Supplemental Table 1: List of genes contained on the custom Steroltalk v2 microarray prepared for this study
|
|
|
- Arline Dickerson
- 6 years ago
- Views:
Transcription
1 Supplemental Table 1: List of genes contained on the custom Steroltalk v2 microarray prepared for this study Gene GenBank No. Description Ch25h NM_ Cholesterol 25-hydroxylase Bile acid synthesis Cyp27a1 NM_ Cytochrome P450, family 27, subfamily a, polypeptide 1 Bile acid synthesis Cyp27b1 AB Cytochrome P450, family 27, subfamily b, polypeptide 1 Bile acid synthesis Cyp39a1 AF Cytochrome P450, family 39, subfamily a, polypeptide 1 Bile acid synthesis Cyp46a1 NM_ Cytochrome P450, family 46, subfamily a, polypeptide 1 Bile acid synthesis Cyp7a1 NM_ Cytochrome P450, family 7, subfamily a, polypeptide 1 Bile acid synthesis Cyp7b1 BC Cytochrome P450, family 7, subfamily b, polypeptide 1 Bile acid synthesis Cyp8b1 NM_ Cytochrome P450, family 8, subfamily b, polypeptide 1 Bile acid synthesis G6Pase NM_ Glucose-6-phosphatase catalytic Carbohydrate metabolism Gck BC Glucokinase Carbohydrate metabolism Gsk3b NM_ Glycogen synthase kinase 3 beta Carbohydrate metabolism Hk1 BC Hexokinase 1 Carbohydrate metabolism Hk2 BC Hexokinase 2 Carbohydrate metabolism Pck1 BC Phosphoenolpyruvate carboxykinase 1, cytosolic Carbohydrate metabolism Pdhb NM_ Pyruvate dehydrogenase (lipoamide) beta Carbohydrate metabolism Pfkm NM_ Phosphofructokinase, muscle Carbohydrate metabolism Pklr NM_ Pyruvate kinase liver and red blood cell Carbohydrate metabolism Pkm2 BC Pyruvate kinase, muscle Carbohydrate metabolism Adipoq BC Adiponectin, C1Q and collagen domain containing Cell signaling Adipor1 NM_ Adiponectin receptor 1 Cell signaling Adipor2 NM_ Adiponectin receptor 2 Cell signaling Arntl2 AY Aryl hydrocarbon receptor nuclear translocator-like 2 Cell signaling Camk1d NM_ Calcium/calmodulin-dependent protein kinase ID Cell signaling Camkk1 AF Calcium/calmodulin-dependent protein kinase kinase 1, alpha Cell signaling Casp3 NM_ Caspase 3, apoptosis-related cysteine peptidase Cell signaling Casp4 NM_ Caspase 4, apoptosis-related cysteine peptidase Cell signaling Cav1 NM_ Caveolin, caveolae protein 1 Cell signaling Egr1 M20157 Early growth response 1 Cell signaling Gckr NM_ Glucokinase regulatory protein Cell signaling Ghrl NM_ Ghrelin Cell signaling Igf1 NM_ Insulin-like growth factor 1 Cell signaling Irs1 AK Insulin receptor substrate 1 Cell signaling Irs2 XM_ Insulin receptor substrate 2 Cell signaling Jak1 BC Janus kinase 1 Cell signaling Jak2 BC Janus kinase 2 Cell signaling Jun BC Jun oncogene Cell signaling Lep U18812 Leptin Cell signaling Lepr BC Leptin receptor Cell signaling Lipe NM_ Lipase, hormone sensitive Cell signaling Mapk8 BC Mitogen activated protein kinase 8 Cell signaling Mapk9 BC Mitogen activated protein kinase 9 Cell signaling Msr1 BC Macrophage scavenger receptor 1 Cell signaling Msr1 AF Macrophage scavenger receptor 1 Cell signaling Nfkbia BC Nuclear factor of kappa light chain gene enhancer in B-cells inhibitor, alpha Cell signaling Nfkbie BC Nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, epsilon Cell signaling Ogt BC O-linked N-acetylglucosamine (GlcNAc) transferase Cell signaling Oprs1 NM_ Opioid receptor, sigma 1 Cell signaling
2 Pik3cg NM_ Phosphoinositide-3-kinase, catalytic, gamma polypeptide Cell signaling Ppp1r3c NM_ Protein phosphatase 1, regulatory (inhibitor) subunit 3C Cell signaling Prkaa1 NM_ Protein kinase, AMP-activated, alpha 1 catalytic subunit Cell signaling Prkaca BC Protein kinase, camp dependent, catalytic, alpha Cell signaling Prkca BC Protein kinase C, alpha Cell signaling Prkcb1 NM_ Protein kinase C, beta 1 Cell signaling Retn NM_ Resistin Cell signaling Serping1 BC Serine (or cysteine) peptidase inhibitor, clade G, member 1 Cell signaling Stat3 BC Signal transducer and activator of transcription 3 Cell signaling Sumo1 BC SMT3 suppressor of mif two 3 homolog 1 (yeast) Cell signaling Sumo2 BC SMT3 suppressor of mif two 3 homolog 2 (yeast) Cell signaling Cyp51 BC Cytochrome P450, family 51 Cholesterol biosynthesis Dhcr24 BC dehydrocholesterol reductase Cholesterol biosynthesis Dhcr7 BC dehydrocholesterol reductase Cholesterol biosynthesis Ebp NM_ Emopamil binding protein Cholesterol biosynthesis Fdft1 NM_ Farnesyl diphosphate farnesyl transferase 1, Squalene synthase Cholesterol biosynthesis Fdps NM_ Farnesyl diphosphate synthase 1 Cholesterol biosynthesis Hmgcr NM_ hydroxy-3-methylglutaryl-Coenzyme A reductase Cholesterol biosynthesis Hmgcs1 NM_ hydroxy-3-methylglutaryl-Coenzyme A synthase 1 Cholesterol biosynthesis Idi1 NM_ Isopentenyl-diphosphate delta isomerase Cholesterol biosynthesis Lss NM_ Lanosterol synthase Cholesterol biosynthesis Mvd NM_ Mevalonate (diphospho) decarboxylase Cholesterol biosynthesis Mvk NM_ Mevalonate kinase Cholesterol biosynthesis Nsdhl BC NAD(P) dependent steroid dehydrogenase-like Cholesterol biosynthesis Pmvk NM_ Phosphomevalonate kinase Cholesterol biosynthesis Sc4mol NM_ Sterol-C4-methyl oxidase-like Cholesterol biosynthesis Sc5d BC Sterol-C5-desaturase homolog Cholesterol biosynthesis Sqle BC Squalene epoxidase Cholesterol biosynthesis Arntl BC Aryl hydrocarbon receptor nuclear translocator-like Circadian regulation Clock AF Circadian locomoter output cycles kaput Circadian regulation Cry1 NM_ Cryptochrome 1 (photolyase-like) Circadian regulation Cry2 NM_ Cryptochrome 2 (photolyase-like) Circadian regulation Dbp BC D site albumin promoter binding protein Circadian regulation Per1 NM_ Period homolog 1 (Drosophila) Circadian regulation Per2 NM_ Period homolog 2 (Drosophila) Circadian regulation Per3 NM_ Period homolog 3 (Drosophila) Circadian regulation Timeless BC Timeless homolog (Drosophila) Circadian regulation Cyp1a1 NM_ Cytochrome P450, family 1, subfamily a, polypeptide 1 Drug metabolism Cyp1a2 NM_ Cytochrome P450, family 1, subfamily a, polypeptide 2 Drug metabolism Cyp1b1 BC Cytochrome P450, family 1, subfamily b, polypeptide 1 Drug metabolism Cyp2a12 NM_ Cytochrome P450, family 2, subfamily a, polypeptide 12 Drug metabolism Cyp2a4 BC Cytochrome P450, family 2, subfamily a, polypeptide 4 Drug metabolism Cyp2b10 AK Cytochrome P450, family 2, subfamily b, polypeptide 10 Drug metabolism Cyp2b13 NM_ Cytochrome P450, family 2, subfamily b, polypeptide 13 Drug metabolism Cyp2b9 NM_ Cytochrome P450, family 2, subfamily b, polypeptide 9 Drug metabolism Cyp2c40 NM_ Cytochrome P450, family 2, subfamily c, polypeptide 40 Drug metabolism Cyp2d22 NM_ Cytochrome P450, family 2, subfamily d, polypeptide 22 Drug metabolism Cyp2e1 NM_ Cytochrome P450, family 2, subfamily e, polypeptide 1 Drug metabolism Cyp2f2 NM_ Cytochrome P450, family 2, subfamily f, polypeptide 2 Drug metabolism
3 Cyp2g1 NM_ Cytochrome P450, family 2, subfamily g, polypeptide 1 Drug metabolism Cyp2j5 NM_ Cytochrome P450, family 2, subfamily j, polypeptide 5 Drug metabolism Cyp2j6 U62295 Cytochrome P450, family 2, subfamily j, polypeptide 6 Drug metabolism Cyp3a11 NM_ Cytochrome P450, family 3, subfamily a, polypeptide 11 Drug metabolism Cyp3a13 NM_ Cytochrome P450, family 3, subfamily a, polypeptide 13 Drug metabolism Cyp3a25 BC Cytochrome P450, family 3, subfamily a, polypeptide 25 Drug metabolism Cyp3a41 NM_ Cytochrome P450, family 3, subfamily a, polypeptide 41 Drug metabolism Acss2 NM_ Acyl-CoA synthetase short-chain family member 2 Fatty acid metabolism Cpt1a BC Carnitine palmitoyltransferase 1a, liver Fatty acid metabolism Cyp4f14 NM_ Cytochrome P450, family 4, subfamily f, polypeptide 14 Fatty acid metabolism Fasn BC Fatty acid synthase Fatty acid metabolism Hmgcl BC hydroxy-3-methylglutaryl-Coenzyme A lyase Fatty acid metabolism Lip1 NM_ Lysosomal acid lipase 1 Fatty acid metabolism Pla2g6 BC Phospholipase A2, group VI Fatty acid metabolism Ptgis NM_ Prostaglandin I2 (prostacyclin) synthase Fatty acid metabolism Scd1 BC Stearoyl-Coenzyme A desaturase 1 Fatty acid metabolism Tbxas1 NM_ Thromboxane A synthase 1, platelet Fatty acid metabolism Alas1 NM_ Aminolevulinic acid synthase 1 Heme metabolism Hmox1 BC Heme oxygenase (decycling) 1 Heme metabolism Hpxn BC Hemopexin Heme metabolism Actb NM_ Actin, beta, cytoplasmic Housekeeping genes Actb NM_ Actin, beta, cytoplasmic Housekeeping genes Gapdh NM_ Glyceraldehyde-3-phosphate dehydrogenase Housekeeping genes Gapdh NM_ Glyceraldehyde-3-phosphate dehydrogenase Housekeeping genes Gapdh NM_ Glyceraldehyde-3-phosphate dehydrogenase Housekeeping genes Ppia NM_ Peptidylprolyl isomerase A Housekeeping genes Rps6ka5 AK Ribosomal protein S6 kinase, polypeptide 5 Housekeeping genes Apoa1 NM_ Apolipopoprotein A-I Lipid transport Apoa2 BC Apolipoprotein A-II Lipid transport Apoa4 BC Apolipoprotein A-IV Lipid transport Apoa5 BC Apolipoprotein A-V Lipid transport Apob XM_ Apolipoprotein B Lipid transport Apoc1 BC Apolipoprotein C-I Lipid transport Apoc2 NM_ Apolipoprotein C-II Lipid transport Apoc3 BC Apolipoprotein C-III Lipid transport Apoe BC Apolipoprotein E Lipid transport Ldlr BC Low density lipoprotein receptor Lipid transport Ldlrap1 NM_ Low density lipoprotein receptor adaptor protein 1 Lipid transport Ar M37890 Androgen receptor Nuclear receptor superfamily Esr1 NM_ Estrogen receptor 1 (alpha) Nuclear receptor superfamily Esr2 NM_ Estrogen receptor 2 (beta) Nuclear receptor superfamily Esrra NM_ Estrogen related receptor, alpha Nuclear receptor superfamily Esrrb NM_ Estrogen related receptor, beta Nuclear receptor superfamily Esrrg NM_ Estrogen-related receptor gamma Nuclear receptor superfamily Hnf4a NM_ Hepatic nuclear factor 4, alpha Nuclear receptor superfamily Hnf4g NM_ Hepatocyte nuclear factor 4, gamma Nuclear receptor superfamily Nr0b1 NM_ Nuclear receptor subfamily 0, group B, member 1 Nuclear receptor superfamily Nr0b2 NM_ Nuclear receptor subfamily 0, group B, member 2 Nuclear receptor superfamily Nr1d2 NM_ Nuclear receptor subfamily 1, group D, member 2 Nuclear receptor superfamily
4 Nr1h2 NM_ liver X receptor beta Nuclear receptor superfamily Nr1h4 NM_ Nuclear receptor subfamily 1, group H, member 4 Nuclear receptor superfamily Nr1i2 NM_ Nuclear receptor subfamily 1, group I, member 2 Nuclear receptor superfamily Nr1i3 NM_ Nuclear receptor subfamily 1, group I, member 3 Nuclear receptor superfamily Nr2c1 NM_ Nuclear receptor subfamily 2, group C, member 1 Nuclear receptor superfamily Nr2c2 NM_ Nuclear receptor subfamily 2, group C, member 2 Nuclear receptor superfamily Nr2e1 NM_ Nuclear receptor subfamily 2, group E, member 1 Nuclear receptor superfamily Nr2e3 NM_ Nuclear receptor subfamily 2, group E, member 3 Nuclear receptor superfamily Nr2f1 NM_ Nuclear receptor subfamily 2, group F, member 1 Nuclear receptor superfamily Nr2f2 NM_ Nuclear receptor subfamily 2, group F, member 2 Nuclear receptor superfamily Nr2f6 NM_ Nuclear receptor subfamily 2, group F, member 6 Nuclear receptor superfamily Nr3c1 NM_ Nuclear receptor subfamily 3, group C, member 1 Nuclear receptor superfamily Nr4a1 NM_ Nuclear receptor subfamily 4, group A, member 1 Nuclear receptor superfamily Nr4a2 NM_ Nuclear receptor subfamily 4, group A, member 2 Nuclear receptor superfamily Nr4a3 NM_ Nuclear receptor subfamily 4, group A, member 3 Nuclear receptor superfamily Nr5a1 NM_ Nuclear receptor subfamily 5, group A, member 1 Nuclear receptor superfamily Nr5a2 NM_ Nuclear receptor subfamily 5, group A, member 2 Nuclear receptor superfamily Nr6a1 AF Nuclear receptor subfamily 6, group A, member 1 Nuclear receptor superfamily Pgr NM_ Progesterone receptor Nuclear receptor superfamily Ppara NM_ Peroxisome proliferator activated receptor alpha Nuclear receptor superfamily Ppard NM_ Peroxisome proliferator activator receptor delta Nuclear receptor superfamily Pparg NM_ Peroxisome proliferator activated receptor gamma Nuclear receptor superfamily Rara NM_ Retinoic acid receptor, alpha Nuclear receptor superfamily Rarb NM_ Retinoic acid receptor, beta Nuclear receptor superfamily Rarg NM_ Retinoic acid receptor, gamma Nuclear receptor superfamily Rora NM_ RAR-related orphan receptor alpha Nuclear receptor superfamily Rorb NM_ RAR-related orphan receptor beta Nuclear receptor superfamily Rorc NM_ RAR-related orphan receptor gamma Nuclear receptor superfamily Rxra NM_ Retinoid X receptor alpha Nuclear receptor superfamily Rxrb NM_ retinoid X receptor beta Nuclear receptor superfamily Rxrg NM_ Retinoid X receptor gamma Nuclear receptor superfamily Thra NM_ thyroid hormone receptor alpha Nuclear receptor superfamily Thrb NM_ thyroid hormone receptor beta Nuclear receptor superfamily Vdr NM_ Vitamin D receptor Nuclear receptor superfamily Acat2 NM_ Acetyl-Coenzyme A acetyltransferase 2 Other Cyp20a1 BC Cytochrome P450, family 20, subfamily A, polypeptide 1 Other Cyp24a1 NM_ Cytochrome P450, family 24, subfamily A, polypeptide 1 Other Cyp26a1 NM_ Cytochrome P450, family 26, subfamily a, polypeptide 1 Other Cyp26b1 NM_ Cytochrome P450, family 26, subfamily b, polypeptide 1 Other Cyp4b1 NM_ Cytochrome P450, family 4, subfamily b, polypeptide 1 Other Gfpt2 BC Glutamine fructose-6-phosphate transaminase 2 Other Icam1 BC Intercellular adhesion molecule Other Nos1 BC Nitric oxide synthase 1, neuronal Other Nos2 BC Nitric oxide synthase 2, inducible, macrophage Other Nos3 BC Nitric oxide synthase 3, endothelial cell Other Orm1 BC Orosomucoid 1 Other Pon1 BC Paraoxonase 1 Other Pon2 NM_ Paraoxonase 2 Other Pten BC Phosphatase and tensin homolog Other
5 Scara3 BC Scavenger receptor class A, member 3 Other Scarb1 NM_ Scavenger receptor class B, member 1 Other Scarb2 BC Scavenger receptor class B, member 2 Other Scp2 BC Sterol carrier protein 2, liver Other Soat1 NM_ Sterol O-acyltransferase 1 Other Soat2 BC Sterol O-acyltransferase 2 Other Star AK Steroidogenic acute regulatory protein Other Uap1 BC UDP-N-acetylglucosamine pyrophosphorylase 1 Other Ucp2 NM_ Uncoupling protein 2 (mitochondrial, proton carrier) Other Vcam1 BC Vascular cell adhesion molecule 1 Other Alb1 BC Albumin 1 Serum proteins Apcs BC Serum amyloid P-component Serum proteins C2 BC Complement component 2 (within H-2S) Serum proteins C3 BC Complement component 3 Serum proteins C4b BC Complement component 4B (Childo blood group) Serum proteins C4bp NM_ Complement component 4 binding protein Serum proteins C9 BC Complement component 9 Serum proteins Crp NM_ C-reactive protein, pentraxin-related Serum proteins Fgb NM_ Fibrinogen, B beta polypeptide Serum proteins Hc M35525 Hemolytic complement Serum proteins Saa1 BC Serum amyloid A 1 Serum proteins Saa2 BC Serum amyloid A 2 Serum proteins Saa3 BC Serum amyloid A 3 Serum proteins Saa4 BC Serum amyloid A 4 Serum proteins Insig1 NM_ Insulin induced gene 1 SREBF signaling pathway Insig2 BC Insulin induced gene 2 SREBF signaling pathway Mbtps1 NM_ Membrane-bound transcription factor peptidase, site 1 SREBF signaling pathway Mbtps2 NM_ Membrane-bound transcription factor peptidase, site 2 SREBF signaling pathway Scap NM_ SREBP cleavage activating protein SREBF signaling pathway Srebf1 NM_ Sterol regulatory element binding factor 1 SREBF signaling pathway Srebf2 NM_ Sterol regulatory element binding factor 2 SREBF signaling pathway Cyp11a1 NM_ Cytochrome P450, family 11, subfamily a, polypeptide 1 Steroid synthesis Cyp11b2 NM_ Cytochrome P450, family 11, subfamily b, polypeptide 2 Steroid synthesis Cyp17a1 NM_ Cytochrome P450, family 17, subfamily a, polypeptide 1 Steroid synthesis Cyp19a1 NM_ Cytochrome P450, family 19, subfamily a, polypeptide 1 Steroid synthesis Cyp21a1 NM_ Cytochrome P450, family 21, subfamily a, polypeptide 1 Steroid synthesis Cebpa BC CCAAT/enhancer binding protein (C/EBP), alpha Transcription regulators Cebpd X61800 CCAAT/enhancer binding protein (C/EBP), delta Transcription regulators Cebpg BC CCAAT/enhancer binding protein (C/EBP), gamma Transcription regulators Cebpz NM_ CCAAT/Enhancer Binding Protein Zeta Transcription regulators Creb1 BC CAMP responsive element binding protein 1 Transcription regulators Crebbp NM_ CREB binding protein Transcription regulators Crem M60285 CAMP responsive element modulator Transcription regulators Crem M60285 CAMP responsive element modulator, transcript tau Transcription regulators Fhl5 AF Four and a half LIM domains 5, also testis CREM activator Transcription regulators Fos NM_ FBJ osteosarcoma oncogene Transcription regulators Foxa1 X74936 Forkhead box A1 Transcription regulators Foxa2 NM_ Forkhead box A2 Transcription regulators Foxo1 NM_ Forkhead box O1 Transcription regulators
6 Hif1a BC Hypoxia inducible factor 1, alpha subunit Transcription regulators Mlxipl NM_ MLX interacting protein-like, old Wbscr14 or Chrebp Transcription regulators Ncoa1 BC Nuclear receptor coactivator 1 Transcription regulators Ncor1 NM_ Nuclear receptor co-repressor 1 Transcription regulators Ncor2 AF Nuclear receptor co-repressor 2 Transcription regulators Nrf1 BC Nuclear respiratory factor 1 Transcription regulators Pcaf BC P300/CBP-associated factor Transcription regulators Ppargc1a NM_ Peroxisome proliferative activated receptor, gamma, coactivator 1 alpha Transcription regulators Ppargc1a NM_ Peroxisome proliferative activated receptor, gamma, coactivator 1 alpha Transcription regulators Ppargc1b NM_ Peroxisome proliferative activated receptor, gamma, coactivator 1 beta Transcription regulators Sirt1 NM_ Sirtuin 1, silent mating type information regulation 2, homolog 1 Transcription regulators Sp1 AF Trans-acting transcription factor 1, Specificity protein 1 Transcription regulators Tbp BC TATA box binding protein Transcription regulators Tcf1 NM_ Transcription factor 1 Transcription regulators Trp53 BC Transformation related protein 53 Transcription regulators Abca1 NM_ ATP-binding cassette, sub-family A (ABC1), member 1 Transporters Abcb11 NM_ ATP-binding cassette, sub-family B (MDR/TAP), member 11 Transporters Abcb1a NM_ ATP-binding cassette, sub-family B (MDR/TAP), member 1A Transporters Abcb1b NM_ ATP-binding cassette, sub-family B (MDR/TAP), member 1B Transporters Abcb4 NM_ ATP-binding cassette, sub-family B (MDR/TAP), member 4 Transporters Abcb7 BC ATP-binding cassette, sub-family B (MDR/TAP), member 7 Transporters Abcc1 NM_ ATP-binding cassette, sub-family C (CFTR/MRP), member 1 Transporters Abcc2 NM_ ATP-binding cassette, sub-family C (CFTR/MRP), member 2 Transporters Abcc3 NM_ ATP-binding cassette, sub-family C (CFTR/MRP), member 3 Transporters Abcg1 AF ATP-binding cassette, sub-family G (WHITE), member 1 Transporters Abcg4 AJ ATP-binding cassette, sub-family G (WHITE), member 4 Transporters Abcg5 NM_ ATP-binding cassette, sub-family G (WHITE), member 5 Transporters Abcg8 NM_ ATP-binding cassette, sub-family G (WHITE), member 8 Transporters Fabp6 NM_ Fatty acid binding protein 6, ileal (gastrotropin) Transporters Slc10a1 BC Solute carrier family 10, member 1 Transporters Slc10a1 BC Solute carrier family 10 (sodium/bile acid cotransporter family), member 1 Transporters Slc10a2 NM_ Solute carrier family 10, member 2 Transporters Slc2a1 BC Solute carrier family 2 (facilitated glucose transporter), member 1 Transporters Slc2a4 BC Solute carrier family 2 (facilitated glucose transporter), member 4 Transporters Slc2a8 BC Solute carrier family 2, (facilitated glucose transporter), member 8 Transporters Slco1a1 AY Solute carrier organic anion transporter family, member 1a1 Transporters Slco1a4 NM_ Solute carrier organic anion transporter family, member 1a4 Transporters Slco1a5 AF Solute carrier organic anion transporter family, member 1a5 Transporters Slco1b2 NM_ Solute carrier organic anion transporter family, member 1b2 Transporters Slco1c1 AY Solute carrier organic anion transporter family, member 1c1 Transporters Slco2b1 BC Solute carrier organic anion transporter family, member 2b1 Transporters
SUPPLEMENTARY FIG. S2. b-galactosidase staining of
 SUPPLEMENTARY FIG. S1. b-galactosidase staining of senescent cells in 500 mg=dl glucose (10 magnification). SUPPLEMENTARY FIG. S3. Toluidine blue staining of chondrogenic-differentiated adipose-tissue-derived
SUPPLEMENTARY FIG. S1. b-galactosidase staining of senescent cells in 500 mg=dl glucose (10 magnification). SUPPLEMENTARY FIG. S3. Toluidine blue staining of chondrogenic-differentiated adipose-tissue-derived
Supplementary Materials Modeling cholesterol metabolism by gene expression profiling in the hippocampus
 Supplementary Materials Modeling cholesterol metabolism by gene expression profiling in the hippocampus Christopher M. Valdez 1, Clyde F. Phelix 1, Mark A. Smith 3, George Perry 1,2, and Fidel Santamaria
Supplementary Materials Modeling cholesterol metabolism by gene expression profiling in the hippocampus Christopher M. Valdez 1, Clyde F. Phelix 1, Mark A. Smith 3, George Perry 1,2, and Fidel Santamaria
Fig. S1. Dose-response effects of acute administration of the β3 adrenoceptor agonists CL316243, BRL37344, ICI215,001, ZD7114, ZD2079 and CGP12177 at
 Fig. S1. Dose-response effects of acute administration of the β3 adrenoceptor agonists CL316243, BRL37344, ICI215,001, ZD7114, ZD2079 and CGP12177 at doses of 0.1, 0.5 and 1 mg/kg on cumulative food intake
Fig. S1. Dose-response effects of acute administration of the β3 adrenoceptor agonists CL316243, BRL37344, ICI215,001, ZD7114, ZD2079 and CGP12177 at doses of 0.1, 0.5 and 1 mg/kg on cumulative food intake
Cholesterol synthesis pathway genes in prostate cancer are transcriptionally downregulated when tissue confounding is minimized
 Rye et al. BMC Cancer (2018) 18:478 https://doi.org/10.1186/s12885-018-4373-y RESEARCH ARTICLE Cholesterol synthesis pathway genes in prostate cancer are transcriptionally downregulated when tissue confounding
Rye et al. BMC Cancer (2018) 18:478 https://doi.org/10.1186/s12885-018-4373-y RESEARCH ARTICLE Cholesterol synthesis pathway genes in prostate cancer are transcriptionally downregulated when tissue confounding
CORRECTION NOTICE. Nat. Chem. Biol. 11, (2015) Sterol metabolism controls T H 17 differentiation by generating endogenous RORγ agonists
 CORRECTION NOTICE Nat. Chem. Biol. 11, 141 147 (2015) Sterol metabolism controls T H 17 differentiation by generating endogenous RORγ agonists Xiao Hu, Yahong Wang, Ling-Yang Hao, Xikui Liu, Chuck A Lesch,
CORRECTION NOTICE Nat. Chem. Biol. 11, 141 147 (2015) Sterol metabolism controls T H 17 differentiation by generating endogenous RORγ agonists Xiao Hu, Yahong Wang, Ling-Yang Hao, Xikui Liu, Chuck A Lesch,
Validated Mouse Quantitative RT-PCR Genes Gene Gene Bank Accession Number Name
 5HT1a NM_008308.4 Serotonin Receptor 1a 5HT1b NM_010482.1 Serotonin Receptor 1b 5HT2a NM_172812.2 Serotonin Receptor 2a ACO-2 NM_080633.2 Aconitase 2 Adora2A NM_009630.02 Adenosine A2a Receptor Aif1 (IbaI)
5HT1a NM_008308.4 Serotonin Receptor 1a 5HT1b NM_010482.1 Serotonin Receptor 1b 5HT2a NM_172812.2 Serotonin Receptor 2a ACO-2 NM_080633.2 Aconitase 2 Adora2A NM_009630.02 Adenosine A2a Receptor Aif1 (IbaI)
Supplemental Table 1 Primer sequences (mouse) used for real-time qrt-pcr studies
 Supplemental Table 1 Primer sequences (mouse) used for real-time qrt-pcr studies Gene symbol Forward primer Reverse primer ACC1 5'-TGAGGAGGACCGCATTTATC 5'-GCATGGAATGGCAGTAAGGT ACLY 5'-GACACCATCTGTGATCTTG
Supplemental Table 1 Primer sequences (mouse) used for real-time qrt-pcr studies Gene symbol Forward primer Reverse primer ACC1 5'-TGAGGAGGACCGCATTTATC 5'-GCATGGAATGGCAGTAAGGT ACLY 5'-GACACCATCTGTGATCTTG
Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes
 Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes The major site of acetoacetate and 3-hydorxybutyrate production is in the liver. 3-hydorxybutyrate is the
Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes The major site of acetoacetate and 3-hydorxybutyrate production is in the liver. 3-hydorxybutyrate is the
Supplementary Table 1. Primer Sequences Used for Quantitative Real-Time PCR
 Supplementary Table 1. Primer Sequences Used for Quantitative Real-Time PCR Gene Forward Primer (5-3 ) Reverse Primer (5-3 ) cadl CTTGGGGGCGCGTCT CTGTTCTTTTGTGCCGTTTCG cyl-coenzyme Dehydrogenase, very
Supplementary Table 1. Primer Sequences Used for Quantitative Real-Time PCR Gene Forward Primer (5-3 ) Reverse Primer (5-3 ) cadl CTTGGGGGCGCGTCT CTGTTCTTTTGTGCCGTTTCG cyl-coenzyme Dehydrogenase, very
Oxidation of Long Chain Fatty Acids
 Oxidation of Long Chain Fatty Acids Dr NC Bird Oxidation of long chain fatty acids is the primary source of energy supply in man and animals. Hibernating animals utilise fat stores to maintain body heat,
Oxidation of Long Chain Fatty Acids Dr NC Bird Oxidation of long chain fatty acids is the primary source of energy supply in man and animals. Hibernating animals utilise fat stores to maintain body heat,
Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction. Marker Sequence (5 3 ) Accession No.
 Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
Supplementary Tables Supplementary Table S1. Primers used for quantitative real-time polymerase chain reaction Marker Sequence (5 3 ) Accession No. Angiopoietin 1, ANGPT1 A CCCTCCGGTGAATATTGGCTGG NM_001146.3
cholesterol structure Cholesterol FAQs Cholesterol promotes the liquid-ordered phase of membranes Friday, October 15, 2010
 cholesterol structure most plasma cholesterol is in the esterified form (not found in cells or membranes) cholesterol functions in all membranes (drives formation of lipid microdomains) cholesterol is
cholesterol structure most plasma cholesterol is in the esterified form (not found in cells or membranes) cholesterol functions in all membranes (drives formation of lipid microdomains) cholesterol is
Lecture #27 Lecturer A. N. Koval
 Lecture #27 Lecturer A. N. Koval Hormones Transduce Signals to Affect Homeostatic Mechanisms Koval A. (C), 2011 2 Lipophilic hormones Classifying hormones into hydrophilic and lipophilic molecules indicates
Lecture #27 Lecturer A. N. Koval Hormones Transduce Signals to Affect Homeostatic Mechanisms Koval A. (C), 2011 2 Lipophilic hormones Classifying hormones into hydrophilic and lipophilic molecules indicates
Mouse Meda-4 : chromosome 5G bp. EST (547bp) _at. 5 -Meda4 inner race (~1.8Kb)
 Supplemental.Figure1 A: Mouse Meda-4 : chromosome 5G3 19898bp I II III III IV V a b c d 5 RACE outer primer 5 RACE inner primer 5 RACE Adaptor ORF:912bp Meda4 cdna 2846bp Meda4 specific 5 outer primer
Supplemental.Figure1 A: Mouse Meda-4 : chromosome 5G3 19898bp I II III III IV V a b c d 5 RACE outer primer 5 RACE inner primer 5 RACE Adaptor ORF:912bp Meda4 cdna 2846bp Meda4 specific 5 outer primer
Gene Expression Multiplexing Menu
 Gene Expression Multiplexing Menu Mouse Rat Human Table of CONTENTS Mouse Inflammatory Plex 2 Rat Stress Plex 3 Growth Regulation Plex 4 Multitox Plex 5 Pharma III Plex 6 Reference Plex 7 Human Transporter
Gene Expression Multiplexing Menu Mouse Rat Human Table of CONTENTS Mouse Inflammatory Plex 2 Rat Stress Plex 3 Growth Regulation Plex 4 Multitox Plex 5 Pharma III Plex 6 Reference Plex 7 Human Transporter
Cholesterol and its transport. Alice Skoumalová
 Cholesterol and its transport Alice Skoumalová 27 carbons Cholesterol - structure Cholesterol importance A stabilizing component of cell membranes A precursor of bile salts A precursor of steroid hormones
Cholesterol and its transport Alice Skoumalová 27 carbons Cholesterol - structure Cholesterol importance A stabilizing component of cell membranes A precursor of bile salts A precursor of steroid hormones
BIOL 158: BIOLOGICAL CHEMISTRY II
 BIOL 158: BIOLOGICAL CHEMISTRY II Lecture 5: Vitamins and Coenzymes Lecturer: Christopher Larbie, PhD Introduction Cofactors bind to the active site and assist in the reaction mechanism Apoenzyme is an
BIOL 158: BIOLOGICAL CHEMISTRY II Lecture 5: Vitamins and Coenzymes Lecturer: Christopher Larbie, PhD Introduction Cofactors bind to the active site and assist in the reaction mechanism Apoenzyme is an
Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array.
 Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array. Gene symbol Gene name TaqMan Assay ID UniGene ID 18S rrna 18S ribosomal RNA Hs99999901_s1 Actb actin, beta Mm00607939_s1
Supplementary Table 1. Genes analysed for expression by angiogenesis gene-array. Gene symbol Gene name TaqMan Assay ID UniGene ID 18S rrna 18S ribosomal RNA Hs99999901_s1 Actb actin, beta Mm00607939_s1
ANSC/NUTR 618 Lipids & Lipid Metabolism
 I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
Nuclear Receptor Profiling
 1. Overview Nuclear Receptors (NRs) act as transcription factors that mediate the effects of certain hormones, drugs and other xenobiotics by modulating the expression of specific genes involved in many
1. Overview Nuclear Receptors (NRs) act as transcription factors that mediate the effects of certain hormones, drugs and other xenobiotics by modulating the expression of specific genes involved in many
Metabolism of cardiac muscle. Dr. Mamoun Ahram Cardiovascular system, 2013
 Metabolism of cardiac muscle Dr. Mamoun Ahram Cardiovascular system, 2013 References This lecture Mark s Basic Medical Biochemistry, 4 th ed., p. 890-891 Hand-out Why is this topic important? Heart failure
Metabolism of cardiac muscle Dr. Mamoun Ahram Cardiovascular system, 2013 References This lecture Mark s Basic Medical Biochemistry, 4 th ed., p. 890-891 Hand-out Why is this topic important? Heart failure
 Acetyl CoA HMG CoA Mevalonate (C6) Dimethylallyl Pyrophosphate isopentenyl Pyrophosphate (C5) Geranyl Pyrophosphate (C10) FarnesylPyrophosphate (C15) Squalene (C30) Lanosterol (C30) 7 Dehydrocholesterol
Acetyl CoA HMG CoA Mevalonate (C6) Dimethylallyl Pyrophosphate isopentenyl Pyrophosphate (C5) Geranyl Pyrophosphate (C10) FarnesylPyrophosphate (C15) Squalene (C30) Lanosterol (C30) 7 Dehydrocholesterol
Receptors Functions and Signal Transduction- L4- L5
 Receptors Functions and Signal Transduction- L4- L5 Faisal I. Mohammed, MD, PhD University of Jordan 1 PKC Phosphorylates many substrates, can activate kinase pathway, gene regulation PLC- signaling pathway
Receptors Functions and Signal Transduction- L4- L5 Faisal I. Mohammed, MD, PhD University of Jordan 1 PKC Phosphorylates many substrates, can activate kinase pathway, gene regulation PLC- signaling pathway
Cholesterol Metabolism
 Cholesterol Metabolism Lippincott s Illustrated Review Chapter 18 Steroid Nucleus 1 2 Cholesterol was isolated from gall bladder stones in 1774 3 Sources and Elimination of Cholesterol Synthesis: 1000
Cholesterol Metabolism Lippincott s Illustrated Review Chapter 18 Steroid Nucleus 1 2 Cholesterol was isolated from gall bladder stones in 1774 3 Sources and Elimination of Cholesterol Synthesis: 1000
CHE 242 Exam 3 Practice Questions
 CHE 242 Exam 3 Practice Questions Glucose metabolism 1. Below is depicted glucose catabolism. Indicate on the pathways the following: A) which reaction(s) of glycolysis are irreversible B) where energy
CHE 242 Exam 3 Practice Questions Glucose metabolism 1. Below is depicted glucose catabolism. Indicate on the pathways the following: A) which reaction(s) of glycolysis are irreversible B) where energy
Table S1 Differentially expressed genes showing > 2 fold changes and p <0.01 for 0.01 mm NS-398, 0.1mM ibuprofen and COX-2 RNAi.
 Table S1 Differentially expressed genes showing > 2 fold changes and p
Table S1 Differentially expressed genes showing > 2 fold changes and p
INTRODUCTORY BIOCHEMISTRY. BI 28 Second Midterm Examination April 3, 2007
 INTRODUCTORY BIOCHEMISTRY BI 28 Second Midterm Examination April 3, 2007 Name SIS # Make sure that your name or SIS # is on every page. This is the only way we have of matching you with your exam after
INTRODUCTORY BIOCHEMISTRY BI 28 Second Midterm Examination April 3, 2007 Name SIS # Make sure that your name or SIS # is on every page. This is the only way we have of matching you with your exam after
Fatty acids synthesis
 Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
INTERACTION DRUG BODY
 INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11464 Supplemental Figure S1. The expression of Vegfb is increased in obese and diabetic mice as compared to lean mice. a-b, Body weight and postprandial blood
SUPPLEMENTARY INFORMATION doi:10.1038/nature11464 Supplemental Figure S1. The expression of Vegfb is increased in obese and diabetic mice as compared to lean mice. a-b, Body weight and postprandial blood
Supplementary Figure 1. Mitochondrial function in skeletal muscle and plasma parameters of STZ mice A D-F
 Supplementary Figure 1. Mitochondrial function in skeletal muscle and plasma parameters of STZ mice A Expression of representative subunits of the mitochondrial respiratory chain was analyzed by real time
Supplementary Figure 1. Mitochondrial function in skeletal muscle and plasma parameters of STZ mice A Expression of representative subunits of the mitochondrial respiratory chain was analyzed by real time
Hmgcoar AGCTTGCCCGAATTGTATGTG TCTGTTGTAACCATGTGACTTC. Cyp7α GGGATTGCTGTGGTAGTGAGC GGTATGGAATCAACCCGTTGTC
 Supplement Table I: primers for Real Time RT-PCR Gene Foward Reverse Hmgcoar AGCTTGCCCGAATTGTATGTG TCTGTTGTAACCATGTGACTTC Cyp7α GGGATTGCTGTGGTAGTGAGC GGTATGGAATCAACCCGTTGTC Cyp27a1 GTGGTCTTATTGGGTACTTGC
Supplement Table I: primers for Real Time RT-PCR Gene Foward Reverse Hmgcoar AGCTTGCCCGAATTGTATGTG TCTGTTGTAACCATGTGACTTC Cyp7α GGGATTGCTGTGGTAGTGAGC GGTATGGAATCAACCCGTTGTC Cyp27a1 GTGGTCTTATTGGGTACTTGC
Mechanistic Toxicology
 SECOND EDITION Mechanistic Toxicology The Molecular Basis of How Chemicals Disrupt Biological Targets URS A. BOELSTERLI CRC Press Tavlor & France Croup CRC Press is an imp^t o* :H Taylor H Francn C'r,,jpi
SECOND EDITION Mechanistic Toxicology The Molecular Basis of How Chemicals Disrupt Biological Targets URS A. BOELSTERLI CRC Press Tavlor & France Croup CRC Press is an imp^t o* :H Taylor H Francn C'r,,jpi
Biologic Oxidation BIOMEDICAL IMPORTAN
 Biologic Oxidation BIOMEDICAL IMPORTAN Chemically, oxidation is defined as the removal of electrons and reduction as the gain of electrons. Thus, oxidation is always accompanied by reduction of an electron
Biologic Oxidation BIOMEDICAL IMPORTAN Chemically, oxidation is defined as the removal of electrons and reduction as the gain of electrons. Thus, oxidation is always accompanied by reduction of an electron
Lecture 16. Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III
 Lecture 16 Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III The Powertrain of Human Metabolism (verview) CARBHYDRATES PRTEINS
Lecture 16 Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III The Powertrain of Human Metabolism (verview) CARBHYDRATES PRTEINS
CARBOHYDRATE METABOLISM 1
 CARBOHYDRATE METABOLISM 1 web 2017 József Mandl Strategy of metabolism 1 Strategy of metabolism to extract energy ( hydrogen ) from the environment to store the energy excess to store hydrogen CH 3 O 2
CARBOHYDRATE METABOLISM 1 web 2017 József Mandl Strategy of metabolism 1 Strategy of metabolism to extract energy ( hydrogen ) from the environment to store the energy excess to store hydrogen CH 3 O 2
Syllabus for BASIC METABOLIC PRINCIPLES
 Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
Respiration. Organisms can be classified based on how they obtain energy: Autotrophs
 Respiration rganisms can be classified based on how they obtain energy: Autotrophs Able to produce their own organic molecules through photosynthesis Heterotrophs Live on organic compounds produced by
Respiration rganisms can be classified based on how they obtain energy: Autotrophs Able to produce their own organic molecules through photosynthesis Heterotrophs Live on organic compounds produced by
Comparison of catabolic and anabolic pathways
 Comparison of catabolic and anabolic pathways Three stages of catabolism Glucose Synthesis of compounds e.g. lactose glycolipids Glucose-6-P Pentosephosphate Pathway Glycolysis Glycogenesis Acetyl-CoA
Comparison of catabolic and anabolic pathways Three stages of catabolism Glucose Synthesis of compounds e.g. lactose glycolipids Glucose-6-P Pentosephosphate Pathway Glycolysis Glycogenesis Acetyl-CoA
Introduction to Detoxification Enzymes. Evolutionary Response to Chemicals in the Environment
 Evolutionary Response to Chemicals in the Environment Introduction to Detoxification Enzymes 1. Introduction to Detoxification enzymes (focusing on cytochrome P450s) 2. Evolutionary response to toxins
Evolutionary Response to Chemicals in the Environment Introduction to Detoxification Enzymes 1. Introduction to Detoxification enzymes (focusing on cytochrome P450s) 2. Evolutionary response to toxins
Chapter 10. Introduction to Nutrition and Metabolism, 3 rd edition David A Bender Taylor & Francis Ltd, London 2002
 Chapter 10 Introduction to Nutrition and Metabolism, 3 rd edition David A Bender Taylor & Francis Ltd, London 2002 Chapter 10: Integration and Control of Metabolism Press the space bar or click the mouse
Chapter 10 Introduction to Nutrition and Metabolism, 3 rd edition David A Bender Taylor & Francis Ltd, London 2002 Chapter 10: Integration and Control of Metabolism Press the space bar or click the mouse
Points 1. Following is the overall reaction catalyzed by the Calvin-Benson cycle:
 BCH 4054 February 22, 2002 HOUR TEST 2 NAME_ Points 1. Following is the overall reaction catalyzed by the Calvin-Benson cycle: CO 2 + 3ATP + 2NADPH 1/3 glyceraldehyde-3-p + 3ADP + 2NADP + Give the structures
BCH 4054 February 22, 2002 HOUR TEST 2 NAME_ Points 1. Following is the overall reaction catalyzed by the Calvin-Benson cycle: CO 2 + 3ATP + 2NADPH 1/3 glyceraldehyde-3-p + 3ADP + 2NADP + Give the structures
Medical Biochemistry Examination II November 9, 2002 Kresge Auditorium Please follow these directions:
 Medical Biochemistry Examination II November 9, 2002 Kresge Auditorium Please follow these directions: 1. Do not begin the exam until all students have received a copy of the exam. You will be instructed
Medical Biochemistry Examination II November 9, 2002 Kresge Auditorium Please follow these directions: 1. Do not begin the exam until all students have received a copy of the exam. You will be instructed
Lipid Metabolism. Catabolism Overview
 Lipid Metabolism Pratt & Cornely, Chapter 17 Catabolism Overview Lipids as a fuel source from diet Beta oxidation Mechanism ATP production Ketone bodies as fuel 1 High energy More reduced Little water
Lipid Metabolism Pratt & Cornely, Chapter 17 Catabolism Overview Lipids as a fuel source from diet Beta oxidation Mechanism ATP production Ketone bodies as fuel 1 High energy More reduced Little water
Under aerobic conditions, pyruvate enters the mitochondria where it is converted into acetyl CoA.
 Under aerobic conditions, pyruvate enters the mitochondria where it is converted into acetyl CoA. Acetyl CoA is the fuel for the citric acid cycle, which processes the two carbon acetyl unit to two molecules
Under aerobic conditions, pyruvate enters the mitochondria where it is converted into acetyl CoA. Acetyl CoA is the fuel for the citric acid cycle, which processes the two carbon acetyl unit to two molecules
March 19 th Batool Aqel
 March 19 th - 2013 6 Batool Aqel Hormones That Bind to Nuclear Receptor Proteins Hormones bind to their receptors.whether the receptor is found in the nucleus or the cytoplasm, at the end they are translocated
March 19 th - 2013 6 Batool Aqel Hormones That Bind to Nuclear Receptor Proteins Hormones bind to their receptors.whether the receptor is found in the nucleus or the cytoplasm, at the end they are translocated
Roles of Lipids. principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular
 Roles of Lipids principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular = Oxidation of fatty acids Central energy-yielding pathway in animals. O
Roles of Lipids principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular = Oxidation of fatty acids Central energy-yielding pathway in animals. O
Integration of Metabolism
 Integration of Metabolism Metabolism is a continuous process. Thousands of reactions occur simultaneously in order to maintain homeostasis. It ensures a supply of fuel, to tissues at all times, in fed
Integration of Metabolism Metabolism is a continuous process. Thousands of reactions occur simultaneously in order to maintain homeostasis. It ensures a supply of fuel, to tissues at all times, in fed
Lecture: 26 OXIDATION OF FATTY ACIDS
 Lecture: 26 OXIDATION OF FATTY ACIDS Fatty acids obtained by hydrolysis of fats undergo different oxidative pathways designated as alpha ( ), beta ( ) and omega ( ) pathways. -oxidation -Oxidation of fatty
Lecture: 26 OXIDATION OF FATTY ACIDS Fatty acids obtained by hydrolysis of fats undergo different oxidative pathways designated as alpha ( ), beta ( ) and omega ( ) pathways. -oxidation -Oxidation of fatty
Summary of fatty acid synthesis
 Lipid Metabolism, part 2 1 Summary of fatty acid synthesis 8 acetyl CoA + 14 NADPH + 14 H+ + 7 ATP palmitic acid (16:0) + 8 CoA + 14 NADP + + 7 ADP + 7 Pi + 7 H20 1. The major suppliers of NADPH for fatty
Lipid Metabolism, part 2 1 Summary of fatty acid synthesis 8 acetyl CoA + 14 NADPH + 14 H+ + 7 ATP palmitic acid (16:0) + 8 CoA + 14 NADP + + 7 ADP + 7 Pi + 7 H20 1. The major suppliers of NADPH for fatty
TNFSF13B tumor necrosis factor (ligand) superfamily, member 13b NF-kB pathway cluster, Enrichment Score: 3.57
 Appendix 2. Highly represented clusters of genes in the differential expression of data. Immune Cluster, Enrichment Score: 5.17 GO:0048584 positive regulation of response to stimulus GO:0050778 positive
Appendix 2. Highly represented clusters of genes in the differential expression of data. Immune Cluster, Enrichment Score: 5.17 GO:0048584 positive regulation of response to stimulus GO:0050778 positive
MITOCHONDRIA LECTURES OVERVIEW
 1 MITOCHONDRIA LECTURES OVERVIEW A. MITOCHONDRIA LECTURES OVERVIEW Mitochondrial Structure The arrangement of membranes: distinct inner and outer membranes, The location of ATPase, DNA and ribosomes The
1 MITOCHONDRIA LECTURES OVERVIEW A. MITOCHONDRIA LECTURES OVERVIEW Mitochondrial Structure The arrangement of membranes: distinct inner and outer membranes, The location of ATPase, DNA and ribosomes The
acid were from Fisher Scientific (Loughborough, UK). Methanol and acetonitrile (HPLC grade) were from VWR (Lutterworth, Leicestershire, UK).
 1 SUPPLEMENTAL MATERIAL 2 SUPPLEMENTARY METHODS 3 4 5 6 7 Materials Unless stated otherwise, all solvents were from Rathburn Chemical Ltd. (Walkerburn, UK) and other chemicals from Sigma-Aldrich (Dorset,
1 SUPPLEMENTAL MATERIAL 2 SUPPLEMENTARY METHODS 3 4 5 6 7 Materials Unless stated otherwise, all solvents were from Rathburn Chemical Ltd. (Walkerburn, UK) and other chemicals from Sigma-Aldrich (Dorset,
2. What is molecular oxygen directly converted into? a. Carbon Dioxide b. Water c. Glucose d. None of the Above
 Biochem 1 Mock Exam 3 Chapter 11: 1. What is glucose completely oxidized into? a. Carbon Dioxide and Water b. Carbon Dioxide and Oxygen c. Oxygen and Water d. Water and Glycogen 2. What is molecular oxygen
Biochem 1 Mock Exam 3 Chapter 11: 1. What is glucose completely oxidized into? a. Carbon Dioxide and Water b. Carbon Dioxide and Oxygen c. Oxygen and Water d. Water and Glycogen 2. What is molecular oxygen
Lipid metabolism in familial hypercholesterolemia
 Lipid metabolism in familial hypercholesterolemia Khalid Al-Rasadi, BSc, MD, FRCPC Head of Biochemistry Department, SQU Head of Lipid and LDL-Apheresis Unit, SQUH President of Oman society of Lipid & Atherosclerosis
Lipid metabolism in familial hypercholesterolemia Khalid Al-Rasadi, BSc, MD, FRCPC Head of Biochemistry Department, SQU Head of Lipid and LDL-Apheresis Unit, SQUH President of Oman society of Lipid & Atherosclerosis
Mitochondria and ATP Synthesis
 Mitochondria and ATP Synthesis Mitochondria and ATP Synthesis 1. Mitochondria are sites of ATP synthesis in cells. 2. ATP is used to do work; i.e. ATP is an energy source. 3. ATP hydrolysis releases energy
Mitochondria and ATP Synthesis Mitochondria and ATP Synthesis 1. Mitochondria are sites of ATP synthesis in cells. 2. ATP is used to do work; i.e. ATP is an energy source. 3. ATP hydrolysis releases energy
number Done by Corrected by Doctor Faisal Al-Khatibe
 number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
Energy storage in cells
 Energy storage in cells Josef Fontana EC - 58 Overview of the lecture Introduction to the storage substances of human body Overview of storage compounds in the body Glycogen metabolism Structure of glycogen
Energy storage in cells Josef Fontana EC - 58 Overview of the lecture Introduction to the storage substances of human body Overview of storage compounds in the body Glycogen metabolism Structure of glycogen
Biology 638 Biochemistry II Exam-2
 Biology 638 Biochemistry II Exam-2 Biol 638, Exam-2 (Code-1) 1. Assume that 16 glucose molecules enter into a liver cell and are attached to a liner glycogen one by one. Later, this glycogen is broken-down
Biology 638 Biochemistry II Exam-2 Biol 638, Exam-2 (Code-1) 1. Assume that 16 glucose molecules enter into a liver cell and are attached to a liner glycogen one by one. Later, this glycogen is broken-down
Glycolysis Part 2. BCH 340 lecture 4
 Glycolysis Part 2 BCH 340 lecture 4 Regulation of Glycolysis There are three steps in glycolysis that have enzymes which regulate the flux of glycolysis These enzymes catalyzes irreversible reactions of
Glycolysis Part 2 BCH 340 lecture 4 Regulation of Glycolysis There are three steps in glycolysis that have enzymes which regulate the flux of glycolysis These enzymes catalyzes irreversible reactions of
LIPID METABOLISM. Sri Widia A Jusman Department of Biochemistry & Molecular Biology FMUI
 LIPID METABOLISM Sri Widia A Jusman Department of Biochemistry & Molecular Biology FMUI Lipid metabolism is concerned mainly with fatty acids cholesterol Source of fatty acids from dietary fat de novo
LIPID METABOLISM Sri Widia A Jusman Department of Biochemistry & Molecular Biology FMUI Lipid metabolism is concerned mainly with fatty acids cholesterol Source of fatty acids from dietary fat de novo
Gene Polymorphisms and Carbohydrate Diets. James M. Ntambi Ph.D
 Gene Polymorphisms and Carbohydrate Diets James M. Ntambi Ph.D Fatty Acids that Flux into Tissue Lipids are from Dietary Sources or are Made De novo from Glucose or Fructose Glucose Fructose Acetyl-CoA
Gene Polymorphisms and Carbohydrate Diets James M. Ntambi Ph.D Fatty Acids that Flux into Tissue Lipids are from Dietary Sources or are Made De novo from Glucose or Fructose Glucose Fructose Acetyl-CoA
NBCE Mock Board Questions Biochemistry
 1. Fluid mosaic describes. A. Tertiary structure of proteins B. Ribosomal subunits C. DNA structure D. Plasma membrane structure NBCE Mock Board Questions Biochemistry 2. Where in the cell does beta oxidation
1. Fluid mosaic describes. A. Tertiary structure of proteins B. Ribosomal subunits C. DNA structure D. Plasma membrane structure NBCE Mock Board Questions Biochemistry 2. Where in the cell does beta oxidation
Chapter 16 - Lipid Metabolism
 Chapter 16 - Lipid Metabolism Fatty acids have four major physiologic roles in the cell: Building blocks of phospholipids and glycolipids Added onto proteins to create lipoproteins, which targets them
Chapter 16 - Lipid Metabolism Fatty acids have four major physiologic roles in the cell: Building blocks of phospholipids and glycolipids Added onto proteins to create lipoproteins, which targets them
Biomarkers for Hypothesis Testing
 Biomarkers for Hypothesis Testing Definition for Drug Development: Biomarker = Any Measure of a Drug Action Proximal to a Clinical Effect Biochemical (PET, MRS & CSF* for CNS drugs) Physiological EEG,
Biomarkers for Hypothesis Testing Definition for Drug Development: Biomarker = Any Measure of a Drug Action Proximal to a Clinical Effect Biochemical (PET, MRS & CSF* for CNS drugs) Physiological EEG,
Developing a clinically feasible personalized medicine approach to pediatric
 Developing a clinically feasible personalized medicine approach to pediatric septic shock Hector R. Wong, Natalie Z. Cvijanovich, Nick Anas, Geoffrey L. Allen, Neal J. Thomas, Michael T. Bigham, Scott
Developing a clinically feasible personalized medicine approach to pediatric septic shock Hector R. Wong, Natalie Z. Cvijanovich, Nick Anas, Geoffrey L. Allen, Neal J. Thomas, Michael T. Bigham, Scott
Biological processes. Mitochondrion Metabolic process Catalytic activity Oxidoreductase
 Full name Glyceraldehyde3 phosphate dehydrogenase Succinatesemialdehyde Glutamate dehydrogenase 1, Alcohol dehydrogenase [NADP+] 2',3'cyclicnucleotide 3' phosphodiesterase Dihydropyrimidinaserelated 2
Full name Glyceraldehyde3 phosphate dehydrogenase Succinatesemialdehyde Glutamate dehydrogenase 1, Alcohol dehydrogenase [NADP+] 2',3'cyclicnucleotide 3' phosphodiesterase Dihydropyrimidinaserelated 2
ANSC/NUTR 618 LIPIDS & LIPID METABOLISM. Triacylglycerol and Fatty Acid Metabolism
 ANSC/NUTR 618 LIPIDS & LIPID METABOLISM II. Triacylglycerol synthesis A. Overall pathway Glycerol-3-phosphate + 3 Fatty acyl-coa à Triacylglycerol + 3 CoASH B. Enzymes 1. Acyl-CoA synthase 2. Glycerol-phosphate
ANSC/NUTR 618 LIPIDS & LIPID METABOLISM II. Triacylglycerol synthesis A. Overall pathway Glycerol-3-phosphate + 3 Fatty acyl-coa à Triacylglycerol + 3 CoASH B. Enzymes 1. Acyl-CoA synthase 2. Glycerol-phosphate
Cholesterol metabolism. Function Biosynthesis Transport in the organism Hypercholesterolemia
 Cholesterol metabolism Function Biosynthesis Transport in the organism Hypercholesterolemia - component of all cell membranes - precursor of bile acids steroid hormones vitamin D Cholesterol Sources: dietary
Cholesterol metabolism Function Biosynthesis Transport in the organism Hypercholesterolemia - component of all cell membranes - precursor of bile acids steroid hormones vitamin D Cholesterol Sources: dietary
Diosgenin, antagonism of LXRs 36 DNA microarray, sesame seed lignan regulation of liver fatty acid metabolism gene expression 12 18, 22, 23
 Subject Index ACC, see Acyl-CoA carboxylase 1 -Acetoxychavicol acetate, inducible nitric oxide synthase suppression in cancer chemoprevention 199, 200 Acetylene carotenoids, anti-inflammatory activity
Subject Index ACC, see Acyl-CoA carboxylase 1 -Acetoxychavicol acetate, inducible nitric oxide synthase suppression in cancer chemoprevention 199, 200 Acetylene carotenoids, anti-inflammatory activity
Glycogen Metabolism Dr. Mohammad Saadeh
 Glycogen Metabolism Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry II Philadelphia University Faculty of pharmacy I. overview Glucose is energy source for Brain.
Glycogen Metabolism Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry II Philadelphia University Faculty of pharmacy I. overview Glucose is energy source for Brain.
respiration mitochondria mitochondria metabolic pathways reproduction can fuse or split DRP1 interacts with ER tubules chapter DRP1 ER tubule
 mitochondria respiration chapter 3-4 shape highly variable can fuse or split structure outer membrane inner membrane cristae intermembrane space mitochondrial matrix free ribosomes respiratory enzymes
mitochondria respiration chapter 3-4 shape highly variable can fuse or split structure outer membrane inner membrane cristae intermembrane space mitochondrial matrix free ribosomes respiratory enzymes
Integration of Metabolism 1. made by: Noor M. ALnairat. Sheet No. 18
 Integration of Metabolism 1 made by: Noor M. ALnairat Sheet No. 18 Data :24/11/2016 SLIDE 2: Metabolism Consist of Highly Interconnected Pathways The basic strategy of catabolic metabolism is to form ATP,
Integration of Metabolism 1 made by: Noor M. ALnairat Sheet No. 18 Data :24/11/2016 SLIDE 2: Metabolism Consist of Highly Interconnected Pathways The basic strategy of catabolic metabolism is to form ATP,
Role of metabolism in Drug-Induced Liver Injury (DILI) Drug Metab Rev. 2007;39(1):
 Role of metabolism in Drug-Induced Liver Injury (DILI) Drug Metab Rev. 2007;39(1):159-234 Drug Metab Rev. 2007;39(1):159-234 Drug Metab Rev. 2007;39(1):159-234 A schematic representation of the most relevant
Role of metabolism in Drug-Induced Liver Injury (DILI) Drug Metab Rev. 2007;39(1):159-234 Drug Metab Rev. 2007;39(1):159-234 Drug Metab Rev. 2007;39(1):159-234 A schematic representation of the most relevant
number Done by Corrected by Doctor Nayef Karadsheh
 number 11 Done by حسام أبو عوض Corrected by Moayyad Al-Shafei Doctor Nayef Karadsheh 1 P a g e General Regulatory Aspects in Metabolism: We can divide all pathways in metabolism to catabolicand anabolic.
number 11 Done by حسام أبو عوض Corrected by Moayyad Al-Shafei Doctor Nayef Karadsheh 1 P a g e General Regulatory Aspects in Metabolism: We can divide all pathways in metabolism to catabolicand anabolic.
Both pathways start with Glucose as a substrate but they differ in the product.
 Glycosis:may occur either with the presence or absence of -Glucose-.So with oxygen we have Aerobic glycolysis-, without the participation of oxygen Anaerobic glycolysis-(it occur in certain places) where
Glycosis:may occur either with the presence or absence of -Glucose-.So with oxygen we have Aerobic glycolysis-, without the participation of oxygen Anaerobic glycolysis-(it occur in certain places) where
Integration Of Metabolism
 Integration Of Metabolism Metabolism Consist of Highly Interconnected Pathways The basic strategy of catabolic metabolism is to form ATP, NADPH, and building blocks for biosyntheses. 1. ATP is the universal
Integration Of Metabolism Metabolism Consist of Highly Interconnected Pathways The basic strategy of catabolic metabolism is to form ATP, NADPH, and building blocks for biosyntheses. 1. ATP is the universal
Ahmad Ulnar. Faisal Nimri ... Dr.Faisal
 24 Ahmad Ulnar Faisal Nimri... Dr.Faisal Fatty Acid Synthesis - Occurs mainly in the Liver (to store excess carbohydrates as triacylglycerols(fat)) and in lactating mammary glands (for the production of
24 Ahmad Ulnar Faisal Nimri... Dr.Faisal Fatty Acid Synthesis - Occurs mainly in the Liver (to store excess carbohydrates as triacylglycerols(fat)) and in lactating mammary glands (for the production of
Fatty Acid and Triacylglycerol Metabolism 1
 Fatty Acid and Triacylglycerol Metabolism 1 Mobilization of stored fats and oxidation of fatty acids Lippincott s Chapter 16 What is the first lecture about What is triacylglycerol Fatty acids structure
Fatty Acid and Triacylglycerol Metabolism 1 Mobilization of stored fats and oxidation of fatty acids Lippincott s Chapter 16 What is the first lecture about What is triacylglycerol Fatty acids structure
Coenzymes, vitamins and trace elements 209. Petr Tůma Eva Samcová
 Coenzymes, vitamins and trace elements 209 Petr Tůma Eva Samcová History and nomenclature of enzymes 1810, Gay-Lussac made an experiment with yeats alter saccharide to ethanol and CO 2 Fermentation From
Coenzymes, vitamins and trace elements 209 Petr Tůma Eva Samcová History and nomenclature of enzymes 1810, Gay-Lussac made an experiment with yeats alter saccharide to ethanol and CO 2 Fermentation From
Published on Second Faculty of Medicine, Charles University (http://www.lf2.cuni.cz )
 Published on Second Faculty of Medicine, Charles University (http://www.lf2.cuni.cz ) Biochemistry Submitted by Marie Havlová on 8. February 2012-0:00 Syllabus of Biochemistry Mechanisms of enzyme catalysis.
Published on Second Faculty of Medicine, Charles University (http://www.lf2.cuni.cz ) Biochemistry Submitted by Marie Havlová on 8. February 2012-0:00 Syllabus of Biochemistry Mechanisms of enzyme catalysis.
Rawan almujaibel. Ayman Musleh. Dr. Nayef
 12 Rawan almujaibel Ayman Musleh Ayman Musleh Dr. Nayef In the previous lecture we talked about digestion and absorption of carbohydrates. In this lecture we will be talking about glycolysis. Glycolysis
12 Rawan almujaibel Ayman Musleh Ayman Musleh Dr. Nayef In the previous lecture we talked about digestion and absorption of carbohydrates. In this lecture we will be talking about glycolysis. Glycolysis
Medical Biochemistry Examination II November 12, 1999 Kresge Auditorium Please follow these directions:
 Medical Biochemistry Examination II November 12, 1999 Kresge Auditorium Please follow these directions: 1. Do not begin the exam until all students have received a copy of the exam. You will be instructed
Medical Biochemistry Examination II November 12, 1999 Kresge Auditorium Please follow these directions: 1. Do not begin the exam until all students have received a copy of the exam. You will be instructed
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function
 Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
Table S9A: List of taurine regulated genes in Bp K96243 Chr 1 (up regulated >=2 fold) Cluster no GENE ID Start Stop Strand Function 1 BPSL0024 26223 26621 + LrgA family BPSL0025 26690 27412 + hypothetical
Fatty Acid and Triacylglycerol Metabolism 1
 Fatty Acid and Triacylglycerol Metabolism 1 Mobilization of stored fats and oxidation of fatty acids Lippincott s Chapter 16 What is the first lecture about What is triacylglycerol Fatty acids structure
Fatty Acid and Triacylglycerol Metabolism 1 Mobilization of stored fats and oxidation of fatty acids Lippincott s Chapter 16 What is the first lecture about What is triacylglycerol Fatty acids structure
Examination III PHRM 836 Biochemistry for Pharmaceutical Sciences II December 19, 2014
 Examination III PHRM 836 Biochemistry for Pharmaceutical Sciences II December 19, 2014 Name: Instructions PHRM 836 Exam III - 1 1. Check your exam to make certain that it has 8 pages including this cover
Examination III PHRM 836 Biochemistry for Pharmaceutical Sciences II December 19, 2014 Name: Instructions PHRM 836 Exam III - 1 1. Check your exam to make certain that it has 8 pages including this cover
MBB 694:407, 115:511 Name Third Exam Niederman, Deis. Please use BLOCK CAPITAL letters like this --- A, B, C, D, E. Not lowercase!
 MBB 694:407, 115:511 Name Third Exam Niederman, Deis Wed. Nov. 22, 2006 Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth two points.
MBB 694:407, 115:511 Name Third Exam Niederman, Deis Wed. Nov. 22, 2006 Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth two points.
Carbohydrate Metabolism I
 Carbohydrate Metabolism I Outline Glycolysis Stages of glycolysis Regulation of Glycolysis Carbohydrate Metabolism Overview Enzyme Classification Dehydrogenase - oxidizes substrate using cofactors as
Carbohydrate Metabolism I Outline Glycolysis Stages of glycolysis Regulation of Glycolysis Carbohydrate Metabolism Overview Enzyme Classification Dehydrogenase - oxidizes substrate using cofactors as
Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section B and ONE question from Section C.
 UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2014-15 FUNDAMENTALS OF CELL BIOLOGY AND BIOCHEMISTRY BIO-4004B Time allowed: 2 hours Answer ALL questions in Section
UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2014-15 FUNDAMENTALS OF CELL BIOLOGY AND BIOCHEMISTRY BIO-4004B Time allowed: 2 hours Answer ALL questions in Section
BCMB 3100 Fall 2013 Exam III
 BCMB 3100 Fall 2013 Exam III 1. (10 pts.) (a.) Briefly describe the purpose of the glycerol dehydrogenase phosphate shuttle. (b.) How many ATPs can be made when electrons enter the electron transport chain
BCMB 3100 Fall 2013 Exam III 1. (10 pts.) (a.) Briefly describe the purpose of the glycerol dehydrogenase phosphate shuttle. (b.) How many ATPs can be made when electrons enter the electron transport chain
Please use BLOCK CAPITAL letters like this --- A, B, C, D, E. Not lowercase! 1. E 10. D 19. E 2. D 11. A 20. A 3. D 12. D 21. A 4. D 13. A 22.
 Molecular Biology and Biochemistry 694:407/115:511 Third Exam Deis Niederman Wednesday, November 22, 2006 Name Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of
Molecular Biology and Biochemistry 694:407/115:511 Third Exam Deis Niederman Wednesday, November 22, 2006 Name Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of
Biosynthesis of Fatty Acids
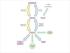 Biosynthesis of Fatty Acids Fatty acid biosynthesis takes place in the cytosol rather than the mitochondria and requires a different activation mechanism and different enzymes and coenzymes than fatty
Biosynthesis of Fatty Acids Fatty acid biosynthesis takes place in the cytosol rather than the mitochondria and requires a different activation mechanism and different enzymes and coenzymes than fatty
GENERAL CHARACTERISTICS OF THE ENDOCRINE SYSTEM FIGURE 17.1
 GENERAL CHARACTERISTICS OF THE ENDOCRINE SYSTEM FIGURE 17.1 1. The endocrine system consists of glands that secrete chemical signals, called hormones, into the blood. In addition, other organs and cells
GENERAL CHARACTERISTICS OF THE ENDOCRINE SYSTEM FIGURE 17.1 1. The endocrine system consists of glands that secrete chemical signals, called hormones, into the blood. In addition, other organs and cells
Chapter 26 Biochemistry 5th edition. phospholipids. Sphingolipids. Cholesterol. db=books&itool=toolbar
 http://www.ncbi.nlm.nih.gov/sites/entrez? db=books&itool=toolbar 1 The surface of a soap bubble is a bilayer formed by detergent molecules 2 Chapter 26 Biochemistry 5th edition phospholipids Sphingolipids
http://www.ncbi.nlm.nih.gov/sites/entrez? db=books&itool=toolbar 1 The surface of a soap bubble is a bilayer formed by detergent molecules 2 Chapter 26 Biochemistry 5th edition phospholipids Sphingolipids
Fatty acid breakdown
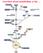 Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Photosynthesis in chloroplasts. Cellular respiration in mitochondria ATP. ATP powers most cellular work
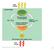 Light energy ECOSYSTEM CO + H O Photosynthesis in chloroplasts Cellular respiration in mitochondria Organic molecules + O powers most cellular work Heat energy 1 becomes oxidized (loses electron) becomes
Light energy ECOSYSTEM CO + H O Photosynthesis in chloroplasts Cellular respiration in mitochondria Organic molecules + O powers most cellular work Heat energy 1 becomes oxidized (loses electron) becomes
Lipid metabolism. Degradation and biosynthesis of fatty acids Ketone bodies
 Lipid metabolism Degradation and biosynthesis of fatty acids Ketone bodies Fatty acids (FA) primary fuel molecules in the fat category main use is for long-term energy storage high level of energy storage:
Lipid metabolism Degradation and biosynthesis of fatty acids Ketone bodies Fatty acids (FA) primary fuel molecules in the fat category main use is for long-term energy storage high level of energy storage:
Activation of sterol regulatory element-binding proteins in mice
 Activation of sterol regulatory element-binding proteins in mice exposed to perfluorooctanoic acid for 28 days Shengmin Yan, Jianshe Wang, and Jiayin Dai* Materials and methods Chemicals and antibodies
Activation of sterol regulatory element-binding proteins in mice exposed to perfluorooctanoic acid for 28 days Shengmin Yan, Jianshe Wang, and Jiayin Dai* Materials and methods Chemicals and antibodies
