Prediction of Post-translational modification sites and Multi-domain protein structure
|
|
|
- Cuthbert Harrell
- 5 years ago
- Views:
Transcription
1 Prediction of Post-translational modification sites and Multi-domain protein structure Robert Newman, Ph.D. Department of Biology Dukka KC, Ph.D. Department of Computational Science and Engineering North Carolina A&T State University Greensboro, NC USA
2 A Brief Overview of Cell Signaling The Akt/PKB Signaling Pathway EMD Calbiochem
3 The Human Kinome Manning et al. (2002)
4 The Human Kinome Manning et al. (2002)
5 The Human Phosphorylome CELLULAR PROTEINS Manning et al. (2002)
6 The Human Phosphorylome CELLULAR PROTEINS Manning et al. (2002)
7 The Human Phosphorylome CELLULAR PROTEINS Manning et al. (2002)
8 The Human Phosphorylome CELLULAR PROTEINS Known P sites: >100,000 Known KSRs: <2,000 Phosite PLUS (Hornbeck et al. (2012))
9
10 Crosstalk Between Cellular Signaling Pathways ROS Signals H2O2 NO O2 Src Akt1 Adapted from Bhattacharyya (2014)
11 Central Theme of the lab Central Theme Elucidation of Protein sequence Structure Function Evolution relationship using various computational approaches Past Projects From sequence to structure Clique based algorithm for protein side-chain packing Refinement of protein structure (KC et al. 2002) From Sequence/structure to function Improvement of protein functional site prediction using phylogenetic motifs and the MINER software (KC, Livesay 2009) From structure to evolution Internal symmetry detection in protein structure (NAR,2014) Evolution of symmetric proteins
12 Outline of the Talk Protein post-translation modification Site prediction Feature Extraction from Protein Sequences (FEPS) Protein Phosphorylation Site Prediction (RF-Phos) Protein Hydroxylation Site Prediction (RF-hydroxysite)
13 Protein Post-translation modification site prediction Post-translational modification (PTM): Chemical modifications occurring in proteins after they are synthesized by ribosomes ( translating mrna into polypeptide chains) PTM increases the functional diversity of the proteome and influence almost all aspects of cell biology. Some PTMS: phosphorylation, hydroxylation, glycosylation, methylation, etc. Identifying and understanding PTMs (as well as sites) is critical in the study of cell biology.
14 Protein Post-translation modification site prediction Discuss our Random Forest based approaches for two of the PTMs site prediction approach RF-Phos: Phosphorylation site prediction approach (Ismail et.al, 2016) RF-Hydroxysite: Hydroxylation site prediction approach (Ismail et al. 2016) Critical thing is extracting the features (numeric values) from protein sequences for Machine Learning Based Approaches
15 FEPS: A tool for feature extraction from protein sequences Ismail, Smith, KC, SubmiLed.
16 FEPS: A tool for feature extraction from protein sequences - Motivation Existing methods: PseAAC (Chou et al, 2008): Computes only one types of features. PROFEAT(Zhu et al. 2011): (10 features) SPiCE (Reinders et al. 2014) ) (18 features) A new tool is required: more features Flexible Easy-to-use With different output formats
17 FEPS: Feature Extraction From Protein Sequence Developed FEPS: web-based bioinformatics tool for sequence-based protein feature extraction available at bcb.ncat.edu/features/ (In Submission) Developed under linux with python + django + JavaScript + mysql. Uses 46 published methods; 6 use any one of the 544 physicochemical properties of amino acids so it makes total of 3304 of different features. Four feature types accept user-defined amino acid indices that make the number of features unlimited. 17
18 Types of Features in FEPS 18
19 FEPS: A tool for feature extraction from protein sequences bcb.ncat.edu/features
20 FEPS: A tool for feature extraction from protein sequences 1- Input Input can be a single files or multiple fasta files Browse to input files Input files
21 FEPS: A tool for feature extraction from protein sequences 2- Feature groups Select one feature type at a time Seven types each with a group of methods Select a method for the list
22 FEPS: A tool for feature extraction from protein sequences 3- Feature type options Tweak feature options
23 FEPS: A tool for feature extraction from protein sequences 3- Physicochemical properties AAindex database Select an amino acid property out of 544
24 FEPS: A tool for feature extraction from protein sequences 3- User-defined indices Amino acid properties User-defined indices
25 FEPS: A tool for feature extraction from protein sequences - Motivation 4- Output options Four file formats are available File formats
26 RF-Phos: Random Forest based protein phosphorylation Site prediction RF-Phos: A Novel General Phosphorylation Site Prediction Tool Based on Random Forest. Hamid D. Ismail, Ahoi Jones, Jung H. Kim, Robert H. Newman, Dukka B. KC., Biomed Res. Intl, ID: ,2016.
27 Phosphorylation Protein synthesis DNA RNA Protein Phosphorylation Addition of a phosphate group to an amino acid residue (Ser, Thr, Tyr) 27
28 Phosphorylation One third of the proteins in the human proteome are substrates for phosphorylation Regulatory functions: Enzymes and receptors are switched off and on Conformational change in structure of enzymes and receptors (activated and deactivated) Plays critical roles in cell cycle, growth, apoptosis and signal transduction pathways. One of the most important post-translational modification sites. Hence, identification of Phosphorylation sites is very important. 28
29 Random Forest: An ensemble classifier Original training data Random Forest (RF) [Breiman et al., 1984] (1) Random bootstrap samples (with replacement) (2) Build decision trees (3) Combine trees
30 Existing methods for Phosphorylation Site Prediction NetPhosK[Blom et al., 2004], GPS2.1 [Xue et al., 2008], NetPhos[Blom et al., 1999], PPRED [Biswas et al., 2010], Musite[Gao et al., 2010], and PhosphoSVM[Dou et al. 2014]. Methods Technique Year NetPhosK ANN 2005 GPS 2.1 PSSM/ GA 2005 NetPhos ANN 2005 Musite SVM 2010 PhosphoSVM SVM 2014 A new method with better performance is highly required.
31 RF-Phos: Random Forest based protein phosphorylation Site prediction Annotated protein sequences with known phosphorylated Ser/Thr/Tyr residues were obtained from P.ELM database [Dinkel et al, 2011]. Annotated fasta format sequence >O S CHK1 86 T CDK2 99 Y ab1 MAQSTATSPDGGTTFEHLWSSLEPDSTYFDLPQSSRG NNEVVGGTDSSMDVFHLEGMTTSVMAQFNLLSSTM DQMSSRAASASPYTPEHAASVPTHSPYAQPSSTF
32 Phosphorylation Site Prediction Redundant sequences (with similarity > 30%) were removed with CD-HIT [Weizhong et al., 2006]. Sliding windows of size 7, 9, 11, 13, 15, 17, 19, 21 were generated from the sequences XXXXXXXXXXXXXXXXX Positive: The windows with experimentally characterized (or +ve) S/T/Y in the middle. XXXX[S/T/Y]XXXX Negative: The windows with -ve S/T/Y in the middle. XXXX[S/T/Y]XXXX 32
33 Phosphorylation Site Prediction: Benchmark Dataset Redundant sequence windows were removed from both +ve windows and ve windows. The remaining windows were used to construct the random forest model. Residue Positive windows (size 9) Negative Before After Used S 20,577 1,554 1,543 T 5, Y 2,
34 Feature Extraction with FEPS Eight types of features were extracted i- Shannon entropy (1 feature) ii- Relative entropy (1feature) iii- Information gain (1 feature) iv- Accessible surface area (9 features) v- Overlapping Properties (90 features) vi- Average cumulative hydrophobicity (4 features) vii- Sequence Features (180 features) viii- Composition, Transition and Distribution (147 features) 34
35 RF- Phos: Overall Algorithm
36 Comparison Metrics (Accuracy) (Precision) (Specificity) (Sensitivity) (F1-score) (Matthew s correlation coefficient)
37 RF- Phos: Random Forest based Predictor for Protein Phosphorylation Sites Comparing RF-Phos to the other methods (10-fold cross validation) Methods Tyrosine AUC Sen (%) Sp (%) MCC NetPhosK GPS Swaminathan NetPhos PPRED Musite PhosphoSVM RF- Phos
38 Phosphorylation Site Prediction Top 10 One of the salient feature of Random Forest is that you can annotate the importance of features 38
39 RF- Phos: Random Forest based Predictor for Protein Phosphorylation Sites
40 RF- Phos: Random Forest based Predictor for Protein Phosphorylation Sites Job ID Download link Predicted sites Annotated sequence
41 RF-Phos: Random Forest based Predictor for Protein Phosphorylation Sites Output screen Output text file
42 RF-Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites Ismail, Newman and KC, RF-Hydroxysite: a random forest based predictor for hydroxylation sites, Molecular Biosystems, 12, , 2016.
43 RF- Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites Addition of OH (Hydroxyl group) Hydroxylase Proline Hydroxyproline Hydroxylase Lysine Hydroxylysine Essential elements of collagen and connective tissues Instability is associated with cancers
44 RF- Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites Existing methods and motivation add reference SVM-based [Hu et al, 2010], ihyd-pseaac[xu et al, 2014], PredHydroxy [Shi, S.P., et al., 2015] Accuracy Methodology Method P K Feature Algorithm SVM- based PSSM SVM ihyd- PseAAC PseAAC LDA PredHydroxy PWAAC + HQI SVM The previous studies found that the evolutionary and physicochemical information are important. There is still room for improvement
45 RF- Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites Position weight amino acid composition (PWAA) High quality physicochemical indices (HQI) Type I entropy Overlapping properties (OP) Average cumulative hydrophobicity Protein disordered region Type II entropy Feature Extraction using FEPS
46 RF- Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites Feature selection (selected ü ) Features P K Position weight amino acid (PWAA) û û Propensity (HQI1) ü û Solvent accessibility (HQI2) û û Alpha-helix frequency (HQI3) ü ü Crystallographic waters (HQI4) ü ü Amino acid composition of MEM (HQI5) ü û Composition of AA in intracellular (HQI6) û û Conformational preference (HQI7) ü û Partition energies (HQI8) ü û Type I entropy (ENTI) ü û Overlapping properties (OP) û û Average cumulative hydrophobicity (ACH) ü ü Protein disordered region (PDR) û û ENTII (Type II entropy) û û
47 RF- Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites Evaluation results (Jackknife) for different window sizes (lysine and proline) 0.80 K P K P K P K P K P K P K P Accu Prec Sens Spec F1sc MCC Residues Window size
48 RF- Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites Evaluation results (Independent sample) K P K P K P K P K P K P K P Accu Prec Sens Spec F1sc MCC AUC
49 RF- Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites Evaluation results Side-by-side comparison (Independent sample window size=15) PredHydroxy RF-Hydroxysite Metrics P L P L Accuracy Sensitivity Specificity MCC
50 RF- Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites RF-Hydroxysite
51 RF- Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites
52 RF- Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites Job id Download link Annotated sequence
53 RF- Hydroxysite: Random Forest based Predictor for Protein Hydroxylation Sites Text file
54 Acknowledgements NCAT Startup Fund NSF Beacon Grant
LIPOPREDICT: Bacterial lipoprotein prediction server
 www.bioinformation.net Server Volume 8(8) LIPOPREDICT: Bacterial lipoprotein prediction server S Ramya Kumari, Kiran Kadam, Ritesh Badwaik & Valadi K Jayaraman* Centre for Development of Advanced Computing
www.bioinformation.net Server Volume 8(8) LIPOPREDICT: Bacterial lipoprotein prediction server S Ramya Kumari, Kiran Kadam, Ritesh Badwaik & Valadi K Jayaraman* Centre for Development of Advanced Computing
Protein Modification Overview DEFINITION The modification of selected residues in a protein and not as a component of synthesis
 Lecture Four: Protein Modification & Cleavage [Based on Chapters 2, 9, 10 & 11 Berg, Tymoczko & Stryer] (Figures in red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Protein Modification
Lecture Four: Protein Modification & Cleavage [Based on Chapters 2, 9, 10 & 11 Berg, Tymoczko & Stryer] (Figures in red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Protein Modification
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A
 Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Steps at which eukaryotic gene expression can be controlled. Cell 7.5
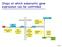 Steps at which eukaryotic gene expression can be controlled Cell 7.5 Protein Variability and Protein Activity Control Aminoacid sequence Three-dimensional shape (conformation) Function Protein processing
Steps at which eukaryotic gene expression can be controlled Cell 7.5 Protein Variability and Protein Activity Control Aminoacid sequence Three-dimensional shape (conformation) Function Protein processing
Ionization of amino acids
 Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
Biochemistry Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology Kharagpur. Lecture -02 Amino Acids II
 Biochemistry Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology Kharagpur Lecture -02 Amino Acids II Ok, we start off with the discussion on amino acids. (Refer Slide Time: 00:48)
Biochemistry Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology Kharagpur Lecture -02 Amino Acids II Ok, we start off with the discussion on amino acids. (Refer Slide Time: 00:48)
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16
 38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
Data mining with Ensembl Biomart. Stéphanie Le Gras
 Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Mal-Lys: prediction of lysine malonylation sites in proteins integrated sequence-based features with mrmr feature selection
 R A P Mal-Lys: prediction of lysine malonylation sites in proteins integrated sequence-based features with mrmr feature selection Yan Xu, Ya-Xin Ding, Jun Ding, Ling-Yun Wu & Yu Xue characterized to be
R A P Mal-Lys: prediction of lysine malonylation sites in proteins integrated sequence-based features with mrmr feature selection Yan Xu, Ya-Xin Ding, Jun Ding, Ling-Yun Wu & Yu Xue characterized to be
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
RNA Secondary Structures: A Case Study on Viruses Bioinformatics Senior Project John Acampado Under the guidance of Dr. Jason Wang
 RNA Secondary Structures: A Case Study on Viruses Bioinformatics Senior Project John Acampado Under the guidance of Dr. Jason Wang Table of Contents Overview RSpredict JAVA RSpredict WebServer RNAstructure
RNA Secondary Structures: A Case Study on Viruses Bioinformatics Senior Project John Acampado Under the guidance of Dr. Jason Wang Table of Contents Overview RSpredict JAVA RSpredict WebServer RNAstructure
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
About This Chapter. Hormones The classification of hormones Control of hormone release Hormone interactions Endocrine pathologies Hormone evolution
 About This Chapter Hormones The classification of hormones Control of hormone release Hormone interactions Endocrine pathologies Hormone evolution Hormones: Function Control Rates of enzymatic reactions
About This Chapter Hormones The classification of hormones Control of hormone release Hormone interactions Endocrine pathologies Hormone evolution Hormones: Function Control Rates of enzymatic reactions
Complexity DNA. Genome RNA. Transcriptome. Protein. Proteome. Metabolites. Metabolome
 DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
The Basics: A general review of molecular biology:
 The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
Lecture 12 (10/11/17) Lecture 12 (10/11/17)
 Lecture 12 (10/11/17) Reading: Ch1; 27-29 Ch5; 157-158, 160-161 Problems: Ch1 (text); 16 Ch4 (text); 1, 2, 3, 4, 6, 7, 8 NEXT Reading: Ch6; 187-189, 204-205 Problems: Ch4 (text); 2, 3 OUTLINE Protein Characterization
Lecture 12 (10/11/17) Reading: Ch1; 27-29 Ch5; 157-158, 160-161 Problems: Ch1 (text); 16 Ch4 (text); 1, 2, 3, 4, 6, 7, 8 NEXT Reading: Ch6; 187-189, 204-205 Problems: Ch4 (text); 2, 3 OUTLINE Protein Characterization
Short polymer. Dehydration removes a water molecule, forming a new bond. Longer polymer (a) Dehydration reaction in the synthesis of a polymer
 HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
Biomolecules: amino acids
 Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
FurinDB: A Database of 20-Residue Furin Cleavage Site Motifs, Substrates and Their Associated Drugs
 Int. J. Mol. Sci. 2011, 12, 1060-1065; doi:10.3390/ijms12021060 OPEN ACCESS Technical Note International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms FurinDB: A Database of 20-Residue
Int. J. Mol. Sci. 2011, 12, 1060-1065; doi:10.3390/ijms12021060 OPEN ACCESS Technical Note International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms FurinDB: A Database of 20-Residue
Previous Class. Today. Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism. Protein Kinase Catalytic Properties
 Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
Chapter 5: The Structure and Function of Large Biological Molecules
 Chapter 5: The Structure and Function of Large Biological Molecules 1. Name the four main classes of organic molecules found in all living things. Which of the four are classified as macromolecules. Define
Chapter 5: The Structure and Function of Large Biological Molecules 1. Name the four main classes of organic molecules found in all living things. Which of the four are classified as macromolecules. Define
CoCoLysis: A Web-Accessible Coiled-Coil Protein Database with Analysis Tools
 CoCoLysis: A Web-Accessible Coiled-Coil Protein Database with Analysis Tools David Brinkmann, Sai Nandoor, Jugal Kalita, Brian Tripet AND Robert Hodges sainandoor@hotmail.com, david.brinkmann@hp.com, kalita@pikespeak.uccs.edu
CoCoLysis: A Web-Accessible Coiled-Coil Protein Database with Analysis Tools David Brinkmann, Sai Nandoor, Jugal Kalita, Brian Tripet AND Robert Hodges sainandoor@hotmail.com, david.brinkmann@hp.com, kalita@pikespeak.uccs.edu
Post-translational modifications of proteins in gene regulation under hypoxic conditions
 203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
SUPPLEMENTARY MATERIAL S-1 INTREPID VARIANTS
 SUPPLEMENTARY MATERIAL S- INTREPID VARIANTS We define different INTREPID variants based on the positional conservation score cons(s, x) which is used to compute the importance score in Equation S-. IMP
SUPPLEMENTARY MATERIAL S- INTREPID VARIANTS We define different INTREPID variants based on the positional conservation score cons(s, x) which is used to compute the importance score in Equation S-. IMP
Page 8/6: The cell. Where to start: Proteins (control a cell) (start/end products)
 Page 8/6: The cell Where to start: Proteins (control a cell) (start/end products) Page 11/10: Structural hierarchy Proteins Phenotype of organism 3 Dimensional structure Function by interaction THE PROTEIN
Page 8/6: The cell Where to start: Proteins (control a cell) (start/end products) Page 11/10: Structural hierarchy Proteins Phenotype of organism 3 Dimensional structure Function by interaction THE PROTEIN
Mature microrna identification via the use of a Naive Bayes classifier
 Mature microrna identification via the use of a Naive Bayes classifier Master Thesis Gkirtzou Katerina Computer Science Department University of Crete 13/03/2009 Gkirtzou K. (CSD UOC) Mature microrna identification
Mature microrna identification via the use of a Naive Bayes classifier Master Thesis Gkirtzou Katerina Computer Science Department University of Crete 13/03/2009 Gkirtzou K. (CSD UOC) Mature microrna identification
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Signal Transduction Pathway Smorgasbord
 Molecular Cell Biology Lecture. Oct 28, 2014 Signal Transduction Pathway Smorgasbord Ron Bose, MD PhD Biochemistry and Molecular Cell Biology Programs Washington University School of Medicine Outline 1.
Molecular Cell Biology Lecture. Oct 28, 2014 Signal Transduction Pathway Smorgasbord Ron Bose, MD PhD Biochemistry and Molecular Cell Biology Programs Washington University School of Medicine Outline 1.
For all of the following, you will have to use this website to determine the answers:
 For all of the following, you will have to use this website to determine the answers: http://blast.ncbi.nlm.nih.gov/blast.cgi We are going to be using the programs under this heading: Answer the following
For all of the following, you will have to use this website to determine the answers: http://blast.ncbi.nlm.nih.gov/blast.cgi We are going to be using the programs under this heading: Answer the following
Enzymes Part III: regulation II. Dr. Mamoun Ahram Summer, 2017
 Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Proteins. Amino acids, structure and function. The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka
 Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Hamby, Stephen Edward (2010) Data mining techniques for protein sequence analysis. PhD thesis, University of Nottingham.
 Hamby, Stephen Edward (2010) Data mining techniques for protein sequence analysis. PhD thesis, University of Nottingham. Access from the University of Nottingham repository: http://eprints.nottingham.ac.uk/11498/1/sehthesis_corrected_003.pdf
Hamby, Stephen Edward (2010) Data mining techniques for protein sequence analysis. PhD thesis, University of Nottingham. Access from the University of Nottingham repository: http://eprints.nottingham.ac.uk/11498/1/sehthesis_corrected_003.pdf
Structure-Function Relationship
 1 P a g e Structure-Function Relationship You have studied the amino acids and their characteristics, but in this part we will study the relation between the structure and the function of protein. Proteins
1 P a g e Structure-Function Relationship You have studied the amino acids and their characteristics, but in this part we will study the relation between the structure and the function of protein. Proteins
Amino acids. Side chain. -Carbon atom. Carboxyl group. Amino group
 PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
* Author to whom correspondence should be addressed; or Tel./Fax:
 Int. J. Mol. Sci. 2014, 15, 7594-7610; doi:10.3390/ijms15057594 Article OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms ihyd-pseaac: Predicting Hydroxyproline
Int. J. Mol. Sci. 2014, 15, 7594-7610; doi:10.3390/ijms15057594 Article OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms ihyd-pseaac: Predicting Hydroxyproline
Problem Set 2 September 18, 2009
 September 18, 2009 General Instructions: 1. You are expected to state all your assumptions and provide step-by-step solutions to the numerical problems. Unless indicated otherwise, the computational problems
September 18, 2009 General Instructions: 1. You are expected to state all your assumptions and provide step-by-step solutions to the numerical problems. Unless indicated otherwise, the computational problems
H C. C α. Proteins perform a vast array of biological function including: Side chain
 Topics The topics: basic concepts of molecular biology elements on Python overview of the field biological databases and database searching sequence alignments phylogenetic trees microarray data analysis
Topics The topics: basic concepts of molecular biology elements on Python overview of the field biological databases and database searching sequence alignments phylogenetic trees microarray data analysis
CHM333 LECTURE 6: 1/25/12 SPRING 2012 Professor Christine Hrycyna AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID:
 AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID: - The R group side chains on amino acids are VERY important. o Determine the properties of the amino acid itself o Determine
AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID: - The R group side chains on amino acids are VERY important. o Determine the properties of the amino acid itself o Determine
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data
 Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Christine Vogel 1, Edward M. Marcotte 1 *
 CALCULATING ABSOLUTE PROTEIN ABUNDANCE FROM MASS SPECTROMETRY BASED PROTEIN EXPRESSION DATA - SUPPLEMENTARY NOTES Christine Vogel 1, Edward M. Marcotte 1 * 1 Center for Systems and Synthetic Biology, Institute
CALCULATING ABSOLUTE PROTEIN ABUNDANCE FROM MASS SPECTROMETRY BASED PROTEIN EXPRESSION DATA - SUPPLEMENTARY NOTES Christine Vogel 1, Edward M. Marcotte 1 * 1 Center for Systems and Synthetic Biology, Institute
Section 1 Proteins and Proteomics
 Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
Gene Regulation Part 2
 Michael Cummings Chapter 9 Gene Regulation Part 2 David Reisman University of South Carolina Other topics in Chp 9 Part 2 Protein folding diseases Most diseases are caused by mutations in the DNA that
Michael Cummings Chapter 9 Gene Regulation Part 2 David Reisman University of South Carolina Other topics in Chp 9 Part 2 Protein folding diseases Most diseases are caused by mutations in the DNA that
Copyright 2008 Pearson Education, Inc., publishing as Pearson Benjamin Cummings
 Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Properties of amino acids in proteins
 Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Biology. Lectures winter term st year of Pharmacy study
 Biology Lectures winter term 2008 1 st year of Pharmacy study 3 rd Lecture Chemical composition of living matter chemical basis of life. Atoms, molecules, organic compounds carbohydrates, lipids, proteins,
Biology Lectures winter term 2008 1 st year of Pharmacy study 3 rd Lecture Chemical composition of living matter chemical basis of life. Atoms, molecules, organic compounds carbohydrates, lipids, proteins,
Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy
 7.4 - Translation 7.4.1 - Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy Each amino acid has a specific trna-activating
7.4 - Translation 7.4.1 - Explain that each trna molecule is recognised by a trna-activating enzyme that binds a specific amino acid to the trna, using ATP for energy Each amino acid has a specific trna-activating
Introduction. Basic Structural Principles PDB
 BCHS 6229 Protein Structure and Function Lecture 1 (October 11, 2011) Introduction Basic Structural Principles PDB 1 Overview Main Goals: Carry out a rapid review of the essentials of protein structure
BCHS 6229 Protein Structure and Function Lecture 1 (October 11, 2011) Introduction Basic Structural Principles PDB 1 Overview Main Goals: Carry out a rapid review of the essentials of protein structure
Biochemistry 15 Doctor /7/2012
 Heme The Heme is a chemical structure that diffracts by light to give a red color. This chemical structure is introduced to more than one protein. So, a protein containing this heme will appear red in
Heme The Heme is a chemical structure that diffracts by light to give a red color. This chemical structure is introduced to more than one protein. So, a protein containing this heme will appear red in
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting
 Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting Question No. 1 of 10 Question 1. Which of the following statements about the nucleus is correct? Question #01 A. The
Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting Question No. 1 of 10 Question 1. Which of the following statements about the nucleus is correct? Question #01 A. The
Protein Structure & Function. University, Indianapolis, USA 3 Department of Molecular Medicine, University of South Florida, Tampa, USA
 Protein Structure & Function Supplement for article entitled MoRFpred, a computational tool for sequence-based prediction and characterization of short disorder-to-order transitioning binding regions in
Protein Structure & Function Supplement for article entitled MoRFpred, a computational tool for sequence-based prediction and characterization of short disorder-to-order transitioning binding regions in
Personalized Colorectal Cancer Survivability Prediction with Machine Learning Methods*
 Personalized Colorectal Cancer Survivability Prediction with Machine Learning Methods* 1 st Samuel Li Princeton University Princeton, NJ seli@princeton.edu 2 nd Talayeh Razzaghi New Mexico State University
Personalized Colorectal Cancer Survivability Prediction with Machine Learning Methods* 1 st Samuel Li Princeton University Princeton, NJ seli@princeton.edu 2 nd Talayeh Razzaghi New Mexico State University
Protein Investigator. Protein Investigator - 3
 Protein Investigator Objectives To learn more about the interactions that govern protein structure. To test hypotheses regarding protein structure and function. To design proteins with specific shapes.
Protein Investigator Objectives To learn more about the interactions that govern protein structure. To test hypotheses regarding protein structure and function. To design proteins with specific shapes.
Evaluating Classifiers for Disease Gene Discovery
 Evaluating Classifiers for Disease Gene Discovery Kino Coursey Lon Turnbull khc0021@unt.edu lt0013@unt.edu Abstract Identification of genes involved in human hereditary disease is an important bioinfomatics
Evaluating Classifiers for Disease Gene Discovery Kino Coursey Lon Turnbull khc0021@unt.edu lt0013@unt.edu Abstract Identification of genes involved in human hereditary disease is an important bioinfomatics
Using SuSPect to Predict the Phenotypic Effects of Missense Variants. Chris Yates UCL Cancer Institute
 Using SuSPect to Predict the Phenotypic Effects of Missense Variants Chris Yates UCL Cancer Institute c.yates@ucl.ac.uk Outline SAVs and Disease Development of SuSPect Features included Feature selection
Using SuSPect to Predict the Phenotypic Effects of Missense Variants Chris Yates UCL Cancer Institute c.yates@ucl.ac.uk Outline SAVs and Disease Development of SuSPect Features included Feature selection
Cell Walls, the Extracellular Matrix, and Cell Interactions (part 1)
 14 Cell Walls, the Extracellular Matrix, and Cell Interactions (part 1) Introduction Many cells are embedded in an extracellular matrix which is consist of insoluble secreted macromolecules. Cells of bacteria,
14 Cell Walls, the Extracellular Matrix, and Cell Interactions (part 1) Introduction Many cells are embedded in an extracellular matrix which is consist of insoluble secreted macromolecules. Cells of bacteria,
Tala Saleh. Ahmad Attari. Mamoun Ahram
 23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
DIRECT IDENTIFICATION OF NEO-EPITOPES IN TUMOR TISSUE
 DIRECT IDENTIFICATION OF NEO-EPITOPES IN TUMOR TISSUE Eustache Paramithiotis PhD Vice President, Biomarker Discovery & Diagnostics 17 March 2016 PEPTIDE PRESENTATION BY MHC MHC I Antigen presentation by
DIRECT IDENTIFICATION OF NEO-EPITOPES IN TUMOR TISSUE Eustache Paramithiotis PhD Vice President, Biomarker Discovery & Diagnostics 17 March 2016 PEPTIDE PRESENTATION BY MHC MHC I Antigen presentation by
If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out.
 Sign In Forgot Password Register username username password password Sign In If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out. ChemWiki
Sign In Forgot Password Register username username password password Sign In If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out. ChemWiki
User Guide. Protein Clpper. Statistical scoring of protease cleavage sites. 1. Introduction Protein Clpper Analysis Procedure...
 User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
Cell Communication. Local and Long Distance Signaling
 Cell Communication Cell to cell communication is essential for multicellular organisms Some universal mechanisms of cellular regulation providing more evidence for the evolutionary relatedness of all life
Cell Communication Cell to cell communication is essential for multicellular organisms Some universal mechanisms of cellular regulation providing more evidence for the evolutionary relatedness of all life
A DATA MINING APPROACH FOR PRECISE DIAGNOSIS OF DENGUE FEVER
 A DATA MINING APPROACH FOR PRECISE DIAGNOSIS OF DENGUE FEVER M.Bhavani 1 and S.Vinod kumar 2 International Journal of Latest Trends in Engineering and Technology Vol.(7)Issue(4), pp.352-359 DOI: http://dx.doi.org/10.21172/1.74.048
A DATA MINING APPROACH FOR PRECISE DIAGNOSIS OF DENGUE FEVER M.Bhavani 1 and S.Vinod kumar 2 International Journal of Latest Trends in Engineering and Technology Vol.(7)Issue(4), pp.352-359 DOI: http://dx.doi.org/10.21172/1.74.048
a. From the grey navigation bar, mouse over Analyze & Visualize and click Annotate Nucleotide Sequences.
 Section D. Custom sequence annotation After this exercise you should be able to use the annotation pipelines provided by the Influenza Research Database (IRD) and Virus Pathogen Resource (ViPR) to annotate
Section D. Custom sequence annotation After this exercise you should be able to use the annotation pipelines provided by the Influenza Research Database (IRD) and Virus Pathogen Resource (ViPR) to annotate
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Protein kinases are enzymes that add a phosphate group to proteins according to the. ATP + protein OH > Protein OPO 3 + ADP
 Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
An Improved Algorithm To Predict Recurrence Of Breast Cancer
 An Improved Algorithm To Predict Recurrence Of Breast Cancer Umang Agrawal 1, Ass. Prof. Ishan K Rajani 2 1 M.E Computer Engineer, Silver Oak College of Engineering & Technology, Gujarat, India. 2 Assistant
An Improved Algorithm To Predict Recurrence Of Breast Cancer Umang Agrawal 1, Ass. Prof. Ishan K Rajani 2 1 M.E Computer Engineer, Silver Oak College of Engineering & Technology, Gujarat, India. 2 Assistant
ML LAId bare. Cambridge Wireless SIG Meeting. Mary-Ann & Phil Claridge 23 November
 ML LAId bare Cambridge Wireless SIG Meeting Mary-Ann & Phil Claridge 23 November 2017 www.mandrel.com @MandrelSystems info@mandrel.com 1 Welcome To Our Toolbox Our Opinionated Views! Data IDE Wrangling
ML LAId bare Cambridge Wireless SIG Meeting Mary-Ann & Phil Claridge 23 November 2017 www.mandrel.com @MandrelSystems info@mandrel.com 1 Welcome To Our Toolbox Our Opinionated Views! Data IDE Wrangling
The Immune Epitope Database Analysis Resource: MHC class I peptide binding predictions. Edita Karosiene, Ph.D.
 The Immune Epitope Database Analysis Resource: MHC class I peptide binding predictions Edita Karosiene, Ph.D. edita@liai.org IEDB Workshop October 29, 2015 Outline Introduction MHC-I peptide binding prediction
The Immune Epitope Database Analysis Resource: MHC class I peptide binding predictions Edita Karosiene, Ph.D. edita@liai.org IEDB Workshop October 29, 2015 Outline Introduction MHC-I peptide binding prediction
Bio 111 Study Guide Chapter 17 From Gene to Protein
 Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
User Guide. Association analysis. Input
 User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
Biochemistry #01 Bone Formation Dr. Nabil Bashir Farah Banyhany
 Biochemistry #01 Bone Formation Dr. Nabil Bashir Farah Banyhany Greetings This lecture is quite detailed, but I promise you will make it through, it just requires your 100% FOCUS! Let s begin. Today s
Biochemistry #01 Bone Formation Dr. Nabil Bashir Farah Banyhany Greetings This lecture is quite detailed, but I promise you will make it through, it just requires your 100% FOCUS! Let s begin. Today s
Chemical Mechanism of Enzymes
 Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
Q1: Circle the best correct answer: (15 marks)
 Q1: Circle the best correct answer: (15 marks) 1. Which one of the following incorrectly pairs an amino acid with a valid chemical characteristic a. Glycine, is chiral b. Tyrosine and tryptophan; at neutral
Q1: Circle the best correct answer: (15 marks) 1. Which one of the following incorrectly pairs an amino acid with a valid chemical characteristic a. Glycine, is chiral b. Tyrosine and tryptophan; at neutral
RNA Processing in Eukaryotes *
 OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
Classification of amino acids: -
 Page 1 of 8 P roteinogenic amino acids, also known as standard, normal or primary amino acids are 20 amino acids that are incorporated in proteins and that are coded in the standard genetic code (subunit
Page 1 of 8 P roteinogenic amino acids, also known as standard, normal or primary amino acids are 20 amino acids that are incorporated in proteins and that are coded in the standard genetic code (subunit
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
A Universal Trend among Proteomes Indicates an Oily Last Common Ancestor. BI Journal Club Aleksander Sudakov
 A Universal Trend among Proteomes Indicates an Oily Last Common Ancestor BI Journal Club 11.03.13 Aleksander Sudakov Used literature Ranjan V. Mannige, Charles L. Brooks, and Eugene I. Shakhnovich. 2012.
A Universal Trend among Proteomes Indicates an Oily Last Common Ancestor BI Journal Club 11.03.13 Aleksander Sudakov Used literature Ranjan V. Mannige, Charles L. Brooks, and Eugene I. Shakhnovich. 2012.
Organic Compounds. Compounds that contain CARBON are called organic. Macromolecules are large organic molecules.
 Macromolecules 1 Organic Compounds Compounds that contain CARBON are called organic. Macromolecules are large organic molecules. 2 Carbon (C) Carbon has 4 electrons in outer shell. Carbon can form covalent
Macromolecules 1 Organic Compounds Compounds that contain CARBON are called organic. Macromolecules are large organic molecules. 2 Carbon (C) Carbon has 4 electrons in outer shell. Carbon can form covalent
Introduction to Biochemistry
 Life is Organized in Increasing Levels of Complexity Introduction to Biochemistry atom simple molecule What is the chemical makeup of living things? macromolecule organ organ system organism organelle
Life is Organized in Increasing Levels of Complexity Introduction to Biochemistry atom simple molecule What is the chemical makeup of living things? macromolecule organ organ system organism organelle
Biomolecules Amino Acids & Protein Chemistry
 Biochemistry Department Date: 17/9/ 2017 Biomolecules Amino Acids & Protein Chemistry Prof.Dr./ FAYDA Elazazy Professor of Biochemistry and Molecular Biology Intended Learning Outcomes ILOs By the end
Biochemistry Department Date: 17/9/ 2017 Biomolecules Amino Acids & Protein Chemistry Prof.Dr./ FAYDA Elazazy Professor of Biochemistry and Molecular Biology Intended Learning Outcomes ILOs By the end
Identification of N-Glycosylation Sites with Sequence and Structural Features Employing Random Forests
 Identification of N-Glycosylation Sites with Sequence and Structural Features Employing Random Forests Shreyas Karnik 1,3, Joydeep Mitra 1, Arunima Singh 1,B.D.Kulkarni 1, V. Sundarajan 2,andV.K.Jayaraman
Identification of N-Glycosylation Sites with Sequence and Structural Features Employing Random Forests Shreyas Karnik 1,3, Joydeep Mitra 1, Arunima Singh 1,B.D.Kulkarni 1, V. Sundarajan 2,andV.K.Jayaraman
LAB#23: Biochemical Evidence of Evolution Name: Period Date :
 LAB#23: Biochemical Evidence of Name: Period Date : Laboratory Experience #23 Bridge Worth 80 Lab Minutes If two organisms have similar portions of DNA (genes), these organisms will probably make similar
LAB#23: Biochemical Evidence of Name: Period Date : Laboratory Experience #23 Bridge Worth 80 Lab Minutes If two organisms have similar portions of DNA (genes), these organisms will probably make similar
Chapter 11 CYTOKINES
 Chapter 11 CYTOKINES group of low molecular weight regulatory proteins secreted by leukocytes as well as a variety of other cells in the body (8~30kD) regulate the intensity and duration of the immune
Chapter 11 CYTOKINES group of low molecular weight regulatory proteins secreted by leukocytes as well as a variety of other cells in the body (8~30kD) regulate the intensity and duration of the immune
Chapter 2 Biosynthesis of Enzymes
 Chapter 2 Biosynthesis of Enzymes 2.1 Basic Enzyme Chemistry 2.1.1 Amino Acids An amino acid is a molecule that has the following formula: The central carbon atom covalently bonded by amino, carboxyl,
Chapter 2 Biosynthesis of Enzymes 2.1 Basic Enzyme Chemistry 2.1.1 Amino Acids An amino acid is a molecule that has the following formula: The central carbon atom covalently bonded by amino, carboxyl,
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Experimental design and workflow utilized to generate the WMG Protein Atlas.
 Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Lecture 2: Glycogen metabolism (Chapter 15)
 Lecture 2: Glycogen metabolism (Chapter 15) First. Fig. 15.1 Review: Animals use glycogen for ENERGY STORAGE. Glycogen is a highly-branched polymer of glucose units: Basic structure is similar to that
Lecture 2: Glycogen metabolism (Chapter 15) First. Fig. 15.1 Review: Animals use glycogen for ENERGY STORAGE. Glycogen is a highly-branched polymer of glucose units: Basic structure is similar to that
Contents. 2 Statistics Static reference method Sampling reference set Statistics Sampling Types...
 Department of Medical Protein Research, VIB, B-9000 Ghent, Belgium Department of Biochemistry, Ghent University, B-9000 Ghent, Belgium http://www.computationalproteomics.com icelogo manual Niklaas Colaert
Department of Medical Protein Research, VIB, B-9000 Ghent, Belgium Department of Biochemistry, Ghent University, B-9000 Ghent, Belgium http://www.computationalproteomics.com icelogo manual Niklaas Colaert
Annotation and Retrieval System Using Confabulation Model for ImageCLEF2011 Photo Annotation
 Annotation and Retrieval System Using Confabulation Model for ImageCLEF2011 Photo Annotation Ryo Izawa, Naoki Motohashi, and Tomohiro Takagi Department of Computer Science Meiji University 1-1-1 Higashimita,
Annotation and Retrieval System Using Confabulation Model for ImageCLEF2011 Photo Annotation Ryo Izawa, Naoki Motohashi, and Tomohiro Takagi Department of Computer Science Meiji University 1-1-1 Higashimita,
Using CART to Mine SELDI ProteinChip Data for Biomarkers and Disease Stratification
 Using CART to Mine SELDI ProteinChip Data for Biomarkers and Disease Stratification Kenna Mawk, D.V.M. Informatics Product Manager Ciphergen Biosystems, Inc. Outline Introduction to ProteinChip Technology
Using CART to Mine SELDI ProteinChip Data for Biomarkers and Disease Stratification Kenna Mawk, D.V.M. Informatics Product Manager Ciphergen Biosystems, Inc. Outline Introduction to ProteinChip Technology
Rajesh Kannangai Phone: ; Fax: ; *Corresponding author
 Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
Amino acids. You are required to know and identify the 20 amino acids : their names, 3 letter abbreviations and their structures.
 Amino acids You are required to know and identify the 20 amino acids : their names, 3 letter abbreviations and their structures. If you wanna make any classification in the world, you have to find what
Amino acids You are required to know and identify the 20 amino acids : their names, 3 letter abbreviations and their structures. If you wanna make any classification in the world, you have to find what
PERFORMANCE MEASURES
 PERFORMANCE MEASURES Of predictive systems DATA TYPES Binary Data point Value A FALSE B TRUE C TRUE D FALSE E FALSE F TRUE G FALSE Real Value Data Point Value a 32.3 b.2 b 2. d. e 33 f.65 g 72.8 ACCURACY
PERFORMANCE MEASURES Of predictive systems DATA TYPES Binary Data point Value A FALSE B TRUE C TRUE D FALSE E FALSE F TRUE G FALSE Real Value Data Point Value a 32.3 b.2 b 2. d. e 33 f.65 g 72.8 ACCURACY
Macromolecules of Life -3 Amino Acids & Proteins
 Macromolecules of Life -3 Amino Acids & Proteins Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Amino Acids Proteins
Macromolecules of Life -3 Amino Acids & Proteins Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Amino Acids Proteins
Chapt 15: Molecular Genetics of Cell Cycle and Cancer
 Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
Chapt 15: Molecular Genetics of Cell Cycle and Cancer Student Learning Outcomes: Describe the cell cycle: steps taken by a cell to duplicate itself = cell division; Interphase (G1, S and G2), Mitosis.
M. Kratzel. Two Examples of interactive elearning
 M. Kratzel Two Examples of interactive elearning I. Amino acid analysis I. Amino acid analysis 1. Hydrolysis 6 M HCl, 110 C 24...72 h = amide bond(s) 2. Derivatization = Phenylisothiocyanate PTC amino
M. Kratzel Two Examples of interactive elearning I. Amino acid analysis I. Amino acid analysis 1. Hydrolysis 6 M HCl, 110 C 24...72 h = amide bond(s) 2. Derivatization = Phenylisothiocyanate PTC amino
