Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes
|
|
|
- Louisa Barrett
- 5 years ago
- Views:
Transcription
1 Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes KM Kortüm 1,8 *, EK Mai 3,4 *, NH Hanafiah 3 *, CX Shi 1, YX Zhu 1, L Bruins 1, S Barrio 1, P Jedlowski 1, M Merz 4, J Xu 3,6, RA Stewart 1, M Andrulis 6, A Jauch 7, J Hillengass 4, H Goldschmidt 4, 5, PL Bergsagel 1, E Braggio 1, AK Stewart 1,2#, and MS Raab 3,4 *contributed equally to this study Supplementary data and information Supplementary Figure 1 Supplementary Figure 2 Supplementary Table 1 Supplementary Table 2 Supplementary methods 1
2 Supplementary Figure 1 Supplementary Figure 1: Genomic mutations, adverse cytogenetic aberrations and treatment characteristics of 50 refractory multiple myeloma patients sorted by drug refractory status and CRBN pathway mutations. To enhance comparability, a second version of Figure 1, sorted by CRBN associated genes (CRBN, IRF4, IKZF-1 and CUL4B, thereafter according to mutation frequency; top to bottom) and patients refractory status immediately prior to sampling (IMIDrefractory, PI-refractory, refractory to other drug classed and not refractory; from left to right) is provided. As in Figure 1, all patients are labeled with individual numbers. Refractory status to immunomodulatory drugs (IMiDs) or proteasome inhibitors (PIs) at any time prior to sampling is shown as well as progressive disease (PD) and refractory (IMiD, PI, other drug classes) status immediately prior to sampling. Information on the adverse cytogenetic aberrations gain 1q21 (> 2 copies), deletion (17p), translocation t(4;14) and t(14;16) are depicted. Genes mutated in at least one case are displayed. Mutation frequencies are displayed by red bars. 2
3 Treatment characteristics, cytogenetic aberrations are color-coded: red = yes; green = no; grey = unknown/not available; refractory drugs immediately prior to sampling: purple = IMiD-refractory; blue = PI-refractory; orange = other drug classes (including cytotoxic agents and antibodies); green = not refractory to treatment immediately prior to sampling. 3
4 Supplementary Figure 2 Patient #32 Patient #24 Patient #12 Supplementary Figure 2: Clonal evolution in 3 MM patients with acquired IMiD resistance at time point 2. In all three cases a subclonal CRBN mutation was identified at that time point that was not detectable at prior, IMiD sensitive time point 1. 4
5 Supplementary Table 1: Supplementary Table 1: Genes included in M 3 Pv3.0 design 5
6 Supplementary Table 2: Variant Table gene patient # chromosomal position coding type VR% transcript Ensembl transcript ID location function protein SIFT Provean PREDICTION PREDICTION (cutoff=0.05) (cutoff=-2.5) XBP1 1 chr22: c.940_958delctgggtatctcaaatctgc INDEL 60% NM_ ENST exonic frameshiftdeletion p.leu314ffs*38 NA NA NA TP53 1 chr17: c.814g>a SNV 90% NM_ ENST exonic missense p.val272met Damaging Neutral probably damaging RASA2 1 chr3: c.643a>t SNV 30% NM_ ENST exonic nonsense p.lys215ter NA NA NA PIM3 1 chr22: c.898_899delinsac SNV 52% NM_ ENST exonic missense p.val300thr Tolerated Neutral benign BRAF 1 chr7: c.1799t>a SNV 64% NM_ ENST exonic missense p.val600glu Damaging Deleterious probably damaging DUSP2 1 chr2: c.719t>g SNV 50% NM_ ENST exonic missense p.ile240arg Damaging Deleterious probably damaging MAFB 1 chr20: c.725a>c SNV 49% NM_ ENST exonic missense p.lys242thr Damaging Deleterious probably damaging KRAS 2 chr12: c.57g>t SNV 50% NM_ ENST exonic missense p.leu19phe Damaging Deleterious probably damaging MAF 2 chr16: c.824a>g SNV 91% NM_ ENST exonic missense p.asn275ser Damaging Deleterious probably damaging IKZF1 2 chr7: c.454g>a SNV 31% NM_ ENST exonic missense p.ala152thr Damaging Deleterious probably damaging TP53 3 chr17: c.412_412delg INDEL 95% NM_ ENST exonic frameshiftdeletion p.ala138fs NA NA NA NRAS 3 chr1: c.183a>c SNV 58% NM_ ENST exonic missense p.gln61his Damaging Deleterious benign RB1 3 chr13: c.958c>t SNV 100% NM_ ENST exonic nonsense p.arg320ter NA NA NA FAM46C 3 chr1: c.569_573delctttg INDEL 51% NM_ ENST exonic frameshiftdeletion p.leu191fs NA NA NA SP140 4 chr2: c.1618a>g SNV 3% NM_ ENST exonic missense p.asn540asp Tolerated Neutral benign CRBN 5 chr3: c _1178delggtatgcctggactgttgcccagtgtaagat INDEL 38% NM_ ENST splicesite_5 frameshiftdeletion p.i393mfs10 NA NA NA KRAS 5 chr12: c.183a>c SNV 27% NM_ ENST exonic missense p.gln61his Damaging Deleterious benign TRAF3 5 chr14: c.1081_1081delg INDEL 38% NM_ ENST exonic frameshiftdeletion p.val361fs NA NA NA CUL4B 6 chrx: c.2221a>g SNV 5% NM_ ENST exonic missense p.lys741glu Damaging Deleterious probably damaging TP53 6 chr17: c.738g>a SNV 14% NM_ ENST exonic missense p.met246ile Damaging Deleterious probably damaging TP53 6 chr17: c.281c>g SNV 7% NM_ ENST exonic nonsense p.ser94ter NA NA NA SHC1 6 chr1: c.904a>c SNV 12% NM_ ENST exonic missense p.ser302arg Damaging Deleterious possibly damaging CUL4B 6 chrx: SNV 18% NM_ ENST splicesite_3 missense NA NA NA KRAS 7 chr12: c.35g>t SNV 95% NM_ ENST exonic missense p.gly12val Damaging Deleterious probably damaging IDH1 7 chr2: c.395g>a SNV 50% NM_ ENST exonic missense p.arg132his Damaging Deleterious benign NRAS 8 chr1: c.35g>c SNV 80% NM_ ENST exonic missense p.gly12ala Damaging Deleterious possibly damaging CDKN2C 8 chr1: c.133a>g SNV 7% NM_ ENST exonic missense p.met45val Damaging Deleterious benign CYLD 8 chr16: c.1096_1096delc INDEL 50% NM_ ENST exonic frameshiftdeletion p.q366nfs*42 NA NA NA BRAF 8 chr7: c.1397g>t SNV 10% NM_ ENST exonic missense p.gly466val Damaging Deleterious probably damaging KRAS 9 chr12: c.37g>t SNV 37% NM_ ENST exonic missense p.gly13cys Damaging Deleterious probably damaging EGR1 9 chr5: c.103c>g SNV 36% NM_ ENST exonic missense p.leu35val Damaging Neutral probably damaging TP53 11 chr17: c.584t>c SNV 13% NM_ ENST exonic missense p.ile195thr Damaging Deleterious probably damaging CRBN 12 chr3: c.979c>t SNV 7% NM_ ENST exonic nonsense p.gln327ter NA NA NA DIS3 12 chr13: c.1460a>t SNV 48% NM_ ENST exonic missense p.asp487val Damaging Deleterious probably damaging NRAS 12 chr1: c.34g>c SNV 50% NM_ ENST exonic missense p.gly12arg Damaging Deleterious possibly damaging PIM1 12 chr6: c.256g>a SNV 70% NM_ ENST exonic missense p.val86met Damaging Neutral possibly damaging NRAS 13 chr1: c.182a>g SNV 20% NM_ ENST exonic missense p.gln61arg Damaging Deleterious benign STAT3 13 chr17: c.28c>g SNV 29% NM_ ENST exonic missense p.leu10val Damaging Neutral probably damaging MYD88 14 chr3: c.666t>a SNV 5% NM_ ENST exonic missense p.ser222arg Damaging Neutral probably damaging CUL4B 15 chrx: c.2459g>c SNV 25% NM_ ENST exonic missense p.arg820thr Damaging Deleterious probably damaging NRAS 15 chr1: c.38g>t SNV 25% NM_ ENST exonic missense p.gly13val Damaging Deleterious probably damaging FAM46C 15 chr1: c.869_873delcttac INDEL 43% NM_ ENST exonic frameshiftdeletion p.tyr291fs NA NA NA Polyphen 2 prediction 6
7 KRAS 16 chr12: c.34g>t SNV 56% NM_ ENST exonic missense p.gly12cys Damaging Deleterious probably damaging IRF4 17 chr6: c.368a>g SNV 47% NM_ ENST exonic missense p.lys123arg Damaging Deleterious probably damaging ATM 17 chr11: INDEL 45% NM_ ENST splicesite_5 frameshiftdeletion NA NA NA CCND1 17 chr11: c.86g>t SNV 41% NM_ ENST exonic missense p.arg29leu Damaging Deleterious benign KRAS 17 chr12: c.351a>t SNV 58% NM_ ENST exonic missense p.lys117asn Damaging Deleterious probably damaging TP53 18 chr17: c.518t>a SNV 13% NM_ ENST exonic missense p.val173glu Damaging Deleterious probably damaging KRAS 19 chr12: c.183a>c SNV 17% NM_ ENST exonic missense p.gln61his Damaging Deleterious benign KRAS 20 chr12: c.183a>c SNV 17% NM_ ENST exonic missense p.gln61his Damaging Deleterious benign TP53 20 chr17: c.524g>a SNV 38% NM_ ENST exonic missense p.arg175his Damaging Deleterious possibly damaging KRAS 21 chr12: c.35g>a SNV 49% NM_ ENST exonic missense p.gly12asp Damaging Deleterious possibly damaging BIRC2 21 chr11: c.184g>t SNV 3% NM_ ENST exonic missense p.val62phe Damaging Deleterious possibly damaging KRAS 22 chr12: c.35g>c SNV 19% NM_ ENST exonic missense p.gly12ala Damaging Deleterious possibly damaging BRAF 23 chr7: c.1780g>a SNV 31% NM_ ENST exonic missense p.asp594asn Damaging Deleterious probably damaging PTPN11 23 chr12: c.1507g>a SNV 39% NM_ ENST exonic missense p.gly503arg Damaging Deleterious probably damaging CRBN 24 chr3: c.1142t>g SNV 7% NM_ ENST exonic missense p.phe381cys Damaging Deleterious probably damaging BRAF 24 chr7: c.1799t>a SNV 27% NM_ ENST exonic missense p.val600glu Damaging Deleterious probably damaging KRAS 25 chr12: c.38g>a SNV 38% NM_ ENST exonic missense p.gly13asp Damaging Deleterious possibly damaging TP53 25 chr17: c.481g>a SNV 3% NM_ ENST exonic missense p.ala161thr Damaging Neutral probably damaging CARD11 26 chr7: c.2471a>c SNV 31% NM_ ENST exonic missense p.asp824ala Tolerated Neutral benign NRAS 27 chr1: c.181c>a SNV 38% NM_ ENST exonic missense p.gln61lys Damaging Deleterious possibly damaging NFKB2 27 chr10: c.1745t>c SNV 21% NM_ ENST exonic missense p.leu582pro Damaging Deleterious probably damaging NFKB2 27 chr10: c.1916a>c SNV 5% NM_ ENST exonic missense p.his639pro Damaging Deleterious probably damaging ATM 28 chr11: c.3378a>c SNV 58% NM_ ENST exonic missense p.lys1126asn Tolerated Neutral possibly damaging NRAS 28 chr1: c.38g>a SNV 54% NM_ ENST exonic missense p.gly13asp Damaging Deleterious benign IRF4 29 chr6: c.176a>g SNV 39% NM_ ENST exonic missense p.lys59arg Tolerated Deleterious probably damaging BIRC3 29 chr11: c.599g>c SNV 29% NM_ ENST exonic missense p.gly200ala Tolerated Neutral possibly damaging KRAS 30 chr12: c.183a>c SNV 58% NM_ ENST exonic missense p.gln61his Damaging Deleterious benign MAX 30 chr14: c.214c>t SNV 77% NM_ ENST exonic nonsense p.gln72* NA NA NA TP53 30 chr17: c.840a>c SNV 41% NM_ ENST exonic missense p.arg280ser Damaging Deleterious probably damaging TRAF3 30 chr14: c.1418t>g SNV 71% NM_ ENST exonic missense p.leu473arg Damaging Deleterious probably damaging SP chr2: c.1711c>t SNV 42% NM_ ENST exonic nonsense p.arg571ter NA NA NA DIS3 31 chr13: c.2325t>g SNV 87% NM_ ENST exonic missense Phe775Leu Damaging Deleterious probably damaging BRAF 31 chr7: c.1799t>a SNV 42% NM_ ENST exonic missense p.val600glu Damaging Deleterious probably damaging CRBN 32 chr3: c.1232c>a SNV 5% NM_ ENST exonic missense p.pro411his Damaging Deleterious probably damaging TLR4 32 chr9: c.1212a>c SNV 9% NM_ ENST exonic missense p.leu404phe Damaging Deleterious probably damaging KRAS 32 chr12: c.35g>a SNV 49% NM_ ENST exonic missense p.gly12asp Damaging Deleterious possibly damaging STAT3 32 chr17: c.394c>g SNV 11% NM_ ENST exonic missense p.pro132ala Tolerated Neutral probably damaging NFKB2 32 chr10: c.2200c>g SNV 5% NM_ ENST exonic missense p.leu734val Damaging Neutral benign BRAF 33 chr7: c.1405g>a SNV 17% NM_ ENST exonic missense p.gly469arg Damaging Deleterious probably damaging CRBN 34 chr3: SNV 21% NM_ ENST splicesite_5 missense NA NA NA BRAF 34 chr7: c.1799t>a SNV 35% NM_ ENST exonic missense p.val600glu Damaging Deleterious probably damaging IGF1R 35 chr15: c.2558c>t SNV 22% NM_ ENST exonic missense p.pro853leu Tolerated Neutral benign NRAS 36 chr1: c.38g>a SNV 13% NM_ ENST exonic missense p.gly13asp Damaging Deleterious benign 7
8 TRAF3 37 chr14: SNV 3% NM_ ENST splicesite_3 missense NA NA NA FAM46C 37 chr1: c.64_77delgttagccggctgca INDEL 25% NM_ ENST exonic frameshiftdeletion p.val22fs NA NA NA IL6ST 37 chr5: c.647c>a SNV 39% NM_ ENST exonic missense p.pro216his Damaging Deleterious probably damaging CUL4B 38 chrx: c.1270g>a SNV 5% NM_ ENST exonic missense p.glu424lys Damaging Deleterious probably damaging NR3C1 38 chr5: c.1832g>a SNV 7% NM_ ENST exonic missense p.arg611lys Damaging Deleterious probably damaging CUL4B 38 chrx: c.1219g>a SNV 5% NM_ ENST exonic missense p.glu407lys Tolerated Neutral benign PSMD1 38 chr2: c.11c>g SNV 8% NM_ ENST exonic missense p.ser4trp Damaging Deleterious probably damaging KRAS 38 chr12: c.34g>a SNV 46% NM_ ENST exonic missense p.gly12ser Damaging Deleterious possibly damaging PIK3CA 38 chr3: c.269g>a SNV 3% NM_ ENST exonic missense p.cys90tyr Damaging Deleterious probably damaging CARD11 38 chr7: c.3424g>a SNV 4% NM_ ENST exonic missense p.glu1142lys Damaging Neutral probably damaging PSMB8 38 chr6: c.92c>t SNV 5% NM_ ENST exonic missense p.thr31met Tolerated Neutral NA ATM 38 chr11: c.1409c>g SNV 6% NM_ ENST exonic nonsense p.ser470ter NA NA NA NR3C1 38 chr5: c.1027c>t SNV 17% NM_ ENST exonic nonsense p.gln343ter NA NA NA NR3C1 38 chr5: c.1375g>a SNV 7% NM_ ENST exonic missense p.gly459arg Damaging Deleterious probably damaging CRBN 38 chr3: c.721_722insgggc INDEL 25% NM_ ENST exonic frameshiftinsertion p.pro241argfs*10 NA NA NA NR3C1 38 chr5: c.1465g>t SNV 8% NM_ ENST exonic nonsense p.glu489ter NA NA NA ATM 39 chr11: c.2572t>c SNV 35% NM_ ENST exonic missense Phe858Leu Tolerated Deleterious possibly damaging FGFR3 40 chr4: c.742c>t SNV 42% NM_ ENST exonic missense p.arg248cys Damaging Deleterious probably damaging NFKBIA 40 chr14: c.650g>a SNV 8% NM_ ENST exonic missense p.gly217asp Damaging Deleterious probably damaging FAM46C 40 chr1: c.274g>a SNV 5% NM_ ENST exonic missense p.asp92asn Damaging Deleterious probably damaging RASA2 40 chr3: c.539g>a SNV 8% NM_ ENST exonic missense p.cys180tyr Damaging Deleterious probably damaging BRAF 41 chr7: c.1497a>c SNV 52% NM_ ENST exonic missense p.lys499asn Damaging Deleterious possibly damaging BRAF 41 chr7: c.2083g>a SNV 39% NM_ ENST exonic missense p.glu695lys Damaging Deleterious benign TP53 41 chr17: c.722c>g SNV 90% NM_ ENST exonic missense p.ser241cys Damaging Deleterious probably damaging NFKB2 42 chr10: c.1336t>g SNV 14% NM_ ENST exonic missense p.tyr446asp Tolerated Neutral possibly damaging KRAS 43 chr12: c.53c>t SNV 70% NM_ ENST exonic missense p.ala18val Damaging Deleterious probably damaging TP53 43 chr17: c.542g>a SNV 7% NM_ ENST exonic missense p.arg181his Damaging Neutral probably damaging BTG1 44 chr12: c.139c>a SNV 38% NM_ ENST exonic missense p.leu47met Tolerated Neutral benign BIRC2 44 chr11: c.488c>g SNV 8% NM_ ENST exonic missense p.ser114cys Damaging Neutral benign TRAF3 44 chr14: c.1678g>c SNV 45% NM_ ENST exonic missense p.val560leu Damaging Deleterious probably damaging CYLD 44 chr16: c.2497c>t SNV 27% NM_ ENST exonic missense p.his833tyr Damaging Deleterious probably damaging NRAS 45 chr1: c.182a>g SNV 22% NM_ ENST exonic missense p.gln61arg Damaging Deleterious benign NFKB2 45 chr10: c.1069g>c SNV 19% NM_ ENST exonic missense p.gly357arg Tolerated Deleterious probably damaging NRAS 46 chr1: c.182a>g SNV 12% NM_ ENST exonic missense p.gln61arg Damaging Deleterious benign TP53 46 chr17: c.329g>t SNV 16% NM_ ENST exonic missense p.arg110leu Damaging Deleterious benign TP53 46 chr17: c.452c>g SNV 7% NM_ ENST exonic missense p.pro151arg Damaging Deleterious probably damaging FAM46C 46 chr1: c.882c>g SNV 16% NM_ ENST exonic missense p.asn294lys Damaging Deleterious benign NRAS 47 chr1: c.182a>g SNV 36% NM_ ENST exonic missense p.gln61arg Damaging Deleterious benign TP53 47 chr17: c.395a>g SNV 6% NM_ ENST exonic missense p.lys132arg Damaging Deleterious probably damaging ATM 48 chr11: c.6067g>a SNV 56% NM_ ENST exonic missense p.gly2023arg Damaging Deleterious NA NRAS 48 chr1: c.183a>t SNV 7% NM_ ENST exonic missense p.gln61his Damaging Deleterious benign KRAS 49 chr12: c.71t>a SNV 14% NM_ ENST exonic missense p.ile24asn Damaging Deleterious probably damaging FGFR3 49 chr4: c.2427a>c SNV 35% NM_ ENST exonic stop_lost p.*809c Damaging Neutral NA DIS3 49 chr13: c.2258a>g SNV 22% NM_ ENST exonic missense p.tyr753cys Damaging Deleterious probably damaging BRAF 49 chr7: c.1780g>a SNV 31% NM_ ENST exonic missense p.asp594asn Damaging Deleterious probably damaging IDH2 49 chr15: c.632a>g SNV 3% NM_ ENST exonic missense p.asn211ser Damaging Neutral benign CDKN1B 49 chr12: c.163_164delgc InDelframeshift 12% NM_ ENST exonic frameshiftdeletion p.ala55glufs*69 NA NA NA IL6ST 50 chr5: g.38954g>a SNV 37% NM_ ENST exonic missense p.ala418thr Tolerated NA benign NRAS 50 chr1: c.35g>a SNV 61% NM_ ENST exonic missense p.gly12asp Damaging Deleterious benign FAM46C 50 chr1: c.448_456delgaccgctgg INDEL 7% NM_ ENST exonic frameshiftdeletion NA Deleterious NA TP53 50 chr17: c.451c>a SNV 40% NM_ ENST exonic missense p.pro151thr Damaging Deleterious possibly damaging 8
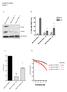
9 Supplementary methods: Details on preparation of the lentiviral vectors for CRBN mutants and wild type: We first created a lenti-vector pcdhpurocrbn to overexpress wild type CRBN by subclone wild type CRBN from pcdhgfpcrbn as previously reported by our group. Then we used pcdhpurocrbn as a template to introduce the CRBN mutations. All mutations were introduced by PCR. In patient #12 (chr3: , genotype G/A) the mutation causes an early termination of transcription. The following pair of primers was used for this patient (forward primer: 5 -cacgctgttttgacctccatagaagattc and reverse primer: 5 - gtctcaggatccttaacattgtttacagcaaagg) to introduce the mutation (stop code added at 3 primer). For creating the CRBN mutation of patient #24 (chr3: genotype A/C) we used two rounds of PCR. In the first round we ran two PCRs with two pair of primers (PCR1: forward primer: 5 -cacgctgttttgacctccatagaagattc and reverse primer: 5 - aggacaccagctgtgttctgtagaa; PCR2: forward primer: 5 - cacagctggtgtcctgggtatgcctggactg and reverse primer: 5 - ACCGGAGCGATCGCAGATCCTTC). The point mutation was introduced by the reverse primer in the PCR1 and forward primer at PCR2). After purification PCR1 and PCR2 was combined together as a template, then amplified by a pair of primers (forward primer: 5 -Cacgctgttttgacctccatagaagattc and reverse primer: 5 - ACCGGAGCGATCGCAGATCCTTC). The mutation of patient #32 was introduced in a way similar with patient #24. Mutated CRBN was cloned into a lenti-vector pcdh-cmv-mcs-ef1-puro (SBI, Mountain View, CA 94043) which was packaged into lentivirus and infected myeloma cells. After selection with puromycin My5 cells over expression either wild type or mutated CRBN were tested response against LEN. Colorimetric assays were also performed to assay drug activity. Cells from 48-hour cultures were pulsed with 10 μl of 5 mg/ml 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrasodium bromide (MTT; Chemicon International Inc, Temecula, CA) to each well for 4 hours, followed by 100 μl isopropanol that contained 0.04 HCl. Absorbance readings at a wavelength of 570 nm were taken on a spectrophotometer. 9
Names for ABO (ISBT 001) Blood Group Alleles
 Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
Names for H (ISBT 018) Blood Group Alleles
 Names for H (ISBT 018) Blood Group Alleles General description: The H blood group system consists of one antigen, H, that is carried on glycolipids and glycoproteins on the RBC membrane, where it is synthesised
Names for H (ISBT 018) Blood Group Alleles General description: The H blood group system consists of one antigen, H, that is carried on glycolipids and glycoproteins on the RBC membrane, where it is synthesised
Supplementary materials and methods
 Supplementary materials and methods Targeted resequencing RNA probes complementary in sequence to the coding exons (+ 2bp) of target genes were designed using e-array (https://earray.chem.agilent.com/earray/)
Supplementary materials and methods Targeted resequencing RNA probes complementary in sequence to the coding exons (+ 2bp) of target genes were designed using e-array (https://earray.chem.agilent.com/earray/)
Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes
 Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes Supplementary Information Jason Flannick*, Nicola L Beer*, Alexander G Bick,
Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes Supplementary Information Jason Flannick*, Nicola L Beer*, Alexander G Bick,
Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia
 Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia Erpelinck-Verschueren, Veronika Rockova, Mathijs Sanders,
Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia Erpelinck-Verschueren, Veronika Rockova, Mathijs Sanders,
Colclough et al., Human Mutation 1
 Region Colclough et al., Human Mutation 1 Supp. Table S1. List of HNF1A mutations Nucleotide Change NM_000545.5 Promoter and 5 UTR Mutations Protein Effect NM_000545.5 Three letter amino acid description
Region Colclough et al., Human Mutation 1 Supp. Table S1. List of HNF1A mutations Nucleotide Change NM_000545.5 Promoter and 5 UTR Mutations Protein Effect NM_000545.5 Three letter amino acid description
Supplementary material PCNT point mutations and familial intracranial aneurysms
 Supplementary material PCNT point mutations and familial intracranial aneurysms Oswaldo Lorenzo-Betancor, MD, PhD 1, Patrick R. Blackburn, PhD 2,3, Emily Edwards, CCRP 4, Rocío Vázquez-do-Campo, MD 4,
Supplementary material PCNT point mutations and familial intracranial aneurysms Oswaldo Lorenzo-Betancor, MD, PhD 1, Patrick R. Blackburn, PhD 2,3, Emily Edwards, CCRP 4, Rocío Vázquez-do-Campo, MD 4,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10833 Supplementary Table 1: Clinical information for 48 whole exome sequencing samples Sample ID Age Gender tumor location Death OS (months) Recurrence PFS
SUPPLEMENTARY INFORMATION doi:10.1038/nature10833 Supplementary Table 1: Clinical information for 48 whole exome sequencing samples Sample ID Age Gender tumor location Death OS (months) Recurrence PFS
GALAFOLD (migalastat) capsules, for oral use Initial U.S. Approval: 2018
 HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
LMM / emerge III Network Reference Sequences October 2016 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene
 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
ALANINE:GLYOXYLATE AMINOTRANSFERASE (AGXT) GENE (PRIMARY HYPEROXALURIA TYPE 1) MUTATION DATA BASE
 ALANINE:GLYOXYLATE AMINOTRANSFERASE (AGXT) GENE (PRIMARY HYPEROXALURIA TYPE 1) REFERENCE SEQUENCES: c.dna: NM_000030 g.dna: NG_008005.1 MUTATION DATA BASE Nomenclature: HGVS guidelines (http://www.hgvs.org/rec.html)
ALANINE:GLYOXYLATE AMINOTRANSFERASE (AGXT) GENE (PRIMARY HYPEROXALURIA TYPE 1) REFERENCE SEQUENCES: c.dna: NM_000030 g.dna: NG_008005.1 MUTATION DATA BASE Nomenclature: HGVS guidelines (http://www.hgvs.org/rec.html)
GALAFOLD (migalastat) capsules, for oral use Initial U.S. Approval: 2018
 HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
De Leeneer et al., Human Mutation 1
 De Leeneer et al., Human Mutation 1 Supp. Table S1A. Overview of the genes included in the workflow Ensembl Gene ID Ensembl Transcript ID Associated Gene Name ENSG00000198691 ENST00000370225 ABCA4 NM_000350.2
De Leeneer et al., Human Mutation 1 Supp. Table S1A. Overview of the genes included in the workflow Ensembl Gene ID Ensembl Transcript ID Associated Gene Name ENSG00000198691 ENST00000370225 ABCA4 NM_000350.2
Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome
 c Indian Academy of Sciences RESEARCH NOTE Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome FENGJIE YANG, YAXIAN CHEN, YU ZHANG, LIRU QIU, YU CHEN and JIANHUA ZHOU Department
c Indian Academy of Sciences RESEARCH NOTE Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome FENGJIE YANG, YAXIAN CHEN, YU ZHANG, LIRU QIU, YU CHEN and JIANHUA ZHOU Department
Genetic Testing and Analysis. (858) MRN: Specimen: Saliva Received: 07/26/2016 GENETIC ANALYSIS REPORT
 GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
analysis in hereditary breast/ovarian cancer families of Portuguese ancestry
 Clin Genet 2015: 88: 41 48 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12441 The role of targeted
Clin Genet 2015: 88: 41 48 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12441 The role of targeted
InheriGen Plus Disease Information
 DISEASE INFORMATION & MUTATIONS TESTED 17α-hydroxylase/17,20-lyase Deficiency CYP17A1 Brazilian 87% < 1 in 112 < 1 in 850 p.arg362cys, p.trp406arg, c.1243+5g>a, p.pro48fs, p.asp487_phe489del, p.tyr32x
DISEASE INFORMATION & MUTATIONS TESTED 17α-hydroxylase/17,20-lyase Deficiency CYP17A1 Brazilian 87% < 1 in 112 < 1 in 850 p.arg362cys, p.trp406arg, c.1243+5g>a, p.pro48fs, p.asp487_phe489del, p.tyr32x
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Intron p Hybrid with p.leu245val p. Gly336Cys. p.leu62phe p. Ala137Val p. Asn152Thr hybrid with p.leu245val p.
 RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM
 Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel
 ACGS, June 2018 The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel Ian M. Frayling Member of Council & Variant Interpretation Committee (VIC) International Society for Gastrointestinal
ACGS, June 2018 The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel Ian M. Frayling Member of Council & Variant Interpretation Committee (VIC) International Society for Gastrointestinal
CHAPTER IV RESULTS Microcephaly General description
 47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
Sample Test Report. Mayo Clinic GeneGuide. Report. Consumer Name DOB: 00/00/0000
 Mayo Clinic GeneGuide Report Consumer Name DOB: // Table Of Contents Mayo Clinic GeneGuide Genetic Test Results Demographics and Ordering Information 3 How to Use This Report 4 Carrier Screening - No variants
Mayo Clinic GeneGuide Report Consumer Name DOB: // Table Of Contents Mayo Clinic GeneGuide Genetic Test Results Demographics and Ordering Information 3 How to Use This Report 4 Carrier Screening - No variants
Supplementary Table 1. PIK3CA mutation in colorectal cancer
 Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures
 Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures Item Type text; Electronic Thesis Authors Chen, Debbie Publisher The University of Arizona. Rights Copyright is held by the
Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures Item Type text; Electronic Thesis Authors Chen, Debbie Publisher The University of Arizona. Rights Copyright is held by the
Hearing Loss Imaging Thyroid function tests TFT (age at test)
 Clinical Information on patients. Unclassified variants are denoted in brackets. EVA is enlarged vestibular aqueducts - Indicates no information. means asian. Patient TFT (age at test) 25215 c.1001+1g>a
Clinical Information on patients. Unclassified variants are denoted in brackets. EVA is enlarged vestibular aqueducts - Indicates no information. means asian. Patient TFT (age at test) 25215 c.1001+1g>a
Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension
 Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension Felix Beuschlein, Sheerazed Boulkroun, Andrea Osswald, Thomas Wieland, Hang N. Nielsen, Urs D. Lichtenauer,
Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension Felix Beuschlein, Sheerazed Boulkroun, Andrea Osswald, Thomas Wieland, Hang N. Nielsen, Urs D. Lichtenauer,
Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss
 Genetics Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss Akhila Alapati, 1 Kerry Goetz, 2 John Suk, 1 Mili Navani,
Genetics Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss Akhila Alapati, 1 Kerry Goetz, 2 John Suk, 1 Mili Navani,
Genotype to phenotype correlations in cartilage oligomeric matrix protein associated chondrodysplasias
 (2014) 22, 1278 1282 & 2014 Macmillan Publishers Limited All rights reserved 1018-4813/14 www.nature.com/ejhg ARTICLE Genotype to phenotype correlations in cartilage oligomeric matrix protein associated
(2014) 22, 1278 1282 & 2014 Macmillan Publishers Limited All rights reserved 1018-4813/14 www.nature.com/ejhg ARTICLE Genotype to phenotype correlations in cartilage oligomeric matrix protein associated
ATM germline heterozygosity does not play a role in CLL initiation but influences rapid disease progression through loss of the remaining ATM allele
 Published Ahead of Print on September 20, 2011, as doi:10.3324/haematol.2011.048827. Copyright 2011 Ferrata Storti Foundation. Early Release Paper ATM germline heterozygosity does not play a role in CLL
Published Ahead of Print on September 20, 2011, as doi:10.3324/haematol.2011.048827. Copyright 2011 Ferrata Storti Foundation. Early Release Paper ATM germline heterozygosity does not play a role in CLL
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Table e-1: Investigation of 33 patients with early onset epilepsy for KCNT1 mutations.
 Table e1: Investigation of 33 patients with early onset epilepsy for KCNT1 mutations. Patient Phenotype Screening Method Diagnostic Karyotype Sanger sequencing NGS Diagnostic Panel WES chromosomal microarray
Table e1: Investigation of 33 patients with early onset epilepsy for KCNT1 mutations. Patient Phenotype Screening Method Diagnostic Karyotype Sanger sequencing NGS Diagnostic Panel WES chromosomal microarray
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
 Int. J. Cancer: 116, 692 702 (2005) ' 2005 Wiley-Liss, Inc. Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
Int. J. Cancer: 116, 692 702 (2005) ' 2005 Wiley-Liss, Inc. Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
p.arg119gly p.arg119his p.ala179thr c.540+1g>a c.617_633+6del Prediction basis structure
 a Missense ATG p.arg119gly p.arg119his p.ala179thr p.ala189val p.gly206trp p.gly206arg p.arg251his p.ala257thr TGA 5 UTR 1 2 3 4 5 6 7 3 UTR Splice Site, Frameshift b Mutation p.gly206trp p.gly4fsx50 c.138+1g>a
a Missense ATG p.arg119gly p.arg119his p.ala179thr p.ala189val p.gly206trp p.gly206arg p.arg251his p.ala257thr TGA 5 UTR 1 2 3 4 5 6 7 3 UTR Splice Site, Frameshift b Mutation p.gly206trp p.gly4fsx50 c.138+1g>a
CIC Edizioni Internazionali
 Next generation sequencing in the identification of a rare genetic disease from preconceptional couple screening to preimplantation genetic diagnosis Claudio Dello Russo 1 Gianluca Di Giacomo 1 Alvaro
Next generation sequencing in the identification of a rare genetic disease from preconceptional couple screening to preimplantation genetic diagnosis Claudio Dello Russo 1 Gianluca Di Giacomo 1 Alvaro
6/12/2018. Disclosures. Clinical Genomics The CLIA Lab Perspective. Outline. COH HopeSeq Heme Panels
 Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH
 Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH Index patients (n=89) FH children Cascade screening (n=82) cdna Mutation Protein 3 0 c.10580g>a p.arg3527gln
Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH Index patients (n=89) FH children Cascade screening (n=82) cdna Mutation Protein 3 0 c.10580g>a p.arg3527gln
Whole Exome Sequenced Characteristics
 Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Germline Variation in Cancer-Susceptibility Genes in a Healthy, Ancestrally Diverse Cohort: Implications for Individual Genome Sequencing
 Germline Variation in Cancer-Susceptibility Genes in a Healthy, Ancestrally Diverse Cohort: Implications for Individual Genome Sequencing Dale L. Bodian, Justine N. McCutcheon, Prachi Kothiyal, Kathi C.
Germline Variation in Cancer-Susceptibility Genes in a Healthy, Ancestrally Diverse Cohort: Implications for Individual Genome Sequencing Dale L. Bodian, Justine N. McCutcheon, Prachi Kothiyal, Kathi C.
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics.
 Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics. Table S2. Gene symbols and data regarding potential oligogenic
Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics. Table S2. Gene symbols and data regarding potential oligogenic
Myeloma Genetics what do we know and where are we going?
 in partnership with Myeloma Genetics what do we know and where are we going? Dr Brian Walker Thames Valley Cancer Network 14 th September 2015 Making the discoveries that defeat cancer Myeloma Genome:
in partnership with Myeloma Genetics what do we know and where are we going? Dr Brian Walker Thames Valley Cancer Network 14 th September 2015 Making the discoveries that defeat cancer Myeloma Genome:
TP53 mutational profile in CLL : A retrospective study of the FILO group.
 TP53 mutational profile in CLL : A retrospective study of the FILO group. Fanny Baran-Marszak Hopital Avicenne Bobigny France 2nd ERIC workshop on TP53 analysis in CLL, Stresa 2017 TP53 abnormalities :
TP53 mutational profile in CLL : A retrospective study of the FILO group. Fanny Baran-Marszak Hopital Avicenne Bobigny France 2nd ERIC workshop on TP53 analysis in CLL, Stresa 2017 TP53 abnormalities :
Supplementary Information. Exome sequencing of serous endometrial tumors identifies recurrent somatic
 Supplementary Information Exome sequencing of serous endometrial tumors identifies recurrent somatic mutations in chromatin-remodeling and ubiquitin ligase complex genes Matthieu Le Gallo 1, Andrea J.
Supplementary Information Exome sequencing of serous endometrial tumors identifies recurrent somatic mutations in chromatin-remodeling and ubiquitin ligase complex genes Matthieu Le Gallo 1, Andrea J.
Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at-risk for hereditary breast cancer and characterization of its genetic ancestry
 /, 2016, Vol. 7, (No. 49), pp: 80465-80481 Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at- for hereditary breast cancer and characterization of its genetic ancestry Gabriela C.
/, 2016, Vol. 7, (No. 49), pp: 80465-80481 Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at- for hereditary breast cancer and characterization of its genetic ancestry Gabriela C.
chr6: _ del HFE NM_ ,744 bp deletion Deletion of entire HFE gene 2
 Table S1. Details of pathogenic and possibly pathogenic variants identified in the HH genes HFE, HFE2, HAMP, TFR2 and SLC40A1 from published reports. Variants have been classified as either 1, possibly
Table S1. Details of pathogenic and possibly pathogenic variants identified in the HH genes HFE, HFE2, HAMP, TFR2 and SLC40A1 from published reports. Variants have been classified as either 1, possibly
Introduction of an NGS gene panel into the Haemato-Oncology MPN service
 Introduction of an NGS gene panel into the Haemato-Oncology MPN service Dr. Anna Skowronska, Dr Jane Bryon, Dr Samuel Clokie, Dr Yvonne Wallis and Professor Mike Griffiths West Midlands Regional Genetics
Introduction of an NGS gene panel into the Haemato-Oncology MPN service Dr. Anna Skowronska, Dr Jane Bryon, Dr Samuel Clokie, Dr Yvonne Wallis and Professor Mike Griffiths West Midlands Regional Genetics
Supplementary Table 2. Identified causative mutations and/or mutation candidates.
 Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
Supplementary Materials for
 www.sciencemag.org/content/354/6319/aaf7000/suppl/dc1 Supplementary Materials for Genetic identification of familial hypercholesterolemia within a single U.S. health care system Noura S. Abul-Husn, Kandamurugu
www.sciencemag.org/content/354/6319/aaf7000/suppl/dc1 Supplementary Materials for Genetic identification of familial hypercholesterolemia within a single U.S. health care system Noura S. Abul-Husn, Kandamurugu
Frequent incidence of BARD1-truncating mutations in germline DNA from triple-negative breast cancer patients
 Clin Genet 2016: 89: 336 340 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12620 Frequent incidence
Clin Genet 2016: 89: 336 340 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12620 Frequent incidence
Supplementary Online Content
 Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
J Clin Oncol 34: by American Society of Clinical Oncology INTRODUCTION
 VOLUME 34 NUMBER 34 DECEMBER 1, 2016 JOURNAL OF CLINICAL ONCOLOGY O R I G I N A L R E P O R T Conflicting Interpretation of Genetic Variants and Cancer Risk by Commercial Laboratories as Assessed by the
VOLUME 34 NUMBER 34 DECEMBER 1, 2016 JOURNAL OF CLINICAL ONCOLOGY O R I G I N A L R E P O R T Conflicting Interpretation of Genetic Variants and Cancer Risk by Commercial Laboratories as Assessed by the
Table S2. Putative mutations identified in the CHH cohort
 Table S. Putative utations identified in the CHH cohort Sapl e Phenotyp e Gene nt change aa change Zyg SIF T PPH MaxEn t In vitro ExA C ExAC NFE Previous report ACMG classificatio n ACMG evidence codes
Table S. Putative utations identified in the CHH cohort Sapl e Phenotyp e Gene nt change aa change Zyg SIF T PPH MaxEn t In vitro ExA C ExAC NFE Previous report ACMG classificatio n ACMG evidence codes
Variant interpretation exercise. ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18
 Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Reverse phenotyping: MCD Detection without clinical diagnosis. Diagnostic: NGS panels versus WES
 Clinical Genetics Leading the way in genetic issues Reverse phenotyping: MCD Detection without clinical diagnosis Diagnostic: NGS panels versus WES Martina Wilke Laboratory Specialist Clinical Genetics
Clinical Genetics Leading the way in genetic issues Reverse phenotyping: MCD Detection without clinical diagnosis Diagnostic: NGS panels versus WES Martina Wilke Laboratory Specialist Clinical Genetics
REVIEWS 1. Genomic complexity of multiple myeloma and its clinical implications
 Genomic complexity of multiple myeloma and its clinical implications Salomon Manier 1,, Karma Z. Salem 1,, Jihye Park 1,, Dan A. Landau 3,, Gad Getz,5 and Irene M. Ghobrial 1, Abstract Multiple myeloma
Genomic complexity of multiple myeloma and its clinical implications Salomon Manier 1,, Karma Z. Salem 1,, Jihye Park 1,, Dan A. Landau 3,, Gad Getz,5 and Irene M. Ghobrial 1, Abstract Multiple myeloma
Mutations of ASXL1 and TET2 in aplastic anemia
 Published Ahead of Print on February 24, 2015, as doi:10.3324/haematol.2014.120931. Copyright 2015 Ferrata Storti Foundation. Mutations of ASXL1 and TET2 in aplastic anemia by Jinbo Huang, Meili Ge, Shihong
Published Ahead of Print on February 24, 2015, as doi:10.3324/haematol.2014.120931. Copyright 2015 Ferrata Storti Foundation. Mutations of ASXL1 and TET2 in aplastic anemia by Jinbo Huang, Meili Ge, Shihong
Supplemental Data. Shin et al. Plant Cell. (2012) /tpc YFP N
 MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
MYC YFP N PIF5 YFP C N-TIC TIC Supplemental Data. Shin et al. Plant Cell. ()..5/tpc..95 Supplemental Figure. TIC interacts with MYC in the nucleus. Bimolecular fluorescence complementation assay using
MUTATION TEST CE IVD. ctnras-braf FEATURES
 ctnras-braf CE IVD TECHNICAL SHEET IDYLLA ctnras-braf MUTATION TEST The Idylla ctnras-braf Mutation Test, performed on the Biocartis Idylla System, is an in vitro diagnostic test for the qualitative detection
ctnras-braf CE IVD TECHNICAL SHEET IDYLLA ctnras-braf MUTATION TEST The Idylla ctnras-braf Mutation Test, performed on the Biocartis Idylla System, is an in vitro diagnostic test for the qualitative detection
Mutation allele frequency threshold does not affect prognostic analysis using nextgeneration sequencing in oral squamous cell carcinoma
 Ma et al. BMC Cancer (2018) 18:758 https://doi.org/10.1186/s12885-018-4481-8 RESEARCH ARTICLE Open Access Mutation allele frequency threshold does not affect prognostic analysis using nextgeneration sequencing
Ma et al. BMC Cancer (2018) 18:758 https://doi.org/10.1186/s12885-018-4481-8 RESEARCH ARTICLE Open Access Mutation allele frequency threshold does not affect prognostic analysis using nextgeneration sequencing
Supplementary Online Content
 Supplementary Online Content Mandelker D, Zhang L, Kemel Y, et al. Mutation detection in patients with advanced cancer by universal sequencing of cancer-related genes in tumor and normal DNA vs guideline-based
Supplementary Online Content Mandelker D, Zhang L, Kemel Y, et al. Mutation detection in patients with advanced cancer by universal sequencing of cancer-related genes in tumor and normal DNA vs guideline-based
Supplementary Materials and Methods
 DD2 suppresses tumorigenicity of ovarian cancer cells by limiting cancer stem cell population Chunhua Han et al. Supplementary Materials and Methods Analysis of publicly available datasets: To analyze
DD2 suppresses tumorigenicity of ovarian cancer cells by limiting cancer stem cell population Chunhua Han et al. Supplementary Materials and Methods Analysis of publicly available datasets: To analyze
CAPP-seq. Davide Rossi, M.D., Ph.D.
 CAPP-seq Davide Rossi, M.D., Ph.D. Hematology IOSI - Oncology Institute of Southern Switzerland IOR - Institute of Oncology Research Bellinzona - Switzerland CAPP-seq capture and sequencing approach Exon
CAPP-seq Davide Rossi, M.D., Ph.D. Hematology IOSI - Oncology Institute of Southern Switzerland IOR - Institute of Oncology Research Bellinzona - Switzerland CAPP-seq capture and sequencing approach Exon
p.r623c p.p976l p.d2847fs p.t2671 p.d2847fs p.r2922w p.r2370h p.c1201y p.a868v p.s952* RING_C BP PHD Cbp HAT_KAT11
 ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
Nature Genetics: doi: /ng Supplementary Figure 1. Alternative splicing events in the 5K panel.
 Supplementary Figure 1 Alternative splicing events in the 5K panel. The majority of cryptic splicing occurred by creation of an AG or GT (Type I). While some other mutations increased the usage of a nearby
Supplementary Figure 1 Alternative splicing events in the 5K panel. The majority of cryptic splicing occurred by creation of an AG or GT (Type I). While some other mutations increased the usage of a nearby
Next Generation Sequencing in Haematological Malignancy: A European Perspective. Wolfgang Kern, Munich Leukemia Laboratory
 Next Generation Sequencing in Haematological Malignancy: A European Perspective Wolfgang Kern, Munich Leukemia Laboratory Diagnostic Methods Cytomorphology Cytogenetics Immunophenotype Histology FISH Molecular
Next Generation Sequencing in Haematological Malignancy: A European Perspective Wolfgang Kern, Munich Leukemia Laboratory Diagnostic Methods Cytomorphology Cytogenetics Immunophenotype Histology FISH Molecular
Supplementary Figure 1
 Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
NIH Public Access Author Manuscript Nat Genet. Author manuscript; available in PMC 2012 June 18.
 NIH Public Access Author Manuscript Published in final edited form as: Nat Genet. ; 43(11): 1119 1126. doi:10.1038/ng.950. Exon capture analysis of G protein-coupled receptors identifies activating mutations
NIH Public Access Author Manuscript Published in final edited form as: Nat Genet. ; 43(11): 1119 1126. doi:10.1038/ng.950. Exon capture analysis of G protein-coupled receptors identifies activating mutations
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP
 Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label)
 July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
Importance of minor TP53 mutated clones in the clinic
 Importance of minor TP53 mutated clones in the clinic Davide Rossi, M.D., Ph.D. Hematology IOSI - Oncology Institute of Southern Switzerland IOR - Institute of Oncology Reserach Bellinzona - Switzerland
Importance of minor TP53 mutated clones in the clinic Davide Rossi, M.D., Ph.D. Hematology IOSI - Oncology Institute of Southern Switzerland IOR - Institute of Oncology Reserach Bellinzona - Switzerland
Research Article Mutation in the Hair Cell Specific Gene POU4F3 Is a Common Cause for Autosomal Dominant Nonsyndromic Hearing Loss in Chinese Hans
 Neural Plasticity Volume 2016, Article ID 9890827, 6 pages http://dx.doi.org/10.1155/2016/9890827 Research Article Mutation in the Hair Cell Specific Gene POU4F3 Is a Common Cause for Autosomal Dominant
Neural Plasticity Volume 2016, Article ID 9890827, 6 pages http://dx.doi.org/10.1155/2016/9890827 Research Article Mutation in the Hair Cell Specific Gene POU4F3 Is a Common Cause for Autosomal Dominant
Exons from 137 genes were targeted for hybrid selection and massively parallel sequencing.
 SUPPLEMENTARY TABLES Supplementary Table 1. Targeted Genes ABL1 ABL2 AKT1 AKT2 AKT3 ALK APC ATM AURKA BCL2 BRAF BRCA1 BRCA2 CCND1 CCNE1 CDC73 CDH1 CDK4 CDK6 CDK8 CDKN1A CDKN2A CEBPA CHEK1 CHEK2 CREBBP
SUPPLEMENTARY TABLES Supplementary Table 1. Targeted Genes ABL1 ABL2 AKT1 AKT2 AKT3 ALK APC ATM AURKA BCL2 BRAF BRCA1 BRCA2 CCND1 CCNE1 CDC73 CDH1 CDK4 CDK6 CDK8 CDKN1A CDKN2A CEBPA CHEK1 CHEK2 CREBBP
Supplementary Figure 1 Overall study design
 Supplementary Figure 1 Overall study design We obtained 19 tumour specimens from 14 men with prostate cancer who harboured pathogenic germline BRCA2-mutations. Germline DNA was available for five patients
Supplementary Figure 1 Overall study design We obtained 19 tumour specimens from 14 men with prostate cancer who harboured pathogenic germline BRCA2-mutations. Germline DNA was available for five patients
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each
 Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
SUPPLEMENTARY DATA REFERENCES
 www.impactjournals.com/oncoscience Oncoscience, Advance Pulications 2016 SUPPLEMENTARY DATA REFERENCES 1. Busch RM, Chapin JS, Mester J, Ferguson L, Haut JS, Frazier TW, Eng C. Cognitive characteristics
www.impactjournals.com/oncoscience Oncoscience, Advance Pulications 2016 SUPPLEMENTARY DATA REFERENCES 1. Busch RM, Chapin JS, Mester J, Ferguson L, Haut JS, Frazier TW, Eng C. Cognitive characteristics
Concurrent Practical Session ACMG Classification
 Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Genetic prognostication and treatment selection in Multiple Myeloma. Kihyun Kim Department of Medicine Sungkyunkwan University Samsung Medical Center
 Genetic prognostication and treatment selection in Multiple Myeloma Kihyun Kim Department of Medicine Sungkyunkwan University Samsung Medical Center The International Congress of BMT 2018 COI disclosure
Genetic prognostication and treatment selection in Multiple Myeloma Kihyun Kim Department of Medicine Sungkyunkwan University Samsung Medical Center The International Congress of BMT 2018 COI disclosure
Fluxion Biosciences and Swift Biosciences Somatic variant detection from liquid biopsy samples using targeted NGS
 APPLICATION NOTE Fluxion Biosciences and Swift Biosciences OVERVIEW This application note describes a robust method for detecting somatic mutations from liquid biopsy samples by combining circulating tumor
APPLICATION NOTE Fluxion Biosciences and Swift Biosciences OVERVIEW This application note describes a robust method for detecting somatic mutations from liquid biopsy samples by combining circulating tumor
Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic Leukodystrophy
 International Journal of Genomics Volume 2018, Article ID 2361068, 9 pages https://doi.org/10.1155/2018/2361068 Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic
International Journal of Genomics Volume 2018, Article ID 2361068, 9 pages https://doi.org/10.1155/2018/2361068 Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Supplementary Figure 1
 Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
Supplementary Figure 1 3 3 3 1 1 Bregma -1.6mm 3 : Bregma Ref) Http://www.mbl.org/atlas165/atlas165_start.html Bregma -.18mm Supplementary Figure 1 Schematic representation of the utilized brain slice
SUPPLEMENTARY INFORMATION. Rare independent mutations in renal salt handling genes contribute to blood pressure variation
 SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
Supplementary Information. Supplementary Figure 1
 Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
Supplementary Information Supplementary Figure 1 1 Supplementary Figure 1. Functional assay of the hcas9-2a-mcherry construct (a) Gene correction of a mutant EGFP reporter cell line mediated by hcas9 or
CITATION FILE CONTENT/FORMAT
 CITATION For any resultant publications using please cite: Matthew A. Field, Vicky Cho, T. Daniel Andrews, and Chris C. Goodnow (2015). "Reliably detecting clinically important variants requires both combined
CITATION For any resultant publications using please cite: Matthew A. Field, Vicky Cho, T. Daniel Andrews, and Chris C. Goodnow (2015). "Reliably detecting clinically important variants requires both combined
WDR62 is associated with the spindle pole and mutated in human microcephaly
 WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester
 Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester dsg6@le.ac.uk CFDNA/CTDNA Circulating-free AS A LIQUID DNA BIOPSY (cfdna) Tumour Biopsy Liquid Biopsy
Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester dsg6@le.ac.uk CFDNA/CTDNA Circulating-free AS A LIQUID DNA BIOPSY (cfdna) Tumour Biopsy Liquid Biopsy
JULY 21, Genetics 101: SCN1A. Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology
 JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
Description of Supplementary Files. File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables
 Description of Supplementary Files File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables Supplementary Figure 1: (A), HCT116 IDH1-WT and IDH1-R132H cells were
Description of Supplementary Files File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables Supplementary Figure 1: (A), HCT116 IDH1-WT and IDH1-R132H cells were
Prevalence of ATM sequence variants in Northern plains American Indian cancer patients
 ORIGINAL RESEARCH ARTICLE published: 30 December 2013 doi: 10.3389/fonc.2013.00318 Prevalence of ATM sequence variants in Northern plains American Indian cancer Daniel G. Petereit 1,2 *, L. Jennifer Hahn
ORIGINAL RESEARCH ARTICLE published: 30 December 2013 doi: 10.3389/fonc.2013.00318 Prevalence of ATM sequence variants in Northern plains American Indian cancer Daniel G. Petereit 1,2 *, L. Jennifer Hahn
Genomic Medicine: What every pathologist needs to know
 Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most
 Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
Supplementary Figure 1 MicroRNA expression in human synovial fibroblasts from different locations. MicroRNA, which were identified by RNAseq as most differentially expressed between human synovial fibroblasts
Phenylalanine hydroxylase deficiency in Mexico: genotype phenotype correlations, BH 4 responsiveness and evidence of a founder effect
 Clin Genet 2015: 88: 62 67 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12444 Phenylalanine hydroxylase
Clin Genet 2015: 88: 62 67 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12444 Phenylalanine hydroxylase
