Names for ABO (ISBT 001) Blood Group Alleles
|
|
|
- Tamsin Mitchell
- 5 years ago
- Views:
Transcription
1 Names for ABO (ISBT 001) blood group alleles v Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically most important system. The ABO gene and its coding exons give rise to one of two principally different glycosyltransferases. The A glycosyltransferase (GTA) catalyzes the addition of a donor substrate, UDP-N-acetylgalactosamine, to an acceptor substrate known as the H antigen. The B glycosyltransferase (GTB) differs by only four amino-acid substitutions from GTA and performs the same enzymatic reaction but uses UDP-galactose as donor substrate. In this way, genetic polymorphism gives rise to two related antigens in this system. Any polymorphism or mutation that s the activity or specificity of the encoded enzyme may therefore alter the ABO phenotype. Alterations that completely abolish enzymic activity give rise to the blood group O phenotype, in which the H antigen remains unconverted and no A or B antigen can be detected. If the genetic alteration decreases the activity of the enzyme, or alters its subcellular location and thereby decreases conversion of H to A or B, a weak A or B subgroup phenotype can result. Furthermore, certain polymorphisms result in promiscuous enzymes that can synthesize both A and B antigen, thereby resulting in the so-called cisab or B(A) phenotypes. The A phenotype is divided into A1 and A2. The former is more prevalent in all populations and has approximately times more A epitopes per red cell. The GTA1 is also better than GTA2 at synthesizing certain forms of A,.e.g. A type and. In addition to the A and B antigens, two other antigens are included in the ABO system, namely A,B and A1. The former is a joint epitope on A or B antigen and is therefore present in both A, B and AB phenotypes. The exact biochemical nature of the A1 antigen has been more controversial but has been proposed to represent A type. Gene name: ABO Number of exons: Initiation codon: Within exon 1 Stop codon: Within exon Entrez Gene ID: 28 LRG sequence: NG_009.1 (genomic) NM_ (transcript) Reference allele: ABO*A1.01 (shaded) Page 1 of 1
2 Names for ABO (ISBT 001) blood group alleles v Molecular bases associated with the A1, A2 and weak A phenotypes Reference allele ABO*A1.01 encodes A glycosyltransferase that synthesizes A antigen. Phenotype Allele name Nucleotide Exon Predicted amino acid A1 ABO*A1.01 A1 ABO*A1.02 c.c>t p.pro1leu A2 ABO*A2.01 c.c>t; c.101delc p.pro1leu; A2 ABO*A2.02 c.10c>t p.arg2trp A2 ABO*A2.0 c.10c>g p.arg2gly A2 ABO*A2.0 c.29a>g; c.2c>g; c.c>t; c.0g>a; c.1c>t; p.arg1gly; p.gly2ser; p.val2met A2 ABO*A2.0 c.c>t; c.1009a>g p.pro1leu; p.arggly A2 ABO*A2.0 c.101delc A2 ABO*A2.0 c.9g>c p.arg180pro A2 ABO*A2.08 c.c>t; c.9g>c p.pro1leu; p.arg180pro A2 ABO*A2.09 c.c>t; c.2g>a; c.101delc p.pro1leu; p.arg1his; A2 ABO*A2.10 c.28t>c; c.c>t A2 ABO*A2.11 c.2c>t; c.c>t A2 ABO*A2.12 c.190g>a; c.2g>a; c.101delc p.trp90arg; p.pro1leu p.pro89leu; p.pro1leu p.valile; p.arg1his; A2 ABO*A2.1 c.c>t; c.2c>t p.pro1leu; p.arg28cys A2 ABO*A2.1 c.10g>t; c.c>t; c.101delc p.valphe; p.arghis; p.pro1leu; A2 ABO*A2.1 c.0c>t; c.c>t p.thr1met; p.pro1leu A2 ABO*A2.18 c.c>t; c.22g>a p.pro1leu; p.arg21gln A2 ABO*A2.19 c.c>t; c.8g>a p.pro1leu; p.glu20lys Page 2 of 1
3 Names for ABO (ISBT 001) blood group alleles v Phenotype Allele name Nucleotide Exon Predicted amino acid A2 ABO*A2.20 c.c>t; p.pro1leu; p.val2met A ABO*A.01 c.81g>a p.asp291asn A ABO*A.02 ; c.101delc p.val2met; A ABO*A.0 c.88c>t p.leu280phe A ABO*A.0 c.c>t; c.9g>a; c.101delc p.arg180his; p.pro1leu; A ABO*A.0 c.820g>a p.val2met A ABO*A.0 c.c>t; c.820g>a p.pro1leu; p.val2met A ABO*A.0 c.c>t; c.c>t p.pro1leu; p.arg29trp Aweak ABO*AW.01 c.0c>t; c.c>t; c.101delc p.thr1met; p.pro1leu; Aweak ABO*AW.02 c.0g>c; c.c>t; c.101delc Aweak ABO*AW.0 c.20g>c; c.c>t; c.101delc p.gly11ala; p.pro1leu; p.arg8thr; p.pro1leu; Aweak ABO*AW.0 c.21c>t p.arg21trp Aweak ABO*AW.0 c.9a>g p.glu22gly Aweak ABO*AW.0 c.02c>g p.arg18gly Aweak ABO*AW.0 c.c>t; c.92c>t; c.101delc p.pro1leu; p.arg198trp; Aweak ABO*AW.08 c.29a>g; c.88c>t; c.2c>g; c.802g>a Aweak ABO*AW.09 c.g>a; c.10g>t; c.188g>a; c.c>t; c.101delc 2 p.proser; p.thr1met; p.arg1gly; p.gly28arg p.ala1thr; p.valphe; p.arghis; p.proser; p.pro1leu; Aweak ABO*AW.10 c.8g>a p.asp22asn Aweak ABO*AW.11 c.2g>a; c.21c>t p.val1met; p.arg21trp Page of 1
4 Names for ABO (ISBT 001) blood group alleles v Phenotype Allele name Nucleotide Exon Predicted amino acid Aweak ABO*AW.12 c.c>t; c.a>g p.pro1leu; p.met18val Aweak ABO*AW.1 c.2t>c 1 p.ala2_met20del Aweak ABO*AW.1 c.c>t; c.99c>a p.pro1leu; p.his2gln Aweak ABO*AW.1 c.+a>t Intron Altered splicing Aweak ABO*AW.1 c.1a>g; c.c>t; c.101delc Aweak ABO*AW.1 c.2c>t; c.c>t; c.101delc Aweak ABO*AW.18 c.t>c; c.c>t; c.101delc 1 p.ala2_met20del; p.pro1leu; p.pro9leu; p.pro1leu; p.ile11thr; p.pro1leu; Aweak ABO*AW.19 c.a>g; c.c>t; c.101delc Aweak ABO*AW.20 c.c>t; c.0g>a; c.101delc Aweak ABO*AW.21 c.c>t; c.0g>c; c.101delc Aweak ABO*AW.22 c.c>t; c.g>a; c.101delc Aweak ABO*AW.2 c.c>t; c.22g>a; c.101delc Aweak ABO*AW.2 c.c>t; c.2c>t; c.101delc Aweak ABO*AW.2 c.c>t; ; c.101delc Aweak ABO*AW.2 c.c>t; c.2g>a; c.101delc p.his1arg; p.pro1leu; p.pro1leu; p.glu20lys; p.pro1leu; p.glu20gln; p.pro1leu;val212met; p.pro1leu; p.arg21gln; p.pro1leu; p.arg28cys; p.pro1leu; p.val2met; p.pro1leu; p.arg1his; Aweak ABO*AW.2 c.2g>a; c.101delc p.arg1his; Aweak ABO*AW.28 c.98+2t>c Intron 1 Altered splicing Aweak ABO*AW.29 c.11t>a p.ile10asn Ax/Aweak ABO*AW.0.01 c.t>a p.phe21ile Ax/Aweak ABO*AW.0.02 c.t>a; c.81g>a p.phe21ile Page of 1
5 Names for ABO (ISBT 001) blood group alleles v Phenotype Allele name Nucleotide Exon Predicted amino acid Ax/Aweak ABO*AW.1.01 c.29a>g; c.t>a; c.81g>a; c.1c>t; p.phe21ile; p.val2met Ax/Aweak ABO*AW c.t>a; c.81g>a; c.1c>t; p.phe21ile; p.val2met Ax/Aweak ABO*AW.2 c.99g>a p.trp2ter Ax/Aweak ABO*AW. c.c>t; c.g>t p.pro1leu; p.trp181cys Ax/Aweak ABO*AW. c.c>t; ; c.1009a>g p.pro1leu; p.val2met; p.arggly Ax/Aweak ABO*AW. c.c>t; c.80c>t p.pro1leu; p.ala28val Ax/Aweak ABO*AW. c.0g>a p.glu20lys Ax/Aweak ABO*AW. c.90a>g p.lys1glu Ax/Aweak ABO*AW.8 c.2g>c p.met12ile Ax/Aweak ABO*AW.9 c.8t>c p.phe129leu Ax/Aweak ABO*AW.0 c.99g>t p.gly1cys Ax/Aweak ABO*AW.1 c.0a>g p.lys12glu Ax/Aweak ABO*AW.2 c.c>t; c.90a>g p.pro1leu; p.asp02gly Ax/Aweak ABO*AW. c.c>t; c.21c>t p.pro1leu; p.arg21trp Afinn/Aweak ABO*AW. c.+a>g Intron Altered splicing Abantu/Aweak ABO*AW. c.20+1delg; c.c>t; c.101delc Intron Altered splicing Am ABO*AM.01 c.c>t; c.1c>t p.pro1leu; p.ala2val Am ABO*AM.02 c.g>a p.val222met Ael ABO*AEL.01 c.80dupg p.phe29valfs*12 Ael ABO*AEL.02 c.c>t; c.t>a; c.81g>a p.pro1leu; p.phe21ile Ael ABO*AEL.0 c.80delg p.phe29serfs*20 Ael ABO*AEL.0 c.+g>a Intron Altered splicing Ael ABO*AEL.0 c.c>t; c.t>c p.pro1leu; p.ile2thr Ael ABO*AEL.0 c.2t>c; c.c>t p.met12thr; p.pro1leu Ael ABO*AEL.0 c.c>t; 81G>A; 1C>T; p.pro1leu; p.val2met Page of 1
6 Names for ABO (ISBT 001) blood group alleles v Phenotype Allele name Nucleotide Exon Predicted amino acid Ael ABO*AEL.08 c.c>t; c.80dupg p.pro1leu; p.phe29valfs*12 The A10-A10 alleles in dbrbc do not give rise to an altered amino acid sequence compared to other alleles, and so are not included here. A108 and A109 are listed as unpublished, and had no phenotype registered in dbrbc. A21 and A21 represent the same coding sequence as ABO*A2.01, but have been registered under other names due to intron polymorphisms. Also, their phenotypes are not given in dbrbc. Some alleles listed above are unpublished, but have been submitted to GenBank/dbRBC. It is also noteworthy that many of the alleles registered as associated with the rare A2 phenotype in Asia (e.g. A2.08, A2.1, A2.1, A2.18 and A2.20) cause amino acid substitutions that have been associated with weaker A subgroups in other studies. In the case of A2.18 and A2.19, the phenotype was given as A, not A2. Page of 1
7 Names for ABO (ISBT 001) blood group alleles v Molecular bases associated with the B and weak B phenotypes The seven B-associated polymorphisms are only shown for the first allele but are present in all the others except ABO*BEL.0, which uses A1.01. Differences compared to ABO*B.01, that encodes B glycosyltransferase, are given. Phenotype Allele name Nucleotide Exon Predicted amino acid B ABO*B.01 c.29a>g; c.2c>g; c.c>t; c.0g>a; c.9c>a; c.80g>c; c.90g>a p.arg1gly; p.gly2ser; p.leu2met; p.gly28ala B ABO*B.02 c.892g>t p.ala298ser B ABO*B.0 c.9c>t p.arg18cys B ABO*B.01 c.10c>t p.arg2trp B ABO*B.02 c.t>a p.phe21ile B ABO*B.0 c.1+g>a Intron Altered splicing B ABO*B.0 c.2g>t p.asp8tyr B ABO*B.0 c.2t>c p.met12thr B ABO*B.0 c.g>a p.asp18asn B ABO*B.0 c.10c>t p.ala1val B ABO*B.08 c.98a>c p.his1pro Bx/Bweak ABO*BW.01 c.81g>a p.asp291asn Bweak ABO*BW.02 c.8c>g p.asp291glu Bweak ABO*BW.0 c.21c>t p.arg21trp Bweak ABO*BW.0 c.8a>g p.asp18gly Bweak ABO*BW.0 c.9g>a p.arg180his Bweak ABO*BW.0 c.10a>g p.lysglu Bweak ABO*BW.0 c.10g>a p.arg2gln Bweak ABO*BW.08 c.8t>g p.met288arg Bweak ABO*BW.09 c.10a>t p.lysmet Bweak ABO*BW.10 c.a>g p.met18val Bweak ABO*BW.11 c.9t>c p.leu22pro Bweak ABO*BW.12 c.28c>t p.pro9leu Page of 1
8 Names for ABO (ISBT 001) blood group alleles v Phenotype Allele name Nucleotide Exon Predicted amino acid Bweak ABO*BW.1 c.2g>a p.val1met Bweak ABO*BW.1 c.a>g p.met189val Bweak ABO*BW.1 c.t>c p.ile192thr Bweak ABO*BW.1 c.8g>a p.asp22asn Bweak ABO*BW.18 c.802g>a p.gly28thr Bweak ABO*BW.19 c.t>a; c.81g>a p.phe21ile Bweak ABO*BW.20 c.81_81insg p.ser2valfs*? Bweak ABO*BW.21 c.88g>c p.gly20arg Bweak ABO*BW.22 c.0g>t p.arg18leu Bweak ABO*BW.2 c.g>c p.arg28pro Bweak ABO*BW.2 c.8g>t p.met18ile Bweak ABO*BW.2 c.10g>a; c.19c>g p.glyarg; p.leu20val Bweak ABO*BW.2 c.g>t 2 p.arg18leu Bweak ABO*BW.2 c.90a>g p.asp02gly Bweak ABO*BW.28 c.1t>c p.trp181arg Bweak ABO*BW.29 c.88c>g p.cys19trp Bweak ABO*BW.0 c.9g>t p.asp2tyr Bweak ABO*BW.1 c.900g>c p.trp00cys Bweak ABO*BW.2 c.808t>a p.phe20ile Bweak ABO*BW. c.0g>a p.val18met Bweak ABO*BW. c.889g>a p.glu29lys Bel ABO*BEL.01 c.1t>g p.met21arg Bel ABO*BEL.02 c.9g>t p.glu22asp Bel ABO*BEL.0 c.02c>t p.arg18trp Bel ABO*BEL.0 c.92g>a p.val18met Bel ABO*BEL.0 c.c>t; c.t>a; c.81g>a; c.1c>t; c.9c>a; c.80g>c; p.pro1leu; p.phe21ile; p.leu2met; p.gly28ala; p.val2met Other variants of B alleles exist, but the ones listed in dbrbc are either based on: 1) lack of one Page 8 of 1
9 Names for ABO (ISBT 001) blood group alleles v of the silent A vs. B SNPs (e.g. B102 has 90G; B10 has C); 2) silent mutations (B109 has 98C>T); ) intron SNPs (e.g. B10, B11, B11, B11); ) a sequence identical to a proven Bweak; ) unpublished (B11-B11). Page 9 of 1
10 Names for ABO (ISBT 001) blood group alleles v Molecular bases associated with cisab and B(A) phenotypes Differences compared to ABO*A1.01 are given. Phenotype Allele name Nucleotide Exon Predicted amino acid cisab ABO*cisAB.01 c.c>t; c.80g>c p.pro1leu; p.gly28ala cisab ABO*cisAB.02 c.2c>g; c.c>t; c.0g>a; c.80g>c cisab ABO*cisAB.0 c.29a>g; c.2c>g; c.c>t; c.00c>t; c.0g>a; c.9c>a; c.80g>c; c.90g>a p.arg1gly; p.gly2ser; p.gly28ala p.arg1gly; p.pro2ser; p.gly2ser; p.leu2met; p.gly28ala cisab ABO*cisAB.0 c.c>t; c.9c>a p.pro1leu; p.leu2met cisab ABO*cisAB.0 c.29a>g; c.2c>g; c.c>t; c.0g>a; c.9c>a; c.90g>a cisab ABO*cisAB.0 c.29a>g; c.c>t; c.0g>a; c.9c>a; c.80g>c; c.90g>a B(A) ABO*BA.01 c.29a>g; c.2c>g; c.9c>a; c.80g>c; c.90g>a B(A) ABO*BA.02 c.29a>g; c.2c>g; c.c>t; c.00c>g; c.0g>a; c.9c>a; c.80g>c; c.90g>a B(A) ABO*BA.0 c.29a>g; c.2c>g; c.c>t; c.9c>a; c.80g>c; c.90g>a B(A) ABO*BA.0 c.29a>g; c.2c>g; c.0a>g; c.c>t; c.0g>a; c.9c>a; c.80g>c; c.90g>a p.arg1gly; p.gly2ser; p.leu2met p.gly2ser; p.leu2met; p.gly28ala p.arg1gly; p.leu2met; p.gly28ala p.arg1gly; p.pro2ala; p.gly2ser; p.leu2met; p.gly28ala p.arg1gly; p.leu2met; p.gly28ala p.arg1gly; p.met21val; p.gly2ser; p.leu2met; p.gly28ala Page 10 of 1
11 Names for ABO (ISBT 001) blood group alleles v Phenotype Allele name Nucleotide Exon Predicted amino acid B(A) ABO*BA.0 c.29a>g; c.2c>g; c.1t>c; c.c>t; c.0g>a; c.9c>a; c.80g>c; c.90g>a B(A) ABO*BA.0 c.29a>g; c.2c>g; c.c>t; c.0g>a; c.9c>a; c.90g>a p.arg1gly; p.met21thr; p.gly2ser; p.leu2met; p.gly28ala p.arg1gly; p.gly2ser; p.leu2met Page 11 of 1
12 Names for ABO (ISBT 001) blood group alleles v Molecular bases associated with O (null) phenotype Differences compared to ABO*A1.01 are given. Phenotype Allele name dbrbc (alt.) name Nucleotide Exon Predicted amino acid O ABO*O O01 (O 1 ) c.21delg O ABO*O O02 (O 1v ) c.10g>t; c.21delg; c.29a>g; c.t>a; c.81g>a; c.1c>t; p.valphe; p.arghis; p.proser; O ABO*O.01.0 O0 c.21delg; c.9t>c O ABO*O.01.0 O0 c.21delg; c.29a>g O ABO*O.01.0 O0, O0 c.21delg; c.t>a; c.81g>a; c.1c>t; O ABO*O.01.0 O0 c.21delg; c.29a>g; c.t>a; c.81g>a; c.21c>t; c.1c>t; O ABO*O O09 c.21delg; c.18c>t; c.c>t O ABO*O O10 c.21delg; C>T O ABO*O O11 c.21delg; c.29a>g; c.2g>a; c.t>a; c.81g>a; c.1c>t; O ABO*O O12 c.21delg; c.29a>g; c.9c>t; c.t>a; c.81g>a; c.1c>t; O ABO*O.01.1 O1 c.21delg; c.29a>g; c.t>a; c.81g>a; c.1c>t; O ABO*O O22 c.21delg c.c>t; c.101delc Page 12 of 1
13 Names for ABO (ISBT 001) blood group alleles v Phenotype Allele name dbrbc (alt.) name Nucleotide Exon Predicted amino acid O ABO*O.01.2 O2 c.21delg; c.29a>g; c.t>a; c.1c>t; ; 10C>T O ABO*O.01.2 O2 c.10g>t; c.21delg; c.29a>g; c.2c>g; c.c>t; c.0g>a; c.9c>a; c.80g>c; c.90g>a O ABO*O.01.2 O2 c.21delg; c.t>c O ABO*O.01.2 O2 c.21delg; c.8c>a O ABO*O.01.2 O2 c.21delg; c.18c>t; c.c>t; c.29c>t O ABO*O O28 c.21delg; c.92a>g p.valphe; p.arghis; O ABO*O O29 c.21delg; c.18c>t O ABO*O.01.1 O1 c.21delg; c.29a>g; c.29g>a; c.t>a; c.81g>a; c.1c>t; O ABO*O.01.2 O2 c.21delg; c.29a>g; c.8c>t; c.t>a; c.81g>a; c.1c>t; O ABO*O.01. O c.21delg; c.29a>g; c.98c>t; c.t>a; c.81g>a; c.1c>t; O ABO*O.01. O c.21delg; c.29a>g; c.1g>a; c.t>a; c.81g>a; c.1c>t; Page 1 of 1
14 Names for ABO (ISBT 001) blood group alleles v Phenotype Allele name dbrbc (alt.) name Nucleotide Exon Predicted amino acid O ABO*O.01. O c.10g>t; c.21delg; c.29a>g; c.81g>a; c.1c>t; O ABO*O.01. O c.10g>t; c.21delg; c.29a>g; c.t>a; c.81g>a; O ABO*O.01.9 O9 c.21delg; c.29a>g; c.81g>a; c.1c>t; O ABO*O.01.0 O0 c.10g>t; c.21delg; c.29a>g; c.t>a; c.81g>a; O ABO*O.01.1 O1 c.21delg; c.29a>g; c.2c>g; c.c>t; c.0g>a; c.9c>a; c.80g>c; c.90g>a O ABO*O.01. O c.21delg; c.29a>g; c.t>a; c.1c>t; O ABO*O.01. O c.21delg; c.t>a; c.1c>t O ABO*O.01. O c.21delg; c.t>a; c.1c>t; O ABO*O.01. O; O0 c.21delg; c.9dela O ABO*O.01. O c.21delg; c.802g>a p.valphe; p.arghis; p.proser; p.valphe; p.arghis; p.proser; p.proser; p.valphe; p.arghis; Page 1 of 1
15 Names for ABO (ISBT 001) blood group alleles v Phenotype Allele name dbrbc (alt.) name Nucleotide Exon Predicted amino acid O ABO*O.01.8 O8 c.21delg; c.29a>g; c.t>a; c.81g>a; c.8c>t; 1C>T; O ABO*O.01.1 O1 c.21delg; c.g>c O ABO*O.01. O c.10g>a; c.10g>t; c.21delg; c.29a>g; c.t>a; c.81g>a; c.1c>t; O ABO*O.01.8 O8 c.10g>t; c.21delg; c.29a>g; c.t>a; c.81g>a; c.1c>t; O ABO*O.01.1 O1 c.21delg; O ABO*O.01. O c.10g>t; c.21delg; c.29a>g; c.2g>a; c.t>a; c.81g>a; c.1c>t; O ABO*O.01. O c.21delg; c.9t>c; c.10_108delagg O ABO*O c.190g>a; c.21delg O ABO*O.01.8 c.10g>t; c.188g>a; c.21delg; c.29a>g O ABO*O O0 (O 2 ) c.g>t; c.29a>g; c.2c>g; c.802g>a 2 p.glyarg; p.valphe; p.arghis; p.proser; p.valphe; p.arghis; p.valphe; p.arghis; p.proser; p.valile; p.valphe; p.arghis; p.arg18leu; p.proser; p.arg1gly; p.gly28arg Page 1 of 1
16 Names for ABO (ISBT 001) blood group alleles v Phenotype Allele name dbrbc (alt.) name Nucleotide Exon Predicted amino acid O ABO*O O8 (O 2-2) c.g>t; c.29a>g; c.2c>g; c.9c>t; c.89g>a; c.802g>a 2 p.arg18leu; p.proser; p.arg1gly; p.arg21cys; p.gly20asp; p.gly28arg O ABO*O.02.0 O9 (O 2-2) c.g>t; c.29a>g; c.2c>g; c.89g>a; c.802g>a 2 p.arg18leu; p.proser; p.arg1gly; p.gly20asp; p.gly28arg O ABO*O.02.0 O0 (O 2 -) c.g>t; c.29a>g; c.88c>t; c.2c>g; c.802g>a 2 p.arg18leu; p.proser; p.thr1met; p.arg1gly; p.gly28arg O ABO*O.0 O08 (O ) c.c>t; c.80dupg; c.101delc p.pro1leu; p.phe29valfs*8 O ABO*O.0.01 O1 (O ) c.8_88insg 2 p.val0glyfs*2 O ABO*O.0.02 O21 (O ) c.8_88insg; c.21delg; c.c>t 2 p.val0glyfs*2 O ABO*O.0 O2 (O ) c.22c>t p.gln108ter O ABO*O.0 O (O ) c.2g>a p.trp181ter O ABO*O.0 O1 (O01) O ABO*O.08 O1 (O02) c.c>t; c.89c>t p.pro1leu; p.ala298val c.92c>a p.tyr09ter O ABO*O O19 (R102) O ABO*O O20 (R10) c.t>a; c.81g>a; c.1c>t; c.29a>g; c.t>a; c.81g>a; c.1c>t; p.phe21ile; p.val2met p.phe21ile; p.val2met O ABO*O.10 O2 c._insg 2 p.phe2valfs* Page 1 of 1
17 Names for ABO (ISBT 001) blood group alleles v Phenotype Allele name dbrbc (alt.) name Nucleotide Exon Predicted amino acid O ABO*O.11 O c.29a>g; c.0_0delcag; c.2c>g; c.c>t; c.0g>a; c.9c>a; c.80g>c; c.90g>a O ABO*O.12 O c.29a>g; c.2c>g; c.g>a; c.c>t; c.0g>a; c.9c>a; c.80g>c; c.90g>a p.gln19del; p.arg1gly; p.gly2ser; p.leu2met; p.gly28ala p.arg1gly; Arg188His; p.gly2ser; p.leu2met; p.gly28ala O ABO*O.1 O8 c.2t>g p.val11gly O ABO*O.1 O9 c.t>a p.val212glu O ABO*O.1 O81 c.9t>c p.tyr2his O ABO*O.1 c.10g>t; c.188g>a; Deletion of exons - - p.? All alleles in which c.21delg occurs are numbered ABO*O.01.XX. Those O alleles that arise from a molecular basis other than c.21delg have been assigned independent ABO*O.XX numbers. Page 1 of 1
Names for H (ISBT 018) Blood Group Alleles
 Names for H (ISBT 018) Blood Group Alleles General description: The H blood group system consists of one antigen, H, that is carried on glycolipids and glycoproteins on the RBC membrane, where it is synthesised
Names for H (ISBT 018) Blood Group Alleles General description: The H blood group system consists of one antigen, H, that is carried on glycolipids and glycoproteins on the RBC membrane, where it is synthesised
Supplementary materials and methods
 Supplementary materials and methods Targeted resequencing RNA probes complementary in sequence to the coding exons (+ 2bp) of target genes were designed using e-array (https://earray.chem.agilent.com/earray/)
Supplementary materials and methods Targeted resequencing RNA probes complementary in sequence to the coding exons (+ 2bp) of target genes were designed using e-array (https://earray.chem.agilent.com/earray/)
Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes
 Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes Supplementary Information Jason Flannick*, Nicola L Beer*, Alexander G Bick,
Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes Supplementary Information Jason Flannick*, Nicola L Beer*, Alexander G Bick,
Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes
 Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes KM Kortüm 1,8 *, EK Mai 3,4 *, NH Hanafiah 3 *, CX Shi 1, YX Zhu 1, L Bruins 1, S Barrio 1,
Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes KM Kortüm 1,8 *, EK Mai 3,4 *, NH Hanafiah 3 *, CX Shi 1, YX Zhu 1, L Bruins 1, S Barrio 1,
Intron p Hybrid with p.leu245val p. Gly336Cys. p.leu62phe p. Ala137Val p. Asn152Thr hybrid with p.leu245val p.
 RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
Colclough et al., Human Mutation 1
 Region Colclough et al., Human Mutation 1 Supp. Table S1. List of HNF1A mutations Nucleotide Change NM_000545.5 Promoter and 5 UTR Mutations Protein Effect NM_000545.5 Three letter amino acid description
Region Colclough et al., Human Mutation 1 Supp. Table S1. List of HNF1A mutations Nucleotide Change NM_000545.5 Promoter and 5 UTR Mutations Protein Effect NM_000545.5 Three letter amino acid description
LMM / emerge III Network Reference Sequences October 2016 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene
 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
GALAFOLD (migalastat) capsules, for oral use Initial U.S. Approval: 2018
 HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
GALAFOLD (migalastat) capsules, for oral use Initial U.S. Approval: 2018
 HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10833 Supplementary Table 1: Clinical information for 48 whole exome sequencing samples Sample ID Age Gender tumor location Death OS (months) Recurrence PFS
SUPPLEMENTARY INFORMATION doi:10.1038/nature10833 Supplementary Table 1: Clinical information for 48 whole exome sequencing samples Sample ID Age Gender tumor location Death OS (months) Recurrence PFS
analysis in hereditary breast/ovarian cancer families of Portuguese ancestry
 Clin Genet 2015: 88: 41 48 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12441 The role of targeted
Clin Genet 2015: 88: 41 48 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12441 The role of targeted
ALANINE:GLYOXYLATE AMINOTRANSFERASE (AGXT) GENE (PRIMARY HYPEROXALURIA TYPE 1) MUTATION DATA BASE
 ALANINE:GLYOXYLATE AMINOTRANSFERASE (AGXT) GENE (PRIMARY HYPEROXALURIA TYPE 1) REFERENCE SEQUENCES: c.dna: NM_000030 g.dna: NG_008005.1 MUTATION DATA BASE Nomenclature: HGVS guidelines (http://www.hgvs.org/rec.html)
ALANINE:GLYOXYLATE AMINOTRANSFERASE (AGXT) GENE (PRIMARY HYPEROXALURIA TYPE 1) REFERENCE SEQUENCES: c.dna: NM_000030 g.dna: NG_008005.1 MUTATION DATA BASE Nomenclature: HGVS guidelines (http://www.hgvs.org/rec.html)
Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia
 Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia Erpelinck-Verschueren, Veronika Rockova, Mathijs Sanders,
Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia Erpelinck-Verschueren, Veronika Rockova, Mathijs Sanders,
Supplementary material PCNT point mutations and familial intracranial aneurysms
 Supplementary material PCNT point mutations and familial intracranial aneurysms Oswaldo Lorenzo-Betancor, MD, PhD 1, Patrick R. Blackburn, PhD 2,3, Emily Edwards, CCRP 4, Rocío Vázquez-do-Campo, MD 4,
Supplementary material PCNT point mutations and familial intracranial aneurysms Oswaldo Lorenzo-Betancor, MD, PhD 1, Patrick R. Blackburn, PhD 2,3, Emily Edwards, CCRP 4, Rocío Vázquez-do-Campo, MD 4,
Sample Test Report. Mayo Clinic GeneGuide. Report. Consumer Name DOB: 00/00/0000
 Mayo Clinic GeneGuide Report Consumer Name DOB: // Table Of Contents Mayo Clinic GeneGuide Genetic Test Results Demographics and Ordering Information 3 How to Use This Report 4 Carrier Screening - No variants
Mayo Clinic GeneGuide Report Consumer Name DOB: // Table Of Contents Mayo Clinic GeneGuide Genetic Test Results Demographics and Ordering Information 3 How to Use This Report 4 Carrier Screening - No variants
CHAPTER IV RESULTS Microcephaly General description
 47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
InheriGen Plus Disease Information
 DISEASE INFORMATION & MUTATIONS TESTED 17α-hydroxylase/17,20-lyase Deficiency CYP17A1 Brazilian 87% < 1 in 112 < 1 in 850 p.arg362cys, p.trp406arg, c.1243+5g>a, p.pro48fs, p.asp487_phe489del, p.tyr32x
DISEASE INFORMATION & MUTATIONS TESTED 17α-hydroxylase/17,20-lyase Deficiency CYP17A1 Brazilian 87% < 1 in 112 < 1 in 850 p.arg362cys, p.trp406arg, c.1243+5g>a, p.pro48fs, p.asp487_phe489del, p.tyr32x
Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH
 Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH Index patients (n=89) FH children Cascade screening (n=82) cdna Mutation Protein 3 0 c.10580g>a p.arg3527gln
Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH Index patients (n=89) FH children Cascade screening (n=82) cdna Mutation Protein 3 0 c.10580g>a p.arg3527gln
Hearing Loss Imaging Thyroid function tests TFT (age at test)
 Clinical Information on patients. Unclassified variants are denoted in brackets. EVA is enlarged vestibular aqueducts - Indicates no information. means asian. Patient TFT (age at test) 25215 c.1001+1g>a
Clinical Information on patients. Unclassified variants are denoted in brackets. EVA is enlarged vestibular aqueducts - Indicates no information. means asian. Patient TFT (age at test) 25215 c.1001+1g>a
Genetic Testing and Analysis. (858) MRN: Specimen: Saliva Received: 07/26/2016 GENETIC ANALYSIS REPORT
 GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
De Leeneer et al., Human Mutation 1
 De Leeneer et al., Human Mutation 1 Supp. Table S1A. Overview of the genes included in the workflow Ensembl Gene ID Ensembl Transcript ID Associated Gene Name ENSG00000198691 ENST00000370225 ABCA4 NM_000350.2
De Leeneer et al., Human Mutation 1 Supp. Table S1A. Overview of the genes included in the workflow Ensembl Gene ID Ensembl Transcript ID Associated Gene Name ENSG00000198691 ENST00000370225 ABCA4 NM_000350.2
Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome
 c Indian Academy of Sciences RESEARCH NOTE Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome FENGJIE YANG, YAXIAN CHEN, YU ZHANG, LIRU QIU, YU CHEN and JIANHUA ZHOU Department
c Indian Academy of Sciences RESEARCH NOTE Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome FENGJIE YANG, YAXIAN CHEN, YU ZHANG, LIRU QIU, YU CHEN and JIANHUA ZHOU Department
Supplementary Materials for
 www.sciencemag.org/content/354/6319/aaf7000/suppl/dc1 Supplementary Materials for Genetic identification of familial hypercholesterolemia within a single U.S. health care system Noura S. Abul-Husn, Kandamurugu
www.sciencemag.org/content/354/6319/aaf7000/suppl/dc1 Supplementary Materials for Genetic identification of familial hypercholesterolemia within a single U.S. health care system Noura S. Abul-Husn, Kandamurugu
The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel
 ACGS, June 2018 The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel Ian M. Frayling Member of Council & Variant Interpretation Committee (VIC) International Society for Gastrointestinal
ACGS, June 2018 The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel Ian M. Frayling Member of Council & Variant Interpretation Committee (VIC) International Society for Gastrointestinal
Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss
 Genetics Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss Akhila Alapati, 1 Kerry Goetz, 2 John Suk, 1 Mili Navani,
Genetics Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss Akhila Alapati, 1 Kerry Goetz, 2 John Suk, 1 Mili Navani,
Genotype to phenotype correlations in cartilage oligomeric matrix protein associated chondrodysplasias
 (2014) 22, 1278 1282 & 2014 Macmillan Publishers Limited All rights reserved 1018-4813/14 www.nature.com/ejhg ARTICLE Genotype to phenotype correlations in cartilage oligomeric matrix protein associated
(2014) 22, 1278 1282 & 2014 Macmillan Publishers Limited All rights reserved 1018-4813/14 www.nature.com/ejhg ARTICLE Genotype to phenotype correlations in cartilage oligomeric matrix protein associated
Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM
 Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures
 Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures Item Type text; Electronic Thesis Authors Chen, Debbie Publisher The University of Arizona. Rights Copyright is held by the
Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures Item Type text; Electronic Thesis Authors Chen, Debbie Publisher The University of Arizona. Rights Copyright is held by the
Re-classification of variants: Implications from the cardiac perspective
 Re-classification of variants: Implications from the cardiac perspective Karen McGuire Oxford Medical Genetics Laboratories, Churchill Hospital, Old Road, Oxford, OX3 7LE. Re-classification New evidence
Re-classification of variants: Implications from the cardiac perspective Karen McGuire Oxford Medical Genetics Laboratories, Churchill Hospital, Old Road, Oxford, OX3 7LE. Re-classification New evidence
chr6: _ del HFE NM_ ,744 bp deletion Deletion of entire HFE gene 2
 Table S1. Details of pathogenic and possibly pathogenic variants identified in the HH genes HFE, HFE2, HAMP, TFR2 and SLC40A1 from published reports. Variants have been classified as either 1, possibly
Table S1. Details of pathogenic and possibly pathogenic variants identified in the HH genes HFE, HFE2, HAMP, TFR2 and SLC40A1 from published reports. Variants have been classified as either 1, possibly
Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic Leukodystrophy
 International Journal of Genomics Volume 2018, Article ID 2361068, 9 pages https://doi.org/10.1155/2018/2361068 Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic
International Journal of Genomics Volume 2018, Article ID 2361068, 9 pages https://doi.org/10.1155/2018/2361068 Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic
Integration Solutions
 Integration Solutions (1) a) With no active glycosyltransferase of either type, an ii individual would not be able to add any sugars to the O form of the lipopolysaccharide. Thus, the only lipopolysaccharide
Integration Solutions (1) a) With no active glycosyltransferase of either type, an ii individual would not be able to add any sugars to the O form of the lipopolysaccharide. Thus, the only lipopolysaccharide
Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at-risk for hereditary breast cancer and characterization of its genetic ancestry
 /, 2016, Vol. 7, (No. 49), pp: 80465-80481 Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at- for hereditary breast cancer and characterization of its genetic ancestry Gabriela C.
/, 2016, Vol. 7, (No. 49), pp: 80465-80481 Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at- for hereditary breast cancer and characterization of its genetic ancestry Gabriela C.
*Correspondence and offprint requests to: Takashi Sekine; ORIGINAL ARTICLE ABSTRACT INTRODUCTION
 Nephrol Dial Transplant (2014) 29: 376 384 doi: 10.1093/ndt/gft394 Advance Access publication 29 September 2013 Japanese Dent disease has a wider clinical spectrum than Dent disease in Europe/USA: genetic
Nephrol Dial Transplant (2014) 29: 376 384 doi: 10.1093/ndt/gft394 Advance Access publication 29 September 2013 Japanese Dent disease has a wider clinical spectrum than Dent disease in Europe/USA: genetic
Concurrent Practical Session ACMG Classification
 Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Supplementary Information. Exome sequencing of serous endometrial tumors identifies recurrent somatic
 Supplementary Information Exome sequencing of serous endometrial tumors identifies recurrent somatic mutations in chromatin-remodeling and ubiquitin ligase complex genes Matthieu Le Gallo 1, Andrea J.
Supplementary Information Exome sequencing of serous endometrial tumors identifies recurrent somatic mutations in chromatin-remodeling and ubiquitin ligase complex genes Matthieu Le Gallo 1, Andrea J.
Table e-1: Investigation of 33 patients with early onset epilepsy for KCNT1 mutations.
 Table e1: Investigation of 33 patients with early onset epilepsy for KCNT1 mutations. Patient Phenotype Screening Method Diagnostic Karyotype Sanger sequencing NGS Diagnostic Panel WES chromosomal microarray
Table e1: Investigation of 33 patients with early onset epilepsy for KCNT1 mutations. Patient Phenotype Screening Method Diagnostic Karyotype Sanger sequencing NGS Diagnostic Panel WES chromosomal microarray
ATM germline heterozygosity does not play a role in CLL initiation but influences rapid disease progression through loss of the remaining ATM allele
 Published Ahead of Print on September 20, 2011, as doi:10.3324/haematol.2011.048827. Copyright 2011 Ferrata Storti Foundation. Early Release Paper ATM germline heterozygosity does not play a role in CLL
Published Ahead of Print on September 20, 2011, as doi:10.3324/haematol.2011.048827. Copyright 2011 Ferrata Storti Foundation. Early Release Paper ATM germline heterozygosity does not play a role in CLL
Supp. Materials and Methods
 Böhm et al., Human Mutation 1 Supp. Materials and Methods Patients Patient selection was based on the clinical presentation and/or histological findings suggesting centronuclear myopathy: muscle weakness
Böhm et al., Human Mutation 1 Supp. Materials and Methods Patients Patient selection was based on the clinical presentation and/or histological findings suggesting centronuclear myopathy: muscle weakness
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Parkinson disease 8, automsomal dominant OMIM number for disease 607060 Disease
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Parkinson disease 8, automsomal dominant OMIM number for disease 607060 Disease
Table S2. Putative mutations identified in the CHH cohort
 Table S. Putative utations identified in the CHH cohort Sapl e Phenotyp e Gene nt change aa change Zyg SIF T PPH MaxEn t In vitro ExA C ExAC NFE Previous report ACMG classificatio n ACMG evidence codes
Table S. Putative utations identified in the CHH cohort Sapl e Phenotyp e Gene nt change aa change Zyg SIF T PPH MaxEn t In vitro ExA C ExAC NFE Previous report ACMG classificatio n ACMG evidence codes
UvA-DARE (Digital Academic Repository) Familial hypercholesterolemia: the Dutch approach Huijgen, R. Link to publication
 UvA-DARE (Digital Academic Repository) Familial hypercholesterolemia: the Dutch approach Huijgen, R. Link to publication Citation for published version (APA): Huijgen, R. (2012). Familial hypercholesterolemia:
UvA-DARE (Digital Academic Repository) Familial hypercholesterolemia: the Dutch approach Huijgen, R. Link to publication Citation for published version (APA): Huijgen, R. (2012). Familial hypercholesterolemia:
Genotype phenotype correlation in a cohort of Portuguese patients comprising the entire spectrum of VWD types: impact of NGS
 Coagulation and Fibrinolysis 17 Genotype phenotype correlation in a cohort of Portuguese patients comprising the entire spectrum of VWD types: impact of NGS Teresa Fidalgo 1 ; Ramon Salvado 1 ; Irene Corrales
Coagulation and Fibrinolysis 17 Genotype phenotype correlation in a cohort of Portuguese patients comprising the entire spectrum of VWD types: impact of NGS Teresa Fidalgo 1 ; Ramon Salvado 1 ; Irene Corrales
Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics.
 Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics. Table S2. Gene symbols and data regarding potential oligogenic
Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics. Table S2. Gene symbols and data regarding potential oligogenic
J Clin Oncol 34: by American Society of Clinical Oncology INTRODUCTION
 VOLUME 34 NUMBER 34 DECEMBER 1, 2016 JOURNAL OF CLINICAL ONCOLOGY O R I G I N A L R E P O R T Conflicting Interpretation of Genetic Variants and Cancer Risk by Commercial Laboratories as Assessed by the
VOLUME 34 NUMBER 34 DECEMBER 1, 2016 JOURNAL OF CLINICAL ONCOLOGY O R I G I N A L R E P O R T Conflicting Interpretation of Genetic Variants and Cancer Risk by Commercial Laboratories as Assessed by the
JULY 21, Genetics 101: SCN1A. Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology
 JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
 Int. J. Cancer: 116, 692 702 (2005) ' 2005 Wiley-Liss, Inc. Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
Int. J. Cancer: 116, 692 702 (2005) ' 2005 Wiley-Liss, Inc. Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
doi: /brain/awp336 Brain 2010: 133;
 doi:10.1093/brain/awp336 Brain 2010: 133; 655 670 655 BRAIN A JOURNAL OF NEUROLOGY Glucose transporter-1 deficiency syndrome: the expanding clinical and genetic spectrum of a treatable disorder Wilhelmina
doi:10.1093/brain/awp336 Brain 2010: 133; 655 670 655 BRAIN A JOURNAL OF NEUROLOGY Glucose transporter-1 deficiency syndrome: the expanding clinical and genetic spectrum of a treatable disorder Wilhelmina
Supplementary Online Content
 Supplementary Online Content Mandelker D, Zhang L, Kemel Y, et al. Mutation detection in patients with advanced cancer by universal sequencing of cancer-related genes in tumor and normal DNA vs guideline-based
Supplementary Online Content Mandelker D, Zhang L, Kemel Y, et al. Mutation detection in patients with advanced cancer by universal sequencing of cancer-related genes in tumor and normal DNA vs guideline-based
Alternative!allele!percentages!(AAP)!in!all!samples! tested!by!smmips.!
 Supplementary,Appendix,! Content, Title, Page,numbers, Supplementary!Figure!1! Cohorts!and!samples! 2! Supplementary!Figure!2! Levels!of!mosaicism!in!PIK3CA!in!the!study!cohort.! 3! Supplementary!Figure!3!
Supplementary,Appendix,! Content, Title, Page,numbers, Supplementary!Figure!1! Cohorts!and!samples! 2! Supplementary!Figure!2! Levels!of!mosaicism!in!PIK3CA!in!the!study!cohort.! 3! Supplementary!Figure!3!
Clinical spectrum and molecular basis of recessive congenital methemoglobinemia in India
 Clin Genet 2015: 87: 62 67 Printed in Singapore. All rights reserved Short Report 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12326 Clinical spectrum
Clin Genet 2015: 87: 62 67 Printed in Singapore. All rights reserved Short Report 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12326 Clinical spectrum
Beta Thalassemia Case Study Introduction to Bioinformatics
 Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Supplementary Table 1. PIK3CA mutation in colorectal cancer
 Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Beta Thalassemia Sami Khuri Department of Computer Science San José State University Spring 2015
 Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Reverse phenotyping: MCD Detection without clinical diagnosis. Diagnostic: NGS panels versus WES
 Clinical Genetics Leading the way in genetic issues Reverse phenotyping: MCD Detection without clinical diagnosis Diagnostic: NGS panels versus WES Martina Wilke Laboratory Specialist Clinical Genetics
Clinical Genetics Leading the way in genetic issues Reverse phenotyping: MCD Detection without clinical diagnosis Diagnostic: NGS panels versus WES Martina Wilke Laboratory Specialist Clinical Genetics
SUPPLEMENTARY DATA REFERENCES
 www.impactjournals.com/oncoscience Oncoscience, Advance Pulications 2016 SUPPLEMENTARY DATA REFERENCES 1. Busch RM, Chapin JS, Mester J, Ferguson L, Haut JS, Frazier TW, Eng C. Cognitive characteristics
www.impactjournals.com/oncoscience Oncoscience, Advance Pulications 2016 SUPPLEMENTARY DATA REFERENCES 1. Busch RM, Chapin JS, Mester J, Ferguson L, Haut JS, Frazier TW, Eng C. Cognitive characteristics
p.arg119gly p.arg119his p.ala179thr c.540+1g>a c.617_633+6del Prediction basis structure
 a Missense ATG p.arg119gly p.arg119his p.ala179thr p.ala189val p.gly206trp p.gly206arg p.arg251his p.ala257thr TGA 5 UTR 1 2 3 4 5 6 7 3 UTR Splice Site, Frameshift b Mutation p.gly206trp p.gly4fsx50 c.138+1g>a
a Missense ATG p.arg119gly p.arg119his p.ala179thr p.ala189val p.gly206trp p.gly206arg p.arg251his p.ala257thr TGA 5 UTR 1 2 3 4 5 6 7 3 UTR Splice Site, Frameshift b Mutation p.gly206trp p.gly4fsx50 c.138+1g>a
Mismatch repair gene mutation spectrum in the Swedish Lynch syndrome population
 ONCOLOGY REPORTS 36: 2823-2835, 2016 Mismatch repair gene mutation spectrum in the Swedish Lynch syndrome population Kristina Lagerstedt-Robinson 1, Anna Rohlin 2,3, Christos Aravidis 4, Beatrice Melin
ONCOLOGY REPORTS 36: 2823-2835, 2016 Mismatch repair gene mutation spectrum in the Swedish Lynch syndrome population Kristina Lagerstedt-Robinson 1, Anna Rohlin 2,3, Christos Aravidis 4, Beatrice Melin
Frequent incidence of BARD1-truncating mutations in germline DNA from triple-negative breast cancer patients
 Clin Genet 2016: 89: 336 340 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12620 Frequent incidence
Clin Genet 2016: 89: 336 340 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12620 Frequent incidence
Supplementary Table 2. Identified causative mutations and/or mutation candidates.
 Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
Novel mutations of PKD genes in Chinese patients suffering from autosomal dominant polycystic kidney disease and seeking assisted reproduction
 He et al. BMC Medical Genetics (2018) 19:186 https://doi.org/10.1186/s12881-018-0693-7 RESEARCH ARTICLE Novel mutations of PKD genes in Chinese patients suffering from autosomal dominant polycystic kidney
He et al. BMC Medical Genetics (2018) 19:186 https://doi.org/10.1186/s12881-018-0693-7 RESEARCH ARTICLE Novel mutations of PKD genes in Chinese patients suffering from autosomal dominant polycystic kidney
Introduction HUMAN GENETICS ORIGINAL PAPER
 J Appl Genetics (2016) 57:175 181 DOI 10.1007/s13353-015-0310-9 HUMAN GENETICS ORIGINAL PAPER Novel sporadic and recurrent mutations in KRT5 and KRT14 genes in Polish epidermolysis bullosa simplex patients:
J Appl Genetics (2016) 57:175 181 DOI 10.1007/s13353-015-0310-9 HUMAN GENETICS ORIGINAL PAPER Novel sporadic and recurrent mutations in KRT5 and KRT14 genes in Polish epidermolysis bullosa simplex patients:
NGS in neurodegenerative disorders. Our first experience
 NGS in neurodegenerative disorders Our first experience Milena Jankovic, PhD Laboratory for genetic and molecular diagnostic of neurological disorders Neurology Clinic, Clinical Center of Serbia University
NGS in neurodegenerative disorders Our first experience Milena Jankovic, PhD Laboratory for genetic and molecular diagnostic of neurological disorders Neurology Clinic, Clinical Center of Serbia University
Supplementary Figure 1
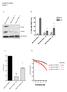 Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
NIH Public Access Author Manuscript Nat Genet. Author manuscript; available in PMC 2012 June 18.
 NIH Public Access Author Manuscript Published in final edited form as: Nat Genet. ; 43(11): 1119 1126. doi:10.1038/ng.950. Exon capture analysis of G protein-coupled receptors identifies activating mutations
NIH Public Access Author Manuscript Published in final edited form as: Nat Genet. ; 43(11): 1119 1126. doi:10.1038/ng.950. Exon capture analysis of G protein-coupled receptors identifies activating mutations
Mutation allele frequency threshold does not affect prognostic analysis using nextgeneration sequencing in oral squamous cell carcinoma
 Ma et al. BMC Cancer (2018) 18:758 https://doi.org/10.1186/s12885-018-4481-8 RESEARCH ARTICLE Open Access Mutation allele frequency threshold does not affect prognostic analysis using nextgeneration sequencing
Ma et al. BMC Cancer (2018) 18:758 https://doi.org/10.1186/s12885-018-4481-8 RESEARCH ARTICLE Open Access Mutation allele frequency threshold does not affect prognostic analysis using nextgeneration sequencing
Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension
 Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension Felix Beuschlein, Sheerazed Boulkroun, Andrea Osswald, Thomas Wieland, Hang N. Nielsen, Urs D. Lichtenauer,
Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension Felix Beuschlein, Sheerazed Boulkroun, Andrea Osswald, Thomas Wieland, Hang N. Nielsen, Urs D. Lichtenauer,
Phenylalanine hydroxylase deficiency in Mexico: genotype phenotype correlations, BH 4 responsiveness and evidence of a founder effect
 Clin Genet 2015: 88: 62 67 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12444 Phenylalanine hydroxylase
Clin Genet 2015: 88: 62 67 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12444 Phenylalanine hydroxylase
Molecular genetics of maple syrup urine disease in the Turkish population
 The Turkish Journal of Pediatrics 2009; 51: 97-102 Original Molecular genetics of maple syrup urine disease in the Turkish population Kerstin Gorzelany 1, Ali Dursun 2, Turgay Coşkun 2, Serap H. Kalkanoğlu-Sivri
The Turkish Journal of Pediatrics 2009; 51: 97-102 Original Molecular genetics of maple syrup urine disease in the Turkish population Kerstin Gorzelany 1, Ali Dursun 2, Turgay Coşkun 2, Serap H. Kalkanoğlu-Sivri
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Genetic and biochemical characteristics of patients with hyperlipidemia who require LDL apheresis
 Genetic and biochemical characteristics of patients with hyperlipidemia who require LDL apheresis Mato Nagel, Ioan Duma, Constantina Vlad, Mandy Benke, Sylvia Nagorka Zentrum für Nephrologie & Stoffwechsel,
Genetic and biochemical characteristics of patients with hyperlipidemia who require LDL apheresis Mato Nagel, Ioan Duma, Constantina Vlad, Mandy Benke, Sylvia Nagorka Zentrum für Nephrologie & Stoffwechsel,
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Morfopoulou S, Brown JR, Davies EG, et al. Human coronavirus
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Morfopoulou S, Brown JR, Davies EG, et al. Human coronavirus
SUPPLEMENTARY INFORMATION. Rare independent mutations in renal salt handling genes contribute to blood pressure variation
 SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
s Unique autosomal recessive variant of palmoplantar keratoderma associated with hearing loss not caused by known mutations*
 154 Case Report s Unique autosomal recessive variant of palmoplantar keratoderma associated with hearing loss not caused by known mutations* Moustafa Abdelaal Hegazi 1,2 Sommen Manou 3 Hazem Sakr 4 Guy
154 Case Report s Unique autosomal recessive variant of palmoplantar keratoderma associated with hearing loss not caused by known mutations* Moustafa Abdelaal Hegazi 1,2 Sommen Manou 3 Hazem Sakr 4 Guy
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP
 Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supplementary Material
 Supplementary Material Mutations in NMNAT1 cause Leber congenital amaurosis with early-onset severe macular and optic atrophy Isabelle Perrault 1, Sylvain Hanein 1, Xavier Zanlonghi 2, Valérie Serre 1,
Supplementary Material Mutations in NMNAT1 cause Leber congenital amaurosis with early-onset severe macular and optic atrophy Isabelle Perrault 1, Sylvain Hanein 1, Xavier Zanlonghi 2, Valérie Serre 1,
Supplementary Online Content
 Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Urea cycle disorders in Spain: an observational, cross-sectional and multicentric study of 104 cases
 Martín-Hernández et al. Orphanet Journal of Rare Diseases 2014, 9:187 RESEARCH Open Access Urea cycle disorders in Spain: an observational, cross-sectional and multicentric study of 104 cases Elena Martín-Hernández
Martín-Hernández et al. Orphanet Journal of Rare Diseases 2014, 9:187 RESEARCH Open Access Urea cycle disorders in Spain: an observational, cross-sectional and multicentric study of 104 cases Elena Martín-Hernández
Primary Osteoporosis in Young Adults: Genetic Basis and Identification of Novel Variants in Causal Genes
 ORIGINAL ARTICLE Primary Osteoporosis in Young Adults: Genetic Basis and Identification of Novel Variants in Causal Genes Corinne Collet, 1,2,3 Agnes Ostertag, 2,3 Manon Ricquebourg, 2,3 Marine Delecourt,
ORIGINAL ARTICLE Primary Osteoporosis in Young Adults: Genetic Basis and Identification of Novel Variants in Causal Genes Corinne Collet, 1,2,3 Agnes Ostertag, 2,3 Manon Ricquebourg, 2,3 Marine Delecourt,
MUTATION TEST CE IVD. ctnras-braf FEATURES
 ctnras-braf CE IVD TECHNICAL SHEET IDYLLA ctnras-braf MUTATION TEST The Idylla ctnras-braf Mutation Test, performed on the Biocartis Idylla System, is an in vitro diagnostic test for the qualitative detection
ctnras-braf CE IVD TECHNICAL SHEET IDYLLA ctnras-braf MUTATION TEST The Idylla ctnras-braf Mutation Test, performed on the Biocartis Idylla System, is an in vitro diagnostic test for the qualitative detection
What do you think of when you here the word genome?
 What do you think of when you here the word genome? What do you think of when you here the word genome? Personal Genomics Outline Review of pre-lab work Genomics and Medicine Case Overview & Assignment
What do you think of when you here the word genome? What do you think of when you here the word genome? Personal Genomics Outline Review of pre-lab work Genomics and Medicine Case Overview & Assignment
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Mutational and phenotypical spectrum of phenylalanine hydroxylase deficiency in Denmark
 Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Molecular Diagnostic Laboratory 18 Sequencing St, Gene Town, ZY Tel: Fax:
 Molecular Diagnostic Laboratory 18 Sequencing St, Gene Town, ZY 01234 Tel: 555-920-3333 Fax: 555-920-3334 www.moldxlaboratory.com Patient Name: Jane Doe Specimen type: Blood, peripheral DOB: 04/05/1990
Molecular Diagnostic Laboratory 18 Sequencing St, Gene Town, ZY 01234 Tel: 555-920-3333 Fax: 555-920-3334 www.moldxlaboratory.com Patient Name: Jane Doe Specimen type: Blood, peripheral DOB: 04/05/1990
EXOME SEQUENCING AS A SECOND-TIER DIAGNOSTIC APPROACH FOR CLINICALLY SUSPECTED DYSFERLINOPATHY PATIENTS
 EXOME SEQUENCING AS A SECOND-TIER DIAGNOSTIC APPROACH FOR CLINICALLY SUSPECTED DYSFERLINOPATHY PATIENTS Marc Bartoli, Jean-Pierre Desvignes, Nicolas Lévy, Martin Krahn To cite this version: Marc Bartoli,
EXOME SEQUENCING AS A SECOND-TIER DIAGNOSTIC APPROACH FOR CLINICALLY SUSPECTED DYSFERLINOPATHY PATIENTS Marc Bartoli, Jean-Pierre Desvignes, Nicolas Lévy, Martin Krahn To cite this version: Marc Bartoli,
TKCC/Garvan Cancer Biology Seminars. Principles of multidisciplinary musculoskeletal tumour management. Prof David Thomas 4 th March 2016
 TKCC/Garvan Cancer Biology Seminars Principles of multidisciplinary musculoskeletal tumour management Prof David Thomas 4 th March 2016 Aims of presentation An state-of-the-art overview of: Australian
TKCC/Garvan Cancer Biology Seminars Principles of multidisciplinary musculoskeletal tumour management Prof David Thomas 4 th March 2016 Aims of presentation An state-of-the-art overview of: Australian
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Identification of Splicing Defects Caused by Mutations in the Dysferlin Gene
 RESEARCH ARTICLE Identification of Splicing Defects Caused by Mutations in the Dysferlin Gene OFFICIAL JOURNAL Virginie Kergourlay, 1,2 Ghadi Raï, 1,2 Gaëlle Blandin, 1,2 David Salgado, 1,2 Christophe
RESEARCH ARTICLE Identification of Splicing Defects Caused by Mutations in the Dysferlin Gene OFFICIAL JOURNAL Virginie Kergourlay, 1,2 Ghadi Raï, 1,2 Gaëlle Blandin, 1,2 David Salgado, 1,2 Christophe
WDR62 is associated with the spindle pole and mutated in human microcephaly
 WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
Genetic characterization of antithrombin, protein C, and protein S deficiencies in Polish patients
 ORIGINAL ARTICLE Genetic characterization of antithrombin, protein C, and protein S deficiencies in Polish patients Ewa Wypasek 1,2, Javier Corral 3, Martine Alhenc Gelas 4, Wojciech Sydor 5, Teresa Iwaniec
ORIGINAL ARTICLE Genetic characterization of antithrombin, protein C, and protein S deficiencies in Polish patients Ewa Wypasek 1,2, Javier Corral 3, Martine Alhenc Gelas 4, Wojciech Sydor 5, Teresa Iwaniec
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
The ABO and Rh system. Dr U. La Rocca 03 th Novembre 2017
 The ABO and Rh system Dr U. La Rocca 03 th Novembre 2017 Main learning endpoints! ü Chemical structure ü Inheritance ü AB0 and Rh antibodies and their importance in transfusion ü Principles of AB0 and
The ABO and Rh system Dr U. La Rocca 03 th Novembre 2017 Main learning endpoints! ü Chemical structure ü Inheritance ü AB0 and Rh antibodies and their importance in transfusion ü Principles of AB0 and
doi: /brain/awq087 Brain 2010: 133; Tyrosine hydroxylase deficiency: a treatable disorder of brain catecholamine biosynthesis
 doi:10.1093/brain/awq087 Brain 2010: 133; 1810 1822 1810 BRAIN A JOURNAL OF NEUROLOGY Tyrosine hydroxylase deficiency: a treatable disorder of brain catecholamine biosynthesis Michèl A. Willemsen, 1 Marcel
doi:10.1093/brain/awq087 Brain 2010: 133; 1810 1822 1810 BRAIN A JOURNAL OF NEUROLOGY Tyrosine hydroxylase deficiency: a treatable disorder of brain catecholamine biosynthesis Michèl A. Willemsen, 1 Marcel
Long-term exposure to Myozyme results in a decrease of anti-drug antibodies in lateonset Pompe disease patients
 Long-term exposure to Myozyme results in a decrease of anti-drug antibodies in lateonset Pompe disease patients Elisa Masat 1, Pascal Laforêt 1,2, Marie De Antonio 3, Guillaume Corre 4, Barbara Perniconi
Long-term exposure to Myozyme results in a decrease of anti-drug antibodies in lateonset Pompe disease patients Elisa Masat 1, Pascal Laforêt 1,2, Marie De Antonio 3, Guillaume Corre 4, Barbara Perniconi
Dr. Ayman Mohsen Mashi, MBBS Consultant Hematology & Blood Transfusion Department Head, Laboratory & Blood Bank King Fahad Central Hospital, Gazan,
 Dr. Ayman Mohsen Mashi, MBBS Consultant Hematology & Blood Transfusion Department Head, Laboratory & Blood Bank King Fahad Central Hospital, Gazan, KSA amashi@moh.gov.sa 24/02/2018 β-thalassemia syndromes
Dr. Ayman Mohsen Mashi, MBBS Consultant Hematology & Blood Transfusion Department Head, Laboratory & Blood Bank King Fahad Central Hospital, Gazan, KSA amashi@moh.gov.sa 24/02/2018 β-thalassemia syndromes
SUPPLEMENTARY DATA. Isolated NIBPL missense mutations that cause Cornelia de Lange syndrome alter MAU2. interaction.
 SUPPLEMENTARY DATA Isolated NIBPL missense mutations that cause Cornelia de Lange syndrome alter MAU2 interaction. Diana Braunholz 1, Melanie Hullings 2, María Concepcion Gil-Rodríguez 1,3, Christopher
SUPPLEMENTARY DATA Isolated NIBPL missense mutations that cause Cornelia de Lange syndrome alter MAU2 interaction. Diana Braunholz 1, Melanie Hullings 2, María Concepcion Gil-Rodríguez 1,3, Christopher
CIC Edizioni Internazionali
 Next generation sequencing in the identification of a rare genetic disease from preconceptional couple screening to preimplantation genetic diagnosis Claudio Dello Russo 1 Gianluca Di Giacomo 1 Alvaro
Next generation sequencing in the identification of a rare genetic disease from preconceptional couple screening to preimplantation genetic diagnosis Claudio Dello Russo 1 Gianluca Di Giacomo 1 Alvaro
Bioinformatics Laboratory Exercise
 Bioinformatics Laboratory Exercise Biology is in the midst of the genomics revolution, the application of robotic technology to generate huge amounts of molecular biology data. Genomics has led to an explosion
Bioinformatics Laboratory Exercise Biology is in the midst of the genomics revolution, the application of robotic technology to generate huge amounts of molecular biology data. Genomics has led to an explosion
