Supplementary Figure 1
|
|
|
- Violet Ball
- 5 years ago
- Views:
Transcription
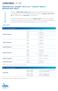
1 Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0 HR Efficiency * * Surviving fraction SF50 (M) HEC1A PTEN +/+ >1.0E-05 HEC1A PTEN -/- -/ E-05 HEC1A PTEN -/- -/ E HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/ KU (M) KU (M) 10-5
2 Supplementary Figure 2 "H2AX Merged HCT116 PTEN+/+ Rad51 - IR HCT116 PTEN-/- Nuclei - IR + IR + IR
3 Supplementary Figure 3 Surviving fraction DU145 MCF-7 HeLa SUM159 BT474 MDA-MB-231 HCC70 PC3 RPMI-7951 SW1783 EC50 SF50 (M) 7.72E E E E A172 UM-UC3 2.94E E-08 MDA-MB E COLO E-09 KU (M)
4 Supplementary Figure 4 EMPTY VECTOR RAD51 cdna RAD51! Tubulin
5 Supplementary Table 1 Cell Line PTEN PIK3CA TP53 BRAF KRAS MLH1 MSH6 CDKN2A A172 * COLO 829 * * * HCC70 * * MDA-MB-468 * * PC3 * * RPMI-7951 * * * * SW1783 * * UM-UC3 * * * * BT474 * * HEC1A * * * HCT116 * * * * MCF7 * * SUM-159 * MDA-MB-231 * * * * DU145 * * * HeLa Cell Line Mutation PTEN PIK3CA TP53 BRAF KRAS MLH1 MSH6 CDKN2A A172 AA p.r55fs*1 CDS c.165_1212del1048 glioma Zygosity Homozygous COLO 829 AA p.? p.v600e p.a68fs*51 CDS c.493_634del142 c.1799t>a c.203_204delcg melanoma Zygosity Homozygous Heterozygous Homozygous HCC70 AA p.f90fs*9 p.r248q CDS c.270delt c.743g>a breast Zygosity Homozygous Homozygous MDA-MB-468 AA p.? p.r273h CDS c.253+1g>t c.818g>a breast Zygosity Homozygous Homozygous PC3 AA p.r55fs*1 p.k139fs*31 CDS c.165_1212del1048 c.414delc prostate Zygosity Homozygous Homozygous RPMI-7951 AA p.? p.s166* p.v600e p.l16r CDS c.1_79del79 c.497c>a c.1799t>a c.47t>g melanoma Zygosity Homozygous Homozygous Heterozygous Homozygous SW1783 AA p.r233* p.r273h CDS c.697c>t c.818g>a glioma Zygosity Homozygous Heterozygous UM-UC3 AA p.m1_*404del p.f113c p.g12c p.m1_*157del CDS c.1_1212del1212 c.338t>g c.34g>t c.1_471del471 bladder Zygosity Homozygous Homozygous Homozygous Homozygous BT-474 AA p.k111n p.e285k CDS c.333g>c c.853g>a breast Zygosity Heterozygous Homozygous HEC1A AA p.g1049r p.r248q p.g12d CDS c.3145g>c c.743g>a c.35g>a endometrium Zygosity Heterozygous Homozygous Homozygous HCT116 AA p.h1047r p.g13d p.s252* p.r24fs*20 CDS c.3140a>g c.38g>a c.755c>a c.68_69insg colorectal Zygosity Heterozygous Heterozygous Homozygous Homozygous MCF7 AA p.e545k p.m1_*157del CDS c.1633g>a c.1_471del471 breast Zygosity Heterozygous Homozygous SUM-159 AA p.h1047l CDS c.3140a>t breast Zygosity Heterozygous MDA-MB-231 AA p.r280k p.g464v p.g13d p.m1_*157del CDS c.839g>a c.1391g>t c.38g>a c.1_471del471 breast Zygosity Homozygous Heterozygous Heterozygous Homozygous DU145 AA p.p223l, p.v274f p.? p.d84y CDS c.668c>t, c.820g>t c.117-2a>t c.250g>t prostate Zygosity Heterozygous Homozygous Homozygous HeLa AA CDS cervix Zygosity
6 Sample Name COSMIC Sample ID Amino Acid Nucleotide Primary Tissue Histology HCC p.f90fs*9 c.270delt breast carcinoma 5839 cell-line S p.k60fs*9 c.180_181ins? breast carcinoma NS MDA- MB p.l70fs*7 c.208_251del44 breast carcinoma NS p.a39fs*5 c.116_117insg central_nervous_system glioma S p.a79fs*20 c.237delc central_nervous_system glioma primary SKMG p.c71fs*6 c.212_255del44 central_nervous_system glioma NS S p.d24fs*20 c.70_71insa central_nervous_system glioma primary S p.d24fs*20 c.70_71insa central_nervous_system glioma primary S p.e43fs*11 c.127delg central_nervous_system glioma NS S p.f56fs*2 c.165_492del328 central_nervous_system glioma primary p.k60* c.178a>t central_nervous_system glioma S p.k66fs*38 c.197_198ins14 central_nervous_system glioma primary S p.k6fs*4 c.17_18delaa central_nervous_system glioma primary S p.l70fs*7 c.210_253del44 central_nervous_system glioma primary S p.l70fs*7 c.210_253del44 central_nervous_system glioma primary SF p.m1_*404del c.1_1212del1212 central_nervous_system glioma cell-line H p.m1_*404del c.1_1212del1212 central_nervous_system glioma cell-line D- 245MG p.m1_*404del c.1_1212del1212 central_nervous_system glioma cell-line D- 336MG p.m1_*404del c.1_1212del1212 central_nervous_system glioma cell-line S p.n12fs*6 c.35_53del19 central_nervous_system glioma primary S p.r14fs*26 c.42_52del11 central_nervous_system glioma primary S p.r15fs*9 c.45_46insga central_nervous_system glioma primary A p.r55fs*1 c.165_1212del1048 central_nervous_system glioma cell-line SW p.r55fs*1 c.165_1212del1048 central_nervous_system glioma cell-line U-87- MG p.v54fs*29 c.160_208del49 central_nervous_system glioma NS S p.y16fs*1 c.47_48insa central_nervous_system glioma NS S p.y46* c.138c>g central_nervous_system glioma primary Pubmed ID Mutation ID Sample Source
7 S p.y65* c.195c>a central_nervous_system glioma primary S p.y65* c.195c>g central_nervous_system glioma primary S p.y76fs*1 c.226_227delta central_nervous_system glioma primary S p.y76fs*1 c.227_228delat central_nervous_system glioma primary S p.y76fs*1 c.227_228delat central_nervous_system glioma primary S p.y88fs*3 c.263_264delat central_nervous_system glioma primary LS p.y88fs*3 c.263_264delat central_nervous_system glioma NS S p.a3fs*14 c.9_30del22 endometrium carcinoma primary S p.c71fs*6 c.213_256del44 endometrium carcinoma primary S p.d24fs*19 c.71_72insc endometrium carcinoma primary S p.d24fs*19 c.71_72insc endometrium carcinoma primary S p.e18fs*5 c.54_57delggat endometrium carcinoma primary p.e73fs*25 c.219_222delaaga endometrium carcinoma S p.e73fs*4 c.219_220delaa endometrium carcinoma primary p.e7fs*3 c.19_20delga endometrium carcinoma NOS p.f56c c.167t>g endometrium carcinoma S p.f90fs*9 c.270delt endometrium carcinoma primary S p.h64fs*9 c.190_191delca endometrium carcinoma primary S p.i32fs*22 c.96delt endometrium carcinoma primary S p.i32fs*22 c.96delt endometrium carcinoma primary S p.i33fs*20 c.97_100delattg endometrium carcinoma primary S p.i33fs*21 c.97dela endometrium carcinoma primary S p.i33fs*21 c.97dela endometrium carcinoma metastasis E p.k6fs*4 c.17_18delaa endometrium carcinoma S p.k6fs*4 c.17_18delaa endometrium carcinoma primary p.k6fs*4 c.17_18delaa endometrium carcinoma S p.l25fs*28 c.75_78delgacc endometrium carcinoma primary p.l57fs*42 c.166_166delt endometrium carcinoma NOS E p.l57fs*6 c.170_171inst endometrium carcinoma
8 S p.l57fs*6 c.170_171inst endometrium carcinoma primary p.l57fs*6 c.170_171inst endometrium carcinoma fixed - NOS S p.n48fs*6 c.143dela endometrium carcinoma primary S p.n63fs*10 c.187_188delaa endometrium carcinoma primary p.n63fs*10 c.187_188delaa endometrium carcinoma p.n63fs*10 c.187_188delaa endometrium carcinoma S p.n63fs*11 c.188_189insa endometrium carcinoma primary S p.n63fs*36 c.187dela endometrium carcinoma primary S p.n63fs*36 c.188dela endometrium carcinoma primary S p.n63fs*36 c.188dela endometrium carcinoma primary S p.n63fs*36 c.188dela endometrium hyperplasia primary S p.n69fs*30 c.206dela endometrium carcinoma primary S p.p30fs*24 c.89delc endometrium carcinoma primary S p.r14fs*10 c.40dela endometrium carcinoma primary S p.r14fs*29 c.41_42insg endometrium carcinoma primary S p.r15fs*28 c.45_46inst endometrium carcinoma primary S p.r55fs*2 c.165_181del17 endometrium carcinoma primary S p.t26fs*28 c.78delc endometrium carcinoma primary S p.v45fs*10 c.134_135insca endometrium carcinoma primary S p.y16fs*21 c.46_65del20 endometrium carcinoma primary S p.y16fs*28 c.46_47inst endometrium carcinoma primary S p.y27fs*1 c.80_90del11 endometrium carcinoma primary S p.y27fs*16 c.80_81delat endometrium carcinoma primary p.y29fs*25 c.83_83delt endometrium carcinoma S p.y68fs*5 c.202_203delta endometrium carcinoma primary S p.y76fs*1 c.227_228delat endometrium carcinoma primary S p.y76fs*1 c.227_228delat endometrium carcinoma primary OPM p.m1_*404del c.1_1212del1212 haematopoietic_and_lymphoid_tissue haematopoietic_neoplasm cell-line S p.m1fs*7 c.1_8delatgacagc haematopoietic_and_lymphoid_tissue lymphoid_neoplasm primary KE p.y27fs*1 c.80_1212del1133 haematopoietic_and_lymphoid_tissue haematopoietic_neoplasm cell-line p.e7fs*3 c.21_22delga large_intestine carcinoma
9 p.f90fs*9 c.270delt large_intestine carcinoma p.l57fs*42 c.170delt large_intestine carcinoma p.l57fs*6 c.170_171inst large_intestine carcinoma S p.c83fs*9 c.247_248inst liver carcinoma primary H p.? c.1-?_492+?del lung carcinoma NS NCI- H p.? c.1-?_492+?del lung carcinoma NS SBC p.m1_*404del c.1_1212del1212 lung carcinoma cell-line NCI- H p.m1_*404del c.1_1212del1212 lung carcinoma cell-line N p.m1_?del fs c.1-?_79+?del lung carcinoma NS N p.m1_?del fs c.1-?_79+?del lung carcinoma NS N p.m1_?del fs c.1-?_79+?del lung carcinoma NS NCI- H p.r55fs*1 c.165_1212del1048 lung carcinoma cell-line LU-134- A p.y27fs*1 c.80_1212del1133 lung carcinoma cell-line S p.l57fs*42 c.170delt meninges meningioma primary p.c71fs*28 c.213_213delt ovary carcinoma fresh - NOS p.f81fs*18 c.241_242tt>g ovary carcinoma NOS RTSG p.k6fs*4 c.17_18delaa ovary carcinoma 4929 cell-line S p.y68fs*5 c.202_203delta ovary carcinoma primary S p.g36fs*18 c.107delg prostate carcinoma primary S p.g44fs*11 c.131_139gcgtataca>acagaaagaca prostate carcinoma primary LNCaP- Clone- FGC p.k6fs*4 c.17_18delaa prostate carcinoma 4929 cell-line LNCaP p.k6fs*4 c.16_17delaa prostate carcinoma NS LNCaP p.k6fs*4 c.16_17delaa prostate carcinoma NS LNCaP p.k6fs*4 c.17_18delaa prostate carcinoma NS S p.q87* c.259c>t prostate carcinoma NS PC p.r55fs*1 c.165_1212del1048 prostate carcinoma cell-line
10 S p.y76fs*1 c.226_227delta prostate carcinoma NS FM p.? c.1-?_492+?del skin malignant_melanoma NS S p.k6fs*4 c.17_18delaa skin malignant_melanoma metastasis G-mel p.m1_?del fs c.1-?_79+?del skin malignant_melanoma NS SW p.m1_*404del c.1_1212del1212 soft_tissue liposarcoma cell-line UM-UC p.m1_*404del c.1_1212del1212 urinary_tract carcinoma cell-line
Whole Exome Sequenced Characteristics
 Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Supplementary Table 2. Identified causative mutations and/or mutation candidates.
 Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label)
 July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM
 Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
UW359 Ovary 3c 3 Serous Recurrent 68 BRCA1 816delGT BRCA1 del exon 1-2. UW417 Ovary 3c 3 Serous Primary 38 BRCA1 1675delA
 Supplementary Table 1. Cases with deleterious germline mutations, somatic HR mutations, and somatic PTEN mutations. ID Site Stage Grade Histology Tumor Age Germline mutation(s) a Somatic HR mutation(s)
Supplementary Table 1. Cases with deleterious germline mutations, somatic HR mutations, and somatic PTEN mutations. ID Site Stage Grade Histology Tumor Age Germline mutation(s) a Somatic HR mutation(s)
Genetic Alteration Panels
 TM Genetic Alteration Panels GENETIC ALTERATION PANELS Table of Contents AKT Genetic Alteration Panel (ATCC No. TCP-1029 )...1 BRAF Genetic Alteration Panel (ATCC No. TCP-1032 )...5 EGFR Genetic Alteration
TM Genetic Alteration Panels GENETIC ALTERATION PANELS Table of Contents AKT Genetic Alteration Panel (ATCC No. TCP-1029 )...1 BRAF Genetic Alteration Panel (ATCC No. TCP-1032 )...5 EGFR Genetic Alteration
Supplementary Table 1. PIK3CA mutation in colorectal cancer
 Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Intron p Hybrid with p.leu245val p. Gly336Cys. p.leu62phe p. Ala137Val p. Asn152Thr hybrid with p.leu245val p.
 RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
Supplementary Information
 Supplementary Information JAK1 truncating mutations in gynecologic cancer define new role of cancerassociated protein tyrosine kinase aberrations Yuan Ren, Yonghong Zhang, Richard Z. Liu, David A. Fenstermacher,
Supplementary Information JAK1 truncating mutations in gynecologic cancer define new role of cancerassociated protein tyrosine kinase aberrations Yuan Ren, Yonghong Zhang, Richard Z. Liu, David A. Fenstermacher,
Supplementary Online Content
 Supplementary Online Content Pearlman R, Frankel WL, Swanson B, et al. Prevalence and spectrum of germline cancer susceptibility gene mutations among patients with early-onset colorectal cancer. JAMA Oncol.
Supplementary Online Content Pearlman R, Frankel WL, Swanson B, et al. Prevalence and spectrum of germline cancer susceptibility gene mutations among patients with early-onset colorectal cancer. JAMA Oncol.
Familial exudative vitreoretinopathy (FEVR) is an inheritable
 Genetics Spectrum of Variants in 389 Chinese Probands With Familial Exudative Vitreoretinopathy Jia-Kai Li, 1 Yian Li, 1 Xiang Zhang, 1 Chun-Li Chen, 2 Yu-Qing Rao, 1 Ping Fei, 1 Qi Zhang, 1 Peiquan Zhao,
Genetics Spectrum of Variants in 389 Chinese Probands With Familial Exudative Vitreoretinopathy Jia-Kai Li, 1 Yian Li, 1 Xiang Zhang, 1 Chun-Li Chen, 2 Yu-Qing Rao, 1 Ping Fei, 1 Qi Zhang, 1 Peiquan Zhao,
Clinical presentation. Duration (Month)
 Supplementary information, Table S1A: Clinical information for 125 PAs. No. Subtype Gender (M/F) Age at diagnosis (year) Clinical presentation Duration (Month) Maximal adenoma diameter(cm) Tumor invasion*
Supplementary information, Table S1A: Clinical information for 125 PAs. No. Subtype Gender (M/F) Age at diagnosis (year) Clinical presentation Duration (Month) Maximal adenoma diameter(cm) Tumor invasion*
Spectrum and Classification of ATP7B Variants in a Large Cohort of Chinese Patients with Wilson s Disease Guides Genetic Diagnosis
 Ivyspring International Publisher 638 Theranostics Research Paper 2016; 6(5): 638-649. doi: 10.7150/thno.14596 Spectrum and Classification of ATP7B Variants in a Large Cohort of Chinese Patients with Wilson
Ivyspring International Publisher 638 Theranostics Research Paper 2016; 6(5): 638-649. doi: 10.7150/thno.14596 Spectrum and Classification of ATP7B Variants in a Large Cohort of Chinese Patients with Wilson
Nature Genetics: doi: /ng Supplementary Figure 1. TCGA data set on HNSCCs reanalyzed in this study.
 Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
The Personalized Cancer Medicine Initiative at Vanderbilt
 The Personalized Cancer Medicine Initiative at Vanderbilt October 26, 2009 William Pao, MD, PhD Assistant Director, Personalized Cancer Medicine Vanderbilt-Ingram Cancer Center Cancer in the United States,
The Personalized Cancer Medicine Initiative at Vanderbilt October 26, 2009 William Pao, MD, PhD Assistant Director, Personalized Cancer Medicine Vanderbilt-Ingram Cancer Center Cancer in the United States,
Species Tumor Type Comment for in vivo work Lead Time for in vivo studies [weeks] MB-49-luc-2 Mouse urinary bladder carcinoma C57BL/6 2
![Species Tumor Type Comment for in vivo work Lead Time for in vivo studies [weeks] MB-49-luc-2 Mouse urinary bladder carcinoma C57BL/6 2 Species Tumor Type Comment for in vivo work Lead Time for in vivo studies [weeks] MB-49-luc-2 Mouse urinary bladder carcinoma C57BL/6 2](/thumbs/88/116309357.jpg) America, Hershey, PA Australia, Melbourne, VIC Europe, Munich info@vivopharm.com www.vivopharm.com Tissue Bladder Species Tumor Type Comment for in vivo work Lead Time for in vivo studies [weeks] MB-49-luc-
America, Hershey, PA Australia, Melbourne, VIC Europe, Munich info@vivopharm.com www.vivopharm.com Tissue Bladder Species Tumor Type Comment for in vivo work Lead Time for in vivo studies [weeks] MB-49-luc-
Recent advances in the molecular pathology of lung cancer: role of EGFR and KRAS mutation testing in treatment selection
 ASIP 2009 USCAP Companion Meeting Molecular Pathology for the Practicing Pathologist Recent advances in the molecular pathology of lung cancer: role of EGFR and KRAS mutation testing in treatment selection
ASIP 2009 USCAP Companion Meeting Molecular Pathology for the Practicing Pathologist Recent advances in the molecular pathology of lung cancer: role of EGFR and KRAS mutation testing in treatment selection
ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS
 ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS 1 This medicinal product is subject to additional monitoring. This will allow quick identification of new safety information. Healthcare professionals are asked
ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS 1 This medicinal product is subject to additional monitoring. This will allow quick identification of new safety information. Healthcare professionals are asked
ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS
 ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS 1 This medicinal product is subject to additional monitoring. This will allow quick identification of new safety information. Healthcare professionals are asked
ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS 1 This medicinal product is subject to additional monitoring. This will allow quick identification of new safety information. Healthcare professionals are asked
PRODUCT MONOGRAPH INCLUDING PATIENT MEDICATION INFORMATION. Migalastat Capsules. 123 mg migalastat (as migalastat hydrochloride)
 PRODUCT MONOGRAPH INCLUDING PATIENT MEDICATION INFORMATION Pr GALAFOLD TM Migalastat Capsules 123 mg migalastat (as migalastat hydrochloride) Various alimentary tract and metabolism products Amicus Therapeutics
PRODUCT MONOGRAPH INCLUDING PATIENT MEDICATION INFORMATION Pr GALAFOLD TM Migalastat Capsules 123 mg migalastat (as migalastat hydrochloride) Various alimentary tract and metabolism products Amicus Therapeutics
Belgian study into von Willebrand Disease (B-Will): First results
 Belgian study into von Willebrand Disease (B-Will): First results Inge Vangenechten VWD Research Unit, Antwerp University Hospital Supported by CSL Behring Chair in von Willebrand Disease, Antwerp University
Belgian study into von Willebrand Disease (B-Will): First results Inge Vangenechten VWD Research Unit, Antwerp University Hospital Supported by CSL Behring Chair in von Willebrand Disease, Antwerp University
Figure S1 Time-dependent down-modulation of HER3 by EZN No Treatment. EZN-3920, 2 μm. Time, h
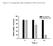 Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Human cell line list Production of Exosome Standards - Cell Lysates
 Human cell line list 2016 - Production of Exosome Standards - Cell Lysates 380 peripheral blood leukemia, pre-b cell 1301 blood leukemia, acute lymphoblastic, T cell 5637 bladder carcinoma 8305C thyroid
Human cell line list 2016 - Production of Exosome Standards - Cell Lysates 380 peripheral blood leukemia, pre-b cell 1301 blood leukemia, acute lymphoblastic, T cell 5637 bladder carcinoma 8305C thyroid
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/12/e1601756/dc1 Supplementary Materials for Syrosingopine sensitizes cancer cells to killing by metformin Don Benjamin, Marco Colombi, Sravanth K. Hindupur, Charles
advances.sciencemag.org/cgi/content/full/2/12/e1601756/dc1 Supplementary Materials for Syrosingopine sensitizes cancer cells to killing by metformin Don Benjamin, Marco Colombi, Sravanth K. Hindupur, Charles
Supplementary Data Supplementary Table S1. IC 50 values of ipatasertib on cell viability in cell lines with
 Supplementary Data Supplementary Table S1. IC 50 values of ipatasertib on cell viability in cell lines with or without alterations in PTEN or PIK3CA. Cell Line Tissue Type Ipatasertib IC 50 (μmol/l) a
Supplementary Data Supplementary Table S1. IC 50 values of ipatasertib on cell viability in cell lines with or without alterations in PTEN or PIK3CA. Cell Line Tissue Type Ipatasertib IC 50 (μmol/l) a
Next generation sequencing based detection of EGFR, KRAS, BRAF, NRAS, PIK3CA, Her 2 and TP53 mutations in patients with non small cell lung cancer
 MOLECULAR MEDICINE REPORTS 18: 2191-2197, 2018 Next generation sequencing based detection of EGFR, KRAS, BRAF, NRAS, PIK3CA, Her 2 and TP53 mutations in patients with non small cell lung cancer CHANGWEN
MOLECULAR MEDICINE REPORTS 18: 2191-2197, 2018 Next generation sequencing based detection of EGFR, KRAS, BRAF, NRAS, PIK3CA, Her 2 and TP53 mutations in patients with non small cell lung cancer CHANGWEN
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/270/270ra6/dc1 Supplementary Materials for Integrated allelic, transcriptional, and phenomic dissection of the cardiac effects of titin truncations
www.sciencetranslationalmedicine.org/cgi/content/full/7/270/270ra6/dc1 Supplementary Materials for Integrated allelic, transcriptional, and phenomic dissection of the cardiac effects of titin truncations
CHAPTER IV RESULTS Microcephaly General description
 47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
America, Hershey, PA Australia, Melbourne, VIC Europe, Munich
 America, Hershey, PA Australia, Melbourne, VIC Europe, Munich info@vivopharm.com www.vivopharm.com Bladder MB-9-luc 2 Mouse urinary bladder carcinoma yes yes yes C57BL/6 2 Bone UMR-106 Rat osteosarcoma
America, Hershey, PA Australia, Melbourne, VIC Europe, Munich info@vivopharm.com www.vivopharm.com Bladder MB-9-luc 2 Mouse urinary bladder carcinoma yes yes yes C57BL/6 2 Bone UMR-106 Rat osteosarcoma
MSI positive MSI negative
 Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Belgian-Czech cooperation in the Brno-VWD study
 Inge Vangenechten I Vangenechten, P Smejkal, O Zapletal, F Bouddount, J Zavrelova, J Blatny, M Penka, JJ Michiels, A Gadisseur. Aim of the study Characterisation of VWD in South Moravia (Czech Republic)
Inge Vangenechten I Vangenechten, P Smejkal, O Zapletal, F Bouddount, J Zavrelova, J Blatny, M Penka, JJ Michiels, A Gadisseur. Aim of the study Characterisation of VWD in South Moravia (Czech Republic)
Names for ABO (ISBT 001) Blood Group Alleles
 Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
Exons from 137 genes were targeted for hybrid selection and massively parallel sequencing.
 SUPPLEMENTARY TABLES Supplementary Table 1. Targeted Genes ABL1 ABL2 AKT1 AKT2 AKT3 ALK APC ATM AURKA BCL2 BRAF BRCA1 BRCA2 CCND1 CCNE1 CDC73 CDH1 CDK4 CDK6 CDK8 CDKN1A CDKN2A CEBPA CHEK1 CHEK2 CREBBP
SUPPLEMENTARY TABLES Supplementary Table 1. Targeted Genes ABL1 ABL2 AKT1 AKT2 AKT3 ALK APC ATM AURKA BCL2 BRAF BRCA1 BRCA2 CCND1 CCNE1 CDC73 CDH1 CDK4 CDK6 CDK8 CDKN1A CDKN2A CEBPA CHEK1 CHEK2 CREBBP
Nature Genetics: doi: /ng Supplementary Figure 1. Alternative splicing events in the 5K panel.
 Supplementary Figure 1 Alternative splicing events in the 5K panel. The majority of cryptic splicing occurred by creation of an AG or GT (Type I). While some other mutations increased the usage of a nearby
Supplementary Figure 1 Alternative splicing events in the 5K panel. The majority of cryptic splicing occurred by creation of an AG or GT (Type I). While some other mutations increased the usage of a nearby
Use of panel tests in place of single gene tests in the cancer genetics clinic
 Clin Genet 2015: 88: 278 282 Printed in Singapore. All rights reserved CLINICAL GENETICS doi: 10.1111/cge.12488 Short Report se of panel tests in place of single gene tests in the cancer genetics clinic
Clin Genet 2015: 88: 278 282 Printed in Singapore. All rights reserved CLINICAL GENETICS doi: 10.1111/cge.12488 Short Report se of panel tests in place of single gene tests in the cancer genetics clinic
How has Molecular Diagnostics changed my practice? Pulmonary Pathology
 How has Molecular Diagnostics changed my practice? Pulmonary Pathology Lynette M. Sholl, M.D. Associate Pathologist Brigham and Women s Hospital Associate Director, Center for Advanced Molecular Diagnostics
How has Molecular Diagnostics changed my practice? Pulmonary Pathology Lynette M. Sholl, M.D. Associate Pathologist Brigham and Women s Hospital Associate Director, Center for Advanced Molecular Diagnostics
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future?
 Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future? Tony Marinakii Purine Research Laboratory Biochemical Sciences Whole exome sequencing No doubt
Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future? Tony Marinakii Purine Research Laboratory Biochemical Sciences Whole exome sequencing No doubt
Supplementary information to:
 Supplementary information to: Digital Sorting of Pure Cell Populations Enables Unambiguous Genetic Analysis of Heterogeneous Formalin-Fixed Paraffin Embedded Tumors by Next Generation Sequencing Authors
Supplementary information to: Digital Sorting of Pure Cell Populations Enables Unambiguous Genetic Analysis of Heterogeneous Formalin-Fixed Paraffin Embedded Tumors by Next Generation Sequencing Authors
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP
 Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each
 Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
Supplementary Online Content
 Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Fluxion Biosciences and Swift Biosciences Somatic variant detection from liquid biopsy samples using targeted NGS
 APPLICATION NOTE Fluxion Biosciences and Swift Biosciences OVERVIEW This application note describes a robust method for detecting somatic mutations from liquid biopsy samples by combining circulating tumor
APPLICATION NOTE Fluxion Biosciences and Swift Biosciences OVERVIEW This application note describes a robust method for detecting somatic mutations from liquid biopsy samples by combining circulating tumor
Research Article Towards a Next-Generation Sequencing Diagnostic Service for Tumour Genotyping: A Comparison of Panels and Platforms
 BioMed Volume 2015, Article ID 478017, 6 pages http://dx.doi.org/10.1155/2015/478017 Research Article Towards a Next-Generation Sequencing Diagnostic Service for Tumour Genotyping: A Comparison of Panels
BioMed Volume 2015, Article ID 478017, 6 pages http://dx.doi.org/10.1155/2015/478017 Research Article Towards a Next-Generation Sequencing Diagnostic Service for Tumour Genotyping: A Comparison of Panels
Summary of Results CARRIER STATUS DNA INSIGHT. Result: Carrier, Heterozygote. Protected Health Information. Propionic Acidemia
 Test Results Reviewed & Approved by: Laboratory Director, Nilesh Dharajiya,.D. SAPLE REPORT CARRIER STATUS DNA INSIGHT PERSONAL DETAILS DOB Jan 1, 19XX ETHNICITY Asian ORDERING HEALTHCARE PROFESSIONAL
Test Results Reviewed & Approved by: Laboratory Director, Nilesh Dharajiya,.D. SAPLE REPORT CARRIER STATUS DNA INSIGHT PERSONAL DETAILS DOB Jan 1, 19XX ETHNICITY Asian ORDERING HEALTHCARE PROFESSIONAL
(a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable
 Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Supplementary Figure 1. Frameshift (FS) mutation in UVRAG. (a) Schematic diagram of the FS mutation of UVRAG in exon 8 containing the highly instable A 10 DNA repeat, generating a premature stop codon
Frequency(%) KRAS G12 KRAS G13 KRAS A146 KRAS Q61 KRAS K117N PIK3CA H1047 PIK3CA E545 PIK3CA E542K PIK3CA Q546. EGFR exon19 NFS-indel EGFR L858R
 Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Genomic sequencing of meningiomas identifies oncogenic SMO and AKT1 mutations
 Genomic sequencing of meningiomas identifies oncogenic SMO and AKT mutations Priscilla K. Brastianos*; Peleg M. Horowitz*; Sandro Santagata; Robert T. Jones; Aaron McKenna; Gad Getz; Keith L. Ligon; Emanuele
Genomic sequencing of meningiomas identifies oncogenic SMO and AKT mutations Priscilla K. Brastianos*; Peleg M. Horowitz*; Sandro Santagata; Robert T. Jones; Aaron McKenna; Gad Getz; Keith L. Ligon; Emanuele
LMM / emerge III Network Reference Sequences October 2016 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene
 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
 Int. J. Cancer: 116, 692 702 (2005) ' 2005 Wiley-Liss, Inc. Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
Int. J. Cancer: 116, 692 702 (2005) ' 2005 Wiley-Liss, Inc. Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
DNA Analysis in Glycogen storage disease
 DNA Analysis in Glycogen storage disease Nick Beauchamp PhD Sheffield, Sheffield Children s NHS Foundation Trust 14th October 2010 Glycogen Synthase Type 0 Glycogen Synthesis and Breakdown Type IV UDPGlucose
DNA Analysis in Glycogen storage disease Nick Beauchamp PhD Sheffield, Sheffield Children s NHS Foundation Trust 14th October 2010 Glycogen Synthase Type 0 Glycogen Synthesis and Breakdown Type IV UDPGlucose
Laboratory, Division of Medical Sciences, National Cancer Centre, Singapore
 Supplementary Information Exome sequencing of liver fluke-associated cholangiocarcinoma Choon Kiat Ong 1,2*, Chutima Subimerb 1,2,3*, Chawalit Pairojkul 3, Sopit Wongkham 3, Ioana Cutcutache 4, Willie
Supplementary Information Exome sequencing of liver fluke-associated cholangiocarcinoma Choon Kiat Ong 1,2*, Chutima Subimerb 1,2,3*, Chawalit Pairojkul 3, Sopit Wongkham 3, Ioana Cutcutache 4, Willie
LavaCell. Fluorescent Cell Stain. (version A) Catalog No
 LavaCell Fluorescent Cell Stain (version A) Catalog No. 15004 Active Motif North America 1914 Palomar Oaks Way, Suite 150 Carlsbad, California 92008, USA Toll free: 877 222 9543 Telephone: 760 431 1263
LavaCell Fluorescent Cell Stain (version A) Catalog No. 15004 Active Motif North America 1914 Palomar Oaks Way, Suite 150 Carlsbad, California 92008, USA Toll free: 877 222 9543 Telephone: 760 431 1263
Names for H (ISBT 018) Blood Group Alleles
 Names for H (ISBT 018) Blood Group Alleles General description: The H blood group system consists of one antigen, H, that is carried on glycolipids and glycoproteins on the RBC membrane, where it is synthesised
Names for H (ISBT 018) Blood Group Alleles General description: The H blood group system consists of one antigen, H, that is carried on glycolipids and glycoproteins on the RBC membrane, where it is synthesised
Converting Novel Therapeutic Models into Early Phase Clinical Trials
 Converting Novel Therapeutic Models into Early Phase Clinical Trials James H. Doroshow, M.D. Deputy Director for Clinical and Translational Research National Cancer Institute, NIH IOM National Cancer Policy
Converting Novel Therapeutic Models into Early Phase Clinical Trials James H. Doroshow, M.D. Deputy Director for Clinical and Translational Research National Cancer Institute, NIH IOM National Cancer Policy
A novel tumor suppressor role for the Notch pathway in bladder cancer
 A novel tumor suppressor role for the Notch pathway in bladder cancer Theodoros Rampias, Paraskevi Vgenopoulou, Margaritis Avgeris, Alexander Polyzos, Konstantinos Stravodimos, Christos Valavanis, Andreas
A novel tumor suppressor role for the Notch pathway in bladder cancer Theodoros Rampias, Paraskevi Vgenopoulou, Margaritis Avgeris, Alexander Polyzos, Konstantinos Stravodimos, Christos Valavanis, Andreas
Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH
 Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH Index patients (n=89) FH children Cascade screening (n=82) cdna Mutation Protein 3 0 c.10580g>a p.arg3527gln
Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH Index patients (n=89) FH children Cascade screening (n=82) cdna Mutation Protein 3 0 c.10580g>a p.arg3527gln
Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia
 Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia Erpelinck-Verschueren, Veronika Rockova, Mathijs Sanders,
Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia Erpelinck-Verschueren, Veronika Rockova, Mathijs Sanders,
(Case 5 A ) c g>a Exon 24 splice acceptor Splicing defect - Splicing defect
 Supplementary table S1. Pathogenic potential of identified DYNC2H1 mutations Family Nucleotide Change Mutation GERP PhyloP SIFT PolyPhen-2 Mutation Taster JATD-1 c. 90443A>G p.d3015g 6.17 5.3 Tolerated
Supplementary table S1. Pathogenic potential of identified DYNC2H1 mutations Family Nucleotide Change Mutation GERP PhyloP SIFT PolyPhen-2 Mutation Taster JATD-1 c. 90443A>G p.d3015g 6.17 5.3 Tolerated
A novel compound heterozygous mutation in SLC5A2 contributes to familial renal glucosuria in a Chinese family, and a review of the relevant literature
 MOLECULAR MEDICINE REPORTS A novel compound heterozygous mutation in SLC5A2 contributes to familial renal glucosuria in a Chinese family, and a review of the relevant literature SHENTANG LI 1, YEYI YANG
MOLECULAR MEDICINE REPORTS A novel compound heterozygous mutation in SLC5A2 contributes to familial renal glucosuria in a Chinese family, and a review of the relevant literature SHENTANG LI 1, YEYI YANG
Description of Supplementary Files. File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables
 Description of Supplementary Files File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables Supplementary Figure 1: (A), HCT116 IDH1-WT and IDH1-R132H cells were
Description of Supplementary Files File Name: Supplementary Information Description: Supplementary Figures and Supplementary Tables Supplementary Figure 1: (A), HCT116 IDH1-WT and IDH1-R132H cells were
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Round Table: Tissue Biopsy versus Liquid Biopsy. César A. Rodríguez Hospital Universitario de Salamanca-IBSAL
 Round Table: Tissue Biopsy versus Liquid Biopsy César A. Rodríguez Hospital Universitario de Salamanca-IBSAL Introduction Classic Advantages of liquid biopsy collection over standard biopsy Standard biopsy
Round Table: Tissue Biopsy versus Liquid Biopsy César A. Rodríguez Hospital Universitario de Salamanca-IBSAL Introduction Classic Advantages of liquid biopsy collection over standard biopsy Standard biopsy
Supplemental Data. Prolonged Rapamycin Treatment Inhibits mtorc2 Assembly and Akt/PKB. Supplemental Experimental Procedures
 Supplemental Data Prolonged Rapamycin Treatment Inhibits mtorc2 Assembly and Akt/PKB Dos D. Sarbassov, Siraj M. Ali, Shomit Sengupta, Joon-Ho Sheen, Peggy P. Hsu, Alex F. Bagley, Andrew L. Markhard, and
Supplemental Data Prolonged Rapamycin Treatment Inhibits mtorc2 Assembly and Akt/PKB Dos D. Sarbassov, Siraj M. Ali, Shomit Sengupta, Joon-Ho Sheen, Peggy P. Hsu, Alex F. Bagley, Andrew L. Markhard, and
Supplementary Figure 1
 A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
Single cell genetic analysis helps validating cytopathological identification of CTCs in patients with Clear Cells Renal Carcinoma
 Single cell genetic analysis helps validating cytopathological identification of CTCs in patients with Clear Cells Renal Carcinoma Patrizia Paterlini Bréchot, MD, Ph.D. Professor of Cell Biology/Oncology
Single cell genetic analysis helps validating cytopathological identification of CTCs in patients with Clear Cells Renal Carcinoma Patrizia Paterlini Bréchot, MD, Ph.D. Professor of Cell Biology/Oncology
GYNplus: A Genetic Test for Hereditary Ovarian and/or Uterine Cancer
 GYNplus: A Genetic Test for Hereditary Ovarian and/or Uterine Cancer Causes of Hereditary Ovarian and Uterine Cancer uterine cancer ovarian cancer Sporadic 75-90% Sporadic 70-80% Hereditary, 5% Lynch syndrome
GYNplus: A Genetic Test for Hereditary Ovarian and/or Uterine Cancer Causes of Hereditary Ovarian and Uterine Cancer uterine cancer ovarian cancer Sporadic 75-90% Sporadic 70-80% Hereditary, 5% Lynch syndrome
SUPPLEMENTARY INFORMATION. Rare independent mutations in renal salt handling genes contribute to blood pressure variation
 SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
GYNplus. genetic testing for hereditary ovarian and/or uterine cancer
 GYNplus genetic testing for hereditary ovarian and/or uterine cancer What Are the Causes of Hereditary Ovarian and Uterine Cancer? uterine cancer ovarian cancer sporadic 70-80% hereditary 5% Lynch syndrome
GYNplus genetic testing for hereditary ovarian and/or uterine cancer What Are the Causes of Hereditary Ovarian and Uterine Cancer? uterine cancer ovarian cancer sporadic 70-80% hereditary 5% Lynch syndrome
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes
 Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes KM Kortüm 1,8 *, EK Mai 3,4 *, NH Hanafiah 3 *, CX Shi 1, YX Zhu 1, L Bruins 1, S Barrio 1,
Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes KM Kortüm 1,8 *, EK Mai 3,4 *, NH Hanafiah 3 *, CX Shi 1, YX Zhu 1, L Bruins 1, S Barrio 1,
TGF-β-SPTBN1-CTCF-regulated tumor suppression in Beckwith- Wiedemann syndrome, a human stem cell disorder
 TGF-β-SPTBN1-CTCF-regulated tumor suppression in Beckwith- Wiedemann syndrome, a human stem cell disorder Jian Chen, Zhi-Xing Yao, Jiun-Sheng Chen, Young Jin Gi, Nina M. Muñoz, Suchin Kundra, H. Franklin
TGF-β-SPTBN1-CTCF-regulated tumor suppression in Beckwith- Wiedemann syndrome, a human stem cell disorder Jian Chen, Zhi-Xing Yao, Jiun-Sheng Chen, Young Jin Gi, Nina M. Muñoz, Suchin Kundra, H. Franklin
Analysis of individual differences in radiosensitivity using genome editing
 3 rd International Symposium on the System of Radiological Protection ICRP2015 Analysis of individual differences in radiosensitivity using genome editing Shinya Matsuura Department of Genetics and Cell
3 rd International Symposium on the System of Radiological Protection ICRP2015 Analysis of individual differences in radiosensitivity using genome editing Shinya Matsuura Department of Genetics and Cell
IntelliGENSM. Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community.
 IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
Multi-omics of 34 colorectal cancer cell lines - a resource for biomedical studies
 Berg et al. Molecular Cancer (2017) 16:116 DOI 10.1186/s12943-017-0691-y RESEARCH Multi-omics of 34 colorectal cancer cell lines - a resource for biomedical studies Open Access Kaja C. G. Berg 1,2, Peter
Berg et al. Molecular Cancer (2017) 16:116 DOI 10.1186/s12943-017-0691-y RESEARCH Multi-omics of 34 colorectal cancer cell lines - a resource for biomedical studies Open Access Kaja C. G. Berg 1,2, Peter
A novel Monoclonal Antibody against Notch1 Targets Leukemiaassociated Mutant Notch1 and Depletes Therapy Resistant Cancer Stem Cells in Solid Tumors
 Supplementary Information:- A novel Monoclonal Antibody against Notch1 Targets Leukemiaassociated Mutant Notch1 and Depletes Therapy Resistant Cancer Stem Cells in Solid Tumors Ankur Sharma 1, Rupali A
Supplementary Information:- A novel Monoclonal Antibody against Notch1 Targets Leukemiaassociated Mutant Notch1 and Depletes Therapy Resistant Cancer Stem Cells in Solid Tumors Ankur Sharma 1, Rupali A
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well
 Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
p.r623c p.p976l p.d2847fs p.t2671 p.d2847fs p.r2922w p.r2370h p.c1201y p.a868v p.s952* RING_C BP PHD Cbp HAT_KAT11
 ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
ARID2 p.r623c KMT2D p.v650fs p.p976l p.r2922w p.l1212r p.d1400h DNA binding RFX DNA binding Zinc finger KMT2C p.a51s p.d372v p.c1103* p.d2847fs p.t2671 p.d2847fs p.r4586h PHD/ RING DHHC/ PHD PHD FYR N
Table S1. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9. Assay (s) Target Name Sequence (5 3 ) Comments
 SUPPLEMENTAL INFORMATION 2 3 Table S. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9 genes. Assay (s) Target Name Sequence (5 3 ) Comments CDC M InfA Forward (NS), CDC M
SUPPLEMENTAL INFORMATION 2 3 Table S. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9 genes. Assay (s) Target Name Sequence (5 3 ) Comments CDC M InfA Forward (NS), CDC M
Co existence of PHF6 and NOTCH1 mutations in adult T-cell acute lymphoblastic leukemia
 16 Co existence of PHF6 and NOTCH1 mutations in adult T-cell acute lymphoblastic leukemia MIN LI 1*, LICHAN XIAO 1*, JINGYAN XU 2*, RUN ZHANG 1, JINGJING GUO 3, JUSTIN OLSON 4, YUJIE WU 1, JIANYONG LI
16 Co existence of PHF6 and NOTCH1 mutations in adult T-cell acute lymphoblastic leukemia MIN LI 1*, LICHAN XIAO 1*, JINGYAN XU 2*, RUN ZHANG 1, JINGJING GUO 3, JUSTIN OLSON 4, YUJIE WU 1, JIANYONG LI
HansaBioMed Catalog 2014 for Exosome Research. X. Human Cell Lines...
 HansaBioMed Catalog 2014 for Exosome Research X..... HansaBioMed Catalog 2014 HUMAN CELL LINES 380 leukemia, pre-b cell 1301 leukemia, acute lymphoblastic, T cell suspension, morphology round, small cells
HansaBioMed Catalog 2014 for Exosome Research X..... HansaBioMed Catalog 2014 HUMAN CELL LINES 380 leukemia, pre-b cell 1301 leukemia, acute lymphoblastic, T cell suspension, morphology round, small cells
Chromosome 15 Chromosome 17 Chromosome 20
 Chromosome 1 Chromosome 6 Chromosome 11 Chromosome 15 Chromosome 17 Chromosome 20 Supplementary figure 1. Regions of suggestive linkage in PVOD families 1,2 and 3. Six regions were identified with suggestive
Chromosome 1 Chromosome 6 Chromosome 11 Chromosome 15 Chromosome 17 Chromosome 20 Supplementary figure 1. Regions of suggestive linkage in PVOD families 1,2 and 3. Six regions were identified with suggestive
MUTATION TEST CE IVD. ctnras-braf FEATURES
 ctnras-braf CE IVD TECHNICAL SHEET IDYLLA ctnras-braf MUTATION TEST The Idylla ctnras-braf Mutation Test, performed on the Biocartis Idylla System, is an in vitro diagnostic test for the qualitative detection
ctnras-braf CE IVD TECHNICAL SHEET IDYLLA ctnras-braf MUTATION TEST The Idylla ctnras-braf Mutation Test, performed on the Biocartis Idylla System, is an in vitro diagnostic test for the qualitative detection
Genetic features of Lynch syndrome in the Israeli population
 Clin Genet 2015: 87: 549 553 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12530 Genetic features
Clin Genet 2015: 87: 549 553 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12530 Genetic features
Spectrum of mutations in monogenic diabetes genes identified from high-throughput DNA sequencing of 6888 individuals
 Bansal et al. BMC Medicine (2017) 15:213 DOI 10.1186/s12916-017-0977-3 RESEARCH ARTICLE Spectrum of mutations in monogenic diabetes genes identified from high-throughput DNA sequencing of 6888 individuals
Bansal et al. BMC Medicine (2017) 15:213 DOI 10.1186/s12916-017-0977-3 RESEARCH ARTICLE Spectrum of mutations in monogenic diabetes genes identified from high-throughput DNA sequencing of 6888 individuals
Molecular and genetic characterization of peroxisome biogenesis disorders Ebberink, M.S.
 UvA-DARE (Digital Academic Repository) Molecular and genetic characterization of peroxisome biogenesis disorders Ebberink, M.S. Link to publication Citation for published version (APA): Ebberink, M. S.
UvA-DARE (Digital Academic Repository) Molecular and genetic characterization of peroxisome biogenesis disorders Ebberink, M.S. Link to publication Citation for published version (APA): Ebberink, M. S.
Variant interpretation exercise. ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18
 Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Dysfunctional Families Annette S. Kim, MD, PhD Associate Professor, Harvard Medical School Brigham and Women s Hospital
 Dsfunctional Families Annette S. Kim, MD, PhD Associate Professor, Harvard Medical School Brigham and Women s Hospital Disclosure of Relevant Financial Relationships USCAP requires that all planners (Education
Dsfunctional Families Annette S. Kim, MD, PhD Associate Professor, Harvard Medical School Brigham and Women s Hospital Disclosure of Relevant Financial Relationships USCAP requires that all planners (Education
Hereditary spastic paraplegia type 35 caused by a novel FA2H mutation
 The Turkish Journal of Pediatrics 2017; 59: 329334 DOI: 10.24953/turkjped.2017.03.016 Case Report Hereditary spastic paraplegia type 35 caused by a novel FA2H mutation Gonca Bektaş¹, Gözde Yeşil², Edibe
The Turkish Journal of Pediatrics 2017; 59: 329334 DOI: 10.24953/turkjped.2017.03.016 Case Report Hereditary spastic paraplegia type 35 caused by a novel FA2H mutation Gonca Bektaş¹, Gözde Yeşil², Edibe
Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3
 Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
Supplemental Figure Legends. Figure S1. ERBB3 mrna levels are elevated in Luminal A breast cancers harboring ERBB3 ErbB3 gene copy number gain. Supplemental Figure S1. ERBB3 mrna levels are elevated in
Colclough et al., Human Mutation 1
 Region Colclough et al., Human Mutation 1 Supp. Table S1. List of HNF1A mutations Nucleotide Change NM_000545.5 Promoter and 5 UTR Mutations Protein Effect NM_000545.5 Three letter amino acid description
Region Colclough et al., Human Mutation 1 Supp. Table S1. List of HNF1A mutations Nucleotide Change NM_000545.5 Promoter and 5 UTR Mutations Protein Effect NM_000545.5 Three letter amino acid description
The Role of Next Generation Sequencing in Solid Tumor Mutation Testing
 The Role of Next Generation Sequencing in Solid Tumor Mutation Testing Allie H. Grossmann MD PhD Department of Pathology, University of Utah Division of Anatomic Pathology & Oncology, ARUP Laboratories
The Role of Next Generation Sequencing in Solid Tumor Mutation Testing Allie H. Grossmann MD PhD Department of Pathology, University of Utah Division of Anatomic Pathology & Oncology, ARUP Laboratories
QHerit Expanded Carrier Screen
 QHerit Expanded Carrier Screen Test Code: 94372 (X) Specimen Requirements: Preferred: 6 ml (4 ml minimum) room-temperature whole blood: 1.5 ml (1 ml minimum) in each of 4 lavender-top (EDTA) or yellow-top
QHerit Expanded Carrier Screen Test Code: 94372 (X) Specimen Requirements: Preferred: 6 ml (4 ml minimum) room-temperature whole blood: 1.5 ml (1 ml minimum) in each of 4 lavender-top (EDTA) or yellow-top
This is a repository copy of Towards a Next-Generation Sequencing Diagnostic Service for Tumour Genotyping: A Comparison of Panels and Platforms.
 This is a repository copy of Towards a Next-Generation Sequencing Diagnostic Service for Tumour Genotyping: A Comparison of Panels and Platforms. White Rose Research Online URL for this paper: http://eprints.whiterose.ac.uk/89503/
This is a repository copy of Towards a Next-Generation Sequencing Diagnostic Service for Tumour Genotyping: A Comparison of Panels and Platforms. White Rose Research Online URL for this paper: http://eprints.whiterose.ac.uk/89503/
Supplementary Information
 Supplementary Information - chimeric fusion transcript in human gastric cancer promotes tumorigenesis through activation of PI3K/AKT signaling Sun Mi Yun, Kwiyeom Yoon, Sunghoon Lee, Eunjeong Kim, Seong-Ho
Supplementary Information - chimeric fusion transcript in human gastric cancer promotes tumorigenesis through activation of PI3K/AKT signaling Sun Mi Yun, Kwiyeom Yoon, Sunghoon Lee, Eunjeong Kim, Seong-Ho
Supplementary Figure 1. Chromatin signature profiling by mass spectrometry.
 Supplementary Information Supplementary Figure 1. Chromatin signature profiling by mass spectrometry. 115 cell lines from the CCLE collection were subjected to chromatin signature profiling by mass spectrometry.
Supplementary Information Supplementary Figure 1. Chromatin signature profiling by mass spectrometry. 115 cell lines from the CCLE collection were subjected to chromatin signature profiling by mass spectrometry.
Long-term exposure to Myozyme results in a decrease of anti-drug antibodies in lateonset Pompe disease patients
 Long-term exposure to Myozyme results in a decrease of anti-drug antibodies in lateonset Pompe disease patients Elisa Masat 1, Pascal Laforêt 1,2, Marie De Antonio 3, Guillaume Corre 4, Barbara Perniconi
Long-term exposure to Myozyme results in a decrease of anti-drug antibodies in lateonset Pompe disease patients Elisa Masat 1, Pascal Laforêt 1,2, Marie De Antonio 3, Guillaume Corre 4, Barbara Perniconi
Collaboration between CEQAS and UK NEQAS for Molecular Genetics Solid Tumour EQAs and variant interpretation
 Collaboration between CEQAS and UK NEQAS for Molecular Genetics Solid Tumour EQAs and variant interpretation Dr Jenni Fairley Deputy Scheme Director (Molecular Pathology), GenQA, Edinburgh, UK From 1 st
Collaboration between CEQAS and UK NEQAS for Molecular Genetics Solid Tumour EQAs and variant interpretation Dr Jenni Fairley Deputy Scheme Director (Molecular Pathology), GenQA, Edinburgh, UK From 1 st
validated somatic mutations detected in pediatric and adult histiocytic neoplasm biopsies based
 Supplementary Figure 1. Somatic mutations detected in histiocytic neoplasms. Number of validated somatic mutations detected in pediatric and adult histiocytic neoplasm biopsies based on whole exome sequencing.
Supplementary Figure 1. Somatic mutations detected in histiocytic neoplasms. Number of validated somatic mutations detected in pediatric and adult histiocytic neoplasm biopsies based on whole exome sequencing.
