Names for H (ISBT 018) Blood Group Alleles
|
|
|
- Rodney Hunter
- 5 years ago
- Views:
Transcription
1 Names for H (ISBT 018) Blood Group Alleles General description: The H blood group system consists of one antigen, H, that is carried on glycolipids and glycoproteins on the RBC membrane, where it is synthesised by the fucosyltransferase product of the FUT1 gene; as well as on glycoproteins on epithelial cells and in body fluids, where it is synthesised by the fucosyltransferase product of the FUT2 gene. In group O individuals, H antigen is the terminal antigen however, in group A and B individuals, the H antigen serves as the precursor structure for A and B blood-group-specific glycosyltransferases. Thus, group O people will test strongly H+ whereas groups A, B and AB will express very little H antigen. Mutations that negatively affect the α2fuct1 enzyme activity (encoded by FUT1) will result in reduced or absent H production (and a concomitant decrease in A and/or B antigens in individuals where those enzymes are encoded). Total absence of H, A and B antigens is called the Oh or Bombay phenotype. Weak expression is referred to as the parabombay phenotype. The enzymes α2fuct1and α2fuct2 are single pass type II membrane glycoproteins in the Golgi. The α2fuct1 protein consists of 365 amino acids and is encoded by FUT1 or H, if analysis is to predict a blood group antigen. The FUT2 gene produces two transcripts; one of 343 amino acids and another more abundant form of 332 amino acids. The longer transcript encodes a protein with approximately one fourth the enzymatic activity and the shorter form is considered to be the active enzyme. Thus, the numbers given below are counted from the second initiating codon. The α2fuct2 protein is encoded by FUT2 or Se, if analysis is to predict a blood group antigen. Gene name: FUT1 Number of exons: 4 Initiation codon: Beginning of exon 4 Stop codon: Within exon 4 Entrez Gene ID: 2523 LRG sequence: NG_ (genomic) NM_ (transcript) Reference allele: FUT1*01 (shaded) Acceptable: H if inferred by hemagglutination/inhibition Page 1 of 5
2 Phenotype Allele name Nucleotide change Exon Predicted amino acid change H+ FUT1*01 H+ FUT1*02 c.35c>t 4 p.ala12val Weak phenotypes H+weak FUT1*01W.01 c.293c>t 4 p.thr98met H+weak FUT1*01W.02 c.328g>a 4 p.ala110thr H+weak FUT1*01W.03 c.349c>t 4 p.his117tyr H+weak FUT1*01W.04 c.442g>t 4 p.asp148tyr H+weak FUT1*01W c.460t>c 4 p.tyr154his H+weak FUT1*01W c.460t>c; c.1042g>a 4 p.tyr154his; p.glu348lys H+weak FUT1*01W.07 c.491t>a 4 p.leu164his H+weak FUT1*01W.08 c.522c>a 4 p.phe174leu H+weak FUT1*01W.09 c.658c>t 4 p.arg220cys H+weak FUT1*01W.10 c.659g>a 4 p.arg220his H+weak FUT1*01W.11 c.661c>t 4 p.arg221cys H+weak FUT1*01W.12 c.682a>g 4 p.met228val H+weak FUT1*01W.13 c.689a>c 4 p.gln230pro H+weak FUT1*01W.14 c.721t>c 4 p.tyr241his H+weak FUT1*01W.15 c.801g>c 4 p.trp267cys H+weak FUT1*01W.16 c.801g>t 4 p.trp267cys H+weak FUT1*01W.17 c.832g>a 4 p.asp278asn H+weak FUT1*01W.18 c.903_904insaac 4 p.asn301_his302insasn H+weak FUT1*01W.19 c.917c>t 4 p.thr306ile H+weak FUT1*01W.20 c.990delg 4 p.pro331glnfs*6 H+weak FUT1*01W.21 c.235g>c 4 p.gly79arg H+weak FUT1*01W.22 c.991c>a 4 p.pro331thr H+weak FUT1*01W.23 c.424c>t 4 p.arg142trp H+weak FUT1*01W.24 c.649g>t 4 p.val217phe H+weak FUT1*01W.25 Identical to FUT*01W.21? c.235g>c 4 p.gly79arg H+weak FUT1*01W.26 c.545g>a 4 p.arg182his Page 2 of 5
3 H+weak FUT1*01W.27 c.958g>a 4 p.gly320arg H+weak FUT1*01W.28 c.896a>c 4 p.gln299pro H+weak FUT1*01W.29 c.655g>c 4 p.val219leu H+weak FUT1*02W.01 c.269g>t 4 p.gly90val H+weak FUT1*02W.02 c.371t>g 4 p.phe124cys H+weak FUT1*02W.03 c.682a>g 4 p.met228val H+weak FUT1*02W.04 c.980a>c 4 p.asn327thr H+weak FUT1*02W.05 c.748c>t 4 p.arg250trp Null alleles H FUT1*01N.01 c.422g>a 4 p.trp141ter H FUT1*01N.02 c.461a>g 4 p.tyr154cys H FUT1*01N.03 c.462c>a 4 p.tyr154ter H FUT1*01N.04 c.513g>c 4 p.trp171cys H FUT1*01N.05 c.538c>t 4 p.gln180ter H FUT1*01N.06 c.551_552delag 4 p.glu184valfs*85 H FUT1*01N.07 c.586c>t 4 p.gln196ter H FUT1*01N.08 c.695g>a 4 p.trp232ter H FUT1*01N.09 c.725t>g 4 p.leu242arg H FUT1*01N.10 c.776t>a 4 p.val259glu H FUT1*01N.11 c.785g>a; c.786c>a 4 p.ser262lys H FUT1*01N.12 c.826c>t 4 p.gln276ter H FUT1*01N.13 c.881_882deltt 4 p.phe294cysfs*40 H FUT1*01N.14 c.944c>t 4 p.ala315val H FUT1*01N.15 c.948c>g 4 p.tyr316ter H FUT1*01N.16 c.980a>c 4 p.asn327thr H FUT1*01N.17 c.1047g>c 4 p.trp349cys H FUT1*01N.18 c.684g>a 4 p.met228ile H FUT1*01N.19 c.694t>c 4 p.trp232arg H FUT1*01N.20 c.768delc 4 p.val257phefs*23 H FUT1*02N.01 c.423g>a 4 p.trp141ter Note that H expression will be masked if a functional A or B allele is also inherited. Also, that H antigen may be weakly detectable on RBCs where FUT*01N homozygosity occurs, due to the adsorption of soluble H antigen synthesized by FUT2. Page 3 of 5
4 Gene name: FUT2 Number of exons: 2 Initiation codon: Beginning of exon 2 Stop codon: Within exon 2 Entrez Gene ID: 2524 LRG sequence: NG_ (genomic) NM_ (transcript) Reference allele: FUT2*01 (shaded) Acceptable: Se if inferred by hemagglutination/inhibition Phenotype (saliva) Allele name Nucleotide change Exon Predicted amino acid change H+ FUT2*01 H+ FUT2*02 c.4g>a 2 p.ala2thr H+ FUT2*03.01 c.40a>g 2 p.ile14val H+ FUT2*03.02 c.40a>g; c.113c>t H+ FUT2*03.03 c.40a>g; c.481g>a 2 p.ile14val; p.ala38val 2 p.ile14val; p.asp161asn H+ FUT2*04 c.379c>t 2 p.arg127cys H+ FUT2*05 c.400g>a 2 p.val134ile H+ FUT2*06 c.481g>a 2 p.asp161asn H+ FUT2*07 c.665g>a 2 p.arg222his H+ FUT2*08 c.685g>a 2 p.val229met H+ FUT2*09 c.716g>a 2 p.arg239gln H+ FUT2*10 c.747_748insgtg 2 p.249_250insval Weak phenotypes H+w FUT2*01W.01 c.278c>t 2 p.ala93val H+w FUT2*01W c.385a>t 2 p.ile129phe H+w FUT2*01W c.385a>t; c.617t>g H+w FUT2*01W c.385a>t; c.841g>a 2 p.ile129phe, p.val206gly 2 p.ile129phe, p.gly281arg H+w FUT2*01W.03 c.853g>a 2 p.ala285thr Null phenotypes Nucleotide polymorphisms H FUT2*01N.01 c.244g>a; c.385a>t 2 p.ala82thr; p.ile129phe H FUT2*01N.02 c.428g>a; 2 p.trp143ter; Page 4 of 5
5 c.739a>g (after termination?) p.gly247ser (after termination?) H FUT2*01N.03 c.569g>a 2 p.arg190his H FUT2*01N.04 c.571c>t 2 p.arg191ter H FUT2*01N.05 c.628c>t 2 p.arg210ter H FUT2*01N.06 c.658c>t 2 p.arg220ter H FUT2*01N.07 c.664c>t 2 p.arg222cys H FUT2*01N.08 c.685_686delgt 2 p.val229glyfs*4 H FUT2*01N.09 c.688_690delgtc 2 p.val230del H FUT2*01N.10 c.400g>a; c.760g>a 2 p.val134ile; p.asp254asn H FUT2*01N.11 c.778delc 2 p.pro260leufs*16 H FUT2*01N.12 c.849g>a 2 p.trp283ter H FUT2*01N.13 c.868 G>A 2 p.gly290arg H FUT2*01N.14 c.950c>t 2 p.pro317leu H FUT2*01N.15 c.302c>t 2 p.pro101leu H FUT2*01N.16 c.960a>g (wrong position?) 2 p.gly247ser (wrong position?) H FUT2*01N.17 c.412g>a 2 p.gly138ser H FUT2*01N.18 c.818c>a 2 p.thr273asn Null phenotypes Gene deletions H FUT2*0N.01 Gene deletion p.0 H FUT2*0N.02 Coding region deleted p.0 H FUT2*0N.03 Fusion gene 1 between FUT2 and Sec1 H FUT2*0N.04 Fusion gene 2 between FUT2 and Sec1 - - Saliva phenotype is shown here to represent secreted H antigen in all body fluids Page 5 of 5
Names for ABO (ISBT 001) Blood Group Alleles
 Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes
 Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes KM Kortüm 1,8 *, EK Mai 3,4 *, NH Hanafiah 3 *, CX Shi 1, YX Zhu 1, L Bruins 1, S Barrio 1,
Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes KM Kortüm 1,8 *, EK Mai 3,4 *, NH Hanafiah 3 *, CX Shi 1, YX Zhu 1, L Bruins 1, S Barrio 1,
Supplementary materials and methods
 Supplementary materials and methods Targeted resequencing RNA probes complementary in sequence to the coding exons (+ 2bp) of target genes were designed using e-array (https://earray.chem.agilent.com/earray/)
Supplementary materials and methods Targeted resequencing RNA probes complementary in sequence to the coding exons (+ 2bp) of target genes were designed using e-array (https://earray.chem.agilent.com/earray/)
Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes
 Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes Supplementary Information Jason Flannick*, Nicola L Beer*, Alexander G Bick,
Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes Supplementary Information Jason Flannick*, Nicola L Beer*, Alexander G Bick,
Intron p Hybrid with p.leu245val p. Gly336Cys. p.leu62phe p. Ala137Val p. Asn152Thr hybrid with p.leu245val p.
 RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
Colclough et al., Human Mutation 1
 Region Colclough et al., Human Mutation 1 Supp. Table S1. List of HNF1A mutations Nucleotide Change NM_000545.5 Promoter and 5 UTR Mutations Protein Effect NM_000545.5 Three letter amino acid description
Region Colclough et al., Human Mutation 1 Supp. Table S1. List of HNF1A mutations Nucleotide Change NM_000545.5 Promoter and 5 UTR Mutations Protein Effect NM_000545.5 Three letter amino acid description
Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia
 Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia Erpelinck-Verschueren, Veronika Rockova, Mathijs Sanders,
Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia Erpelinck-Verschueren, Veronika Rockova, Mathijs Sanders,
GALAFOLD (migalastat) capsules, for oral use Initial U.S. Approval: 2018
 HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
Hearing Loss Imaging Thyroid function tests TFT (age at test)
 Clinical Information on patients. Unclassified variants are denoted in brackets. EVA is enlarged vestibular aqueducts - Indicates no information. means asian. Patient TFT (age at test) 25215 c.1001+1g>a
Clinical Information on patients. Unclassified variants are denoted in brackets. EVA is enlarged vestibular aqueducts - Indicates no information. means asian. Patient TFT (age at test) 25215 c.1001+1g>a
analysis in hereditary breast/ovarian cancer families of Portuguese ancestry
 Clin Genet 2015: 88: 41 48 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12441 The role of targeted
Clin Genet 2015: 88: 41 48 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12441 The role of targeted
CHAPTER IV RESULTS Microcephaly General description
 47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
LMM / emerge III Network Reference Sequences October 2016 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene
 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
GALAFOLD (migalastat) capsules, for oral use Initial U.S. Approval: 2018
 HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10833 Supplementary Table 1: Clinical information for 48 whole exome sequencing samples Sample ID Age Gender tumor location Death OS (months) Recurrence PFS
SUPPLEMENTARY INFORMATION doi:10.1038/nature10833 Supplementary Table 1: Clinical information for 48 whole exome sequencing samples Sample ID Age Gender tumor location Death OS (months) Recurrence PFS
Sample Test Report. Mayo Clinic GeneGuide. Report. Consumer Name DOB: 00/00/0000
 Mayo Clinic GeneGuide Report Consumer Name DOB: // Table Of Contents Mayo Clinic GeneGuide Genetic Test Results Demographics and Ordering Information 3 How to Use This Report 4 Carrier Screening - No variants
Mayo Clinic GeneGuide Report Consumer Name DOB: // Table Of Contents Mayo Clinic GeneGuide Genetic Test Results Demographics and Ordering Information 3 How to Use This Report 4 Carrier Screening - No variants
De Leeneer et al., Human Mutation 1
 De Leeneer et al., Human Mutation 1 Supp. Table S1A. Overview of the genes included in the workflow Ensembl Gene ID Ensembl Transcript ID Associated Gene Name ENSG00000198691 ENST00000370225 ABCA4 NM_000350.2
De Leeneer et al., Human Mutation 1 Supp. Table S1A. Overview of the genes included in the workflow Ensembl Gene ID Ensembl Transcript ID Associated Gene Name ENSG00000198691 ENST00000370225 ABCA4 NM_000350.2
Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH
 Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH Index patients (n=89) FH children Cascade screening (n=82) cdna Mutation Protein 3 0 c.10580g>a p.arg3527gln
Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH Index patients (n=89) FH children Cascade screening (n=82) cdna Mutation Protein 3 0 c.10580g>a p.arg3527gln
Supplementary material PCNT point mutations and familial intracranial aneurysms
 Supplementary material PCNT point mutations and familial intracranial aneurysms Oswaldo Lorenzo-Betancor, MD, PhD 1, Patrick R. Blackburn, PhD 2,3, Emily Edwards, CCRP 4, Rocío Vázquez-do-Campo, MD 4,
Supplementary material PCNT point mutations and familial intracranial aneurysms Oswaldo Lorenzo-Betancor, MD, PhD 1, Patrick R. Blackburn, PhD 2,3, Emily Edwards, CCRP 4, Rocío Vázquez-do-Campo, MD 4,
Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM
 Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss
 Genetics Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss Akhila Alapati, 1 Kerry Goetz, 2 John Suk, 1 Mili Navani,
Genetics Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss Akhila Alapati, 1 Kerry Goetz, 2 John Suk, 1 Mili Navani,
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
ALANINE:GLYOXYLATE AMINOTRANSFERASE (AGXT) GENE (PRIMARY HYPEROXALURIA TYPE 1) MUTATION DATA BASE
 ALANINE:GLYOXYLATE AMINOTRANSFERASE (AGXT) GENE (PRIMARY HYPEROXALURIA TYPE 1) REFERENCE SEQUENCES: c.dna: NM_000030 g.dna: NG_008005.1 MUTATION DATA BASE Nomenclature: HGVS guidelines (http://www.hgvs.org/rec.html)
ALANINE:GLYOXYLATE AMINOTRANSFERASE (AGXT) GENE (PRIMARY HYPEROXALURIA TYPE 1) REFERENCE SEQUENCES: c.dna: NM_000030 g.dna: NG_008005.1 MUTATION DATA BASE Nomenclature: HGVS guidelines (http://www.hgvs.org/rec.html)
Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome
 c Indian Academy of Sciences RESEARCH NOTE Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome FENGJIE YANG, YAXIAN CHEN, YU ZHANG, LIRU QIU, YU CHEN and JIANHUA ZHOU Department
c Indian Academy of Sciences RESEARCH NOTE Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome FENGJIE YANG, YAXIAN CHEN, YU ZHANG, LIRU QIU, YU CHEN and JIANHUA ZHOU Department
Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures
 Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures Item Type text; Electronic Thesis Authors Chen, Debbie Publisher The University of Arizona. Rights Copyright is held by the
Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures Item Type text; Electronic Thesis Authors Chen, Debbie Publisher The University of Arizona. Rights Copyright is held by the
InheriGen Plus Disease Information
 DISEASE INFORMATION & MUTATIONS TESTED 17α-hydroxylase/17,20-lyase Deficiency CYP17A1 Brazilian 87% < 1 in 112 < 1 in 850 p.arg362cys, p.trp406arg, c.1243+5g>a, p.pro48fs, p.asp487_phe489del, p.tyr32x
DISEASE INFORMATION & MUTATIONS TESTED 17α-hydroxylase/17,20-lyase Deficiency CYP17A1 Brazilian 87% < 1 in 112 < 1 in 850 p.arg362cys, p.trp406arg, c.1243+5g>a, p.pro48fs, p.asp487_phe489del, p.tyr32x
Supplementary Materials for
 www.sciencemag.org/content/354/6319/aaf7000/suppl/dc1 Supplementary Materials for Genetic identification of familial hypercholesterolemia within a single U.S. health care system Noura S. Abul-Husn, Kandamurugu
www.sciencemag.org/content/354/6319/aaf7000/suppl/dc1 Supplementary Materials for Genetic identification of familial hypercholesterolemia within a single U.S. health care system Noura S. Abul-Husn, Kandamurugu
chr6: _ del HFE NM_ ,744 bp deletion Deletion of entire HFE gene 2
 Table S1. Details of pathogenic and possibly pathogenic variants identified in the HH genes HFE, HFE2, HAMP, TFR2 and SLC40A1 from published reports. Variants have been classified as either 1, possibly
Table S1. Details of pathogenic and possibly pathogenic variants identified in the HH genes HFE, HFE2, HAMP, TFR2 and SLC40A1 from published reports. Variants have been classified as either 1, possibly
Genetic Testing and Analysis. (858) MRN: Specimen: Saliva Received: 07/26/2016 GENETIC ANALYSIS REPORT
 GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
Genotype to phenotype correlations in cartilage oligomeric matrix protein associated chondrodysplasias
 (2014) 22, 1278 1282 & 2014 Macmillan Publishers Limited All rights reserved 1018-4813/14 www.nature.com/ejhg ARTICLE Genotype to phenotype correlations in cartilage oligomeric matrix protein associated
(2014) 22, 1278 1282 & 2014 Macmillan Publishers Limited All rights reserved 1018-4813/14 www.nature.com/ejhg ARTICLE Genotype to phenotype correlations in cartilage oligomeric matrix protein associated
Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at-risk for hereditary breast cancer and characterization of its genetic ancestry
 /, 2016, Vol. 7, (No. 49), pp: 80465-80481 Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at- for hereditary breast cancer and characterization of its genetic ancestry Gabriela C.
/, 2016, Vol. 7, (No. 49), pp: 80465-80481 Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at- for hereditary breast cancer and characterization of its genetic ancestry Gabriela C.
Table e-1: Investigation of 33 patients with early onset epilepsy for KCNT1 mutations.
 Table e1: Investigation of 33 patients with early onset epilepsy for KCNT1 mutations. Patient Phenotype Screening Method Diagnostic Karyotype Sanger sequencing NGS Diagnostic Panel WES chromosomal microarray
Table e1: Investigation of 33 patients with early onset epilepsy for KCNT1 mutations. Patient Phenotype Screening Method Diagnostic Karyotype Sanger sequencing NGS Diagnostic Panel WES chromosomal microarray
The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel
 ACGS, June 2018 The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel Ian M. Frayling Member of Council & Variant Interpretation Committee (VIC) International Society for Gastrointestinal
ACGS, June 2018 The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel Ian M. Frayling Member of Council & Variant Interpretation Committee (VIC) International Society for Gastrointestinal
Supplementary Table 1. PIK3CA mutation in colorectal cancer
 Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Mutation allele frequency threshold does not affect prognostic analysis using nextgeneration sequencing in oral squamous cell carcinoma
 Ma et al. BMC Cancer (2018) 18:758 https://doi.org/10.1186/s12885-018-4481-8 RESEARCH ARTICLE Open Access Mutation allele frequency threshold does not affect prognostic analysis using nextgeneration sequencing
Ma et al. BMC Cancer (2018) 18:758 https://doi.org/10.1186/s12885-018-4481-8 RESEARCH ARTICLE Open Access Mutation allele frequency threshold does not affect prognostic analysis using nextgeneration sequencing
Alternative!allele!percentages!(AAP)!in!all!samples! tested!by!smmips.!
 Supplementary,Appendix,! Content, Title, Page,numbers, Supplementary!Figure!1! Cohorts!and!samples! 2! Supplementary!Figure!2! Levels!of!mosaicism!in!PIK3CA!in!the!study!cohort.! 3! Supplementary!Figure!3!
Supplementary,Appendix,! Content, Title, Page,numbers, Supplementary!Figure!1! Cohorts!and!samples! 2! Supplementary!Figure!2! Levels!of!mosaicism!in!PIK3CA!in!the!study!cohort.! 3! Supplementary!Figure!3!
Re-classification of variants: Implications from the cardiac perspective
 Re-classification of variants: Implications from the cardiac perspective Karen McGuire Oxford Medical Genetics Laboratories, Churchill Hospital, Old Road, Oxford, OX3 7LE. Re-classification New evidence
Re-classification of variants: Implications from the cardiac perspective Karen McGuire Oxford Medical Genetics Laboratories, Churchill Hospital, Old Road, Oxford, OX3 7LE. Re-classification New evidence
Supplementary Online Content
 Supplementary Online Content Mandelker D, Zhang L, Kemel Y, et al. Mutation detection in patients with advanced cancer by universal sequencing of cancer-related genes in tumor and normal DNA vs guideline-based
Supplementary Online Content Mandelker D, Zhang L, Kemel Y, et al. Mutation detection in patients with advanced cancer by universal sequencing of cancer-related genes in tumor and normal DNA vs guideline-based
Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic Leukodystrophy
 International Journal of Genomics Volume 2018, Article ID 2361068, 9 pages https://doi.org/10.1155/2018/2361068 Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic
International Journal of Genomics Volume 2018, Article ID 2361068, 9 pages https://doi.org/10.1155/2018/2361068 Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic
ATM germline heterozygosity does not play a role in CLL initiation but influences rapid disease progression through loss of the remaining ATM allele
 Published Ahead of Print on September 20, 2011, as doi:10.3324/haematol.2011.048827. Copyright 2011 Ferrata Storti Foundation. Early Release Paper ATM germline heterozygosity does not play a role in CLL
Published Ahead of Print on September 20, 2011, as doi:10.3324/haematol.2011.048827. Copyright 2011 Ferrata Storti Foundation. Early Release Paper ATM germline heterozygosity does not play a role in CLL
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Parkinson disease 8, automsomal dominant OMIM number for disease 607060 Disease
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Parkinson disease 8, automsomal dominant OMIM number for disease 607060 Disease
Supp. Materials and Methods
 Böhm et al., Human Mutation 1 Supp. Materials and Methods Patients Patient selection was based on the clinical presentation and/or histological findings suggesting centronuclear myopathy: muscle weakness
Böhm et al., Human Mutation 1 Supp. Materials and Methods Patients Patient selection was based on the clinical presentation and/or histological findings suggesting centronuclear myopathy: muscle weakness
J Clin Oncol 34: by American Society of Clinical Oncology INTRODUCTION
 VOLUME 34 NUMBER 34 DECEMBER 1, 2016 JOURNAL OF CLINICAL ONCOLOGY O R I G I N A L R E P O R T Conflicting Interpretation of Genetic Variants and Cancer Risk by Commercial Laboratories as Assessed by the
VOLUME 34 NUMBER 34 DECEMBER 1, 2016 JOURNAL OF CLINICAL ONCOLOGY O R I G I N A L R E P O R T Conflicting Interpretation of Genetic Variants and Cancer Risk by Commercial Laboratories as Assessed by the
UvA-DARE (Digital Academic Repository) Familial hypercholesterolemia: the Dutch approach Huijgen, R. Link to publication
 UvA-DARE (Digital Academic Repository) Familial hypercholesterolemia: the Dutch approach Huijgen, R. Link to publication Citation for published version (APA): Huijgen, R. (2012). Familial hypercholesterolemia:
UvA-DARE (Digital Academic Repository) Familial hypercholesterolemia: the Dutch approach Huijgen, R. Link to publication Citation for published version (APA): Huijgen, R. (2012). Familial hypercholesterolemia:
Supplementary Information. Exome sequencing of serous endometrial tumors identifies recurrent somatic
 Supplementary Information Exome sequencing of serous endometrial tumors identifies recurrent somatic mutations in chromatin-remodeling and ubiquitin ligase complex genes Matthieu Le Gallo 1, Andrea J.
Supplementary Information Exome sequencing of serous endometrial tumors identifies recurrent somatic mutations in chromatin-remodeling and ubiquitin ligase complex genes Matthieu Le Gallo 1, Andrea J.
Genotype phenotype correlation in a cohort of Portuguese patients comprising the entire spectrum of VWD types: impact of NGS
 Coagulation and Fibrinolysis 17 Genotype phenotype correlation in a cohort of Portuguese patients comprising the entire spectrum of VWD types: impact of NGS Teresa Fidalgo 1 ; Ramon Salvado 1 ; Irene Corrales
Coagulation and Fibrinolysis 17 Genotype phenotype correlation in a cohort of Portuguese patients comprising the entire spectrum of VWD types: impact of NGS Teresa Fidalgo 1 ; Ramon Salvado 1 ; Irene Corrales
Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension
 Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension Felix Beuschlein, Sheerazed Boulkroun, Andrea Osswald, Thomas Wieland, Hang N. Nielsen, Urs D. Lichtenauer,
Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension Felix Beuschlein, Sheerazed Boulkroun, Andrea Osswald, Thomas Wieland, Hang N. Nielsen, Urs D. Lichtenauer,
Reverse phenotyping: MCD Detection without clinical diagnosis. Diagnostic: NGS panels versus WES
 Clinical Genetics Leading the way in genetic issues Reverse phenotyping: MCD Detection without clinical diagnosis Diagnostic: NGS panels versus WES Martina Wilke Laboratory Specialist Clinical Genetics
Clinical Genetics Leading the way in genetic issues Reverse phenotyping: MCD Detection without clinical diagnosis Diagnostic: NGS panels versus WES Martina Wilke Laboratory Specialist Clinical Genetics
Integration Solutions
 Integration Solutions (1) a) With no active glycosyltransferase of either type, an ii individual would not be able to add any sugars to the O form of the lipopolysaccharide. Thus, the only lipopolysaccharide
Integration Solutions (1) a) With no active glycosyltransferase of either type, an ii individual would not be able to add any sugars to the O form of the lipopolysaccharide. Thus, the only lipopolysaccharide
doi: /brain/awp336 Brain 2010: 133;
 doi:10.1093/brain/awp336 Brain 2010: 133; 655 670 655 BRAIN A JOURNAL OF NEUROLOGY Glucose transporter-1 deficiency syndrome: the expanding clinical and genetic spectrum of a treatable disorder Wilhelmina
doi:10.1093/brain/awp336 Brain 2010: 133; 655 670 655 BRAIN A JOURNAL OF NEUROLOGY Glucose transporter-1 deficiency syndrome: the expanding clinical and genetic spectrum of a treatable disorder Wilhelmina
Table S2. Putative mutations identified in the CHH cohort
 Table S. Putative utations identified in the CHH cohort Sapl e Phenotyp e Gene nt change aa change Zyg SIF T PPH MaxEn t In vitro ExA C ExAC NFE Previous report ACMG classificatio n ACMG evidence codes
Table S. Putative utations identified in the CHH cohort Sapl e Phenotyp e Gene nt change aa change Zyg SIF T PPH MaxEn t In vitro ExA C ExAC NFE Previous report ACMG classificatio n ACMG evidence codes
Concurrent Practical Session ACMG Classification
 Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Morfopoulou S, Brown JR, Davies EG, et al. Human coronavirus
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Morfopoulou S, Brown JR, Davies EG, et al. Human coronavirus
Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
 Int. J. Cancer: 116, 692 702 (2005) ' 2005 Wiley-Liss, Inc. Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
Int. J. Cancer: 116, 692 702 (2005) ' 2005 Wiley-Liss, Inc. Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
Beta Thalassemia Sami Khuri Department of Computer Science San José State University Spring 2015
 Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Novel mutations of PKD genes in Chinese patients suffering from autosomal dominant polycystic kidney disease and seeking assisted reproduction
 He et al. BMC Medical Genetics (2018) 19:186 https://doi.org/10.1186/s12881-018-0693-7 RESEARCH ARTICLE Novel mutations of PKD genes in Chinese patients suffering from autosomal dominant polycystic kidney
He et al. BMC Medical Genetics (2018) 19:186 https://doi.org/10.1186/s12881-018-0693-7 RESEARCH ARTICLE Novel mutations of PKD genes in Chinese patients suffering from autosomal dominant polycystic kidney
PIK3CA-associated developmental disorders exhibit distinct classes of mutations with variable expression and tissue distribution
 PIK3CA-associated developmental disorders exhibit distinct classes of mutations with variable expression and tissue distribution Ghayda Mirzaa, 1,2 Andrew E. Timms, 3 Valerio Conti, 4 Evan August Boyle,
PIK3CA-associated developmental disorders exhibit distinct classes of mutations with variable expression and tissue distribution Ghayda Mirzaa, 1,2 Andrew E. Timms, 3 Valerio Conti, 4 Evan August Boyle,
Correlation of phenotype with genotype and protein structure in RYR1-related disorders
 https://doi.org/10.1007/s00415-018-9033-2 ORIGINAL COMMUNICATION Correlation of phenotype with genotype and protein structure in RYR1-related disorders Joshua J. Todd 1 Vatsala Sagar 2 Tokunbor A. Lawal
https://doi.org/10.1007/s00415-018-9033-2 ORIGINAL COMMUNICATION Correlation of phenotype with genotype and protein structure in RYR1-related disorders Joshua J. Todd 1 Vatsala Sagar 2 Tokunbor A. Lawal
Supplementary Table 2. Identified causative mutations and/or mutation candidates.
 Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
MUTATION TEST CE IVD. ctnras-braf FEATURES
 ctnras-braf CE IVD TECHNICAL SHEET IDYLLA ctnras-braf MUTATION TEST The Idylla ctnras-braf Mutation Test, performed on the Biocartis Idylla System, is an in vitro diagnostic test for the qualitative detection
ctnras-braf CE IVD TECHNICAL SHEET IDYLLA ctnras-braf MUTATION TEST The Idylla ctnras-braf Mutation Test, performed on the Biocartis Idylla System, is an in vitro diagnostic test for the qualitative detection
Frequent incidence of BARD1-truncating mutations in germline DNA from triple-negative breast cancer patients
 Clin Genet 2016: 89: 336 340 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12620 Frequent incidence
Clin Genet 2016: 89: 336 340 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12620 Frequent incidence
Beta Thalassemia Case Study Introduction to Bioinformatics
 Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Downloaded from:
 Ryan, NS; Nicholas, JM; Weston, PS; Liang, Y; Lashley, T; Guerreiro, R; Adamson, G; Kenny, J; Beck, J; Chavez-Gutierrez, L; de Strooper, B; Revesz, T; Holton, J; Mead, S; Rossor, MN; Fox, NC (2016) Clinical
Ryan, NS; Nicholas, JM; Weston, PS; Liang, Y; Lashley, T; Guerreiro, R; Adamson, G; Kenny, J; Beck, J; Chavez-Gutierrez, L; de Strooper, B; Revesz, T; Holton, J; Mead, S; Rossor, MN; Fox, NC (2016) Clinical
The ABO and Rh system. Dr U. La Rocca 03 th Novembre 2017
 The ABO and Rh system Dr U. La Rocca 03 th Novembre 2017 Main learning endpoints! ü Chemical structure ü Inheritance ü AB0 and Rh antibodies and their importance in transfusion ü Principles of AB0 and
The ABO and Rh system Dr U. La Rocca 03 th Novembre 2017 Main learning endpoints! ü Chemical structure ü Inheritance ü AB0 and Rh antibodies and their importance in transfusion ü Principles of AB0 and
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
*Correspondence and offprint requests to: Takashi Sekine; ORIGINAL ARTICLE ABSTRACT INTRODUCTION
 Nephrol Dial Transplant (2014) 29: 376 384 doi: 10.1093/ndt/gft394 Advance Access publication 29 September 2013 Japanese Dent disease has a wider clinical spectrum than Dent disease in Europe/USA: genetic
Nephrol Dial Transplant (2014) 29: 376 384 doi: 10.1093/ndt/gft394 Advance Access publication 29 September 2013 Japanese Dent disease has a wider clinical spectrum than Dent disease in Europe/USA: genetic
NIH Public Access Author Manuscript Nat Genet. Author manuscript; available in PMC 2012 June 18.
 NIH Public Access Author Manuscript Published in final edited form as: Nat Genet. ; 43(11): 1119 1126. doi:10.1038/ng.950. Exon capture analysis of G protein-coupled receptors identifies activating mutations
NIH Public Access Author Manuscript Published in final edited form as: Nat Genet. ; 43(11): 1119 1126. doi:10.1038/ng.950. Exon capture analysis of G protein-coupled receptors identifies activating mutations
WDR62 is associated with the spindle pole and mutated in human microcephaly
 WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
SUPPLEMENTARY DATA REFERENCES
 www.impactjournals.com/oncoscience Oncoscience, Advance Pulications 2016 SUPPLEMENTARY DATA REFERENCES 1. Busch RM, Chapin JS, Mester J, Ferguson L, Haut JS, Frazier TW, Eng C. Cognitive characteristics
www.impactjournals.com/oncoscience Oncoscience, Advance Pulications 2016 SUPPLEMENTARY DATA REFERENCES 1. Busch RM, Chapin JS, Mester J, Ferguson L, Haut JS, Frazier TW, Eng C. Cognitive characteristics
Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics.
 Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics. Table S2. Gene symbols and data regarding potential oligogenic
Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics. Table S2. Gene symbols and data regarding potential oligogenic
TKCC/Garvan Cancer Biology Seminars. Principles of multidisciplinary musculoskeletal tumour management. Prof David Thomas 4 th March 2016
 TKCC/Garvan Cancer Biology Seminars Principles of multidisciplinary musculoskeletal tumour management Prof David Thomas 4 th March 2016 Aims of presentation An state-of-the-art overview of: Australian
TKCC/Garvan Cancer Biology Seminars Principles of multidisciplinary musculoskeletal tumour management Prof David Thomas 4 th March 2016 Aims of presentation An state-of-the-art overview of: Australian
s Unique autosomal recessive variant of palmoplantar keratoderma associated with hearing loss not caused by known mutations*
 154 Case Report s Unique autosomal recessive variant of palmoplantar keratoderma associated with hearing loss not caused by known mutations* Moustafa Abdelaal Hegazi 1,2 Sommen Manou 3 Hazem Sakr 4 Guy
154 Case Report s Unique autosomal recessive variant of palmoplantar keratoderma associated with hearing loss not caused by known mutations* Moustafa Abdelaal Hegazi 1,2 Sommen Manou 3 Hazem Sakr 4 Guy
CIC Edizioni Internazionali
 Next generation sequencing in the identification of a rare genetic disease from preconceptional couple screening to preimplantation genetic diagnosis Claudio Dello Russo 1 Gianluca Di Giacomo 1 Alvaro
Next generation sequencing in the identification of a rare genetic disease from preconceptional couple screening to preimplantation genetic diagnosis Claudio Dello Russo 1 Gianluca Di Giacomo 1 Alvaro
The proteasome inhibitor bortezomib reduced cholesterol accumulation in fibroblasts from Niemann Pick type C patients carrying missense mutations
 The proteasome inhibitor bortezomib reduced cholesterol accumulation in fibroblasts from Niemann Pick type C patients carrying missense mutations Judit Macıas-Vidal 1,2,3, Marisa Giros 1,2,3, Martina Guerrero
The proteasome inhibitor bortezomib reduced cholesterol accumulation in fibroblasts from Niemann Pick type C patients carrying missense mutations Judit Macıas-Vidal 1,2,3, Marisa Giros 1,2,3, Martina Guerrero
What do you think of when you here the word genome?
 What do you think of when you here the word genome? What do you think of when you here the word genome? Personal Genomics Outline Review of pre-lab work Genomics and Medicine Case Overview & Assignment
What do you think of when you here the word genome? What do you think of when you here the word genome? Personal Genomics Outline Review of pre-lab work Genomics and Medicine Case Overview & Assignment
Supplementary Figure 1
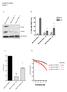 Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
Genetic and biochemical characteristics of patients with hyperlipidemia who require LDL apheresis
 Genetic and biochemical characteristics of patients with hyperlipidemia who require LDL apheresis Mato Nagel, Ioan Duma, Constantina Vlad, Mandy Benke, Sylvia Nagorka Zentrum für Nephrologie & Stoffwechsel,
Genetic and biochemical characteristics of patients with hyperlipidemia who require LDL apheresis Mato Nagel, Ioan Duma, Constantina Vlad, Mandy Benke, Sylvia Nagorka Zentrum für Nephrologie & Stoffwechsel,
Supplementary Online Content
 Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Mismatch repair gene mutation spectrum in the Swedish Lynch syndrome population
 ONCOLOGY REPORTS 36: 2823-2835, 2016 Mismatch repair gene mutation spectrum in the Swedish Lynch syndrome population Kristina Lagerstedt-Robinson 1, Anna Rohlin 2,3, Christos Aravidis 4, Beatrice Melin
ONCOLOGY REPORTS 36: 2823-2835, 2016 Mismatch repair gene mutation spectrum in the Swedish Lynch syndrome population Kristina Lagerstedt-Robinson 1, Anna Rohlin 2,3, Christos Aravidis 4, Beatrice Melin
Variant interpretation exercise. ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18
 Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Use of panel tests in place of single gene tests in the cancer genetics clinic
 Clin Genet 2015: 88: 278 282 Printed in Singapore. All rights reserved CLINICAL GENETICS doi: 10.1111/cge.12488 Short Report se of panel tests in place of single gene tests in the cancer genetics clinic
Clin Genet 2015: 88: 278 282 Printed in Singapore. All rights reserved CLINICAL GENETICS doi: 10.1111/cge.12488 Short Report se of panel tests in place of single gene tests in the cancer genetics clinic
Long-term exposure to Myozyme results in a decrease of anti-drug antibodies in lateonset Pompe disease patients
 Long-term exposure to Myozyme results in a decrease of anti-drug antibodies in lateonset Pompe disease patients Elisa Masat 1, Pascal Laforêt 1,2, Marie De Antonio 3, Guillaume Corre 4, Barbara Perniconi
Long-term exposure to Myozyme results in a decrease of anti-drug antibodies in lateonset Pompe disease patients Elisa Masat 1, Pascal Laforêt 1,2, Marie De Antonio 3, Guillaume Corre 4, Barbara Perniconi
Dr Rosline Hassan Haematology Department, School of Medical Sciences, Universiti Sains Malaysia, Kelantan
 Dr Rosline Hassan Haematology Department, School of Medical Sciences, Universiti Sains Malaysia, Kelantan THE FIRST ASEAN FEDERATION OF HAEMATOLOGY AND THE VIIITH MALAYSIAN NATIONAL HAEMATOLOGY SCIENTIFIC
Dr Rosline Hassan Haematology Department, School of Medical Sciences, Universiti Sains Malaysia, Kelantan THE FIRST ASEAN FEDERATION OF HAEMATOLOGY AND THE VIIITH MALAYSIAN NATIONAL HAEMATOLOGY SCIENTIFIC
MUTATIONS, MUTAGENESIS, AND CARCINOGENESIS. (Start your clickers)
 MUTATIONS, MUTAGENESIS, AND CARCINOGENESIS (Start your clickers) How do mutations arise? And how do they affect a cell and its organism? Mutations: heritable changes in genes Mutations occur in DNA But
MUTATIONS, MUTAGENESIS, AND CARCINOGENESIS (Start your clickers) How do mutations arise? And how do they affect a cell and its organism? Mutations: heritable changes in genes Mutations occur in DNA But
NGS in neurodegenerative disorders. Our first experience
 NGS in neurodegenerative disorders Our first experience Milena Jankovic, PhD Laboratory for genetic and molecular diagnostic of neurological disorders Neurology Clinic, Clinical Center of Serbia University
NGS in neurodegenerative disorders Our first experience Milena Jankovic, PhD Laboratory for genetic and molecular diagnostic of neurological disorders Neurology Clinic, Clinical Center of Serbia University
p.arg119gly p.arg119his p.ala179thr c.540+1g>a c.617_633+6del Prediction basis structure
 a Missense ATG p.arg119gly p.arg119his p.ala179thr p.ala189val p.gly206trp p.gly206arg p.arg251his p.ala257thr TGA 5 UTR 1 2 3 4 5 6 7 3 UTR Splice Site, Frameshift b Mutation p.gly206trp p.gly4fsx50 c.138+1g>a
a Missense ATG p.arg119gly p.arg119his p.ala179thr p.ala189val p.gly206trp p.gly206arg p.arg251his p.ala257thr TGA 5 UTR 1 2 3 4 5 6 7 3 UTR Splice Site, Frameshift b Mutation p.gly206trp p.gly4fsx50 c.138+1g>a
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Targeted next-generation sequencing reveals MODY in up to 6.5% of antibody-negative diabetes cases listed in the Norwegian Childhood Diabetes Registry
 Diabetologia (2017) 60:625 635 DOI 10.1007/s00125-016-4167-1 ARTICLE Targeted next-generation sequencing reveals MODY in up to 6.5% of antibody-negative diabetes cases listed in the Norwegian Childhood
Diabetologia (2017) 60:625 635 DOI 10.1007/s00125-016-4167-1 ARTICLE Targeted next-generation sequencing reveals MODY in up to 6.5% of antibody-negative diabetes cases listed in the Norwegian Childhood
Whole Exome Sequenced Characteristics
 Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Integrated analysis of cancer-related pathways affected by genetic and epigenetic alterations in gastric cancer
 Gastric Cancer (2015) 18:65 76 DOI 10.1007/s10120-014-0348-0 ORIGINAL ARTICLE Integrated analysis of cancer-related pathways affected by genetic and epigenetic alterations in gastric cancer Yukie Yoda
Gastric Cancer (2015) 18:65 76 DOI 10.1007/s10120-014-0348-0 ORIGINAL ARTICLE Integrated analysis of cancer-related pathways affected by genetic and epigenetic alterations in gastric cancer Yukie Yoda
The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation
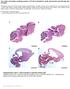 The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation Anita Becker-Heck#, Irene Zohn#, Noriko Okabe#, Andrew Pollock#, Kari Baker Lenhart,
The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation Anita Becker-Heck#, Irene Zohn#, Noriko Okabe#, Andrew Pollock#, Kari Baker Lenhart,
Culture Density (OD600) 0.1. Culture Density (OD600) Culture Density (OD600) Culture Density (OD600) Culture Density (OD600)
 A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP
 Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label)
 July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
Urea cycle disorders in Spain: an observational, cross-sectional and multicentric study of 104 cases
 Martín-Hernández et al. Orphanet Journal of Rare Diseases 2014, 9:187 RESEARCH Open Access Urea cycle disorders in Spain: an observational, cross-sectional and multicentric study of 104 cases Elena Martín-Hernández
Martín-Hernández et al. Orphanet Journal of Rare Diseases 2014, 9:187 RESEARCH Open Access Urea cycle disorders in Spain: an observational, cross-sectional and multicentric study of 104 cases Elena Martín-Hernández
 www.lessonplansinc.com Topic: Protein Synthesis - Sentence Activity Summary: Students will simulate transcription and translation by building a sentence/polypeptide from words/amino acids. Goals & Objectives:
www.lessonplansinc.com Topic: Protein Synthesis - Sentence Activity Summary: Students will simulate transcription and translation by building a sentence/polypeptide from words/amino acids. Goals & Objectives:
Supplemental Data. Shin et al. Plant Cell. (2012) /tpc YFP N
 MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
Case Report Congenital Insensitivity to Pain: A Case Report and Review of the Literature
 Case Reports in Neurological Medicine, Article ID 141953, 4 pages http://dx.doi.org/10.1155/2014/141953 Case Report Congenital Insensitivity to Pain: A Case Report and Review of the Literature Leema Reddy
Case Reports in Neurological Medicine, Article ID 141953, 4 pages http://dx.doi.org/10.1155/2014/141953 Case Report Congenital Insensitivity to Pain: A Case Report and Review of the Literature Leema Reddy
Brief Communication Diagnostic Genetics
 Brief Communication Diagnostic Genetics Ann Lab Med 2015;35:535-539 http://dx.doi.org/10.3343/alm.2015.35.5.535 ISSN 2234-3806 eissn 2234-3814 CYP21A2 Mutation Analysis in Korean Patients With Congenital
Brief Communication Diagnostic Genetics Ann Lab Med 2015;35:535-539 http://dx.doi.org/10.3343/alm.2015.35.5.535 ISSN 2234-3806 eissn 2234-3814 CYP21A2 Mutation Analysis in Korean Patients With Congenital
Clinical spectrum and molecular basis of recessive congenital methemoglobinemia in India
 Clin Genet 2015: 87: 62 67 Printed in Singapore. All rights reserved Short Report 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12326 Clinical spectrum
Clin Genet 2015: 87: 62 67 Printed in Singapore. All rights reserved Short Report 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12326 Clinical spectrum
