Atomic insight into the CD4 binding-induced conformational
|
|
|
- Crystal James
- 5 years ago
- Views:
Transcription
1 Atomic insight into the CD4 binding-induced conformational changes in HIV-1 gp120 Shang-Te D. Hsu and Alexandre M. J. J. Bonvin* Bijvoet Center for Biomolecular Research, Utrecht University, Padualaan 8, 3584CH Utrecht, The Netherlands Running title: CD4 binding-induced conformational changes in HIV-1 gp120 *Corresponding author: Tel: Fax:
2 Summary The entry of HIV-1 into a target cell requires gp120 and receptor CD4 as well as coreceptor CCR5/CXCR4 recognition events associated with conformational changes of the involved proteins. The binding of CD4 to gp120 is the initiation step of the whole process involving structural rearrangements that are crucial for subsequent pathways. Despite the wealth of knowledge about the gp120/cd4 interactions, details of the conformational changes occurring at this stage remain elusive. We have performed molecular dynamics simulations in explicit solvent based on the gp120/cd4/cd4i crystal structure in conjunction with modeled V3 and V4 loops to gain insight into the dynamics of the binding process. Three differentiated interaction modes between CD4 and gp120 were found, which involve electrostatics, hydrogen bond and van der Waals networks. A binding funnel model is proposed based on the dynamical nature of the binding interface together with a CD4-attraction gradient centered in gp120 at the CD4-Phe43-binding cavity. Distinct dynamical behaviors of free and CD4-bound gp120 were monitored, which likely represent the ground and pre-fusogenic states, respectively. The transition between these states revealed concerted motions in gp120 leading to: i) loop contractions around the CD4-Phe43- insertion cavity; ii) stabilization of the four-stranded bridging sheet structure; and iii) translocation and clustering of the V3 loop and the bridging sheet leading to the formation of the coreceptor binding site. Our results provide new insight into the dynamic of the underlying molecular recognition mechanism that complements the biochemical and structural studies. Keywords: HIV-1 gp120, CD4, molecular dynamics, conformational change, binding funnel 2
3 Introduction The entry of Human Immunodeficiency Virus type-1 (HIV-1) into a target cell is an extremely intricate process (for reviews, see ref 1-4), which requires the recognition of the host cell receptor CD4 and coreceptors, mainly CCR5 and CXCR4, by the viral envelope glycoprotein (Env), consisting of gp120 and gp41. Env is organized into trimeric spike structures on the surface of the virion 5 anchored in the membrane by the transmembrane subunit gp41 while gp120 is presented at the surface as the frontier for the recognition process. The initiation step involves binding of CD4 to gp120 which induces conformational changes in gp leading to the exposure and/or formation of antigenic epitopes recognized by the coreceptors. 9,10 Binding to the coreceptors takes place subsequently and gp41 undergoes conformational changes that mediate the fusion process by formation of the fusogenic structure bringing viral and target cell membranes into close vicinity. This complex mechanism involves a series of structural rearrangements in which conformational dynamics plays a crucial role. Over the past decades, a large number of crucial elements involved in this system, in particular gp120, have been unraveled using a variety of approaches. 1,11 The recent determination of the crystallographic structure of the core of gp120 in complex with CD4 and the CD4-induced antibody (CD4i), which recognizes the coreceptor binding site, provided tremendous structural insights. 12,13 Despite the highly variable sequence of HIV-1 gp120 throughout evolution, the topology of the core has been retained with some degree of sequence diversity. 12,13 It contains several disulphide bridges that buckle the flexible hypervariable loops to form knots at the periphery of the rigid core and basically define their boundaries. The heavy glycosylation at the silent face 3
4 and the mobile variable loops together provide a shielding umbrella to evade immune system neutralization. 1 CD4 is bound to gp120 predominantly via electrostatic interactions with a large, but mismatched interface. An unusually protuberant Phe43 of CD4 inserts into a receptive hole of gp making up a knob-and-socket interaction. 8 Evidence from a thermodynamics study has suggested that the binding of CD4 to gp120 induces substantial structural rearrangements primarily in the core structure, a truncated form of gp120 that was used in the crystallographic studies where the flexible N- and C-termini as well the variable loops, V1-V3, are absent (see Figure 1). 14 It is now generally accepted that the gp120/cd4 complex formation leads to the accessibility of the coreceptor binding site. The third variable (V3) loop, in particular, plays a central role here since it contains not only the principle neutralizing determinant (PND) 15 but also the main coreceptor determinant (CXCR4 versus CCR5 usage) for HIV-1 at this stage. 3 X-ray crystallographic and Nuclear Magnetic Resonance (NMR) studies of various types of V3 loop fragments have shown a high degree of structural heterogeneity, which may thus provide means for HIV-1 to evade neutralization by the immune system. 17 On one hand, biochemical studies have established the outline of the fusion process revealing its vast diversity and sophistication; on the other hand, results from the structural studies have provided static snapshots of the interaction between gp120 and its interacting partners at atomic resolution. The dynamics involved in this mechanism, however, remains elusive. We have performed molecular dynamics (MD) simulations to investigate the conformational changes in gp120 induced upon complexation of CD4. By comparing the dynamical properties of gp120 in the free form and bound to CD4, we will show that CD4 binding substantially reduces the dynamical motions of gp120. An extensive 4
5 intermolecular hydrogen bond network is formed at the interface and around the Phe43 binding cavity. We monitored motions around the CD4 binding site that are correlated to the V3 loop and trimerization interface, which likely represent the initial steps of the subsequent structural rearrangements required for coreceptor binding and gp41 mediated membrane fusion. Material and Methods Generation o the starting structures The starting structure of gp120 with the truncated LAI strain sequence (SWISS-PROT accession number P03377) was obtained from SWISS-MODEL 23 using structural homologs as template (PDB entries 1G9M 12, 1G9N 13 and 1GC1 for the gp120 core and 1CE4 for the V3 loop 24 ). The particular strain was chosen because of ongoing collaborations with experimental groups (I. Braakman, Utrecht University and B. Berkhout, University of Amsterdam, personal communication). The primary sequence of gp120 of LAI differs from HxBc2 by only six amino acids in the core region (98.1% sequence identity, see Figure 1). Core gp120 is defined as the construct present in the crystal structure with the truncated V1-V3 and N- and C-termini. The V4 loop ( ), for which electron density is missing in all crystal structures, and the V3 loop, which has been proposed to undergo significant rearrangements upon CD4 binding, were modeled onto the g120 core, together with the six core mutations of the LAI strain, using the default settings of SWISS-MODEL. 23 For the V3 loop, a NMR structure (residues ; PDB ID: 1CE4) was taken for docking onto the core structure. This modeling and in particular the six point mutations are expected to have only minor effect on the overall structure of gp120. For example, the crystal structures of two HIV-1 variants, HxBc2 (1G9M) and YU2 (1G9N) with a sequence 5
6 identity of 85.9%, show backbone (N, C and Ca) root-mean-square deviation (RMSD) of only 1.16 Å. The structural variation in core gp120 between LAI and HxBc2 which share 98.1% sequence identity should thus be minimal and our starting model for the simulations can be considered accurate. For the simulation of the complex form (Cplx), the LAI gp120 model (residues ) was superimposed onto the crystal structure of the complex (1G9M) to obtain the relative orientation with respect to the D1 domain of CD4 (CD4-D1, residues 1-99). Monomeric gp120 (Freegp120) and CD4-D1 (Free-CD4) taken from the structure of the complex were used as starting structures in the free simulations for comparison. The starting conformations of gp120 in the free and complex simulations were thus identical and correspond to the CD4-bound form. Molecular dynamics simulation setup and structural analysis The GROMACS 3.0 molecular dynamics package 25 was used with the GROMOS 43A1 force field. 26 Starting structures of the free gp120 (Free-gp120), the free CD4 D1 domain (Free-CD4) and the gp120/cd4 D1 domain complex (Cplx) were individually solvated using the single point charge (SPC) water model 27 in rectangular periodic boxes with a 1.4 nm solute-wall minimum distance. After a first steepest descent energy minimization with positional restraints on the solute, seven, three and ten chloride anions were introduced in Free-gp120, Free-CD4 and Cplx, respectively, to obtain electro-neutralized systems. The resulting systems comprised 89608, and atoms for Free-gp120, Free-CD4 and Cplx, respectively. A second energy minimization was then performed, followed by five successive 20 ps MD runs with decreasing positional restraints force constants on the solutes (K posres = 1000, 1000, 100, 10 and 0 kj mol -1 nm -2 ) prior to the 10 ns production runs. A two femtosecond 6
7 time step was used for the integration of the equations of motion. Solute, solvent and counter-ions were independently coupled to a reference temperature bath at 300K with a coupling constant t T of 0.1 ps 28. The pressure was maintained by weakly coupling the system to an external pressure bath at one atmosphere with a coupling constant t P of 0.5 ps. Non-bonded interactions were calculated using twin range cutoffs of 0.8 and 1.4 nm. Long range electrostatic interactions beyond the cutoff were treated with the generalized reaction field model using a dielectric constant of The non-bonded pair list was updated every 10 steps. The LINCS algorithm 30 was used for bond length constraining in conjunction with dummy atoms for the aromatic rings and amino group in side chains. 31 Owing to the large system sizes ca 2 ns were required to reach equilibrium. The analysis was performed therefore on the 2-10 ns segments for all systems. Equilibrated trajectories (2-10 ns) of free and CD4-bound gp120 were merged for essential dynamics analysis. 32 After diagonalizing the backbone atom (N, C and Ca) mass-weighted covariance matrix the primary principle component (the first eigenmode) was extracted and identified as the transition mode between the two states. From a projection of the merged trajectory along this eigenmode the structures corresponding to the two extremes were obtained. The b-turn conformations in the V3 loop were compared with several experimental structures of this loop (PDB entries: 1CE4 24, 1B03 18, 1QNZ 19, 1AI1 16, 1F58 and 2F58 17 ). For all types of b-turns a minimum distance between the Ca(i) and Ca(i+3) of 0.7 nm was required. The criteria for backbone torsion f/y angles for type II b-turn are: f(i+1) = -60º, y(i+1) = 120º, f(i+2) = 80º and y(i+2) = 0º and those for type VIII 7
8 b-turn were: f(i+1) = -60º, y(i+1) = -30º, f(i+2) = -120º and y(i+2) = 120º, each torsion angle has a ±45º tolerance. Results In order to investigate the conformational changes in gp120 associated with CD4 binding, three molecular dynamics (MD) simulations of 10 ns production were performed in explicit solvent for the following systems: free gp120 (Free-gp120), the free D1 domain of CD4 (Free-CD4) and the gp120/cd4 D1 domain complex (Cplx). All systems showed stable trajectories with conserved secondary and tertiary structures (Table 1). Stability and equilibration of the systems was monitored by foloowing the RMSDs as a function of time (Figure 2). The loop regions in free gp120, namely V1-V5, give rise to larger structural deviations than in the complex form (Figure 2 and Table 1). The larger RMSDs of the entire gp120 backbone in the free form primarily stem from the re-positioning of the loops near the CD4 binding site as can be readily seen by comparing the snapshots of the trajectories of gp120 in the free and CD4 bound forms (Figure 3). The snapshots also illustrate that the V3 loop, which started from an identical initial configuration in the free and complex simulations, evolves into distinct configurations with well-separated spatial distributions in the two simulations. In addition, substantial reduction in the amplitude of loop motions, in particular in the V1/V2 stem, can also be observed (this will be discussed in detail in the following section). Given the compactness of CD4-D1, its RMSDs from the crystal structure are, in both runs, small and constant regardless of the presence of gp120 (see Table 1). Despite the different dynamical behavior of gp120 s flexible loops, which is reflected both in RMSDs and per residue fluctuations, the internal energies of both gp120 and CD4 are not much perturbed 8
9 upon binding (Table 2). Also worth noting is that the energy of binding (E complex E free gp120 E free CD4 ) amounts to 130 kj/mol (considering only protein-protein and protein-solvent interactions and thus assuming that no significant changes are taking place in the bulk solvent and counter ions). This is in fair agreement with the change in enthalpy upon complexation of gp120/cd4 (-62±2 kj/mol at 310K) reported by Doyle and co-workers. 14 Substantial reduction of gp120 loop mobility and conformational change upon CD4 binding A clear difference in mobility is found between the variable loops of gp120, V1-V5, and the conserved core region. Binding of CD4 substantially decreases the motions of the loops interacting with CD4, especially the V1/V2 stem and the V5 loop. It also induces a remarkable repositioning of the V3 loop that remains highly mobile in both states (Figure 3). Analysis of the backbone RMS fluctuations, expressed in terms of crystallographic temperature factor (B-factor), as a function of residue sequence for Free-gp120 and Cplx (Figure 4) reveals that most of the high-mobility sites (except V3 and V4) coincide with the CD4 binding-site. The calculated B-factors of gp120 match well with the crystallographic data with the exception of the V1/V2 stem. In CD4, however, they are significantly smaller than the experimental ones. Discrepancies in the overall B-factors amplitude are also present between the two crystal structures of free CD4, 1CDH 33 and 3CD4 34, where the latter shows similar sequential patterns but 30-40% larger overall values. Although a quantitative comparison between the simulated and experimental B-factors may not be applicable due to the difference between experimental and simulation conditions, 35 it is apparent 9
10 that the observed reduction in loop mobility upon binding near Phe43 in CD4-D1 is consistent with the experimental findings (Figure 4 inset). The sizeable reduction of mobility upon binding is primarily determined by the intermolecular contacts in the crystal structure, e.g. the V1/V2 stem, D and V5 regions as well as the C3 and C4 regions that contact the protruding CD4-Phe The attenuation of the loop motions arises, to a large extent, from the formation of many intermolecular hydrogen bonds, most of which are also present in the crystal structure (Table 3, Figure 4 and 5). The large loop motions in gp120 observed in both the free and complex simulations are consistent with the fact that the crystallization of gp120 was only possible when the variable loops, i.e. V1-V3, were truncated (Figure 1); the introduction of CD4 was necessary to further stabilize the ternary complex. Despite its shorter loop length, resulting in higher structural restriction, the V4 loop was still disordered and unresolved in the crystal structures, 12,13 probably because of its intrinsic conformational heterogeneity and remoteness from the CD4 binding interface. Intermolecular hydrogen bond network and Phe43 cavity The increase in rigidity observed in the complex originates primarily from the formation of intermolecular hydrogen bonds. Although many of them are already present in the crystal structure, some are only formed during the molecular dynamics simulation. Our simulation allows, in addition, to assess the dynamical nature of these critical interactions. This can be done by analysing their occurrence throughout the trajectory. Many long-lived intermolecular hydrogen bonds bridge the tips of the mobile loops of gp120 interacting with CD4 and a hemisphere of CD4-D1 (Figure 5). This hydrogen bond network nicely encloses the docking cavity of Phe43, which is 10
11 the key element for the binding mechanism. A large number of side chain-side chain or side chain-backbone hydrogen bonds cover a large area of the complex interface in a nonspecific manner. For instance, the side chain hydroxyl group of S42 in CD4 is able to form several hydrogen bonds by hopping among various backbone hydrogen bond acceptors in gp120 that are in its close proximity (Table 3). In addition, there are a few stable backbone-backbone hydrogen bonds in the vicinity of the Phe43 cavity (Table 3) involving residues of gp120 (S365, G366, G367 and D368) that are well conserved among various HIV-1 strains. The importance of these residues has been highlighted by structure-based mutagenic studies in which mutations of S365, G366 and D368 in gp120 diminished the affinity of the CD4-binding site specific antibody b Overall, the residues involved in these intermolecular hydrogen bonds gave rise to an average intermolecular Coulomb s electrostatic energy of 624±124 kj/mol, accounting for 61% of the total intermolecular Coulomb s electrostatic energy (Table2). These observations allow us to propose a functional role for the hydrogen bond network: the nonspecific, loose, flexible and wide-spread hydrogen bonds and saltbridges and the specific, tight, rigid and confined ones generate an affinity gradient that drives the insertion of the Phe43 phenyl ring into its binding cavity in gp120. Once inserted, it is locked in its knob-and-socket geometry by the specific hydrogen bond network in the proximity of the binding pocket. The hydrophobic contacts between the CD4 Phe43 side chain and the gp120 cavity then provides the short-range stabilizing factor. These contacts are extremely well maintained during the simulation bearing very small structural variations (Table 4); translation and rotation of the 11
12 phenyl ring with respect to the cavity were limited within 0.2 nm and 60º, respectively (data not shown). CD4 binding induced bridging sheet stabilization The so-called bridging sheet is a four-stranded b-sheet that consists of b2 and b3 in the V1/V2 stem and b20 and b21 in the C3 region. It interposes between the inner and outer domains of gp120 and is thought to be flexible and disordered in the absence of CD4. Recent thermodynamic studies on the binding of gp120 and CD4 have demonstrated unprecedented large entropy changes, which were proposed to be possibly accounted for by a stabilization of the bridging sheet and a restriction of the interdomain motions upon CD4 binding. Detailed structural and dynamical information is, however, still lacking. The conformational changes of the bridging sheet upon removal of CD4 were monitored in silico by RMSD matrix analysis (Figure 6 lower right panel). The bridging sheet undergoes a series of transient structural changes with a major one at about 8 ns, after which it adopts a conformation somewhat similar to the ones in the ns segment. The same analysis on the simulation of the complex showed little change throughout the 10 ns simulation (Figure 6 upper left panel). Several interstrand backbone hydrogen bonds between the antiparallel b2 and b21 that delimit the inner and outer domains of gp120 also show different hydrogen-acceptor distance distribution in the two simulations: While the total numbers of the interstrand backbone hydrogen bonds throughout the simulations are the same, for Free-gp120, the hydrogen bond length distribution peaks at nm with a half width of nm whereas in Cplx it peaks at nm with a narrower half width of nm. The 12
13 shorter hydrogen bond distances in the complex are indicative of an increased stability of the bridging sheet upon CD4 binding while it remains flexible otherwise. This is consistent with the model proposed in the thermodynamic studies mentioned previously. Conformational changes and polymorphism of the V3 loop upon CD4 binding The V3 loop has been targeted as a prominent candidate to tackle the viral fusion problem. Based on structural, 12 mutagenic 11 and antigenic analyses, 37 coreceptor binding was proposed to require the V3 loop in concert with the bridging sheet. These form the coreceptor binding site that is created only upon CD4 binding. A large panel of monoclonal antibodies (MAbs) based on V3 loop fragments has been elicited against several primary isolates from different HIV-1 clades (subtypes) Several V3 loop-derived peptides have also been devised to inhibit viral entry into target cell in a coreceptor-specific manner Successful vaccine design has however been hindered by the underlying structural polymorphism of the V3 loop. The wellconserved region, GPGR, can indeed adopt various type of b-turns and is found in various configurations when bound to different MAbs (extensive discussion can be found in ref. 20) ,22,44 Note that previous structural studies were only performed on V3 loop fragments and little is thus known about its conformation when attached to the core structure. We therefore investigated the dynamics and conformation of the V3 loop both in free gp120 and in the CD4-bound form. As mentioned above the V3 loop is highly mobile and occupies an ample space in the vicinity of the coreceptor binding site. Different localizations in the free and CD4- bound states can be clearly distinguished in the corresponding trajectories (Figure 3). 13
14 Relocation of the V3 loop, upon CD4 binding, towards the basal part of the bridging sheet agrees with the model proposed by Sodroski and co-workers based on their mutagenesis study on the coreceptor CCR5 binding. 11 In their comprehensive study, the removal of the V3 loop (D ) was found to abolish the coreceptor CCR5 binding completely. Many residues (K121, T123, K207, L317, E381, F383, I420, K421, Q422, P438, R440 and G441) were also identified to result in more than 90% loss in CCR5 binding when mutated. Comparing the solvent accessible surface area (SAS) of these residues in the free and complex simulations reveals an increase of SAS in the complex form only for I420 (15%) and Q422 (31%) while a small decrease is observed for R440 (8%). Other residues are either unaffected or buried in both the free and complex form by the V3 loop. The changes in SAS of the above three residues amount, however, to a gain of ca 110 Å 2, which may be an indication of the proposed exposure of the binding site. The clustering of the V3 loop and the bridging sheet can also be inferred from the lowering of the electrostatic Coulomb s energy between the V3 loop as a whole (residues ) and the subset of residues that are important for CCR5 binding: -438±60 kj/mol in the complex form versus -345±101 kj/mol in the free form. The abundant basic residues of the V3 loop (six arginines and two lysines, see Figure 1) are thought to facilitate the recruitment of the generally acidic chemokine coreceptor subsequent to the binding of CD They generate a positive electrostatic potential that occupies an enormous space centered at the basal region of the V3 loop. Moving from the free to the complex form relocates the positive potential toward the coreceptor binding site while neglecting the V3 loop drastically reduces this positive electrostatic potential (Figure 7) 14
15 Structurally, the V3 loop shows a high propensity of b-hairpin formation in our simulations but the position of the turn is poorly defined (Figure 8). The conserved GPGR sequence shows several transient b-turns, including type II and type VIII b- turns in the two simulations. Other tight turns were found at RGPR, QRGP and IQRG, as a result of one to three residue upstream shifts. Comparison with the available GPGR structures (see Material and Methods) gives backbone RMSDs between 0.2 and 0.6 nm with the various antibody-bound structures irrespective of the effect of CD4 complexation (data not shown). Essential dynamics analysis reveals concerted loop motions in gp120 upon CD4 binding The complexity of molecular motions observed in MD simulations can be simplified by essential dynamics analysis, 32 whereby sets of essential and correlated motions are extracted. Such an analysis allows distinguishing high frequency local motions, which are restrained and harmonic in nature and contain less structural information, from essential, large amplitude global motions. In order to focus on the motions associated with CD4 binding such an analysis was performed on merged trajectories from the free and complex form of gp120 and of CD4-D1. Two distinct distributions along the first eigenvector, i.e., with the largest eigenvalue, can be found, which correspond to the well separated free and complex states of gp120 (Figure 9A & B). Already the second eigenvector of the two trajectories shows overlap in positional fluctuation displacement suggesting similar motions between the two states. Narrow Gaussian displacement distributions appear after the first few eigenvectors suggesting restrained harmonic motions common to the two states. Projections of the two extremes of the first eigenvector along the merged trajectory on the average structure 15
16 illustrate the major difference between the two states (Figure 9C). Note that the extrapolation between the two extremes does not represent the transition pathway but merely highlights the structural differences. In gp120, a correlated loop-contraction primarily around the CD4 binding site is revealed with, in particular, a pronounced closure of the lids of the Phe43 cavity (Figure 9C). This correlated motion increases the curvature of the CD4 binding site leading to a gain in interfacial complementarity. The concerted contraction involves primarily the V1/V2 stem, V5 and, to a less extent, the C3 and C4 regions. These finger-like structures, in particular the V1/V2 stem and V5, are in a much more open and relaxed state in the free form. The most prominent transition occurred in the V3 loop, which is driven by CD4 binding towards the basal region of the bridging sheet. In contrast to the substantial conformational changes in gp120 no major structural transition upon binding was identified for CD4-D1 suggesting that the principle modes of motion are not affected by complexation and that only their amplitudes are reduced. Discussion We have shown by MD simulations that CD4 binding reduces the mobility of various loops of gp120 around the binding site and induces conformational changes that effectively lead to the wrapping up of a hemisphere of CD4-D1. While substantial changes occur in gp120, in CD4-D1, only the Phe43 containing loop showed a slight reduction of mobility upon complexation. The intermolecular interactions can be categorized into three levels: nonspecific side chain-side chain or side chain-backbone hydrogen bonds (or salt-bridges), specific backbone-backbone hydrogen bonds and 16
17 hydrophobic contacts between both partners resulting in the insertion of CD4 Phe43 into the receptive cavity in gp120. The large-amplitude motions of the V1/V2 stem and V3 and V4 loops may provide a shielding umbrella that masks the CD4 binding site in the monomeric form of gp120 as proposed by Kwong et al. based on their thermodynamic finding. 48 The observed entropy lost upon binding of CD4 to gp120 14,48 is a clear indication of a reduction of flexibility and is in line with our observations. When CD4 is attracted into the vicinity of the binding site, the mobility of the V1/V2 stem, V5 loop and D reduces. The extensive intermolecular hydrogen bond formation and van der Waals contacts at the gp120/cd4 interface result in the closure of C3 and C4 regions leading to the formation of the required Phe43 binding cavity. They also reduce the overall mobility in the regions involved in the intermolecular contacts between both partners. The formation of the extensive intermolecular hydrogen bonds as well as the stabilization of the bridging sheet may account for the experimentally observed entropic loss, 14,48 which is compensated by a gain in enthalpy through the intermolecular interactions described here. In contrast, the V3 loop remains flexible after its relocation induced by the binding of CD4 and no entropy cost is thus paid for this process. These observations are in agreement with the experimental findings that the main entropic changes occur within the core structure of gp One can summarize the above observations into a "binding funnel" model where dynamics and different interaction modes are coupled leading to efficient recognition and specific affinity. 49 During the search of receptor CD4, gp120 is constantly subject to immune system attack. A defined binding pocket based on rigid body docking 17
18 would require exhausting geometry search while antibodies neutralization via structural epitope recognition is also taking place. Gp120 devised, therefore, a multilevel binding mechanism as such that the conformation with the specific binding cavity is embedded in an ensemble of less selective conformations facilitating target search: i) the charged residues located in the highly mobile loops generate long range electrostatic attraction that guide CD4 to its binding interface consisting of a nonspecific hydrogen bond network bearing less structural selectivity and energetic trapping; ii) the loose side chain-side chain hydrogen bonding that confines gp120 and CD4 into close proximity leads to the formation of specific backbone-backbone hydrogen bonds and subtle changes in the loops around CD4 lead to the closure of the lid of the receptive cavity; iii) while the specific hydrogen bonds are formed, the Phe43 phenyl ring plugs into its binding cavity. Only one molecule that perfectly matches with the last two steps, CD4, will be able to accomplish the structural rearrangements that weld the complex structure together, thereby completing the first stage of the fusion process: the gp120/cd4 recognition. It should be noted that we did not simulate the binding process and therefore no kinetic or free energy data could be obtained from the current study. Therefore, our binding funnel model cannot be interpreted in term of a free energy surface of binding; it only presents a geometrical and chronological view of the binding process. The ability of anti-cd4-binding-site MAbs to neutralize HIV-1 has been associated with their affinity for the trimeric envelope virion structure, rather than monomeric gp Recent thermodynamics study also suggests that the monomeric gp120 may act as decoys to the immune system. 48 Most antibodies but b12 and 4KG5 are in fact elicited against the flexible monomeric gp120 with suboptimal angle of approach to 18
19 the CD4 binding site such that they are sterically hindered by the interaction between the variable loops in gp The ability to bind oligomeric gp120 is in fact the determinant to neutralize the virus. 51,52 On the one hand, the interpretation of our observed conformational changes is confounded by the limited knowledge about the trimeric state of gp120, the monomeric gp120 crystal structure being the only structural template available to date. On the other hand, CD4 is the unique element for the initiation of the CD4-dependent viral entry pathway. The formation of the gp120/cd4 complex is apparently the common ground for recognition, regardless of the oligomeric state. This actually demonstrates the remarkable plasticity of gp120 where the quaternary constraint imposed by the trimerization of the envelope proteins may well be generated via the binding to CD4 alone, assuming that the structure of the monomeric gp120/cd4 complex resembles that in the oligomeric state. Although not presented here, we do observe correlated motions upon CD4 binding on the back-side of gp120 which has been proposed to be involved in the trimerization process, either directly or more likely via interactions with the transmembrane trimerization domain gp41. Under the assumption of an unique gp120/cd4 complex structure, the gp120/cd4/cd4i crystal structure has provided invaluable information about the gp120/cd4 interface, which has led to a large number of structural and biochemical studies such as structural mapping by monoclonal antibodies. 36,53 Yet, it still lacks the dynamical information regarding the conformational changes occurring in gp120 upon CD4 binding, the limiting factor to the subsequent viral entry process. Some intermolecular hydrogen bonds, mostly mobile side chain-side chain hydrogen bonding (Figure 5), were only identified in our simulation. The geometry and rigidity of the well-defined CD4-Phe43 cavity, which is composed of some mobile residues in the C3 and C4 regions, is only induced upon CD4-binding (Figure 3). Furthermore, 19
20 formation of the complimentary gp120/cd4 interface requires correlated loop contractions as revealed by essential dynamics analysis. This process also leads to a stabilization of the bridging sheet while the two b-hairpins forming it (the V1/V2 stem and C3) are somewhat parted in the free form. Correlated motions at the putative trimerization interface are also observed. We can only speculate that the perturbation at the trimerization interface, propagated from the gp120/cd4 interface, may trigger the release of tension stored within the coiled-coil structure of gp41 as proposed in the spring-loaded model. 54 Conclusion Our simulations of free and CD4-bound gp120 represent two distinct states, namely the relaxed ground state and the contracted, excited or pre-fusogenic state of gp120, respectively. Large amplitude loop motions in gp120 are attenuated and show concerted contraction upon CD4 binding. The associated entropic cost is compensated by the wealth of intermolecular interactions at the CD4 interface ranging from nonspecific to specific hydrogen bonds and hydrophobic contacts around and at the Phe43 binding cavity, in agreement with the large enthalpy/entropy compensation measured experimentally. This differentiated mode of interaction from long-range electrostatic attraction via non-specific and specific short-range interactions, in combination with the dynamical nature of the system, allowed us to propose a binding funnel model for the gp120/cd4 interaction. In addition, the complexation also drives clustering of the V3 loop and the bridging sheet that generates a highly positive electrostatic attraction gradient for subsequent coreceptor binding. In line with the proposed model deduced from the mutagenesis mapping of the coreceptor binding site, 11 our results provide a plausible explanation for the functional relationship 20
21 between the V3 loop and CD4-binding and the otherwise inefficient CD4-independent entry pathway (reviewed in ref 3). We have here demonstrated the sophisticated plasticity of gp120, from a rigid core to a floppy exterior shielding that provides recognition specificity without compromising the capacity to evade attack from the immune system. Acknowledgements The authors thank Prof. Ineke Braakman and Eelco van Anken (Department of Bio- Organic Chemistry, Bijvoet Center, Utrecht University, The Netherlands) and Prof. Ben Berkhout (Department of Human Retrovirology, Academic Medical Center, University of Amsterdam, The Netherlands) for careful reading and helpful discussions. This work was sponsored by the National Computing Facilities (NCF) for the use of supercomputer facilities, with financial support from the Netherlands Organization for Scientific Research (NWO). Supplementary material Two molecular movies showing orthogonal views of the primary transition of gp120 from free to bound form extracted via essential dynamic analysis are available as supplementary material in QuickTime format. The CD4 D1 domain is shown in transparency with Phe43 in yellow space-filling to indicate the position of the binding cavity. The binding of the D1 domain of CD4 is only a cartoon to highlight the resulting motion in gp120 and is not a result of the simulations. 21
22 References 1. Wyatt R, Sodroski J. The HIV-1 envelope glycoproteins: Fusogens, antigens, and immunogens. Science 1998;280: Chan DC, Kim PS. HIV entry and its inhibition. Cell 1998;93: Berger EA, Murphy PM, Farber JM. Chemokine receptors as HIV-1 coreceptors: Roles in viral entry, tropism, and disease. Annu. Rev. Immunol. 1999;17: Poignard P, Saphire EO, Parren P, Burton DR. GP120: Biologic aspects of structural features. Annu. Rev. Immunol. 2001;19: Kwong PD, Wyatt R, Sattentau QJ, Sodroski J, Hendrickson WA. Oligomeric modeling and electrostatic analysis of the gp120 envelope glycoprotein of human immunodeficiency virus. J. Virol. 2000;74: Sattentau QJ, Moore JP. Conformational-Changes Induced in the Human-Immunodeficiency- Virus Envelope Glycoprotein by Soluble Cd4 Binding. J. Exp. Med. 1991;174: Thali M, Moore JP, Furman C, Charles M, Ho DD, Robinson J, Sodroski J. Characterization of Conserved Human-Immunodeficiency-Virus Type-1 Gp120 Neutralization Epitopes Exposed Upon Gp120-Cd4 Binding. J. Virol. 1993;67: Moore JP, Binley J. HIV - Envelope's letters boxed into shape. Nature 1998;393: Wu LJ, Gerard NP, Wyatt R, Choe H, Parolin C, Ruffing N, Borsetti A, Cardoso AA, Desjardin E, Newman W, Gerard C, Sodroski J. CD4-induced interaction of primary HIV-1 gp120 glycoproteins with the chemokine receptor CCR-5. Nature 1996;384: Trkola A, Dragic T, Arthos J, Binley JM, Olson WC, Allaway GP, ChengMayer C, Robinson J, Maddon PJ, Moore JP. CD4-dependent, antibody-sensitive interactions between HIV-1 and its co-receptor CCR-5. Nature 1996;384: Rizzuto CD, Wyatt R, Hernandez-Ramos N, Sun Y, Kwong PD, Hendrickson WA, Sodroski J. A conserved HIV gp120 glycoprotein structure involved in chemokine receptor binding. Science 1998;280: Kwong PD, Wyatt R, Robinson J, Sweet RW, Sodroski J, Hendrickson WA. Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody. Nature 1998;393:
23 13. Kwong PD, Wyatt R, Majeed S, Robinson J, Sweet RW, Sodroski J, Hendrickson WA. Structures of HIV-1 gp120 envelope glycoproteins from laboratory-adapted and primary isolates. Structure 2000;8: Myszka DG, Sweet RW, Hensley P, Brigham-Burke M, Kwong PD, Hendrickson WA, Wyatt R, Sodroski J, Doyle ML. Energetics of the HIV gp120-cd4 binding reaction. Proc. Natl. Acad. Sci. U. S. A. 2000;97: Rusche JR, Javaherian K, McDanal C, Petro J, Lynn DL, Grimaila R, Langlois A, Gallo RC, Arthur LO, Fischinger PJ, Bolognesi DP, Putney SD, Matthews TJ. Antibodies That Inhibit Fusion of Human Immunodeficiency Virus- Infected Cells Bind a 24-Amino Acid Sequence of the Viral Envelope, Gp120. Proc. Natl. Acad. Sci. U. S. A. 1988;85: Ghiara JB, Ferguson DC, Satterthwait AC, Dyson HJ, Wilson IA. Structure-based design of a constrained peptide mimic of the HIV-1 V3 loop neutralization site. J. Mol. Biol. 1997;266: Stanfield RL, Cabezas E, Satterthwait AC, Stura EA, Profy AT, Wilson IA. Dual conformations for the HIV-1 gp120 V3 loop in complexes with different neutralizing Fabs. Struct. Fold. Des. 1999;7: Tugarinov V, Zvi A, Levy R, Anglister J. A cis proline turn linking two beta-hairpin strands in the solution structure of an antibody-bound HIV-1(IIIB) V3 peptide. Nat. Struct. Biol. 1999;6: Tugarinov V, Zvi A, Levy R, Hayek Y, Matsushita S, Anglister J. NMR structure of an antigp120 antibody complex with a V3 peptide reveals a surface important for co-receptor binding. Struct. Fold. Des. 2000;8: Sharon M, Kessler N, Levy R, Zolla-Pazner S, Gorlach M, Anglister J. Alternative Conformations of HIV-1 V3 Loops Mimic beta Hairpins in Chemokines, Suggesting a Mechanism for Coreceptor Selectivity. Structure 2003;11: Chandrasekhar K, Profy AT, Dyson HJ. Solution Conformational Preferences of Immunogenic Peptides Derived from the Principal Neutralizing Determinant of the Hiv- 1 Envelope Glycoprotein Gp120. Biochemistry 1991;30:
24 22. Rini JM, Stanfield RL, Stura EA, Salinas PA, Profy AT, Wilson IA. Crystal-Structure of a Human-Immunodeficiency-Virus Type-1 Neutralizing Antibody, 50.1, in Complex with Its V3 Loop Peptide Antigen. Proc. Natl. Acad. Sci. U. S. A. 1993;90: Guex N, Peitsch MC. SWISS-MODEL and the Swiss-PdbViewer: An environment for comparative protein modeling. Electrophoresis 1997;18: Vranken WF, Budesinsky M, Fant F, Boulez K, Borremans FAM. The Complete Consensus V3 Loop Peptide of the Envelope Protein Gp120 of Hiv-1 Shows Pronounced Helical Character in Solution. FEBS Lett. 1995;374: Lindahl E, Hess B, van der Spoel D. GROMACS 3.0: a package for molecular simulation and trajectory analysis. J. Mol. Model. 2001;7: Daura X, Mark AE, van Gunsteren WF. Parametrization of aliphatic CHn united atoms of GROMOS96 force field. J. Comput. Chem. 1998;19: Berendsen HC, Postma JPM, van Gunsteren WF, Hermans J. Interaction models for water in relation to protein hydration. Intermolecular Forces (Pullman, B., Ed.), Reidel Publishing Company, Dordrecht pp 28. Berendsen HJC, Postma JPM, van Gunsteren WF, Di Nola A, Haak JR. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 1984;81: Tironi IG, Sperb R, Smith PE, van Gunsteren WF. A Generalized Reaction Field Method for Molecular-Dynamics Simulations. J. Chem. Phys. 1995;102: Hess B, Bekker H, Berendsen HJC, Fraaije J. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem. 1997;18: Feenstra KA, Hess B, Berendsen HJC. Improving efficiency of large time-scale molecular dynamics simulations of hydrogen-rich systems. J. Comput. Chem. 1999;20: Amadei A, Linssen ABM, Berendsen HJC. Essential Dynamics of Proteins. Proteins 1993;17: Ryu SE, Kwong PD, Truneh A, Porter TG, Arthos J, Rosenberg M, Dai XP, Xuong NH, Axel R, Sweet RW, et al. Crystal structure of an HIV-binding recombinant fragment of human CD4. Nature 1990;348:
25 34. Wang JH, Yan YW, Garrett TPJ, Liu JH, Rodgers DW, Garlick RL, Tarr GE, Husain Y, Reinherz EL, Harrison SC. Atomic-Structure of a Fragment of Human Cd4 Containing 2 Immunoglobulin-Like Domains. Nature 1990;348: Hünenberger PH, Mark AE, van Gunsteren WF. Fluctuation and Cross-Correlation Analysis of Protein Motions Observed in Nanosecond Molecular-Dynamics Simulations. J. Mol. Biol. 1995;252: Saphire EO, Parren P, Pantophlet R, Zwick MB, Morris GM, Rudd PM, Dwek RA, Stanfield RL, Burton DR, Wilson IA. Crystal structure of a neutralizing human IgG against HIV-1: A template for vaccine design. Science 2001;293: Wyatt R, Kwong PD, Desjardins E, Sweet RW, Robinson J, Hendrickson WA, Sodroski JG. The antigenic structure of the HIV gp120 envelope glycoprotein. Nature 1998;393: Nyambi PN, Gorny MK, Bastiani L, van der Groen G, Williams C, Zolla-Pazner S. Mapping of epitopes exposed on intact human immunodeficiency virus type 1 (HIV-1) virions: A new strategy for studying the immunologic relatedness of HIV-1. J. Virol. 1998;72: Gorny MK, Xu JY, Karwowska S, Buchbinder A, Zollapazner S. Repertoire of Neutralizing Human Monoclonal-Antibodies Specific for the V3 Domain of Hiv-1 Gp120. J. Immunol. 1993;150: Gorny MK, Williams C, Volsky B, Revesz K, Cohen S, Polonis VR, Honnen WJ, Kayman SC, Krachmarov C, Pinter A, Zolla-Pazner S. Human monoclonal antibodies specific for conformation-sensitive epitopes of V3 neutralize human immunodeficiency virus type 1 primary isolates from various clades. J. Virol. 2002;76: Basmaciogullari S, Babcock GJ, Van Ryk D, Wojtowicz W, Sodroski J. Identification of conserved and variable structures in the human immunodeficiency virus gp120 glycoprotein of importance for CXCR4 binding. J. Virol. 2002;76: Sakaida H, Hori T, Yonezawa A, Sato A, Isaka Y, Yoshie O, Hattori T, Uchiyama T. T-tropic human immunodeficiency virus type 1 (HIV-1) derived V3 loop peptides directly bind to CXCR-4 and inhibit T-tropic HIV- 1 infection. J. Virol. 1998;72: Verrier F, Borman AM, Brand D, Girard M. Role of the HIV type 1 glycoprotein 120 V3 loop in determining coreceptor usage. Aids Res. Hum. Retrovir. 1999;15:
26 44. Ghiara JB, Stura EA, Stanfield RL, Profy AT, Wilson IA. Crystal-Structure of the Principal Neutralization Site of Hiv-1. Science 1994;264: Dragic T, Trkola A, Lin SW, Nagashima KA, Kajumo F, Zhao L, Olson WC, Wu LJ, Mackay CR, Allaway GP, Sakmar TP, Moore JP, Maddon PJ. Amino-terminal substitutions in the CCR5 coreceptor impair gp120 binding and human immunodeficiency virus type 1 entry. J. Virol. 1998;72: Farzan M, Choe H, Vaca L, Martin K, Sun Y, Desjardins E, Ruffing N, Wu LJ, Wyatt R, Gerard N, Gerard C, Sodroski J. A tyrosine-rich region in the N terminus of CCR5 is important for human immunodeficiency virus type 1 entry and mediates an association between gp120 and CCR5. J. Virol. 1998;72: Rabut GEE, Konner JA, Kajumo F, Moore JP, Dragic T. Alanine substitutions of polar and nonpolar residues in the amino-terminal domain of CCR5 differently impair entry of macrophage- and dualtropic isolates of human immunodeficiency virus type 1. J. Virol. 1998;72: Kwong PD, Doyle ML, Casper DJ, Cicala C, Leavitt SA, Majeed S, Steenbeke TD, Venturi M, Chaiken I, Fung M, Katinger H, Parren P, Robinson J, Van Ryk D, Wang LP, Burton DR, Freire E, Wyatt R, Sodroski J, Hendrickson WA, Arthos J. HIV-1 evades antibody-mediated neutralization through conformational masking of receptor-binding sites. Nature 2002;420: Tsai CJ, Kumar S, Ma BY, Nussinov R. Folding funnels, binding funnels, and protein function. Protein Sci. 1999;8: Zwick MB, Kelleher R, Jensen R, Labrijn AF, Wang M, Quinnan GV, Parren P, Burton DR. A novel human antibody against human immunodeficiency virus type 1 gp120 is V1, V2, and V3 loop dependent and helps delimit the epitope of the broadly neutralizing antibody immunoglobulin G1 b12. J. Virol. 2003;77: Roben P, Moore JP, Thali M, Sodroski J, Barbas CF, Burton DR. Recognition Properties of a Panel of Human Recombinant Fab Fragments to the Cd4 Binding-Site of Gp120 That Show Differing Abilities to Neutralize Human-Immunodeficiency-Virus Type-1. J. Virol. 1994;68:
27 52. Sattentau QJ, Moore JP. Human-Immunodeficiency-Virus Type-1 Neutralization Is Determined by Epitope Exposure on the Gp120 Oligomer. J. Exp. Med. 1995;182: Xiang SH, Kwong PD, Gupta R, Rizzuto CD, Casper DJ, Wyatt R, Wang LP, Hendrickson WA, Doyle ML, Sodroski J. Mutagenic stabilization and/or disruption of a CD4-bound state reveals distinct conformations of the human immunodeficiency virus type 1 gp120 envelope glycoprotein. J. Virol. 2002;76: Eckert DM, Kim PS. Mechanisms of viral membrane fusion and its inhibition. Annu. Rev. Biochem. 2001;70: Kraulis PJ. Molscript - a Program to Produce Both Detailed and Schematic Plots of Protein Structures. J. Appl. Crystallogr. 1991;24: Merritt EA, Bacon DJ. Raster3D: Photorealistic molecular graphics. Macromolecular Crystallography, Pt B, pp 57. Koradi R, Billeter M, Wüthrich K. MOLMOL: a program for display and analysis of macromolecular structures. J. Mol. Graphics 1996;14: Nicholls A, Sharp KA, Honig B. Protein Folding and Association - Insights from the Interfacial and Thermodynamic Properties of Hydrocarbons. Proteins 1991;11:
28 Figure legends Figure 1. Primary sequence of the truncated form of LAI strain with the core structure and the V3 loop used in the current MD simulation with the numbering system according to the HXBc2 isolate. The secondary structure elements are numbered according to the X-ray core structure (PDB entry 1G9M, chain G). 12 The truncated V3 loop and the disordered V4 loop that are absent in the coordinates of HXBc2 gp120 are indicated by dashes along the sequence of HXBc2. The truncated N- and C-termini are omitted for clarity. Black blocks depict differences in sequence composition of LAI versus HXBc2 that are present in the simulations Figure 2. Time evolution of the RMSD from the starting structure during the 10 ns simulations of gp120 and CD4. A. Free-gp120; B. gp120 in Cplx; C. Free-CD4; D. CD4-D1 in Cplx. The backbone (N, C and Ca) RMSDs of the secondary structure elements defined in the crystal structures and the complete backbone are shown in solid and dashed lines, respectively. The all atom (including hydrogen atoms) RMSDs are shown in gray lines. The RMSDs were calculated every 2 ps after superimposition on the backbone of the secondary structure element and are plotted as a running average over a 50 ps window. Figure 3. Snapshots of the MD simulations. Left: Free gp120; Right: Gp120 in complex with CD4 D1 domain. The backbone Ca traces are taken every 200 ps from the MD trajectories between 2-10 ns. The structures are superimposed on the secondary structure backbones of gp120. Gp120 is colored from black at the N- terminus to light gray at the C-terminus. CD4-D1 structure ensemble from the Cplx simulation is shown in white. Loops that showed significant differences in 28
29 conformation and/or flexibility are indicated (C3 is composed of b15 and a3; C4 is composed of b20 and b21, which form the four-stranded bridging sheet with b2 and b3 in the V1/V2 stem). Figures were generated using Molscript 55 and Raster3D. 56 Figure 4. Comparison of gp120 Ca atoms temperature factors (B-factors) calculated from the RMS fluctuations in the 2-10 ns simulations of Free-gp120 (dashed lines) and Cplx (solid lines). The B-factors for CD4-D1 are shown in insets at a different scale. Experimental B-factors taken from the ternary gp120/cd4/cd4i crystal structure (PDB entry 1G9M) 12 are shown in gray area. The B-factors of CD4-D1 in the free form (light gray area overlaid onto the complex one) are taken from the free CD4 structure (PDB entry 3CD4) 34. They are comparable to the B-factors in the complex except for the Phe43 binding site Note that the V3 loop was truncated and the electron density of the V4 loop was missing. Residues forming stable intermolecular hydrogen bonds in the complex form are indicated with open circles along the horizontal axes (for definitions, see Table 3). Figure 5. Mapping of the intermolecular hydrogen bond network onto the intermolecular contact interfaces. Left panel: The contact van der Waals surfaces of gp120 and CD4-D1. Heavy atoms of gp120 and CD4-D1 that have intermolecular distances within 5 Å in the starting structure are colored red. The side chain of CD4- Phe43 is shown in yellow and as sticks in gp120 to indicate the binding cavity. Right panel: Residues involved in intermolecular hydrogen bonds. The types of hydrogen bonding is color-coded as: gold: side chain-side chain hydrogen bond; orange: side chain-side chain and/or side chain-backbone hydrogen bond; red: backbone-backbone and/or side chain-backbone hydrogen bonds (for details, see Table 3). Residues that 29
30 form the receptive CD4-Phe43 cavity (E370, N425, W427 and G473) are colored green, except G473, which is also involved in intermolecular backbone-backbone hydrogen bonding. Note that several residues that do not have intermolecular contacts in the starting structure form intermolecular hydrogen bonds during the molecaulr dynamics simulations (gp120: K97, V127 and A129; CD4: K22, D53 and E87). The figures were generated using MolMol. 57 Figure 6. Pair-wise backbone RMSD matrix of the bridging sheet in the free (lower right panel) and the complex form (upper left panel). The four b-strands are defined as b2: ; b3: ; b20: ; b21: Each colored dot represents a positional RMSD between two conformations taken from the respective trajectories indicated on the axes and is color-coded accordingly to the scale shown on the right. The RMSDs were calculated on backbone atoms of the bridging sheet (b2, b3, b20 and b21) after superimposition on the backbone atoms of b2 and b3. Conformations were taken every 10 ps. An equilibrated conformational sampling period is found when an off-diagonal region shows a continuous low RMSD (blue to green). Figure 7. Electrostatic potential surface of gp120. A. Free gp120. B Gp120 in complex with CD4-D1. CD4-D1 is labeled and is shown in van der Waals surface. C. Gp120 in the complex form without CD4-D1 and the V3 loop. The core structure of gp120 is identical to that in B and CD4-D1 and the V3 loop were manually removed from the complex structure. The red and blue surfaces correspond to electrostatic potentials of 2 and +2 kt/e, respectively. They were calculated using GRASP 58 with protein and solvent dielectric constants of 2 and 80, respectively. The ionic strength was set to zero. The structures are taken from the snapshots at 6 ns both in the free 30
31 and complex trajectories, which represent the most populated configuration of the V3 loop based on RMSD matrix analysis. The CD4 and CCR5 binding sites are indicated by yellow and pink arrows, respectively. Figure 8. Secondary structure evolutions of the V3 loop (residues ) as a function of the simulation time. The position of the conserved residues, GPGR, is indicated at the right end of each diagram. Figure 9. CD4 binding-induced conformational changes of gp120 extracted via essential dynamics analysis. A. Projections of the total atomic fluctuations of the merged trajectory (free:0-8 ns and complex: 8-16 ns) along the selected eigenvectors. The eigenvectors are numbered with respect to the amplitude of displacement in descending order. B. Histograms of the distributions of the displacement of each eigenvector. Distinct distribution can only be found in the first eigenvector. Note that the scale is different for the 50 th eigenvector (bottom row). C. Projections of the two extremes of the first eigenvector along the merged trajectory onto the average structure. The linear extrapolation between the two extremes is colored from blue (free) to red (complex) to highlight the primary structural differences between the two states. The C3 and C4 regions of gp120 that are part of the Phe43 cavity show significant closure of their lid upon CD4 binding. The V3 loop shows the largest conformational change. The view from the back side of the structure after 180º rotation (bottom) reveals propagated structural perturbations at the interior of gp120, involving b17, which is adjacent to the Phe43 cavity as well as the two b-strands, b12 and b13 that connect the V3 loop. 31
32 Table 1. Structural statistics (2-10 ns) Average backbone RMSD a with respect to the starting structure (nm) Gp120 Gp120 CD4 CD4 Gp120+CD4 Gp120+CD4 all bb 2º bb all bb 2º bb all bb 2º bb Free 0.42 (0.02) 0.16 (0.02) 0.10 (0.02) 0.08 (0.01) Cplx 0.37 (0.02) 0.14 (0.01) 0.11 (0.01) 0.09 (0.01) 0.40 (0.02) 0.35 (0.02) Secondary structure elements statistics b (number of residues) b sheet a helix Turn Gp120 initial c Free 96(7) 56(3) 12 (4) Cplx 111(7) 50(2) 13 (3) CD4 initial c Free 47(3) 5(1) 11 (3) Cplx 47(4) 6(2) 12 (3) a. The backbone (bb) positional RMSD values were calculated with respect to the starting structure (based on PDB entry 1G9M, 12 see Materials and Methods) after superposition on the respective secondary structure elements (2º) as defined by DSSP ( The high RMSDs of gp120+cd4 in the complex are due to relative motions of the two proteins while separate analysis of the individual molecule shows small deviations from the starting crystal structure. The larger deviations of the overall backbone in gp120 is due to the high mobility of the V3 loop. Standard deviations are indicated in parentheses. b. The secondary structure content was calculated with DSSP excluding the modeled V3 and V4 loops. c. The total numbers of residues of gp120 and CD4 in our simulations are 346 and 99, respectively (cf. Material and Methods). 32
33 Table 2. Statistics of non-bonded energies a (2-10 ns) Coulomb's electrostatic energy (kj/mol) Free gp120 Gp120 internal (363) Gp120-solvent (674) Total (766) Free CD4 CD4 internal (314) CD4-solvent (497) Total (588) Gp120/CD4 complex Gp120 internal (416) Gp120-solvent (699) CD4 internal (289) CD4-solvent (503) CD4-gp (177) Total (1015) Lennard-Jones (van der Waals) energy (kj/mol) Free gp120 Gp120 internal (115) Gp120-solvent (144) Total (184) Free CD4 CD4 internal (51) CD4-solvent -671 (78) Total (93) Gp120/CD4 complex Gp120 internal (103) Gp120-solvent (137) CD4 internal (50) CD4-solvent -477 (25) CD4-gp (76) Total (196) a. The non-bonded energies were calculated with the GROMOS96 force field 26 using a twin range cutoff of 0.8 and 1.4 nm with a reaction field correction (see Material and Methods). The energies are the sum of short (SR) and long range (LR) terms; 1-4 terms were not included. Standard deviations are indicated in parentheses. 33
34 Table 3 Intermolecular hydrogen bond and salt bridge statistics a (2-10 ns) Intermolecular hydrogen bond CD4 Gp120 Occurrence (%) Backbone - backbone F43 N G473 O 11.5 L44 O D368 N 67.8 K46 N S365 O 14.2 K46 N G366 O 15.7 Side chain backbone K35 Nz A281 O 15.9 S42 Og Q428 O 69.9 S42 Og E429 O 61.5 S42 Og E429 N 31.2 L44 N D368 Od 92.6 D53 Od2 G367 N 11.1 R59 Nh1 E429 O 45.9 R59 Nh1 V430 O 55.2 Q64 Ne2 C126 O 92.4 Q64 Ne2 V127 O 18.0 Q64 Ne2 A129 b O 29.3 N66 Nd2 A129 b O 25.8 Side chain side chain K22 Nz E429 Cd 77.8 Q25 Ne2 D474 Od 93.8 K29 Nz N280 Od 83.6 R59 Ne D368 Od 80.7 Q64 Ne2 S131 Og 15.2 E87 Cd K97 Nz 61.1 D88 Cg K97 Nz 14.8 a. Intermolecular hydrogen bonds and salt bridges (side chain-side chain hydrogen bonds) occurring for more than 10% during the 2-10 ns period are listed. A hydrogen bond is considered to exist when the donor-hydrogenacceptor angle is larger than 120º and the donor-acceptor distance is smaller than 0.28 nm. b. A129 is one of the linker residues GAG that were used in the crystal construct (1G9M) to replace the V1/V2 loops ( )
35 Table 4. Statistics of CD4 Phe43-gp120 cavity native contacts (%) a (2-10 ns) Residue Atom Occurrence E370 Cg E370 Cd 99.5 N425 Cb 94.1 N425 C 94.1 N425 O 99.0 W427 N 99.1 W427 Ca W427 Cb 83.2 G473 Ca 99.2 G473 C 98.9 G473 O a. Eleven heavy atoms of gp120 have interatomic distances with any atom of the CD4 Phe43 side chain within 0.35 nm in the crystal structure (1G9M). 12 A native contact during the simulation is considered to exist if the distance between the listed atoms and the center of the Phe43 phenyl ring is less than 0.6 nm. 35
36 LAI 83 EVVLVNVTEN FNMWKNDMVE QMHEDIISLW DQSLKPCVKL TPLCVGAGSC NTSVITQACP 206 HxBc2 83 EVVLVNVTEN FNMWKNDMVE QMHEDIISLW DQSLKPCVKL TPLCVGAGSC NTSVITQACP 206 LAI 207 KVSFEPIPIH YCAPAGFAIL KCNNKTFNGT GPCTNVSTVQ CTHGIRPVVS TQLLLNGSLA 266 HxBc2 207 KVSFEPIPIH YCAPAGFAIL KCNNKTFNGT GPCTNVSTVQ CTHGIRPVVS TQLLLNGSLA 266 LAI 267 EEEVVIRSAN FTDNAKTIIV QLNQSVEINC TRPNNNTRKS IRIQRGPGRA FVTIGKIGNM 326 HxBc2 267 EEEVVIRSVN FTDNAKTIIV QLNTSVEINC T GAG LAI 327 RQAHCNISRA KWNATLKQIA SKLREQFGNN KTIIFKQSSG GDPEIVTHSF NCGGEFFYCN 386 HxBc HCNISRA KWNNTLKQIA SKLREQFGNN KTIIFKQSSG GDPEIVTHSF NCGGEFFYCN 386 α4 β13 β1 α2 LAI 387 STQLFNSTWF NSTWSTEGSN NTEGSDTITL PCRIKQFINM WQEVGKAMYA PPISGQIRCS 446 HxBc2 387 STQLFNSTWF N GSDTITL PCRIKQIINM WQKVGKAMYA PPISGQIRCS 446 LAI 447 SNITGLLLTR DGGNNNNGSE IFRPGGGDMR DNWRSELYKY KVVKIE 492 HxBc2 447 SNITGLLLTR DGGNSNNESE IFRPGGGDMR DNWRSELYKY KVVKIE 492 α1 β4 β5 β6 β7 β8 β9 β10 β11 β12 β14 β15 α3 β16 β17 β18 V4 β19 β20 β21 β22 β23 V5 β24 α5 β25 β2 V1/V2 stem V3 β3 Figure 1 Hsu & Bonvin
37 gp120 free RMSD (nm) A B gp120 cplx RMSD (nm) CD4 free RMSD (nm) C D CD4 cplx RMSD (nm) Time (ps) Time (ps) Figure 2 Hsu & Bonvin
38 D V5 V5 D CD4 N C C3 C4 N C C4 V1/V2 stem V1/V2 stem Free V3 Cplx V3 Figure 3 Hsu & Bonvin
39
40 K97 Q428 W427 G473 E429 V127 A129 A281 N280 D474 G126 N425 S365 G366 G367 E370 D368 o 180 E87 K29 K22 Q25 K35 S42 F43 Q64 L44 K46 R59 D53 Figure 5 Hsu & Bonvin
41 Cplx RMSD (nm) ns 0 Free 10 Figure 6 Hsu & Bonvin
42 A B CD4 C V3 V3 Figure 7 Hsu & Bonvin
43 330 Cplx Free GPRG GPRG Time (ns) Figure 8 Hsu & Bonvin
44 A displacement(nm) displacement(nm) displacement(nm) displacement(nm) displacement(nm) eigenvector eigenvector eigenvector eigenvector eigenvector time(ps) Frequency Frequency Frequency Frequency Frequency B cplx free displacement(nm) C V1/V2 stem Cplx Free V4 C CD4 binding site b12/ b13 N C4 bridging sheet b17 V3 N V5 C3 V3 C V4 180 o V1/V2 stem Figure 9 Hsu & Bonvin
Entropy Calculation of HIV-1 Env gp120, its Receptor CD4, and their Complex: An Analysis of Configurational Entropy Changes upon Complexation
 Biophysical Journal Volume 88 January 2005 15 24 15 Entropy Calculation of HIV-1 Env gp120, its Receptor CD4, and their Complex: An Analysis of Configurational Entropy Changes upon Complexation Shang-Te
Biophysical Journal Volume 88 January 2005 15 24 15 Entropy Calculation of HIV-1 Env gp120, its Receptor CD4, and their Complex: An Analysis of Configurational Entropy Changes upon Complexation Shang-Te
HIV Anti-HIV Neutralizing Antibodies
 ,**/ The Japanese Society for AIDS Research The Journal of AIDS Research : HIV HIV Anti-HIV Neutralizing Antibodies * Junji SHIBATA and Shuzo MATSUSHITA * Division of Clinical Retrovirology and Infectious
,**/ The Japanese Society for AIDS Research The Journal of AIDS Research : HIV HIV Anti-HIV Neutralizing Antibodies * Junji SHIBATA and Shuzo MATSUSHITA * Division of Clinical Retrovirology and Infectious
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
From Antibody to Vaccine a Tale of Structural Biology and Epitope Scaffolds
 Dale and Betty Bumpers Vaccine Research Center National Institute of Allergy and Infectious Diseases National Institutes of Health Department of Health and Human Services From Antibody to Vaccine a Tale
Dale and Betty Bumpers Vaccine Research Center National Institute of Allergy and Infectious Diseases National Institutes of Health Department of Health and Human Services From Antibody to Vaccine a Tale
Supplementary Information
 Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Low ds/dn Does Not Correlate With High Variation of Amino Acid Sequences Along the gp120 Protein Structure
 Low ds/dn Does Not Correlate With High Variation of Amino Acid Sequences Along the gp120 Protein Structure Zach Goldstein & Jordan Detamore BIOL 368: Bioinformatics Laboratory Department of Biology Loyola
Low ds/dn Does Not Correlate With High Variation of Amino Acid Sequences Along the gp120 Protein Structure Zach Goldstein & Jordan Detamore BIOL 368: Bioinformatics Laboratory Department of Biology Loyola
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Structural Insights into HIV-1 Neutralization by Broadly Neutralizing Antibodies PG9 and PG16
 Structural Insights into HIV-1 Neutralization by Broadly Neutralizing Antibodies PG9 and PG16 Robert Pejchal, Laura M. Walker, Robyn L. Stanfield, Wayne C. Koff, Sanjay K. Phogat, Pascal Poignard, Dennis
Structural Insights into HIV-1 Neutralization by Broadly Neutralizing Antibodies PG9 and PG16 Robert Pejchal, Laura M. Walker, Robyn L. Stanfield, Wayne C. Koff, Sanjay K. Phogat, Pascal Poignard, Dennis
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Bioinformatics for molecular biology
 Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
YUMI YAMAGUCHI-KABATA AND TAKASHI GOJOBORI* Center for Information Biology, National Institute of Genetics, Mishima , Japan
 JOURNAL OF VIROLOGY, May 2000, p. 4335 4350 Vol. 74, No. 9 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Reevaluation of Amino Acid Variability of the Human
JOURNAL OF VIROLOGY, May 2000, p. 4335 4350 Vol. 74, No. 9 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Reevaluation of Amino Acid Variability of the Human
Conformational Flexibility of the Peptide Hormone Ghrelin in Solution and Lipid Membrane Bound: A Molecular Dynamics Study
 Open Access Article The authors, the publisher, and the right holders grant the right to use, reproduce, and disseminate the work in digital form to all users. Journal of Biomolecular Structure & Dynamics,
Open Access Article The authors, the publisher, and the right holders grant the right to use, reproduce, and disseminate the work in digital form to all users. Journal of Biomolecular Structure & Dynamics,
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody
 CD4 binding induces conformational changes in the gp120 glycoprotein, some of which involve the exposure and/or formation of a binding site for specific chemokine receptors. These chemokine receptors,
CD4 binding induces conformational changes in the gp120 glycoprotein, some of which involve the exposure and/or formation of a binding site for specific chemokine receptors. These chemokine receptors,
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B
 3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
Identification of Mutation(s) in. Associated with Neutralization Resistance. Miah Blomquist
 Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
Chapter VI. Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase
 Chapter VI Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase Chapter VI Abstract We report the study of T12A Cyt c CcP complex using isothermal titration calorimetry (ITC),
Chapter VI Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase Chapter VI Abstract We report the study of T12A Cyt c CcP complex using isothermal titration calorimetry (ITC),
CD4 T Cell Decline Is Not Associated With Amino Acid Changes in HIV-1 gp120
 CD4 T Cell Decline Is Not Associated With Amino Acid Changes in HIV-1 gp120 Colin Wikholm and Isai Lopez BIOL 368: Bioinformatics Laboratory Department of Biology Loyola Marymount University November 15,
CD4 T Cell Decline Is Not Associated With Amino Acid Changes in HIV-1 gp120 Colin Wikholm and Isai Lopez BIOL 368: Bioinformatics Laboratory Department of Biology Loyola Marymount University November 15,
Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen
 Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen David Lutje Hulsik Unit for Virus Host Cell Interaction UMI 3265 University Joseph Fourier-EMBL-CNRS, Grenoble Env catalyzed
Crystal structure of the neutralizing antibody HK20 in complex with its gp41 antigen David Lutje Hulsik Unit for Virus Host Cell Interaction UMI 3265 University Joseph Fourier-EMBL-CNRS, Grenoble Env catalyzed
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
Molecular Dynamics of HIV-1 Reverse Transcriptase
 Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
Pacific Symposium on Biocomputing 12:40-50(2007)
 PREDICTING STRUCTURE AND DYNAMICS OF LOOSELY- ORDERED PROTEIN COMPLEXES: INFLUENZA HEMAGGLUTININ FUSION PEPTIDE PETER M. KASSON Medical Scientist Training Program Stanford University, Stanford CA 94305
PREDICTING STRUCTURE AND DYNAMICS OF LOOSELY- ORDERED PROTEIN COMPLEXES: INFLUENZA HEMAGGLUTININ FUSION PEPTIDE PETER M. KASSON Medical Scientist Training Program Stanford University, Stanford CA 94305
EMERGING ISSUES IN THE HUMORAL IMMUNE RESPONSE TO HIV. (Summary of the recommendations from an Enterprise Working Group)
 AIDS Vaccine 07, Seattle, August 20-23, 2007 EMERGING ISSUES IN THE HUMORAL IMMUNE RESPONSE TO HIV (Summary of the recommendations from an Enterprise Working Group) The Working Group Reston, Virginia,
AIDS Vaccine 07, Seattle, August 20-23, 2007 EMERGING ISSUES IN THE HUMORAL IMMUNE RESPONSE TO HIV (Summary of the recommendations from an Enterprise Working Group) The Working Group Reston, Virginia,
Coarse grained simulations of Lipid Bilayer Membranes
 Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
On the conformational properties of amylose and cellulose oligomers in solution
 On the conformational properties of amylose and cellulose oligomers in solution Author Winger, Moritz, Christen, Markus, F. van Gunsteren, Wilfred Published 29 Journal Title International Journal of Carbohydrate
On the conformational properties of amylose and cellulose oligomers in solution Author Winger, Moritz, Christen, Markus, F. van Gunsteren, Wilfred Published 29 Journal Title International Journal of Carbohydrate
Supporting Information
 Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Lecture 10 More about proteins
 Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
a) The statement is true for X = 400, but false for X = 300; b) The statement is true for X = 300, but false for X = 200;
 1. Consider the following statement. To produce one molecule of each possible kind of polypeptide chain, X amino acids in length, would require more atoms than exist in the universe. Given the size of
1. Consider the following statement. To produce one molecule of each possible kind of polypeptide chain, X amino acids in length, would require more atoms than exist in the universe. Given the size of
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study
 Biophysical Journal Volume 96 February 2009 785 797 785 Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study Charles H. Davis and Max L. Berkowitz * Department
Biophysical Journal Volume 96 February 2009 785 797 785 Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study Charles H. Davis and Max L. Berkowitz * Department
proteins Fluctuation dynamics analysis of gp120 envelope protein reveals a topologically based communication network
 proteins STRUCTURE O FUNCTION O BIOINFORMATICS Fluctuation dynamics analysis of gp120 envelope protein reveals a topologically based communication network Indira Shrivastava 1 and Judith M. LaLonde 2 1
proteins STRUCTURE O FUNCTION O BIOINFORMATICS Fluctuation dynamics analysis of gp120 envelope protein reveals a topologically based communication network Indira Shrivastava 1 and Judith M. LaLonde 2 1
We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field
 Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Antigen Receptor Structures October 14, Ram Savan
 Antigen Receptor Structures October 14, 2016 Ram Savan savanram@uw.edu 441 Lecture #8 Slide 1 of 28 Three lectures on antigen receptors Part 1 (Today): Structural features of the BCR and TCR Janeway Chapter
Antigen Receptor Structures October 14, 2016 Ram Savan savanram@uw.edu 441 Lecture #8 Slide 1 of 28 Three lectures on antigen receptors Part 1 (Today): Structural features of the BCR and TCR Janeway Chapter
Structural Plasticity and Conformational Transitions of HIV Envelope Glycoprotein gp120
 Structural Plasticity and Conformational Transitions of HIV Envelope Glycoprotein gp120 Anil Korkut 1, Wayne A. Hendrickson 1,2,3 * 1 Department of Biochemistry and Molecular Biophysics, Columbia University,
Structural Plasticity and Conformational Transitions of HIV Envelope Glycoprotein gp120 Anil Korkut 1, Wayne A. Hendrickson 1,2,3 * 1 Department of Biochemistry and Molecular Biophysics, Columbia University,
BCH Graduate Survey of Biochemistry
 BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 10 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html
BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 10 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html
Supplementary information for Effects of Stretching Speed on. Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular
 Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Protein Secondary Structure
 Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
PHAR3316 Pharmacy biochemistry Exam #2 Fall 2010 KEY
 1. How many protons is(are) lost when the amino acid Asparagine is titrated from its fully protonated state to a fully deprotonated state? A. 0 B. 1 * C. 2 D. 3 E. none Correct Answer: C (this question
1. How many protons is(are) lost when the amino acid Asparagine is titrated from its fully protonated state to a fully deprotonated state? A. 0 B. 1 * C. 2 D. 3 E. none Correct Answer: C (this question
Lesson 5 Proteins Levels of Protein Structure
 Lesson 5 Proteins Levels of Protein Structure Primary 1º Structure The primary structure is simply the sequence of amino acids in a protein. Chains of amino acids are written from the amino terminus (N-terminus)
Lesson 5 Proteins Levels of Protein Structure Primary 1º Structure The primary structure is simply the sequence of amino acids in a protein. Chains of amino acids are written from the amino terminus (N-terminus)
Analysis of protein modeling for envelope glycoprotein GP120 for HIV via bioinformatics approaches
 International Research Journal of Virology Vol. 1(1), pp. 002-006, March, 2014. www.premierpublishers.org ISSN: 2326-7193x IRJV Research Article Analysis of protein modeling for envelope glycoprotein GP120
International Research Journal of Virology Vol. 1(1), pp. 002-006, March, 2014. www.premierpublishers.org ISSN: 2326-7193x IRJV Research Article Analysis of protein modeling for envelope glycoprotein GP120
Dynamic contact network between ribosomal subunits enables rapid large-scale rotation during spontaneous translocation
 Published online 24 June 2015 Nucleic Acids Research, 2015, Vol. 43, No. 14 6747 6760 doi: 10.1093/nar/gkv649 Dynamic contact network between ribosomal subunits enables rapid large-scale rotation during
Published online 24 June 2015 Nucleic Acids Research, 2015, Vol. 43, No. 14 6747 6760 doi: 10.1093/nar/gkv649 Dynamic contact network between ribosomal subunits enables rapid large-scale rotation during
SAM Teacher s Guide Four Levels of Protein Structure
 SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
Citation for published version (APA): Von Eije, K. J. (2009). RNAi based gene therapy for HIV-1, from bench to bedside
 UvA-DARE (Digital Academic Repository) RNAi based gene therapy for HIV-1, from bench to bedside Von Eije, K.J. Link to publication Citation for published version (APA): Von Eije, K. J. (2009). RNAi based
UvA-DARE (Digital Academic Repository) RNAi based gene therapy for HIV-1, from bench to bedside Von Eije, K.J. Link to publication Citation for published version (APA): Von Eije, K. J. (2009). RNAi based
The challenge of an HIV vaccine from the antibody perspective. Dennis Burton The Scripps Research Institute
 The challenge of an HIV vaccine from the antibody perspective Dennis Burton The Scripps Research Institute AIDS Pandemic Nov 2005 North America 1.2 million [650 000 1.8 million] Caribbean 300 000 [200
The challenge of an HIV vaccine from the antibody perspective Dennis Burton The Scripps Research Institute AIDS Pandemic Nov 2005 North America 1.2 million [650 000 1.8 million] Caribbean 300 000 [200
Supporting material. Membrane permeation induced by aggregates of human islet amyloid polypeptides
 Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Inter-Species Cross-Seeding: Stability and Assembly of Rat - Human Amylin Aggregates. Workalemahu M. Berhanu
 Inter-Species Cross-Seeding: Stability and Assembly of Rat - Human Amylin Aggregates Workalemahu M. Berhanu University of Oklahoma Department of Chemistry and Biochemistry STRUCTURE OF AMYLOIDS & ROLE
Inter-Species Cross-Seeding: Stability and Assembly of Rat - Human Amylin Aggregates Workalemahu M. Berhanu University of Oklahoma Department of Chemistry and Biochemistry STRUCTURE OF AMYLOIDS & ROLE
Proteins and their structure
 Proteins and their structure Proteins are the most abundant biological macromolecules, occurring in all cells and all parts of cells. Proteins also occur in great variety; thousands of different kinds,
Proteins and their structure Proteins are the most abundant biological macromolecules, occurring in all cells and all parts of cells. Proteins also occur in great variety; thousands of different kinds,
NIH Public Access Author Manuscript J Am Chem Soc. Author manuscript; available in PMC 2008 September 29.
 NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
Entry of enveloped viruses into cells requires transformation
 Energetics of the HIV gp120-cd4 binding reaction David G. Myszka*, Raymond W. Sweet, Preston Hensley, Michael Brigham-Burke, Peter D. Kwong, Wayne A. Hendrickson **, Richard Wyatt, Joseph Sodroski, and
Energetics of the HIV gp120-cd4 binding reaction David G. Myszka*, Raymond W. Sweet, Preston Hensley, Michael Brigham-Burke, Peter D. Kwong, Wayne A. Hendrickson **, Richard Wyatt, Joseph Sodroski, and
The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method
 Journal of Physics: Conference Series PAPER OPEN ACCESS The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method To cite this article: Sri
Journal of Physics: Conference Series PAPER OPEN ACCESS The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method To cite this article: Sri
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Copyright Mark Brandt, Ph.D. 46
 Examples of tein Structures tein types teins fall into three general classes, based on their overall three-dimensional structure and on their functional role: fibrous, membrane, and globular. Fibrous proteins
Examples of tein Structures tein types teins fall into three general classes, based on their overall three-dimensional structure and on their functional role: fibrous, membrane, and globular. Fibrous proteins
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Loyola Marymount University Department of Biology October 25, Zach, Matt, Mia, Will
 Alexander M. Andrianov & Ivan V. Anishchenko (2009) Computational Model of the HIV-1 Subtype A V3 Loop: Study on the Conformational Mobility for Structure Based Anti-AIDS Drug Design, Journal of Biomolecular
Alexander M. Andrianov & Ivan V. Anishchenko (2009) Computational Model of the HIV-1 Subtype A V3 Loop: Study on the Conformational Mobility for Structure Based Anti-AIDS Drug Design, Journal of Biomolecular
Simulationen von Lipidmembranen
 Simulationen von Lipidmembranen Thomas Stockner Thomas.stockner@meduniwien.ac.at Summary Methods Force Field MD simulations Membrane simulations Application Oxidized lipids Anesthetics Molecular biology
Simulationen von Lipidmembranen Thomas Stockner Thomas.stockner@meduniwien.ac.at Summary Methods Force Field MD simulations Membrane simulations Application Oxidized lipids Anesthetics Molecular biology
The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation
 www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
Crystallization-grade After D After V3 cocktail. Time (s) Time (s) Time (s) Time (s) Time (s) Time (s)
 Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Nafith Abu Tarboush DDS, MSc, PhD
 Nafith Abu Tarboush DDS, MSc, PhD natarboush@ju.edu.jo www.facebook.com/natarboush Protein conformation Many conformations are possible for proteins due to flexibility of amino acids linked by peptide
Nafith Abu Tarboush DDS, MSc, PhD natarboush@ju.edu.jo www.facebook.com/natarboush Protein conformation Many conformations are possible for proteins due to flexibility of amino acids linked by peptide
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Proteins consist of joined amino acids They are joined by a Also called an Amide Bond
 Lecture Two: Peptide Bond & Protein Structure [Chapter 2 Berg, Tymoczko & Stryer] (Figures in Red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Proteins consist of joined amino acids
Lecture Two: Peptide Bond & Protein Structure [Chapter 2 Berg, Tymoczko & Stryer] (Figures in Red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Proteins consist of joined amino acids
Structural Flexibility and Functional Valence of CD4-IgG2 (PRO 542): Potential for Cross-Linking Human Immunodeficiency Virus Type 1 Envelope Spikes
 JOURNAL OF VIROLOGY, July 2001, p. 6682 6686 Vol. 75, No. 14 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.14.6682 6686.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Structural
JOURNAL OF VIROLOGY, July 2001, p. 6682 6686 Vol. 75, No. 14 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.14.6682 6686.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Structural
Nonsynonymous Amino Acid Mutations in gp120 Binding Sites are Related to Progression of HIV-1
 Nonsynonymous Amino Acid Mutations in gp120 Binding Sites are Related to Progression of HIV-1 Matthew Allegretti and Anindita Varshneya BIOL 368: Bioinformatics Laboratory Loyola Marymount University November
Nonsynonymous Amino Acid Mutations in gp120 Binding Sites are Related to Progression of HIV-1 Matthew Allegretti and Anindita Varshneya BIOL 368: Bioinformatics Laboratory Loyola Marymount University November
Energetics of dendrimer binding to HIV-1 gp120- CD4 complex and mechanismic aspects of its role as an entry-inhibitor
 Journal of Physics: Conference Series PAPER OPEN ACCESS Energetics of dendrimer binding to HIV-1 gp120- CD4 complex and mechanismic aspects of its role as an entry-inhibitor Related content - Charged nanoparticle
Journal of Physics: Conference Series PAPER OPEN ACCESS Energetics of dendrimer binding to HIV-1 gp120- CD4 complex and mechanismic aspects of its role as an entry-inhibitor Related content - Charged nanoparticle
In Silico Vaccine Design Based on Molecular Simulations of Rhinovirus Chimeras Presenting HIV-1 gp41 Epitopes
 doi:10.1016/j.jmb.2008.10.089 J. Mol. Biol. (2009) 385, 675 691 Available online at www.sciencedirect.com In Silico Vaccine Design Based on Molecular Simulations of Rhinovirus Chimeras Presenting HIV-1
doi:10.1016/j.jmb.2008.10.089 J. Mol. Biol. (2009) 385, 675 691 Available online at www.sciencedirect.com In Silico Vaccine Design Based on Molecular Simulations of Rhinovirus Chimeras Presenting HIV-1
Predicting antigenic sites of hemagglutinin-neuraminidase glycoprotein Newcastle disease virus
 Romanian Biotechnological Letters Vol. 14, No.4, 2009, pp. 4589-4596 Copyright 2008 Bucharest University Printed in Romania. All rights reserved Romanian Society of Biological Sciences ORIGINAL PAPER Predicting
Romanian Biotechnological Letters Vol. 14, No.4, 2009, pp. 4589-4596 Copyright 2008 Bucharest University Printed in Romania. All rights reserved Romanian Society of Biological Sciences ORIGINAL PAPER Predicting
Ionization of amino acids
 Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
2005 LANDES BIOSCIENCE. DO NOT DISTRIBUTE.
 [Human Vaccines 1:2, 45-60; March/April 2005]; 2005 Landes Bioscience Review Role of Neutralizing Antibodies in Protective Immunity Against HIV Indresh K. Srivastava* Jeffrey B. Ulmer Susan W. Barnett
[Human Vaccines 1:2, 45-60; March/April 2005]; 2005 Landes Bioscience Review Role of Neutralizing Antibodies in Protective Immunity Against HIV Indresh K. Srivastava* Jeffrey B. Ulmer Susan W. Barnett
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Antigen Recognition by T cells
 Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
Antigen Recognition by T cells TCR only recognize foreign Ags displayed on cell surface These Ags can derive from pathogens, which replicate within cells or from pathogens or their products that cells
H C. C α. Proteins perform a vast array of biological function including: Side chain
 Topics The topics: basic concepts of molecular biology elements on Python overview of the field biological databases and database searching sequence alignments phylogenetic trees microarray data analysis
Topics The topics: basic concepts of molecular biology elements on Python overview of the field biological databases and database searching sequence alignments phylogenetic trees microarray data analysis
Molecular Mechanics Studies of Enzyme Evolutionary Mechanisms
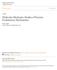 Clemson University TigerPrints All Dissertations Dissertations 1-2010 Molecular Mechanics Studies of Enzyme Evolutionary Mechanisms Manoj Singh Clemson University, mksingh@clemson.edu Follow this and additional
Clemson University TigerPrints All Dissertations Dissertations 1-2010 Molecular Mechanics Studies of Enzyme Evolutionary Mechanisms Manoj Singh Clemson University, mksingh@clemson.edu Follow this and additional
Rama Abbady. Odai Bani-Monia. Diala Abu-Hassan
 5 Rama Abbady Odai Bani-Monia Diala Abu-Hassan Lipid Rafts Lipid rafts are aggregates (accumulations) of sphingolipids. They re semisolid clusters (10-200 nm) of cholesterol and sphingolipids (sphingomyelin
5 Rama Abbady Odai Bani-Monia Diala Abu-Hassan Lipid Rafts Lipid rafts are aggregates (accumulations) of sphingolipids. They re semisolid clusters (10-200 nm) of cholesterol and sphingolipids (sphingomyelin
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes
 Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Secondary Structure North 72nd Street, Wauwatosa, WI Phone: (414) Fax: (414) dmoleculardesigns.com
 Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
Proteins. (b) Protein Structure and Conformational Change
 Proteins (b) Protein Structure and Conformational Change Protein Structure and Conformational Change Proteins contain the elements carbon (C), hydrogen (H), oxygen (O2) and nitrogen (N2) Some may also
Proteins (b) Protein Structure and Conformational Change Protein Structure and Conformational Change Proteins contain the elements carbon (C), hydrogen (H), oxygen (O2) and nitrogen (N2) Some may also
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature19102 Supplementary Discussion Benzothiazepine Binding in Ca V Ab Diltiazem and other benzothiazepines inhibit Ca V 1.2 channels in a frequency-dependent manner consistent with pore block
doi:10.1038/nature19102 Supplementary Discussion Benzothiazepine Binding in Ca V Ab Diltiazem and other benzothiazepines inhibit Ca V 1.2 channels in a frequency-dependent manner consistent with pore block
MBB 694:407, 115:511. Please use BLOCK CAPITAL letters like this --- A, B, C, D, E. Not lowercase!
 MBB 694:407, 115:511 First Test Severinov/Deis Tue. Sep. 30, 2003 Name Index number (not SSN) Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions
MBB 694:407, 115:511 First Test Severinov/Deis Tue. Sep. 30, 2003 Name Index number (not SSN) Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions
Structural Analysis of TCRpMHC Complexes Using Computational Tools. Feroze Mohideen Briarcliff High School
 Structural Analysis of TCRpMHC Complexes Using Computational Tools Feroze Mohideen Briarcliff High School TCR-pMHC Complexes Peptide Structure of TCR-pMHC complex PDB AO7 Crossreactivity is the ability
Structural Analysis of TCRpMHC Complexes Using Computational Tools Feroze Mohideen Briarcliff High School TCR-pMHC Complexes Peptide Structure of TCR-pMHC complex PDB AO7 Crossreactivity is the ability
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
PHARMACOPHORE MODELING
 CHAPTER 9 PHARMACOPHORE MODELING 9.1 PHARMACOPHORES AS VIRTUAL SCREENING TOOLS The pharmacophore concept has been widely used over the past decades for rational drug design approaches and has now been
CHAPTER 9 PHARMACOPHORE MODELING 9.1 PHARMACOPHORES AS VIRTUAL SCREENING TOOLS The pharmacophore concept has been widely used over the past decades for rational drug design approaches and has now been
Supplementary Information: A Critical. Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers
 Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes.
 Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes. Alexander Kantardjiev 1 and Pavletta Shestakova 1 1 Institute of Organic Chemistry with
Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes. Alexander Kantardjiev 1 and Pavletta Shestakova 1 1 Institute of Organic Chemistry with
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
Going Nowhere Fast: Lentivirus genetic sequence evolution does not correlate with phenotypic evolution.
 Going Nowhere Fast: Lentivirus genetic sequence evolution does not correlate with phenotypic evolution. Brian T. Foley, PhD btf@lanl.gov HIV Genetic Sequences, Immunology, Drug Resistance and Vaccine Trials
Going Nowhere Fast: Lentivirus genetic sequence evolution does not correlate with phenotypic evolution. Brian T. Foley, PhD btf@lanl.gov HIV Genetic Sequences, Immunology, Drug Resistance and Vaccine Trials
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Study on Different types of Structure based Properties in Human Membrane Proteins
 2017 IJSRST Volume 3 Issue 7 Print ISSN: 2395-6011 Online ISSN: 2395-602X Themed Section: Science and Technology Study on Different types of Structure based Properties in Human Membrane Proteins S. C.
2017 IJSRST Volume 3 Issue 7 Print ISSN: 2395-6011 Online ISSN: 2395-602X Themed Section: Science and Technology Study on Different types of Structure based Properties in Human Membrane Proteins S. C.
ARV Mode of Action. Mode of Action. Mode of Action NRTI. Immunopaedia.org.za
 ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
Protein Modeling Event
 Protein Modeling Event School Name: School Number: Team Member 1: Team Member 2: : Pre-Build Score: On-Site Build Score: Test Score: Tie Breaker: Total: Final Rank: Part I: Pre-Build (40% of total score)
Protein Modeling Event School Name: School Number: Team Member 1: Team Member 2: : Pre-Build Score: On-Site Build Score: Test Score: Tie Breaker: Total: Final Rank: Part I: Pre-Build (40% of total score)
Supporting Information: Revisiting partition in. hydrated bilayer systems
 Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
A V3 Loop-Dependent gp120 Element Disrupted by CD4 Binding Stabilizes the Human Immunodeficiency Virus Envelope Glycoprotein Trimer
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for 4-2010 A V3 Loop-Dependent gp120 Element Disrupted by CD4 Binding Stabilizes
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for 4-2010 A V3 Loop-Dependent gp120 Element Disrupted by CD4 Binding Stabilizes
Introduction to Proteomics Dr. Sanjeeva Srivastava Department of Biosciences and Bioengineering Indian Institute of Technology - Bombay
 Introduction to Proteomics Dr. Sanjeeva Srivastava Department of Biosciences and Bioengineering Indian Institute of Technology - Bombay Lecture 01 Introduction to Amino Acids Welcome to the proteomic course.
Introduction to Proteomics Dr. Sanjeeva Srivastava Department of Biosciences and Bioengineering Indian Institute of Technology - Bombay Lecture 01 Introduction to Amino Acids Welcome to the proteomic course.
PRC2 crystal clear. Matthieu Schapira
 PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
