nucleotide binding status of RagA mutants. Wild type and mutant forms of MycRagA was transfected into HEK293 cells and the transfected cells were labeled with 32 Pphosphate. MycRagA was immunoprecipitated
Figure S1 Nucleotide binding status of RagA mutants. Wild type and mutant forms of MycRagA was transfected into HEK293 cells and the transfected cells were labeled with 32 Pphosphate. MycRagA was immunoprecipitated
Cellular Neurobiology BIPN140. 1st Midterm Exam October 18 th, Tuesday Material covered: Lectures 1-6 & Reading
 Cellular Neurobiology BIPN140 1st Midterm Exam October 18 th, Tuesday Material covered: Lectures 1-6 & Reading Review session October 17 th 3500 Pacitic Hall, 6-8 pm (access code is 127895) Come with questions!
Cellular Neurobiology BIPN140 1st Midterm Exam October 18 th, Tuesday Material covered: Lectures 1-6 & Reading Review session October 17 th 3500 Pacitic Hall, 6-8 pm (access code is 127895) Come with questions!
P G K R P E W M G W L K P R G G A V N Y A R P L Q G R V T M T R D V Y S D T A F
 Supplementary Figure 1 VRC01 45-08-110497H 45-08-212510H 45-08-511533H 45-08-541880H Q V Q L V Q S G G Q M K K P G E S M R I S C R A S G Y E F I D C T L N W I R L A CAGGTGCAGCTGGTGCAGTCTGGGGGTCAGATGAAGAAGCCTGGCGAGTCGATGAGAATTTCTTGTCGGGCTTCTGGATATGAATTTATTGATTGTACGCTAAATTGGATTCGTCTGGCC
Supplementary Figure 1 VRC01 45-08-110497H 45-08-212510H 45-08-511533H 45-08-541880H Q V Q L V Q S G G Q M K K P G E S M R I S C R A S G Y E F I D C T L N W I R L A CAGGTGCAGCTGGTGCAGTCTGGGGGTCAGATGAAGAAGCCTGGCGAGTCGATGAGAATTTCTTGTCGGGCTTCTGGATATGAATTTATTGATTGTACGCTAAATTGGATTCGTCTGGCC
Genetics of Sudden Cardiac Death. Geoffrey Pitt Ion Channel Research Unit Duke University. Disclosures: Grant funding from Medtronic.
 Genetics of Sudden Cardiac Death Geoffrey Pitt Ion Channel Research Unit Duke University Disclosures: Grant funding from Medtronic Duke U N I V E R S I T Y Sudden Cardiac Death High incidence 50-100 per
Genetics of Sudden Cardiac Death Geoffrey Pitt Ion Channel Research Unit Duke University Disclosures: Grant funding from Medtronic Duke U N I V E R S I T Y Sudden Cardiac Death High incidence 50-100 per
iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers).
 Supplementary Materials Supplementary Methods iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers). IDH1 R132H and R132L Forward:
Supplementary Materials Supplementary Methods iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers). IDH1 R132H and R132L Forward:
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/317/5841/183/dc1 Supporting Online Material for Astrocytes Potentiate Transmitter Release at Single Hippocampal Synapses Gertrudis Perea and Alfonso Araque* *To whom
www.sciencemag.org/cgi/content/full/317/5841/183/dc1 Supporting Online Material for Astrocytes Potentiate Transmitter Release at Single Hippocampal Synapses Gertrudis Perea and Alfonso Araque* *To whom
The E1784K mutation in SCN5A is associated with mixed clinical phenotype of type 3 long QT syndrome
 Research article The E1784K mutation in SCN5A is associated with mixed clinical phenotype of type 3 long QT syndrome Naomasa Makita, 1 Elijah Behr, 2 Wataru Shimizu, 3 Minoru Horie, 4 Akihiko Sunami, 5
Research article The E1784K mutation in SCN5A is associated with mixed clinical phenotype of type 3 long QT syndrome Naomasa Makita, 1 Elijah Behr, 2 Wataru Shimizu, 3 Minoru Horie, 4 Akihiko Sunami, 5
ICAAC/IDSA DC, Oct. 26, 2008
 Tenofovir (TDF)- or Abacavir (ABC)-selected Minority Subpopulations in Viremic Subjects Detected by Ultra-deep Sequencing R. T. D Aquila 1, E. Rouse 2, J. Horton 2, A. Kheshti 1,3, S. Raffanti 1,3, K.
Tenofovir (TDF)- or Abacavir (ABC)-selected Minority Subpopulations in Viremic Subjects Detected by Ultra-deep Sequencing R. T. D Aquila 1, E. Rouse 2, J. Horton 2, A. Kheshti 1,3, S. Raffanti 1,3, K.
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Functional Effects of Two Voltage-Gated Sodium Channel Mutations That Cause Generalized Epilepsy with Febrile Seizures Plus Type 2
 The Journal of Neuroscience, October 1, 2001, 21(19):7481 7490 Functional Effects of Two Voltage-Gated Sodium Channel Mutations That Cause Generalized Epilepsy with Febrile Seizures Plus Type 2 Jay Spampanato,
The Journal of Neuroscience, October 1, 2001, 21(19):7481 7490 Functional Effects of Two Voltage-Gated Sodium Channel Mutations That Cause Generalized Epilepsy with Febrile Seizures Plus Type 2 Jay Spampanato,
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
The State of the Molecular Autopsy for Sudden Death in the Young
 The State of the Molecular Autopsy for Sudden Death in the Young Michael J. Ackerman, MD, PhD Windland Smith Rice Cardiovascular Genomics Research Professor Professor of Medicine, Pediatrics, and Pharmacology
The State of the Molecular Autopsy for Sudden Death in the Young Michael J. Ackerman, MD, PhD Windland Smith Rice Cardiovascular Genomics Research Professor Professor of Medicine, Pediatrics, and Pharmacology
Resistencias & Epidemiología. Eva Poveda Division of Clinical Virology INIBIC-Complexo Hospitalario Universitario de A Coruña
 Resistencias & Epidemiología Eva Poveda Division of Clinical Virology INIBIC-Complexo Hospitalario Universitario de A Coruña Rapid Evolution of HCV Regimens: Easier to take/tolerate, Short Duration, Pangenotypic,
Resistencias & Epidemiología Eva Poveda Division of Clinical Virology INIBIC-Complexo Hospitalario Universitario de A Coruña Rapid Evolution of HCV Regimens: Easier to take/tolerate, Short Duration, Pangenotypic,
Amino Acid Substitutions in the Pore Helix of GluR6 Control Inhibition by Membrane Fatty Acids
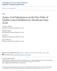 Washington University School of Medicine Digital Commons@Becker Open Access Publications 2008 Amino Acid Substitutions in the Pore Helix of GluR6 Control Inhibition by Membrane Fatty Acids Timothy J. Wilding
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2008 Amino Acid Substitutions in the Pore Helix of GluR6 Control Inhibition by Membrane Fatty Acids Timothy J. Wilding
