Integrated Genomic Characterization of Papillary Thyroid Carcinoma
|
|
|
- Marlene Francis
- 6 years ago
- Views:
Transcription
1 Integrated Genomic Characterization of Papillary Thyroid Carcinoma Resource The Cancer Genome Atlas Research Network 1, * 1 The Cancer Genome Atlas Program Office, National Cancer Institute at NIH, 31 Center Drive, Bldg. 31, Suite 3A20, Bethesda, MD 20892, USA *Correspondence: giordano@umich.edu (T.J.G.), gadgetz@broadinstitute.org (G.G.) This is an open access article under the CC BY-NC-ND license ( SUMMARY Papillary thyroid carcinoma (PTC) is the most common type of thyroid cancer. Here, we describe the genomic landscape of 496 PTCs. We observed a low frequency of somatic alterations (relative to other carcinomas) and extended the set of known PTC driver alterations to include EIF1AX, PPM1D, and CHEK2 and diverse gene fusions. These discoveries reduced the fraction of PTC cases with unknown oncogenic driver from 25% to 3.5%. Combined analyses of genomic variants, gene expression, and methylation demonstrated that different driver groups lead to different pathologies with distinct signaling and differentiation characteristics. Similarly, we identified distinct molecular subgroups of BRAF-mutant tumors, and multidimensional analyses highlighted a potential involvement of oncomirs in less-differentiated subgroups. Our results propose a reclassification of thyroid cancers into molecular subtypes that better reflect their underlying signaling and differentiation properties, which has the potential to improve their pathological classification and better inform the management of the disease. INTRODUCTION The incidence of thyroid cancer has increased 3-fold over the past 30 years (Chen et al., 2009) and the prevalence of different histologies and genetic profiles has changed over time (Jung et al., 2014). All thyroid cancers, except medullary carcinoma, are derived from follicular cells that comprise the simple unicellular epithelium of normal thyroid. Eighty percent of all thyroid cancers are papillary thyroid carcinomas (PTCs), named for their papillary histological architecture. In addition, PTCs encompass several subtypes, including the follicular variant (FV), characterized by a predominantly follicular growth pattern. PTCs are usually curable with 5-year survival of over 95% (Hay et al., 2002); however, occasionally they dedifferentiate into more aggressive and lethal thyroid cancers. Current treatment involves surgery, thyroid hormone, and radioactive iodine (RAI) therapy (which exploits thyroid follicular cells avidity for iodine). Previous genetic studies report a high frequency (70%) of activating somatic alterations of genes encoding effectors in the mitogen-activated protein kinase (MAPK) signaling pathway, including point mutations of BRAF and the RAS genes (Cohen et al., 2003; Kimura et al., 2003; Lemoine et al., 1988; Suárez et al., 1988), as well as fusions involving the RET (Grieco et al., 1990) and NTRK1 tyrosine kinases (Pierotti et al., 1995). These mutations are almost always mutually exclusive (Soares et al., 2003), suggesting similar or redundant downstream effects. The various MAPK pathway alterations are strongly associated with distinct clinicopathological characteristics (Adeniran et al., 2006), and gene expression (Giordano et al., 2005) and DNA methylation profiles (Ellis et al., 2014). Mutations in members of the phosphoinositide 3-kinase (PI3K) pathway, such as PTEN, PIK3CA, and AKT1, have also been reported at low frequencies (Xing, 2013). We present The Cancer Genome Atlas (TCGA) project results from a comprehensive multiplatform analysis of 496 PTCs, the largest cohort studied to date. Clinically aggressive thyroid cancers (poorly and undifferentiated carcinomas) were excluded to maximally develop the compendium of tumor-initiating alterations. While excluding histological types of aggressive tumors limited some aspects of the study, the homogeneous PTC cohort allowed robust correlative analyses of multidimensional molecular data. The relatively quiet PTC genome allowed us to assess the signaling and differentiation consequences of the common drivers. Furthermore, the cohort allowed us to define integrated molecular subtypes that correspond to histology, signaling, differentiation state, and risk assessment. We put forth that our results will lead to improved clinicopathologic classification and management of patients. RESULTS Samples, Clinical Data, and Analytical Approach Tumor samples and matched germline DNA from blood or normal thyroid from 496 patients included 324 (69.4%) classical-type (CT), 99 (21.2%) follicular-variant (FV), 35 (7.5%) tall cell variant (TCV), 9 (2.0%) uncommon PTC variants, and 29 without histological annotation, primarily from nonirradiated patients (Table S1A available online). We estimated risk of tumor recurrence based on the 2009 American Thyroid Association guidelines (American Thyroid Association Guidelines Taskforce on Thyroid Nodules and Differentiated Thyroid Cancer et al., 2009), and assessed mortality risk using MACIS 676 Cell 159, , October 23, 2014 ª2014 The Authors
2 scores (Hay et al., 1993) (see Supplemental Information). We generated comprehensive and high-quality molecular data at TCGA genome sequencing and characterization centers with one proteomic and six genomic platforms (Table S1B) and analyzed the data at multiple genomic data analysis centers (Table S1C). Although 496 primary tumors were studied, the number of informative cases varied across platforms, mostly for technical reasons, and 390 tumors were analyzed on all major platforms (SNP arrays, exomes, RNA-seq, mirna-seq, and DNA methylation). Our analysis strategy consisted of three parts, each based on integrating several molecular data sets. First, we identified somatic mutations that included single nucleotide variants, small insertions and deletions, gene fusions, and copy-number alterations in order to characterize the genomic landscape of PTC and identify driver events in cases without any previously known driver (i.e., the so-called dark matter of the PTC genome). Next, we focused on the consequences of the driver mutations. We developed a gene expression signature of samples with these mutations and characterized tumors based on this signature. We then determined the differential signaling consequences of BRAF V600E and RAS mutations, the most common pathogenic mutations in the cohort, using protein and mrna expression data. Finally, we used the molecular data to derive molecular classifications of PTC and integrated these classifications with data related to genotype, signaling, differentiation, and risk. Somatic Single Nucleotide Variants and Insertions and Deletions Whole exome DNA sequencing of 402 (of 496) tumor/normal pairs (average depth-of-coverage; for tumors, for normals) showed a low somatic mutation density (0.41 nonsynonymous mutations per Mb, on average) (Figures 1A and S1A; Table S3) relative to other cancers (Lawrence et al., 2014; Lawrence et al., 2013). Mutation density correlated with age (Pearson correlation p = , Figure S1B), risk of recurrence (Kruskal-Wallis test p = ), and MACIS score (Pearson correlation p = , Figure S1C). To compensate for the confounding effect of age at diagnosis, we regressed the effect of age from mutation density and found that the association of risk with age-corrected mutation density remained (Kruskal-Wallis test p = ); MACIS scores did not (p = 0.19, Figure S1C). This association was maintained for the CT cohort (p = ), but not for other variants (Figure S1D). The correlation between mutation density and age suggests age should be used as a continuous variable in risk stratification (Bischoff et al., 2013), instead of a threshold of 45 years used in many staging systems. Mutation densities were not associated with other variables, like genotype or radiation exposure (Mann-Whitney p = and p = 0.173, respectively; Figure S1E). TCVs had the highest mutation density (Figure S1F), consistent with their known aggressive behavior. Five BRAF V600E mutant tumors with aggressive histologic features had higher mutation densities (Figures S1E and S1G). Ten tumors with the highest mutation densities (>1/Mb) were enriched for mutations associated with the APOBEC process (Roberts et al., 2013) (Mann-Whitney p = ), similar to bladder cancer (Cancer Genome Atlas Research Network, 2014). The relatively large number of patients in this study and the low background mutation density provides the statistical power to detect significantly mutated genes (SMGs) in as low as 3% of cases (>90% power for 90% of genes) (Lawrence et al., 2014). MutSig (Lawrence et al., 2013) detected seven SMGs (q < 0.1) (Figure 1C and Table S2) with four of the seven in < 3% of patients. SMGs included the MAPK-related genes, BRAF, NRAS, HRAS, and KRAS, which were virtually mutually exclusive (Figure 1C, Fisher s exact test p = , MEMo [Ciriello et al., 2012] corrected p < 0.01, Table S4A) in 300/402 (74.6%) patients. The 248 (61.7%) BRAF mutations were mostly V600E substitutions (Figures 1C and S2A and Table S3C). Somatic single nucleotide variants (SSNVs) were identified in 52 patients (12.9%) within codons 12 and 61 of RAS genes (Figures 1C and S2B and Table S3D). We observed strong associations between BRAF and RAS mutation status and histology, with BRAF V600E characterizing CT and TCV and RAS mutations characterizing FV (Figures 1B and 1C and Table S3E). These observations confirm the critical role of MAPK pathway alterations in PTC and illustrate that having more than one mutation confers no clonal advantage. MutSig also identified EIF1AX (eukaryotic translation initiation factor 1A, X-linked) as significantly mutated (q = ; Figure 1C and Table S3B). EIF1AX encodes a protein that mediates transfer of Met-tRNAf to 40S ribosomal subunits to form the 40S preinitiation complex for protein translation. Six (1.5%) mutations (Figure S2C) were identified in tumors lacking other known driver mutations (Fisher s exact test of EIF1AX versus RAS/ BRAF, p = 0.013) with one exception: one case with three driver mutations in KRAS, BRAF, and EIF1AX; here the KRAS mutation was clonal (cancer cell fraction estimated at 100% with 95% CI [56% 100%]), while EIF1AX and BRAF were subclonal at 76% (38% 100%) and 53% (22% 94%) cancer cell fractions, respectively (see Supplemental Information). The near-mutual exclusivity of EIF1AX alterations with MAPK pathway mutations (Figure 1C), together with recurrent mutations in other tumors (Forbes et al., 2011; Martin et al., 2013), suggests that EIF1AX is a novel cancer gene in PTC. The two remaining SMGs (PPM1D, CHEK2, Figures S2D and S2E) encode interacting proteins related to DNA repair (Oliva-Trastoy et al., 2007) and occurred concomitant with MAPK-pathway driver mutations (Figure 1C). Germline PPM1D mutations are associated with breast and ovarian cancer predisposition and may impair p53 function (Kleiblova et al., 2013). Although not statistically significant, there were 8 additional DNA repair-related mutations in 26 (6.5%) tumors, all mutually exclusive (Figure S3A, MEMo corrected p = 0.24). Tumors carrying these mutations had a significantly higher median mutation density (Table S5A, Mann Whitney p = 0.022), although not after age-adjustment (p = 0.111), and were enriched with high-risk patients (Fisher s exact test p = 0.018). These observations, together with our finding of a mutated network of FANCA-associated genes (see Integrated Analysis of Somatic Alterations section), suggest that acquisition of a defect in DNA repair represents a mechanism for development of aggressive PTC. Cell 159, , October 23, 2014 ª2014 The Authors 677
3 A B C D E F Figure 1. Landscape of Genomic Alterations in 402 Papillary Thyroid Carcinomas (A) Mutation density (mutations/mb) across the cohort. (B) Tumor purity, patient age, gender, history of radiation exposure, risk of recurrence, MACIS score, histological type, and BRS score. (C) Number and frequency of recurrent mutations in genes (left) ranked by MutSig significance (right), gene-sample matrix of mutations (middle) with TERT promoter mutations (bottom). (D) Number and frequency of fusion events (left), gene-sample matrix of fusions across the cohort (middle). (E) Number and frequency of SCNAs (left), chromosome-sample matrix of SCNAs across the cohort (middle) with focal deletions in BRAF and PTEN (bottom), GISTIC2 significance (right). (F) Driving variant types across the cohort. Samples were sorted by driving variant type with dark matter on the left. See also Figures S1, S2, S3, S4 and Tables S1, S2, S3, S4A, S5A, S5B, and S5E. We also observed alterations in other cancer genes, pathways and functional groups (chromatin remodeling, PI3K, WNT and tumor suppressor genes). Although not statistically significant, they are known to play a role in thyroid cancer pathogenesis and progression (Xing, 2013). We identified 93 mutations within 57 epigenetic regulatory genes in 80/402 (20.0%) tumors, 9 of which possessed more than one mutation (Figure S3D). Mutations in MLL (1.7%), ARID1B (1.0%), and MLL3 (1%) were most frequent (Figures 1C and S3D). In the PI3K and PPARg pathways, we observed 20 nearly mutually exclusive mutations in PTEN, AKT1/2, and PAX8/PPARG (Figure S3E), representing 4.5% (18/402) of cases. Mutations of five WNT pathway-related genes were found in 6/402 (1.5%) tumors (Figure S3B), a lower frequency than reported for aggressive tumor types (Xing, 2013). Mutations of tumor suppressor genes (TP53, RB1, NF1/ 2, MEN1, and PTEN) were identified in 15/402 (3.7%) tumors (Figure S3C). In addition, two genes that may be tumor suppressors were near significance: (1) ZFHX3, a zinc finger homeobox transcription factor (Minamiya et al., 2012), in 7/402 (1.7%) tumors (q = 0.79); and (2) BDP1, which may regulate AKT signaling (Woiwode et al., 2008), in 5/402 (1.2%) tumors (q = 0.58). Finally, mutations in thyroid-related genes were infrequent: 11/402 (2.7%) mutations in thyroglobulin, 2/402 (0.5%) mutations in thyroid-stimulating hormone receptor (TSHR). No mutations in thyroid hormone receptor genes (TRHA and TRHB) were found. We identified TERT promoter mutations in 36 (9.4%) of 384 informative tumors, with 27 (7.0%) C228T, 1 (0.3%) C228A, and 8 (2.1%) C250T substitutions. These mutations were present 678 Cell 159, , October 23, 2014 ª2014 The Authors
4 A B C D Figure 2. TERT Promoter Mutations and Clonality Assessment of Driver Mutations (A C) Association of TERT promoter mutations with (A) risk of recurrence, (B) MACIS score, and (C) thyroid differentiation score (TDS). See also Table S2. (D) Mutation cancer cell fraction distribution. The majority of all mutations, including driver mutations BRAF, NRAS, HRAS, KRAS, and EIF1AX, have a calculated cancer cell fraction close to 1.0, indicating their presence all tumor cells. in PTCs of all histological types. TERT promoter mutations had modest association with mutation drivers (Fisher s exact test p = 0.029) and arm-level somatic copy-number alterations (p = 0.023), but not BRAF mutations or gene fusions. They showed strong associations with older age, MACIS scores (Kruskal- Wallis test p = , and p = , Figure 2B, respectively), and high risk of recurrence (Fisher s exact test p= , Figure 2A), within the entire cohort, and these associations remained within the BRAF V600E tumors. Finally, TERT promoter mutations occurred in less-differentiated PTCs (lower TDS values, see Signaling and Differentiation section) (Kruskal-Wallis test p = , Figure 2C). These associations are consistent with published results (Melo et al., 2014; Xing et al., 2014) and suggest that molecular diagnostic assessment of TERT promoter mutations may be used to identify high risk patients. A recent report suggested that BRAF V600E mutations in PTC are often present only in a small subset of the cancer cells (Guerra et al., 2012), information relevant to therapeutic application of BRAF inhibitors. To address this issue, we used the ABSOLUTE package (Carter et al., 2012) to estimate the cancer cell fraction (CCF) of mutations in BRAF, NRAS, HRAS, KRAS, and EIF1AX (see Supplemental Information). In our study, all of these driver mutations were present in the majority of tumor cells (Figure 2D), i.e., they are largely clonal. Our results instead confirm other studies showing homogeneous cancer cell immunohistochemical staining with BRAF V600E -specific antibodies (Ghossein et al., 2013). We performed targeted sequencing validation experiments on a subset of 318 tumors at 333 mutated sites. Two hundred sixty-four of 265 (99.6%) of mutations in driver genes were confirmed. We also carried out targeted validation on a random set of 54 mutations, of which 49 (91%) were confirmed, for an overall validation rate of 96%. Gene Fusions Chromosomal rearrangements and translocations contribute to PTC pathogenesis (Xing, 2013). We identified both known and novel fusions, including new partners of previously described fusions, in 74 (15.3%) of 484 informative cases, based on multiple platforms (Figure 3). Fusions were mutually exclusive with each other and with BRAF, RAS, and EIF1AX mutations (Fisher s exact test p = ; Figures 1C, 1D and S3F). Fusionpositive tumors were associated with younger age of diagnosis (Wilcoxon rank-sum test p = 0.005), but not with risk of recurrence (Fisher s exact test p = 0.55) or age-corrected mutation density (rank-sum test p = 0.341, Table S5A). RET fusions were most frequent (33/484, 6.8%) (Table S5B and Figure S3F); however, less frequent than previously reported in sporadic or radiation-associated PTC (Ricarte-Filho et al., 2013). We identified four novel unique RET fusions that preserved the kinase domain (Figure 3A and Table S5B). Fusions involving BRAF have been identified in PTC (Ciampi et al., 2005) and other cancers. In melanoma, these events define a molecular subclass with distinct response to MEK inhibition (Hutchinson et al., 2013). In our cohort, we identified 13/484 (2.7%) BRAF fusions with diverse gene partners (Figure 3A and Table S5B); three tumors exhibited the SND1/BRAF gene fusion seen in a gastric cancer cell line (Lee et al., 2012). Some fusions supported BRAF signaling with expression and conservation of its kinase domain (MKRN1/BRAF), while others suggested an alternative activating mechanism. Six of nine BRAF fusions were validated by independent PCR experiments, while PCR evidence for the other three was inconclusive. These diverse fusions, together with the BRAF point mutations and indels described above, illustrate the various possible mechanisms of activating BRAF and highlight its oncogenic importance in PTC. Cell 159, , October 23, 2014 ª2014 The Authors 679
5 Figure 3. Candidate Driver Gene Fusions in Papillary Thyroid Carcinoma (A) RNA expression fusion plots for representative novel candidate genes involving RET, BRAF, ALK, NTRK3, and LTK fusions. Each gene in the fusion plot is drawn 5 0 to 3 0, exon specific relative expression data are represented with low (blue) and high expression (red), and the kinase domain is mapped with a green box. The pairs of numbers across the links indicate the number of split reads and paired-end supporting reads from RNA-seq. (B) Circos plots ( ofret, BRAF, and NTRK3 fusions. Red links represent recurrent fusions, black nonrecurrent. See also Figures S3F and Table S5B. We identified PAX8/PPARG fusions in 4/484 (0.8%) tumors. Originally found in follicular carcinomas (Kroll et al., 2000), PAX8/PPARG translocations have been reported with low frequency in PTC, especially FV. ETV6/NTRK3 and RBPMS/ NTRK3 fusions were uncovered in 6/484 (1.2%) tumors. These fusions are more prevalent in radiation-induced thyroid cancers but have lower prevalence in sporadic PTC (Ricarte-Filho et al., 2013). THADA fusions were identified in 6/484 (1.2%) tumors. Fusions involving ALK presented in 4/484 (0.8%) tumors, including EML4/ALK (Figure 3A), which is observed in lung adenocarcinomas and rare thyroid cancers (Kelly et al., 2014), suggesting opportunities for targeted inhibition of ALK. In addition, two cases had FGFR2 fusions and two cases had nonrecurrent fusions of MET and LTK (Table S4B). Somatic Copy-Number Alterations Somatic copy-number alterations (SCNAs) were identified in 135 (27.2%) of 495 informative tumors. These 135 cases were significantly enriched in cases with no driver mutation or fusion (Fisher s exact test p = ; Figures 1C, 1D, and 1E), suggesting that SCNAs may also drive PTC. Arm-level alterations occurred significantly more frequently in FV than in CT subtypes (Figure S4A) (FDR < 0.1, p < 0.008), providing evidence for a close relationship between FV and follicular neoplasms in which SCNAs are more common (Wreesmann et al., 2004a). Unsupervised clustering of chromosomal arm-level alterations defined four distinct classes (Figure S4B). The largest class (72.9%) lacked significant gains or losses (SCNA-quiet), reflecting the highly differentiated nature of PTC; this group was not enriched for any particular genotype or histologic type. A second class (9.9%) was characterized by an isolated loss of 22q (SCNA-22q-del), a region that includes NF2 and CHEK2 and reported to be lost with significant frequency in PTC (Kjellman et al., 2001). Seventy tumors had 22q loss and five tumors possessed CHEK2 mutations, with 4 cases containing both mutations (p = ). The SCNA-22q-del cohort contained few TCV and was enriched for FV (p < 0.05) (Figure S4C). This result suggests that loss of CHEK2 and/or the NF2 tumor suppressor may be important in PTC, particularly in the FV subtype. A third class (14.8%) was characterized by a few SCNA events and gain of 1q (SCNA-low-1q-amp), was enriched for TCV (p < ) and BRAF mutations (p < 0.05) (Figure S4C), and was associated with significantly higher MACIS scores (p < ) (Figure S4D), risk profiles (Figure S4E), and tumor stage (Figure S4F), consistent with reports of 1q gains in aggressive PTC (Wreesmann et al., 2004b). The final and smallest class (2.4%) was defined by a higher frequency of focal gains and losses (SCNA-high). We found few significantly recurring focal alterations using GISTIC2 (Mermel et al., 2011). In 5/13 tumors with BRAF fusions, we detected by SNP array the resulting focal alterations at 7q34. Five tumors with 10q23.31 deletions lacked PTEN expression (Figure S4G). We also found single case amplifications containing oncogenic driver genes (e.g., FGFR3). Together, these results, in the context of SSNVs and fusions, suggest that SCNAs may represent both tumor-initiating and progressionrelated events in PTC. 680 Cell 159, , October 23, 2014 ª2014 The Authors
6 Integrated Analyses of Somatic Alterations Next, we sought to identify additional genes that may harbor driver mutations (point mutations, fusions or copy-number changes) that did not meet significance, by searching within protein-protein interaction networks for subnetworks enriched with frequently mutated genes. To this end, we applied the HotNet2 algorithm (M.D.M. Leiserson, F. Vandin, H.T. Wu, J.R. Dobson, J.V. Eldridge, J.L. Thomas, A. Papoutsaki, Y. Kim, B. Niu, M. McLellan, M.S. Lawrence, A.G. Perez, D. Tamborero, Y. Cheng, G.A. Ryslik, N. Lopez-Bigas, G. Getz, L. Ding, and B.J. Raphael, unpublished data) and identified 17 significantly mutated subnetworks (p < 0.004, Table S5C and Figure S5A). As expected, the largest subnetwork (16 genes) included four known members of the MAPK signaling pathway (BRAF and three RAS genes) and 12 additional genes (Figure S5B). Some of these additional genes (e.g., RAP1GAP) displayed mutations that were mutually exclusive with BRAF (SSNVs, indels and fusions) and RAS, and may represent additional MAPK drivers. Other mutated genes in this subnetwork (e.g., PIK3CA) overlapped with BRAF V600E and may alter the biology of these cancers by activating PI3K signaling, a hypothesis that requires validation. Other HotNet2 subnetworks significantly overlapped with known pathways (Table S5C and Figures S5C S5E). Identification of the ECM-receptor interactions pathway is consistent with the role of ECM microenvironment in PTC (Nucera et al., 2011). The finding of a FANCA-associated protein complex subnetwork provides additional evidence for DNA repair playing a role in PTC. We used network-based stratification (NBS) (Hofree et al., 2013) to discover three somatic mutation-based PTC subtypes (NBS1-3, Figures S6A S6C). As expected, a strong association with histologic subtype was found, with a significant association between NBS1 and FV histology (Figure S6B, b, Fisher s exact test p < ). The subtypes were also significantly associated with lymph node status, extrathyroidal extension, stage and risk of recurrence (Figure S6B, a, c e, Fisher s exact test p = , , , , respectively), as well as other molecular characteristics (Figure S6C). In order to identify somatic events that characterize each subnetwork, we applied the HotNet algorithm to each NBS subtype (Figure S6D and Table S5D). Subtype NBS1 was associated with perturbations in RAS, PTEN, PPARG, and TSHR. Subtype NBS2 was associated with alterations in RET and related genes such as NTRK3. Subtype NBS3 appears to be predominantly BRAF V600E -associated. These results confirm three broad classes of PTC determined by the common drivers. We also looked for the presence of viral pathogens in PTC using two independent methods to assess the RNA-seq data: PathSeq (Kostic et al., 2011) and BioBloom Tools (Chu et al., 2014). We identified two tumors with hepatitis B virus (HBV) and one tumor with human papillomavirus 45 (HPV45) at relative frequencies exceeding 0.1 viral reads per million human reads (RPM) for PathSeq and 0.2 RPM for BBT (see Supplemental Information and Tables S4G S4I), indicating that viral pathogens are unlikely significant contributors to PTC pathogenesis. Dark Matter Summary Starting with 402 cases with informative exome DNA sequence data, we examined tumors that lacked apparent driver mutations, so-called dark matter samples, for the presence of novel potential driver alterations. SSNVs involving drivers accounted for 299 (73.6%) cases. Mutually exclusive fusions increased the number of cases with drivers to 358 (89.0%). Three mutually exclusive focal deletions (2 PTEN and 1 BRAF) brought the number of cases to 361 (89.8%). Mutually exclusive arm-level SCNAs were present in 27 additional cases, which were mostly FV. Although we cannot pinpoint the driving genes, if we assume that some of these SCNAs indeed act as drivers, the total number of cases with apparent drivers increased to 96.5%, leaving 14 (3.5%) as dark matter cases. By reviewing events in these cases, we observed additional potential drivers such as APC, ATM, NF1, and SPOP, mutations of chromatin remodeling genes (e.g., MLL) and potential gene fusions (Table S5E). Including these events and considering arm-level SCNAs as drivers, we have identified putative cancer drivers in 397/402 PTCs (98.8%). Signaling and Differentiation PTC is a MAPK-driven cancer that has two mutually exclusive drivers with distinct signaling consequences: BRAF V600E and mutated RAS. Tumors driven by BRAF V600E do not respond to the negative feedback from ERK to RAF (since it signals as a monomer), resulting in high MAPK-signaling (Pratilas et al., 2009). Conversely, tumors driven by RAS and RTK fusions signal via RAF dimers that respond to ERK feedback, resulting in lower MAPK-signaling. This differential signaling results in profound phenotypic differences. For example, expression of genes responsible for iodine uptake and metabolism are greatly reduced in BRAF V600E tumors, in contrast to the RAF-dimer tumors in which expression of these genes is largely preserved (Durante et al., 2007). These observations, together with the relatively low number of other genomic alterations, allow for a clear view of the signaling and transcriptional outputs of these two primary drivers. The distinct profile of expression of genes involved in thyroid hormone biosynthesis that we observed between BRAF V600E and RAS-driven tumors is recapitulated closely in mouse PTC models induced by knock-in mutations of Braf V600E or Hras G12V (Charles et al., 2011; Franco et al., 2011), suggesting that these arise as a consequence of the constitutive activation of these drivers. To explore these relationships across our cohort, we developed a BRAF V600E -RAS score (BRS) to quantify the extent to which the gene expression profile of a given tumor resembles either the BRAF V600E or RAS mutant profiles. Using 391 samples with both exome and RNA sequencing data, we compared BRAF V600E -mutated and RAS-mutated tumors to derive a 71- gene signature. Correlations with this signature were used to derive a continuous measure ( 1 to +1) with BRAF V600E -like (BVL) PTCs being negative and RAS-like (RL) PTCs positive (see Supplemental Information). As expected, this signature showed strong separation of the BRAF V600E and RAS mutant tumors (Figures S7A and S7B). We then used the BRS as a reference continuous scale from most BVL to most RL to interrogate the signaling consequences of the other, less common, mutations (Figures 4A and 4B). All BRAF mutations other than BRAF V600E exhibited RL behavior, including one BRAF K601E, a splice-site mutation and three indels. This is consistent with previous observations that BRAF K601E Cell 159, , October 23, 2014 ª2014 The Authors 681
7 Figure 4. The BRAF V600E -RAS Score (A E) Thyroid samples (A) (n = 391) were ranked by BRAF V600E -RAS score (BRS), with BRAF V600E -like and RAS-like samples having negative ( 1 to 0) and positive scores (0 to 1), respectively. The BRS is strongly associated with: (B) driver mutation status; (C) thyroid differentiation score (TDS); (D) single data-type clusters; and (E) histology and follicular fraction. The RAS-like samples (normalized score > 0, in red on the top bar) consistently emerged as a distinct subgroup characterized by a higher TDS. See also Figures S6, S7A, and S7B and Tables S2 and S4B. occurs in FV tumors that are mostly RL-PTCs (Park et al., 2013). All of the BRAF fusions were BVL. Four of the six EIF1AX mutations were RL, one neutral, and one weakly BVL. All of the PAX8/ PPARG fusions were RL, consistent with their prevalence in follicular-patterned tumors. Nearly all of the RET fusions were weakly BVL, and the NTRK1/3 and ALK fusions were largely neutral. Next, we focused on thyroid differentiation, which plays a central role in thyroid cancer. We summarized the expression levels of 16 thyroid metabolism and function genes (Table S5F), which were highly correlated across our cohort, and produced a single metric, designated the Thyroid differentiation score (TDS). The TDS and BRS measures were highly correlated across all tumors (Spearman = 0.78, p = ), despite being derived from different gene sets. This correlation was mainly driven by RL-PTCs having relatively high TDS values. The BRAF V600E PTC cohort, considered a homogeneous group in numerous studies, showed a wide range of TDS values (Figure 5), and maintained the TDS and BRS correlation, albeit to a lesser degree (Spearman = 0.38, p = , Figures 4, 5, and S7C S7E). To gain insights regarding the observed TDS variation, we identified other genes whose expression levels correlated with TDS across all tumors and within the BRAF V600E cohort. We discovered that the TDS was associated with global expression changes, significantly correlated and anticorrelated with thousands of genes in both cohorts (Figures S7F and S7G). We obtained similar observations using previously reported DNA microarray data (Giordano et al., 2005) (Figure S7H). Next, to test whether differences in the TDS within the BRAF V600E mutant cohort were associated with subtle architectural changes, we histologically graded the tumors (Figure S7I) and showed that TDS was indeed correlated with grade (Kruskal-Wallis test p = , Figure S7J). TDS also correlated with risk (p = ) and MACIS (Spearman correlation p = ) but only weakly with tumor purity (p = )(Figure S7J), indicating that the observed differences in TDS and global expression levels were not strongly influenced by variations in levels of tumor stroma or lymphocyte infiltration. These results support the validity of the TDS and BRS and illustrate that, while independently derived, these measures reflect similar biological properties that are profoundly reflected by gene expression in PTC. Although the TDS was correlated with many genes, among the most correlated were several genes with cancer relevance 682 Cell 159, , October 23, 2014 ª2014 The Authors
8 Figure 5. Role of Thyroid Differentiation in Papillary Thyroid Carcinomas Thyroid differentiation score (TDS) across the cohort with tumors sorted by driver mutation and TDS. Below TDS are the BRAF V600E -RAS score (BRS), ERK signature, histological type, MACIS score, risk of recurrence, driver mutations, and gene expression data for nine thyroid genes used to derive the TDS (TG, TPO, SLC26A4 [pendrin], SLC5A5 [Na/I symporter], SLC5A8 [apical iodide transporter], DIO1, DIO2, DUOX1, and DUOX2), four selected mrnas correlated to TDS, and three selected mirs correlated to TDS. Featured mrna (except for 16 thyroid genes) and mirna genes were selected based on Spearman correlation to TDS in the BRAF V600E cohort (*) and the full cohort (**) (see Supplemental Information). See also Figures S7C S7J and Table S5F. (TFF3, KIT, PVRL4, and FHL1, q = , , , and , respectively, Figures 5, S7F, and S7G). Among mirs with cancer relevance, mir-21, mir- 146b, and mir-204 were highly correlated with the TDS (Figure 5, S7F, and S7G), with mir-21 being the most negatively correlated mir in both the entire and BRAF V600E cohorts. mir-21 and mir-146b are oncogenic mirs in several tumor types (Di Leva et al., 2014) and mir-204 is downregulated in several tumors types and may be a tumor suppressor (Imam et al., 2012). We subsequently identified variable expression of these mirs in distinct mir clusters with different TDS and BRS values (see Molecular Classification section). Our data demonstrate significant gene expression variation across the BRAF V600E cohort, which may account for the range of differentiation observed and may explain the uncertainty regarding the prognostic and predictive power of BRAF V600E mutation (Xing et al., 2013). Dedifferentiation likely plays a role in dampening responses to RAI therapy and is consistent with observations that RAI-refractory metastases are enriched for BRAF V600E mutants (Sabra et al., 2013). Although other factors are likely involved, our results support the view that potent constitutive activation of the MAPK transcriptional output by oncogenic BRAF V600E downregulates the expression of genes involved in iodine metabolism (Chakravarty et al., 2011; Franco et al., 2011). Of note, the loss of differentiation within the BRAF V600E cohort is likely smaller than that observed in histologically aggressive thyroid cancers. An integrated view of our TDS analysis (Figure 5) summarizes how RL-PTCs result in highly differentiated tumors enriched for follicular histology with distinct gene expression and DNA methylation patterns. Conversely, BVL-PTCs result in predominantly less-differentiated tumors enriched for classical and tall cell histology, with distinct gene expression and DNA methylation patterns. Further assessment of BRS and TDS might be useful in the setting of a clinical trial and even pathology practice using immunohistochemistry. To better understand the downstream signaling effects of the main driver events (BRAF V600E, RAS), we examined mrna expression and protein and phosphoprotein levels of various signaling pathways. We used a 52-gene signature derived by inhibiting MEK in a BRAF V600E melanoma cell line (Pratilas et al., 2009) to assess the ERK (and MAPK) activation level. BRS was highly correlated to ERK activation level (i.e., ERK score, see Supplemental Information); BVL-PTCs showed overactivation of the pathway (Figure S8A) and increased expression of DUSP genes (Figure 6). Note that, although the two scores are highly correlated, they were derived independently and have no genes in common. In wild-type cells, ERK induces a negative feedback on effectors upstream in the pathway, resulting in impairment of RAF dimerization. Because BRAF V600E signals as a monomer, it is insensitive to feedback leading to ERK overactivation (Poulikakos et al., 2010). RET fusions consistently had BVL phenotype, high MAPK activity and were associated with low ps338-craf and Cell 159, , October 23, 2014 ª2014 The Authors 683
9 A B Figure 6. Downstream Signaling of BVL and RL PTCs (A) MAPK and PI3K pathways are differentially activated in the BVL and RL PTCs. (B) BRAF V600E -mutated cases show robust activation of MAPK signaling resulting in higher output of the ERK transcriptional program, represented in particular by DUSP (DUSP4, 5 and 6) mrnas. This may be due to insensitivity of BRAF V600E to ERK inhibitory feedback. By contrast, RAS-like tumors activated both MAPK and PI3K/AKT signaling, as shown by higher pakt levels in these tumors. The mechanism by which RAS-like tumors activated MAPK signaling was distinct from that of BRAF V600E tumors, as they had higher CRAF phosphorylation, consistent with engagement of RAF dimers. Paradoxically, RL-PTCs had higher phosphorylation of the ERK substrate p90rsk, which was associated with mtor activation, likely through phosphorylation and consequent inhibition of TSC2. RL-PTCs also showed activation of an antiapoptotic program, characterized by S112-BAD phosphorylation (a target of P90RSK) and BCL2 overexpression. See also Figures S8 and Tables S4B and S4F. ps299-araf (Figure S8B), consistent with preferential signaling via BRAF homodimers (Mitsutake et al., 2006). RL-PTCs had concurrent activation of PI3K/AKT and MAPK signaling, the latter mostly through c-raf phosphorylation (Figure 6). Despite having lower MAPK activity than BVL-PTCs, RL-PTCs showed significantly higher phosphorylation of p90rsk, a direct ERK substrate. Activation of p90rsk was associated with robust inhibition of TSC2, a distinguishing hallmark of the two functional classes, and likely to induce mtor. Elevated p90rsk in the RL-PTCs was also associated with phosphorylation of its substrate BAD, and with concurrent BCL2 overexpression, leading to antiapoptotic signaling (Figure 6 and Table S4B). Finally, we used TieDIE (Paull et al., 2013) to assess differential pathway activation between BVL-PTC and RL-PTC (see Supplemental Information). This approach identified the small GTPase RHEB, a known regulator of mtor activity (Groenewoud and Zwartkruis, 2013), as a contributing factor to the differences observed between BVL-PTC and RL-PTC (Figure S8C, Table S4F and Data S1). These findings confirm many of the known signaling changes induced by BRAF V600E and RAS mutations in PTC, provide a framework for MAPK downstream activity in tumors with other driver alterations, and importantly shed new light on the role of p90rsk as a crucial crossroad for MAPK, mtor, and BCL2 signaling in RAS-driven tumors. Molecular Classification The comprehensive and multiplatform molecular data and large sample size in this study provide an opportunity to derive and refine the classification of PTC into molecular subtypes and associate them with clinically relevant parameters. To this end, we leveraged the BRS and TDS measures to inform the relationships between tumor cluster, histology, genotype, signaling, and differentiation. Applying unsupervised clustering methods to four genomic data sets yielded a different number of subtypes for each data set: five for mrna expression, six for mir expression, four for DNA methylation, and four for protein expression (Figures S9A S9D). All clustering results were consistent with two meta-clusters that separated the BRAF V600E -driven tumors (BVL-PTCs) from ones with RAS mutations (RL-PTCs), recapitulating the BRS-partitioning and association with histological subtypes (Figures 4 and S9A). We used StratomeX (Streit et al., 2014) to visually highlight these relationships (see THCA publication page, thca_2014/); in particular, the significant distinction between BVL-PTCs and RL-PTCs that is evident in each of the molecular data sets (Table S5G). We also applied SuperCluster (see Supplemental Information) to the four genomic data sets, which supported the overarching separation of BVL-PTCs and RL- PTCs (Figure S9E). Next, we focused on the internal structure reported by different data types for these two meta-clusters. The RL-PTC group was associated with a single cluster in all data sets except DNA methylation (described below). Overall, this group was characterized by FV histology, relatively low risk of recurrence, distinct mrna expression profiles with lower expression of 684 Cell 159, , October 23, 2014 ª2014 The Authors
10 Figure 7. Unsupervised Clusters for mirna-seq Data Heatmap showing discriminatory mirs (5p or 3p mature strands) with the largest 6% of metagene matrix scores (see Supplemental Information), as well as mir-204-5p, 221-3p, and 222-3p, which were highlighted in correlations to BRS and TDS scores (see Figure S10D). The scalebar shows log 2 normalized (reads-per-million, RPM), mediancentered mir abundance. mir names in red are discussed in the text. Gray vertical lines in the clinical information tracks mark samples without clinical data, and in the mutation tracks gray lines identify samples without sequence data. See also Figures S9, S10, and Tables S4C, S4D, S4E, S5G, and S6. immune response genes, higher expression of mir-182-5p and mir-183-5p, low levels of fibronectin, VHL, and CHK2 proteins, and high expression of claudin-7, TIGAR, and BRCA2 proteins (Figures 7, S9B and S9D). RL tumors were highly differentiated and associated with younger patients. The DNA methylation data partitioned these tumors into two clusters: the larger, termed Meth-follicular, showed few methylation changes compared to normal thyroid, while the other, termed the Meth-CpG Island cluster, was characterized by hypermethylation of a large number of CpG sites in islands and shores (Figure S9C). The significance of this distinction is unclear, although the Meth-CpG Island cluster tended to have high tumor purity with less lymphocyte infiltration and stromal cells (Figure S9C, d). The different data sets partitioned the BVL-PTC group into different numbers of subtypes (from two based on DNA methylation data to five based on mir data), which did not overlap with each other such as to form a fully consistent lower level partitioning of BVL-PTCs (Figures 4 and S9). Regardless of which data set was used, the clusters were significantly different based on parameters like proportions of driver mutations and gene fusions, mutational densities, histological and risk profiles, age, and BRS and TDS values (Table S5G). The most striking relationship between subtypes was that mrna-cluster 5 was nearly fully embedded within mir-cluster 6 (86/106 mrna-cluster 5 tumors were part of the 144 tumors in mir-cluster 6; Fisher s exact test p = ). This mrna cluster, which we termed tall cell-like, contained most of the TCV tumors (74%), had the highest frequency of BRAF V600E mutations (78%), and the lowest BRS and TDS values (i.e., the strongest BVL phenotype and least differentiated). Since the tall cell-like cluster was associated with more advanced stage (see THCA publication page) and higher risk (Table S5G), this may be clinically relevant. We again used StratomeX to highlight the relationships between mrna and mir clusters with recurrence risk and histology (see THCA publication page). The tall cell-like cluster was also identified by SuperCluster (Figure S9E). Integrated mir Analysis Given the increased role of mirs in determining cancer phenotype, we further examined the association between mir expression, molecular subtypes, and clinical parameters. In addition to mir-182 and mir-183 in the RL-PTC group, several other cancer-relevant mirs were relatively abundant in other mir clusters (Figures 7 and S9B). These included oncomirs (mir-21 and mir-146b) and tumor suppressor mirs (let-7 family, mir-204, and mir-375). OncomiRs mir-221 and mir-222 were reported to play a role in PTC aggressiveness (Mardente et al., 2012) and, in our data, were associated with less-differentiated tumors (Figures 7 and S10D, b). We focused integrative analysis on mir-21, mir-146b, and mir-204 because they were epigenetically regulated, correlated with BRS and TDS, and/or differentially expressed between PTCs and normal thyroid, as well as between clusters derived Cell 159, , October 23, 2014 ª2014 The Authors 685
11 from mirna-seq data (Figures S9B, S10A, S10B, and S10F). mir-21 expression correlated with highly variable DNA methylation (Figure S10C), defined the tall cell-like mrna and DNA methylation Meth-classical-1 clusters and was highly correlated with low BRS and TDS values (Figures 7 and S10D). mir-21 is a regulator of several cancer-related genes (Di Leva et al., 2014). In our data, using Regulome Explorer s pairwise associations, its expression was anticorrelated with expression of cancerpromoting genes and regulators of apoptosis (e.g., PDCD4)(Figure S10E). PDCD4 is a mir-21 target gene reported to function as a tumor suppressor in diverse tumors (Zhu et al., 2008). These observations raise the possibility that increased mir-21 expression via epigenetic dysregulation may contribute to the clinically aggressive nature of this BVL-PTC subcluster and may partly explain the aggressive nature of TCV. mir-146b expression exhibited similar patterns of differential expression and correlations to DNA methylation, BRS, and TDS (Figures 7, S10C, and S10D), and likely influenced expression of, for example, IRAK1, KIT and TRAF6 (Figure S10E). These results are consistent with observations that mir-146b is associated with risk of recurrence and promotes cell migration and invasion (Chou et al., 2013). mir-204 expression, while less influenced by DNA methylation, was preferentially lost in mir clusters 5 and 6, the two BVL-PTC subclusters with the lowest BRS and TDS values (Figures 7 and S9B). This is consistent with data from other tumors, i.e., mir-204 functions as a tumor suppressor and high levels suppress cell migration, invasion, and EMT (Qiu et al., 2013). These results suggest that loss of mir-204 may also contribute to aggressive PTCs with BRAF V600E mutations. Collectively, our results are consistent with prior studies and suggest that mirs may regulate fundamental aspects of the PTC phenotype, i.e., signaling, differentiation, invasion and metastasis, by finetuning gene expression. DISCUSSION This study illustrates the dominant role and mutually exclusive nature of driving somatic genetic alterations, be they SSNVs, indels, or fusions, in the MAPK and PI3K pathways in PTC. The relative low overall density of somatic mutations may be the biological basis for the indolent clinical behavior of PTC. We discovered new driver mutations in PTC, either entirely novel in this cancer (EIF1AX) or novel alterations of known drivers (RET, BRAF and ALK fusions). As a result of these discoveries, the dark matter of the PTC genome has been reduced substantially from 25% to less than 4%, which should have profound consequences for preoperative cancer diagnosis in thyroid nodules. Molecular testing of mutation hotspots, rearrangements, and gene expression using fine-needle aspiration specimens has become an effective diagnostic tool to more precisely select patients for thyroid surgery (Alexander et al., 2012; Nikiforov et al., 2011), thereby reducing the number of thyroidectomies done for benign nodules and tumors (Nikiforov et al., 2013), and determining the extent of initial thyroid surgery (i.e., lobectomy versus total thyroidectomy) (Yip et al., 2014). Through these advances, molecular diagnostics has improved the care of patients with thyroid nodules and cancer. Our expansion of the PTC somatic genetic landscape has the potential to even further enhance the care of these patients. This study also offers conclusive evidence that mutated BRAF and other driving mutations are clonal events present in the majority of cells within tumors and identified novel fusion partners of oncogenes (e.g., RET, BRAF, and ALK), expanding the biological basis for targeted therapy. Beyond the driver mutations, we discovered individual genes (CHEK2, ATM, and TERT) and sets of functionally related genes (chromatin remodeling) with alterations or expression patterns (mir-21 and mir-146b) that define clinically-relevant subclasses and may contribute to loss of differentiation and tumor progression. Specifically, increased expression of mir-21 was associated with a known aggressive form of PTC (tall cell variant) and may be a critical event in its pathogenesis. Similarly, TERT promoter mutations identified a subset of aggressive, less-differentiated PTCs, consistent with recent reports (Melo et al., 2014; Xing et al., 2014). Our study also indicates that BRAF V600E PTC represents a diverse group of tumors, consisting of at least four molecular subtypes, with variable degrees of thyroid differentiation. Collectively, our results suggest that BRAF V600E PTC should not be considered a homogeneous group in clinical studies and that future studies should include molecular components designed to capture the breadth of genetic diversity among PTCs. We demonstrate striking signaling differences in RAS- and BRAF V600E -driven PTCs. In particular, BVL-PTCs signal preferentially through MAPK while RL-PTCs signal through both MAPK and PI3K. The relative simplicity of the PTC genome, with dominant mutually exclusive driving events, together with the large cohort and comprehensive data analyzed in this study enabled us to clearly dissect these signaling differences. Our overarching conclusion is that RL-PTCs and BVL-PTCs are fundamentally different in their genomic, epigenomic, and proteomic profiles. This is consistent with their known histological differences and the published literature. However, the breadth and depth of our integrative findings have wide implications for basic pathobiology, tumor classification schemes, and traditional and targeted therapies. This view is supported by recent data suggesting differential response to a MEK inhibitor related to thyroid cancer genotype (Ho et al., 2013). We feel, based on the strength of our multidimensional genomic findings, that a pathologic reclassification of follicular-patterned thyroid lesions is justified. There was a time when follicular-patterned PTCs (i.e., RL-PTCs) were classified as follicular carcinomas. Perhaps the time has come to revise the classification of thyroid cancer to reunite the FV of PTC with follicular carcinomas. Moreover, a refined classification scheme that more accurately reflects the genotypic and phenotypic differences between and within RL and BVL PTCs would lead to more precise surgical and medical therapy, especially as thyroid cancer therapy enters the realm of precision medicine. EXPERIMENTAL PROCEDURES Tumor and normal thyroid samples were obtained from patients with approval from local institutional review boards. DNA, RNA, and protein were purified and distributed throughout the TCGA network. In total, 496 primary tumors and 8 metastatic tumors with associated clinicopathologic data were assayed on at least one molecular profiling platform. Platforms included exome 686 Cell 159, , October 23, 2014 ª2014 The Authors
12 and whole genome DNA sequencing, RNA sequencing, mirna sequencing, SNP arrays, DNA methylation arrays, and reverse phase protein arrays. Integrated multiplatform analyses were performed. The data and analysis results can be explored through the Broad Institute GDAC portal ( org/ /c17p8wzg) and FireBrowse portal ( cohort=thca), Memorial Sloan Kettering Cancer Center cbioportal ( TieDIE ( MBatch batch effects assessor ( Regulome Explorer ( and Next-Generation Clustered Heat Maps ( See also Supplemental Information and the THCA publication page ( nci.nih.gov/docs/publications/thca_2014/). Data Access The primary and processed data used to generate the analyses presented here can be downloaded by registered users from The Cancer Genome Atlas at The primary sequence files are deposited in CGHub ( and all other molecular, clinical and pathological data are deposited at the Data Coordinating Center (DCC) for public access ( cghub.ucsc.edu/ and /) and the TCGA Data Portal ( SUPPLEMENTAL INFORMATION Supplemental Information includes Extended Experimental Procedures, ten figures, six tables, and one data file and can be found with this article online at CONSORTIA The members of The Cancer Genome Atlas Research Network for this project are Nishant Agrawal, Rehan Akbani, B. Arman Aksoy, Adrian Ally, Harindra Arachchi, Sylvia L. Asa, J. Todd Auman, Miruna Balasundaram, Saianand Balu, Stephen B. Baylin, Madhusmita Behera, Brady Bernard, Rameen Beroukhim, Justin A. Bishop, Aaron D. Black, Tom Bodenheimer, Lori Boice, Moiz S. Bootwalla, Jay Bowen, Reanne Bowlby, Christopher A. Bristow, Robin Brookens, Denise Brooks, Robert Bryant, Elizabeth Buda, Yaron S.N. Butterfield, Tobias Carling, Rebecca Carlsen, Scott L. Carter, Sally E. Carty, Timothy A. Chan, Amy Y. Chen, Andrew D. Cherniack, Dorothy Cheung, Lynda Chin, Juok Cho, Andy Chu, Eric Chuah, Kristian Cibulskis, Giovanni Ciriello, Amanda Clarke, Gary L. Clayman, Leslie Cope, John A. Copland, Kyle Covington, Ludmila Danilova, Tanja Davidsen, John A. Demchok, Daniel DiCara, Noreen Dhalla, Rajiv Dhir, Sheliann S. Dookran, Gideon Dresdner, Jonathan Eldridge, Greg Eley, Adel K. El-Naggar, Stephanie Eng, James A. Fagin, Timothy Fennell, Robert L. Ferris, Sheila Fisher, Scott Frazer, Jessica Frick, Stacey B. Gabriel, Ian Ganly, Jianjiong Gao, Levi A. Garraway, Julie M. Gastier-Foster, Gad Getz, Nils Gehlenborg, Ronald Ghossein, Richard A. Gibbs, Thomas J. Giordano, Karen Gomez-Hernandez, Jonna Grimsby, Benjamin Gross, Ranabir Guin, Angela Hadjipanayis, Hollie A. Harper, D. Neil Hayes, David I. Heiman, James G. Herman, Katherine A. Hoadley, Matan Hofree, Robert A. Holt, Alan P. Hoyle, Franklin W. Huang, Mei Huang, Carolyn M. Hutter, Trey Ideker, Lisa Iype, Anders Jacobsen, Stuart R. Jefferys, Corbin D. Jones, Steven J.M. Jones, Katayoon Kasaian, Electron Kebebew, Fadlo R. Khuri, Jaegil Kim, Roger Kramer, Richard Kreisberg, Raju Kucherlapati, David J. Kwiatkowski, Marc Ladanyi, Phillip H. Lai, Peter W. Laird, Eric Lander, Michael S. Lawrence, Darlene Lee, Eunjung Lee, Semin Lee, William Lee, Kristen M. Leraas, Tara M. Lichtenberg, Lee Lichtenstein, Pei Lin, Shiyun Ling, Jinze Liu, Wenbin Liu, Yingchun Liu, Virginia A. LiVolsi, Yiling Lu, Yussanne Ma, Harshad S. Mahadeshwar, Marco A. Marra, Michael Mayo, David G. McFadden, Shaowu Meng, Matthew Meyerson, Piotr A. Mieczkowski, Michael Miller, Gordon Mills, Richard A. Moore, Lisle E. Mose, Andrew J. Mungall, Bradley A. Murray, Yuri E. Nikiforov, Michael S. Noble, Akinyemi I. Ojesina, Taofeek K. Owonikoko, Bradley A. Ozenberger, Angeliki Pantazi, Michael Parfenov, Peter J. Park, Joel S. Parker, Evan O. Paull, Chandra Sekhar Pedamallu, Charles M. Perou, Jan F. Prins, Alexei Protopopov, Suresh S. Ramalingam, Nilsa C. Ramirez, Ricardo Ramirez, Benjamin J. Raphael, W. Kimryn Rathmell, Xiaojia Ren, Sheila M. Reynolds, Esther Rheinbay, Matthew D. Ringel, Michael Rivera, Jeffrey Roach, A. Gordon Robertson, Mara W. Rosenberg, Matthew Rosenthal, Sara Sadeghi, Gordon Saksena, Chris Sander, Netty Santoso, Jacqueline E. Schein, Nikolaus Schultz, Steven E. Schumacher, Raja R. Seethala, Jonathan Seidman, Yasin Senbabaoglu, Sahil Seth, Samantha Sharpe, Kenna R. Mills Shaw, John P. Shen, Ronglai Shen, Steven Sherman, Margi Sheth, Yan Shi, Ilya Shmulevich, Gabriel L. Sica, Janae V. Simons, Rileen Sinha, Payal Sipahimalani, Robert C. Smallridge, Heidi J. Sofia, Matthew G. Soloway, Xingzhi Song, Carrie Sougnez, Chip Stewart, Petar Stojanov, Joshua M. Stuart, S. Onur Sumer, Yichao Sun, Barbara Tabak, Angela Tam, Donghui Tan, Jiabin Tang, Roy Tarnuzzer, Barry S. Taylor, Nina Thiessen, Leigh Thorne, Vésteinn Thorsson, R. Michael Tuttle, Christopher B. Umbricht, David J. Van Den Berg, Fabio Vandin, Umadevi Veluvolu, Roel G.W. Verhaak, Michelle Vinco, Doug Voet, Vonn Walter, Zhining Wang, Scot Waring, Paul M. Weinberger, Nils Weinhold, John N. Weinstein, Daniel J. Weisenberger, David Wheeler, Matthew D. Wilkerson, Jocelyn Wilson, Michelle Williams, Daniel A. Winer, Lisa Wise, Junyuan Wu, Liu Xi, Andrew W. Xu, Liming Yang, Lixing Yang, Travis I. Zack, Martha A. Zeiger, Dong Zeng, Jean Claude Zenklusen, Ni Zhao, Hailei Zhang, Jianhua Zhang, Jiashan (Julia) Zhang, Wei Zhang, Erik Zmuda, Lihua Zou. AUTHOR CONTRIBUTIONS Project leaders: G.G and T.J.G. Analysis coordinator: C.S. Data coordinator: J.C. Manuscript coordinator: T.J.G. Project coordinators: M.S. and J.Z. Clinical expertise and data analysis: S.L.A., J.B., J.A.F., I.G., T.J.G., E.K., L.I., K.L., T.M.L., D.G.M., A.P., M.D.R., R.C.S., C.B.U., Y.E.N., and M.A.Z. Supplemental pathology review: S.L.A., T.J.G., R.G., J.A.B., and Y.E.N. DNA sequence and copy-number analysis: A.D.C., J.C., G.G., K.K., J.K., D.J.K., L.L., B.A.M., E.R., M.R., G.S., C.S., and C.S. TERT promoter sequencing and analysis: C.S., C.S., J.G., F.W.H., L.A.G., S.D.D., Y.E.N., A.H., R.K., and G.G. Gene fusions: A.H., K.A.H., R.K., C.S., and T.J.G. Viral sequence detection: A.I.O., M.M., C.S.P., S.S., and Y.M. DNA methylation analysis: L.C. and L.D. mrna analysis: K.A.H., C.S., V.W., N.Z., and A.H. mirna analysis: L.I. and A.G.R. Pathway/Network/Integrated analysis: G.C., J.A.F., N.G., M.H., L.I., E.O.P., B.J.R., M.D.R., A.G.R., J.M.S., N.S., Y.S., and F.V. Dark matter analysis: J.C., C.S., and T.J.G. Thyroid differentiation analysis: J.K., L.D., L.I., A.G.R., C.S., G.G., and T.J.G. RPPA analysis: R.A., W.L., Y.L., and G.B.M. Supercluster: R.A. Batch effect analysis: R.A., S.L., and J.N.W. Writing committee: T.J.G., G.G., C.S., J.A.F., S.L.A., D.G.M., Y.E.N., M.D.R., I.G., R.C.S., and M.A.Z. Manuscript review: M.M. and R.K. ACKNOWLEDGMENTS We are grateful to all the patients and families who contributed to this study, to Chris Gunter for editing, and Margi Sheth and Jiashan (Julia) Zhang for project management. Thomas Giordano thanks his colleagues who covered his clinical duties. Supported by the following grants from the United States National Institutes of Health: 5U24CA143799, 5U24CA143835, 5U24CA143840, 5U24CA143843, 5U24CA143845, 5U24CA143848, 5U24CA143858, 5U24CA143866, 5U24CA143867, 5U24CA143882, 5U24CA143883, 5U24CA144025, U54HG003067, U54HG003079, and U54HG003273, P30CA Yuri Nikiforov is a consultant for Quest Diagnostics. Raju Kucherlapati serves on the Board of Directors of KEW Group, Inc. Daniel J. Weisenberger is a consultant for Zymo Research Corporation. Received: May 11, 2014 Revised: September 16, 2014 Accepted: September 23, 2014 Published: October 23, 2014 REFERENCES Adeniran, A.J., Zhu, Z., Gandhi, M., Steward, D.L., Fidler, J.P., Giordano, T.J., Biddinger, P.W., and Nikiforov, Y.E. (2006). Correlation between genetic Cell 159, , October 23, 2014 ª2014 The Authors 687
13 alterations and microscopic features, clinical manifestations, and prognostic characteristics of thyroid papillary carcinomas. Am. J. Surg. Pathol. 30, Alexander, E.K., Kennedy, G.C., Baloch, Z.W., Cibas, E.S., Chudova, D., Diggans, J., Friedman, L., Kloos, R.T., LiVolsi, V.A., Mandel, S.J., et al. (2012). Preoperative diagnosis of benign thyroid nodules with indeterminate cytology. N. Engl. J. Med. 367, American Thyroid Association (ATA) Guidelines Taskforce on Thyroid Nodules and Differentiated Thyroid Cancer, Cooper, D.S., Doherty, G.M., Haugen, B.R., Kloos, R.T., Lee, S.L., Mandel, S.J., Mazzaferri, E.L., McIver, B., Pacini, F., Schlumberger, M., et al. (2009). Revised American Thyroid Association management guidelines for patients with thyroid nodules and differentiated thyroid cancer. Thyroid 19, Bischoff, L.A., Curry, J., Ahmed, I., Pribitkin, E., and Miller, J.L. (2013). Is above age 45 appropriate for upstaging well-differentiated papillary thyroid cancer? Endocr. Pract. 19, Cancer Genome Atlas Research Network (2014). Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, Carter, S.L., Cibulskis, K., Helman, E., McKenna, A., Shen, H., Zack, T., Laird, P.W., Onofrio, R.C., Winckler, W., Weir, B.A., et al. (2012). Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, Chakravarty, D., Santos, E., Ryder, M., Knauf, J.A., Liao, X.H., West, B.L., Bollag, G., Kolesnick, R., Thin, T.H., Rosen, N., et al. (2011). Small-molecule MAPK inhibitors restore radioiodine incorporation in mouse thyroid cancers with conditional BRAF activation. J. Clin. Invest. 121, Charles, R.P., Iezza, G., Amendola, E., Dankort, D., and McMahon, M. (2011). Mutationally activated BRAF(V600E) elicits papillary thyroid cancer in the adult mouse. Cancer Res. 71, Chen, A.Y., Jemal, A., and Ward, E.M. (2009). Increasing incidence of differentiated thyroid cancer in the United States, Cancer 115, Chou, C.K., Yang, K.D., Chou, F.F., Huang, C.C., Lan, Y.W., Lee, Y.F., Kang, H.Y., and Liu, R.T. (2013). Prognostic implications of mir-146b expression and its functional role in papillary thyroid carcinoma. J. Clin. Endocrinol. Metab. 98, E196 E205. Chu, J., Sadeghi, S., Raymond, A., Jackman, S.D., Nip, K.M., Mar, R., Mohamadi, H., Butterfield, Y.S., Robertson, A.G., and Birol, I. (2014). BioBloom tools: fast, accurate and memory-efficient host species sequence screening using bloom filters. Bioinformatics. Published online August 20, dx.doi.org/ /bioinformatics/btu558. Ciampi, R., Knauf, J.A., Kerler, R., Gandhi, M., Zhu, Z., Nikiforova, M.N., Rabes, H.M., Fagin, J.A., and Nikiforov, Y.E. (2005). Oncogenic AKAP9- BRAF fusion is a novel mechanism of MAPK pathway activation in thyroid cancer. J. Clin. Invest. 115, Ciriello, G., Cerami, E., Sander, C., and Schultz, N. (2012). Mutual exclusivity analysis identifies oncogenic network modules. Genome Res. 22, Cohen, Y., Xing, M., Mambo, E., Guo, Z., Wu, G., Trink, B., Beller, U., Westra, W.H., Ladenson, P.W., and Sidransky, D. (2003). BRAF mutation in papillary thyroid carcinoma. J. Natl. Cancer Inst. 95, Di Leva, G., Garofalo, M., and Croce, C.M. (2014). MicroRNAs in cancer. Annu. Rev. Pathol. 9, Durante, C., Puxeddu, E., Ferretti, E., Morisi, R., Moretti, S., Bruno, R., Barbi, F., Avenia, N., Scipioni, A., Verrienti, A., et al. (2007). BRAF mutations in papillary thyroid carcinomas inhibit genes involved in iodine metabolism. J. Clin. Endocrinol. Metab. 92, Ellis, R.J., Wang, Y., Stevenson, H.S., Boufraqech, M., Patel, D., Nilubol, N., Davis, S., Edelman, D.C., Merino, M.J., He, M., et al. (2014). Genome-wide methylation patterns in papillary thyroid cancer are distinct based on histological subtype and tumor genotype. J. Clin. Endocrinol. Metab. 99, E329 E337. Forbes, S.A., Bindal, N., Bamford, S., Cole, C., Kok, C.Y., Beare, D., Jia, M., Shepherd, R., Leung, K., Menzies, A., et al. (2011). COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 39 (Database issue), D945 D950. Franco, A.T., Malaguarnera, R., Refetoff, S., Liao, X.H., Lundsmith, E., Kimura, S., Pritchard, C., Marais, R., Davies, T.F., Weinstein, L.S., et al. (2011). Thyrotrophin receptor signaling dependence of Braf-induced thyroid tumor initiation in mice. Proc. Natl. Acad. Sci. USA 108, Ghossein, R.A., Katabi, N., and Fagin, J.A. (2013). Immunohistochemical detection of mutated BRAF V600E supports the clonal origin of BRAF-induced thyroid cancers along the spectrum of disease progression. J. Clin. Endocrinol. Metab. 98, E1414 E1421. Giordano, T.J., Kuick, R., Thomas, D.G., Misek, D.E., Vinco, M., Sanders, D., Zhu, Z., Ciampi, R., Roh, M., Shedden, K., et al. (2005). Molecular classification of papillary thyroid carcinoma: distinct BRAF, RAS, and RET/PTC mutationspecific gene expression profiles discovered by DNA microarray analysis. Oncogene 24, Grieco, M., Santoro, M., Berlingieri, M.T., Melillo, R.M., Donghi, R., Bongarzone, I., Pierotti, M.A., Della Porta, G., Fusco, A., and Vecchio, G. (1990). PTC is a novel rearranged form of the ret proto-oncogene and is frequently detected in vivo in human thyroid papillary carcinomas. Cell 60, Groenewoud, M.J., and Zwartkruis, F.J. (2013). Rheb and mammalian target of rapamycin in mitochondrial homoeostasis. Open boil. 3, Guerra, A., Sapio, M.R., Marotta, V., Campanile, E., Rossi, S., Forno, I., Fugazzola, L., Budillon, A., Moccia, T., Fenzi, G., and Vitale, M. (2012). The primary occurrence of BRAF(V600E) is a rare clonal event in papillary thyroid carcinoma. J. Clin. Endocrinol. Metab. 97, Hay, I.D., Bergstralh, E.J., Goellner, J.R., Ebersold, J.R., and Grant, C.S. (1993). Predicting outcome in papillary thyroid carcinoma: development of a reliable prognostic scoring system in a cohort of 1779 patients surgically treated at one institution during 1940 through Surgery 114, , discussion Hay, I.D., Thompson, G.B., Grant, C.S., Bergstralh, E.J., Dvorak, C.E., Gorman, C.A., Maurer, M.S., McIver, B., Mullan, B.P., Oberg, A.L., et al. (2002). Papillary thyroid carcinoma managed at the Mayo Clinic during six decades ( ): temporal trends in initial therapy and long-term outcome in 2444 consecutively treated patients. World J. Surg. 26, Ho, A.L., Grewal, R.K., Leboeuf, R., Sherman, E.J., Pfister, D.G., Deandreis, D., Pentlow, K.S., Zanzonico, P.B., Haque, S., Gavane, S., et al. (2013). Selumetinib-enhanced radioiodine uptake in advanced thyroid cancer. N. Engl. J. Med. 368, Hofree, M., Shen, J.P., Carter, H., Gross, A., and Ideker, T. (2013). Networkbased stratification of tumor mutations. Nat. Methods 10, Hutchinson, K.E., Lipson, D., Stephens, P.J., Otto, G., Lehmann, B.D., Lyle, P.L., Vnencak-Jones, C.L., Ross, J.S., Pietenpol, J.A., Sosman, J.A., et al. (2013). BRAF fusions define a distinct molecular subset of melanomas with potential sensitivity to MEK inhibition. Clin. Cancer Res. 19, Imam, J.S., Plyler, J.R., Bansal, H., Prajapati, S., Bansal, S., Rebeles, J., Chen, H.I., Chang, Y.F., Panneerdoss, S., Zoghi, B., et al. (2012). Genomic loss of tumor suppressor mirna-204 promotes cancer cell migration and invasion by activating AKT/mTOR/Rac1 signaling and actin reorganization. PLoS ONE 7, e Jung, C.K., Little, M.P., Lubin, J.H., Brenner, A.V., Wells, S.A., Jr., Sigurdson, A.J., and Nikiforov, Y.E. (2014). The increase in thyroid cancer incidence during the last four decades is accompanied by a high frequency of BRAF mutations and a sharp increase in RAS mutations. J. Clin. Endocrinol. Metab. 99, E276 E285. Kelly, L.M., Barila, G., Liu, P., Evdokimova, V.N., Trivedi, S., Panebianco, F., Gandhi, M., Carty, S.E., Hodak, S.P., Luo, J., et al. (2014). Identification of the transforming STRN-ALK fusion as a potential therapeutic target in the aggressive forms of thyroid cancer. Proc. Natl. Acad. Sci. USA 111, Kimura, E.T., Nikiforova, M.N., Zhu, Z., Knauf, J.A., Nikiforov, Y.E., and Fagin, J.A. (2003). High prevalence of BRAF mutations in thyroid cancer: genetic evidence for constitutive activation of the RET/PTC-RAS-BRAF signaling pathway in papillary thyroid carcinoma. Cancer Res. 63, Cell 159, , October 23, 2014 ª2014 The Authors
14 Kjellman, P., Lagercrantz, S., Höög, A., Wallin, G., Larsson, C., and Zedenius, J. (2001). Gain of 1q and loss of 9q21.3-q32 are associated with a less favorable prognosis in papillary thyroid carcinoma. Genes Chromosomes Cancer 32, Kleiblova, P., Shaltiel, I.A., Benada, J., Sevcík, J., Pechácková, S., Pohlreich, P., Voest, E.E., Dundr, P., Bartek, J., Kleibl, Z., et al. (2013). Gain-of-function mutations of PPM1D/Wip1 impair the p53-dependent G1 checkpoint. J. Cell Biol. 201, Kostic, A.D., Ojesina, A.I., Pedamallu, C.S., Jung, J., Verhaak, R.G., Getz, G., and Meyerson, M. (2011). PathSeq: software to identify or discover microbes by deep sequencing of human tissue. Nat. Biotechnol. 29, Kroll, T.G., Sarraf, P., Pecciarini, L., Chen, C.J., Mueller, E., Spiegelman, B.M., and Fletcher, J.A. (2000). PAX8-PPARgamma1 fusion oncogene in human thyroid carcinoma [corrected]. Science 289, Lawrence, M.S., Stojanov, P., Polak, P., Kryukov, G.V., Cibulskis, K., Sivachenko, A., Carter, S.L., Stewart, C., Mermel, C.H., Roberts, S.A., et al. (2013). Mutational heterogeneity in cancer and the search for new cancerassociated genes. Nature 499, Lawrence, M.S., Stojanov, P., Mermel, C.H., Robinson, J.T., Garraway, L.A., Golub, T.R., Meyerson, M., Gabriel, S.B., Lander, E.S., and Getz, G. (2014). Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, Lee, N.V., Lira, M.E., Pavlicek, A., Ye, J., Buckman, D., Bagrodia, S., Srinivasa, S.P., Zhao, Y., Aparicio, S., Rejto, P.A., et al. (2012). A novel SND1-BRAF fusion confers resistance to c-met inhibitor PF in GTL16 cells through [corrected] MAPK activation. PLoS ONE 7, e Lemoine, N.R., Mayall, E.S., Wyllie, F.S., Farr, C.J., Hughes, D., Padua, R.A., Thurston, V., Williams, E.D., and Wynford-Thomas, D. (1988). Activated ras oncogenes in human thyroid cancers. Cancer Res. 48, Mardente, S., Mari, E., Consorti, F., Di Gioia, C., Negri, R., Etna, M., Zicari, A., and Antonaci, A. (2012). HMGB1 induces the overexpression of mir-222 and mir-221 and increases growth and motility in papillary thyroid cancer cells. Oncol. Rep. 28, Martin, M., Maßhöfer, L., Temming, P., Rahmann, S., Metz, C., Bornfeld, N., van de Nes, J., Klein-Hitpass, L., Hinnebusch, A.G., Horsthemke, B., et al. (2013). Exome sequencing identifies recurrent somatic mutations in EIF1AX and SF3B1 in uveal melanoma with disomy 3. Nat. Genet. 45, Melo, M., da Rocha, A.G., Vinagre, J., Batista, R., Peixoto, J., Tavares, C., Celestino, R., Almeida, A., Salgado, C., Eloy, C., et al. (2014). TERT promoter mutations are a major indicator of poor outcome in differentiated thyroid carcinomas. J. Clin. Endocrinol. Metab. 99, E754 E765. Mermel, C.H., Schumacher, S.E., Hill, B., Meyerson, M.L., Beroukhim, R., and Getz, G. (2011). GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41. Minamiya, Y., Saito, H., Ito, M., Imai, K., Konno, H., Takahashi, N., Motoyama, S., and Ogawa, J. (2012). Suppression of Zinc Finger Homeobox 3 expression in tumor cells decreases the survival rate among non-small cell lung cancer patients. Cancer Biomark. 11, Mitsutake, N., Miyagishi, M., Mitsutake, S., Akeno, N., Mesa, C., Jr., Knauf, J.A., Zhang, L., Taira, K., and Fagin, J.A. (2006). BRAF mediates RET/PTCinduced mitogen-activated protein kinase activation in thyroid cells: functional support for requirement of the RET/PTC-RAS-BRAF pathway in papillary thyroid carcinogenesis. Endocrinology 147, Nikiforov, Y.E., Ohori, N.P., Hodak, S.P., Carty, S.E., LeBeau, S.O., Ferris, R.L., Yip, L., Seethala, R.R., Tublin, M.E., Stang, M.T., et al. (2011). Impact of mutational testing on the diagnosis and management of patients with cytologically indeterminate thyroid nodules: a prospective analysis of 1056 FNA samples. J. Clin. Endocrinol. Metab. 96, Nikiforov, Y.E., Yip, L., and Nikiforova, M.N. (2013). New strategies in diagnosing cancer in thyroid nodules: impact of molecular markers. Clin. Cancer Res. 19, Nucera, C., Lawler, J., and Parangi, S. (2011). BRAF(V600E) and microenvironment in thyroid cancer: a functional link to drive cancer progression. Cancer Res. 71, Oliva-Trastoy, M., Berthonaud, V., Chevalier, A., Ducrot, C., Marsolier- Kergoat, M.C., Mann, C., and Leteurtre, F. (2007). The Wip1 phosphatase (PPM1D) antagonizes activation of the Chk2 tumour suppressor kinase. Oncogene 26, Park, J.Y., Kim, W.Y., Hwang, T.S., Lee, S.S., Kim, H., Han, H.S., Lim, S.D., Kim, W.S., Yoo, Y.B., and Park, K.S. (2013). BRAF and RAS mutations in follicular variants of papillary thyroid carcinoma. Endocr. Pathol. 24, Paull, E.O., Carlin, D.E., Niepel, M., Sorger, P.K., Haussler, D., and Stuart, J.M. (2013). Discovering causal pathways linking genomic events to transcriptional states using Tied Diffusion Through Interacting Events (TieDIE). Bioinformatics 29, Pierotti, M.A., Bongarzone, I., Borrello, M.G., Mariani, C., Miranda, C., Sozzi, G., and Greco, A. (1995). Rearrangements of TRK proto-oncogene in papillary thyroid carcinomas. J. Endocrinol. Invest. 18, Poulikakos, P.I., Zhang, C., Bollag, G., Shokat, K.M., and Rosen, N. (2010). RAF inhibitors transactivate RAF dimers and ERK signalling in cells with wild-type BRAF. Nature 464, Pratilas, C.A., Taylor, B.S., Ye, Q., Viale, A., Sander, C., Solit, D.B., and Rosen, N. (2009). (V600E)BRAF is associated with disabled feedback inhibition of RAF-MEK signaling and elevated transcriptional output of the pathway. Proc. Natl. Acad. Sci. USA 106, Qiu, Y.H., Wei, Y.P., Shen, N.J., Wang, Z.C., Kan, T., Yu, W.L., Yi, B., and Zhang, Y.J. (2013). mir-204 inhibits epithelial to mesenchymal transition by targeting slug in intrahepatic cholangiocarcinoma cells. Cell. Physiol. Biochem. 32, Ricarte-Filho, J.C., Li, S., Garcia-Rendueles, M.E., Montero-Conde, C., Voza, F., Knauf, J.A., Heguy, A., Viale, A., Bogdanova, T., Thomas, G.A., et al. (2013). Identification of kinase fusion oncogenes in post-chernobyl radiation-induced thyroid cancers. J. Clin. Invest. 123, Roberts, S.A., Lawrence, M.S., Klimczak, L.J., Grimm, S.A., Fargo, D., Stojanov, P., Kiezun, A., Kryukov, G.V., Carter, S.L., Saksena, G., et al. (2013). An APOBEC cytidine deaminase mutagenesis pattern is widespread in human cancers. Nat. Genet. 45, Sabra, M.M., Dominguez, J.M., Grewal, R.K., Larson, S.M., Ghossein, R.A., Tuttle, R.M., and Fagin, J.A. (2013). Clinical outcomes and molecular profile of differentiated thyroid cancers with radioiodine-avid distant metastases. J. Clin. Endocrinol. Metab. 98, E829 E836. Soares, P., Trovisco, V., Rocha, A.S., Lima, J., Castro, P., Preto, A., Máximo, V., Botelho, T., Seruca, R., and Sobrinho-Simões, M. (2003). BRAF mutations and RET/PTC rearrangements are alternative events in the etiopathogenesis of PTC. Oncogene 22, Streit, M., Lex, A., Gratzl, S., Partl, C., Schmalstieg, D., Pfister, H., Park, P.J., and Gehlenborg, N. (2014). Guided visual exploration of genomic stratifications in cancer. Nat. Methods 11, Suárez, H.G., Du Villard, J.A., Caillou, B., Schlumberger, M., Tubiana, M., Parmentier, C., and Monier, R. (1988). Detection of activated ras oncogenes in human thyroid carcinomas. Oncogene 2, Woiwode, A., Johnson, S.A., Zhong, S., Zhang, C., Roeder, R.G., Teichmann, M., and Johnson, D.L. (2008). PTEN represses RNA polymerase III-dependent transcription by targeting the TFIIIB complex. Mol. Cell. Biol. 28, Wreesmann, V.B., Ghossein, R.A., Hezel, M., Banerjee, D., Shaha, A.R., Tuttle, R.M., Shah, J.P., Rao, P.H., and Singh, B. (2004a). Follicular variant of papillary thyroid carcinoma: genome-wide appraisal of a controversial entity. Genes Chromosomes Cancer 40, Wreesmann, V.B., Sieczka, E.M., Socci, N.D., Hezel, M., Belbin, T.J., Childs, G., Patel, S.G., Patel, K.N., Tallini, G., Prystowsky, M., et al. (2004b). Genome-wide profiling of papillary thyroid cancer identifies MUC1 as an independent prognostic marker. Cancer Res. 64, Xing, M. (2013). Molecular pathogenesis and mechanisms of thyroid cancer. Nat. Rev. Cancer 13, Cell 159, , October 23, 2014 ª2014 The Authors 689
15 Xing, M., Alzahrani, A.S., Carson, K.A., Viola, D., Elisei, R., Bendlova, B., Yip, L., Mian, C., Vianello, F., Tuttle, R.M., et al. (2013). Association between BRAF V600E mutation and mortality in patients with papillary thyroid cancer. JAMA 309, Xing, M., Liu, R., Liu, X., Murugan, A.K., Zhu, G., Zeiger, M.A., Pai, S., and Bishop, J. (2014). BRAF V600E and TERT Promoter Mutations Cooperatively Identify the Most Aggressive Papillary Thyroid Cancer With Highest Recurrence. J. Clin. Oncol. 32, Yip, L., Wharry, L.I., Armstrong, M.J., Silbermann, A., McCoy, K.L., Stang, M.T., Ohori, N.P., LeBeau, S.O., Coyne, C., Nikiforova, M.N., et al. (2014). A clinical algorithm for fine-needle aspiration molecular testing effectively guides the appropriate extent of initial thyroidectomy. Ann. Surg. 260, Zhu, S., Wu, H., Wu, F., Nie, D., Sheng, S., and Mo, Y.Y. (2008). MicroRNA-21 targets tumor suppressor genes in invasion and metastasis. Cell Res. 18, Cell 159, , October 23, 2014 ª2014 The Authors
16 Supplemental Information SUPPLEMENTAL REFERENCES Altschul, S.F., Madden, T.L., Schäffer, A.A., Zhang, J., Zhang, Z., Miller, W., and Lipman, D.J. (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, Ashburner, M., Ball, C.A., Blake, J.A., Botstein, D., Butler, H., Cherry, J.M., Davis, A.P., Dolinski, K., Dwight, S.S., Eppig, J.T., et al.; The Gene Ontology Consortium (2000). Gene ontology: tool for the unification of biology. Nat. Genet. 25, Benjamini, Y., and Hochbeg, Y. (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc., B 57, 289. Berger, M.F., Hodis, E., Heffernan, T.P., Deribe, Y.L., Lawrence, M.S., Protopopov, A., Ivanova, E., Watson, I.R., Nickerson, E., Ghosh, P., et al. (2012). Melanoma genome sequencing reveals frequent PREX2 mutations. Nature 485, Bibikova, M., Barnes, B., Tsan, C., Ho, V., Klotzle, B., Le, J.M., Delano, D., Zhang, L., Schroth, G.P., Gunderson, K.L., et al. (2011). High density DNA methylation array with single CpG site resolution. Genomics 98, Byers, L.A., Diao, L., Wang, J., Saintigny, P., Girard, L., Peyton, M., Shen, L., Fan, Y., Giri, U., Tumula, P.K., et al. (2013). An epithelial-mesenchymal transition gene signature predicts resistance to EGFR and PI3K inhibitors and identifies Axl as a therapeutic target for overcoming EGFR inhibitor resistance. Clin. Cancer Res. 19, Cancer Genome Atlas, N.; Cancer Genome Atlas Network (2012). Comprehensive molecular portraits of human breast tumours. Nature 490, Cancer Genome Atlas Research Network (2011). Integrated genomic analyses of ovarian carcinoma. Nature 474, Cancer Genome Atlas Research Network (2013). Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499, Chen, K., Wallis, J.W., McLellan, M.D., Larson, D.E., Kalicki, J.M., Pohl, C.S., McGrath, S.D., Wendl, M.C., Zhang, Q., Locke, D.P., et al. (2009). BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nat. Methods 6, Chiang, D.Y., Getz, G., Jaffe, D.B., O Kelly, M.J., Zhao, X., Carter, S.L., Russ, C., Nusbaum, C., Meyerson, M., and Lander, E.S. (2009). High-resolution mapping of copy-number alterations with massively parallel sequencing. Nat. Methods 6, Cibulskis, K., Lawrence, M.S., Carter, S.L., Sivachenko, A., Jaffe, D., Sougnez, C., Gabriel, S., Meyerson, M., Lander, E.S., and Getz, G. (2013). Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, Cibulskis, K., McKenna, A., Fennell, T., Banks, E., DePristo, M., and Getz, G. (2011). ContEst: estimating cross-contamination of human samples in next-generation sequencing data. Bioinformatics 27, Cingolani, P., Platts, A., Wang, L., Coon, M., Nguyen, T., Wang, L., Land, S.J., Lu, X., and Ruden, D.M. (2012). A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, Coombes, K., Neeley, E.S., and Joy, C. (2011). SuperCurve: SuperCurve Package. R package version Costello, M., Pugh, T.J., Fennell, T.J., Stewart, C., Lichtenstein, L., Meldrim, J.C., Fostel, J.L., Friedrich, D.C., Perrin, D., Dionne, D., et al. (2013). Discovery and characterization of artifactual mutations in deep coverage targeted capture sequencing data due to oxidative DNA damage during sample preparation. Nucleic Acids Res. 41, e67. Dabney, A.R. (2006). ClaNC: point-and-click software for classifying microarrays to nearest centroids. Bioinformatics 22, Das, J., and Yu, H. (2012). HINT: High-quality protein interactomes and their applications in understanding human disease. BMC Syst. Biol. 6, 92. de Hoon, M.J., Imoto, S., Nolan, J., and Miyano, S. (2004). Open source clustering software. Bioinformatics 20, Drier, Y., Lawrence, M.S., Carter, S.L., Stewart, C., Gabriel, S.B., Lander, E.S., Meyerson, M., Beroukhim, R., and Getz, G. (2013). Somatic rearrangements across cancer reveal classes of samples with distinct patterns of DNA breakage and rearrangement-induced hypermutability. Genome Res. 23, Dulak, A.M., Stojanov, P., Peng, S., Lawrence, M.S., Fox, C., Stewart, C., Bandla, S., Imamura, Y., Schumacher, S.E., Shefler, E., et al. (2013). Exome and wholegenome sequencing of esophageal adenocarcinoma identifies recurrent driver events and mutational complexity. Nat. Genet. 45, Forbes, S.A., Tang, G., Bindal, N., Bamford, S., Dawson, E., Cole, C., Kok, C.Y., Jia, M., Ewing, R., Menzies, A., et al. (2010). COSMIC (the Catalogue of Somatic Mutations in Cancer): a resource to investigate acquired mutations in human cancer. Nucleic Acids Res. 38 (Database issue), D652 D657. Gaujoux, R., and Seoighe, C. (2010). A flexible R package for nonnegative matrix factorization. BMC Bioinformatics 11, 367. Getz, G., Höfling, H., Mesirov, J.P., Golub, T.R., Meyerson, M., Tibshirani, R., and Lander, E.S. (2007). Comment on The consensus coding sequences of human breast and colorectal cancers. Science 317, Gonzalez-Angulo, A.M., Hennessy, B.T., Meric-Bernstam, F., Sahin, A., Liu, W., Ju, Z., Carey, M.S., Myhre, S., Speers, C., Deng, L., et al. (2011). Functional proteomics can define prognosis and predict pathologic complete response in patients with breast cancer. Clin. Proteomics 8, 11. Helwak, A., Kudla, G., Dudnakova, T., and Tollervey, D. (2013). Mapping the human mirna interactome by CLASH reveals frequent noncanonical binding. Cell 153, Hennessy, B.T., Lu, Y., Gonzalez-Angulo, A.M., Carey, M.S., Myhre, S., Ju, Z., Davies, M.A., Liu, W., Coombes, K., Meric-Bernstam, F., et al. (2010). A Technical Assessment of the Utility of Reverse Phase Protein Arrays for the Study of the Functional Proteome in Non-microdissected Human Breast Cancers. Clin. Proteomics 6, Hennessy, B.T., Lu, Y., Poradosu, E., Yu, Q., Yu, S., Hall, H., Carey, M.S., Ravoori, M., Gonzalez-Angulo, A.M., Birch, R., et al. (2007). Pharmacodynamic markers of perifosine efficacy. Clinical cancer research 13, Hollants, S., Redeker, E.J., and Matthijs, G. (2012). Microfluidic amplification as a tool for massive parallel sequencing of the familial hypercholesterolemia genes. Clin. Chem. 58, Hsu, S.D., Tseng, Y.T., Shrestha, S., Lin, Y.L., Khaleel, A., Chou, C.H., Chu, C.F., Huang, H.Y., Lin, C.M., Ho, S.Y., et al. (2014). mirtarbase update 2014: an information resource for experimentally validated mirna-target interactions. Nucleic Acids Res. 42 (Database issue), D78 D85. Hu, J., He, X., Baggerly, K.A., Coombes, K.R., Hennessy, B.T., and Mills, G.B. (2007). Non-parametric quantification of protein lysate arrays. Bioinformatics 23, Huang, da W., Sherman, B.T., Zheng, X., Yang, J., Imamichi, T., Stephens, R., and Lempicki, R.A. (2009). Extracting biological meaning from large gene lists with DAVID. Current protoc. bioinformatics Chapter 13, Unit Cell 159, , October 23, 2014 ª2014 The Authors S1
17 Huang, F.W., Hodis, E., Xu, M.J., Kryukov, G.V., Chin, L., and Garraway, L.A. (2013). Highly recurrent TERT promoter mutations in human melanoma. Science 339, Kanehisa, M., and Goto, S. (2000). KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, Korn, J.M., Kuruvilla, F.G., McCarroll, S.A., Wysoker, A., Nemesh, J., Cawley, S., Hubbell, E., Veitch, J., Collins, P.J., Darvishi, K., et al. (2008). Integrated genotype calling and association analysis of SNPs, common copy number polymorphisms and rare CNVs. Nat. Genet. 40, Landau, D.A., Carter, S.L., Stojanov, P., McKenna, A., Stevenson, K., Lawrence, M.S., Sougnez, C., Stewart, C., Sivachenko, A., Wang, L., et al. (2013). Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 152, Lee, I., Blom, U.M., Wang, P.I., Shim, J.E., and Marcotte, E.M. (2011). Prioritizing candidate disease genes by network-based boosting of genome-wide association data. Genome Res. 21, Lee, W.P., Stromberg, M.P., Ward, A., Stewart, C., Garrison, E.P., and Marth, G.T. (2014). MOSAIK: a hash-based algorithm for accurate next-generation sequencing short-read mapping. PLoS ONE 9, e Lex, A., Streit, M., Schulz, H.-J., Partl, C., Schmalstieg, D., Park, P.J., and Gehlenborg, N. (2012). StratomeX: Visual Analysis of Large-Scale Heterogeneous Genomics Data for Cancer Subtype Characterization. Comput. Graph. Forum 31, Li, B., and Dewey, C.N. (2011). RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323. Li, H., and Durbin, R. (2010). Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, Li, J., and Tibshirani, R. (2013). Finding consistent patterns: a nonparametric approach for identifying differential expression in RNA-Seq data. Stat. Methods Med. Res. 22, Liang, J., Shao, S.H., Xu, Z.X., Hennessy, B., Ding, Z., Larrea, M., Kondo, S., Dumont, D.J., Gutterman, J.U., Walker, C.L., et al. (2007). The energy sensing LKB1- AMPK pathway regulates p27(kip1) phosphorylation mediating the decision to enter autophagy or apoptosis. Nat. Cell Biol. 9, Liu, Y., Hayes, D.N., Nobel, A., and Marron, J.S. (2008). Statistical significance of clustering for high-dimension, low-smaple size data. J Am. Stat. Assn. 103, Lohr, J.G., Stojanov, P., Lawrence, M.S., Auclair, D., Chapuy, B., Sougnez, C., Cruz-Gordillo, P., Knoechel, B., Asmann, Y.W., Slager, S.L., et al. (2012). Discovery and prioritization of somatic mutations in diffuse large B-cell lymphoma (DLBCL) by whole-exome sequencing. Proc. Natl. Acad. Sci. USA 109, Luc, P.V., and Tempst, P. (2004). PINdb: a database of nuclear protein complexes from human and yeast. Bioinformatics 20, McCarroll, S.A., Kuruvilla, F.G., Korn, J.M., Cawley, S., Nemesh, J., Wysoker, A., Shapero, M.H., de Bakker, P.I., Maller, J.B., Kirby, A., et al. (2008). Integrated detection and population-genetic analysis of SNPs and copy number variation. Nat. Genet. 40, Mullokandov, G., Baccarini, A., Ruzo, A., Jayaprakash, A.D., Tung, N., Israelow, B., Evans, M.J., Sachidanandam, R., and Brown, B.D. (2012). High-throughput assessment of microrna activity and function using microrna sensor and decoy libraries. Nat. Methods 9, Neeley, E.S., Kornblau, S.M., Coombes, K.R., and Baggerly, K.A. (2009). Variable slope normalization of reverse phase protein arrays. Bioinformatics 25, Olshen, A.B., Venkatraman, E.S., Lucito, R., and Wigler, M. (2004). Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, R-Core-Team (2012). R: A language and environment for statistical computing (Vienna, Austria: R Foundation for Statistical Computing). Robinson, M.D., McCarthy, D.J., and Smyth, G.K. (2010). edger: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, Saldanha, A.J. (2004). Java Treeview extensible visualization of microarray data. Bioinformatics 20, Saunders, C.T., Wong, W.S., Swamy, S., Becq, J., Murray, L.J., and Cheetham, R.K. (2012). Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, Smigielski, E.M., Sirotkin, K., Ward, M., and Sherry, S.T. (2000). dbsnp: a database of single nucleotide polymorphisms. Nucleic Acids Res. 28, Stransky, N., Egloff, A.M., Tward, A.D., Kostic, A.D., Cibulskis, K., Sivachenko, A., Kryukov, G.V., Lawrence, M.S., Sougnez, C., McKenna, A., et al. (2011). The mutational landscape of head and neck squamous cell carcinoma. Science 333, Subramanian, A., Tamayo, P., Mootha, V.K., Mukherjee, S., Ebert, B.L., Gillette, M.A., Paulovich, A., Pomeroy, S.L., Golub, T.R., Lander, E.S., and Mesirov, J.P. (2005). Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, Tay, Y., Rinn, J., and Pandolfi, P.P. (2014). The multilayered complexity of cerna crosstalk and competition. Nature 505, Thorvaldsdóttir, H., Robinson, J.T., and Mesirov, J.P. (2013). Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, Tibes, R., Qiu, Y., Lu, Y., Hennessy, B., Andreeff, M., Mills, G.B., and Kornblau, S.M. (2006). Reverse phase protein array: validation of a novel proteomic technology and utility for analysis of primary leukemia specimens and hematopoietic stem cells. Mol. Cancer Ther. 5, Torres-García, W., Zheng, S., Sivachenko, A., Vegesna, R., Wang, Q., Yao, R., Berger, M.F., Weinstein, J.N., Getz, G., and Verhaak, R.G. (2014). PRADA: pipeline for RNA sequencing data analysis. Bioinformatics 30, Tusher, V.G., Tibshirani, R., and Chu, G. (2001). Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, Vandin, F., Clay, P., Upfal, E., and Raphael, B.J. (2012). Discovery of mutated subnetworks associated with clinical data in cancer. Pacific Symposium on Biocomputing Pacific Symposium on Biocomputing, Vaske, C.J., Benz, S.C., Sanborn, J.Z., Earl, D., Szeto, C., Zhu, J., Haussler, D., and Stuart, J.M. (2010). Inference of patient-specific pathway activities from multidimensional cancer genomics data using PARADIGM. Bioinformatics 26, i237 i245. Wang, K., Singh, D., Zeng, Z., Coleman, S.J., Huang, Y., Savich, G.L., He, X., Mieczkowski, P., Grimm, S.A., Perou, C.M., et al. (2010). MapSplice: accurate mapping of RNA-seq reads for splice junction discovery. Nucleic Acids Res. 38, e178. S2 Cell 159, , October 23, 2014 ª2014 The Authors
18 Wilkerson, M.D., and Hayes, D.N. (2010). ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26, Yang, L., Luquette, L.J., Gehlenborg, N., Xi, R., Haseley, P.S., Hsieh, C.H., Zhang, C., Ren, X., Protopopov, A., Chin, L., et al. (2013). Diverse mechanisms of somatic structural variations in human cancer genomes. Cell 153, Zack, T.I., Schumacher, S.E., Carter, S.L., Cherniack, A.D., Saksena, G., Tabak, B., Lawrence, M.S., Zhang, C.Z., Wala, J., Mermel, C.H., et al. (2013). Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, Cell 159, , October 23, 2014 ª2014 The Authors S3
19 (legend on next page) S4 Cell 159, , October 23, 2014 ª2014 The Authors
20 Figure S1. Exome DNA Sequencing and Mutation Density, Related to Figure 1. (A) Mutation density and effective coverage by exome DNA sequencing. Top: Mutation density in coding regions according to mutation type (nonsynonymous in blue, synonymous in red, and dbsnp build 134 in green). The 402 exomes samples are sorted from high to low mutation density, showing 11 PC tumors with a nonsynonymous mutation density exceeding 1/Mb and a steadily decreasing trend down to one sample with no coding mutations at all. The proportions of synonymous mutations and dbsnp mutations are consistent with 25% and 5% respectively over the spectrum. Bottom: Estimated number of covered exome bases, sorted in the same order. Covered bases are counted as those with at least 143 tumor read depth and 83 normal read depth. The median of 29.3 Mb covered bases corresponds to 89% of the total bases on the exome target region. The coverage spectrum has a standard deviation of 0.62 Mb, but there is no trend in coverage for high or low mutation density tumors. (B) Correlation of nonsynonymous mutation density and patient age (Pearson correlation = 0.442, p = ). The mean mutation density across the entire cohort was mutations/mb. The mutation density was least-squares fit to a linear model (density = a(years)+b), excluding the outlier tumor marked in red with mutation density > 2/Mb. The resulting fit (dashed gray line) had slope [ to ] mutations/(mb year) and intercept [ ] mutations/mb (95% confidence intervals). The residuals of this fit were used as the age corrected mutation density when quantifying how the mutation density is correlated to other parameters in this study. (C) Mutation density correlations to risk and MACIS for all histologic types. Upper left: Nonsynonymous mutation density versus MACIS for 359 tumors (with both exome data and a MACIS score) with super-imposed least-square fit showing a positive Pearson correlation of 0.40 with p = Upper right: Nonsynonymous Mutation density age fit residual versus MACIS for 359 tumors with least-square fit showing small correlation of 0.07 with p = Lower left: Nonsynonymous mutation density versus risk category boxplot for 368 tumors. There is a significant difference in the nonsynonymous mutation density across the three risk categories (Kruskal-Wallis test p = Lower right: Nonsynonymous mutation density age fit residual versus risk category boxplot for 368 tumors. The mean synonymous mutation density age fit residual increases with risk from for low risk tumors to for high risk tumors, a significant trend across the three risk categories (Kruskal-Wallis test p = ). (D) Mutation density correlations to risk and MACIS for classical type PTCs. Upper left: Nonsynonymous mutation density versus MACIS for 250 CT tumors (with both exome data and clinical information) with super-imposed least-square fit showing a positive Pearson correlation of 0.42 with p = Upper right: Nonsynonymous Mutation density age fit residual versus MACIS for 250 tumors with least-square fit showing small correlation of 0.06 with p = Lower left: Nonsynonymous mutation density versus risk category boxplot for 258 tumors. There is a significant difference in the nonsynonymous mutation density across the three risk categories (Kruskal-Wallis test p = Lower right: Nonsynonymous mutation density age fit residual versus risk category boxplot for 258 tumors. The mean nonsynonymous mutation density age fit residual increases with risk from for low risk tumors to 0.20 for high-risk tumors, a significant trend across the three risk categories (Kruskal-Wallis test p = ). (E) Mutation density according to BRAF, RAS and non-braf-ras driver mutation status. Several tumors, all BRAF V600E, displayed outlier mutation densities (top), but there was no significant difference in the nonsynonymous mutation density (p = Mann-Whitney rank-sum test) or the age-corrected mutation density (p = 0.380) between the BRAF- and RAS-mutated tumors (bottom). (F) Nonsynonymous mutation density by PTC histological type. The tall cell variant (TCV) cohort displayed a significantly higher mutation density (0.50 mut/mb) compared to CT (classical) (0.40 mut/mb) and FV (follicular) (0.40 mut/mb) (Kruskal-Wallis test p = 0.058), although when the mutation density is corrected for age the differences become insignificant (Kruskal-Wallis test p = 0.64). (G) Histopathology of five tumors that have outlier mutation densities and TC morphology and/or complex papillary growth patterns. H&E. Cell 159, , October 23, 2014 ª2014 The Authors S5
21 Figure S2. Amino Acid Positions of Mutations in Several Top Significantly Mutated Genes, Related To Figure 1 (A) BRAF. (B) KRAS. (C) EIF1AX mutations in the THCA cohort (top) as well as the in the COSMIC database (middle) and in uveal melanoma (bottom) according to Martin et al. (Martin et al., 2013). Cancer cell fraction (CCF) and mutation type are shown as pie charts where red represents the CCF. All of the EIF1AX mutations have cancer cell fractions (CCFs) > 0.75 (see Mutation Clonality Analysis section in Extended Experimental Procedures). (D) PPM1D. (E) CHEK2. S6 Cell 159, , October 23, 2014 ª2014 The Authors
22 (legend on next page) Cell 159, , October 23, 2014 ª2014 The Authors S7
23 Figure S3. Somatic Mutations in Relevant Cancer Genes and Gene Fusions, Related to Figures 1 and 3 (A) Somatic mutations within DNA repair-related genes. Twenty-six tumors contained 26 mutually exclusive mutations across 8 genes (MEM0 corrected p = 0.24), as shown. Purple, SSNV; orange, gene fusion. BRS: red, RAS-like; blue, BRAF V600E -like. (B) Somatic mutations of WNT pathway genes. Six tumors contained mutually exclusive mutations involving 5 wnt-related genes. Blue, SSNV; orange, fusion. (C) Somatic mutations of known tumor-suppressor genes. Fifteen tumors contained 15 mutually exclusive mutations of 6 well known tumor-suppressor genes. Blue, SSNV; orange, fusion; red, deletion; green, deletion+ssnv. (D) Somatic mutations within chromatin remodeling genes. Mutations in genes involved in epigenetic regulation of gene expression, i.e., chromatin remodeling genes, were frequent, although not frequent enough to be significant on their own. A total of 80 tumors contained 93 mutations across 59 epigenetic regulatory genes, as shown. Mutations in a subset of 49 chromatin modifier genes were significantly mutually exclusive (MEMo corrected p < 0.02, Table S4A). Seventy-one tumors had a single mutation, 7 tumors had 2 mutations, 1 tumor had 3 mutations, and 1 tumor had 5 mutations. As a group, tumors with mutations in chromatin remodeling genes had a significantly higher median age-corrected mutation density (Table S5A, Wilcoxon rank-sum p = ). These tumors did not display a distinctive global methylation pattern, as they were represented in roughly equal percentages within DNA methylation subtypes (see Molecular Classification). Blue, SSNV; orange, gene fusion. (E) Somatic mutations of common PI3K pathways genes and PPARg. HRAS, NRAS and KRAS SSNVs, RET fusions, PAX8/PPARG fusions, PTEN deletions and are close to mutually exclusive with each other and with BRAF alterations. Purple, SSNV; orange, fusion; red, deletion; green, deletion+ssnv. Histology: light blue, CT; red, FV, dark blue, TCV. BRS: red, RAS-like; blue, BRAF V600E -like. (F) Gene fusions found in the PTC cohort (n = 484) grouped by fusion partner (RET, PPARG, BRAF, NTRK1, NTRK3, ALK, THADA, FGFR2, MET and LTK). Risk of recurrence, histology (CT, blue; FV, red; TC dark blue; gray, na) and BRS (RAS-like, red; BRAF V600E -like, dark blue; gray, BRS not available) are shown. S8 Cell 159, , October 23, 2014 ª2014 The Authors
24 Figure S4. Somatic Copy-Number Alterations, Related to Figure 1 (A) Comparison of frequency of arm-level alterations in different histological types. FV had many statistically significant gains and losses compared to CT and TCV. (legend continued on next page) Cell 159, , October 23, 2014 ª2014 The Authors S9
25 (B) Clustering of SCNA data defined 4 groups of PTC, designated Quiet, 22q, Many SCNAs, and Some SCNAs. Histology, risk and BRAF/RAS genotype are shown. Specific associations between SCNA features and selected clinicopathologic features are shown in Figures S4C S4F. (C) Associations between histology, BRAF/RAS genotype and 22q deletions and 1q amplifications. Tumors with 22q Del had a higher percentage of FV and no TCV (Fisher exact test p < 0.05), a higher percentage of BRAF-wild-type tumors (p < 0.01), and a higher percentage of RAS mutations (p < ). Tumors with 1q amplification had a higher percentage of TCV and CT without FV (p < ), a higher percentage of BRAF mutations (p < 0.05), and no statistically significant differences between tumors with and without RAS mutations. (D) Associations between 22q deletion and 1q amplification and MACIS score. There were no statistically significant differences in MACIS score between tumors with and without 22q deletion. However, differences were observed when looking only at CT (p < ). MACIS scores were significantly higher in tumors with 1q amplification, both when considering all PTC (p < ) and only BRAF mutants (p < ). (E) Associations between risk and 1q amplification and 22q deletion. The risk profile of tumors was significantly higher in tumors with 1q amplifications (p < 0.01). There were no significant differences in risk between tumors with and without 22q deletions. (F) Associations between tumor stage and 1q amplification and 22q deletion. Stage was significantly higher in tumors with 1q amplifications (p < ). There were no significant differences in tumor stage between tumors with and without 22q deletion. (G) Correlation between PTEN copy-number alterations and expression. One tumor, TCGA-ET-A4KN, contained a PTEN intron deletion and low levels of PTEN expression. S10 Cell 159, , October 23, 2014 ª2014 The Authors
26 Figure S5. HotNet2 Analysis (A) Seven of the subnetworks (MAPK, ECM-receptor, complement and coagulation cascade, FANCA-associated, VEGF signaling pathway, SNF2h/cohesion, and NOD-like receptor signaling) identified by HotNet. See Table S5C. (B) Mutations in the MAPK subnetwork identified by HotNet (p = 0.03). Inactivating mutations are defined as frame shift indels; and nonstop, nonsense, and splice site mutations. The number of mutated samples for each gene is reported. Sample types are defined according to the BRAF V600E -RAS score described in the Signaling and Thyroid Differentiation section of Extended Experimental Procedures. (C) Mutations in the ECM-receptor subnetwork identified by HotNet (p = ). For details see legend of (B). (D) Mutations in the complement and coagulase cascade subnetwork identified by HotNet (p = ). For details see legend of (B). (E) Mutations in the FANCA-associated subnetwork identified by HotNet (p = ). For details see legend of (B). Cell 159, , October 23, 2014 ª2014 The Authors S11
27 Figure S6. Network Based Stratification, Related to Figure 4 (A) Intrinsic validation measures of subtype quality. (a) Cophenetic correlation, (b) Mean Silhouette, (c) Weighted graph modularity score. (d) Visualization of the coclustering matrices for different values of K. (B) Phenotype-NBS subtype associations. Association with (a) extrathyroidal extension, (b) histological type, (c) pathological T stage, (d) lymph node status (pathological N stage), and (e) risk of recurrence category. The numbers in each cell in a table correspond to the number of patients in that subtype-phenotype pair, and color corresponds to a Pearson residual which quantifies the deviation from expected count for that cell in the table. (C) Associations between NBS cluster and (a) BRS, (b) ERK score and (c) TDS. Boxplot figure, red line corresponds to medians, and box limits to upper and lower quartiles in the data. (D) Characteristic subnetwork of NBS subtypes. Each subnetwork is a module detected by using HotNet on patient profiles from a single NBS subtype (a subtype NBS1, b subtype NBS2, and c subtype NBS3). Bold outlines indicate known cancer genes from the COSMIC gene census. Blue edges indicate the interaction is included in the HumanNet gene network, gray edges are inferred influence edges. Color of a node corresponds to the number of individuals with a mutation in the gene within that subtype. S12 Cell 159, , October 23, 2014 ª2014 The Authors
28 (legend on next page) Cell 159, , October 23, 2014 ª2014 The Authors S13
29 Figure S7. The BRAF V600E -RAS Score and Thyroid Differentiation Score, Related to Figures 4, 5, and 7 (A) mrna expression of the 71 genes used to derive the BRAF V600E -RAS score (BRS) across 391 PTC samples. Samples are ranked based on their BRS value ranging from 1 (dark blue) indicating the most BRAF V600E -like cases to 1 (red) indicating the most RAS-like cases. (B) Principal component analysis of 391 PTCs using the 71 genes used to derive the BRAF V600E -RAS score (BRS). (C) Correlation of BRAF V600E -RAS score (BRS) and Thyroid Differentiation Score (TDS). Left: Correlation across all tumors (n = 391). Tumors are annotated by genotype. Right: Correlation across the BRAF V600E cohort of tumors (n = 228). The TDS was derived using mrna expression levels of the 16 genes in Table S5F. (D) Pairwise correlations (red) and anticorrelations (blue) across the full cohort (n = 390), measured by Spearman correlation coefficients among the TDS, BRS, mrna expression levels of 16 thyroid genes, and expression levels of mrna and mirna genes highlighted in Figure 5. (E) Pairwise correlations (red) and anticorrelations (blue) across the BRAF V600E cohort (n = 228), measured by Spearman correlation coefficients among the TDS, BRS, mrna expression levels of 16 thyroid genes, and expression levels of mrna and mirna genes highlighted in Figure 5. (F) mrnas (top) and mirnas (bottom) correlated to the TDS across the full cohort. The 16 thyroid genes of the TDS are highlighted by red circles, while selected mrna and mirna genes in Figure 5 are highlighted by blue circles. In both (top) and (bottom), the x axis represents Spearman correlation coefficients and y axis represents Log10 (False Discovery Rate) (FDR). (G) mrnas (top) and mirnas (bottom) correlated to the TDS across the BRAF-V600E cohort. For details see legend to (D). (H) mrna genes correlated to the TDS in (top) 51 PTC of all genotypes and (bottom) 26 BRAF V600E PTC samples using previously published DNA microarray data (Giordano et al., 2005). (I) Histologic grading of BRAF V600E tumors. Representative tumors in each grade are shown. H&E. (J) TDS correlations within the BRAF V600E cohort. Top left: Correlation between TDS and architectural grade. Tumors were architecturally graded using a 3-grade system as described in Extended Experimental Procedures. Top right: Correlation between TDS and risk of recurrence. Bottom left: Correlation between TDS and MACIS score. Bottom right: Correlation between TDS and tumor purity. S14 Cell 159, , October 23, 2014 ª2014 The Authors
30 (legend on next page) Cell 159, , October 23, 2014 ª2014 The Authors S15
31 Figure S8. The ERK Score and Signaling Consequences, Related to Figure 6 (A) Heatmap of the 52 genes comprising the ERK-activity signature in Pratilas et al. (Pratilas et al., 2009), ordered from left to right according to BRS values (from 1 to 1). The vertical bar on the left shows the direction of regulation: red lines show genes that are downregulated and blue lines show genes that are upregulated upon MEK inhibition. (B) Expression levels of ps338-craf and ps299-araf from RPPA data, according to mutation group. (C) TieDIE networks connecting BRAF/RAS mutated genes to transcription factors and signaling proteins with altered activity in tumor samples. Left: The core network of genes connecting mutant genes NRAS and BRAF to inputs generated with RNA-Seq (ERK1-2-active complex) with paths of length up to 3. Nodes correspond to proteins or complexes in the TieDIE solution; the size of the label text indicates the influence attributed to each node, in relation to the three input data sets. Solid lines indicate transcriptional regulation and dashed lines indicate protein regulation, and dotted lines indicate HPRD-PPI interactions or component associations. Each tick-mark in each circle represents a single patient sample. Circles represent genomic perturbations, PARADIGM inferred activities, RPPA and expression for proteins and complexes. Outer ring represents mutation status for BRAF and NRAS genes; for all other genes it represents the PARADIGM inferred activity levels. Second most outer ring represents RPPA activity level, and the inner ring represents the gene expression. All rings are sorted in the same order and according to the presence of a mutation in BRAF or RAS-related genes (NRAS/KRAS/HRAS/EIF1AX), and secondarily according to the activity of BRAF within mutant samples, as measured by RPPA. Right: The larger TieDIE network generated by allowing all paths between BRAF/NRAS to RNA- Seq generated inputs with paths of length up to 4. S16 Cell 159, , October 23, 2014 ª2014 The Authors
32 (legend on next page) Cell 159, , October 23, 2014 ª2014 The Authors S17
33 Figure S9. Molecular Classification of PTC, Related to Figures 4, 5, and 7 (A) Gene expression heatmap with subtype and mutation annotations. Unsupervised consensus clustering (Wilkerson) of RNaseq data from 482 tumor samples detected five RNA subtypes. The heatmap shows the expression levels of 755 classifier genes identified by ClaNC (Dabney, 2006) whose expression patterns characterize the RNA subtypes. Genes appear in rows and are clustered hierarchically. Samples appear in columns and are organized according to RNA subtype, mirna subtype, methylation subtype, copy-number subtype, RAS mutation status (HRAS, NRAS, KRAS), and BRAF mutation or fusion status. (B) Unsupervised consensus clustering of mirna-seq 5p and 3p mature strand data for 495 tumor samples. (a) A schematic of the 5p and 3p abundance data used (indicated by the gray triangle). Pri = primary transcript. Pre = pre-mirna. (b) Profiles of cophenetic correlation coefficient and consensus average silhouette width from the rank survey, and the red/blue consensus membership heatmap for a six-cluster solution. (c) Normalized abundance heatmap for the six-group solution, with histology and driver mutation/fusion covariates (as in Figure 7). The heatmap shows discriminatory mirs with the largest 6% of metagene matrix scores (Gaujoux and Seoighe, 2010), and mir-204-5p, 221-3p and 222-3p, which were included because they were highlighted from correlations to scores (see Figure S10D). The scalebar shows log2 normalized (reads-per-million, RPM), median-centered mir abundance. mir names in red are discussed in the main text. Gray vertical lines in the histology covariate tracks mark samples with no clinical data, and in the driver mutation/fusion covariate tracks mark samples with no whole exome sequence data. d) Distributions of purity from ABSOLUTE (Carter et al., 2012). e) Distribution of RNaseq-based EMT scores (Byers et al., 2013). E = epithelial, M = mesenchymal. Median scores for all mir-based clusters were weakly negative, i.e., epithelial. f) Distributions of RPM abundance for selected mirs. (C) Unsupervised clusters of DNA methylation data. (a) The heat map shows beta values of 496 PTC samples in columns that are ordered by four clusters and top 1% of the most variable CpG loci in rows. The 56 matched normal samples are shown on the right. (b) Association of DNA methylation clusters with driver mutations (shown in black) and fusions (shown in green), as well as with histology. p values on the right are derived from Fisher s exact test. Mutation and fusion data are available for 393 samples. (c) Association with groupings derived on the basis of mrna, mirna, and protein expression data. p values on the right are derived from Fisher s exact test. (d) Tumor purity, percent of lymphocyte infiltration and stromal cells are depicted in boxplots, colors correspond to the top bar of the heat map on panel a. (D) Heat map of unsupervised hierarchical clustering of 368 tumor samples (columns) and 181 antibodies (rows) for RPPA data. The annotation bars (shown on top) were not used for clustering. The legend for the annotation bars is shown on the left. The leftmost (red) cluster has predominantly follicular samples, and is depleted in BRAF mutations. The other three clusters are enriched in BRAF mutations, but depleted in NRAS, HRAS and EIF1AX mutations. They are predominantly CT samples by histology. Various markers for the clusters are highlighted by arrows (in the map, blue color means low expression, white means medium, and red means high expression). (E) Overall superclusters derived from clusters on individual data types. The columns contain samples. The rows contain sample cluster memberships from multiple data types. Different colors represent different clusters within a row. The annotation bars on top of the heat map show clinical associations of the super clusters (not used to derive the clusters). The figure shows two super clusters, a RAS-like cluster showing predominantly RAS mutants (left) and a BRAF V600E -like cluster showing predominantly BRAF V600E mutants (right). The figure shows very good agreement between the various data types with respect to RAS/BRAF status. S18 Cell 159, , October 23, 2014 ª2014 The Authors
34 (legend on next page) Cell 159, , October 23, 2014 ª2014 The Authors S19
Dr Catherine Woolnough, Hospital Scientist, Chemical Pathology, Royal Prince Alfred Hospital. NSW Health Pathology University of Sydney
 Dr Catherine Woolnough, Hospital Scientist, Chemical Pathology, Royal Prince Alfred Hospital NSW Health Pathology University of Sydney Thyroid Cancer TC incidence rates in NSW Several subtypes - Papillary
Dr Catherine Woolnough, Hospital Scientist, Chemical Pathology, Royal Prince Alfred Hospital NSW Health Pathology University of Sydney Thyroid Cancer TC incidence rates in NSW Several subtypes - Papillary
Genomic landsccape of poorly differentiated and anaplastic thyroid carcinomas: Clues for better classification, risk stratification and therapy
 ACCME/Disclosures The USCAP requires that anyone in a position to influence or control the content of CME disclose any relevant financial relationship WITH COMMERCIAL INTERESTS which they or their spouse/partner
ACCME/Disclosures The USCAP requires that anyone in a position to influence or control the content of CME disclose any relevant financial relationship WITH COMMERCIAL INTERESTS which they or their spouse/partner
Molecular Diagnostics in Thyroid Tumors
 USCAP 2011 Endocrine Pathology Society Companion Meeting Molecular Diagnostics in Thyroid Tumors Yuri E. Nikiforov, M.D., Ph.D. Department of Pathology University of Pittsburgh Medical Center Outline Overview
USCAP 2011 Endocrine Pathology Society Companion Meeting Molecular Diagnostics in Thyroid Tumors Yuri E. Nikiforov, M.D., Ph.D. Department of Pathology University of Pittsburgh Medical Center Outline Overview
Clinical and Molecular Approach to Using Thyroid Needle Biopsy for Nodular Disease
 Clinical and Molecular Approach to Using Thyroid Needle Biopsy for Nodular Disease Robert L. Ferris, MD, PhD Department of Otolaryngology/Head and Neck Surgery and Yuri E. Nikiforov, MD, PhD Division of
Clinical and Molecular Approach to Using Thyroid Needle Biopsy for Nodular Disease Robert L. Ferris, MD, PhD Department of Otolaryngology/Head and Neck Surgery and Yuri E. Nikiforov, MD, PhD Division of
Dilemmas in Cytopathology and Histopathology
 Dilemmas in Cytopathology and Histopathology Yuri E. Nikiforov, MD, PhD Division of Molecular & Genomic Pathology University of Pittsburgh Medical Center, USA Objectives Discuss new WHO classification
Dilemmas in Cytopathology and Histopathology Yuri E. Nikiforov, MD, PhD Division of Molecular & Genomic Pathology University of Pittsburgh Medical Center, USA Objectives Discuss new WHO classification
Section 2 Original Policy Date 2013 Last Review Status/Date September 1, 2014
 Policy Number 2.04.82 Molecular Markers in Fine Needle Aspirates of the Thyroid Medical Policy Section 2 Original Policy Date 2013 Last Review Status/Date September 1, 2014 Disclaimer Our medical policies
Policy Number 2.04.82 Molecular Markers in Fine Needle Aspirates of the Thyroid Medical Policy Section 2 Original Policy Date 2013 Last Review Status/Date September 1, 2014 Disclaimer Our medical policies
Biomarker development in the era of precision medicine. Bei Li, Interdisciplinary Technical Journal Club
 Biomarker development in the era of precision medicine Bei Li, 23.08.2016 Interdisciplinary Technical Journal Club The top ten highest-grossing drugs in the United States help between 1 in 25 and 1 in
Biomarker development in the era of precision medicine Bei Li, 23.08.2016 Interdisciplinary Technical Journal Club The top ten highest-grossing drugs in the United States help between 1 in 25 and 1 in
Frequency(%) KRAS G12 KRAS G13 KRAS A146 KRAS Q61 KRAS K117N PIK3CA H1047 PIK3CA E545 PIK3CA E542K PIK3CA Q546. EGFR exon19 NFS-indel EGFR L858R
 Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies
 Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Markers in Thyroid Nodule Evaluation. Yuri E. Nikiforov, MD, PhD Division of Molecular & Genomic Pathology University of Pittsburgh Medical Center
 Markers in Thyroid Nodule Evaluation Yuri E. Nikiforov, MD, PhD Division of Molecular & Genomic Pathology University of Pittsburgh Medical Center Disclosures Quest Diagnostics (consultant) UPMC/CBLPath
Markers in Thyroid Nodule Evaluation Yuri E. Nikiforov, MD, PhD Division of Molecular & Genomic Pathology University of Pittsburgh Medical Center Disclosures Quest Diagnostics (consultant) UPMC/CBLPath
AD (Leave blank) TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients
 AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
Insights from Sequencing the Melanoma Exome
 Insights from Sequencing the Melanoma Exome Michael Krauthammer, MD PhD, December 2 2015 Yale University School Yof Medicine 1 2012 Exome Screens and Results Exome Sequencing of 108 sun-exposed melanomas
Insights from Sequencing the Melanoma Exome Michael Krauthammer, MD PhD, December 2 2015 Yale University School Yof Medicine 1 2012 Exome Screens and Results Exome Sequencing of 108 sun-exposed melanomas
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Osamu Tetsu, MD, PhD Associate Professor Department of Otolaryngology-Head and Neck Surgery School of Medicine, University of California, San
 Osamu Tetsu, MD, PhD Associate Professor Department of Otolaryngology-Head and Neck Surgery School of Medicine, University of California, San Francisco Lung Cancer Classification Pathological Classification
Osamu Tetsu, MD, PhD Associate Professor Department of Otolaryngology-Head and Neck Surgery School of Medicine, University of California, San Francisco Lung Cancer Classification Pathological Classification
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
Case year old female presented with asymmetric enlargement of the left lobe of the thyroid
 Case 4 22 year old female presented with asymmetric enlargement of the left lobe of the thyroid gland. No information available relative to a prior fine needle aspiration biopsy. A left lobectomy was performed.
Case 4 22 year old female presented with asymmetric enlargement of the left lobe of the thyroid gland. No information available relative to a prior fine needle aspiration biopsy. A left lobectomy was performed.
Case 4 Diagnosis 2/21/2011 TGB
 Case 4 22 year old female presented with asymmetric enlargement of the left lobe of the thyroid gland. No information available relative to a prior fine needle aspiration biopsy. A left lobectomy was performed.
Case 4 22 year old female presented with asymmetric enlargement of the left lobe of the thyroid gland. No information available relative to a prior fine needle aspiration biopsy. A left lobectomy was performed.
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
The Bethesda Indeterminate Categories: An Update to Diagnosis and Molecular Testing
 William C. Faquin, MD, PhD Professor of Pathology Harvard Medical School Director, Head and Neck Pathology Massachusetts Eye and Ear Massachusetts General Hospital The Bethesda Indeterminate Categories:
William C. Faquin, MD, PhD Professor of Pathology Harvard Medical School Director, Head and Neck Pathology Massachusetts Eye and Ear Massachusetts General Hospital The Bethesda Indeterminate Categories:
Supplementary Figure 1. Copy Number Alterations TP53 Mutation Type. C-class TP53 WT. TP53 mut. Nature Genetics: doi: /ng.
 Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Genomic Medicine: What every pathologist needs to know
 Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
ASC Companion Meeting at the 2017 USCAP: Ancillary Molecular Testing in "Indeterminate. Thyroid Nodules: How Far Have We Come?
 ASC Companion Meeting at the 2017 USCAP: Ancillary Molecular Testing in "Indeterminate Thyroid Nodules: How Far Have We Come? William C. Faquin, MD, PhD, Massachusetts General Hospital, Boston, MA The
ASC Companion Meeting at the 2017 USCAP: Ancillary Molecular Testing in "Indeterminate Thyroid Nodules: How Far Have We Come? William C. Faquin, MD, PhD, Massachusetts General Hospital, Boston, MA The
EXAMPLE. - Potentially responsive to PI3K/mTOR and MEK combination therapy or mtor/mek and PKC combination therapy. ratio (%)
 Dr Kate Goodhealth Goodhealth Medical Clinic 123 Address Road SUBURBTOWN NSW 2000 Melanie Citizen Referring Doctor Your ref Address Dr John Medico 123 Main Street, SUBURBTOWN NSW 2000 Phone 02 9999 9999
Dr Kate Goodhealth Goodhealth Medical Clinic 123 Address Road SUBURBTOWN NSW 2000 Melanie Citizen Referring Doctor Your ref Address Dr John Medico 123 Main Street, SUBURBTOWN NSW 2000 Phone 02 9999 9999
Computer Science, Biology, and Biomedical Informatics (CoSBBI) Outline. Molecular Biology of Cancer AND. Goals/Expectations. David Boone 7/1/2015
 Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Research Strategy: 1. Background and Significance
 Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Molecular Markers in Fine Needle Aspirates of the Thyroid
 Molecular Markers in Fine Needle Aspirates of the Thyroid Policy Number: 2.04.78 Last Review: 3/2014 Origination: 3/2013 Next Review: 3/2015 Policy Blue Cross and Blue Shield of Kansas City (Blue KC) will
Molecular Markers in Fine Needle Aspirates of the Thyroid Policy Number: 2.04.78 Last Review: 3/2014 Origination: 3/2013 Next Review: 3/2015 Policy Blue Cross and Blue Shield of Kansas City (Blue KC) will
Corporate Medical Policy
 Corporate Medical Policy Molecular Analysis for Targeted Therapy for Non-Small Cell Lung File Name: Origination: Last CAP Review: Next CAP Review: Last Review: molecular_analysis_for_targeted_therapy_for_non_small_cell_lung_cancer
Corporate Medical Policy Molecular Analysis for Targeted Therapy for Non-Small Cell Lung File Name: Origination: Last CAP Review: Next CAP Review: Last Review: molecular_analysis_for_targeted_therapy_for_non_small_cell_lung_cancer
04/09/2018. Follicular Thyroid Tumors Updates in Classification & Practical Tips. Dissecting Indeterminants. In pursuit of the low grade malignancy
 Follicular Thyroid Tumors Updates in Classification & Practical Tips Jennifer L. Hunt, MD, MEd Aubrey J. Hough Jr, MD, Endowed Professor of Pathology Chair of Pathology and Laboratory Medicine University
Follicular Thyroid Tumors Updates in Classification & Practical Tips Jennifer L. Hunt, MD, MEd Aubrey J. Hough Jr, MD, Endowed Professor of Pathology Chair of Pathology and Laboratory Medicine University
The mutations that drive cancer. Paul Edwards. Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge
 The mutations that drive cancer Paul Edwards Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge Previously on Cancer... hereditary predisposition Normal Cell Slightly
The mutations that drive cancer Paul Edwards Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge Previously on Cancer... hereditary predisposition Normal Cell Slightly
3/27/2017. Disclosure of Relevant Financial Relationships. Each year over 550,000 thyroid FNAs are performed in the U.S.!!! THYROID FNA: THE GOOD NEWS
 Disclosure of Relevant Financial Relationships William C. Faquin, MD, PhD Director, Head and Neck Pathology Massachusetts Eye and Ear Massachusetts General Hospital Professor of Pathology Harvard Medical
Disclosure of Relevant Financial Relationships William C. Faquin, MD, PhD Director, Head and Neck Pathology Massachusetts Eye and Ear Massachusetts General Hospital Professor of Pathology Harvard Medical
NGS in tissue and liquid biopsy
 NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
Expanded View Figures
 Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
Characterisation of structural variation in breast. cancer genomes using paired-end sequencing on. the Illumina Genome Analyser
 Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Reviewers' comments: Reviewer #1 (Remarks to the Author):
 Reviewers' comments: Reviewer #1 (Remarks to the Author): In this study the authors analysed 18 deep penetrating nevi for oncogenic genomic changes (single nucleotide variations, insertions/deletions,
Reviewers' comments: Reviewer #1 (Remarks to the Author): In this study the authors analysed 18 deep penetrating nevi for oncogenic genomic changes (single nucleotide variations, insertions/deletions,
Let s Make Sense of Present & Predict Future. In Light of Past 1/12/2016
 The New Diagnostic Paradigms in Thyroid Surgical Pathology and Affects on Reporting of Thyroid Fine Needle Aspiration Specimens Deliberations, Criticisms & Discussions Zubair W. Baloch, MD, PhD. Professor
The New Diagnostic Paradigms in Thyroid Surgical Pathology and Affects on Reporting of Thyroid Fine Needle Aspiration Specimens Deliberations, Criticisms & Discussions Zubair W. Baloch, MD, PhD. Professor
Looking Beyond the Standard-of- Care : The Clinical Trial Option
 1 Looking Beyond the Standard-of- Care : The Clinical Trial Option Terry Mamounas, M.D., M.P.H., F.A.C.S. Medical Director, Comprehensive Breast Program UF Health Cancer Center at Orlando Health Professor
1 Looking Beyond the Standard-of- Care : The Clinical Trial Option Terry Mamounas, M.D., M.P.H., F.A.C.S. Medical Director, Comprehensive Breast Program UF Health Cancer Center at Orlando Health Professor
Results and Discussion of Receptor Tyrosine Kinase. Activation
 Results and Discussion of Receptor Tyrosine Kinase Activation To demonstrate the contribution which RCytoscape s molecular maps can make to biological understanding via exploratory data analysis, we here
Results and Discussion of Receptor Tyrosine Kinase Activation To demonstrate the contribution which RCytoscape s molecular maps can make to biological understanding via exploratory data analysis, we here
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs.
 Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK
 CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
ACCME/Disclosures. Questions to Myself? 4/11/2016
 The New Diagnostic Paradigms in Thyroid Surgical Pathology and Affects on Reporting of Thyroid Fine-Needle Aspiration Specimens Deliberations, Criticisms & Discussions Zubair W. Baloch, MD, PhD. Professor
The New Diagnostic Paradigms in Thyroid Surgical Pathology and Affects on Reporting of Thyroid Fine-Needle Aspiration Specimens Deliberations, Criticisms & Discussions Zubair W. Baloch, MD, PhD. Professor
Clinical Grade Genomic Profiling: The Time Has Come
 Clinical Grade Genomic Profiling: The Time Has Come Gary Palmer, MD, JD, MBA, MPH Senior Vice President, Medical Affairs Foundation Medicine, Inc. Oct. 22, 2013 1 Why We Are Here A Shared Vision At Foundation
Clinical Grade Genomic Profiling: The Time Has Come Gary Palmer, MD, JD, MBA, MPH Senior Vice President, Medical Affairs Foundation Medicine, Inc. Oct. 22, 2013 1 Why We Are Here A Shared Vision At Foundation
MicroRNA expression profiling and functional analysis in prostate cancer. Marco Folini s.c. Ricerca Traslazionale DOSL
 MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium
 Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium Ulrike Peters Fred Hutchinson Cancer Research Center University of Washington U01-CA137088-05, PI: Peters
Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium Ulrike Peters Fred Hutchinson Cancer Research Center University of Washington U01-CA137088-05, PI: Peters
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Genomic tests to personalize therapy of metastatic breast cancers. Fabrice ANDRE Gustave Roussy Villejuif, France
 Genomic tests to personalize therapy of metastatic breast cancers Fabrice ANDRE Gustave Roussy Villejuif, France Future application of genomics: Understand the biology at the individual scale Patients
Genomic tests to personalize therapy of metastatic breast cancers Fabrice ANDRE Gustave Roussy Villejuif, France Future application of genomics: Understand the biology at the individual scale Patients
THYROID TUMOR DIAGNOSIS: MARKER OF THE MONTH CLUB
 THYROID TUMOR DIAGNOSIS: MARKER OF THE MONTH CLUB CHARACTERISTIC OF THE IDEAL TUMOR MARKER Specific Sensitive Easy to perform Easy to interpret Adaptable to FNA Reasonable cost (CHEAP) THYROID TUMOR MARKERS
THYROID TUMOR DIAGNOSIS: MARKER OF THE MONTH CLUB CHARACTERISTIC OF THE IDEAL TUMOR MARKER Specific Sensitive Easy to perform Easy to interpret Adaptable to FNA Reasonable cost (CHEAP) THYROID TUMOR MARKERS
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
microrna Presented for: Presented by: Date:
 microrna Presented for: Presented by: Date: 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions
microrna Presented for: Presented by: Date: 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions
Tumor suppressor genes D R. S H O S S E I N I - A S L
 Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
Comprehensive Genomic Profiling, in record time. Accurate. Clinically Proven. Fast.
 Comprehensive Genomic Profiling, in record time Accurate. ly Proven. Fast. PCDx advantages Comprehensive genomic profiling, in record time PCDx Comprehensive Genomic Profiling (CGP) provides precise information
Comprehensive Genomic Profiling, in record time Accurate. ly Proven. Fast. PCDx advantages Comprehensive genomic profiling, in record time PCDx Comprehensive Genomic Profiling (CGP) provides precise information
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making
 Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making November 20, 2014 Capturing Value in Next Generation Sequencing Symposium Douglas Johnson MD, MSCI Vanderbilt-Ingram
Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making November 20, 2014 Capturing Value in Next Generation Sequencing Symposium Douglas Johnson MD, MSCI Vanderbilt-Ingram
Disclosures Genomic testing in lung cancer
 Disclosures Genomic testing in lung cancer No disclosures Objectives Understand how FISH and NGS provide complementary data for the evaluation of lung cancer Recognize the challenges of performing testing
Disclosures Genomic testing in lung cancer No disclosures Objectives Understand how FISH and NGS provide complementary data for the evaluation of lung cancer Recognize the challenges of performing testing
SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer.
 Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
2015 American Thyroid Association Thyroid Nodule and Cancer Guidelines
 2015 American Thyroid Association Thyroid Nodule and Cancer Guidelines Angela M. Leung, MD, MSc, ECNU November 5, 2016 Outline Workup of nontoxic thyroid nodule(s) Ultrasound FNAB Management of FNAB results
2015 American Thyroid Association Thyroid Nodule and Cancer Guidelines Angela M. Leung, MD, MSc, ECNU November 5, 2016 Outline Workup of nontoxic thyroid nodule(s) Ultrasound FNAB Management of FNAB results
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Activation of cellular proto-oncogenes to oncogenes. How was active Ras identified?
 Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
METASTATIC COLORECTAL CANCER: TUMOR MUTATIONAL ANALYSIS AND ITS IMPACT ON CHEMOTHERAPY SUMA SATTI, MD
 METASTATIC COLORECTAL CANCER: TUMOR MUTATIONAL ANALYSIS AND ITS IMPACT ON CHEMOTHERAPY SUMA SATTI, MD INTRODUCTION Second leading cause of cancer related death in the United States. 136,830 cases in 2014
METASTATIC COLORECTAL CANCER: TUMOR MUTATIONAL ANALYSIS AND ITS IMPACT ON CHEMOTHERAPY SUMA SATTI, MD INTRODUCTION Second leading cause of cancer related death in the United States. 136,830 cases in 2014
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
POORLY DIFFERENTIATED, HIGH GRADE AND ANAPLASTIC CARCINOMAS: WHAT IS EVERYONE TALKING ABOUT?
 POORLY DIFFERENTIATED, HIGH GRADE AND ANAPLASTIC CARCINOMAS: WHAT IS EVERYONE TALKING ABOUT? AGGRESSIVE THYROID CANCERS PAPILLARY CARCINOMA CERTAIN SUBTYPES POORLY DIFFERENTIATED CARCINOMA HIGH GRADE DIFFERENTIATED
POORLY DIFFERENTIATED, HIGH GRADE AND ANAPLASTIC CARCINOMAS: WHAT IS EVERYONE TALKING ABOUT? AGGRESSIVE THYROID CANCERS PAPILLARY CARCINOMA CERTAIN SUBTYPES POORLY DIFFERENTIATED CARCINOMA HIGH GRADE DIFFERENTIATED
Next generation histopathological diagnosis for precision medicine in solid cancers
 Next generation histopathological diagnosis for precision medicine in solid cancers from genomics to clinical application Aldo Scarpa ARC-NET Applied Research on Cancer Department of Pathology and Diagnostics
Next generation histopathological diagnosis for precision medicine in solid cancers from genomics to clinical application Aldo Scarpa ARC-NET Applied Research on Cancer Department of Pathology and Diagnostics
The Pathology of Neoplasia Part II
 The Pathology of Neoplasia Part II February 2018 PAUL BOGNER, MD A S S O C I A T E P R O F E S S O R O F O N C O L O G Y P A T H O L O G Y A N D D E R M A T O L O G Y Clinical goals of cancer pathology
The Pathology of Neoplasia Part II February 2018 PAUL BOGNER, MD A S S O C I A T E P R O F E S S O R O F O N C O L O G Y P A T H O L O G Y A N D D E R M A T O L O G Y Clinical goals of cancer pathology
Evolution of Pathology
 1 Traditional pathology Molecular pathology 2 Evolution of Pathology Gross Pathology Cellular Pathology Morphologic Pathology Molecular/Predictive Pathology Antonio Benivieni (1443-1502): First autopsy
1 Traditional pathology Molecular pathology 2 Evolution of Pathology Gross Pathology Cellular Pathology Morphologic Pathology Molecular/Predictive Pathology Antonio Benivieni (1443-1502): First autopsy
Transform genomic data into real-life results
 CLINICAL SUMMARY Transform genomic data into real-life results Biomarker testing and targeted therapies can drive improved outcomes in clinical practice New FDA-Approved Broad Companion Diagnostic for
CLINICAL SUMMARY Transform genomic data into real-life results Biomarker testing and targeted therapies can drive improved outcomes in clinical practice New FDA-Approved Broad Companion Diagnostic for
Expanded View Figures
 EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
American Society of Cytopathology Companion Society Symposium Uses and Misuses of Ancillary Tests in Cytopathology
 American Society of Cytopathology Companion Society Symposium Uses and Misuses of Ancillary Tests in Cytopathology Zubair W. Baloch. MD, PhD. Professor of Pathology, UPENN Medical Center Perelman School
American Society of Cytopathology Companion Society Symposium Uses and Misuses of Ancillary Tests in Cytopathology Zubair W. Baloch. MD, PhD. Professor of Pathology, UPENN Medical Center Perelman School
SUPPLEMENTARY APPENDIX
 SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
Refining Prognosis of Early Stage Lung Cancer by Molecular Features (Part 2): Early Steps in Molecularly Defined Prognosis
 5/17/13 Refining Prognosis of Early Stage Lung Cancer by Molecular Features (Part 2): Early Steps in Molecularly Defined Prognosis Johannes Kratz, MD Post-doctoral Fellow, Thoracic Oncology Laboratory
5/17/13 Refining Prognosis of Early Stage Lung Cancer by Molecular Features (Part 2): Early Steps in Molecularly Defined Prognosis Johannes Kratz, MD Post-doctoral Fellow, Thoracic Oncology Laboratory
Introduction to Cancer Bioinformatics and cancer biology. Anthony Gitter Cancer Bioinformatics (BMI 826/CS 838) January 20, 2015
 Introduction to Cancer Bioinformatics and cancer biology Anthony Gitter Cancer Bioinformatics (BMI 826/CS 838) January 20, 2015 Why cancer bioinformatics? Devastating disease, no cure on the horizon Major
Introduction to Cancer Bioinformatics and cancer biology Anthony Gitter Cancer Bioinformatics (BMI 826/CS 838) January 20, 2015 Why cancer bioinformatics? Devastating disease, no cure on the horizon Major
609G: Concepts of Cancer Genetics and Treatments (3 credits)
 Master of Chemical and Life Sciences Program College of Computer, Mathematical, and Natural Sciences 609G: Concepts of Cancer Genetics and Treatments (3 credits) Text books: Principles of Cancer Genetics,
Master of Chemical and Life Sciences Program College of Computer, Mathematical, and Natural Sciences 609G: Concepts of Cancer Genetics and Treatments (3 credits) Text books: Principles of Cancer Genetics,
Cancer. The fundamental defect is. unregulated cell division. Properties of Cancerous Cells. Causes of Cancer. Altered growth and proliferation
 Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Cancer The fundamental defect is unregulated cell division. Properties of Cancerous Cells Altered growth and proliferation Loss of growth factor dependence Loss of contact inhibition Immortalization Alterated
Dr Yvonne Wallis Consultant Clinical Scientist West Midlands Regional Genetics Laboratory
 Dr Yvonne Wallis Consultant Clinical Scientist West Midlands Regional Genetics Laboratory Personalised Therapy/Precision Medicine Selection of a therapeutic drug based on the presence or absence of a specific
Dr Yvonne Wallis Consultant Clinical Scientist West Midlands Regional Genetics Laboratory Personalised Therapy/Precision Medicine Selection of a therapeutic drug based on the presence or absence of a specific
Introduction to Cancer Biology
 Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Introduction to Cancer Biology Robin Hesketh Multiple choice questions (choose the one correct answer from the five choices) Which ONE of the following is a tumour suppressor? a. AKT b. APC c. BCL2 d.
Development of Carcinoma Pathways
 The Construction of Genetic Pathway to Colorectal Cancer Moriah Wright, MD Clinical Fellow in Colorectal Surgery Creighton University School of Medicine Management of Colon and Diseases February 23, 2019
The Construction of Genetic Pathway to Colorectal Cancer Moriah Wright, MD Clinical Fellow in Colorectal Surgery Creighton University School of Medicine Management of Colon and Diseases February 23, 2019
Developments in small cell lung cancer G. Giaccone, MD PhD Chief, Medical Oncology Branch and Affiliates National Cancer Institute Bethesda MD USA
 Developments in small cell lung cancer G. Giaccone, MD PhD Chief, Medical Oncology Branch and Affiliates National Cancer Institute Bethesda MD USA Geneva April 20, 2012 Neuroendocrine tumors of lung Typical
Developments in small cell lung cancer G. Giaccone, MD PhD Chief, Medical Oncology Branch and Affiliates National Cancer Institute Bethesda MD USA Geneva April 20, 2012 Neuroendocrine tumors of lung Typical
New Drug development and Personalized Therapy in The Era of Molecular Medicine
 New Drug development and Personalized Therapy in The Era of Molecular Medicine Ramesh K. Ramanathan MD Virginia G. Piper Cancer Center Translational Genomics Research Institute Scottsdale, AZ Clinical
New Drug development and Personalized Therapy in The Era of Molecular Medicine Ramesh K. Ramanathan MD Virginia G. Piper Cancer Center Translational Genomics Research Institute Scottsdale, AZ Clinical
National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant
 National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant Reporting Period July 1, 2011 June 30, 2012 Formula Grant Overview The National Surgical
National Surgical Adjuvant Breast and Bowel Project (NSABP) Foundation Annual Progress Report: 2009 Formula Grant Reporting Period July 1, 2011 June 30, 2012 Formula Grant Overview The National Surgical
Supplementary Figure 1: Tissue of Origin analysis on 152 cell lines. (a) Heatmap representation of the 30 Tissue scores for the 152 cell lines.
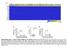 Supplementary Figure 1: Tissue of Origin analysis on 152 cell lines. (a) Heatmap representation of the 30 Tissue scores for the 152 cell lines. The scores summarize the global expression of the tissue
Supplementary Figure 1: Tissue of Origin analysis on 152 cell lines. (a) Heatmap representation of the 30 Tissue scores for the 152 cell lines. The scores summarize the global expression of the tissue
Ultrasound-Guided Fine-Needle Aspiration of Thyroid Nodules: New events
 Ultrasound-Guided Fine-Needle Aspiration of Thyroid Nodules: New events Sandrine Rorive, M.D., PhD. Erasme Hospital - Université Libre de Bruxelles (ULB) INTRODUCTION The assessment of thyroid nodules
Ultrasound-Guided Fine-Needle Aspiration of Thyroid Nodules: New events Sandrine Rorive, M.D., PhD. Erasme Hospital - Université Libre de Bruxelles (ULB) INTRODUCTION The assessment of thyroid nodules
Oncogenes and Tumor Suppressors MCB 5068 November 12, 2013 Jason Weber
 Oncogenes and Tumor Suppressors MCB 5068 November 12, 2013 Jason Weber jweber@dom.wustl.edu Oncogenes & Cancer DNA Tumor Viruses Simian Virus 40 p300 prb p53 Large T Antigen Human Adenovirus p300 E1A
Oncogenes and Tumor Suppressors MCB 5068 November 12, 2013 Jason Weber jweber@dom.wustl.edu Oncogenes & Cancer DNA Tumor Viruses Simian Virus 40 p300 prb p53 Large T Antigen Human Adenovirus p300 E1A
Nature Medicine: doi: /nm.3967
 Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Nature Genetics: doi: /ng Supplementary Figure 1. Clinical timeline for the discovery WES cases.
 Supplementary Figure 1 Clinical timeline for the discovery WES cases. This illustrates the timeline of the disease events during the clinical course of each patient s disease, further indicating the available
Supplementary Figure 1 Clinical timeline for the discovery WES cases. This illustrates the timeline of the disease events during the clinical course of each patient s disease, further indicating the available
Molecular Markers in Fine Needle Aspirates of the Thyroid
 Medical Policy Manual Genetic Testing, Policy No. 49 Molecular Markers in Fine Needle Aspirates of the Thyroid Next Review: April 2019 Last Review: June 2018 Effective: July 1, 2018 IMPORTANT REMINDER
Medical Policy Manual Genetic Testing, Policy No. 49 Molecular Markers in Fine Needle Aspirates of the Thyroid Next Review: April 2019 Last Review: June 2018 Effective: July 1, 2018 IMPORTANT REMINDER
Clasificación Molecular del Cáncer de Próstata. JM Piulats
 Clasificación Molecular del Cáncer de Próstata JM Piulats Introduction The Gleason score is the major method for prostate cancer tissue grading and the most important prognostic factor in this disease.
Clasificación Molecular del Cáncer de Próstata JM Piulats Introduction The Gleason score is the major method for prostate cancer tissue grading and the most important prognostic factor in this disease.
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Evaluation and Management of Thyroid Nodules. Nick Vernetti, MD, FACE Palm Medical Group Las Vegas, Nevada
 Evaluation and Management of Thyroid Nodules Nick Vernetti, MD, FACE Palm Medical Group Las Vegas, Nevada Disclosure Consulting Amgen Speaking Amgen Objectives Understand the significance of incidental
Evaluation and Management of Thyroid Nodules Nick Vernetti, MD, FACE Palm Medical Group Las Vegas, Nevada Disclosure Consulting Amgen Speaking Amgen Objectives Understand the significance of incidental
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature23291 Supplementary discussion part 1 Our model suggests that, in tumours that express Class 3 BRAF mutants and activated RAS, both BRAF MUT /RAF WT heterodimers and WT RAF dimers are
doi:10.1038/nature23291 Supplementary discussion part 1 Our model suggests that, in tumours that express Class 3 BRAF mutants and activated RAS, both BRAF MUT /RAF WT heterodimers and WT RAF dimers are
IntelliGENSM. Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community.
 IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis
 The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
Medical Policy Manual. Topic: Molecular Markers in Fine Needle Aspirates of the Thyroid. Date of Origin: April 2013
 Medical Policy Manual Topic: Molecular Markers in Fine Needle Aspirates of the Thyroid Date of Origin: April 2013 Section: Genetic Testing Last Reviewed Date: April 2014 Policy No: 49 Effective Date: July
Medical Policy Manual Topic: Molecular Markers in Fine Needle Aspirates of the Thyroid Date of Origin: April 2013 Section: Genetic Testing Last Reviewed Date: April 2014 Policy No: 49 Effective Date: July
MET skipping mutation, EGFR
 New NSCLC biomarkers in clinical research: detection of MET skipping mutation, EGFR T790M, and other important biomarkers Fernando López-Ríos Laboratorio de Dianas Terapéuticas Hospital Universitario HM
New NSCLC biomarkers in clinical research: detection of MET skipping mutation, EGFR T790M, and other important biomarkers Fernando López-Ríos Laboratorio de Dianas Terapéuticas Hospital Universitario HM
MATCHMAKING IN ONCOLOGY CHALLENGES AND COMBINATION STRATEGY FOR NOVEL TARGETED AGENTS
 MATCHMAKING IN ONCOLOGY CHALLENGES AND COMBINATION STRATEGY FOR NOVEL TARGETED AGENTS DR. RACHEL E. LAING*, OLIVIER LESUEUR*, STAN HOPKINS AND DR. SEAN X. HU MAY 2012 Recent advances in oncology, such
MATCHMAKING IN ONCOLOGY CHALLENGES AND COMBINATION STRATEGY FOR NOVEL TARGETED AGENTS DR. RACHEL E. LAING*, OLIVIER LESUEUR*, STAN HOPKINS AND DR. SEAN X. HU MAY 2012 Recent advances in oncology, such
number Done by Corrected by Doctor Maha Shomaf
 number 19 Done by Waseem Abo-Obeida Corrected by Abdullah Zreiqat Doctor Maha Shomaf Carcinogenesis: the molecular basis of cancer. Non-lethal genetic damage lies at the heart of carcinogenesis and leads
number 19 Done by Waseem Abo-Obeida Corrected by Abdullah Zreiqat Doctor Maha Shomaf Carcinogenesis: the molecular basis of cancer. Non-lethal genetic damage lies at the heart of carcinogenesis and leads
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Molecular biopathology of thyroid tumors
 Molecular biopathology of thyroid tumors Philippe Vielh MD, PhD, FIAC Director of Cytopathology Deputy Director of Anatomic Pathology National Health Laboratory of Luxembourg Past President of the International
Molecular biopathology of thyroid tumors Philippe Vielh MD, PhD, FIAC Director of Cytopathology Deputy Director of Anatomic Pathology National Health Laboratory of Luxembourg Past President of the International
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
