Identification of Causal Genetic Drivers of Human Disease through Systems-Level Analysis of Regulatory Networks
|
|
|
- Sabina Goodwin
- 5 years ago
- Views:
Transcription
1 Biologists are from Venus, Mathematicians are from Mars, They cosegregate on Earth, And conditionally associate to create a DIGGIT. Identification of Causal Genetic Drivers of Human Disease through Systems-Level Analysis of Regulatory Networks JAMES C. CHEN MARIANO J. ALVAREZ FLAMINIA TALOS HARSHIL DHRUV
2 Motivation 1. Identification of Driver Mutations is usually performed with statistical models. 2. These models can identify only the highly penetrant and frequent driver events. To achieve statistical power (in context of multiple hypothesis-testing correction), these models need large cohorts and/or large effect sizes. 3. Moreover, these models typically do not provide mechanistic insight. 4. On the other hand, Gene-based biochemical studies can provide insight into regulatory mechanisms but do not scale.
3 Problem Can we identify genetic determinants of a disease: Can we go beyond the highly penetrant and frequent driver events While remaining statistically rigorous Without using extremely large cohorts Can such an algorithm provide mechanistic insight into the process by which these genetic determinants play out their effect?
4 Idea 1. Overall Idea: Diverse alteration patterns induce common aberrant signals. These signals converge on regulatory modules and associated MR proteins that represent key regulatory bottlenecks. Dysregulation of these bottlenecks is both necessary and sufficient for disease initiation/progression. Once MR proteins and modules representing regulatory bottlenecks are identified, driver genetic events must be harbored either by these MRs or by their upstream pathways. 2. Algorithm can identify these driver genetic events by systematically exploring regulatory/signaling networks upstream of these MR genes: Approach is likely to collapse the number of testable hypotheses. Approach may provide regulatory clues to help elucidate associated mechanisms. Solution: DIGGIT: Driver Gene Inference by Genetical-Genomics and Mutual Information
5 DIGGIT: Summary of Findings 1. Combining cellular networks, gene expression, and genomic data (DIGGIT) finds novel driver mutations. 2. Uncovered KLHL9 deletions as upstream activators of two previously established Master Regulators of the subtype, C/EBPβ and C/EBPδ. 3. KLHL9 deletions predict mesenchymal transformation and poorest prognosis in GBM. 4. KLHL9 post-translationally regulates CEBPβ/δ. 5. Rescue of KLHL9 expression inhibits tumor growth by inducing degradation of C/EBP proteins and abrogating the mesenchymal signature. 6. DIGGIT can be used on any genetic disease with matched expression and genomic data.
6 MES-GBM: An Ideal Candidate 1. Glioblastoma Multiforme (GBM) is the most common human brain malignancy. 2. Virtually incurable, very aggressive and deadly - average survival of months post-diagnosis. 3. Three subtypes associated with expression of mesenchymal, proliferative, and proneural (PN) genes. 4. MES-GBM has the worst prognosis. 5. Despite multiple studies, genetic determinants of MES-GBM are largely elusive. 6. Provides an ideal context to test this rationale, as its established genetic determinants account for < 25% of the patients.
7 Link to Prior Work 1. In 2010 (The Transcriptional Network for Mesenchymal Transformation of Brain Tumours), reported that aberrant co-activation of the transcription factors (TFs) C/EBPβ, C/EBPδ, and STAT3 is necessary and sufficient to induce mesenchymal reprogramming in GBM. 2. This suggested that this TF module represents an obligate pathway or regulatory bottleneck between driver alterations and aberrant mesenchymal program activity. 3. Hypothesize that the genetic drivers of MES-GBM are either among these genes or in their upstream pathways.
8 Mutual Information Slides borrowed from University of Wisconsin, Madison (CS 760) University of Illinois, Chicago (ECE 534)
9 Entropy
10 Entropy
11 Entropy: Example The Entropy of a randomly selected letter in an English document is about 4.11 bits. Assuming its probability is as given in the table, we obtain this number by averaging log 1/p i (shown in the fourth column) under the probability distribution (third column)
12 Entropy is Important
13 Mutual Information
14 Mutual Information and Entropy
15 Conditional Mutual Information
16 Mutual Information and Correlation Correlation: 1. Correlation measures the linear relationship or monotonic relationship (e.g. Pearson's correlation or Spearman's correlation) between two variables, X and Y. Mutual Information: 1. Mutual information is more general and measures the reduction of uncertainty in Y after observing X. 2. It is the KL distance between the joint density and the product of the individual densities. 3. So MI can measure non-monotonic relationships and other more complicated relationships.
17 DIGGIT Methods / Process
18 DIGGIT: Overall Process 1. 5-step pipeline process 2. Inputs: Large set of Gene Expression Profiles (GEPD) Sample matched Genetic Variant Profiles (GVPD) Accurate and comprehensive repertoire of cell-contextspecific molecular interactions (Interactome) Overall flowchart of the DIGGIT pipeline. Green: Use of MR Inference results Red arrows: Use of F-CNVGs results Blue arrows: MINDy/aQTL analysis results 3. Output: A p-value ranked list of candidate driver F-CNVGs.
19 Step-0: ARACNE 1. ARACNE (Algorithm for the Reconstruction of Accurate Cellular Networks), a novel algorithm, uses microarray expression profiles to reverse engineer human regulatory network. 2. Specifically designed to scale up to the complexity of regulatory networks in mammalian cells, yet general enough to address a wider range of network deconvolution problems. 3. This method uses an information theoretic approach (Mutual Information) to eliminate the vast majority of indirect interactions typically inferred by pairwise analysis. 4. On synthetic datasets, ARACNE achieves extremely low error rates and significantly outperforms established methods, such as Relevance Networks and Bayesian Networks. 5. DIGGIT uses ARACNE to reverse engineer the cellular network (Interactome) from GEPD
20 Step-1: MR Analysis Objective: Identify candidate MRs as TFs that activate over-expressed and repress under-expr genes. Inputs: 1. Context specific regulatory network (Interactome) rev-engineered from GEPD set 2. Gene expression signature of interest Results: Identified 6 MR genes - C/EBPβ, C/EBPδ, STAT3, BHLHB2, RUNX1, and FOSL2 1. Inferred using the MARINa algorithm. 2. One MR (blue circle) is represented in the panel. 3. Grey circles represent the repertoire of genetic alterations that may be associated with the phenotype 4. Those within the two diagonal lines (funnel) represent alterations in pathways upstream of the MR. 5. The red circle represents a bona fide causal driver alteration.
21 Step-2: F-CNVG Analysis Objective: Identify candidate functional CNVs (F-CNVGs). Inputs: 1. GEPD & sample matched GVPD. Results: Identified 1,486 candidate F-CNVGs. Inferred F-CNVGs included most genes previously reported as GBM drivers (14/18 > 88%). 1. F-CNVGs are determined by association analysis of copy number and gene expression. 2. Select copy-number alterations (CNVGs) whose ploidy is informative of gene expression as candidate functional CNVs (F-CNVGs). 3. Assessed based on (1) mutual information (MI) between copy number and expression or (2) differential expression in wild-type (WT) versus amplified/deleted samples. 4. Removes a large number of genes whose expression is not affected by ploidy. 5. The insert shows two examples: (a) an example of no dependency between copy number and expression and not selected as a candidate F-CNVG and (b) an example with highly significant dependency and thus selected as a candidate F-CNVG.
22 Step-3: MINDy Analysis Objective: Identify F-CNVGs that are candidate post-translational modulators of MR activity. Inputs: MR list(step 1) & F-CNVG list (step 2). Output: Generates a p value-ranked list of candidate F-CNVGs in pathways upstream of MR genes. Results: Identified 92 statistically significant candidate MES-MR modulators. 1. Use Conditional Mutual Information: Compute the cmi I[MR;T M], where M is a candidate modulator gene and T is an ARACNe-inferred MR-target gene. 2. Blue arrows represent physical signal-transduction interactions upstream of the MR. 3. Green arrows represent one specific M MR T triplet tested by MINDy, as an example. 4. MINDy does not infer the blue arrows but only the fact that a protein is an upstream modulator of MR activity.
23 CMI in MINDy Analysis
24 Step-4: aqtl Analysis Objective: Identify F-CNVGs whose alterations cosegregate with aberrant MR activity. Inputs: MR list (step 1), F-CNVG list (step 2), GEPD data set, and the Interactome. Output: Generates a p value-ranked list of candidate F-CNVG-aQTL. Results: 125 out of 1,486 F-CNVGs from step 2 were inferred as aqtls. 1. Activity quantitative trait loci (aqtl) are inferred based on the statistical significance of the MI between copy number and MR activity. 2. Differential MR activity is inferred from their differential target expression, using a single-sample version of MARINa. Cosegregation computed (shown by the blue arrows). 3. The vertical gradient rectangle shows all genes sorted from the most over-expressed (red) to the most underexpressed (blue), when comparing samples with copy-number alterations in a gene (Gene X) (thick red lines) to WT samples (thin black lines). 4. If MR targets significantly cosegregate with the differential expression signature (i.e., if positively regulated and repressed MR targets, shown as red and blue bars, are over- and under-expressed, respectively, as shown), then Gene X alterations are likely to affect MR-activity.
25 Step-5: Conditional Association Analysis Objective: Identify F-CNVGs that abrogate all other associations with the phenotype (e.g., the MES- GBM subtype) when samples harboring their alterations are removed from the analysis. Inputs: MINDy/aQTL-prioritized F-CNVGs (steps 3/4), a phenotypic classifier, and GEPD data set Output: Generates a final p value-ranked list of candidate driver F-CNVGs Results: C/EBPδ and KLHL9 abrogated assciation of all other F-CNVGs, while remaining significant Conditional analysis discarded CDKN2A, a well-established tumor suppressor 1. Use conditional association analysis 2. Each cell shows the statistical significance of the association between the i-th gene (rows) and the phenotype of interest (as a heatmap), when considering only samples that have no alterations in the j-th gene (columns). 3. For instance, when conditioning on G3, no other gene is significantly associated with the subtype, whereas G3 is still significantly associated with the subtype when conditioning on G1, G2, or G4. 4. This suggests that G3 is a bona fide driver gene.
26 DIGGIT Integrative Analysis Infers Candidate MES-GBM Driver Mutations DIGGIT analysis of pathways upstream of MES-GBM MRs identifies CEBPδ amplification and KLHL9 deletions as candidate genetic determinants of the GBM-MES subtype. p values shown represent the integrated p value of the aqtl and MINDy steps. Co-mutated F-CNVGs are shown as a network, with distance between connected nodes inversely proportional to the statistical significance of their cosegregation, as assessed by Fisher s exact test (FET). Only statistically significant pairs are shown (p = 0.05, corrected), with amplifications and deletions represented as blue and red nodes, respectively.
27 DIGGIT Integrative Analysis Infers Candidate MES-GBM Driver Mutations Conditional association analysis for the two main co-segregating mutation clusters identified by DIGGIT. Color scale in the matrix cell (i,j) represents the strength of association ( log10(p)) between the i-th F- CNVG (row) and the MES subtype, conditional to removing samples with alterations in the j-th F-CNVG (column). Effect size of DIGGIT-inferred genetic determinants of the MES-GBM subtype. Classical GBM oncogenes are shown only as a reference, for comparison purposes.
28 Key Takeaways 1 2 Only C/EBPδ and KLHL9 abrogated association of all other F-CNVGs, while remaining significant when conditioning on other F-CNVGs Conditional analysis discarded CDKN2A (a well-established tumor suppressor located proximally to KLHL9) as a candidate causal driver of MES-GBM. 3 4 C/EBPδ amp and KLHL9 / events account for 48% of TCGA MES-GBM samples Along with independent deletions/mutations of NF1 covering an additional 8%, these may constitute the most common subtype drivers.
29 Association of KLHL9 Deletions Is Confirmed in an Independent Cohort 1. Tested whether association of KLHL9 deletions with poor prognosis could be validated in an independent cohort. 2. Analyzed 63 FFPEs, representing 40 poor-prognosis (survival < 35 weeks) and 23 goodprognosis (survival > 130 weeks) GBM samples. 3. Quantitative genomic PCR revealed higher frequency of homozygous KLHL9 deletions in poor-prognosis (21/40) versus good-prognosis samples (4/23) (p = by FET). Even higher frequency (>50%) than in TCGA samples (38%). 4. IHC staining of 10 KLHL9 / and 10 KLHL9 WT confirmed association with aberrant C/EBPβ and C/EBPδ protein expression in vivo (odds ratio 12.25, p = 0.028). 5. Confirms KLHL9 / events as poor-prognosis biomarkers and their association with aberrant MES-MR activity in vivo. No KLHL9 missense or nonsense mutations were detected.
30 Association of KLHL9 Deletions Is Confirmed in an Independent Cohort 1. Kaplan-Meier analysis of GBM samples in TCGA. 2. Patients with KLHL9 / and C/EBPδ Amp events are shown as a red curve 3. Proneural subtype patients are shown as a black curve 4. KLHL9 WT /CEBPδ WT samples are shown as a blue curve 5. Kaplan-Meier p values are shown, including p1 (red versus blue) and p2 (red versus black). 6. Survival for patients with each specific genotype is shown as vertical bars below the plot.
31 C/EBPδ and KLHL9 Alterations Are Predictive of Poor Prognosis in Multiple Tumors 1. Assessed whether C/EBPδ Amp and KLHL9 / events may be predictive of poor prognosis in GBM and other tumors. 2. In GBM, Kaplan-Meier analysis revealed significantly worse prognosis for patients harboring C/EBPδ Amp and KLHL9 / alterations, compared to either good-prognosis or C/EBPδ WT /KLHL9 WT patients. 3. None of the patients with these alterations survived longer than 36 weeks post-diagnosis, and patients harboring both events had the worst overall prognosis, suggesting a cooperative effect. 4. Thus, C/EBPδ Amp and KLHL9 / represent genetic biomarkers of poor prognosis, independent of subtype classification. 5. Kaplan-Meier analysis revealed that KLHL9 homozygous deletions and missense/nonsense mutations are associated with the worst prognosis also in Lung (LuAd) and Ovarian (OvCa) adenocarcinomas independent of CDKN2A status. 6. Gene set enrichment analysis (GSEA) confirmed aberrant C/EBPβ and/or C/EBPδ activity in KLHL9 / samples, suggesting a possible pan-cancer role of KLHL9 deletions via aberrant C/EBP activity.
32 1. Enrichment analysis of CEBPB and CEBPD ARACNe-inferred targets in genes differentially expressed in KLHL9 / versus KLHL9 WT samples. 2. Results for both lung adenocarcinoma (LuAd) and ovarian cancer (OV) are shown. 3. This analysis confirms that C/EBP protein activity is aberrantly increased by loss of KLHL9 function in multiple tumor types.
33 Kaplan-Meier analysis of the association between KLHL9 / alterations and poor prognosis in lung and serous ovarian adenocarcinoma, respectively. Analysis of inferred differential activity of C/EBPβ and C/EBPδ in KLHL9 / samples.
34 Ectopic (Unusual) KLHL9 Expression in GBM Cells Abrogates (stops) C/EBPβ and C/EBPδ Abundance 1. To mechanistically elucidate KLHL9-mediated regulation of established MES-MRs (C/EBPβ, C/EBPδ, and STAT3), rescued KLHL9 expression in homozygously deleted cells. 2. Used cell lines SF210 and SF763 cells labeled as KLHL9 / ;CDKN2A / ;C/EBP WT. 3. RNA-seq profiling revealed significant differential expression of ARACNe-inferred C/EBPβ and C/EBPδ targets by GSEA compared to controls. 4. Involved significant down-regulation of established MES markers: CHI3L1/YKL40, LIF, FOSL2, ACTA2, and FN1. 5. Observed significant reduction in C/EBPδ and more modest decrease in C/EBPβ protein levels. 6. Levels of phospho-stat3, representing the transcriptionally active isoform, were also reduced. 7. Exogenous expression of P16/INK4A (CDKN2A) had no effect on either C/EBPβ or C/EBPδ protein expression or on MES signature genes. 8. Results show that rescue of KLHL9 expression collapses the MES-GBM signature by downregulating C/EBPβ and C/EBPδ at the protein level.
35 Proteasomal Degradation of C/EBPb and C/EBPd Depends on KLHL9- Mediated Polyubiquitylation 1. KLHL9 s has a putative function as an adaptor of Cul3-based E3 ubiquitin ligase. 2. Tested KLHL9 s role in mediating polyubiquitylation-dependent proteasomal degradation of C/EBPb and C/EBPd. 3. Measured degradation and relative half-life of C/EBPb and C/EBPd following rescue of KLHL9 expression in SF MG-132-mediated proteasome inhibition abrogated C/EBPb and C/EBPd degradation, confirming that KLHL9 is required for their proteasomal processing.
36 More Mechanistic Insights 1. KLHL9 Mediates Polyubiquitylation of C/EBPb and C/EBPd Isoforms Confirmed that proteasomal degradation of C/EBPs depends on KLHL9-mediated interaction with the CUL3 E3 ligase complex Confirmed that KLHL9-mediated C/EBP regulation depends on a functional KLHL9-CUL3 E3 ligase complex 2. KLHL9 Expression Delays Exit from S Phase in Glioma Cells Confirmed that rescue of KLHL9 expression delays the cell cycle by imposing a late S/G2 checkpoint. 3. KLHL9 Expression in KLHL9 -/- Patient-Derived GBM Tumors Reduces Growth in Orthotopic Xenografts Experiments show that in vitro cell-cycle-dependent reduction in proliferative potential, induced by ectopic KLHL9 expression in human cell cultures, is recapitulated in vivo and induces retardation in tumor growth.
37 Unbiased Inference of Driver Alterations in BRCA and AD 1. Analysis of sample-matched CNV/expression data from the TCGA breast cancer (BRCA) cohort. Compiled a list of 25 alterations from a literature search of validated CNV alterations linked to BRCA tumorigenesis Final step (Conditional Association Analysis) yielded 35 F-CNVGs Of these, 19 (76%) could be matched in the 25-gene literature compiled list Only 5 of these were statistically significant by Genome Wide Association Studies (GWAS) 2. Analysis of sample-matched SNP/expression data from a recent integrative study of Alzheimer s disease. DIGGIT identified 13 F-SNPs significant by conditional association analysis. Among these, TYROBP was ranked 1st (p = 4.2 x ), achieving higher significance than even APOE, ranked 9th (p = 2.0 x )
Computational Inferences of Mutations Driving Mesenchymal Differentiation in Glioblastoma
 Computational Inferences of Mutations Driving Mesenchymal Differentiation in Glioblastoma James Chen Submitted in partial fulfillment of the requirements for the Doctor of Philosophy Degree in the Graduate
Computational Inferences of Mutations Driving Mesenchymal Differentiation in Glioblastoma James Chen Submitted in partial fulfillment of the requirements for the Doctor of Philosophy Degree in the Graduate
Glioblastoma Multiforme
 Glioblastoma Multiforme Highly malignant, invasive, difficult-to-treat primary brain tumor" " Frequency: 9,000 cases/year (peak age, 55 65 years)" " Recurrence: rapid growth; size may double every 10 days"
Glioblastoma Multiforme Highly malignant, invasive, difficult-to-treat primary brain tumor" " Frequency: 9,000 cases/year (peak age, 55 65 years)" " Recurrence: rapid growth; size may double every 10 days"
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
Package diggitdata. April 11, 2019
 Type Package Title Example data for the diggit package Version 1.14.0 Date 2014-08-29 Author Mariano Javier Alvarez Package diggitdata April 11, 2019 Maintainer Mariano Javier Alvarez
Type Package Title Example data for the diggit package Version 1.14.0 Date 2014-08-29 Author Mariano Javier Alvarez Package diggitdata April 11, 2019 Maintainer Mariano Javier Alvarez
HALLA KABAT * Outreach Program, mircore, 2929 Plymouth Rd. Ann Arbor, MI 48105, USA LEO TUNKLE *
 CERNA SEARCH METHOD IDENTIFIED A MET-ACTIVATED SUBGROUP AMONG EGFR DNA AMPLIFIED LUNG ADENOCARCINOMA PATIENTS HALLA KABAT * Outreach Program, mircore, 2929 Plymouth Rd. Ann Arbor, MI 48105, USA Email:
CERNA SEARCH METHOD IDENTIFIED A MET-ACTIVATED SUBGROUP AMONG EGFR DNA AMPLIFIED LUNG ADENOCARCINOMA PATIENTS HALLA KABAT * Outreach Program, mircore, 2929 Plymouth Rd. Ann Arbor, MI 48105, USA Email:
Breast cancer. Risk factors you cannot change include: Treatment Plan Selection. Inferring Transcriptional Module from Breast Cancer Profile Data
 Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
SUPPLEMENTARY APPENDIX
 SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs.
 Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
Supplementary Figure 1: Comparison of acgh-based and expression-based CNA analysis of tumors from breast cancer GEMMs. (a) CNA analysis of expression microarray data obtained from 15 tumors in the SV40Tag
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis
 The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Nature Genetics: doi: /ng Supplementary Figure 1. HOX fusions enhance self-renewal capacity.
 Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Supplementary Figure 1 HOX fusions enhance self-renewal capacity. Mouse bone marrow was transduced with a retrovirus carrying one of three HOX fusion genes or the empty mcherry reporter construct as described
Identification of Tissue Independent Cancer Driver Genes
 Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
User Guide. Association analysis. Input
 User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser
 Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser Melissa S. Cline 1*, Brian Craft 1, Teresa Swatloski 1, Mary Goldman 1, Singer Ma 1, David Haussler 1, Jingchun Zhu 1 1 Center for Biomolecular
Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser Melissa S. Cline 1*, Brian Craft 1, Teresa Swatloski 1, Mary Goldman 1, Singer Ma 1, David Haussler 1, Jingchun Zhu 1 1 Center for Biomolecular
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Expanded View Figures
 Molecular Systems iology Tumor CNs reflect metabolic selection Nicholas Graham et al Expanded View Figures Human primary tumors CN CN characterization by unsupervised PC Human Signature Human Signature
Molecular Systems iology Tumor CNs reflect metabolic selection Nicholas Graham et al Expanded View Figures Human primary tumors CN CN characterization by unsupervised PC Human Signature Human Signature
Understanding Genotype- Phenotype relations in Cancer via Network Approaches
 AlgoCSB Algorithmic Methods in Computational and Systems Biology Understanding Genotype- Phenotype relations in Cancer via Network Approaches Teresa Przytycka NIH / NLM / NCBI Phenotypes Journal Wisla
AlgoCSB Algorithmic Methods in Computational and Systems Biology Understanding Genotype- Phenotype relations in Cancer via Network Approaches Teresa Przytycka NIH / NLM / NCBI Phenotypes Journal Wisla
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Supplementary Information
 Supplementary Information Guided Visual Exploration of Genomic Stratifications in Cancer Marc Streit 1,6, Alexander Lex 2,6, Samuel Gratzl¹, Christian Partl³, Dieter Schmalstieg³, Hanspeter Pfister², Peter
Supplementary Information Guided Visual Exploration of Genomic Stratifications in Cancer Marc Streit 1,6, Alexander Lex 2,6, Samuel Gratzl¹, Christian Partl³, Dieter Schmalstieg³, Hanspeter Pfister², Peter
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Supplementary Information. Supplementary Figures
 Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
Supplementary. properties of. network types. randomly sampled. subsets (75%
 Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
Machine-Learning on Prediction of Inherited Genomic Susceptibility for 20 Major Cancers
 Machine-Learning on Prediction of Inherited Genomic Susceptibility for 20 Major Cancers Sung-Hou Kim University of California Berkeley, CA Global Bio Conference 2017 MFDS, Seoul, Korea June 28, 2017 Cancer
Machine-Learning on Prediction of Inherited Genomic Susceptibility for 20 Major Cancers Sung-Hou Kim University of California Berkeley, CA Global Bio Conference 2017 MFDS, Seoul, Korea June 28, 2017 Cancer
Introduction. Introduction
 Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Computer Science, Biology, and Biomedical Informatics (CoSBBI) Outline. Molecular Biology of Cancer AND. Goals/Expectations. David Boone 7/1/2015
 Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Supplementary Materials for
 www.sciencemag.org/content/355/6332/eaai8478/suppl/dc1 Supplementary Materials for Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq Andrew S. Venteicher,
www.sciencemag.org/content/355/6332/eaai8478/suppl/dc1 Supplementary Materials for Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq Andrew S. Venteicher,
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
Research Strategy: 1. Background and Significance
 Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Nature Genetics: doi: /ng Supplementary Figure 1. Somatic coding mutations identified by WES/WGS for 83 ATL cases.
 Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Nature Medicine: doi: /nm.4439
 Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
Supplemental Information. Integrated Genomic Analysis of the Ubiquitin. Pathway across Cancer Types
 Cell Reports, Volume 23 Supplemental Information Integrated Genomic Analysis of the Ubiquitin Pathway across Zhongqi Ge, Jake S. Leighton, Yumeng Wang, Xinxin Peng, Zhongyuan Chen, Hu Chen, Yutong Sun,
Cell Reports, Volume 23 Supplemental Information Integrated Genomic Analysis of the Ubiquitin Pathway across Zhongqi Ge, Jake S. Leighton, Yumeng Wang, Xinxin Peng, Zhongyuan Chen, Hu Chen, Yutong Sun,
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer
 Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Genetic alterations of histone lysine methyltransferases and their significance in breast cancer Supplementary Materials and Methods Phylogenetic tree of the HMT superfamily The phylogeny outlined in the
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
Nature Genetics: doi: /ng Supplementary Figure 1. Phenotypic characterization of MES- and ADRN-type cells.
 Supplementary Figure 1 Phenotypic characterization of MES- and ADRN-type cells. (a) Bright-field images showing cellular morphology of MES-type (691-MES, 700-MES, 717-MES) and ADRN-type (691-ADRN, 700-
Supplementary Figure 1 Phenotypic characterization of MES- and ADRN-type cells. (a) Bright-field images showing cellular morphology of MES-type (691-MES, 700-MES, 717-MES) and ADRN-type (691-ADRN, 700-
Computational Investigation of Homologous Recombination DNA Repair Deficiency in Sporadic Breast Cancer
 University of Massachusetts Medical School escholarship@umms Open Access Articles Open Access Publications by UMMS Authors 11-16-2017 Computational Investigation of Homologous Recombination DNA Repair
University of Massachusetts Medical School escholarship@umms Open Access Articles Open Access Publications by UMMS Authors 11-16-2017 Computational Investigation of Homologous Recombination DNA Repair
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
SUPPLEMENTARY FIGURES: Supplementary Figure 1
 SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Bredel M, Scholtens DM, Yadav AK, et al. NFKBIA deletion in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Bredel M, Scholtens DM, Yadav AK, et al. NFKBIA deletion in
MicroRNA expression profiling and functional analysis in prostate cancer. Marco Folini s.c. Ricerca Traslazionale DOSL
 MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
User s Manual Version 1.0
 User s Manual Version 1.0 #639 Longmian Avenue, Jiangning District, Nanjing,211198,P.R.China. http://tcoa.cpu.edu.cn/ Contact us at xiaosheng.wang@cpu.edu.cn for technical issue and questions Catalogue
User s Manual Version 1.0 #639 Longmian Avenue, Jiangning District, Nanjing,211198,P.R.China. http://tcoa.cpu.edu.cn/ Contact us at xiaosheng.wang@cpu.edu.cn for technical issue and questions Catalogue
COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION
 COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION Pierre Martinez, Nicholas McGranahan, Nicolai Juul Birkbak, Marco Gerlinger, Charles Swanton* SUPPLEMENTARY INFORMATION SUPPLEMENTARY
COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION Pierre Martinez, Nicholas McGranahan, Nicolai Juul Birkbak, Marco Gerlinger, Charles Swanton* SUPPLEMENTARY INFORMATION SUPPLEMENTARY
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplemental Figure S1. RANK expression on human lung cancer cells.
 Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
Supplemental Figure S1. RANK expression on human lung cancer cells. (A) Incidence and H-Scores of RANK expression determined from IHC in the indicated primary lung cancer subgroups. The overall expression
Biomarker development in the era of precision medicine. Bei Li, Interdisciplinary Technical Journal Club
 Biomarker development in the era of precision medicine Bei Li, 23.08.2016 Interdisciplinary Technical Journal Club The top ten highest-grossing drugs in the United States help between 1 in 25 and 1 in
Biomarker development in the era of precision medicine Bei Li, 23.08.2016 Interdisciplinary Technical Journal Club The top ten highest-grossing drugs in the United States help between 1 in 25 and 1 in
IPA Advanced Training Course
 IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
About OMICS Group Conferences
 About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
About OMICS Group OMICS Group International is an amalgamation of Open Access publications and worldwide international science conferences and events. Established in the year 2007 with the sole aim of
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
The Cancer Genome Atlas & International Cancer Genome Consortium
 The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
Integrative Omics for The Systems Biology of Complex Phenotypes
 Integrative Omics for The Systems Biology of Complex Phenotypes Mehmet Koyutürk Case Western Reserve University (1) Electrical Engineering and Computer Science (2) Center for Proteomics and Bioinformatics
Integrative Omics for The Systems Biology of Complex Phenotypes Mehmet Koyutürk Case Western Reserve University (1) Electrical Engineering and Computer Science (2) Center for Proteomics and Bioinformatics
Supplemental Information. Molecular, Pathological, Radiological, and Immune. Profiling of Non-brainstem Pediatric High-Grade
 Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
Expanded View Figures
 Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer.
 Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1. Copy Number Alterations TP53 Mutation Type. C-class TP53 WT. TP53 mut. Nature Genetics: doi: /ng.
 Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium
 Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium Ulrike Peters Fred Hutchinson Cancer Research Center University of Washington U01-CA137088-05, PI: Peters
Integration of Cancer Genome into GECCO- Genetics and Epidemiology of Colorectal Cancer Consortium Ulrike Peters Fred Hutchinson Cancer Research Center University of Washington U01-CA137088-05, PI: Peters
cis-regulatory enrichment analysis in human, mouse and fly
 cis-regulatory enrichment analysis in human, mouse and fly Zeynep Kalender Atak, PhD Laboratory of Computational Biology VIB-KU Leuven Center for Brain & Disease Research Laboratory of Computational Biology
cis-regulatory enrichment analysis in human, mouse and fly Zeynep Kalender Atak, PhD Laboratory of Computational Biology VIB-KU Leuven Center for Brain & Disease Research Laboratory of Computational Biology
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
ARTICLE RESEARCH. Macmillan Publishers Limited. All rights reserved
 Extended Data Figure 6 Annotation of drivers based on clinical characteristics and co-occurrence patterns. a, Putative drivers affecting greater than 10 patients were assessed for enrichment in IGHV mutated
Extended Data Figure 6 Annotation of drivers based on clinical characteristics and co-occurrence patterns. a, Putative drivers affecting greater than 10 patients were assessed for enrichment in IGHV mutated
DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK
 CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Figure S2. Distribution of acgh probes on all ten chromosomes of the RIL M0022
 96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
Gene expression profiling predicts clinical outcome of prostate cancer. Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M.
 SUPPLEMENTARY DATA Gene expression profiling predicts clinical outcome of prostate cancer Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M. Hoffman, William L. Gerald Table of Contents
SUPPLEMENTARY DATA Gene expression profiling predicts clinical outcome of prostate cancer Gennadi V. Glinsky, Anna B. Glinskii, Andrew J. Stephenson, Robert M. Hoffman, William L. Gerald Table of Contents
Supplementary Figure 1: LUMP Leukocytes unmethylabon to infer tumor purity
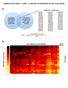 Supplementary Figure 1: LUMP Leukocytes unmethylabon to infer tumor purity A Consistently unmethylated sites (30%) in 21 cancer types 174,696
Supplementary Figure 1: LUMP Leukocytes unmethylabon to infer tumor purity A Consistently unmethylated sites (30%) in 21 cancer types 174,696
Review (V1) - The Hallmarks of Cancer
 Review (V1) - The Hallmarks of Cancer Robert A. Weinberg 1 Review (V1) - The Hallmarks of Cancer 2 Review (V1) - Number of somatic mutations in human cancers Top: children vs. adults Numbers in parentheses
Review (V1) - The Hallmarks of Cancer Robert A. Weinberg 1 Review (V1) - The Hallmarks of Cancer 2 Review (V1) - Number of somatic mutations in human cancers Top: children vs. adults Numbers in parentheses
Review (V1) - The Hallmarks of Cancer
 Review (V1) - The Hallmarks of Cancer Robert A. Weinberg 1 Review (V1) - The Hallmarks of Cancer 2 Review (V1) - Number of somatic mutations in human cancers Top: children vs. adults Numbers in parentheses
Review (V1) - The Hallmarks of Cancer Robert A. Weinberg 1 Review (V1) - The Hallmarks of Cancer 2 Review (V1) - Number of somatic mutations in human cancers Top: children vs. adults Numbers in parentheses
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD
 Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
CS2220 Introduction to Computational Biology
 CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Nature Immunology: doi: /ni Supplementary Figure 1. RNA-Seq analysis of CD8 + TILs and N-TILs.
 Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Supplementary Figure 1 RNA-Seq analysis of CD8 + TILs and N-TILs. (a) Schematic representation of the tumor and cell types used for the study. HNSCC, head and neck squamous cell cancer; NSCLC, non-small
Cellecta Overview. Started Operations in 2007 Headquarters: Mountain View, CA
 Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Putative effectors for prognosis in lung adenocarcinoma are ethnic and gender specific
 /, Advance Publications 2015 Putative effectors for prognosis in lung adenocarcinoma are ethnic and gender specific Andrew Woolston 1, Nardnisa Sintupisut 1, Tzu-Pin Lu 2, Liang-Chuan Lai 3, Mong-Hsun
/, Advance Publications 2015 Putative effectors for prognosis in lung adenocarcinoma are ethnic and gender specific Andrew Woolston 1, Nardnisa Sintupisut 1, Tzu-Pin Lu 2, Liang-Chuan Lai 3, Mong-Hsun
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Update on Genetic Testing for Melanoma
 Update on Genetic Testing for Melanoma Emily Y. Chu, M.D., Ph.D. Assistant Professor of Dermatology & Pathology and Laboratory Medicine Hospital of the University of Pennsylvania February 18, 2018 AAD
Update on Genetic Testing for Melanoma Emily Y. Chu, M.D., Ph.D. Assistant Professor of Dermatology & Pathology and Laboratory Medicine Hospital of the University of Pennsylvania February 18, 2018 AAD
Linking Proteomic and Transcriptional Data through the Interactome and Epigenome Reveals a Map of Oncogeneinduced
 Linking Proteomic and Transcriptional Data through the Interactome and Epigenome Reveals a Map of Oncogeneinduced Signaling The MIT Faculty has made this article openly available. Please share how this
Linking Proteomic and Transcriptional Data through the Interactome and Epigenome Reveals a Map of Oncogeneinduced Signaling The MIT Faculty has made this article openly available. Please share how this
Using Network Flow to Bridge the Gap between Genotype and Phenotype. Teresa Przytycka NIH / NLM / NCBI
 Using Network Flow to Bridge the Gap between Genotype and Phenotype Teresa Przytycka NIH / NLM / NCBI Journal Wisla (1902) Picture from a local fare in Lublin, Poland Genotypes Phenotypes Journal Wisla
Using Network Flow to Bridge the Gap between Genotype and Phenotype Teresa Przytycka NIH / NLM / NCBI Journal Wisla (1902) Picture from a local fare in Lublin, Poland Genotypes Phenotypes Journal Wisla
Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics, 2010
 Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics, 2010 C.J.Vaske et al. May 22, 2013 Presented by: Rami Eitan Complex Genomic
Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics, 2010 C.J.Vaske et al. May 22, 2013 Presented by: Rami Eitan Complex Genomic
Molecular Markers. Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC
 Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Nature Genetics: doi: /ng Supplementary Figure 1. PCA for ancestry in SNV data.
 Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Introduction to Cancer Bioinformatics and cancer biology. Anthony Gitter Cancer Bioinformatics (BMI 826/CS 838) January 20, 2015
 Introduction to Cancer Bioinformatics and cancer biology Anthony Gitter Cancer Bioinformatics (BMI 826/CS 838) January 20, 2015 Why cancer bioinformatics? Devastating disease, no cure on the horizon Major
Introduction to Cancer Bioinformatics and cancer biology Anthony Gitter Cancer Bioinformatics (BMI 826/CS 838) January 20, 2015 Why cancer bioinformatics? Devastating disease, no cure on the horizon Major
Latent Factor Analysis to Discover Pathway-Associated Putative Segmental Aneuploidies in Human Cancers
 Latent Factor Analysis to Discover Pathway-Associated Putative Segmental Aneuploidies in Human Cancers Joseph E. Lucas 1 *, Hsiu-Ni Kung 1,2, Jen-Tsan A. Chi 1,2 * 1 Institute for Genome Sciences and Policy,
Latent Factor Analysis to Discover Pathway-Associated Putative Segmental Aneuploidies in Human Cancers Joseph E. Lucas 1 *, Hsiu-Ni Kung 1,2, Jen-Tsan A. Chi 1,2 * 1 Institute for Genome Sciences and Policy,
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Results and Discussion of Receptor Tyrosine Kinase. Activation
 Results and Discussion of Receptor Tyrosine Kinase Activation To demonstrate the contribution which RCytoscape s molecular maps can make to biological understanding via exploratory data analysis, we here
Results and Discussion of Receptor Tyrosine Kinase Activation To demonstrate the contribution which RCytoscape s molecular maps can make to biological understanding via exploratory data analysis, we here
PBZ FT01_PBZ FT01_TZ FT01_NZ. interface zone (I) tumor zone (TZ) necrotic zone (NZ)
 Oncotarget, Supplementary Materials www.impactjournals.com/oncotarget/ SUPPLEMENTRY FLES ndividuals factor map (P) FT_ FT_ FT_ Dim (.%) Dim (.%) >% peripheral brain zone () around % interface zone () FT
Oncotarget, Supplementary Materials www.impactjournals.com/oncotarget/ SUPPLEMENTRY FLES ndividuals factor map (P) FT_ FT_ FT_ Dim (.%) Dim (.%) >% peripheral brain zone () around % interface zone () FT
The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response
 Abstract # 319030 Poster # F.9 The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response Jozsef Karman, Brian Johnston, Sofija Miljovska,
Abstract # 319030 Poster # F.9 The epigenetic landscape of T cell subsets in SLE identifies known and potential novel drivers of the autoimmune response Jozsef Karman, Brian Johnston, Sofija Miljovska,
SUPPLEMENTARY INFORMATION In format provided by Javier DeFelipe et al. (MARCH 2013)
 Supplementary Online Information S2 Analysis of raw data Forty-two out of the 48 experts finished the experiment, and only data from these 42 experts are considered in the remainder of the analysis. We
Supplementary Online Information S2 Analysis of raw data Forty-two out of the 48 experts finished the experiment, and only data from these 42 experts are considered in the remainder of the analysis. We
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Genomic tests to personalize therapy of metastatic breast cancers. Fabrice ANDRE Gustave Roussy Villejuif, France
 Genomic tests to personalize therapy of metastatic breast cancers Fabrice ANDRE Gustave Roussy Villejuif, France Future application of genomics: Understand the biology at the individual scale Patients
Genomic tests to personalize therapy of metastatic breast cancers Fabrice ANDRE Gustave Roussy Villejuif, France Future application of genomics: Understand the biology at the individual scale Patients
