Difference imaging reveals ordered regions of RNA in turnip yellow mosaic virus B Böttcher* and RA Crowther
|
|
|
- Justin Randall
- 6 years ago
- Views:
Transcription
1 Research Article 387 Difference imaging reveals ordered regions of RNA in turnip yellow mosaic virus B Böttcher* and RA Crowther Background: Turnip yellow mosaic virus (TYMV) is a small icosahedral plant virus with a capsid containing 180 subunits arranged with hexamer pentamer clustering. Cross-linking studies have indicated extensive contacts between RNA and coat protein, suggesting that substantial parts of the RNA might be icosahedrally ordered. Results: Comparison of maps computed to a Fourier cut-off of 1.5 nm from electron micrographs of ice-embedded specimens of TYMV and of empty capsids produced by freeze-thawing reveals strong inner features around the threefold axes in the virus but not in the empty capsid. Internal features of subunit packing indicate that interhexamer contacts are closer than those between pentamers and hexamers and that pentamer density in the empty capsid is reduced relative to that in the virus. Conclusions: The differences between virus and empty capsid indicate that substantial parts of the RNA are icosahedrally ordered and that the exit of RNA on freeze-thawing is accompanied by the loss of at least one pentamer unit. Address: Medical Research Council Laboratory of Molecular Biology, Hills Road, Cambridge, CB2 2QH, UK. *Corresponding author. Key words: cryo-electron microscopy, difference imaging, image processing, turnip yellow mosaic virus, RNA organization Received: 15 Dec 1995 Revisions requested: 8 Jan 1996 Revisions received: 7 Feb 1996 Accepted: 15 Feb 1996 Structure 15 April 1996, 4: Current Biology Ltd ISSN Introduction Turnip yellow mosaic virus (TYMV) is a member of the tymovirus family. It has a positive-stranded RNA genome with 6318 bases of which 96.9% are coding. The capsid is built of 180 copies of a 20 kda coat protein organized with icosahedral T=3 symmetry and clustered in hexamers and pentamers [1]. The mean outer radius of the capsid was measured to be between 14 nm (by X-ray small-angle scattering [2]) and 15 nm (by cryo-electron microscopy) [3]. Neutron small-angle scattering showed that most of the RNA is buried inside a radius of nm [4,5]. From X-ray diffraction it was predicted that significant parts of the RNA penetrate deeply into the protein shell at the positions of the 32 hexamer and pentamer morphological units, extending out to a radius of 12.5 nm [6]. The viral RNA contains about 38% cytosine and can be cross-linked at ph 7.3 after bisulphite treatment and ultraviolet irradiation mainly to two sites on the coat protein [7]. In the cross-linking experiments almost all of the protein subunits reacted with viral RNA. Upon alkaline treatment or bulk freezing, the intact virus swells by 3 4% before complete release of RNA [4,8]: between five and nine coat protein subunits are lost [9,10]. The remaining protein maintains a spherical shell, which at neutral ph has the same outer dimensions as the virus [8]. It is likely that a similar loss of subunits occurs in vivo during infection of plants [11]. In this paper we present three-dimensional maps of TYMV and of the empty capsid computed to a Fourier cut-off of about 1.5 nm from electron micrographs of unstained frozen specimens. The maps confirm previous models of the overall organization of subunits in the outer part of the capsid but show that the subunits are dimer-clustered at inner radii. The difference map reveals the presence of ordered regions of RNA around the threefold axes and shows the sites of contact between the RNA and the inner surface of the capsid protein. Internal features of subunit packing suggest closer interactions between neighbouring hexamer units than between a pentamer and the surrounding hexamers. The density associated with a pentamer unit is lower in empty capsids than in the virus, indicating that exit of the RNA on freeze-thawing is accompanied by the loss of at least one pentamer unit. Results Micrographs Figure 1 shows micrographs of TYMV and of empty protein capsids. The virus particles have a circular shape and appear to be filled (Fig. 1a). Empty capsids, which have released their RNA after freezing and thawing, have a more ring-shaped appearance (Fig. 1b). Some of these rings appear to be broken or incomplete but many of them seem to be intact. In all micrographs used for reconstruction of TYMV, the rod-shaped tobacco mosaic virus (TMV) was also present (Fig. 1). The TMV particles were used to assess the quality of each micrograph (see Table 1). The TMV also allowed the determination of the size of TYMV and the empty capsids absolutely and independently. It was thus
2 388 Structure 1996, Vol 4 No 4 Figure 1 Table 1 Summary of imaging conditions and quality of micrographs used for TYMV reconstruction. Micrograph number Sum of TYMV transforms Defocus (nm) Ctf zeros (nm):* observed 1.43, , , ,1.87 calculated 1.44, , , ,1.88 No. of TYMV particles included Electron dose (e nm 2 ) TMV amplitude ratio 3/6 after ctf correction Image quality was monitored by the ratio of the amplitudes of the 3rd to the 6th layerline of the best TMV on each micrograph after correction for the ctf. Electron diffraction indicates a true value for the ratio of amplitudes of 0.75 [21]. *For simplicity, only the positions of the first two zeros of the ctf are listed. Micrographs of unstained specimens of (a) TYMV and (b) empty capsids produced by freeze-thawing. Rod-shaped TMV particles are present in both images for reference. Micrographs were taken at the Hitachi HF2000 at 200 kv and magnification with the defocus in (a) =2810 nm and in (b) =1780 nm. Scale bar: 50 nm. possible to check whether the empty shells had a different radius from that of the virus particles. In general micrographs taken at the Hitachi microscope show little contrast. However, it was possible to determine the orientation of these (sometimes barely visible) particles by using a set of reliable reference projections (see the Materials and methods section). Three-dimensional maps Three-dimensional maps of the virus and of the empty capsid are shown as surface representations in Figure 2. The maps, including data from 180 particles for the virus (Fig. 2a) and 56 particles for the empty capsid (Fig. 2b), were calculated to a Fourier cut-off of 1/1.5 nm 1. The reconstructions show an icosahedral shell with T=3 symmetry. The 180 subunits are arranged in 32 knobs, 12 at the fivefold symmetry axes and 20 at the threefold symmetry axes. Indentations in the centres of the knobs at the threefold and fivefold symmetry axes can be observed. The maximum diameter of the particles is 30.1 nm for the knobs at the threefold axes and 31.2 nm at the fivefold axes. Small differences are apparent in the surface representations of the virus and the empty capsid, which we attribute to the much smaller number of particles used in the reconstruction of the latter. The difference map shows strong features at inner radii (Fig. 2c), which we attribute to ordered RNA (see below). When contoured at the same level, the difference map at outer radii (Fig. 2d) shows only tiny features around the pentamers. Spherical sections of the reconstructions were calculated and projected. The 0.5 nm thick spherical slices were spaced radially by 0.5 nm starting at an outer radius of 16 nm and ending at an inner radius of 8.5 nm. Selected sections are shown in Figure 3. In the empty capsid, the density is confined to radii between 11 nm and 15.5 nm (Fig. 3, bottom row), whereas in the virus, density can also be observed at smaller radii (Fig. 3, top row). The spherical sections of the empty capsid and the virus show that densities corresponding to the knobs at the fivefold symmetry axes protrude about 0.5 nm further out than densities corresponding to the knobs at the threefold symmetry axes. The knobs join together to form a continuous shell at an outer radius of about 13 nm. On the inner surface of the capsid (radius about 11.5 nm), the subunits become dimer-clustered. The two subunits forming the strict dimer are disposed
3 Research Article Ordered regions of RNA in TYMV Böttcher and Crowther 389 Figure 2 Surface representation of three-dimensional maps of TYMV. (a) Intact virus. (b) Empty capsid. Difference maps of virus minus empty capsid showing (c) radii nm and (d) radii nm. The maps, which are viewed down a strict twofold axis, were calculated to a Fourier cut-off of 1/1.5 nm 1. All four plots were made at the same density threshold for the surface. approximately towards the fivefold axes (see Fig. 4a). At radii between 12 nm and 12.5 nm (see Fig. 4b) four subunits from neighbouring hexamers (two from each) interlock closely across the strict twofold axes, whereas the neighbouring subunits from the pentamers are excluded from this interaction. The knobs at the fivefold symmetry axes of the virus and of the empty capsid appear to be hollow with a small opening towards the outside of the virus. The opening widens towards the inside. The subunits at the tips of the knobs point towards each other at a radius of 15 nm. Measured relative to the density in hexamers, the density Figure 3 Spherical slices of the three-dimensional maps of TYMV. The intact virus is shown in the top row and the equivalent sections of the empty capsid in the bottom row. The slices are 0.5 nm thick and 1 nm apart starting at an outer radius of 15 nm, as indicated in the lower row. Density corresponding to protein and RNA is shown in white.
4 390 Structure 1996, Vol 4 No 4 Figure 4 Two particular spherical slices of a threedimensional map of TYMV. The map was calculated to a Fourier cut-off of 1/1.15 nm 1 and the sections correspond to radii of (a) 10.9 nm and (b) 12.2 nm. The sections were selected because the subunit packing and contacts between RNA and protein are displayed clearly. of pentamers in empty capsids is 86% of that of pentamers in the virus, indicating that at least one pentamer has been lost in the empty capsids. At the threefold symmetry axes, the knobs have a hole or area of very low protein density in the centre running from the outside of the virus or empty capsid towards the inside, with its narrowest part at a radius of about 13.5 nm. The most prominent features in the virus map at smaller radii are hollow triangles of density centred at the threefold Figure 5 symmetry axes (Fig. 4a). These triangles start at an inner radius of about 9.5 nm and are still faintly visible at an outer radius of 12 nm. No such features are present in the empty capsid and the background density in the empty capsid is on average much lower than in the virus. These hollow triangles centred on the threefold axes, which we believe correspond to ordered RNA, are the dominant features in the difference map computed by subtracting the capsid density from the virus density (Fig. 2c). Radial bridges of density extend from these triangles to the inner surface of the capsid at a radius of about nm, as can be seen clearly when the difference map is superimposed on the map of the virus (Fig. 5). Radial density Radial density plots computed from the three-dimensional maps are shown in Figure 6. Protein density peaks occur at a radius of 12.5 nm in the virus reconstruction (Fig. 6a), as well as in the empty capsid reconstruction (Fig. 6b). The width of the protein peak is about 3 nm. For the virus reconstruction there is an inner peak at a radius of 9.5 nm, which is missing in the radial density of the empty capsid. This peak can be seen clearly in a plot of the difference between the radial densities of the virus and empty capsid reconstructions (Fig. 6c). Superposition of the difference map (virus minus empty capsid) and the virus map. The surface representation of the difference map (see Fig. 2c) is placed inside the TYMV map (see Fig. 2a) but with one quarter of the TYMV map cut away. The density ascribed to RNA is represented in blue, the protein density in yellow. Contrast transfer function correction In all reconstructions, images from micrographs with different defocuses were combined. Thus, information missing at the zeros of the contrast transfer function (ctf) in one micrograph was recovered from other micrographs included in the reconstruction. Nevertheless, a relatively strong Fresnel fringe remained which cannot be explained by the Fourier cut-off alone. Correction for 5 6% amplitude contrast according to [12] reduces, but does not eliminate, the Fresnel fringe. The remaining fringes are likely to be due to wrong correction at very low spatial frequencies, at which frequencies the expected contrast at all defocus values is small.
5 Research Article Ordered regions of RNA in TYMV Böttcher and Crowther 391 Figure 6 Figure 7 Assessment of the effective resolution of the maps. Weighted phase residual (right-hand scale, light blue and yellow traces), and Fourier shell correlation coefficient (left-hand scale, red and dark blue traces) plotted against Fourier spacing for TYMV (light and dark blue) and empty capsid (yellow and red). The significance level for the Fourier shell correlation coefficient is indicated by a dotted grey line. Radial density distributions in the three-dimensional maps. (a) TYMV. (b) Empty capsid. (c) The difference plot between virus and empty capsid density. Resolution The effective resolution was determined by dividing the particles into two sets (for both the virus and the empty capsid), computing two independent three-dimensional maps of each, and determining the Fourier shell cross correlation and phase residuals [13]. The phase-residual plots and the Fourier shell cross-correlation plots are approximately mirror related and reach the 85 phase residual and significance level respectively at about the same Fourier spacing (Fig. 7), as found in a recent comparison of the two measures [14]. Both methods to determine resolution basically yield the same result. In the maps used for difference imaging, a Fourier cut-off of 1/1.5 nm 1 was applied. At this Fourier spacing, both maps had a weighted phase residual of less than 70 giving quite reliable information. The resolution of the map of empty capsids is worse than that of the complete virus particle (Fig. 7). This difference probably reflects the smaller number of particles included in the empty capsid map rather than disruption of the capsid organization. Indeed, if the number of particles in the virus reconstruction is reduced to that in the empty capsid reconstruction, almost the same plots are obtained for the virus as for empty capsid (data not shown). Thus, the resolution is strongly dependent on the number of particles included in the reconstruction. The TMV particles in the same micrograph show a clear sixth layerline (spacing 1/1.15 nm 1 ) in optical diffraction, which indicates that the micrographs should be suitable for determining the structure of TYMV to at least 1/1.15 nm 1, if a sufficient number of particles is included. Discussion Protein organization in the capsid The maps computed from the unstained TYMV particles show the same external hexamer pentamer clustering as was found in negatively stained specimens by model building [1] or by computation [15]. However the use of unstained specimens permits the visualization of internal structures, in this case the inner dimer clustering of the protein subunits and a set of features that we believe, on the basis of differences between virus particles and empty capsids, correspond to RNA. Intact virus and empty capsids have the same maximal outer dimensions of nm, in good agreement with a previous cryo-electron microscopy study [3]. The knobs at the fivefold axes extend slightly further than those at the threefold axes, as was found in a map computed from negatively stained specimens [15]. The apparent radii of TYMV and its empty capsid were determined by photon correlation spectroscopy [8]: the complete virus yielded an average radius of 14.6 nm, somewhat smaller than the maximal dimensions reported here. Upon alkaline treatment, TYMV loses about six
6 392 Structure 1996, Vol 4 No 4 subunits [10] and swells by about 4% [8] before complete release of its RNA. Upon return to ph 7.0, the average radius decreases again to 14.6 nm [8]. TYMV also loses between five and nine subunits and releases its RNA upon freeze-thawing [9]. It is likely that the mechanisms in both cases (alkaline treatment and freeze-thawing) are similar. It is therefore expected that viruses and empty capsids have the same outer dimensions under the imaging conditions chosen, in agreement with our observations here. No swelling of the virus is apparent at 100 K, in contrast to that observed by neutron small-angle scattering of bulk-frozen TYMV at 80 K [4]. At outer radii (13 15 nm), the subunits interact strongly to form hexamer and pentamer units. At smaller radii ( nm), the subunits join to form a more continuous shell of density protecting the RNA. On the inner surface of the capsid the subunits are disposed in a dimerclustered manner, leaving large holes around the fivefold and threefold axes (Fig. 3, bottom row, 11 nm radius). The details of subunit packing support the idea that it is pentamer units that are lost on freeze-thawing. Four subunits from neighbouring hexamer units (two from each) seem to interlock across the strict twofold axis at a radius of nm to form a closely packed feature, whereas the subunits from the pentamers stay separate (Fig. 4b). Thus, except for interaction at internal radii, the pentamer units are clearly separated from the frame composed of hexamerically arranged subunits. It is therefore likely that upon freeze-thawing a knob at one of the fivefold symmetry axes is lost rather than one from one of the threefold axes. This is supported by the finding, that relative to the hexamers, the protein density of the pentamers in the empty capsids at radii between 13.5 nm and 15 nm is 86% of that observed in the virus. This corresponds to an average loss of about eight subunits, which is in agreement with [9]. This indicates that, on average, the virus loses at least one pentamer morphological unit and that it is unlikely that any loss of knobs occurs at the threefold axes. A similar loss of subunits occurs during uncoating in vivo [11]. Difference imaging shows ordered RNA The difference plot of the radial density distributions between viruses and empty capsids shows no substantial amount of density at radii greater than 11 nm, which is in agreement with earlier small-angle neutron scattering data collected from viruses at room temperature, for which no RNA could be found at radii greater than nm [4,5]. The strongest features in the difference map between the virus and the empty capsid are hollow triangles of density centred on each of the threefold axes. These triangles of density, which we ascribe to segments of ordered RNA, extend outwards to partly fill the threefold holes on the inside of the capsid. The overall length of the edge of the triangle is about 5.5 nm and the thickness about 2 nm. At a radius of 11 nm each triangle of density appears to make close contact with one subunit in each of the three neighbouring strict dimers and a glancing contact with one subunit in each of the three neighbouring local dimers (Fig. 4a). At a larger radius ( nm) there are radial bridges of density to the inner surface of the capsid (Fig. 5). These may be RNA or perhaps loops of protein interacting with the RNA, which are ordered in the virus but disordered in the empty shell. The difference map also shows a weaker feature on the fivefold axes. While this might represent residual noise in the map, there is good reason to think that it arises from partly ordered RNA. The hole on the fivefold axes is only 3.5 nm in diameter, which could accommodate only one double-helical RNA stem. The contacts made by the triangles of RNA at the threefold axes leave one protein subunit in each local dimer unsatisfied, with the equivalent contact surface pointing towards the fivefold axis. If one out of five of these sites around the fivefold axes were occupied on a random basis, averaging would give rise to a weak feature on the fivefold axes. Our observations agree with the general model of Klug et al. [6], who proposed that RNA bunches penetrate the capsid at the fivefold and threefold axes to an effective outer radius of 12.5 nm. This coincides with the radius of the radial bridges of density at the threefold axes in our map. However, the ordered RNA only touches the protein and is clearly not embedded in the protein capsid (Fig. 5), as proposed earlier [6]. The contact occurs at two distinct sites on the protein subunit, one at a smaller radius of about 11 nm where the hollow RNA triangles make a sliding contact to the surrounding protein subunits at the strict and local twofold axes, and the other at a radius of about nm in the form of radial bridges. These two sites might correspond to the two regions in the protein subunit that were identified by cross-linking experiments [7]. Biological implications The capsid of turnip yellow mosaic virus (TYMV) consists of 180 copies of a single coat protein that forms a spherical shell around the 6.3 kb genomic RNA. The coat protein extends from an inner radius of about 11 nm to an outer radius of about 15.5 nm. At outer radii, the subunits are clustered into 12 pentamer and 20 hexamer morphological units, which lie at the icosahedral fivefold and threefold axes respectively. TYMV is a good model for studying protein RNA interactions in viruses, because cross-linking experiments have demonstrated that specific contacts occur between RNA and protein. By comparing empty capsids (produced by freeze-thawing) with intact
7 Research Article Ordered regions of RNA in TYMV Böttcher and Crowther 393 virus particles, we have shown that at the threefold axes a hollow triangle of RNA makes two contacts to each of the surrounding subunits but does not otherwise penetrate the capsid protein. The dimensions of the RNA triangle are sufficient to accommodate a double-helical RNA rod at each side, oriented tangentially to the inner surface of the capsid. A weaker feature seen at the fivefold axes could be a partially ordered double-helical RNA rod arranged perpendicular to the surface of the capsid. The capsid is formed of only one type of subunit. These subunits must be arranged in such a way that a stable spherical shell can be maintained whilst permitting release of RNA during infection. The release appears to occur by loss of some protein subunits. (In vitro, freeze-thawing or alkali treatment initiates a similar protein loss.) Different kinds of interactions between subunits are probably responsible for allowing these apparently conflicting requirements to be met. At outer radii, hexamer pentamer clustering occurs but at inner radii the subunits are dimerclustered across the strict and local twofold axes. In between these radial extremes the subunits from the pentamers are clearly separated from the subunits of the hexamers, with the latter forming an interlocked arrangement of four subunits across the strict twofold axes, involving two subunits from each hexamer. This shows that a single type of subunit is able to make geometrically very different contacts with neighbouring subunits. From the observed arrangement of the subunits and the relative density distribution in the empty capsid and the virus, it appears that after freeze-thawing one or two pentamers are lost. In contrast, the subunits forming the hexamers at outer radii build a very stable framework at inner radii which maintains the integrity of the spherical shell. Presumably this provides a mechanism whereby a stable capsid can, under relatively mild conditions, release the RNA during infection. Materials and methods Sample preparation Before freezing, TYMV was mixed with TMV to act as an internal calibration. Virus concentrations in the mixed sample were about mg ml 1 for each virus. Samples were frozen according to [16] with a freezing device as described in [17]. The device was placed in a cold room and the chamber was humidified by two water-soaked sponges to prevent evaporation from the sample. The grids were glow discharged in air and used within 1 h. The sample (2 l) was applied onto a 400 mesh copper-rhodium grid (Agar Scientific, Stansted, UK) coated with a perforated carbon film. Some of the grids were covered with an additional very thin carbon film to reduce charging and allow thinner films of ice. Most of the sample was removed by blotting with two layers of filter paper (Whatman, Maidstone, UK) for s. Immediately after blotting, the grid was plunged into liquid ethane by a spring-driven device. The ethane was prevented from freezing by a heater. Samples were also prepared without the heater. No difference with respect to the vitrification of the water was observed. Empty capsids were prepared according to [3]. The virus suspension (2 l) was applied to a grid, which was dipped into liquid nitrogen until the solution was completely frozen. The sample was then allowed to thaw on the grid at 4 C, after which the grid was prepared as described above. Electron microscopy Micrographs were taken under low-dose conditions at nominal magnifications of and at a Philips EM420 (Philips Electron Optics, Eindhoven, The Netherlands) operating at 120 kv or at at a Hitachi HF2000 (Hitachi Ltd., Tokyo, Japan) microscope operating at 200 kv which was equipped with a field emission filament. At both microscopes Gatan cryo stages (Gatan Inc., Pleasanton, CA) were used. A temperature between 95 K and 100 K was maintained. Images were recorded on Kodak SO163 film (Eastman Kodak Company, Rochester, NY), which was developed for 12 min in full strength Kodak D19 developer. Image processing TMV particles were scanned with a modified Joyce Loebl densitometer (Joyce Loebl, Gateshead, UK) to judge the quality of the micrographs. A step size of 10 m was used corresponding to approximately 0.17 nm per pixel at specimen level. Parts of the TMV were boxed with a narrow box with a length between 100 nm and 150 nm. After background subtraction, a Fourier transformation was computed. From the transform the intensities of the third and sixth layer lines were determined. The intensities were corrected for the contrast transfer function (ctf) and the envelope function due to spatial coherence [18] and used to judge the quality of the micrographs used. TYMV particles were scanned using either a Nikon or a Joyce Loebl densitometer. A step size of 20 m/pixel was used. For calibration of the magnification, TMV on the same image was scanned using the same settings of the densitometer. A Fourier transform was computed of the TMV. From the positions of the third and sixth layerlines (1/2.3 nm 1 and 1/1.15 nm 1 ) the magnification was calculated and particles from different images scaled respectively. The centres of the TYMV particles were first determined by cross correlation to a Gaussian ring with the same diameter as the virus and the width of the ring roughly corresponding to the width of the capsid. Subsequently, the origins of new particles were determined by cross correlation to a sum of eight different projections of an existing map. The particles were boxed with a circular box with a radius of 20.8 nm and the pre-determined centre. The background was subtracted and the grey value distribution was normalized. For the first reconstruction, focus pairs taken at magnification and a defocus of about 1000 nm and 2000 nm at the Philips EM420 were chosen. The orientations of the higher defocused particles were determined using selfcommon lines [19]. The orientations found were then used for images of smaller defocus. The reliability of the orientations was judged by computation of a low-resolution two-particle three-dimensional reconstruction between a reference particle and each other particle in the data set [20]. The preservation of the threefold symmetry in the projections along the threefold symmetry axis was used as the main criterion for accepting an orientation. Then a map was calculated from these particles. Further refinement of the origin and angles of view was performed using crosscommon lines to three projections of the calculated map [20]. With the current set of programs for making three-dimensional maps of icosahedral viruses, it is difficult to incorporate proper ctf corrections for combining data from micrographs taken at different defocuses. We have partly overcome this difficulty by computing separate three-dimensional maps from images at each particular defocus and by then combining these three-dimensional maps with appropriate corrections for the defocus, although this approach does not deal with astigmatism. The correction is done by computing the three-dimensional Fourier transform of each map and then estimating the true value of each frequency component by a weighted combination of the observed
8 394 Structure 1996, Vol 4 No 4 values of that component. The contrast transfer function, c, incorporating amplitude and phase contrast is of the form: c = a cos + (1 a 2 ) sin, where a is the fractional amplitude contrast and χ is the phase shift. If F i is the observed value of a particular frequency component in the transform of the i th map, then the true value of this component F is estimated by: c i i w i F i F = i c i2 w i F i + f Here c i is the value of the ctf at this frequency in the i th transform and w i is an overall fractional weight ( w i =1) for the i th map which reflects its reliability (e.g. the number of particles included). The term f, akin to the signal-to-noise ratio in a Wiener filter, prevents over amplification of terms when the rest of the denominator is small. In our case where we combine images with several different defocuses, f has its principal effect at very low spatial frequencies, where it sets the average density level in the final three-dimensional map computed by back transformation. f was adjusted to give an average radial density value of zero inside and outside the empty capsids. The same filter was used for the corresponding virus reconstructions. For the amplitude contrast, a value of 5% was used for images from the Hitachi. When using projections of the computed map as a basis for determining or refining the orientation parameters of particles, it is important for the computed projections to have a defocus applied that matches that of the image. This can be done, as above, by using the three-dimensional transform but in this case simply multiplying by the appropriate ctf and back-transforming, before computing the projections. Existing maps were used as references to find the orientations of particles in micrographs taken under different conditions. For example, the map of the images taken at the Philips EM420 at was scaled and then used as a reference to find the first orientations of the particles from micrographs taken at the Hitachi HF2000 at 200 kv and a magnification of The resulting orientations were used to calculate a preliminary map from these particles. For further refinement of the orientations, only maps resulting from micrographs taken at the Hitachi microscope were used as reference. Difference maps Reconstructions were calculated for empty capsids and the complete virus to the same resolution. In order to determine the density scaling of the virus relative to the empty capsid, it was necessary to determine whether subunits had been lost from the empty capsid. This was done by integrating the density over radii greater than 13 nm in a pentamer unit and a hexamer unit in the maps of both the virus and the empty capsid. The ratio of pentamer density to hexamer density was lower in the empty capsid than the virus indicating that pentamer units had been lost in the empty capsid. The density of the empty capsid reconstruction was first scaled to give the same integral peak for the radial density distribution for radii between 11.5 nm and 15.5 nm. Before subtraction from the virus map the density of the empty capsid map was weighted by a factor of 1.05 to adjust for the subunit loss. The features in the difference map were not strongly dependent on the precise way in which the scaling was done. References 1. Finch, J.T. & Klug, A. (1966). Arrangement of protein subunits and the distribution of nucleic acid in turnip yellow mosaic virus: II. Electron microscopic studies. J. Mol. Biol. 15, Schmidt, P., Kaesberg, P. & Beeman, W.W. (1954). Small-angle X-ray scattering from turnip yellow mosaic virus. Biochim. Biophys. Acta 14, Adrian, M., Timmins, P.A. & Witz, J. (1992). In vivo decapsidation of turnip yellow mosaic virus investigated by cryo-electron microscopy: a model for the decapsidation of a small isometric virus. J. Gen. Virol. 73, Witz, J., Timmins, P.A. & Adrian, M. (1993). Organization of turnip yellow mosaic virus investigated by neutron small-angle scattering at 80 K: an intermediate state preceding decapsidation of the virion? Proteins 17, Jacrot, B., Chauvin, C. & Witz, J. (1977). Comparative neutron smallangle scattering study of small spherical RNA viruses. Nature 266, Klug, A., Longley, W. & Leberman, R. (1966). Arrangement of protein subunits and the distribution of nucleic acid in turnip yellow mosaic virus: I. X-ray diffraction studies. J. Mol. Biol. 15, Ehresmann, B., Briand, J.-P., Reinbolt, J. & Witz, J. (1980). Identification of binding sites of turnip yellow mosaic virus protein and RNA by crosslinks induced in situ. Eur. J. Biochem. 108, Keeling, J., Collins, E.R. & Matthews, R.E.F. (1979). Behaviour of turnip yellow mosaic virus nucleoproteins under alkaline conditions. Virology 97, Katouzian-Safadi, M., Berthet-Colominas, C., Witz, J. & Kruse, J. (1983). Evidence for the presence of a hole in the capsid of turnip yellow mosaic virus after RNA release by freezing and thawing: decapsidation of turnip yellow mosaic virus in vitro. Eur. J. Biochem. 137, Keeling, J. & Matthews, R.E.F. (1982). Mechanism for release of RNA from turnip yellow mosaic virus at high ph. Virology 119, Matthews, R.E.F. & Witz, J. (1985). Uncoating of turnip yellow mosaic virus RNA in vivo. Virology 144, Toyoshima, C., Yonekura, K. & Sasabe, H. (1993). Contrast transfer for frozen-hydrated specimens II. Amplitude contrast at very low frequencies. Ultramicroscopy 48, van Heel, M. (1987). Similarity measures between images. Ultramicroscopy 21, de la Fraga, L.G., Dopazo, J. & Carazo, J.M. (1995). Confidence limits for resolution estimation in image averaging by random subsampling. Ultramicroscopy 60, Mellema, J.E. & Amos, L.A. (1972). Three-dimensional image reconstruction of turnip yellow mosaic virus. J. Mol. Biol. 72, Dubochet, J., et al., & Schultz, P. (1988). Cryo-electron microscopy of vitrified specimens. Quart. Rev. Biophys. 21, Bellare, J.R., Davis, H.T., Scriven, L.E. & Talmon, Y. (1988). Controlled environment vitrification system: an improved sample preparation technique. J. Electron Microsc. Tech. 10, Reimer L. (1993). Transmission Electron Microscopy. pp , Springer Verlag, Berlin and Heidelberg. 19. Crowther, R.A. (1971). Procedures for three-dimensional reconstruction of spherical viruses by Fourier synthesis from electron micrographs. Philos. Trans. R. Soc. (Lond) B 261, Crowther, R.A., et al., & Pumpens, P. (1994). Three-dimensional structure of hepatitis B virus core particles determined by electron cryomicroscopy. Cell 77, Cyrklaff, M. & Kühlbrandt, W. (1994). High-resolution electron microscopy of biological specimens in cubic ice. Ultramicroscopy 55, Resolution For an estimate of resolution, data sets were divided randomly into two equal subsets. From each subset a map was calculated. Fourier shell correlation coefficients and weighted phase residuals were calculated between maps according to [13] (equations 3 and 6, respectively). Acknowledgements We thank Dr PJG Butler for providing TMV and TYMV and Dr JT Finch for comments on the manuscript. BB was supported by an EMBO long term fellowship (ALTF ) and an EEC human capital and mobility grant (ERB4001GT931348).
Science Supporting Online Material.
 - S1 - Science Supporting Online Material. Grünewald et al., Three-dimensional Structure of Herpes Simplex Virus from Cryoelectron Tomography. Materials and Methods Strains, cultivation and purification
- S1 - Science Supporting Online Material. Grünewald et al., Three-dimensional Structure of Herpes Simplex Virus from Cryoelectron Tomography. Materials and Methods Strains, cultivation and purification
Proteins and symmetry
 Proteins and symmetry Viruses (symmetry) Viruses come in many shapes, sizes and compositions All carry genomic nucleic acid (RNA or DNA) Structurally and genetically the simplest are the spherical viruses
Proteins and symmetry Viruses (symmetry) Viruses come in many shapes, sizes and compositions All carry genomic nucleic acid (RNA or DNA) Structurally and genetically the simplest are the spherical viruses
Structural biology of viruses
 Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Ultrastructure of Mycoplasmatales Virus laidlawii x
 J. gen. Virol. (1972), I6, 215-22I Printed in Great Britain 2I 5 Ultrastructure of Mycoplasmatales Virus laidlawii x By JUDY BRUCE, R. N. GOURLAY, AND D. J. GARWES R. HULL* Agricultural Research Council,
J. gen. Virol. (1972), I6, 215-22I Printed in Great Britain 2I 5 Ultrastructure of Mycoplasmatales Virus laidlawii x By JUDY BRUCE, R. N. GOURLAY, AND D. J. GARWES R. HULL* Agricultural Research Council,
capsomeres: direct evidence from cryo-electron-microscopy
 The capsid of small papova viruses contains 72 pentameric capsomeres: direct evidence from cryo-electron-microscopy of simian virus 40 Timothy S. Baker,* Jacqueline Drak,* and Minou Bina* *Department of
The capsid of small papova viruses contains 72 pentameric capsomeres: direct evidence from cryo-electron-microscopy of simian virus 40 Timothy S. Baker,* Jacqueline Drak,* and Minou Bina* *Department of
Virology. *Viruses can be only observed by electron microscope never by light microscope. The size of the virus: nm in diameter.
 Virology We are going to start with general introduction about viruses, they are everywhere around us; in food; within the environment; in direct contact to etc.. They may cause viral infection by itself
Virology We are going to start with general introduction about viruses, they are everywhere around us; in food; within the environment; in direct contact to etc.. They may cause viral infection by itself
STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES
 STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES Introduction Viruses are noncellular genetic elements that use a living cell for their replication and have an extracellular state. Viruses
STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES Introduction Viruses are noncellular genetic elements that use a living cell for their replication and have an extracellular state. Viruses
The protein stoichiometry of viral capsids via tiling theory
 The protein stoichiometry of viral capsids via tiling theory REIDUN TWAROCK Centre for Mathematical Science City University Northampton Square, London EC1V 0HB UNITED KINGDOM Abstract: - A vital part of
The protein stoichiometry of viral capsids via tiling theory REIDUN TWAROCK Centre for Mathematical Science City University Northampton Square, London EC1V 0HB UNITED KINGDOM Abstract: - A vital part of
Virus Structure. Characteristics of capsids. Virus envelopes. Virion assembly John Wiley & Sons, Inc. All rights reserved.
 Virus Structure Characteristics of capsids Virus envelopes Virion assembly Capsids package viral genomes and transmit them to a new host cell Capsid rigid, symmetrical container composed of viral protein
Virus Structure Characteristics of capsids Virus envelopes Virion assembly Capsids package viral genomes and transmit them to a new host cell Capsid rigid, symmetrical container composed of viral protein
Overview of virus life cycle
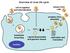 Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
PHARMACEUTICAL MICROBIOLOGY JIGAR SHAH INSTITUTE OF PHARMACY NIRMA UNIVERSITY
 PHARMACEUTICAL MICROBIOLOGY JIGAR SHAH INSTITUTE OF PHARMACY NIRMA UNIVERSITY VIRUS - HISTORY In 1886, the Dutch Chemist Adolf Mayer showed TMD In 1892, the Russian Bactriologist Dimtri Iwanowski isolate
PHARMACEUTICAL MICROBIOLOGY JIGAR SHAH INSTITUTE OF PHARMACY NIRMA UNIVERSITY VIRUS - HISTORY In 1886, the Dutch Chemist Adolf Mayer showed TMD In 1892, the Russian Bactriologist Dimtri Iwanowski isolate
Composition & Architecture of plant viruses. P.N. Sharma Department of Plant Pathology, CSK HPKV, Palampur (H.P.)
 Composition & Architecture of plant viruses P.N. Sharma Department of Plant Pathology, CSK HPKV, Palampur (H.P.) Plant Viruses Classification, Morphology, Genome, and Structure Importance Detailed knowledge
Composition & Architecture of plant viruses P.N. Sharma Department of Plant Pathology, CSK HPKV, Palampur (H.P.) Plant Viruses Classification, Morphology, Genome, and Structure Importance Detailed knowledge
An Externalized Polypeptide Partitions between Two Distinct Sites on Genome-Released Poliovirus Particles
 REFERENCES CONTENT ALERTS An Externalized Polypeptide Partitions between Two Distinct Sites on Genome-Released Poliovirus Particles Jun Lin, Naiqian Cheng, Marie Chow, David J. Filman, Alasdair C. Steven,
REFERENCES CONTENT ALERTS An Externalized Polypeptide Partitions between Two Distinct Sites on Genome-Released Poliovirus Particles Jun Lin, Naiqian Cheng, Marie Chow, David J. Filman, Alasdair C. Steven,
Viral structure م.م رنا مشعل
 Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Structure of viruses
 Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2018 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2018 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Structure of viruses
 Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2017 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2017 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Structural vs. nonstructural proteins
 Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Supplementary Figures
 Supplementary Figures Spatial arrangement Variation in the morphology of central NCs (shape x size) x Variation in the morphology of satellite NCs (shape x size) x Variations in the spatial arrangement
Supplementary Figures Spatial arrangement Variation in the morphology of central NCs (shape x size) x Variation in the morphology of satellite NCs (shape x size) x Variations in the spatial arrangement
Dr. Gary Mumaugh. Viruses
 Dr. Gary Mumaugh Viruses Viruses in History In 1898, Friedrich Loeffler and Paul Frosch found evidence that the cause of foot-and-mouth disease in livestock was an infectious particle smaller than any
Dr. Gary Mumaugh Viruses Viruses in History In 1898, Friedrich Loeffler and Paul Frosch found evidence that the cause of foot-and-mouth disease in livestock was an infectious particle smaller than any
General Virology I. Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department
 General Virology I Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department ١ General Virology I Lecture Outline Introduction istory Definition
General Virology I Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department ١ General Virology I Lecture Outline Introduction istory Definition
CONTENTS. 1. Introduction. 4. Virology. 2. Virus Structure. 5. Virus and Medicine. 3. Virus Replication. 6. Review
 CONTENTS 1. Introduction 4. Virology 2. Virus Structure 5. Virus and Medicine 3. Virus Replication 6. Review We have all gotten viruses from bacteria, plants to animals. Viruses cause colds, flu, warts
CONTENTS 1. Introduction 4. Virology 2. Virus Structure 5. Virus and Medicine 3. Virus Replication 6. Review We have all gotten viruses from bacteria, plants to animals. Viruses cause colds, flu, warts
Chapter 6- An Introduction to Viruses*
 Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
ELECTRON MICROSCOPIC STUDIES ON EQUINE ENCEPHALOSIS VIRUS
 Onderstepoort]. vet. Res. 40 (2), 53-58 (1973) ELECTRON MICROSCOPIC STUDIES ON EQUINE ENCEPHALOSIS VIRUS G. LECATSAS, B. J. ERASMUS and H. J. ELS, Veterinary Research Institute, Onderstepoort ABSTRACT
Onderstepoort]. vet. Res. 40 (2), 53-58 (1973) ELECTRON MICROSCOPIC STUDIES ON EQUINE ENCEPHALOSIS VIRUS G. LECATSAS, B. J. ERASMUS and H. J. ELS, Veterinary Research Institute, Onderstepoort ABSTRACT
Supplementary Figure 1. Sample preparation schematic. First (Stage I), square islands of MoO 3 are prepared by either photolithography followed by
 Supplementary Figure 1. Sample preparation schematic. First (Stage I), square islands of MoO 3 are prepared by either photolithography followed by thermal evaporation and liftoff or by a process where
Supplementary Figure 1. Sample preparation schematic. First (Stage I), square islands of MoO 3 are prepared by either photolithography followed by thermal evaporation and liftoff or by a process where
Introductory Virology. Ibrahim Jamfaru School of Medicine UHAS
 Introductory Virology Ibrahim Jamfaru School of Medicine UHAS Lecture outline Definition of viruses and general characteristics Structure of virus (virion) Chemical composition of viruses Virus morphology
Introductory Virology Ibrahim Jamfaru School of Medicine UHAS Lecture outline Definition of viruses and general characteristics Structure of virus (virion) Chemical composition of viruses Virus morphology
Sindbis Virus Glycoproteins Form a Regular Icosahedral Surface Lattice
 JOURNAL OF VIROLOGY, July 1975, p. 141-145 Copyright i 1975 American Society for Microbiology Vol. 16, No. 1 Printed in U.S.A. Sindbis Virus Glycoproteins Form a Regular Icosahedral Surface Lattice C.-H.
JOURNAL OF VIROLOGY, July 1975, p. 141-145 Copyright i 1975 American Society for Microbiology Vol. 16, No. 1 Printed in U.S.A. Sindbis Virus Glycoproteins Form a Regular Icosahedral Surface Lattice C.-H.
Reviewers' Comments: Reviewer #2: Remarks to the Author:
 Reviewers' Comments: Reviewer #1: Remarks to the Author: This manuscript presents cryo-em studies of a giant virus Paramecium bursaria chlorella virus 1 (PBCV-1), which is the first cryo-em structure of
Reviewers' Comments: Reviewer #1: Remarks to the Author: This manuscript presents cryo-em studies of a giant virus Paramecium bursaria chlorella virus 1 (PBCV-1), which is the first cryo-em structure of
The Structure of Viruses of the Newcastle Disease- Mumps-Influenza (Myxovirus) Group
 680 * VALENTINE, R. C. & ISAACS, A. (1957). J. gen. Microbiol. 16, 680-685 The Structure of Viruses of the Newcastle Disease- Mumps-Influenza (Myxovirus) Group BY R. C. VALENTINE AND A. IsAAcS National
680 * VALENTINE, R. C. & ISAACS, A. (1957). J. gen. Microbiol. 16, 680-685 The Structure of Viruses of the Newcastle Disease- Mumps-Influenza (Myxovirus) Group BY R. C. VALENTINE AND A. IsAAcS National
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
علم األحياء الدقيقة Microbiology Introduction to Virology & Immunology
 علم األحياء الدقيقة Microbiology Introduction to Virology & Immunology What is a virus? Viruses may be defined as acellular organisms whose genomes consist of nucleic acid (DNA or RNA), and which obligatory
علم األحياء الدقيقة Microbiology Introduction to Virology & Immunology What is a virus? Viruses may be defined as acellular organisms whose genomes consist of nucleic acid (DNA or RNA), and which obligatory
Three-dimensional structure of a membrane-containing virus
 Proc. Natl. Acad. Sci. USA Vol. 90, pp. 9095-9099, October 1993 Microbiology Three-dimensional structure of a membrane-containing virus ANGEL M. PAREDES*, DENNIS T. BROWN*t, ROSALBA ROTHNAGELt, WAH CHIU*,
Proc. Natl. Acad. Sci. USA Vol. 90, pp. 9095-9099, October 1993 Microbiology Three-dimensional structure of a membrane-containing virus ANGEL M. PAREDES*, DENNIS T. BROWN*t, ROSALBA ROTHNAGELt, WAH CHIU*,
Lecture 5 (Ch6) - Viruses. Virus Characteristics. Viral Host Range
 Lecture 5 (Ch6) - Viruses Topics Characteristics Structure/Classification Multiplication Cultivation and replication Non-viral infectious agents Treatment 1 Virus Characteristics obligate intracellular
Lecture 5 (Ch6) - Viruses Topics Characteristics Structure/Classification Multiplication Cultivation and replication Non-viral infectious agents Treatment 1 Virus Characteristics obligate intracellular
Cell Overview. Hanan Jafar BDS.MSc.PhD
 Cell Overview Hanan Jafar BDS.MSc.PhD THE CELL is made of: 1- Nucleus 2- Cell Membrane 3- Cytoplasm THE CELL Formed of: 1. Nuclear envelope 2. Chromatin 3. Nucleolus 4. Nucleoplasm (nuclear matrix) NUCLEUS
Cell Overview Hanan Jafar BDS.MSc.PhD THE CELL is made of: 1- Nucleus 2- Cell Membrane 3- Cytoplasm THE CELL Formed of: 1. Nuclear envelope 2. Chromatin 3. Nucleolus 4. Nucleoplasm (nuclear matrix) NUCLEUS
1.4 Page 1 Cell Membranes S. Preston 1
 AS Unit 1: Basic Biochemistry and Cell Organisation Name: Date: Topic 1.3 Cell Membranes and Transport Page 1 1.3 Cell Membranes and Transport from your syllabus l. Cell Membrane Structure 1. Read and
AS Unit 1: Basic Biochemistry and Cell Organisation Name: Date: Topic 1.3 Cell Membranes and Transport Page 1 1.3 Cell Membranes and Transport from your syllabus l. Cell Membrane Structure 1. Read and
Cryo-electron microscopy of hepatitis B virions reveals variability in envelope capsid interactions
 The EMBO Journal (2007) 26, 4160 4167 & 2007 European Molecular Biology Organization All Rights Reserved 0261-4189/07 www.embojournal.org Cryo-electron microscopy of hepatitis B virions reveals variability
The EMBO Journal (2007) 26, 4160 4167 & 2007 European Molecular Biology Organization All Rights Reserved 0261-4189/07 www.embojournal.org Cryo-electron microscopy of hepatitis B virions reveals variability
Observations on the Geometric Distribution of Micro-structural Features in Cortical Bone
 Observations on the Geometric Distribution of Micro-structural Features in Cortical Bone I. S. Hage, R. F. Hamade Abstract This paper reports on observations of the distribution of micro-structural features
Observations on the Geometric Distribution of Micro-structural Features in Cortical Bone I. S. Hage, R. F. Hamade Abstract This paper reports on observations of the distribution of micro-structural features
Nature Neuroscience doi: /nn Supplementary Figure 1. Characterization of viral injections.
 Supplementary Figure 1 Characterization of viral injections. (a) Dorsal view of a mouse brain (dashed white outline) after receiving a large, unilateral thalamic injection (~100 nl); demonstrating that
Supplementary Figure 1 Characterization of viral injections. (a) Dorsal view of a mouse brain (dashed white outline) after receiving a large, unilateral thalamic injection (~100 nl); demonstrating that
Herpes simplex virus L particles contain spherical membrane-enclosed inclusion vesicles
 Journal of General Virology (1994), 75, 1749-1753. Printed in Great Britain 1749 Herpes simplex virus L particles contain spherical membrane-enclosed inclusion vesicles J. F. Szildgyi 1. and J. Berriman
Journal of General Virology (1994), 75, 1749-1753. Printed in Great Britain 1749 Herpes simplex virus L particles contain spherical membrane-enclosed inclusion vesicles J. F. Szildgyi 1. and J. Berriman
BIOLOGY 3A LABORATORY Morphology and the shape and form of biological structures
 BIOLOGY 3A LABORATORY Morphology and the shape and form of biological structures For the harmony of the world is made manifest in form and number, and the heart and soul and all the poetry of Natural philosophy
BIOLOGY 3A LABORATORY Morphology and the shape and form of biological structures For the harmony of the world is made manifest in form and number, and the heart and soul and all the poetry of Natural philosophy
Cryo-electron microscopy reconstruction shows poliovirus 135S. particles poised for membrane interaction and RNA release
 JVI Accepts, published online ahead of print on 20 November 2013 J. Virol. doi:10.1128/jvi.01949-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. Cryo-electron microscopy reconstruction
JVI Accepts, published online ahead of print on 20 November 2013 J. Virol. doi:10.1128/jvi.01949-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. Cryo-electron microscopy reconstruction
Peter A ~imrnind*, David wild2+ and Jean witz3
 The three-dimensional distribution of RNA and protein in the interior of tomato bushy stunt virus: a neutron low-resolution single-crystal diffraction study Peter A ~imrnind*, David wild2+ and Jean witz3
The three-dimensional distribution of RNA and protein in the interior of tomato bushy stunt virus: a neutron low-resolution single-crystal diffraction study Peter A ~imrnind*, David wild2+ and Jean witz3
Is action potential threshold lowest in the axon?
 Supplementary information to: Is action potential threshold lowest in the axon? Maarten H. P. Kole & Greg J. Stuart Supplementary Fig. 1 Analysis of action potential (AP) threshold criteria. (a) Example
Supplementary information to: Is action potential threshold lowest in the axon? Maarten H. P. Kole & Greg J. Stuart Supplementary Fig. 1 Analysis of action potential (AP) threshold criteria. (a) Example
PROCEDURE Follow the instructions as you proceed through the Click and Learn, and answer the questions in the spaces provided.
 ABOUT THIS WORKSHEET This worksheet complements the Click and Learn developed in conjunction with the 2016 documentary, Spillover: Zika, Ebola & Beyond (http://www.hhmi.org/biointeractive/virusexplorer).
ABOUT THIS WORKSHEET This worksheet complements the Click and Learn developed in conjunction with the 2016 documentary, Spillover: Zika, Ebola & Beyond (http://www.hhmi.org/biointeractive/virusexplorer).
Chapter 19: Viruses. 1. Viral Structure & Reproduction. What exactly is a Virus? 11/7/ Viral Structure & Reproduction. 2.
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Lab Tuesday: Virus Diseases
 Lab Tuesday: Virus Diseases Quiz for Bacterial Pathogens lab (pp 69-75) and Biocontrol of Crown Gall (p. 115-119), Observation of Viral Movement in Plants (p. 121), and Intro section for Viruses (pp. 77-79).
Lab Tuesday: Virus Diseases Quiz for Bacterial Pathogens lab (pp 69-75) and Biocontrol of Crown Gall (p. 115-119), Observation of Viral Movement in Plants (p. 121), and Intro section for Viruses (pp. 77-79).
Supplementary Figure S1 Black silicon and dragonfly wing nanotopography.
 Supplementary Figure S1 Black silicon and dragonfly wing nanotopography. Representative low-magnification scanning electron micrographs of a) Black silicon (bsi) and b) Diplacodes bipunctata dragonfly
Supplementary Figure S1 Black silicon and dragonfly wing nanotopography. Representative low-magnification scanning electron micrographs of a) Black silicon (bsi) and b) Diplacodes bipunctata dragonfly
Viruses 101., and concluded that living organisms do not crystallize. In other words,.
 Viruses 101 In 1897, Dutch scientist called tiny particles in the liquid extracted from a plant disease, which is the Latin word for. In 1935, American biochemist isolated crystals of, and concluded that
Viruses 101 In 1897, Dutch scientist called tiny particles in the liquid extracted from a plant disease, which is the Latin word for. In 1935, American biochemist isolated crystals of, and concluded that
Chapter 19: Viruses. 1. Viral Structure & Reproduction. 2. Bacteriophages. 3. Animal Viruses. 4. Viroids & Prions
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Coronaviruses cause acute, mild upper respiratory infection (common cold).
 Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
Introduction. The goal of TRUS QA is to ensure your system can do all of this accurately.
 TRUS QA Workshop Introduction The goals of using TRUS in prostate brachytherapy Visualize the prostate Need the US to penetrate deeply enough Need sufficient grey scale resolution to be able to visualize
TRUS QA Workshop Introduction The goals of using TRUS in prostate brachytherapy Visualize the prostate Need the US to penetrate deeply enough Need sufficient grey scale resolution to be able to visualize
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Electron Cryomicroscopy and Computer Image Processing Techniques
 Cryo-EM and Computer Image Processing 9 2 Electron Cryomicroscopy and Computer Image Processing Techniques Use in Structure Function Studies of Rotavirus B. V. Venkataram Prasad and Mary K. Estes 1. Introduction
Cryo-EM and Computer Image Processing 9 2 Electron Cryomicroscopy and Computer Image Processing Techniques Use in Structure Function Studies of Rotavirus B. V. Venkataram Prasad and Mary K. Estes 1. Introduction
9700 BIOLOGY. Mark schemes should be read in conjunction with the question paper and the Principal Examiner Report for Teachers.
 CAMBRIDGE INTERNATIONAL EXAMINATIONS GCE Advanced Subsidiary Level and GCE Advanced Level MARK SCHEME for the May/June 2013 series 9700 BIOLOGY 9700/31 Paper 31 (Advanced Practical Skills 1), maximum raw
CAMBRIDGE INTERNATIONAL EXAMINATIONS GCE Advanced Subsidiary Level and GCE Advanced Level MARK SCHEME for the May/June 2013 series 9700 BIOLOGY 9700/31 Paper 31 (Advanced Practical Skills 1), maximum raw
Noninvasive Blood Glucose Analysis using Near Infrared Absorption Spectroscopy. Abstract
 Progress Report No. 2-3, March 31, 1999 The Home Automation and Healthcare Consortium Noninvasive Blood Glucose Analysis using Near Infrared Absorption Spectroscopy Prof. Kamal Youcef-Toumi Principal Investigator
Progress Report No. 2-3, March 31, 1999 The Home Automation and Healthcare Consortium Noninvasive Blood Glucose Analysis using Near Infrared Absorption Spectroscopy Prof. Kamal Youcef-Toumi Principal Investigator
This week s topic will be: Evidence for the Fluid Mosaic Model. Developing theories, testing hypotheses and techniques for visualizing cells
 Tutorials, while not mandatory, will allow you to improve your final grade in this course. Thank you for your attendance to date. These notes are not a substitute for the discussions that we will have
Tutorials, while not mandatory, will allow you to improve your final grade in this course. Thank you for your attendance to date. These notes are not a substitute for the discussions that we will have
Sum of Neurally Distinct Stimulus- and Task-Related Components.
 SUPPLEMENTARY MATERIAL for Cardoso et al. 22 The Neuroimaging Signal is a Linear Sum of Neurally Distinct Stimulus- and Task-Related Components. : Appendix: Homogeneous Linear ( Null ) and Modified Linear
SUPPLEMENTARY MATERIAL for Cardoso et al. 22 The Neuroimaging Signal is a Linear Sum of Neurally Distinct Stimulus- and Task-Related Components. : Appendix: Homogeneous Linear ( Null ) and Modified Linear
Three-dimensional Structure of Rhesus Rotavirus by Cryoelectron Microscopy and Image Reconstruction
 Three-dimensional Structure of Rhesus Rotavirus by Cryoelectron Microscopy and Image Reconstruction Mark Yeager,* Kelly A. Dryden,* Norman H. Olson,* Harry B. Greenberg, and Timothy S. Baker* * Scripps
Three-dimensional Structure of Rhesus Rotavirus by Cryoelectron Microscopy and Image Reconstruction Mark Yeager,* Kelly A. Dryden,* Norman H. Olson,* Harry B. Greenberg, and Timothy S. Baker* * Scripps
triplelayered unit for the plasma cell membrane component; two such Model parameters are assigned to three varieties of peripheral nerve
 STRUCTURAL PARAMETERS OF NERVE MYELIN* BY C. R. WORTHINGTON DEPARTMENT OF PHYSICS AND BIOPHYSICS RESEARCH DIVISION, UNIVERSITY OF MICHIGAN, ANN ARBOR Communicated by J. L. Oncley, April 7, 1969 Abstract.-Low-angle
STRUCTURAL PARAMETERS OF NERVE MYELIN* BY C. R. WORTHINGTON DEPARTMENT OF PHYSICS AND BIOPHYSICS RESEARCH DIVISION, UNIVERSITY OF MICHIGAN, ANN ARBOR Communicated by J. L. Oncley, April 7, 1969 Abstract.-Low-angle
Two-dimensional Electron Crystallography
 Two-dimensional Electron Crystallography Thomas Braun, University of Basel, Basel, Switzerland Andreas Engel, University of Basel, Basel, Switzerland Electron crystallography allows the structural analysis
Two-dimensional Electron Crystallography Thomas Braun, University of Basel, Basel, Switzerland Andreas Engel, University of Basel, Basel, Switzerland Electron crystallography allows the structural analysis
Detailed Syllabus. Waves and Modern Physics Spring 2011
 Detailed Syllabus Waves and Modern Physics Spring 2011 Prof. Antonio Badolato Date Lecture Topics; Chapters from Books (G = Giancoli) Homework (HW) and Recitation (R) Assignments 12 Jan, Wed Oscillations:
Detailed Syllabus Waves and Modern Physics Spring 2011 Prof. Antonio Badolato Date Lecture Topics; Chapters from Books (G = Giancoli) Homework (HW) and Recitation (R) Assignments 12 Jan, Wed Oscillations:
polyhedroses of the alfalfa caterpillar and other insects. Grote, and the alfalfa caterpillar, Colias philodice eurytheme Bdvl., were used.
 A DEMONSTRATION OF THE NATURE OF POLYHEDRA USING ALKALINE SOLUTIONS' KENNETH M. HUGHES University of California, Berkeley, California Received for publication November 1, 1949 The composition of the polyhedral
A DEMONSTRATION OF THE NATURE OF POLYHEDRA USING ALKALINE SOLUTIONS' KENNETH M. HUGHES University of California, Berkeley, California Received for publication November 1, 1949 The composition of the polyhedral
BIOL*1090 Introduction To Molecular and Cellular Biology Fall 2014
 Last time... BIOL*1090 Introduction To Molecular and Cellular Biology Fall 2014 Lecture 3 - Sept. 15, 2014 Viruses Biological Membranes Karp 7th ed: Chpt. 4; sections 4-1, 4-3 to 4-7 1 2 VIRUS Non-cellular
Last time... BIOL*1090 Introduction To Molecular and Cellular Biology Fall 2014 Lecture 3 - Sept. 15, 2014 Viruses Biological Membranes Karp 7th ed: Chpt. 4; sections 4-1, 4-3 to 4-7 1 2 VIRUS Non-cellular
Theta sequences are essential for internally generated hippocampal firing fields.
 Theta sequences are essential for internally generated hippocampal firing fields. Yingxue Wang, Sandro Romani, Brian Lustig, Anthony Leonardo, Eva Pastalkova Supplementary Materials Supplementary Modeling
Theta sequences are essential for internally generated hippocampal firing fields. Yingxue Wang, Sandro Romani, Brian Lustig, Anthony Leonardo, Eva Pastalkova Supplementary Materials Supplementary Modeling
Overview: Chapter 19 Viruses: A Borrowed Life
 Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
Nature Methods: doi: /nmeth Supplementary Figure 1. Activity in turtle dorsal cortex is sparse.
 Supplementary Figure 1 Activity in turtle dorsal cortex is sparse. a. Probability distribution of firing rates across the population (notice log scale) in our data. The range of firing rates is wide but
Supplementary Figure 1 Activity in turtle dorsal cortex is sparse. a. Probability distribution of firing rates across the population (notice log scale) in our data. The range of firing rates is wide but
18.2. Viral Structure and Reproduction. Viruses differ in shape and in ways of entering
 18.2 Viral Structure and Reproduction VOCABULARY bacteriophage lytic infection lysogenic infection prophage compare the structures of viruses to cells, describe viral reproduction, and describe the role
18.2 Viral Structure and Reproduction VOCABULARY bacteriophage lytic infection lysogenic infection prophage compare the structures of viruses to cells, describe viral reproduction, and describe the role
LECTURE 13. Dr. Teresa D. Golden University of North Texas Department of Chemistry
 LECTURE 13 Dr. Teresa D. Golden University of North Texas Department of Chemistry Goniometer circle - centered at the sample, with the x-ray source and detector on the circumference of the circle. Focusing
LECTURE 13 Dr. Teresa D. Golden University of North Texas Department of Chemistry Goniometer circle - centered at the sample, with the x-ray source and detector on the circumference of the circle. Focusing
(From The Rockefeller Institute) Materials and Methods. Observations with the Electron Microscope
 ELECTRON MICROSCOPE STUDY OF THE DEVELOPMENT OF THE PAPILLOMA VIRUS IN THE SKIN OF THE RABBIT* BY ROBERT S. STONE,~ M.D., RICHARD E. SHOPE, M.D., DAN H. MOORE, P,~.D. (From The Rockefeller Institute) PLATES
ELECTRON MICROSCOPE STUDY OF THE DEVELOPMENT OF THE PAPILLOMA VIRUS IN THE SKIN OF THE RABBIT* BY ROBERT S. STONE,~ M.D., RICHARD E. SHOPE, M.D., DAN H. MOORE, P,~.D. (From The Rockefeller Institute) PLATES
Part I. Content: History of Viruses. General properties of viruses. Viral structure. Viral classifications. Virus-like agents.
 Viruses Part I Content: History of Viruses. General properties of viruses. Viral structure. Viral classifications. Virus-like agents. History Through the 1800s, many scientists discovered that something
Viruses Part I Content: History of Viruses. General properties of viruses. Viral structure. Viral classifications. Virus-like agents. History Through the 1800s, many scientists discovered that something
The Zombies of the Scientific Community Viruses and Other Acellular Infectious Agents. Acellular Agents
 viruses protein and nucleic acid viroids RNA virusoids RNA prions proteins The Zombies of the Scientific Community Viruses and Other Acellular Infectious Agents Acellular Agents Viruses major cause of
viruses protein and nucleic acid viroids RNA virusoids RNA prions proteins The Zombies of the Scientific Community Viruses and Other Acellular Infectious Agents Acellular Agents Viruses major cause of
Symmetry-mismatch reconstruction of genomes and associated proteins within icosahedral viruses using cryo-em
 Biophys Rep 2016, 2(1):25 32 DOI 10.1007/s41048-016-0024-5 Biophysics Reports PROTOCOL Symmetry-mismatch reconstruction of genomes and associated proteins within icosahedral viruses using cryo-em Xiaowu
Biophys Rep 2016, 2(1):25 32 DOI 10.1007/s41048-016-0024-5 Biophysics Reports PROTOCOL Symmetry-mismatch reconstruction of genomes and associated proteins within icosahedral viruses using cryo-em Xiaowu
3D Imaging of Biological Specimens by Electron Microscopy
 3D Imaging of Biological Specimens by Electron Microscopy Focus on Microscopy Hannover, May 2010 Elke Spiess, Wim Voorhout and Wim Busing FEI Company The world of decreasing dimension From van Leeuwenhoek
3D Imaging of Biological Specimens by Electron Microscopy Focus on Microscopy Hannover, May 2010 Elke Spiess, Wim Voorhout and Wim Busing FEI Company The world of decreasing dimension From van Leeuwenhoek
Kinematic Analysis of Lumbar Spine Undergoing Extension and Dynamic Neural Foramina Cross Section Measurement
 Copyright c 2008 ICCES ICCES, vol.7, no.2, pp.57-62 Kinematic Analysis of Lumbar Spine Undergoing Extension and Dynamic Neural Foramina Cross Section Measurement Yongjie Zhang 1, Boyle C. Cheng 2, Changho
Copyright c 2008 ICCES ICCES, vol.7, no.2, pp.57-62 Kinematic Analysis of Lumbar Spine Undergoing Extension and Dynamic Neural Foramina Cross Section Measurement Yongjie Zhang 1, Boyle C. Cheng 2, Changho
Custom-Made Products / Scientific Research Instruments
 Synchrotron Radiation Instruments Double Crystal Monochromator A double crystal monochromator is an instrument that extracts light of a specific wavelength using crystal diffraction. Light of various wavelengths
Synchrotron Radiation Instruments Double Crystal Monochromator A double crystal monochromator is an instrument that extracts light of a specific wavelength using crystal diffraction. Light of various wavelengths
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supporting Information for:
 Supporting Information for: A Robust Liposomal Platform for Direct Colorimetric Detection of Sphingomyelinase Enzyme and Inhibitors Margaret N. Holme, 1,2,3,4 Subinoy Rana, 1,2,3,5 Hanna M. G. Barriga,
Supporting Information for: A Robust Liposomal Platform for Direct Colorimetric Detection of Sphingomyelinase Enzyme and Inhibitors Margaret N. Holme, 1,2,3,4 Subinoy Rana, 1,2,3,5 Hanna M. G. Barriga,
Chair of Medical Biology, Microbiology, Virology, and Immunology STRUCTURE, CLASSIFICATION AND PHYSIOLOGY OF VIRUSES
 Chair of Medical Biology, Microbiology, Virology, and Immunology STRUCTURE, CLASSIFICATION AND PHYSIOLOGY OF VIRUSES Viruses are small obligate intracellular parasites, which by definition contain either
Chair of Medical Biology, Microbiology, Virology, and Immunology STRUCTURE, CLASSIFICATION AND PHYSIOLOGY OF VIRUSES Viruses are small obligate intracellular parasites, which by definition contain either
C-Fern TM Investigations:
 C-Fern TM Investigations: Chemical Attraction C-Fern Sperm Chemotaxis Kit Instructor Version Introduction In this manual, a vertical bar to the left of text indicates material included in the Instructor
C-Fern TM Investigations: Chemical Attraction C-Fern Sperm Chemotaxis Kit Instructor Version Introduction In this manual, a vertical bar to the left of text indicates material included in the Instructor
LESSON 1.4 WORKBOOK. Viral sizes and structures. Workbook Lesson 1.4
 Eukaryotes organisms that contain a membrane bound nucleus and organelles. Prokaryotes organisms that lack a nucleus or other membrane-bound organelles. Viruses small, non-cellular (lacking a cell), infectious
Eukaryotes organisms that contain a membrane bound nucleus and organelles. Prokaryotes organisms that lack a nucleus or other membrane-bound organelles. Viruses small, non-cellular (lacking a cell), infectious
Viruses defined acellular organisms genomes nucleic acid replicate inside host cells host metabolic machinery ribosomes
 The Viruses Viruses Viruses may be defined as acellular organisms whose genomes consist of nucleic acid, obligately replicate inside host cells using host metabolic machinery and ribosomes to form a pool
The Viruses Viruses Viruses may be defined as acellular organisms whose genomes consist of nucleic acid, obligately replicate inside host cells using host metabolic machinery and ribosomes to form a pool
Cell Membrane Structure (1.3) IB Diploma Biology
 Cell Membrane Structure (1.3) IB Diploma Biology Essential idea: The structure of biological membranes makes them fluid and dynamic http://www.flickr.com/photos/edsweeney/6346198056/ 1.3.1 Phospholipids
Cell Membrane Structure (1.3) IB Diploma Biology Essential idea: The structure of biological membranes makes them fluid and dynamic http://www.flickr.com/photos/edsweeney/6346198056/ 1.3.1 Phospholipids
OCTAGONAL NUCLEAR PORES
 OCTAGONAL NUCLEAR PORES JOSEPH G. GALL From tile Department of Biology, Yale University, New Haven, Connecticut ABSTRACT Negative staining of isolated nuclear envelopes by phosphotungstate shows that the
OCTAGONAL NUCLEAR PORES JOSEPH G. GALL From tile Department of Biology, Yale University, New Haven, Connecticut ABSTRACT Negative staining of isolated nuclear envelopes by phosphotungstate shows that the
AET-treated normal red cells (PNH-like cells)
 J. clin. Path., 1971, 24, 677-684 Electron microscope study of PNH red cells and AET-treated normal red cells (PNH-like cells) S. M. LEWIS, G. LAMBERTENGHI, S. FERRONE, AND G. SIRCHIA From the Department
J. clin. Path., 1971, 24, 677-684 Electron microscope study of PNH red cells and AET-treated normal red cells (PNH-like cells) S. M. LEWIS, G. LAMBERTENGHI, S. FERRONE, AND G. SIRCHIA From the Department
The Structure of the Rotavirus Inner Capsid Studied by Electron Microscopy of Chemically Disrupted Particles
 J. gen. Virol. (1986), 67, 1721-1725. Printed in Great Britain 1721 Key words: rotavirus/capsid structure~chemical degradation The Structure of the Rotavirus Inner Capsid Studied by Electron Microscopy
J. gen. Virol. (1986), 67, 1721-1725. Printed in Great Britain 1721 Key words: rotavirus/capsid structure~chemical degradation The Structure of the Rotavirus Inner Capsid Studied by Electron Microscopy
Some living things are made of ONE cell, and are called. Other organisms are composed of many cells, and are called. (SEE PAGE 6)
 Section: 1.1 Question of the Day: Name: Review of Old Information: N/A New Information: We tend to only think of animals as living. However, there is a great diversity of organisms that we consider living
Section: 1.1 Question of the Day: Name: Review of Old Information: N/A New Information: We tend to only think of animals as living. However, there is a great diversity of organisms that we consider living
19 2 Viruses Slide 1 of 34
 1 of 34 What Is a Virus? What Is a Virus? Viruses are particles of nucleic acid, protein, and in some cases, lipids. Viruses can reproduce only by infecting living cells. 2 of 34 What Is a Virus? Viruses
1 of 34 What Is a Virus? What Is a Virus? Viruses are particles of nucleic acid, protein, and in some cases, lipids. Viruses can reproduce only by infecting living cells. 2 of 34 What Is a Virus? Viruses
Discrimination and Generalization in Pattern Categorization: A Case for Elemental Associative Learning
 Discrimination and Generalization in Pattern Categorization: A Case for Elemental Associative Learning E. J. Livesey (el253@cam.ac.uk) P. J. C. Broadhurst (pjcb3@cam.ac.uk) I. P. L. McLaren (iplm2@cam.ac.uk)
Discrimination and Generalization in Pattern Categorization: A Case for Elemental Associative Learning E. J. Livesey (el253@cam.ac.uk) P. J. C. Broadhurst (pjcb3@cam.ac.uk) I. P. L. McLaren (iplm2@cam.ac.uk)
WHY? Viruses are considered non-living because they do:
 Viruses What is a Virus? Non-living particle WHY? Viruses are considered non-living because they do: NOT Carry out metabolism NOT Grow or develop NOT Replicate without the help of a living cell (host).
Viruses What is a Virus? Non-living particle WHY? Viruses are considered non-living because they do: NOT Carry out metabolism NOT Grow or develop NOT Replicate without the help of a living cell (host).
An active unpleasantness control system for indoor noise based on auditory masking
 An active unpleasantness control system for indoor noise based on auditory masking Daisuke Ikefuji, Masato Nakayama, Takanabu Nishiura and Yoich Yamashita Graduate School of Information Science and Engineering,
An active unpleasantness control system for indoor noise based on auditory masking Daisuke Ikefuji, Masato Nakayama, Takanabu Nishiura and Yoich Yamashita Graduate School of Information Science and Engineering,
Convergence Principles: Information in the Answer
 Convergence Principles: Information in the Answer Sets of Some Multiple-Choice Intelligence Tests A. P. White and J. E. Zammarelli University of Durham It is hypothesized that some common multiplechoice
Convergence Principles: Information in the Answer Sets of Some Multiple-Choice Intelligence Tests A. P. White and J. E. Zammarelli University of Durham It is hypothesized that some common multiplechoice
Observations on the Structure of the Nucleocapsids of some Paramyxoviruses
 J. gen. Virol. (I97O), 6, 14i-i5o Printed in Great Britain I4I Observations on the Structure of the Nucleocapsids of some Paramyxoviruses By J. T. FINCH Medical Research Council Laboratory of Molecular
J. gen. Virol. (I97O), 6, 14i-i5o Printed in Great Britain I4I Observations on the Structure of the Nucleocapsids of some Paramyxoviruses By J. T. FINCH Medical Research Council Laboratory of Molecular
Structural dynamics of viral nanomachines
 TECHNOLOGY Structural dynamics of viral nanomachines Joanna Kam 1 *, Andrew C. Demmert 1 *, Justin R. Tanner 1, Sarah M. McDonald 1 & Deborah F. Kelly 1 Rotavirus double-layered particles (DLPs) are formed
TECHNOLOGY Structural dynamics of viral nanomachines Joanna Kam 1 *, Andrew C. Demmert 1 *, Justin R. Tanner 1, Sarah M. McDonald 1 & Deborah F. Kelly 1 Rotavirus double-layered particles (DLPs) are formed
Parallel-Axis Gear Terminology
 Parallel-Axis Gear Terminology For more detailed coverage of this subject, consult ANSI/AGMA Standard 1012-F90; Gear Nomenclature, Definitions with Terms and Symbols Active Profile- that part of the gear
Parallel-Axis Gear Terminology For more detailed coverage of this subject, consult ANSI/AGMA Standard 1012-F90; Gear Nomenclature, Definitions with Terms and Symbols Active Profile- that part of the gear
Microbiology Chapter 7 Viruses
 Microbiology Chapter 7 Viruses 7:1 Viral Structure and Classification VIRUS: a biological particle composed of genetic material (DNA or RNA) encased in a protein coat CAPSID: protein coat surrounding a
Microbiology Chapter 7 Viruses 7:1 Viral Structure and Classification VIRUS: a biological particle composed of genetic material (DNA or RNA) encased in a protein coat CAPSID: protein coat surrounding a
Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions
 Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions 11/20/2017 MDufilho 1 Characteristics of Viruses Viruses Minuscule, acellular, infectious agent having either DNA or RNA Cause infections
Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions 11/20/2017 MDufilho 1 Characteristics of Viruses Viruses Minuscule, acellular, infectious agent having either DNA or RNA Cause infections
Nodavirus Coat Protein Imposes Dodecahedral RNA Structure Independent of Nucleotide Sequence and Length
 JOURNAL OF VIROLOGY, Mar. 2004, p. 2897 2905 Vol. 78, No. 6 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.6.2897 2905.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Nodavirus
JOURNAL OF VIROLOGY, Mar. 2004, p. 2897 2905 Vol. 78, No. 6 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.6.2897 2905.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Nodavirus
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Virology Introduction. Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment.
 DEVH Virology Introduction Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment. Definitions Virology: The science which study the
DEVH Virology Introduction Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment. Definitions Virology: The science which study the
count the strands of Hb S molecules present in cross sections of to improve the resolution in cross sections of the fibers, we have
 Proc. Nati. Acad. Sci. USA Vol 76, No. 3, pp. 1140-1144, March 1979 Biochemistry Cross-sectional views of hemoglobin S fibers by electron microscopy and computer modeling (sickle cell hemoglobin fibers/thin
Proc. Nati. Acad. Sci. USA Vol 76, No. 3, pp. 1140-1144, March 1979 Biochemistry Cross-sectional views of hemoglobin S fibers by electron microscopy and computer modeling (sickle cell hemoglobin fibers/thin
