Calcium and Cyclic AMP signals differentially regulate CREB function via a Rap1-ERK pathway
|
|
|
- Paulina Cole
- 5 years ago
- Views:
Transcription
1 JBC Papers in Press. Published on August 18, 2000 as Manuscript M Calcium and Cyclic AMP signals differentially regulate CREB function via a Rap1-ERK pathway Savraj S. Grewal, Daniel M. Fass, Hong Yao, Cindy L. Ellig, Richard H. Goodman, Philip J.S. Stork Vollum Institute and Department of Cell and Developmental Biology, Oregon Health Sciences University, 3181 SW Sam Jackson Park Road, Portland, OR To whom correspondence should be addressed. Philip Stork, Vollum Institute, Oregon Health Sciences University, L474, SW Sam Jackson Park Road, Portland, OR Tele; , Fax: stork@ohsu.edu. Running title: Rap1-ERK regulation of CREB Copyright 2000 by The American Society for Biochemistry and Molecular Biology, Inc.
2 Other authors: Savraj S. Grewal: Vollum Institute, Oregon Health Sciences University, L474, SW Sam Jackson Park Road, Portland, OR Tele; , Fax: Daniel M. Fass: Vollum Institute, Oregon Health Sciences University, L474, SW Sam Jackson Park Road, Portland, OR Tele; , Fax: Hong Yao: Vollum Institute, Oregon Health Sciences University, L474, SW Sam Jackson Park Road, Portland, OR Tele; , Fax: Cindy L. Ellig: Vollum Institute, Oregon Health Sciences University, L474, SW Sam Jackson Park Road, Portland, OR Tele; , Fax: Richard H. Goodman: Vollum Institute, Oregon Health Sciences University, L474, SW Sam Jackson Park Road, Portland, OR Tele; , Fax:
3 SUMMARY Two major intracellular signals that regulate neuronal function are calcium and cyclic AMP (camp). In many cases, the actions of these two second messengers involve long-term changes in gene expression. One well studied target of both calcium and camp signaling is the transcription factor CREB. Multiple signaling pathways have been shown to contribute to the regulation of CREB-dependent transcription, including both protein kinase A (PKA)- and mitogen activated p rotein/ extracellular signal- r egulated k inase (MAP kinase or ERK)- dependent kinase cascades. We have previously described a mechanism by which camp and calcium influx may stimulate ERKs in neuronal cells. This pathway involves the PKAdependent activation of the Ras-related small G-protein, Rap1, and subsequent stimulation of the neuronal Raf isoform, B-Raf. In this study, we examined the contribution of the Rap1-ERK pathway to the control of gene transcription by calcium influx and camp. Using the PC12 cell model system we found that calcium influx and camp both stimulated CREB-dependent transcription via a Rap1-ERK pathway, but this regulation occurred through distinct mechanisms. Calcium-mediated phosphorylation of CREB through the PKA-Rap1-ERK pathway. In contrast, camp phosphorylated CREB via PKA directly but required a Rap1-ERK pathway to activate a component downstream of CREB phosphorylation and CBP recruitment. These data suggest that the Rap1/B-Raf signaling pathway may have an important role in the regulation of CREB-dependent gene expression. INTRODUCTION Two major intracellular signals that regulate neuronal function are calcium and cyclic adenosine monophosphate (camp). For example, both activation of G-protein coupled receptors linked to adenylate cyclase and stimulation of calcium influx via either receptoror voltage-operated calcium channels can exert rapid actions on neurotransmission and 3
4 neuronal excitability (1,2). However, in many cases, the actions of these two second messengers involve slower, long-term changes in gene expression (3,4). These effects are often mediated by stimulation of intracellular signal transduction pathways that regulate the activity of multiple transcription factors. One well studied target of both calcium and camp is the transcription factor camp-responsive element binding protein (CREB) (5). Regulation of this protein may be important in mediating changes in synaptic plasticity and neuronal survival (6-8). CREB binds as a dimer to a conserved cyclic-amp response element (CRE) found in the promoters of numerous eukaryotic genes (4). Phosphorylation of serine-133 is a critical event in CREB activation (9), which induces an increase in CREB transactivation potential by allowing the recruitment and binding to co-activators such as CREB-binding protein (CBP) (10,11). Early studies identified PKA as a major physiological kinase responsible for Ser-133 phosphorylation (12). Consequently, one prevailing model of CREB regulation is that, following increases in intracellular camp levels and activation of PKA, the catalytic subunit of PKA translocates into the nucleus and phosphorylates CREB, leading to the stimulation of gene transcription (13). Subsequent studies have identified additional CREB kinases, including members of the calcium/calmodulin-dependent kinase (CaMK) family (14-18) and the extracellular signal-regulated kinase (ERK)-stimulated RSK and MSK kinases (19-21). These kinase families have been reported to mediate the actions of both calcium and growth factors on CREB phosphorylation. However, it is becoming increasingly clear that signaling events in addition to CREB phosphorylation are required for full CREB-dependent transcription (5). For example, processes such as recruitment and regulation of co-activators and coupling to the basal transcription machinery may present additional targets for kinase action. Indeed, the activity of the transcriptional co-activator CBP has been reported to be regulated by a variety of kinases including PKA, CaMKIV and ERK, possibly by direct phosphorylation (11,22-28). As 4
5 such, it is likely that the multiple actions of different signaling pathways may ultimately contribute to the stimulation of full CREB-dependent transcription by both calcium and camp. We have previously described a mechanism by which camp and calcium influx may stimulate ERKs in neuronal cells (29,30). This pathway involves the PKA-dependent activation of the Ras-related small G-protein, Rap1, and subsequent stimulation of the neuronal Raf isoform, B-Raf. Activation of this pathway results in robust stimulation of ERKs and ERK-dependent gene expression (29-31). Both Rap1 and, in particular, B-Raf are highly expressed within the nervous system. Hence, stimulation of a Rap1-ERK pathway may contribute to neuronal camp and calcium signaling (30,32,33). Moreover, this pathway may allow for cross talk between PKA and ERK signaling systems. The downstream consequences of stimulation of the Rap1-ERK pathway by calcium and camp are not fully understood. However, given the ability of both PKA and ERK to regulate CREB activity, it is possible some of the observed actions of PKA occur via cross talk through the Rap1-ERK pathway. In this study, we examined the contribution of the Rap1-ERK pathway to the control of gene transcription by calcium influx and camp. Using the PC12 cell model system, we found that both stimulation of calcium influx and elevation of intracellular camp levels stimulated CREB-dependent transcription via the PKA-dependent Rap1-ERK pathway. Interestingly, this appears to occur through distinct mechanisms. Calcium used a PKA- Rap1-ERK pathway to mediate phosphorylation of CREB. In marked contrast, camp phosphorylated CREB via PKA directly but required a Rap1-ERK pathway to mediate an event that was downstream from Ser-133 phosphorylation and CBP recruitment to achieve full transcription. These data suggest a revised model for camp regulation of CREB in 5
6 which the co-ordinate action of both PKA-dependent phosphorylation of CREB and Rap1- ERK stimulation of a downstream target are required for full transcription. EXPERIMENTAL PROCEDURES Materials - PC12-GR5 cells were kindly provided by R. Nishi, Oregon Health Sciences University, Portland, Oregon. Forskolin, 8-CPT-cAMP and PD98059 were purchased from Cal Biochem (Riverside, CA). Phosphorylation-specific and phosphorylation stateindependent rat polyclonal CREB antibodies were purchased from New England Biolabs (Beverly, MA). Polyclonal antibody to B-Raf(C19) was purchased from Santa Cruz Biotechnology Inc. Cell culture - PC12 cells were maintained in DMEM (Dulbecco-Modified Eagle Medium) plus 10% horse serum and 5% fetal calf serum on 100 mm plates to 50-60% confluence at 37o C in 5% CO 2 prior to harvesting. NIH3T3 and Hek293 cells were maintained in DMEM plus 10 % fetal calf serum. For luciferase assays and western blotting, cells were maintained in low serum (0.2% fetal calf serum) media for 16 hours at 37 o C in 5% CO 2 prior to treatment with various reagents. All inhibitors were added 20 minutes before treatment. Plasmids and Transfections - Fifty percent-confluent PC12 cells were co-transfected with the indicated cdnas using a Superfect transfection kit (Qiagen) according to the manufacturer instructions. Hek293 and NIH3T3 cells were transfected using Lipofectamine (Gibco BRL) according to the manufacturer instructions. The vector pcdna3 (Invitrogen Corp.) was added to each set of transfections as appropriate to ensure that each plate received the same amount of DNA. Following transfection, cells were allowed to recover for twenty-fours before being maintained in low serum media and 6
7 treated, as described. The following plasmids were used with the amounts used per 60mm dish indicated in brackets: PKI (1 or 3µg), GAL4-CREB (2µg), catalytic subunit of PKA (cpka; 4µg), Rap1GAP1 (2 or 4µg), 5XGal-E1b-TATA-luciferase (gal-luciferase, 2µg), 5XCRE-luciferase (1µg), Fos-luciferase, (1µg), CREB-DIEDML (5 µg), GAL4-CREB- DIEDML (2 µg). Western blotting - Cells were treated for appropriate times as indicated and then harvested in boiling SDS sample buffer. Cell lysates were boiled for additional five minutes and proteins resolved by SDS-PAGE. Western blotting with phospho-specific and phosphorylation state-independent antibodies was performed as per manufacturer instructions. Immunofluorescence assays - PC12 cells were treated as described and after the indicated times fixed in paraformaldeyde. Following permeablization in methanol and blocking in 5% normal goat serum (NGS), cells were incubated in primary antibody (1:2500 in 5%NGS) overnight at 4 C. Cells were incubated in secondary antibody (1:2500 Alexa 546 conjugated anti-rabbit ) for one hour at room temperature. Cells were visualized using a Leica DMRB microscope. The intensity of CREB immunofluorescence was measured using NIH Image software. Briefly, signal intensity was quantitated as nuclear pixel density above background, and was expressed as fold increase over unstimulated cells. Luciferase reporter gene assays - Cells were treated with the appropriate stimuli for four to five hours. Cells were then lysed and equal protein amounts of lysate per condition were assayed for luciferase as described. All experiments were performed with at least three independently treated plates per condition. RESULTS 7
8 Calcium and camp stimulate CRE-dependent transcription via a PKA-dependent Rap1- ERK pathway. We initially began examining the regulation of CREB function in PC12 cells using a CRE reporter system consisting of five reiterated CREs controlling the expression of luciferase (34). Following transfection of the reporter into cells, we found that KCl depolarizationmediated opening of L-type calcium channels induced a marked stimulation of CREdependent transcription (Fig.1). Similarly, elevation of intracellular camp levels by treatment with the adenylate cyclase activator, forskolin, also stimulated transcription (Fig.1). In both cases, the increase in CRE-dependent transcription was reversed by either co-transfection of the selective PKA inhibitor, PKI (Fig.1A), or pre-treatment with the selective MEK inhibitor, PD98059 (Fig.1B). These data suggest that both calcium influx and camp require PKA and ERK signaling events to regulate CREB function. We have previously demonstrated that, in PC12 cells, calcium and camp activate a PKA-dependent Rap1/B-Raf signaling pathway to stimulate ERKs (29,30). Therefore, the requirement for both PKA and ERK in the regulation of CRE-dependent transcription may reflect activation of this pathway. We investigated this using a specific inhibitor of Rap1, Rap1GAP1 (35). This protein stimulates the conversion of Rap1 from its GTP-bound, active, to GDPbound, inactive state. Previous work has demonstrated that overexpression of Rap1GAP1 proteins inhibit Rap signaling in PC12 cells (36). In addition, using an Elk-gal reporter system to monitor ERK activation, we found that the activation of Elk-dependent transcription by both KCl and forskolin in PC12 cells was inhibited by co-transfection of Rap1GAP1 (Fig. 2A). Using the CRE-luciferase reporter system, we also found that the increase in transcription mediated by both KCl and forskolin was inhibited by Rap1GAP1 (Fig. 2B). These data suggest that both calcium influx and camp regulate CREBdependent transcription via activation of a PKA-dependent Rap1-ERK signaling cascade. 8
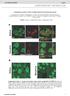
9 Calcium influx stimulates CREB Ser-133 phosphorylation via a PKA-Rap1-ERK pathway. CREB transcriptional activity is stimulated by the phosphorylation of Ser133. Numerous studies have demonstrated that this can occur through either PKA- or ERK-dependent pathways. Phosphorylation of Ser-133 may therefore be a target for Rap1-ERK signaling in the regulation of CRE activity. We examined the phosphorylation of CREB by calcium and camp in PC12 cells using an antibody that recognizes phospho-ser133 CREB (Fig. 3). Both KCl and forskolin induced a rapid and sustained phosphorylation of CREB as assayed by western blot. In both cases this was reversed by pretreatment with the PKA inhibitor, H89 (Fig. 3). In contrast, the MEK inhibitor PD98059 blocked KCl-, but not forskolin-induced phosphorylation (Fig. 3). We also examined CREB phosphorylation by immunofluoresence. Both KCl and forskolin induced an increase in phospho-creb immunoreactivity (Fig. 4). These effects were reversed by transfection of the selective PKA inhibitor, PKI (Fig. 4B). Furthermore, as we observed with the western blots, PD98059 only blocked the increase CREB phosphorylation induced by KCl (Fig. 4C). These data suggest that the requirement for Rap1-ERK action in CREB activation by calcium probably reflects an ERK-dependent phosphorylation of CREB. Activation of ERK signaling was not necessary for camp-mediated phosphorylation. In contrast, camp appeared to induce CREB phosphorylation via a direct PKA-dependent mechanism, consistent with the well described model of transcriptional regulation by camp. Activation of a Rap1-ERK pathway is a conserved component of camp-dependent regulation of CREB transcription. The previous data indicate that camp uses a Rap1-ERK pathway to control CREdependent transcription, but that this regulation occurs at a site distinct from CREB Ser-133 phosphorylation. It is possible that this requirement for ERK activity is a function of the reporter system used. We therefore examined transcriptional regulation in a different promoter context, using a GAL4-CREB/ gal-luciferase reporter system. PC12 cells were 9
10 co-transfected with a 5Xgal-luciferase reporter plasmid and a GAL4-CREB protein in which the CREB bzip domain was replaced with an N-terminal GAL4 DNA binding and dimerization domain. Previous studies have demonstrated that this reporter system provides a measure of CREB-dependent transcription (14,37,38). Treatment of cells with the camp analogue, 8-CPT-cAMP or forskolin, or transfection of the PKA catalytic subunit (cpka) all stimulated GAL4-CREB activity in PC12 cells (Fig. 5). As observed with the CRE-luciferase reporter, these actions were reversed by the MEK inhibitor, PD98059 (Fig. 5A, B). Inhibition of Rap1 activity by co-transfection of Rap1GAP1 also prevented forskolin-mediated stimulation of GAL4-CREB activity (Fig. 5C). We next examined whether the requirement for a Rap1-ERK pathway in camp signaling was specific to PC12 cells. We have previously demonstrated that the ability of camp to activate a Rap1-dependent signaling pathway to ERKs is dependent on the expression of B- Raf (30). Thus, we examined the regulation of CREB in two cell lines, Hek293 and NIH3T3, which differ in their expression of B-Raf. Hek293 cells express high levels of B-Raf. Elevation of camp by either direct stimulation of adenylate cyclase or activation of β-adrenergic receptors activates ERKs in these cells (data not shown). Using the CREluciferase reporter assay, we found that both forskolin and the β-adrenergic receptor agonist, isoproterenol, markedly stimulated CRE-dependent transcription (Fig. 6A). Moreover, these effects were reversed by PD98059 (Fig. 6A). In contrast, the phosphorylation of CREB Ser-133 induced by forskolin and isoproteronol was not blocked by PD98059 (Fig. 6B). Thus, as in PC12 cells, camp stimulation of CRE-dependent transcription requires an ERK signaling component at a site distinct from CREB phosphorylation. We also investigated CRE-dependent transcription in the NIH3T3 cell line. PKA is unable to activate ERKs in these cells due to the absence of the Raf isoform, B-Raf (30). Transfection of B-Raf into these cells allows PKA to stimulate ERKs (30). In the absence of B-Raf, both forskolin and cpka were able to activate CRE-dependent 10
11 transcription (Fig. 7). However, transfection of B-Raf potentiated the stimulatory actions of both forskolin and cpka while having no effects on the basal levels of transcription (Fig. 7A, B). These actions were independent of CREB phosphorylation: examination by western blot revealed that B-Raf had no effect on Ser-133 phosphorylation mediated by cpka (Fig. 7C). Together, these data suggest that the ability of PKA to activate a Rap1- ERK pathway is important for full CREB-dependent transcription by camp. The Rap1-ERK pathway acts downstream from CREB Ser-133 phosphorylation and CBP recruitment to mediate full stimulation of CREB-dependent transcription by camp. The data described above suggest that a Rap1-ERK pathway contributes to the regulation of camp-mediated gene expression at a site downstream from CREB Ser-133 phosphorylation. One potential target for Rap1-ERK signaling is the control of co-activator function. For example, studies have demonstrated that CBP, a co-activator for CREB, can be regulated by a variety of kinase cascades including both PKA- and ERK-dependent signaling pathways. We examined this by using a recently described mutant of CREB, CREB-DIEDML. This mutant contains a substitution of six non-conserved amino acids (DIEDML) from the transcription factor sterol-responsive element binding protein (SREBP) that replace Ser-133 and the surrounding five amino acids (RRPSYR) in CREB(39). The resulting molecule allows constitutive, phosphorylation-independent binding to the co-activator CBP (39). Transfection of this CREB-DIEDML into F9 teratocarcinoma cells was previously shown to lead to constitutive stimulation of CREdependent transcription (39). We found that in PC12 cells, CREB-DIEDML also stimulated both CRE- and Fos-dependent transcription in an extracellular-signal-independent manner (Fig. 8A). Fusion of the activation domain of CREB-DIEDML to the DNA binding domain of GAL4 allows it to induce constitutive stimulation of a gal-luciferase reporter in a signalindependent manner in PC12 cells ((39) and Fig. 8B). Forskolin treatment potentiated GAL4-CREB-DIEDML activity further (Fig. 8B). Interestingly, as observed with the 11
12 GAL4-CREB protein, this stimulation by forskolin was reversed by PD98059 (Fig. 8B). Calcium influx was unable to stimulate GAL4-CREB activity (data not shown). These findings are consistent with previous studies using GAL reporter systems, in which GAL4- CREB fusion proteins lacking the CREB bzip domain are unresponsive to calcium influx (14,38). As a result, we were unable to evaluate the actions of calcium influx on CREB- DIEDML activity. DISCUSSION In this study, we have demonstrated that activation of a Rap1-ERK signaling pathway in PC12 cells participates in the regulation of CREB-dependent transcription by both calcium influx and camp. However, this pathway seems to be used in distinct ways. Calciumdependent phosphorylation of CREB requires Rap1-ERK signaling. In contrast, the camp-dependent stimulation of the Rap1-ERK pathway appears to contribute to transcription at step downstream of CBP recruitment (Fig. 9). Previous studies have demonstrated that a variety of kinase signaling pathways can differentially contribute to calcium influx-mediated phosphorylation of CREB in PC12 cells and neurons (5). The exact mechanism used by calcium may depend on both cell-type and stimulus-specific factors. For example, in hippocampal cells, CaMKIV is reported to be a major calcium-regulated CREB kinase (17). In other cells, where CaMKIV is not expressed, ERK-dependent pathways may predominate. A recent study reported that calcium-mediated phosphorylation of CREB in PC12 cells occurred via the ERK dependent activation of the CREB kinase, RSK-2 (21). PKA was also required for regulating the nuclear translocation of ERK/RSK2, a prerequisite for CREB phosphorylation (21). Our findings are consistent with a similar ERK-dependent mechanism of CREB phosphorylation following calcium influx. We also identify a 12
13 requirement for PKA in this event. However, we suggest that this may reflect the ability of PKA to stimulate ERKs. While the Ras-ERK pathway is an important target for calcium signaling in many cell types, our previous studies have indicated that PKA-dependent activation of a Rap1/B-Raf signaling pathway is the predominant mechanism by which calcium influx stimulates ERKs in the PC12 model system used in this report (29). A similar pathway may also be used in hippocampal cells (29,40). Ultimately, PKA signaling may play multiple roles in the regulation of ERK dependent pathways that lead to CREB phosphorylation following calcium influx. A major finding of this study is that camp requires a Rap1-ERK signaling pathway to stimulate CREB-dependent transcription. Unlike the situation with calcium, this requirement for ERK activation appears not to involve its ability to mediate CREB Ser-133 phosphorylation. We find that stimulation of intracellular camp levels can lead to CREB phosphorylation via PKA, presumably through a direct action on Ser-133. Such an action has classically been thought to account for the ability of camp to stimulate CRE-dependent transcription (4). Indeed, phosphorylation of CREB is often considered to be a measure of transcriptional activation by camp. Our data suggest that stimulation of a Rap1-ERK pathway may represent an additional required component of CREB regulation by camp. Importantly, this requirement for Rap1-ERK signaling was seen in multiple cell-types and in different promoter contexts. In addition, hormonal stimulation of β-adrenergic receptors coupled to adenylate cyclases used a similar mechanism to stimulate transcription. We therefore propose a revised model of camp regulation of CREB in which both PKAmediated Ser-133 phosphorylation and stimulation of a PKA-Rap1-ERK pathway are required for full CREB-dependent transcription. Rap1 is highly expressed in neurons. Moreover, B-Raf expression is mainly restricted to neuronal and neuroendocrine cells. Therefore, this pathway is a potential regulator of neuronal CREB activity. Interestingly, previous reports have demonstrated that, in the context of calcium regulation of c-fos 13
14 transcription in PC12 cells, a PKA-dependent signaling event distinct from CREB phosphorylation contributes to c-fos gene activation (41,42). It is possible that these findings reflect a similar activation of Rap1-ERK signaling as described here. An important consideration is the mechanism by which the Rap1-ERK pathway controls transcription downstream of CREB phosphorylation. One potential target is the transcriptional co-activator, CBP. CBP is a phosphoprotein that can potentially be regulated by a variety of kinase signaling pathways. However, it has been difficult to discriminate between mechanisms responsible for CREB phosphorylation versus CBP activation. One common approach has been to use gene-reporter assay systems in which GAL4-CBP fusion proteins target CBP to synthetic Gal promoters. Using these methods, studies have demonstrated that CBP-dependent transcriptional activity can be stimulated via a variety of signaling kinases including CaMKIV, PKA and ERKs (11,22-28). However, it is unclear how well the GAL4-CBP fusion recapitulates the regulation of native CBP. In this report, we have examined the actions of camp using a CREB mutant, CREB- DIEDML that contains a six amino acid substitution within its kinase inducible domain (39). Importantly, this mutant can bind CBP and stimulate CRE-dependent transcription in a phosphorylation-independent manner (39). We show that stimulation of camp/pka signaling can further increase the transcriptional activity of this mutant. This increase appears to be indirect, involving PKA-dependent activation of the Rap1-ERK pathway. Given that CREB-DIEDML can constitutively bind to CBP and recruit it to the promoter, it is possible these effects involve modification of CBP activity itself. For example, some studies have demonstrated that the transcriptional activity of a GAL4-CBP fusion protein can be increased by ERKs (27,28,43). Furthermore, PKA has been shown to augment GAL4-CBP function even when the sole consensus PKA phosphorylation site in CBP is mutated (11,44). These data are therefore also consistent with an indirect action of PKA, 14
15 possibly via Rap1-ERK signaling. Intriguingly, in one study, the stimulatory actions of PKA on CBP were only observed in PC12 cells, a cell line that expresses B-Raf and in which camp can therefore stimulate a Rap1-ERK pathway (45). In contrast, no effect of PKA was seen in COS-7 cells that express little or no B-Raf (45). Some reports have suggested that stimulation of Ras-ERK signaling cannot increase CBP activity (22,24). In particular, Ras-dependent activation of RSK2 may actually antagonize CBP-dependent transcription (46). These differences in the reported actions of ERK signaling may reflect contrasting roles for Ras- versus Rap1-dependent signaling. Thus, one model is that a Rap1-ERK pathway can increase CBP activity, whereas Ras signaling, possibly via RSK2, exerts an inhibitory effect. Given that a major action of PKA is to inhibit Ras signaling (47,48), camp modulation of CBP activity may reflect the net action of PKA-dependent inhibition of Ras and PKA-mediated activation of a Rap1-ERK pathway. CBP is also a co-activator for a variety of other transcription factors, e.g. Smads (49,50), STATs (51), nuclear hormone receptors (52). Therefore, regulation of its function through the Rap1-ERK signaling pathway may also provide a mechanism by which camp can control transcription independently of CREB. An interesting finding of this study is that one signaling pathway (i.e. Rap1-ERK) can have distinct consequences on regulation of transcription depending on the type of signal (i.e. calcium influx versus camp). Moreover, stimulation of PKA activity is central to regulation of CREB by both calcium and camp, but through different downstream effects. These particular effects may reflect stimulation of discrete spatial or subcellular pools of PKA (53). For example, since Rap1 is membrane bound, PKA signaling at the membrane is probably required for activation of Rap1/B-Raf signaling and thus can regulate both calcium-induced phosphorylation of CREB and camp-mediated actions on CBP function through the Rap1-ERK pathway. In contrast, phosphorylation of CREB by camp 15
16 probably involves a direct nuclear action of PKA. A similar scenario has been proposed for calcium signaling where distinct transcriptional responses are induced by cytoplasmic versus nuclear pools of calcium (54,55). These differential actions of PKA may be important in neurons where the intensity and spatial regulation of PKA activation may determine pathways used to stimulate CREB. For example discrete stimulation in dendrites may not be sufficient to lead to robust elevation of nuclear PKA, but could stimulate a Rap1-ERK pathway to phosphorylate CREB (56). In contrast, robust elevation of camp levels in the cell body could lead to direct nuclear PKA-mediated phosphorylation of CREB. In conclusion, we find that stimulation of a PKA-dependent Rap1-ERK signaling pathway is important for the regulation of CREB function by both calcium and camp signals. Moreover, this mechanism of regulation may be important in controlling the neuronal action of CREB. Because CREB is a major regulator of processes such as synaptic plasticity and neuronal survival (5,57), we propose that activation of Rap1-ERK signaling may be important in mediating these neuronal events. 16
17 FIGURE LEGENDS FIG 1. Depolarization and forskolin stimulate CRE reporter via a PKA- and ERK-dependent mechanism. (A) PC12 cells were transfected with a CRE-luciferase reporter and either 0, 1, or 3 µg PKI as indicated. Cells were then treated with either KCl (60 mm) or forskolin (10µM) for four hours as indicated and then harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells. Each bar represents the mean +/- SEM of at least three independent treatments. (B) PC12 cells were transfected with a CRE-luciferase reporter. Cells were pretreated with 0, 20, 50, or 100 µm PD98059 as indicated. Cells were stimulated with either KCl (60 mm) or forskolin (10µM) for four hours as indicated and then harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells. Each bar represents the mean +/- SEM of three independent treatments. FIG 2. Depolarization and forskolin stimulate both Elk-1- and CREdependent transcription via a Rap1-ERK pathway. (A) PC12 cells were transfected with a gal-elk1 plasmid and gal-luciferase reporter, and 0, 2 or 4 µg Rap1GAP1 as indicated. Cells were then treated with either KCl (60 mm) or forskolin (10µM) for four hours as indicated and then harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells. Each bar represents the mean +/- SEM of at least three independent treatments. (B) PC12 cells were transfected with a CREluciferase reporter and either 0, 2, or 4 µg Rap1GAP1 as indicated. Cells were stimulated with either KCl (60 mm) or forskolin (10µM) for four hours as indicated and then harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells. Each bar represents the mean +/- SEM of three independent treatments. 17
18 FIG 3. Depolarization, but not forskolin, stimulates CREB Ser-133 phosphorylation via an ERK-dependent pathway. PC12 cells were pretreated with either H89 (10µM) or PD98059 (50µM) as indicated. They were then stimulated with either KCl (60 mm) or forskolin (10µM) for 15min to 4 hours. At each timepoint, cell lysates were collected in SDS sample buffer, resolved by SDS-PAGE and analyzed for phospho-creb by western blotting (upper panels). The blots were then stripped and reprobed for CREB (lower panels). FIG 4. Depolarization- and forskolin dependent stimulation of phospho- CREB immunofluoresence. (A) PC12 cells were transfected with either empty vector or PKI, and GFP as a marker, as indicated. Following stimulation with either KCl (60mM) or forskolin (10µM), cells were processed for phospho-creb immunoreactivity as described. The arrowheads identify representative transfected cells. Left panels, Hoechst staining of nuclei; middle panel, GFP transfected cells; right panel, phospho-creb immunofluorescence. (B) The intensity of phospho-creb immunofluorescence in transfected cells was measured and expressed as fold increase over basal, unstimulated cells. Data are expressed as mean +/- SEM (n>50 cells per condition). (C) PC12 cells were pretreated with PD98059 (50µM) as indicated. Following stimulation with either KCl (60mM) or forskolin (10µM), cells were processed for phospho-creb immunoreactivity as described. The intensity of phospho-creb immunofluorescence in transfected cells was measured and expressed as fold increase over basal, unstimulated cells. Data are expressed as mean +/- SEM (n>50 cells per condition). FIG 5. Cyclic AMP stimulates CREB-dependent transcription via a Rap1- ERK pathway. (A) PC12 cells were transfected with a GAL4-CREB plasmid and a galluciferase reporter. Cells were pretreated with PD98059 (50µM) as indicated and then stimulated with either the camp analog, 8-CPT-cAMP (175µM) or forskolin (10µM) for 18
19 four hours, before being harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells. Each bar represents the mean +/- SEM of three independent treatments. (B) PC12 cells were transfected with a GAL4-CREB plasmid, a gal-luciferase reporter, and either empty vector or cpka as indicated. Cells were incubated with PD98059 (50µM) for four hours as indicated and then harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells. Each bar represents the mean +/- SEM of three independent treatments. (C) PC12 cells were transfected with a GAL4- CREB plasmid, a gal-luciferase reporter, and either empty vector or Rap1GAP1 as indicated. Cells were left untreated or stimulated with forskolin (10µM) for four hours as indicated and then harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells. Each bar represents the mean +/- SEM of three independent treatments. FIG 6. Cyclic AMP regulates CREB activity via Rap1-ERK signaling in Hek293 cells. (A) Hek293 cells were transfected with a 5XCRE-luciferase reporter. Cells were pretreated with PD98059 (50µM), as indicated, and then stimulated with either forskolin (10µM) or the β-adrenergic receptor agonist, isoproterenol (10µM), for four hours, before being harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells. Each bar represents the mean +/- SEM of three independent treatments. (B) Hek293 cells were pretreated with PD98059 (50µM), as indicated, and then stimulated with either forskolin (10µM) or the β-adrenergic receptor agonist, isoproterenol (10µM) for 15min. Cell lysates were collected in SDS sample buffer, resolved by SDS-PAGE and either phospho-creb (upper panel) or CREB (lower panel) western blotting performed. FIG 7. Cyclic AMP regulates CREB activity via Rap1/B-Raf signaling in NIH3T3 cells. (A) NIH3T3 cells were co-transfected with a 5XCRE-luciferase reporter 19
20 and either empty vector or B-Raf as indicated. Cells were then left untreated or stimulated with forskolin (10µM) for four hours as indicated, before being harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells. Each bar represents the mean +/- SEM of three independent treatments. (B) NIH3T3 cells were cotransfected with a 5XCRE-luciferase reporter and combinations of empty vector, cpka and B-Raf as indicated. Twenty-four hours later, cells were harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells. Each bar represents the mean +/- SEM of three independent treatments. (C) NIH3T3 cells were co-transfected with combinations of empty vector, cpka and B-Raf as indicated. Twenty-four hours later, cell lysates were collected in SDS sample buffer, resolved by SDS-PAGE and B-Raf (upper panel), phospho-creb (middle panel) or CREB (lower panel) western blotting performed. FIG 8. Cyclic AMP regulation of CREB-DIEDML. (A) PC12 cells were cotransfected with either vector or CREB-DIEDML as indicated and either a CRE-luciferase (left bar graph) or Fos-luciferase (58) (right bar graph) reporter. Twenty-four hours later, cells were harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells. Each bar represents the mean +/- SEM of three independent treatments. (B) PC12 cells were co-transfected with either GAL4-CREB or GAL4-CREB-DIEDML as indicated and a gal-luciferase reporter. Cells were pretreated with PD98059 (50µM) as indicated and then stimulated with forskolin (10µM) for four hours, before being harvested for luciferase assay. Data are expressed as fold increase over basal, unstimulated cells transfected with the gal-luciferase reporter only. Each bar represents the mean +/- SEM of three independent treatments. FIG 9. A putative model of CREB regulation by calcium influx and camp in PC12 cells. Calcium influx via L-type channels (left) stimulates CREB-dependent transcription via activation of a PKA-dependent Rap1-ERK pathway. The requirement for 20
21 this pathway probably reflects a role for ERK signaling in calcium-induced CREB phosphorylation. Elevation of intracellular camp levels (right) also stimulates CREB function through the PKA-dependent Rap1-ERK pathway. In this case, camp phosphorylates CREB directly via PKA and utilizes the Rap1-ERK pathway to stimulate a component downstream of CBP recruitment. This latter step may reflect an action of ERK signaling on CBP function. A similar step may also be utilized by calcium signaling. 21
22 REFERENCES 1. O'Connor, V., Augustine, G. J., and Betz, H. (1994) Cell 76, Wickman, K., and Clapham, D. E. (1995) Physiol. Rev. 75, Ghosh, A., and Greenberg, M. E. (1995) Science 268, Montminy, M. (1997) Annu. Rev. Biochem. 66, Shaywitz, A. J., and Greenberg, M. E. (1999) Annu. Rev. Biochem. 68, Martin, K. C., and Kandel, E. R. (1996) Neuron 17, Riccio, A., Ahn, S., Davenport, C. M., Blendy, J. A., and Ginty, D. D. (1999) Science 286, Bonni, A., Brunet, A., West, A. E., Datta, S. R., Takasu, M. A., and Greenberg, M. E. (1999) Science 286, Yamamoto, K. K., Gonzalez, G. A., Biggs, W. H. d., and Montminy, M. R. (1988) Nature 334, Chrivia, J. C., Kwok, R. P., Lamb, N., Hagiwara, M., Montminy, M. R., and Goodman, R. H. (1993) Nature 365, Kwok, R. P., Lundblad, J. R., Chrivia, J. C., Richards, J. P., Bachinger, H. P., Brennan, R. G., Roberts, S. G., Green, M. R., and Goodman, R. H. (1994) Nature 370, Gonzalez, G. A., and Montminy, M. R. (1989) Cell 59, Hagiwara, M., Brindle, P., Harootunian, A., Armstrong, R., Rivier, J., Vale, W., Tsien, R., and Montminy, M. R. (1993) Mol. Cell. Biol. 13, Sheng, M., Thompson, M. A., and Greenberg, M. E. (1991) Science 252, Sun, P., Enslen, H., Myung, P. S., and Maurer, R. A. (1994) Genes Dev. 8,
23 16. Matthews, R. P., Guthrie, C. R., Wailes, L. M., Zhao, X., Means, A. R., and McKnight, G. S. (1994) Mol. Cell. Biol. 14, Deisseroth, K., Bito, H., and Tsien, R. W. (1996) Neuron 16, Dash, P. K., Karl, K. A., Colicos, M. A., Prywes, R., and Kandel, E. R. (1991) Proc. Natl. Acad. Sci. USA 88, Xing, J., Ginty, D. D., and Greenberg, M. E. (1996) Science 273, Deak, M., Clifton, A. D., Lucocq, L. M., and Alessi, D. R. (1998) EMBO J. 17, Impey, S., Obrietan, K., Wong, S. T., Poser, S., Yano, S., Wayman, G., Deloulme, J. C., Chan, G., and Storm, D. R. (1998) Neuron 21, Chawla, S., Hardingham, G. E., Quinn, D. R., and Bading, H. (1998) Science 281, Ait-Si-Ali, S., Carlisi, D., Ramirez, S., Upegui-Gonzalez, L. C., Duquet, A., Robin, P., Rudkin, B., Harel-Bellan, A., and Trouche, D. (1999) Biochem. Biophys. Res. Commun. 262, Hu, S. C., Chrivia, J., and Ghosh, A. (1999) Neuron 22, Hardingham, G. E., Chawla, S., Cruzalegui, F. H., and Bading, H. (1999) Neuron 22, Xu, L., Lavinsky, R. M., Dasen, J. S., Flynn, S. E., McInerney, E. M., Mullen, T. M., Heinzel, T., Szeto, D., Korzus, E., Kurokawa, R., Aggarwal, A. K., Rose, D. W., Glass, C. K., and Rosenfeld, M. G. (1998) Nature 395, Liu, Y. Z., Chrivia, J. C., and Latchman, D. S. (1998) J. Biol. Chem. 273, Liu, Y. Z., Thomas, N. S., and Latchman, D. S. (1999) Neuroreport 10, Grewal, S. S., Horgan, A. M., York, R. D., Withers, G. S., Banker, G. A., and Stork, P. J. (2000) J. Biol. Chem. 275,
24 30. Vossler, M. R., Yao, H., York, R. D., Pan, M. G., Rim, C. S., and Stork, P. J. (1997) Cell 89, York, R. D., Yao, H., Dillon, T., Ellig, C. L., Eckert, S. P., McCleskey, E. W., and Stork, P. J. (1998) Nature 392, Grewal, S. S., York, R. D., and Stork, P. J. (1999) Curr. Opin. Neurobiol. 9, Dugan, L. L., Kim, J. S., Zhang, Y., Bart, R. D., Sun, Y., Holtzman, D. M., and Gutmann, D. H. (1999) J. Biol. Chem. 274, Chen, W., Shields, T. S., Stork, P. J. S., and Cone, R. D. (1995) Analytical Biochem. 226, Rubinfeld, B., Munemitsu, S., Clark, R., Conroy, L., Watt, K., Crosier, W. J., McCormick, F., and Polakis, P. (1991) Cell 65, Jordan, J. D., Carey, K. D., Stork, P. J., and Iyengar, R. (1999) J. Biol. Chem. 274, Hagiwara, M., Alberts, A., Brindle, P., Meinkoth, J., Feramisco, J., Deng, T., Karin, M., Shenolikar, S., and Montminy, M. (1992) Cell 70, Bonni, A., Ginty, D. D., Dudek, H., and Greenberg, M. E. (1995) Mol. Cell. Neurosci. 6, Cardinaux, J. R., Notis, J. C., Zhang, Q., Vo, N., Craig, J. C., Fass, D. M., Brennan, R. G., and Goodman, R. H. (2000) Mol. Cell. Biol. 20, Martin, K. C., Michael, D., Rose, J. C., Barad, M., Casadio, A., Zhu, H., and Kandel, E. R. (1997) Neuron 18, Ginty, D. D., Glowacka, D., Bader, D. S., Hidaka, H., and Wagner, J. A. (1991) J. Biol. Chem. 266, Thompson, M. A., Ginty, D. D., Bonni, A., and Greenberg, M. E. (1995) J. Biol. Chem. 270,
25 43. Janknecht, R., and Nordheim, A. (1996) Biochem. Biophys. Res. Commun. 228, Zanger, K., Cohen, L. E., Hashimoto, K., Radovick, S., and Wondisford, F. E. (1999) Mol. Endocrinol. 13, Swope, D. L., Mueller, C. L., and Chrivia, J. C. (1996) J. Biol. Chem. 271, Nakajima, T., Fukamizu, A., Takahashi, J., Gage, F. H., Fisher, T., Blenis, J., and Montminy, M. R. (1996) Cell 86, Cook, S. J., and McCormick, F. (1993) Science 262, Wu, J., Dent, P., Jelinek, T., Wolfman, A., Weber, M. J., and Sturgill, T. W. (1993) Science 262, Janknecht, R., Wells, N. J., and Hunter, T. (1998) Genes Dev. 12, Feng, X. H., Zhang, Y., Wu, R. Y., and Derynck, R. (1998) Genes Dev. 12, Kurokawa, R., Kalafus, D., Ogliastro, M. H., Kioussi, C., Xu, L., Torchia, J., Rosenfeld, M. G., and Glass, C. K. (1998) Science 279, Kamei, Y., Xu, L., Heinzel, T., Torchia, J., Kurokawa, R., Gloss, B., Lin, S. C., Heyman, R. A., Rose, D. W., Glass, C. K., and Rosenfeld, M. G. (1996) Cell 85, Colledge, M., and Scott, J. D. (1999) Trends Cell Biol. 9, Hardingham, G. E., Chawla, S., Johnson, C. M., and Bading, H. (1997) Nature 385, Ginty, D. D. (1997) Neuron 18, Roberson, E. D., English, J. D., Adams, J. P., Selcher, J. C., Kondratick, C., and Sweatt, J. D. (1999) J. Neurosci. 19, Walton, M. R., and Dragunow, I. (2000) Trends Neurosci. 23,
26 58. Visvader, J., Sassone-Corsi, P., and Verma, I. (1988) Proc. Natl. Acad. Sci. USA 85,
27 (A) (B) fold increase CRE-luciferase fold increase CRE-luciferase :µg PKI KCl forskolin :µm PD98059 KCl forskolin
28
29
30
31
32
33
34
35
36 Calcium and Cyclic AMP signals differentially regulate CREB function via a Rap1-ERK pathway Savraj S. Grewal, Daniel M. Fass, Hong Yao, Cindy L. Ellig, Richard H. Goodman and Philip J. S. Stork J. Biol. Chem. published online August 18, 2000 Access the most updated version of this article at doi: /jbc.M Alerts: When this article is cited When a correction for this article is posted Click here to choose from all of JBC's alerts
Ethanol Stimulates camp-responsive Element (CRE)-Mediated Transcription via CRE-Binding Protein and camp-dependent Protein Kinase
 0022-3565/02/3011-66 70$7.00 THE JOURNAL OF PHARMACOLOGY AND EXPERIMENTAL THERAPEUTICS Vol. 301, No. 1 Copyright 2002 by The American Society for Pharmacology and Experimental Therapeutics 4616/967855
0022-3565/02/3011-66 70$7.00 THE JOURNAL OF PHARMACOLOGY AND EXPERIMENTAL THERAPEUTICS Vol. 301, No. 1 Copyright 2002 by The American Society for Pharmacology and Experimental Therapeutics 4616/967855
REGULATION OF CREB PHOSPHORYLATION BY camp AND CA 2+ IN PAROTID ACINAR CELLS
 Vol. 43, No. 3, October 1997 BIOCHEMISTRY end MOLECULAR BIOLOGY INTERNATIONAL Pages 563-570 REGULATION OF CREB PHOSPHORYLATION BY camp AND CA 2+ IN PAROTID ACINAR CELLS Taishin Takuma TM, Yoshifumi Tajima
Vol. 43, No. 3, October 1997 BIOCHEMISTRY end MOLECULAR BIOLOGY INTERNATIONAL Pages 563-570 REGULATION OF CREB PHOSPHORYLATION BY camp AND CA 2+ IN PAROTID ACINAR CELLS Taishin Takuma TM, Yoshifumi Tajima
Lecture 15. Signal Transduction Pathways - Introduction
 Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
Part-4. Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death
 Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
Part-4 Cell cycle regulatory protein 5 (Cdk5) A novel target of ERK in Carb induced cell death 95 1. Introduction The process of replicating DNA and dividing cells can be described as a series of coordinated
Electrical Stimulation Control Nerve Regeneration via the p38 Mitogen-activated Protein Kinase and CREB
 Electrical Stimulation Control Nerve Regeneration via the p38 Mitogen-activated Protein Kinase and CREB Kenji Kawamura, Yoshio Kano. Kibi International University, Takahashi-city, Japan. Disclosures: K.
Electrical Stimulation Control Nerve Regeneration via the p38 Mitogen-activated Protein Kinase and CREB Kenji Kawamura, Yoshio Kano. Kibi International University, Takahashi-city, Japan. Disclosures: K.
Cellular Signaling Pathways. Signaling Overview
 Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
Biol403 MAP kinase signalling
 Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
CREB Transcriptional Activity in Neurons Is Regulated by Multiple, Calcium-Specific Phosphorylation Events
 Neuron, Vol. 34, 221 233, April 11, 2002, Copyright 2002 by Cell Press CREB Transcriptional Activity in Neurons Is Regulated by Multiple, Calcium-Specific Phosphorylation Events Jon M. Kornhauser, 1 Christopher
Neuron, Vol. 34, 221 233, April 11, 2002, Copyright 2002 by Cell Press CREB Transcriptional Activity in Neurons Is Regulated by Multiple, Calcium-Specific Phosphorylation Events Jon M. Kornhauser, 1 Christopher
Molecular Mechanisms Underlying Activity- Dependent Regulation of BDNF Expression
 Molecular Mechanisms Underlying Activity- Dependent Regulation of BDNF Expression Perry B. Shieh, Anirvan Ghosh Department of Neuroscience, Johns Hopkins University School of Medicine, 725 N. Wolfe St.,
Molecular Mechanisms Underlying Activity- Dependent Regulation of BDNF Expression Perry B. Shieh, Anirvan Ghosh Department of Neuroscience, Johns Hopkins University School of Medicine, 725 N. Wolfe St.,
Regulation of cell function by intracellular signaling
 Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
Revision. camp pathway
 االله الرحمن الرحيم بسم Revision camp pathway camp pathway Revision camp pathway Adenylate cyclase Adenylate Cyclase enzyme Adenylate cyclase catalyses the formation of camp from ATP. Stimulation or inhibition
االله الرحمن الرحيم بسم Revision camp pathway camp pathway Revision camp pathway Adenylate cyclase Adenylate Cyclase enzyme Adenylate cyclase catalyses the formation of camp from ATP. Stimulation or inhibition
Sarah Jaar Marah Al-Darawsheh
 22 Sarah Jaar Marah Al-Darawsheh Faisal Mohammad Receptors can be membrane proteins (for water-soluble hormones/ligands) or intracellular (found in the cytosol or nucleus and bind to DNA, for lipid-soluble
22 Sarah Jaar Marah Al-Darawsheh Faisal Mohammad Receptors can be membrane proteins (for water-soluble hormones/ligands) or intracellular (found in the cytosol or nucleus and bind to DNA, for lipid-soluble
Computational Biology I LSM5191
 Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture 6 Notes: Control Systems in Gene Expression Pulling it all together: coordinated control of transcriptional regulatory molecules Simple Control:
Computational Biology I LSM5191 Aylwin Ng, D.Phil Lecture 6 Notes: Control Systems in Gene Expression Pulling it all together: coordinated control of transcriptional regulatory molecules Simple Control:
Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression
 Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
SUPPLEMENTARY INFORMATION. Supplementary Figures S1-S9. Supplementary Methods
 SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
BIPN 140 Problem Set 6
 BIPN 140 Problem Set 6 1) The hippocampus is a cortical structure in the medial portion of the temporal lobe (medial temporal lobe in primates. a) What is the main function of the hippocampus? The hippocampus
BIPN 140 Problem Set 6 1) The hippocampus is a cortical structure in the medial portion of the temporal lobe (medial temporal lobe in primates. a) What is the main function of the hippocampus? The hippocampus
Principles of Genetics and Molecular Biology
 Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras
 Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
Bioluminescence Resonance Energy Transfer (BRET)-based studies of receptor dynamics in living cells with Berthold s Mithras Tarik Issad, Ralf Jockers and Stefano Marullo 1 Because they play a pivotal role
BIPN 140 Problem Set 6
 BIPN 140 Problem Set 6 1) Hippocampus is a cortical structure in the medial portion of the temporal lobe (medial temporal lobe in primates. a) What is the main function of the hippocampus? The hippocampus
BIPN 140 Problem Set 6 1) Hippocampus is a cortical structure in the medial portion of the temporal lobe (medial temporal lobe in primates. a) What is the main function of the hippocampus? The hippocampus
Protein kinases are enzymes that add a phosphate group to proteins according to the. ATP + protein OH > Protein OPO 3 + ADP
 Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
The elements of G protein-coupled receptor systems
 The elements of G protein-coupled receptor systems Prostaglandines Sphingosine 1-phosphate a receptor that contains 7 membrane-spanning domains a coupled trimeric G protein which functions as a switch
The elements of G protein-coupled receptor systems Prostaglandines Sphingosine 1-phosphate a receptor that contains 7 membrane-spanning domains a coupled trimeric G protein which functions as a switch
Signal Transduction: G-Protein Coupled Receptors
 Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
G-Protein Signaling. Introduction to intracellular signaling. Dr. SARRAY Sameh, Ph.D
 G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
MCB*4010 Midterm Exam / Winter 2008
 MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
Ionotropic glutamate receptors (iglurs)
 Ionotropic glutamate receptors (iglurs) GluA1 GluA2 GluA3 GluA4 GluN1 GluN2A GluN2B GluN2C GluN2D GluN3A GluN3B GluK1 GluK2 GluK3 GluK4 GluK5 The general architecture of receptor subunits Unique properties
Ionotropic glutamate receptors (iglurs) GluA1 GluA2 GluA3 GluA4 GluN1 GluN2A GluN2B GluN2C GluN2D GluN3A GluN3B GluK1 GluK2 GluK3 GluK4 GluK5 The general architecture of receptor subunits Unique properties
Membrane associated receptor transfers the information. Second messengers relay information
 Membrane associated receptor transfers the information Most signals are polar and large Few of the signals are nonpolar Receptors are intrinsic membrane proteins Extracellular and intracellular domains
Membrane associated receptor transfers the information Most signals are polar and large Few of the signals are nonpolar Receptors are intrinsic membrane proteins Extracellular and intracellular domains
By the name of Allah
 By the name of Allah Receptors function and signal transduction ( Hormones and receptors Types) We were talking about receptors of the neurotransmitters; we have 2 types of receptors: 1- Ionotropic receptors
By the name of Allah Receptors function and signal transduction ( Hormones and receptors Types) We were talking about receptors of the neurotransmitters; we have 2 types of receptors: 1- Ionotropic receptors
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cognitive Enhancement Strategies. Florian Plattner, James A. Bibb
 Cognitive Enhancement Strategies Florian Plattner, James A. Bibb A decline in memory and cognitive function is a natural aspect of aging. In addition, cognitive deficits are comorbid with many mental disorders
Cognitive Enhancement Strategies Florian Plattner, James A. Bibb A decline in memory and cognitive function is a natural aspect of aging. In addition, cognitive deficits are comorbid with many mental disorders
The relation of transcription to memory formation
 Vol. 50 No. 3/2003 775 782 QUARTERLY Review The relation of transcription to memory formation Edward Korzus University of California San Diego, Department of Neurosciences, 9500 Gilman Drive (M/C 0986),
Vol. 50 No. 3/2003 775 782 QUARTERLY Review The relation of transcription to memory formation Edward Korzus University of California San Diego, Department of Neurosciences, 9500 Gilman Drive (M/C 0986),
Neurotransmitter Systems II Receptors. Reading: BCP Chapter 6
 Neurotransmitter Systems II Receptors Reading: BCP Chapter 6 Neurotransmitter Systems Normal function of the human brain requires an orderly set of chemical reactions. Some of the most important chemical
Neurotransmitter Systems II Receptors Reading: BCP Chapter 6 Neurotransmitter Systems Normal function of the human brain requires an orderly set of chemical reactions. Some of the most important chemical
Biol220 Cell Signalling Cyclic AMP the classical secondary messenger
 Biol220 Cell Signalling Cyclic AMP the classical secondary messenger The classical secondary messenger model of intracellular signalling A cell surface receptor binds the signal molecule (the primary
Biol220 Cell Signalling Cyclic AMP the classical secondary messenger The classical secondary messenger model of intracellular signalling A cell surface receptor binds the signal molecule (the primary
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature06994 A phosphatase cascade by which rewarding stimuli control nucleosomal response A. Stipanovich*, E. Valjent*, M. Matamales*, A. Nishi, J.H. Ahn, M. Maroteaux, J. Bertran-Gonzalez,
doi: 10.1038/nature06994 A phosphatase cascade by which rewarding stimuli control nucleosomal response A. Stipanovich*, E. Valjent*, M. Matamales*, A. Nishi, J.H. Ahn, M. Maroteaux, J. Bertran-Gonzalez,
Exendin-4-mediated cell proliferation in beta-cells: focusing on the transcription factors
 Exendin-4-mediated cell proliferation in beta-cells: focusing on the transcription factors Seo-Yoon Chang, Jae Min Cho, Yang-Hyeok Jo, Myung-Jun Kim Departments of Physiology, College of Medicine, The
Exendin-4-mediated cell proliferation in beta-cells: focusing on the transcription factors Seo-Yoon Chang, Jae Min Cho, Yang-Hyeok Jo, Myung-Jun Kim Departments of Physiology, College of Medicine, The
Cell Communication. Cell Communication. Communication between cells requires: ligand: the signaling molecule
 Cell Communication Cell Communication Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the ligand binds (may be on the plasma membrane or within
Cell Communication Cell Communication Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the ligand binds (may be on the plasma membrane or within
2013 W. H. Freeman and Company. 12 Signal Transduction
 2013 W. H. Freeman and Company 12 Signal Transduction CHAPTER 12 Signal Transduction Key topics: General features of signal transduction Structure and function of G protein coupled receptors Structure
2013 W. H. Freeman and Company 12 Signal Transduction CHAPTER 12 Signal Transduction Key topics: General features of signal transduction Structure and function of G protein coupled receptors Structure
Lecture: CHAPTER 13 Signal Transduction Pathways
 Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
D 2 dopamine receptors induce mitogen-activated protein kinase and camp response element-binding protein phosphorylation in neurons
 Proc. Natl. Acad. Sci. USA Vol. 96, pp. 11607 11612, September 1999 Neurobiology D 2 dopamine receptors induce mitogen-activated protein kinase and camp response element-binding protein phosphorylation
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 11607 11612, September 1999 Neurobiology D 2 dopamine receptors induce mitogen-activated protein kinase and camp response element-binding protein phosphorylation
Chapter 11. Cell Communication. Signal Transduction Pathways
 Chapter 11 Cell Communication Signal Transduction Pathways Signal-Transduction Pathway Signal on a cell s surface is converted into a specific cellular response Local signaling (short distance) - Paracrine
Chapter 11 Cell Communication Signal Transduction Pathways Signal-Transduction Pathway Signal on a cell s surface is converted into a specific cellular response Local signaling (short distance) - Paracrine
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
www.sciencesignaling.org/cgi/content/full/6/283/ra57/dc1 Supplementary Materials for JNK3 Couples the Neuronal Stress Response to Inhibition of Secretory Trafficking Guang Yang,* Xun Zhou, Jingyan Zhu,
Chapter 9. Cellular Signaling
 Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Enhancement of synaptic transmission by cyclic AMP modulation of presynaptic I h channels. Vahri Beaumont and Robert S. Zucker
 Enhancement of synaptic transmission by cyclic AMP modulation of presynaptic I h channels Vahri Beaumont and Robert S. Zucker Background I h channels discovered in 1976 (Noma A. and Irisawa H.) Voltage-gated
Enhancement of synaptic transmission by cyclic AMP modulation of presynaptic I h channels Vahri Beaumont and Robert S. Zucker Background I h channels discovered in 1976 (Noma A. and Irisawa H.) Voltage-gated
Receptors and Drug Action. Dr. Subasini Pharmacology Department Ishik University, Erbil
 Receptors and Drug Action Dr. Subasini Pharmacology Department Ishik University, Erbil Receptors and Drug Action Receptor Receptor is defined as a macromolecule or binding site located on the surface or
Receptors and Drug Action Dr. Subasini Pharmacology Department Ishik University, Erbil Receptors and Drug Action Receptor Receptor is defined as a macromolecule or binding site located on the surface or
Chapter 5 CREB Glycosylation Moderates Phosphorylation-Dependent CREB Activity in Cultured Pancreatic Cell Lines.
 79 Chapter 5 CREB Glycosylation Moderates Phosphorylation-Dependent CREB Activity in Cultured Pancreatic Cell Lines. Summary. In vitro studies of CREB glycosylation found that the transactivation potential
79 Chapter 5 CREB Glycosylation Moderates Phosphorylation-Dependent CREB Activity in Cultured Pancreatic Cell Lines. Summary. In vitro studies of CREB glycosylation found that the transactivation potential
Mechanisms of Hormone Action
 Mechanisms of Hormone Action General principles: 1. Signals act over different ranges. 2. Signals have different chemical natures. 3. The same signal can induce a different response in different cells.
Mechanisms of Hormone Action General principles: 1. Signals act over different ranges. 2. Signals have different chemical natures. 3. The same signal can induce a different response in different cells.
Supplementary Table 1. List of primers used in this study
 Supplementary Table 1. List of primers used in this study Gene Forward primer Reverse primer Rat Met 5 -aggtcgcttcatgcaggt-3 5 -tccggagacacaggatgg-3 Rat Runx1 5 -cctccttgaaccactccact-3 5 -ctggatctgcctggcatc-3
Supplementary Table 1. List of primers used in this study Gene Forward primer Reverse primer Rat Met 5 -aggtcgcttcatgcaggt-3 5 -tccggagacacaggatgg-3 Rat Runx1 5 -cctccttgaaccactccact-3 5 -ctggatctgcctggcatc-3
Biochemie 4. Cell communication - GPCR
 Biochemie 4 Cell communication - GPCR 1 Lecture outline General principles - local and long-distance signaling - classes of receptors - molecular switches and second messengers Receptor tyrosine kinases
Biochemie 4 Cell communication - GPCR 1 Lecture outline General principles - local and long-distance signaling - classes of receptors - molecular switches and second messengers Receptor tyrosine kinases
Yanping Li, Maho Takahashi, and Philip J. S. Stork 1 From the Vollum Institute, Oregon Health and Science University, Portland, Oregon 97239
 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 288, NO. 38, pp. 27646 27657, September 20, 2013 2013 by The American Society for Biochemistry and Molecular Biology, Inc. Published in the U.S.A. Ras-mutant Cancer
THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 288, NO. 38, pp. 27646 27657, September 20, 2013 2013 by The American Society for Biochemistry and Molecular Biology, Inc. Published in the U.S.A. Ras-mutant Cancer
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION
 Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
Growth and Differentiation Phosphorylation Sampler Kit
 Growth and Differentiation Phosphorylation Sampler Kit E 0 5 1 0 1 4 Kits Includes Cat. Quantity Application Reactivity Source Akt (Phospho-Ser473) E011054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit
Growth and Differentiation Phosphorylation Sampler Kit E 0 5 1 0 1 4 Kits Includes Cat. Quantity Application Reactivity Source Akt (Phospho-Ser473) E011054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit
Goals and Challenges of Communication. Communication and Signal Transduction. How Do Cells Communicate?
 Goals and Challenges of Communication Reaching (only) the correct recipient(s) Imparting correct information Timeliness Causing the desired effect Effective termination Communication and Signal Transduction
Goals and Challenges of Communication Reaching (only) the correct recipient(s) Imparting correct information Timeliness Causing the desired effect Effective termination Communication and Signal Transduction
3. Results. 3.1 Elevated osmolality is required for the DBcAMP-elicited expression of robust AQP2 protein levels in IMCD cells
 3. Results 3.1 Elevated osmolality is required for the DcAMPelicited expression of robust AQP2 protein levels in IMCD cells Primary cultured IMCD cells, derived from rat inner medullae, were used as a
3. Results 3.1 Elevated osmolality is required for the DcAMPelicited expression of robust AQP2 protein levels in IMCD cells Primary cultured IMCD cells, derived from rat inner medullae, were used as a
Principles of cell signaling Lecture 4
 Principles of cell signaling Lecture 4 Johan Lennartsson Molecular Cell Biology (1BG320), 2014 Johan.Lennartsson@licr.uu.se 1 Receptor tyrosine kinase-induced signal transduction Erk MAP kinase pathway
Principles of cell signaling Lecture 4 Johan Lennartsson Molecular Cell Biology (1BG320), 2014 Johan.Lennartsson@licr.uu.se 1 Receptor tyrosine kinase-induced signal transduction Erk MAP kinase pathway
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation.
 Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Leen Osama, Lujain Hamdan, Osama Mohd, Razi Kittaneh... Faisal Mohammad
 23 Leen Osama, Lujain Hamdan, Osama Mohd, Razi Kittaneh... Faisal Mohammad Revision of previous lectures G-proteins coupled receptors mechanism: When a hormone binds to G-protein coupled receptor, GTP
23 Leen Osama, Lujain Hamdan, Osama Mohd, Razi Kittaneh... Faisal Mohammad Revision of previous lectures G-proteins coupled receptors mechanism: When a hormone binds to G-protein coupled receptor, GTP
condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1%
 FIGURE LEGENDS Supplementary Fig 1 (A) sumoylation pattern detected under denaturing condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1% SDS in the presence and absence
FIGURE LEGENDS Supplementary Fig 1 (A) sumoylation pattern detected under denaturing condition. Left panel, the HCT-116 cells were lysed with RIPA buffer containing 0.1% SDS in the presence and absence
ab SREBP-2 Translocation Assay Kit (Cell-Based)
 ab133114 SREBP-2 Translocation Assay Kit (Cell-Based) Instructions for Use For analysis of translocation of SREBP-2 into nuclei. This product is for research use only and is not intended for diagnostic
ab133114 SREBP-2 Translocation Assay Kit (Cell-Based) Instructions for Use For analysis of translocation of SREBP-2 into nuclei. This product is for research use only and is not intended for diagnostic
Vets 111/Biov 111 Cell Signalling-2. Secondary messengers the cyclic AMP intracellular signalling system
 Vets 111/Biov 111 Cell Signalling-2 Secondary messengers the cyclic AMP intracellular signalling system The classical secondary messenger model of intracellular signalling A cell surface receptor binds
Vets 111/Biov 111 Cell Signalling-2 Secondary messengers the cyclic AMP intracellular signalling system The classical secondary messenger model of intracellular signalling A cell surface receptor binds
Cell Communication. Cell Communication. Cell Communication. Cell Communication. Cell Communication. Chapter 9. Communication between cells requires:
 Chapter 9 Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the receptor binds -may be on the plasma membrane or within the cell 2 There are four
Chapter 9 Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the receptor binds -may be on the plasma membrane or within the cell 2 There are four
Kirschner, 2005). Briefly, parallel MEF cultures were isolated from single littermate
 SUPPLEMENTAL MATERIALS AND METHODS Generation of MEFs, osteoblasts and cell culture Prkar1a -/- and control MEFs were generated and cultured as described (Nadella and Kirschner, 2005). Briefly, parallel
SUPPLEMENTAL MATERIALS AND METHODS Generation of MEFs, osteoblasts and cell culture Prkar1a -/- and control MEFs were generated and cultured as described (Nadella and Kirschner, 2005). Briefly, parallel
Psych 181: Dr. Anagnostaras
 Psych 181: Dr. Anagnostaras Lecture 5 Synaptic Transmission Introduction to synaptic transmission Synapses (Gk., to clasp or join) Site of action of most psychoactive drugs 6.5 1 Synapses Know basic terminology:
Psych 181: Dr. Anagnostaras Lecture 5 Synaptic Transmission Introduction to synaptic transmission Synapses (Gk., to clasp or join) Site of action of most psychoactive drugs 6.5 1 Synapses Know basic terminology:
CREB1 (phospho S133) Transcription Factor Assay Kit
 CREB1 (phospho S133) Transcription Factor Assay Kit Catalog Number KA1308 96 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle
CREB1 (phospho S133) Transcription Factor Assay Kit Catalog Number KA1308 96 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle
Hormones and Signal Transduction. Dr. Kevin Ahern
 Dr. Kevin Ahern Signaling Outline Signaling Outline Background Signaling Outline Background Membranes Signaling Outline Background Membranes Hormones & Receptors Signaling Outline Background Membranes
Dr. Kevin Ahern Signaling Outline Signaling Outline Background Signaling Outline Background Membranes Signaling Outline Background Membranes Hormones & Receptors Signaling Outline Background Membranes
Plasma membranes. Plasmodesmata between plant cells. Gap junctions between animal cells Cell junctions. Cell-cell recognition
 Cell Communication Cell Signaling Cell-to-cell communication is essential for multicellular organisms Communicate by chemical messengers Animal and plant cells have cell junctions that directly connect
Cell Communication Cell Signaling Cell-to-cell communication is essential for multicellular organisms Communicate by chemical messengers Animal and plant cells have cell junctions that directly connect
Bio 111 Study Guide Chapter 11 Cell Communication
 Bio 111 Study Guide Chapter 11 Cell Communication BEFORE CLASS: Reading: Read the introduction on p. 210, and for Concept 11.1, read from the first full paragraph on p. 212. Read all of Concept 11.2. Pay
Bio 111 Study Guide Chapter 11 Cell Communication BEFORE CLASS: Reading: Read the introduction on p. 210, and for Concept 11.1, read from the first full paragraph on p. 212. Read all of Concept 11.2. Pay
Propagation of the Signal
 OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
EFFECTORS IMPLICATED IN THE AC1 INHIBITORY EFFECT ON CELL PROLIFERATION IN PANCREATIC CANCER CELLS
 EFFECTORS IMPLICATED IN THE AC1 INHIBITORY EFFECT ON CELL PROLIFERATION IN PANCREATIC CANCER CELLS VIDYA MEDEPALLI ADVISOR: Maria Eugenia Sabbatini, PhD PHI KAPPA PHI RESEARCH CONFERENCE MARCH 18 TH, 2016
EFFECTORS IMPLICATED IN THE AC1 INHIBITORY EFFECT ON CELL PROLIFERATION IN PANCREATIC CANCER CELLS VIDYA MEDEPALLI ADVISOR: Maria Eugenia Sabbatini, PhD PHI KAPPA PHI RESEARCH CONFERENCE MARCH 18 TH, 2016
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Data Sheet. Notch Pathway Reporter Kit Catalog # 60509
 Data Sheet Notch Pathway Reporter Kit Catalog # 60509 6042 Cornerstone Court W, Ste B Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. NOTCH signaling
Data Sheet Notch Pathway Reporter Kit Catalog # 60509 6042 Cornerstone Court W, Ste B Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. NOTCH signaling
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Chairoungdua et al., http://www.jcb.org/cgi/content/full/jcb.201002049/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Expression of CD9 and CD82 inhibits Wnt/ -catenin
Supplemental material Chairoungdua et al., http://www.jcb.org/cgi/content/full/jcb.201002049/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Expression of CD9 and CD82 inhibits Wnt/ -catenin
KEY CONCEPT QUESTIONS IN SIGNAL TRANSDUCTION
 Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
FUNDAMENTALS OF BIOCHEMISTRY, CELL BIOLOGY AND BIOPHYSICS Vol. I - Biochemistry of Vitamins, Hormones and Other Messenger Molecules - Chris Whiteley
 BIOCHEMISTRY OF VITAMINS, HORMONES AND OTHER MESSENGER MOLECULES Chris Whiteley Department of Biochemistry and Microbiology, Rhodes University, Grahamstown, South Africa Keywords: phosphorylation, phosphorylase,
BIOCHEMISTRY OF VITAMINS, HORMONES AND OTHER MESSENGER MOLECULES Chris Whiteley Department of Biochemistry and Microbiology, Rhodes University, Grahamstown, South Africa Keywords: phosphorylation, phosphorylase,
INTERACTION DRUG BODY
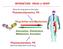 INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
2.5. AMPK activity
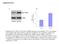 Supplement Fig. A 3 B phos-ampk 2.5 * Control AICAR AMPK AMPK activity (Absorbance at 45 nm) 2.5.5 Control AICAR Supplement Fig. Effects of AICAR on AMPK activation in macrophages. J774. macrophages were
Supplement Fig. A 3 B phos-ampk 2.5 * Control AICAR AMPK AMPK activity (Absorbance at 45 nm) 2.5.5 Control AICAR Supplement Fig. Effects of AICAR on AMPK activation in macrophages. J774. macrophages were
Acetylation of CREB by CBP enhances CREB-dependent transcription
 JBC Papers in Press. Published on February 2, 23 as Manuscript M35462 Acetylation of CREB by CBP enhances CREB-dependent transcription Qing Lu 1, Amanda E. Hutchins 1, Colleen M. Doyle 1, James R. Lundblad
JBC Papers in Press. Published on February 2, 23 as Manuscript M35462 Acetylation of CREB by CBP enhances CREB-dependent transcription Qing Lu 1, Amanda E. Hutchins 1, Colleen M. Doyle 1, James R. Lundblad
Chapter 15: Signal transduction
 Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
MEK1 Assay Kit 1 Catalog # Lot # 16875
 MEK1 Assay Kit 1 Kit Components Assay Dilution Buffer (ADB), Catalog # 20-108. Three vials, each containing 1.0ml of assay dilution buffer (20mM MOPS, ph 7.2, 25mM ß-glycerol phosphate, 5mM EGTA, 1mM sodium
MEK1 Assay Kit 1 Kit Components Assay Dilution Buffer (ADB), Catalog # 20-108. Three vials, each containing 1.0ml of assay dilution buffer (20mM MOPS, ph 7.2, 25mM ß-glycerol phosphate, 5mM EGTA, 1mM sodium
a b G75 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server.
 a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
Synaptic plasticityhippocampus. Neur 8790 Topics in Neuroscience: Neuroplasticity. Outline. Synaptic plasticity hypothesis
 Synaptic plasticityhippocampus Neur 8790 Topics in Neuroscience: Neuroplasticity Outline Synaptic plasticity hypothesis Long term potentiation in the hippocampus How it s measured What it looks like Mechanisms
Synaptic plasticityhippocampus Neur 8790 Topics in Neuroscience: Neuroplasticity Outline Synaptic plasticity hypothesis Long term potentiation in the hippocampus How it s measured What it looks like Mechanisms
Biosignals, Chapter 8, rearranged, Part I
 Biosignals, Chapter 8, rearranged, Part I Nicotinic Acetylcholine Receptor: A Ligand-Binding Ion Channel Classes of Receptor Proteins in Eukaryotes, Heterotrimeric G Proteins Signaling View the Heterotrimeric
Biosignals, Chapter 8, rearranged, Part I Nicotinic Acetylcholine Receptor: A Ligand-Binding Ion Channel Classes of Receptor Proteins in Eukaryotes, Heterotrimeric G Proteins Signaling View the Heterotrimeric
Supplementary Material and Methods
 Online Supplement Kockx et al, Secretion of Apolipoprotein E from Macrophages 1 Supplementary Material and Methods Cloning of ApoE-GFP Full-length human apoe3 cdna (pcdna3.1/zeo + -apoe) was kindly provided
Online Supplement Kockx et al, Secretion of Apolipoprotein E from Macrophages 1 Supplementary Material and Methods Cloning of ApoE-GFP Full-length human apoe3 cdna (pcdna3.1/zeo + -apoe) was kindly provided
ORIGINAL PAPER DISRUPTION OF CELL SPREADING BY THE ACTIVATION OF MEK/ERK PATHWAY IS DEPENDENT ON AP-1 ACTIVITY
 Nagoya J. Med. Sci. 72. 139 ~ 144, 1 ORIGINAL PAPER DISRUPTION OF CELL SPREADING BY THE ACTIVATION OF MEK/ERK PATHWAY IS DEPENDENT ON AP-1 ACTIVITY FENG XU, SATOKO ITO, MICHINARI HAMAGUCHI and TAKESHI
Nagoya J. Med. Sci. 72. 139 ~ 144, 1 ORIGINAL PAPER DISRUPTION OF CELL SPREADING BY THE ACTIVATION OF MEK/ERK PATHWAY IS DEPENDENT ON AP-1 ACTIVITY FENG XU, SATOKO ITO, MICHINARI HAMAGUCHI and TAKESHI
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Human PKA (Protein Kinase A) Activity Assay Kit
 Human PKA (Protein Kinase A) Activity Assay Kit CATALOG NO: IRAAKT2532 LOT NO: SAMPLE INTENDED USE The PKA (Protein Kinase A) Activity kit is designed to quantitatively measure PKA activity in a variety
Human PKA (Protein Kinase A) Activity Assay Kit CATALOG NO: IRAAKT2532 LOT NO: SAMPLE INTENDED USE The PKA (Protein Kinase A) Activity kit is designed to quantitatively measure PKA activity in a variety
Supplemental Information. Menin Deficiency Leads to Depressive-like. Behaviors in Mice by Modulating. Astrocyte-Mediated Neuroinflammation
 Neuron, Volume 100 Supplemental Information Menin Deficiency Leads to Depressive-like Behaviors in Mice by Modulating Astrocyte-Mediated Neuroinflammation Lige Leng, Kai Zhuang, Zeyue Liu, Changquan Huang,
Neuron, Volume 100 Supplemental Information Menin Deficiency Leads to Depressive-like Behaviors in Mice by Modulating Astrocyte-Mediated Neuroinflammation Lige Leng, Kai Zhuang, Zeyue Liu, Changquan Huang,
SUPPLEMENTARY INFORMATION
 Supplementary Figure 1. Normal AMPAR-mediated fepsp input-output curve in CA3-Psen cdko mice. Input-output curves, which are plotted initial slopes of the evoked fepsp as function of the amplitude of the
Supplementary Figure 1. Normal AMPAR-mediated fepsp input-output curve in CA3-Psen cdko mice. Input-output curves, which are plotted initial slopes of the evoked fepsp as function of the amplitude of the
Lecture 9: Cell Communication I
 02.05.10 Lecture 9: Cell Communication I Multicellular organisms need to coordinate cellular functions in different tissues Cell-to-cell communication is also used by single celled organisms to signal
02.05.10 Lecture 9: Cell Communication I Multicellular organisms need to coordinate cellular functions in different tissues Cell-to-cell communication is also used by single celled organisms to signal
Xenoestrogen-induced Regulation of EZH2 and Histone Methylation via Non-Genomic Estrogen
 Xenoestrogen-induced Regulation of EZH2 and Histone Methylation via Non-Genomic Estrogen Receptor Signaling to PI3K/AKT Tiffany G. Bredfeldt, Kristen L. Greathouse, Stephen H. Safe, Mien-Chie Hung, Mark
Xenoestrogen-induced Regulation of EZH2 and Histone Methylation via Non-Genomic Estrogen Receptor Signaling to PI3K/AKT Tiffany G. Bredfeldt, Kristen L. Greathouse, Stephen H. Safe, Mien-Chie Hung, Mark
Model Answer. M.Sc. Zoology (First Semester) Examination Paper LZT 103 (Endocrinology)
 Model Answer M.Sc. Zoology (First Semester) Examination-2013 Paper LZT 103 (Endocrinology) Section A 1. (i) d (ii) b (iii) b (iv) c (v) c (vi) a (vii) c (viii) a (ix) d (x) b Section B Q.2 Answer Hormonal
Model Answer M.Sc. Zoology (First Semester) Examination-2013 Paper LZT 103 (Endocrinology) Section A 1. (i) d (ii) b (iii) b (iv) c (v) c (vi) a (vii) c (viii) a (ix) d (x) b Section B Q.2 Answer Hormonal
Synaptic Transmission: Ionic and Metabotropic
 Synaptic Transmission: Ionic and Metabotropic D. Purves et al. Neuroscience (Sinauer Assoc.) Chapters 5, 6, 7. C. Koch. Biophysics of Computation (Oxford) Chapter 4. J.G. Nicholls et al. From Neuron to
Synaptic Transmission: Ionic and Metabotropic D. Purves et al. Neuroscience (Sinauer Assoc.) Chapters 5, 6, 7. C. Koch. Biophysics of Computation (Oxford) Chapter 4. J.G. Nicholls et al. From Neuron to
FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643
 Data Sheet FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643 Background The PI3K/AKT signaling pathway is essential for cell growth and survival. Disruption of this pathway or its regulation has been linked
Data Sheet FOXO Reporter Kit PI3K/AKT Pathway Cat. #60643 Background The PI3K/AKT signaling pathway is essential for cell growth and survival. Disruption of this pathway or its regulation has been linked
BIOLOGY. Cell Communication CAMPBELL. Reece Urry Cain Wasserman Minorsky Jackson. Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick
 CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 11 Cell Communication Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Cellular Messaging Cells can signal to
CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 11 Cell Communication Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Cellular Messaging Cells can signal to
