Supplementary Table 2. Identified causative mutations and/or mutation candidates.
|
|
|
- Elvin Kristopher Jefferson
- 5 years ago
- Views:
Transcription
1 Supplementary Table 2. Identified causative mutations and/or mutation candidates. Nonsense mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the mutation detected families 1 EYA1 NM_ NP_ c.634c>t p.q212x AR AB GJB2 NM_ NP_ c.408c>a p.y136x AR, sporadic AB MIA NM_ NP_ c.178c>t p.r60x AD AB MYO6 NM_ NP_ c.1975c>t p.r659x AD AB TMPRSS3 NM_ NP_ c.607c>a p.q203x AR AB Splicing junction mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the mutation detected families 1 KCNQ1 NM_ NP_ c g>t p.q530fs sporadic AB SLC26A4 NM_ NP_ c.919-2a>g p.v306fs AR, sporadic AB Insertion/Deletion mutations base change aa change Average depth Result of next generation in 432 patient Hereditary form of the mutation detected families 1 GJB2 NM_ NP_ c.36insg p.v12fs sporadic AB GJB2 NM_ NP_ c del16 p.g59fs AR, sporadic AB GJB2 NM_ NP_ c.235delc p.l79fs AR, sporadic, unknown AB GJB2 NM_ NP_ c delat p.h100fs AR, sporadic AB MYO15A NM_ NP_ c insc p.p393fs sporadic AB MYH9 NM_ NP_ c.3016insgag p.t1006fs AR AB TECTA NM_ NP_ c.597delg p.l199fs sporadic AB839001
2 Missense mutations base change aa change Average depth Result of next generation in 432 patient Polyphen2 score Hereditary form of the mutation detected families 1 ACTG1 NM_ NP_ c.485c>t p.t162m sporadic AB APOD NM_ NP_ c.91c>t p.p31s AD AB ATP6V1B1 NM_ NP_ c.599t>c p.i200t unknown AB BSND NM_ NP_ c.893g>a p.g298e sporadic AB CCS NM_ NP_ c.167c>t p.t56i sporadic AB CCS NM_ NP_ c.212g>a p.r71q sporadic AB CCS NM_ NP_ c.586c>t p.r196c sporadic AB CDH23 NM_ NP_ c.719c>t p.p240l AD/Mit, AR, sporadic AB CDH23 NM_ NP_ c.2866g>a p.e956k sporadic AB CDH23 NM_ NP_ c.4249c>t p.r1417w unknown AB CDH23 NM_ NP_ c.4346g>a p.g1449d sporadic AB CDH23 NM_ NP_ c.4762c>t p.r1588w AR, AD, sporadic AB CDH23 NM_ NP_ c.7145g>a p.r2382q sporadic AB CDH23 NM_ NP_ c.8734g>a p.g2912s sporadic AB CLU NM_ NP_ c.382g>a p.v128i AR AB COL11A1 NM_ NP_ c.560c>t p.t187m AD/Mit AB COL11A1 NM_ NP_ c.2614t>a p.f872i AD, sporadic AB COL11A1 NM_ NP_ c.2771c>t p.p924l sporadic AB COL11A2 NM_ NP_ c.688g>t p.g230w AD/Mit, sporadic AB COL11A2 NM_ NP_ c.2002c>t p.p668s AD/Mit, AR, sporadic AB COL11A2 NM_ NP_ c.3392g>a p.r1131q AD AB COL11A2 NM_ NP_ c.4265c>t p.p1422l sporadic AB COL2A1 NM_ NP_ c.4148c>t p.t1383m unknown AB COL2A1 NM_ NP_ c.4196a>g p.y1399c sporadic AB COL4A3 NM_ NP_ c.469g>c p.g157r AD AB COL4A3 NM_ NP_ c.1295c>t p.p432l AD, sporadic AB COL4A3 NM_ NP_ c.2827g>a p.g943r sporadic AB COL4A3 NM_ NP_ c.4928g>a p.r1643k AD AB COL4A4 NM_ NP_ c.2045a>g p.d682g unknown AB COL4A4 NM_ NP_ c.2872c>g p.p958a sporadic AB COL4A4 NM_ NP_ c.4316g>a p.g1439d AR AB839032
3 32 COL4A5 NM_ NP_ c.2858g>t p.g953v AD, AD/Mit, sporadic AB COL4A5 NM_ NP_ c.3044g>a p.g1015e AD/Mit AB COL4A5 NM_ NP_ c.4003c>t p.p1335s sporadic AB COL9A1 NM_ NP_ c.2395g>c p.g799r sporadic AB COL9A3 NM_ NP_ c.1361g>a p.g454e sporadic AB COL9A3 NM_ NP_ c.1649c>t p.p550l AD AB DIAPH1 NM_ NP_ c.2032c>t p.p678s AD/Mit AB EDN3 NM_ NP_ c.554c>t p.t185m sporadic AB EDNRB NM_ NP_ c.311a>t p.n104i sporadic AB ESRRB NM_ NP_ c.475c>t p.r159c sporadic AB ESRRB NM_ NP_ c.536g>a p.r179h sporadic AB EYA1 NM_ NP_ c.403g>a p.g135s AD/Mit, AD, sporadic AB EYA1 NM_ NP_ c.671g>t p.g224v sporadic AB EYA1 NM_ NP_ c.1286a>g p.d429g AD/Mit AB FBXO2 NM_ NP_ c.286g>a p.g96r AD/Mit AB GJB2 NM_ NP_ c.23c>t p.t8m unknown AB GJB2 NM_ NP_ c.109g>a p.v37i sporadic AB GJB2 NM_ NP_ c.134g>a p.g45e AR, sporadic, AD/Mit/AR AB GJB2 NM_ NP_ c.257c>g p.t86r unknown AB GJB2 NM_ NP_ c.427c>t p.r143w sporadic AB GJB3 NM_ NP_ c.250g>a p.v84i sporadic AB GJB3 NM_ NP_ c.437t>c p.l146p unknown AB GJB3 NM_ NP_ c.459g>t p.w153c sporadic AB GJB6 NM_ NP_ c.556a>g p.t186a sporadic AB GPR98 NM_ NP_ c.913t>c p.y305h sporadic AB GPR98 NM_ NP_ c.1797a>t p.r599s AR AB GPR98 NM_ NP_ c.1804g>a p.a602t AR, sporadic AB GPR98 NM_ NP_ c.2039a>g p.d680g sporadic AB GPR98 NM_ NP_ c.4703g>a p.s1568n sporadic AB GPR98 NM_ NP_ c.6469g>a p.v2157m sporadic AB GPR98 NM_ NP_ c.12335t>g p.l4112w sporadic AB GPR98 NM_ NP_ c.12554g>c p.g4185a sporadic AB GPR98 NM_ NP_ c.12704a>g p.y4235c sporadic AB GPR98 NM_ NP_ c.13763g>a p.g4588e sporadic AB GPR98 NM_ NP_ c.13996a>g p.i4666v AD AB GPR98 NM_ NP_ c.14319a>g p.i4773m sporadic AB839068
4 68 GPR98 NM_ NP_ c.18602a>c p.n6201h AD/Mit, sporadic AB ISLR NM_ NP_ c.146c>t p.p49l AD AB KCNQ4 NM_ NP_ c.2014g>a p.v672m sporadic AB KIAA1199 NM_ NP_ c.653g>a p.r218h sporadic AB LHFP NM_ NP_ c.299c>t p.a100v sporadic AB LRP1 NM_ NP_ c.6493g>a p.g2165r AD/Mit AB LRP1 NM_ NP_ c.8801c>t p.s2934l sporadic AB LRP1 NM_ NP_ c.10802c>t p.a3601v sporadic AB LRP1 NM_ NP_ c.11780g>a p.r3927h sporadic AB LRP1 NM_ NP_ c.12346c>t p.h4116y AR AB MARVELD2 NM_ NP_ c.166c>t p.p56s sporadic AB MARVELD2 NM_ NP_ c.1115g>a p.r372q sporadic AB MARVELD2 NM_ NP_ c.1641t>a p.d547e sporadic AB MYH14 NM_ NP_ c.2593a>c p.k865q sporadic AB MYH14 NM_ NP_ c.4804g>a p.e1602k sporadic AB MYH9 NM_ NP_ c.2404c>t p.r802w sporadic AB MYH9 NM_ NP_ c.4352c>t p.a1451v AD AB MYO15A NM_ NP_ c.514c>t p.l172f AD/Mit AB MYO15A NM_ NP_ c.554g>a p.g185d AD AB MYO15A NM_ NP_ c.613t>c p.f205l sporadic AB MYO15A NM_ NP_ c.671a>g p.y224c sporadic AB MYO15A NM_ NP_ c.4216g>a p.e1406k sporadic AB MYO15A NM_ NP_ c.4322g>t p.g1441v sporadic AB MYO15A NM_ NP_ c.4828g>a p.e1610k AR AB MYO15A NM_ NP_ c.4888c>g p.r1630g sporadic AB MYO15A NM_ NP_ c.5117g>t p.g1706v sporadic AB MYO15A NM_ NP_ c.9478c>t p.l3160f AD, sporadic AB MYO15A NM_ NP_ c.9781a>t p.n3261y sporadic AB MYO15A NM_ NP_ c.10263c>g p.i3421m sporadic AB MYO1A NM_ NP_ c.2303g>a p.r768q sporadic AB MYO3A NM_ NP_ c.426t>g p.h142q sporadic AB MYO3A NM_ NP_ c.848a>c p.q283p sporadic AB MYO3A NM_ NP_ c.1324c>a p.h442n AD, AD/AR/Mit, sporadic AB MYO3A NM_ NP_ c.1819g>t p.v607f sporadic AB MYO6 NM_ NP_ c.614g>a p.r205q AD AB MYO6 NM_ NP_ c.3814c>t p.r1272w sporadic AB839104
5 104 MYO7A NM_ NP_ c.2023c>t p.r675c AD, sporadic AB MYO7A NM_ NP_ c.4806g>a p.e1603k AD, sporadic AB MYO7A NM_ NP_ c.6235c>t p.r2079w sporadic AB NDP NM_ NP_ c.58g>a p.g20r sporadic AB OTOF NM_ NP_ c.157g>a p.a53t sporadic AB OTOF NM_ NP_ c.3683g>t p.r1228l sporadic AB OTOF NM_ NP_ c.4417g>t p.g1473c sporadic AB OTOF NM_ NP_ c.5408a>c p.e1803a AR AB PCDH15 NM_ NP_ c.298g>a p.g100r AD/Mit AB PCDH15 NM_ NP_ c.833g>a p.r278h sporadic AB PCDH15 NM_ NP_ c.944c>t p.p315l sporadic AB PCDH15 NM_ NP_ c.2528c>a p.a843d unknown AB PCDH15 NM_ NP_ c.2884c>t p.r962c AR, sporadic AB PCDH15 NM_ NP_ c.3451g>a p.g1151r sporadic AB POU4F3 NM_ NP_ c.736c>a p.p246t sporadic AB RDX NM_ NP_ c.869t>c p.l290p unknown AB SLC17A8 NM_ NP_ c.1120g>t p.a374s AD, sporadic AB SLC26A4 NM_ NP_ c.439a>g p.m147v sporadic AB SLC26A4 NM_ NP_ c.1229c>t p.t410m AR, sporadic AB SLC26A4 NM_ NP_ c.1579a>c p.t527p sporadic AB SLC26A4 NM_ NP_ c.2168a>g p.h723r AD, AD/Mit, AR, sporadic AB STRC NM_ NP_ c.2914c>t p.r972w sporadic AB STRC NM_ NP_ c.4622g>a p.r1541q sporadic AB TCOF1 NM_ NP_ c.1045a>g p.s349g sporadic AB TCOF1 NM_ NP_ c.2344c>g p.q782e sporadic AB TCOF1 NM_ NP_ c.2666g>a p.r889h AR AB TECTA NM_ NP_ c.1471c>t p.r491c sporadic, unknown AB TECTA NM_ NP_ c.3511g>a p.v1171m AD, sporadic AB TECTA NM_ NP_ c.4198c>t p.h1400y AD AB TECTA NM_ NP_ c.4315c>a p.l1439i AD/Mit AB TMPRSS3 NM_ NP_ c.212t>c p.f71s sporadic AB TMPRSS3 NM_ NP_ c.280g>a p.g94r sporadic, unknown AB TMPRSS3 NM_ NP_ c.316c>t p.r106c AD/Mit, sporadic AB TMPRSS3 NM_ NP_ c.1159g>a p.a387t AR AB TRIOBP NM_ NP_ c.154g>a p.d52n sporadic AB TRIOBP NM_ NP_ c.4840g>t p.g1614c AD/Mit, AR AB839140
6 140 TRIOBP NM_ NP_ c.5519g>a p.r1840h AR AB TRIOBP NM_ NP_ c.6860g>a p.r2287h AR AB USH1C NM_ NP_ c.188g>a p.r63q unknown AB USH1C NM_ NP_ c.2191c>t p.r731w sporadic AB USH2A NM_ NP_ c.206g>t p.s69i sporadic AB USH2A NM_ NP_ c.1678c>g p.p560a sporadic AB USH2A NM_ NP_ c.5608c>t p.r1870w sporadic AB USH2A NM_ NP_ c.1877g>a p.r626q sporadic AB USH2A NM_ NP_ c.4070c>t p.t1357m AR, sporadic AB USH2A NM_ NP_ c.4616c>t p.t1539i sporadic AB USH2A NM_ NP_ c.7000a>g p.n2334d AD/AR/Mit AB USH2A NM_ NP_ c.7068t>g p.n2356k AD/Mit, sporadic, unknown AB USH2A NM_ NP_ c.9259g>a p.v3087i AD AB USH2A NM_ NP_ c.10852g>a p.g3618s sporadic AB USH2A NM_ NP_ c.10904c>a p.t3635n AD/Mit AB USH2A NM_ NP_ c.10999a>c p.t3667p sporadic AB USH2A NM_ NP_ c.14017t>c p.y4673h AR AB USH2A NM_ NP_ c.15233c>g p.p5078r AD AB USP11 NM_ NP_ c.484g>a p.a162t sporadic AB WFS1 NM_ NP_ c.449c>t p.a150v AD/Mit AB WFS1 NM_ NP_ c.908t>c p.l303p AD AB WNK2 NM_ NP_ c.947c>t p.t316m sporadic AB WNK2 NM_ NP_ c.1703c>t p.p568l sporadic AB WNK2 NM_ NP_ c.5951c>t p.p1984l sporadic AB839164
Whole Exome Sequenced Characteristics
 Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
Supplementary Tables Supplementary Table 1: Patient characteristics of 45 whole exome sequenced HNSCC tumors Whole Exome Sequenced Characteristics Tumors (n=45) Age, years Median (range) 61.0 (19-90) Sex,
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label)
 July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
July 2015 Assay ID Assay Name Gene COSMIC ID Amino acid change Nucleotide change Wild type allele (VIC label) Mutant allele (FAM label) p AHLJ090 AKT1_33765 AKT1 33765 p.e17k c.49g>a C T 2 AHBKFRM BRAF_473
Spectrum and Classification of ATP7B Variants in a Large Cohort of Chinese Patients with Wilson s Disease Guides Genetic Diagnosis
 Ivyspring International Publisher 638 Theranostics Research Paper 2016; 6(5): 638-649. doi: 10.7150/thno.14596 Spectrum and Classification of ATP7B Variants in a Large Cohort of Chinese Patients with Wilson
Ivyspring International Publisher 638 Theranostics Research Paper 2016; 6(5): 638-649. doi: 10.7150/thno.14596 Spectrum and Classification of ATP7B Variants in a Large Cohort of Chinese Patients with Wilson
Clinical presentation. Duration (Month)
 Supplementary information, Table S1A: Clinical information for 125 PAs. No. Subtype Gender (M/F) Age at diagnosis (year) Clinical presentation Duration (Month) Maximal adenoma diameter(cm) Tumor invasion*
Supplementary information, Table S1A: Clinical information for 125 PAs. No. Subtype Gender (M/F) Age at diagnosis (year) Clinical presentation Duration (Month) Maximal adenoma diameter(cm) Tumor invasion*
Supplementary Online Content
 Supplementary Online Content Pearlman R, Frankel WL, Swanson B, et al. Prevalence and spectrum of germline cancer susceptibility gene mutations among patients with early-onset colorectal cancer. JAMA Oncol.
Supplementary Online Content Pearlman R, Frankel WL, Swanson B, et al. Prevalence and spectrum of germline cancer susceptibility gene mutations among patients with early-onset colorectal cancer. JAMA Oncol.
Supplementary Figure 1
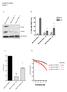 Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
Familial exudative vitreoretinopathy (FEVR) is an inheritable
 Genetics Spectrum of Variants in 389 Chinese Probands With Familial Exudative Vitreoretinopathy Jia-Kai Li, 1 Yian Li, 1 Xiang Zhang, 1 Chun-Li Chen, 2 Yu-Qing Rao, 1 Ping Fei, 1 Qi Zhang, 1 Peiquan Zhao,
Genetics Spectrum of Variants in 389 Chinese Probands With Familial Exudative Vitreoretinopathy Jia-Kai Li, 1 Yian Li, 1 Xiang Zhang, 1 Chun-Li Chen, 2 Yu-Qing Rao, 1 Ping Fei, 1 Qi Zhang, 1 Peiquan Zhao,
Belgian study into von Willebrand Disease (B-Will): First results
 Belgian study into von Willebrand Disease (B-Will): First results Inge Vangenechten VWD Research Unit, Antwerp University Hospital Supported by CSL Behring Chair in von Willebrand Disease, Antwerp University
Belgian study into von Willebrand Disease (B-Will): First results Inge Vangenechten VWD Research Unit, Antwerp University Hospital Supported by CSL Behring Chair in von Willebrand Disease, Antwerp University
GALAFOLD (migalastat) capsules, for oral use Initial U.S. Approval: 2018
 HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
Feedback of results. Report via to NTGMC inbox. Review by GMC clinician
 Genetic deafness Maria Bitner-Glindzicz Genetics and Genomic Medicine Programme UCL Institute of Child Health, UCL Ear Institute, and Great Ormond Street Hospital for Children Feedback of results Report
Genetic deafness Maria Bitner-Glindzicz Genetics and Genomic Medicine Programme UCL Institute of Child Health, UCL Ear Institute, and Great Ormond Street Hospital for Children Feedback of results Report
DNA Analysis in Glycogen storage disease
 DNA Analysis in Glycogen storage disease Nick Beauchamp PhD Sheffield, Sheffield Children s NHS Foundation Trust 14th October 2010 Glycogen Synthase Type 0 Glycogen Synthesis and Breakdown Type IV UDPGlucose
DNA Analysis in Glycogen storage disease Nick Beauchamp PhD Sheffield, Sheffield Children s NHS Foundation Trust 14th October 2010 Glycogen Synthase Type 0 Glycogen Synthesis and Breakdown Type IV UDPGlucose
GALAFOLD (migalastat) capsules, for oral use Initial U.S. Approval: 2018
 HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
The Genetics of Usher Syndrome
 The Genetics of Usher Syndrome Heidi L. Rehm, PhD, FACMG Assistant Professor of Pathology, BWH and HMS Director, Laboratory for Molecular Medicine, PCPGM DNA is Highly Compacted into Chromosomes The DNA
The Genetics of Usher Syndrome Heidi L. Rehm, PhD, FACMG Assistant Professor of Pathology, BWH and HMS Director, Laboratory for Molecular Medicine, PCPGM DNA is Highly Compacted into Chromosomes The DNA
Laboratory, Division of Medical Sciences, National Cancer Centre, Singapore
 Supplementary Information Exome sequencing of liver fluke-associated cholangiocarcinoma Choon Kiat Ong 1,2*, Chutima Subimerb 1,2,3*, Chawalit Pairojkul 3, Sopit Wongkham 3, Ioana Cutcutache 4, Willie
Supplementary Information Exome sequencing of liver fluke-associated cholangiocarcinoma Choon Kiat Ong 1,2*, Chutima Subimerb 1,2,3*, Chawalit Pairojkul 3, Sopit Wongkham 3, Ioana Cutcutache 4, Willie
Belgian-Czech cooperation in the Brno-VWD study
 Inge Vangenechten I Vangenechten, P Smejkal, O Zapletal, F Bouddount, J Zavrelova, J Blatny, M Penka, JJ Michiels, A Gadisseur. Aim of the study Characterisation of VWD in South Moravia (Czech Republic)
Inge Vangenechten I Vangenechten, P Smejkal, O Zapletal, F Bouddount, J Zavrelova, J Blatny, M Penka, JJ Michiels, A Gadisseur. Aim of the study Characterisation of VWD in South Moravia (Czech Republic)
GENETIC TESTING FOR HEREDITARY HEARING LOSS
 GENETIC TESTING FOR HEREDITARY HEARING LOSS Non-Discrimination Statement and Multi-Language Interpreter Services information are located at the end of this document. Coverage for services, procedures,
GENETIC TESTING FOR HEREDITARY HEARING LOSS Non-Discrimination Statement and Multi-Language Interpreter Services information are located at the end of this document. Coverage for services, procedures,
Research Article Mutation Analysis of the Common Deafness Genes in Patients with Nonsyndromic Hearing Loss in Linyi by SNPscan Assay
 BioMed Research International Volume 2016, Article ID 1302914, 7 pages http://dx.doi.org/10.1155/2016/1302914 Research Article Mutation Analysis of the Common Deafness Genes in Patients with Nonsyndromic
BioMed Research International Volume 2016, Article ID 1302914, 7 pages http://dx.doi.org/10.1155/2016/1302914 Research Article Mutation Analysis of the Common Deafness Genes in Patients with Nonsyndromic
Supplementary Table 1. PIK3CA mutation in colorectal cancer
 Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Liao X et al. PIK3CA Mutation in Colorectal Cancer. Page 1 Supplementary Table 1. PIK3CA mutation in colorectal cancer Exon Domain Nucleotide change* Amino acid change* cases 9 Helical c.1621t>a p.e541t
Nature Genetics: doi: /ng Supplementary Figure 1. Alternative splicing events in the 5K panel.
 Supplementary Figure 1 Alternative splicing events in the 5K panel. The majority of cryptic splicing occurred by creation of an AG or GT (Type I). While some other mutations increased the usage of a nearby
Supplementary Figure 1 Alternative splicing events in the 5K panel. The majority of cryptic splicing occurred by creation of an AG or GT (Type I). While some other mutations increased the usage of a nearby
Supplementary Information. Exome sequencing identifies distinct mutational patterns in liver fluke related and noninfection-related
 Supplementary Information Exome sequencing identifies distinct mutational patterns in liver fluke related and noninfection-related bile duct cancers Waraporn Chan-on 1,2,3*, Maarja-Liisa Nairismägi 1,2*,
Supplementary Information Exome sequencing identifies distinct mutational patterns in liver fluke related and noninfection-related bile duct cancers Waraporn Chan-on 1,2,3*, Maarja-Liisa Nairismägi 1,2*,
Spectrum of mutations in monogenic diabetes genes identified from high-throughput DNA sequencing of 6888 individuals
 Bansal et al. BMC Medicine (2017) 15:213 DOI 10.1186/s12916-017-0977-3 RESEARCH ARTICLE Spectrum of mutations in monogenic diabetes genes identified from high-throughput DNA sequencing of 6888 individuals
Bansal et al. BMC Medicine (2017) 15:213 DOI 10.1186/s12916-017-0977-3 RESEARCH ARTICLE Spectrum of mutations in monogenic diabetes genes identified from high-throughput DNA sequencing of 6888 individuals
A novel compound heterozygous mutation in SLC5A2 contributes to familial renal glucosuria in a Chinese family, and a review of the relevant literature
 MOLECULAR MEDICINE REPORTS A novel compound heterozygous mutation in SLC5A2 contributes to familial renal glucosuria in a Chinese family, and a review of the relevant literature SHENTANG LI 1, YEYI YANG
MOLECULAR MEDICINE REPORTS A novel compound heterozygous mutation in SLC5A2 contributes to familial renal glucosuria in a Chinese family, and a review of the relevant literature SHENTANG LI 1, YEYI YANG
Nature Genetics: doi: /ng Supplementary Figure 1. TCGA data set on HNSCCs reanalyzed in this study.
 Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
Summary of Results CARRIER STATUS DNA INSIGHT. Result: Carrier, Heterozygote. Protected Health Information. Propionic Acidemia
 Test Results Reviewed & Approved by: Laboratory Director, Nilesh Dharajiya,.D. SAPLE REPORT CARRIER STATUS DNA INSIGHT PERSONAL DETAILS DOB Jan 1, 19XX ETHNICITY Asian ORDERING HEALTHCARE PROFESSIONAL
Test Results Reviewed & Approved by: Laboratory Director, Nilesh Dharajiya,.D. SAPLE REPORT CARRIER STATUS DNA INSIGHT PERSONAL DETAILS DOB Jan 1, 19XX ETHNICITY Asian ORDERING HEALTHCARE PROFESSIONAL
Table S4. List of point mutations found by WES in SMZL patients. * DNA extracted from frozen tissue *Method of detection: Variant detected using
 Table S4. List of point mutations found by WES in SMZL patients. * DNA extracted from frozen tissue *Method of detection: Variant detected using RAMSES (R) or CNAG method (C) or both (R/C) **Validation
Table S4. List of point mutations found by WES in SMZL patients. * DNA extracted from frozen tissue *Method of detection: Variant detected using RAMSES (R) or CNAG method (C) or both (R/C) **Validation
(Case 5 A ) c g>a Exon 24 splice acceptor Splicing defect - Splicing defect
 Supplementary table S1. Pathogenic potential of identified DYNC2H1 mutations Family Nucleotide Change Mutation GERP PhyloP SIFT PolyPhen-2 Mutation Taster JATD-1 c. 90443A>G p.d3015g 6.17 5.3 Tolerated
Supplementary table S1. Pathogenic potential of identified DYNC2H1 mutations Family Nucleotide Change Mutation GERP PhyloP SIFT PolyPhen-2 Mutation Taster JATD-1 c. 90443A>G p.d3015g 6.17 5.3 Tolerated
The Personalized Cancer Medicine Initiative at Vanderbilt
 The Personalized Cancer Medicine Initiative at Vanderbilt October 26, 2009 William Pao, MD, PhD Assistant Director, Personalized Cancer Medicine Vanderbilt-Ingram Cancer Center Cancer in the United States,
The Personalized Cancer Medicine Initiative at Vanderbilt October 26, 2009 William Pao, MD, PhD Assistant Director, Personalized Cancer Medicine Vanderbilt-Ingram Cancer Center Cancer in the United States,
Genomic Profiling on an Unselected Solid Tumor Population Reveals a Highly Mutated Wnt/β-Catenin Pathway Associated with Oncogenic EGFR Mutations
 Article Genomic Profiling on an Unselected Solid Tumor Population Reveals a Highly Mutated Wnt/β-Catenin Pathway Associated with Oncogenic EGFR Mutations Jingrui Jiang 1, *, Alexei Protopopov 2, Ruobai
Article Genomic Profiling on an Unselected Solid Tumor Population Reveals a Highly Mutated Wnt/β-Catenin Pathway Associated with Oncogenic EGFR Mutations Jingrui Jiang 1, *, Alexei Protopopov 2, Ruobai
Mutational and phenotypical spectrum of phenylalanine hydroxylase deficiency in Denmark
 Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Supplementary Information
 Supplementary Information JAK1 truncating mutations in gynecologic cancer define new role of cancerassociated protein tyrosine kinase aberrations Yuan Ren, Yonghong Zhang, Richard Z. Liu, David A. Fenstermacher,
Supplementary Information JAK1 truncating mutations in gynecologic cancer define new role of cancerassociated protein tyrosine kinase aberrations Yuan Ren, Yonghong Zhang, Richard Z. Liu, David A. Fenstermacher,
Prevalence of Hearing Impairment
 Prevalence of Hearing Impairment 28 million Americans 2 million profoundly deaf 1/1000 congenitally deaf 1/3 impaired by age 65 1/2 impaired by age 80 NIDCD National Strategic Research Plan, 1989 Genetic
Prevalence of Hearing Impairment 28 million Americans 2 million profoundly deaf 1/1000 congenitally deaf 1/3 impaired by age 65 1/2 impaired by age 80 NIDCD National Strategic Research Plan, 1989 Genetic
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/20119 holds various files of this Leiden University dissertation. Author: Lotta, Luca Andrea Title: Pathophysiology of thrombotic thrombocytopenic purpura
Cover Page The handle http://hdl.handle.net/1887/20119 holds various files of this Leiden University dissertation. Author: Lotta, Luca Andrea Title: Pathophysiology of thrombotic thrombocytopenic purpura
Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM
 Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
Recent advances in the molecular pathology of lung cancer: role of EGFR and KRAS mutation testing in treatment selection
 ASIP 2009 USCAP Companion Meeting Molecular Pathology for the Practicing Pathologist Recent advances in the molecular pathology of lung cancer: role of EGFR and KRAS mutation testing in treatment selection
ASIP 2009 USCAP Companion Meeting Molecular Pathology for the Practicing Pathologist Recent advances in the molecular pathology of lung cancer: role of EGFR and KRAS mutation testing in treatment selection
Stem Cell Therapy for Acquired Hearing Loss in Children; FDA-Approved Study. Linda Baumgartner, CCC-SLP, Cert.AVT James Baumgartner, MD
 Stem Cell Therapy for Acquired Hearing Loss in Children; FDA-Approved Study Linda Baumgartner, CCC-SLP, Cert.AVT James Baumgartner, MD Stem Cell Basics I'll never grow up, never grow up, never grow up
Stem Cell Therapy for Acquired Hearing Loss in Children; FDA-Approved Study Linda Baumgartner, CCC-SLP, Cert.AVT James Baumgartner, MD Stem Cell Basics I'll never grow up, never grow up, never grow up
A Natural History Study of Subjects with X-linked Retinoschisis in Anticipation of a Phase I/II Gene Therapy Trial
 A Natural History Study of Subjects with X-linked Retinoschisis in Anticipation of a Phase I/II Gene Therapy Trial Mark E. Pennesi, MD/PhD Associate Professor X-linked Retinoschisis Background Rare X-Linked
A Natural History Study of Subjects with X-linked Retinoschisis in Anticipation of a Phase I/II Gene Therapy Trial Mark E. Pennesi, MD/PhD Associate Professor X-linked Retinoschisis Background Rare X-Linked
UW359 Ovary 3c 3 Serous Recurrent 68 BRCA1 816delGT BRCA1 del exon 1-2. UW417 Ovary 3c 3 Serous Primary 38 BRCA1 1675delA
 Supplementary Table 1. Cases with deleterious germline mutations, somatic HR mutations, and somatic PTEN mutations. ID Site Stage Grade Histology Tumor Age Germline mutation(s) a Somatic HR mutation(s)
Supplementary Table 1. Cases with deleterious germline mutations, somatic HR mutations, and somatic PTEN mutations. ID Site Stage Grade Histology Tumor Age Germline mutation(s) a Somatic HR mutation(s)
Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future?
 Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future? Tony Marinakii Purine Research Laboratory Biochemical Sciences Whole exome sequencing No doubt
Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future? Tony Marinakii Purine Research Laboratory Biochemical Sciences Whole exome sequencing No doubt
Comprehensive genetic testing for hearing and vision loss
 Comprehensive genetic testing for hearing and vision loss Hearing and vision loss can result from both genetic and non-genetic etiologies In general, there is a genetic basis for up to 50% of prelingual
Comprehensive genetic testing for hearing and vision loss Hearing and vision loss can result from both genetic and non-genetic etiologies In general, there is a genetic basis for up to 50% of prelingual
TGF-β-SPTBN1-CTCF-regulated tumor suppression in Beckwith- Wiedemann syndrome, a human stem cell disorder
 TGF-β-SPTBN1-CTCF-regulated tumor suppression in Beckwith- Wiedemann syndrome, a human stem cell disorder Jian Chen, Zhi-Xing Yao, Jiun-Sheng Chen, Young Jin Gi, Nina M. Muñoz, Suchin Kundra, H. Franklin
TGF-β-SPTBN1-CTCF-regulated tumor suppression in Beckwith- Wiedemann syndrome, a human stem cell disorder Jian Chen, Zhi-Xing Yao, Jiun-Sheng Chen, Young Jin Gi, Nina M. Muñoz, Suchin Kundra, H. Franklin
Negative feedback-defective PRPS1 mutants drive thiopurine. resistance in relapsed childhood ALL
 SUPPLEMENTARY INFORMATION Negative feedback-defective PRPS1 mutants drive thiopurine resistance in relapsed childhood ALL Benshang Li 1,5,, Hui Li 1,5,, Yun Bai 2,, Renate Kirschner-Schwabe 3,10, Jun J
SUPPLEMENTARY INFORMATION Negative feedback-defective PRPS1 mutants drive thiopurine resistance in relapsed childhood ALL Benshang Li 1,5,, Hui Li 1,5,, Yun Bai 2,, Renate Kirschner-Schwabe 3,10, Jun J
X-exome Sequencing of 405 Unresolved Families Identifies Seven Novel Intellectual Disability Genes
 X-exome Sequencing of 405 Unresolved Families Identifies Seven Novel Intellectual Disability Genes Hao Hu, 1,41 Stefan A. Haas, 2,41 Jamel Chelly, 3,4 Hilde Van Esch, 5 Martine Raynaud, 6,7,8 Arjan P.M.
X-exome Sequencing of 405 Unresolved Families Identifies Seven Novel Intellectual Disability Genes Hao Hu, 1,41 Stefan A. Haas, 2,41 Jamel Chelly, 3,4 Hilde Van Esch, 5 Martine Raynaud, 6,7,8 Arjan P.M.
Corporate Medical Policy
 Corporate Medical Policy Genetic Testing for Hereditary Hearing Loss File Name: Origination: Last CAP Review: Next CAP Review: Last Review: genetic_testing_for_hereditary_hearing_loss 10/2013 7/2018 7/2019
Corporate Medical Policy Genetic Testing for Hereditary Hearing Loss File Name: Origination: Last CAP Review: Next CAP Review: Last Review: genetic_testing_for_hereditary_hearing_loss 10/2013 7/2018 7/2019
Exomes and Beyond Addressing the Genome with OneSeq. Madhuri Hegde, PhD, FACMG Emory University Emory Genetics Laboratory Atlanta, GA
 Exomes and Beyond Addressing the Genome with OneSeq Madhuri Hegde, PhD, FACMG Emory University Emory Genetics Laboratory Atlanta, GA Technologies to Detect Various Types of Mutations RESOLUTION Targeted
Exomes and Beyond Addressing the Genome with OneSeq Madhuri Hegde, PhD, FACMG Emory University Emory Genetics Laboratory Atlanta, GA Technologies to Detect Various Types of Mutations RESOLUTION Targeted
Whole exome sequencing Gene package Hearing impairment version 2,
 Whole Exome Sequencing Gene package Hearing impairment, version 2, 23 9 2016 Technical information After DNA was enriched using Agilent Sureselect Clinical Research Exome (CRE) Capture, samples were run
Whole Exome Sequencing Gene package Hearing impairment, version 2, 23 9 2016 Technical information After DNA was enriched using Agilent Sureselect Clinical Research Exome (CRE) Capture, samples were run
Genetic Testing for Hereditary Hearing Loss Section 2.0 Medicine Subsection 2.04 Pathology/Laboratory
 2.04.87 Genetic Testing for Hereditary Hearing Loss Section 2.0 Medicine Subsection 2.04 Pathology/Laboratory Effective Date 1/30/2015 Original Policy Date 1/30/2015 Next Review Date January 2016 Description
2.04.87 Genetic Testing for Hereditary Hearing Loss Section 2.0 Medicine Subsection 2.04 Pathology/Laboratory Effective Date 1/30/2015 Original Policy Date 1/30/2015 Next Review Date January 2016 Description
Detection of BRCA1/2 mutations in breast cancer patients from Thailand and Pakistan
 Clin Genet 2012: 82: 594 598 Printed in Singapore. All rights reserved Letter to the Editor 2012 John Wiley & Sons A/S. Published by Blackwell Publishing Ltd CLINICAL GENETICS doi: 10.1111/j.1399-0004.2012.01869.x
Clin Genet 2012: 82: 594 598 Printed in Singapore. All rights reserved Letter to the Editor 2012 John Wiley & Sons A/S. Published by Blackwell Publishing Ltd CLINICAL GENETICS doi: 10.1111/j.1399-0004.2012.01869.x
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Generation of cetuximab-resistant cells and analysis of singlecell clones. Cetuximab-sensitive cells (LIM1215 and OXCO-2) were chronically treated with cetuximab
Supplementary Figure 1 Supplementary Figure 1: Generation of cetuximab-resistant cells and analysis of singlecell clones. Cetuximab-sensitive cells (LIM1215 and OXCO-2) were chronically treated with cetuximab
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Nangalia J, Massie CE, Baxter EJ, et al. Somatic CALR s in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Nangalia J, Massie CE, Baxter EJ, et al. Somatic CALR s in
Intron p Hybrid with p.leu245val p. Gly336Cys. p.leu62phe p. Ala137Val p. Asn152Thr hybrid with p.leu245val p.
 RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
Hereditary hemorrhagic telangiectasia: ENG and ALK-1 mutations in Dutch patients
 Hum Genet (2005) 116: 8 16 DOI 10.1007/s00439-004-1196-5 ORIGINAL INVESTIGATION T. G. W. Letteboer Æ R. A. Zewald Æ E. J. Kamping G. de Haas Æ J. J. Mager Æ R. J. Snijder Æ D. Lindhout F. A. M. Hennekam
Hum Genet (2005) 116: 8 16 DOI 10.1007/s00439-004-1196-5 ORIGINAL INVESTIGATION T. G. W. Letteboer Æ R. A. Zewald Æ E. J. Kamping G. de Haas Æ J. J. Mager Æ R. J. Snijder Æ D. Lindhout F. A. M. Hennekam
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary information
 Supplementary information! Recurrent activating mutation in PRKACA in cortisol-producing adrenal tumors Gerald Goh, Ute I. Scholl, James M. Healy, Murim Choi, Manju L. Prasad, Carol Nelson- Williams, John
Supplementary information! Recurrent activating mutation in PRKACA in cortisol-producing adrenal tumors Gerald Goh, Ute I. Scholl, James M. Healy, Murim Choi, Manju L. Prasad, Carol Nelson- Williams, John
Protocol. Genetic Testing for Nonsyndromic Hearing Loss
 Protocol Genetic Testing for Nonsyndromic Hearing Loss (20487) Medical Benefit Effective Date: 04/01/14 Next Review Date: 01/15 Preauthorization Yes Review Dates: 01/14 The following Protocol contains
Protocol Genetic Testing for Nonsyndromic Hearing Loss (20487) Medical Benefit Effective Date: 04/01/14 Next Review Date: 01/15 Preauthorization Yes Review Dates: 01/14 The following Protocol contains
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP
 Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Distinct molecular abnormalities underlie unique clinical features of essential thrombocythemia. in children
 Distinct molecular abnormalities underlie unique clinical features of essential thrombocythemia in children Supplementary methods Patients and diagnostic criteria The study was approved by the hospital-based
Distinct molecular abnormalities underlie unique clinical features of essential thrombocythemia in children Supplementary methods Patients and diagnostic criteria The study was approved by the hospital-based
Names for ABO (ISBT 001) Blood Group Alleles
 Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
UKGTN Testing Criteria Test name: Syndromic and Non Syndromic Hearing Loss 95 Gene Panel
 UKGTN Testing Criteria Test name: Syndromic and Non Syndromic Hearing Loss 95 Gene Panel Approved name and symbol of disorder/condition(s): See Appendix 1 Approved name and symbol of gene(s): See Appendix
UKGTN Testing Criteria Test name: Syndromic and Non Syndromic Hearing Loss 95 Gene Panel Approved name and symbol of disorder/condition(s): See Appendix 1 Approved name and symbol of gene(s): See Appendix
Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes
 Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes KM Kortüm 1,8 *, EK Mai 3,4 *, NH Hanafiah 3 *, CX Shi 1, YX Zhu 1, L Bruins 1, S Barrio 1,
Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes KM Kortüm 1,8 *, EK Mai 3,4 *, NH Hanafiah 3 *, CX Shi 1, YX Zhu 1, L Bruins 1, S Barrio 1,
Whole exome sequencing Gene package Hearing impairment version 3.1,
 Whole Exome Sequencing Gene package Hearing impairment, version 3.1, 22 11 2017 Technical information DNA was enriched using Agilent SureSelect Clinical Research Exome V2 capture and paired end sequenced
Whole Exome Sequencing Gene package Hearing impairment, version 3.1, 22 11 2017 Technical information DNA was enriched using Agilent SureSelect Clinical Research Exome V2 capture and paired end sequenced
SUPPLEMENTARY INFORMATION
 Somatic ERCC2 Mutations Are Associated with a Distinct Genomic Signature in Urothelial Tumors Jaegil Kim, Kent W Mouw, Paz Polak, Lior Z Braunstein, Atanas Kamburov, Grace Tiao, David J Kwiatkowski, Jonathan
Somatic ERCC2 Mutations Are Associated with a Distinct Genomic Signature in Urothelial Tumors Jaegil Kim, Kent W Mouw, Paz Polak, Lior Z Braunstein, Atanas Kamburov, Grace Tiao, David J Kwiatkowski, Jonathan
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
A novel tumor suppressor role for the Notch pathway in bladder cancer
 A novel tumor suppressor role for the Notch pathway in bladder cancer Theodoros Rampias, Paraskevi Vgenopoulou, Margaritis Avgeris, Alexander Polyzos, Konstantinos Stravodimos, Christos Valavanis, Andreas
A novel tumor suppressor role for the Notch pathway in bladder cancer Theodoros Rampias, Paraskevi Vgenopoulou, Margaritis Avgeris, Alexander Polyzos, Konstantinos Stravodimos, Christos Valavanis, Andreas
Supplementary appendix
 Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Wells JM, Farris RF, Gosdin TA, et al. Pulmonary
Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Wells JM, Farris RF, Gosdin TA, et al. Pulmonary
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
ClinGen Hearing Loss Variant Curation Expert Panel ACMG/AMP Classification Rules for Hearing Loss SUMMARY OF CLASSIFICATION CRITERIA
 PATHOGENIC CRITERIA RULE RULE DESCRIPTION ClinGen Hearing Loss Variant Curation Expert Panel ACMG/AMP Classification Rules for Hearing Loss SUMMARY OF CLASSIFICATION CRITERIA PVS1 PVS1_Strong PVS1_Moderate
PATHOGENIC CRITERIA RULE RULE DESCRIPTION ClinGen Hearing Loss Variant Curation Expert Panel ACMG/AMP Classification Rules for Hearing Loss SUMMARY OF CLASSIFICATION CRITERIA PVS1 PVS1_Strong PVS1_Moderate
Usher Syndrome and Progressive Hearing Loss
 Usher Syndrome and Progressive Hearing Loss Margaret A. Kenna, MD, MPH Otolaryngology and Communication Enhancement Boston Children s Hospital Professor of Otology and Laryngology Harvard Medical School
Usher Syndrome and Progressive Hearing Loss Margaret A. Kenna, MD, MPH Otolaryngology and Communication Enhancement Boston Children s Hospital Professor of Otology and Laryngology Harvard Medical School
Hereditary spastic paraplegia type 35 caused by a novel FA2H mutation
 The Turkish Journal of Pediatrics 2017; 59: 329334 DOI: 10.24953/turkjped.2017.03.016 Case Report Hereditary spastic paraplegia type 35 caused by a novel FA2H mutation Gonca Bektaş¹, Gözde Yeşil², Edibe
The Turkish Journal of Pediatrics 2017; 59: 329334 DOI: 10.24953/turkjped.2017.03.016 Case Report Hereditary spastic paraplegia type 35 caused by a novel FA2H mutation Gonca Bektaş¹, Gözde Yeşil², Edibe
Genetic Testing for Hereditary Hearing Loss
 Protocol Genetic Testing for Hereditary Hearing Loss (20487) Medical Benefit Effective Date: 01/01/18 Next Review Date: 11/18 Preauthorization Yes Review Dates: 01/14, 11/14, 11/15, 11/16, 11/17 Preauthorization
Protocol Genetic Testing for Hereditary Hearing Loss (20487) Medical Benefit Effective Date: 01/01/18 Next Review Date: 11/18 Preauthorization Yes Review Dates: 01/14, 11/14, 11/15, 11/16, 11/17 Preauthorization
LMM / emerge III Network Reference Sequences October 2016 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene
 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
This is the published version of a paper published in BMC Cardiovascular Disorders. Citation for the original published paper (version of record):
 http://www.diva-portal.org This is the published version of a paper published in BMC Cardiovascular Disorders. Citation for the original published paper (version of record): Stattin, E., Boström, I., Winbo,
http://www.diva-portal.org This is the published version of a paper published in BMC Cardiovascular Disorders. Citation for the original published paper (version of record): Stattin, E., Boström, I., Winbo,
Names for H (ISBT 018) Blood Group Alleles
 Names for H (ISBT 018) Blood Group Alleles General description: The H blood group system consists of one antigen, H, that is carried on glycolipids and glycoproteins on the RBC membrane, where it is synthesised
Names for H (ISBT 018) Blood Group Alleles General description: The H blood group system consists of one antigen, H, that is carried on glycolipids and glycoproteins on the RBC membrane, where it is synthesised
Spectrum of somatically acquired mutations identified by combining WES and genome-wide DNA array analysis in the discovery cohort of 30 JMML cases.
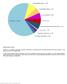 Supplementary Figure 1 Spectrum of somatically acquired mutations identified by combining WES and genome-wide DNA array analysis in the discovery cohort of 30 JMML cases. A total of 85 somatically acquired
Supplementary Figure 1 Spectrum of somatically acquired mutations identified by combining WES and genome-wide DNA array analysis in the discovery cohort of 30 JMML cases. A total of 85 somatically acquired
FEP Medical Policy Manual
 FEP Medical Policy Manual Effective Date: July 15, 2018 Related Policies: 2.04.102 Whole Exome and Whole Genome Sequencing for Diagnosis of Genetic Disorders Genetic Testing for Hereditary Hearing Loss
FEP Medical Policy Manual Effective Date: July 15, 2018 Related Policies: 2.04.102 Whole Exome and Whole Genome Sequencing for Diagnosis of Genetic Disorders Genetic Testing for Hereditary Hearing Loss
Unravelling the genetic basis of simplex Retinitis Pigmentosa cases
 SUPPLEMENTARY INFORMATION Unravelling the genetic basis simplex Retinitis Pigmentosa cases Nereida Bravo-Gil 1,2#, María González-del Pozo 1,2#, Marta Martín-Sánchez 1, Cristina Méndez-Vidal 1,2, Enrique
SUPPLEMENTARY INFORMATION Unravelling the genetic basis simplex Retinitis Pigmentosa cases Nereida Bravo-Gil 1,2#, María González-del Pozo 1,2#, Marta Martín-Sánchez 1, Cristina Méndez-Vidal 1,2, Enrique
SUPPLEMENTARY INFORMATION. Rare independent mutations in renal salt handling genes contribute to blood pressure variation
 SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
CHAPTER IV RESULTS Microcephaly General description
 47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
Supplementary Online Content
 Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Moyamoya disease (MMD) is a vasculopathy characterized
 CLINICAL ARTICLE J Neurosurg 126:1106 1113, 2017 RNF213 as the major susceptibility gene for Chinese patients with moyamoya disease and its clinical relevance *Qian Zhang, MD, 1 Yaping Liu, PhD, 2 Dong
CLINICAL ARTICLE J Neurosurg 126:1106 1113, 2017 RNF213 as the major susceptibility gene for Chinese patients with moyamoya disease and its clinical relevance *Qian Zhang, MD, 1 Yaping Liu, PhD, 2 Dong
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Usher syndrome (USH) is an autosomal recessive disorder
 Biochemistry and Molecular Biology Microarray-Based Mutation Analysis of 183 Spanish Families with Usher Syndrome Teresa Jaijo, 1,2 Elena Aller, 1,2 Gema García-García, 1,2 María J. Aparisi, 1 Sara Bernal,
Biochemistry and Molecular Biology Microarray-Based Mutation Analysis of 183 Spanish Families with Usher Syndrome Teresa Jaijo, 1,2 Elena Aller, 1,2 Gema García-García, 1,2 María J. Aparisi, 1 Sara Bernal,
Nature Genetics: doi: /ng Supplementary Figure 1. TNFAIP3-associated haplotypes in family 1.
 Supplementary Figure 1 TNFAIP3-associated haplotypes in family 1. The p.leu227* mutation (shown as a star) arose de novo in the first affected member of the family (P1). Red haplotypes carry the TNFAIP3
Supplementary Figure 1 TNFAIP3-associated haplotypes in family 1. The p.leu227* mutation (shown as a star) arose de novo in the first affected member of the family (P1). Red haplotypes carry the TNFAIP3
Spectrum of USH2A Mutations in Scandinavian Patients with Usher Syndrome Type II
 MUTATIO I BRIEF HUMA MUTATIO Mutation in Brief #995(2008) Online pectrum of UH2A Mutations in candinavian Patients with Usher yndrome Type II Bo reyer, Vigdis Brox 2, Lisbeth Tranebjærg -4, Thomas Rosenberg
MUTATIO I BRIEF HUMA MUTATIO Mutation in Brief #995(2008) Online pectrum of UH2A Mutations in candinavian Patients with Usher yndrome Type II Bo reyer, Vigdis Brox 2, Lisbeth Tranebjærg -4, Thomas Rosenberg
Abstract. Introduction
 Molecular Epidemiology and Functional Assessment of Novel Allelic Variants of SLC26A4 in Non-Syndromic Hearing Loss Patients with Enlarged Vestibular Aqueduct in China Yongyi Yuan 1,4., Weiwei Guo 1.,
Molecular Epidemiology and Functional Assessment of Novel Allelic Variants of SLC26A4 in Non-Syndromic Hearing Loss Patients with Enlarged Vestibular Aqueduct in China Yongyi Yuan 1,4., Weiwei Guo 1.,
Detection of BRCA1 and BRCA2 germline mutations in Japanese population using next-generation sequencing
 ORIGINAL ARTICLE Detection of BRCA1 and BRCA2 germline mutations in Japanese population using next-generation sequencing Yosuke Hirotsu 1, Hiroshi Nakagomi 2, Ikuko Sakamoto 3, Kenji Amemiya 1,4, Hitoshi
ORIGINAL ARTICLE Detection of BRCA1 and BRCA2 germline mutations in Japanese population using next-generation sequencing Yosuke Hirotsu 1, Hiroshi Nakagomi 2, Ikuko Sakamoto 3, Kenji Amemiya 1,4, Hitoshi
b 0.50 Synonymous Subtype
 omatic mutation count Mutations per Mb a 80 * b 0.50 ynonymous 60 0.40 Non-synonymous 40 0.30 0.20 20 0.10 c 0 Fibroadenoma Phyllodes 0.00 ubtype Benign Borderline MAL upplementary Figure 1 Landscape of
omatic mutation count Mutations per Mb a 80 * b 0.50 ynonymous 60 0.40 Non-synonymous 40 0.30 0.20 20 0.10 c 0 Fibroadenoma Phyllodes 0.00 ubtype Benign Borderline MAL upplementary Figure 1 Landscape of
Supporting Information
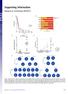 Supporting Information Rampal et al. 10.1073/pnas.1407792111 Fig. S1. Genetic events in leukemic transformation of chronic-phase MPNs. (A) Survival of post-mpn AML patients according to mutational status
Supporting Information Rampal et al. 10.1073/pnas.1407792111 Fig. S1. Genetic events in leukemic transformation of chronic-phase MPNs. (A) Survival of post-mpn AML patients according to mutational status
Mutations in Danish patients with long QT syndrome and the identification of a large
 Christiansen et al. BMC Medical Genetics 2014, 15:31 RESEARCH ARTICLE Open Access Mutations in Danish patients with long QT syndrome and the identification of a large founder family with p.f29l in KCNH2
Christiansen et al. BMC Medical Genetics 2014, 15:31 RESEARCH ARTICLE Open Access Mutations in Danish patients with long QT syndrome and the identification of a large founder family with p.f29l in KCNH2
Genetic Hearing Loss in Children
 Genetic Hearing Loss in Children José Faibes Lubianca & Ricardo Godinho The prevalence of genetic hearing loss reaches very high numbers. In developed countries, about 50% of the cases of pre-lingual severe
Genetic Hearing Loss in Children José Faibes Lubianca & Ricardo Godinho The prevalence of genetic hearing loss reaches very high numbers. In developed countries, about 50% of the cases of pre-lingual severe
Mucopolysaccharidosis type IIIB may predominantly present with an attenuated clinical phenotype
 J Inherit Metab Dis (2010) 33:759 767 DOI 10.1007/s10545-010-9199-y ORIGINAL ARTICLE Mucopolysaccharidosis type IIIB may predominantly present with an attenuated clinical phenotype Marlies J. Valstar &
J Inherit Metab Dis (2010) 33:759 767 DOI 10.1007/s10545-010-9199-y ORIGINAL ARTICLE Mucopolysaccharidosis type IIIB may predominantly present with an attenuated clinical phenotype Marlies J. Valstar &
POLICY PRODUCT VARIATIONS DESCRIPTION/BACKGROUND RATIONALE DEFINITIONS BENEFIT VARIATIONS DISCLAIMER REFERENCES CODING INFORMATION POLICY HISTORY
 Original Issue Date (Created): November 26, 2013 Most Recent Review Date (Revised): November 26, 2013 Effective Date: February 01, 2014 POLICY PRODUCT VARIATIONS DESCRIPTION/BACKGROUND RATIONALE DEFINITIONS
Original Issue Date (Created): November 26, 2013 Most Recent Review Date (Revised): November 26, 2013 Effective Date: February 01, 2014 POLICY PRODUCT VARIATIONS DESCRIPTION/BACKGROUND RATIONALE DEFINITIONS
Genotype phenotype correlations for hearing impairment: Approaches to management
 Clin Genet 2014: 85: 514 523 Printed in Singapore. All rights reserved Review 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12339 Genotype phenotype
Clin Genet 2014: 85: 514 523 Printed in Singapore. All rights reserved Review 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12339 Genotype phenotype
Original Article Genetic mutations of hotspots in patients with nonsyndromic deafness in Fujian, China
 Int J Clin Exp Pathol 2017;10(6):7067-7074 www.ijcep.com /ISSN:1936-2625/IJCEP0054265 Original Article Genetic mutations of hotspots in patients with nonsyndromic deafness in Fujian, China Lihua Wu 1,
Int J Clin Exp Pathol 2017;10(6):7067-7074 www.ijcep.com /ISSN:1936-2625/IJCEP0054265 Original Article Genetic mutations of hotspots in patients with nonsyndromic deafness in Fujian, China Lihua Wu 1,
Variable Clinical Expression in Patients with Mosaicism for KCNQ2 Mutations
 Variable Clinical Expression in Patients with Mosaicism for KCNQ2 Mutations Mathieu Milh, 1,2,3 Caroline Lacoste, 1,2,4 Pierre Cacciagli, 1,2,4 Affef Abidi, 1,2 Julie Sutera-Sardo, 1,2,3 Ilias Tzelepis,
Variable Clinical Expression in Patients with Mosaicism for KCNQ2 Mutations Mathieu Milh, 1,2,3 Caroline Lacoste, 1,2,4 Pierre Cacciagli, 1,2,4 Affef Abidi, 1,2 Julie Sutera-Sardo, 1,2,3 Ilias Tzelepis,
Exome Sequencing Reveals a Novel PRPS1 Mutation in a Family with CMTX5 without Optic Atrophy
 CASE REPORT J Clin Neurol 2013;9:283-288 Print ISSN 1738-6586 / On-line ISSN 2005-5013 http://dx.doi.org/10.3988/jcn.2013.9.4.283 Open Access Exome Sequencing Reveals a Novel PRPS1 Mutation in a Family
CASE REPORT J Clin Neurol 2013;9:283-288 Print ISSN 1738-6586 / On-line ISSN 2005-5013 http://dx.doi.org/10.3988/jcn.2013.9.4.283 Open Access Exome Sequencing Reveals a Novel PRPS1 Mutation in a Family
Table S1. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9. Assay (s) Target Name Sequence (5 3 ) Comments
 SUPPLEMENTAL INFORMATION 2 3 Table S. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9 genes. Assay (s) Target Name Sequence (5 3 ) Comments CDC M InfA Forward (NS), CDC M
SUPPLEMENTAL INFORMATION 2 3 Table S. Oligonucleotides used for the in-house RT-PCR assays targeting the M, H7 or N9 genes. Assay (s) Target Name Sequence (5 3 ) Comments CDC M InfA Forward (NS), CDC M
Supplemental information
 Supplemental information Frequent reconstitution of IDH2 mutant clonal multilineage hematopoiesis following chemotherapy for acute myeloid leukemia Daniel H. Wiseman, Emma L. Williams, Deepti P. Wilks,
Supplemental information Frequent reconstitution of IDH2 mutant clonal multilineage hematopoiesis following chemotherapy for acute myeloid leukemia Daniel H. Wiseman, Emma L. Williams, Deepti P. Wilks,
Supplementary Figure 1
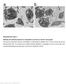 Supplementary Figure 1 Platelets from affected patients are comparable to controls on electron micrograph. Thin-section transmission electron micrographs of representative platelets from a normal control
Supplementary Figure 1 Platelets from affected patients are comparable to controls on electron micrograph. Thin-section transmission electron micrographs of representative platelets from a normal control
ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS
 ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS 1 This medicinal product is subject to additional monitoring. This will allow quick identification of new safety information. Healthcare professionals are asked
ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS 1 This medicinal product is subject to additional monitoring. This will allow quick identification of new safety information. Healthcare professionals are asked
Hereditary deafness and phenotyping in humans
 Hereditary deafness and phenotyping in humans Maria Bitner-Glindzicz Unit of Clinical and Molecular Genetics, Institute of Child Health, London, UK Correspondence to: Dr Maria Bitner-Glindzicz, Unit of
Hereditary deafness and phenotyping in humans Maria Bitner-Glindzicz Unit of Clinical and Molecular Genetics, Institute of Child Health, London, UK Correspondence to: Dr Maria Bitner-Glindzicz, Unit of
