Supplementary Information
|
|
|
- Marjory Gibson
- 5 years ago
- Views:
Transcription
1 Supplementary Information A de novo paradigm for mental retardation Lisenka ELM Vissers 1 *, Joep de Ligt 1 *, Christian Gilissen 1, Irene Janssen 1, Marloes Steehouwer 1, Petra de Vries 1, Bart van Lier 1, Peer Arts 1, Nienke Wieskamp 1, Marisol del Rosario 1, Bregje WM van Bon 1, Alexander Hoischen 1, Bert BA de Vries 1, Han G Brunner 1#, Joris A Veltman 1# 1 Department of Human Genetics - 855, Nijmegen Centre for Molecular Life Sciences and Institute for Genetic and Metabolic Disorders, Radboud University Nijmegen Medical Centre, PO Box 9101, 6500 HB Nijmegen, The Netherlands. *These authors contributed equally to this work # These authors jointly directed this work To whom correspondence should be addressed Correspondence to: Prof. dr. Han G. Brunner or Dr. ir. Joris A. Veltman Department of Human Genetics 855 Radboud University Nijmegen Medical Centre PO Box 9101, 6500 HB Nijmegen, The Netherlands h.brunner@antrg.umcn.nl j.veltman@antrg.umcn.nl Telephone: Fax:
2 Supplementary note regarding clinical patient description MR Trio 1. This boy was born a term after an uneventful pregnancy with a weight of 3850g (60 th centile) to a healthy 35-year-old mother and 38-year-old father. He was the first child of non-consanguineous parent and had a healthy sister. At 6 months hypotonia was noted and physical therapy started. Development was delayed. He could sit at 18 months and walk at 3 years and spoke 10 words only at the age of 5. At the age of 4 years his developmental level was tested which showed a 2 years delay. He has moderately severe mental retardation. On physical examination at the age of 2 years he had a normal height of 85.2 cm (15 th centile) and head circumference of 47.5 cm (15 th centile). He had mild (facial) dysmorphism consisting of prominent forehead, plagiocephalic skull, a hypotonic face with down slanting palpebral fissures and short broad hands and feet. Additional studies included a chromosome analysis by 250 K SNP array, FMR1 repeat expansion (DNA level), metabolic studies, and MRI of the brain. These revealed no abnormalities. MR Trio 2. This boy was born at 38 weeks as the first child of a non-consanguineous couple (maternal age 32 and paternal age 30 years) after an uncomplicated pregnancy with a weight of 3460g (50 th centile). He had a younger healthy sister and a nephew with a congenital heart defect. Initially he had feeding problems due to unspecified swallowing difficulties. He walked at 18 months. At the age of 5 years he used a total of 5 words only. He attended special schooling for children with severe learning difficulties. At the age of 5 years he had a height of 110 cm (15 th centile) and head circumference of 54 cm (90 th centile). He had a left epicanthic fold and mild downslanting palpebral fissures without any additional dysmorphic features. Further studies including chromosome analysis (250 K SNP array), FMR1 gene repeat expansion (DNA level), metabolic studies and MRI of the brain revealed no abnormalities. MR Trio 3. At 36 weeks pregnancy, growth retardation and unilateral dilatation of the renal pelvis were noted. After a caesarian section because of breech position this boy was born with a low birth weight of 2010g (- 2.5 SD). His non-consanguineous parents had 3 older healthy children (two girls and a boy). Parental ages at time of birth of the proband were 33 (mother) and 36 years (father). There were no neonatal problems, except for mild hypoglycemia and hyperbilirubinemia, treated with UV therapy. As a neonate feeding remained slow and difficult and the pyelectasy was confirmed with ultrasound. His development was delayed. He could sit after 1 year and walk after 2 years 9 months. He spoke his first words at 2 years and his level was 21 months at the age of 4 years 5 months, consistent with moderate mental retardation. At the age of 33 months his height was 84 cm (-3 SD) and head circumference 48.5 cm (15 th centile). He had telecanthus, down slanting palpebral fissures, high nasal bridge, and a broad mouth. The dysmorphic features were not suggestive of any specific syndrome diagnosis according to an experienced clinical geneticist (B.B.A.d.V.). Additional studies including chromosome analysis (250 K SNP array), FMR1 analysis (DNA level), metabolic studies and ophthalmological investigations revealed no abnormalities. 2
3 MR Trio 4. This girl was born after 36 weeks following an uncomplicated pregnancy with a weight of 3660g (98 th centile) as the first child of non-consanguineous parents (maternal age 32 and paternal age 31 year). Her motor development was delayed due to hypotonia. With support of physical therapy, she sat at 14 months and walked at 2 years. She spoke her first words after 2 years and at the age of 3 years 8 months she used 30 words. At that age she had a height of 100 cm (30 th centile) and a head circumference of 47.5 cm (10 th centile). She had thin blond hair, periorbital fullness and a broad nose with full tip. No syndrome diagnosis was made. Further studies including chromosome analysis (250 K SNP array), FMR1 repeat expansion (DNA level), metabolic studies and MRI investigation of the brain which revealed no abnormalities. MR Trio 5. This boy was born at 41w 2d after an uncomplicated pregnancy with a low birth weight of 2600g (3 rd centile) as the only child of a non-consanguineous couple (maternal age 32 and paternal age 35 years). His development was clearly delayed. He walked at 17 months but at the age of 6 years he still could not speak. His behavior was happy and he seldom cried. On physical examination at the age of 2y 10m he had a low normal height of 91 cm (5 th centile) and his head circumference was 49.2 cm (25 th centile). He had thin fair hair, upward slanting palpebral fissures, epicanthic folds, almond shaped eyes, proximal implant of the thumbs and a scrotal raphe. An initial clinical suspicion of Angelman syndrome led to methylation study of the 15q11-q13 region and mutation analysis of UBE3A. These revealed no aberrations. Further studies including chromosome analysis (250 K SNP array), FMR1 repeat expansion (DNA level) and metabolic studies and ophthalmological investigations revealed no abnormalities. MR Trio 6. This boy was born at 38w 3d after an uncomplicated pregnancy by cesarean section with a weight of 3000g (40 th centile) as the second child of a non-consanguineous couple (maternal age 29 and paternal age 27 years). His development was delayed. He sat at 15 months, walked at 22 months and spoke his first words after 21 months and at the age of 3 years used 20 words only. At the age of 21 months his height was 80.5 cm (5 th centile) and head circumference 47.3 cm (15 th centile). He had no evident facial dysmorphisms. Further studies including chromosome analysis (250 K SNP array), FMR1 repeat expansion (DNA level), metabolic studies and MRI investigation of the brain revealed no abnormalities. MR Trio 7. This 19-year-old male was the second child of non-consanguineous parents and born at 40 weeks after an uncomplicated pregnancy with a weight of 3500g (50 th centile). Parental ages at time of birth were 28 and 29 years for mother and father, respectively. Early motor development was slow but within normal range. Speech development was severely delayed, he spoke his first words at the age of 4 years. He attended special school and his developmental level was 3-3,5 years at the age of 19 years. He is severely retarded. He showed variable aggressive behavior. On physical examination at the age of 19 years, a low-normal height of 172 cm (5 th centile) and normal head circumference of 56 cm (15 th centile) was noted. He had minor facial dysmorphisms; upward slanting palpebral fissures, slightly everted ears, thin upper lip with full everted lower lip without additional physical features. Further studies including 3
4 chromosome analysis (250 K microarray), FMR1 expansion (chromosome level), EHMT1 gene and metabolic studies revealed no abnormalities. MR Trio 8. This girl was born at 36w 5d after an uncomplicated pregnancy by cesarean section due to breech position with a weight of 2800g (50 th centile) as the only child of a non-consanguineous couple (maternal age 32 and paternal age 29 years). Her motor development was delayed due to hypotonia, she sat at 19 months. MRI of the brain revealed mild delay of myelinization at 10 months. A muscle biopsy at the age of 20 months did not show any abnormalities. At the age of 4 ½ months she did not speak any words. At age 4 years epilepsy was diagnosed. On physical examination at the age of 11 months she had a normal height of 70 cm (10 th centile) and head circumference of 46.3 cm (60 th centile). She had a high forehead, upward slanting palpebral fissures, mild hypertelorism, epicantical folds, low nasal bridge and full lips with everted lower lips. A general, predominantly axial, hypotonia was present. Further studies, including chromosome analysis (250 K SNP array), DNA-analysis of FMR1, myotonic dystrophy, Prader Willi syndrome, as well as metabolic studies and ophthalmological investigation, all revealed no abnormalities. MR Trio 9. This boy was born at 39w 1d after an uncomplicated pregnancy with a weight of 3520g (50 th centile) as the second child of a non-consanguineous couple (maternal age 31 and paternal age 37 years). There were neonatal feeding difficulties. His development was delayed. He sat at 12 months, walked at 18 months and spoke his first words at 2 years. At the age of 3 2/12 years he used 5-10 words only. He was moderately mentally retarded. On physical examination at that age he had a normal height of 103 cm (75 th centile) and head circumference of 50.5 cm (50 th centile). He had mild (facial) dysmorphism consisting of upward slanting palpebral fissures, short palpebral fissures, flattened upper helices and broad nose with full tip. Further studies including chromosome analysis (250 K SNP array), FMR1 repeat expansion (DNA level) and metabolic studies revealed no abnormalities. MR Trio 10. This boy was born after a normal gestation at 39w 6d with a weight of 3270g (25 th centile) as the second child of non-consanguineous parents (maternal age 29 and paternal age 31 years). At 6 months, delayed development was noted., He sat at months, walked at months and at the age of 4 years he only used 4 words and his developmental level at that age was months. On physical examination at the age of 2 4/12 years he had a height of 89.5 cm (25 th centile) and head circumference of 49 cm (30 th centile). He had a metopic ridge, hypotelorism (ICD 2cm, < 3 rd centile), mild ptosis, short palpebral fissures and a flat palate. He had a broad, uncertain gait. Further studies, including G-banded chromosomes and FMR1 repeat expansion analysis (DNA level) were normal. Additional metabolic and chromosome analysis (250K SNP array) revealed no abnormalities. 4
5 Supplementary Tables Supplementary Table 1. Overview of exome-sequencing performance reads sequenced reads mapped bps mapped % mapping to targeted sequence* MR trio 1 Mother 75,539,970 62,421,598 2,994,163, Father 60,482,344 49,757,641 2,359,196, Patient 83,750,821 63,968,695 2,993,659, MR trio 2 Mother 61,213,042 45,635,581 2,129,184, Father 81,822,660 64,920,388 3,101,222, Patient 81,274,402 63,413,997 2,998,226, MR trio 3 Mother 91,218,082 65,444,907 3,111,727, Father 90,482,575 66,637,781 3,174,885, Patient 89,089,332 63,253,491 2,987,977, MR trio 4 Mother 89,715,373 63,797,713 3,045,658, Father 80,613,693 64,548,447 3,100,745, Patient 84,917,415 65,900,711 3,147,306, MR trio 5 Mother 95,997,554 75,047,477 3,635,298, Father 85,743,467 61,089,261 2,939,733, Patient 72,400,831 55,557,595 2,658,661, MR trio 6 Mother 77,312,957 66,715,554 3,208,045, Father 82,262,575 63,187,587 3,006,761, Patient 74,889,401 57,633,832 2,769,859, MR trio 7 Mother 88,884,506 67,665,146 3,207,116, Father 93,174,731 72,970,692 3,483,792, Patient 76,030,490 58,908,643 2,789,555, MR trio 8 Mother 82,506,855 66,109,620 3,185,522, Father 55,116,325 45,653,877 2,163,335, Patient 81,579,330 63,654,731 3,047,698,
6 MR trio 9 Mother 88,622,619 69,950,138 3,363,461, Father 95,568,766 82,036,334 3,985,154, Patient 83,296,156 62,687,172 3,032,856, MR trio 10 Mother 90,342,631 73,134,022 3,537,828, Father 95,285,830 79,698,200 3,875,693, Patient 87,139,147 67,783,275 3,255,909, Over all 30 individuals Median 83,523,489 64,258,571 3,074,222, Min 55,116,325 45,635,581 2,129,184, Max 95,997,554 82,036,334 3,985,154, *Defined as the sum of bases mapping to the SureSelect bait and one read length (50 bp) away from the Sure Select bait. 6
7 Supplementary table 2: All potential de novo variants per trio followed up by Sanger sequencing The columns represent and should be interpreted as follows: Chromosome: chromosome Start: genomic start position of variant base(s), mapped on hg18 End: genomic end position of variant base(s), mapped on hg18 Reference: wildtype sequence for the genomic position Variant: variant sequence for the genomic position Reads: number of reads for the genomic position (e.g. total coverage) Variant reads: number of reads showing the variant base(s) Unique starts: number of variant reads with an unique start position % variation: the percentage of reads that show the variant (based on variant reads/reads). Low covered variants (<15%) are hereby distinguished from heterozygous (15-80%) and homozygous/hemizygous mutations (>80%). Gene name: gene in which the variant occurs Gene ID: NM used to annotate the variants Total exons: total number of exons known to be included in the gene ID used Variant in exon: exon in which the variant occurred Reference aa: wildtype amino acid for this genomic position Variant aa: variant amino acid ( * denotes premature stop) PhyloP score: conservation score for the genomic nucleotide the variant starts Grantham score: conservation score indicating the physiochemical changes due to amino acid change Cells with red shading and text in bold represent validated and de novo mutations, in gray shading and gray text represent validated but inherited variants, and cells without shading and gray text represent variants that could not be validated in the patients, hence parents were not tested. Notes: For indels, unique start sites cannot be determined and are therefore not presented For frameshift variants, the first two amino acid changes are provided Grantham scores for nonsense and frameshift variations are not provided. Nonsense variants do not have a Grantham score. Grantham scores for frameshift variants are not representative of the change, as in many cases a premature stop occurs downstream of the mutation. 7
8 Supplementary Table 2 continued Chromosome MR trio 1 Start position End position Reference Variant Reads Variant reads Unique starts % Variation Gene name Gene id Total exons Variation in exon Ref erence aa Variant aa phylop Score Grantham Score chr A C DYNC1H1 NM_ H P chr T G ALG5 NM_ K T chr G A LEMD2 NM_ A V chr G A ZNF599 NM_ L F MR trio 2 chrx C T RAB39B NM_ W * chr G A CLCN3 NM_ C Y chr CTCTCTCTCAC CTCTCTCAC MAGI1 NM_ E D chr A G KIT NM_ N D chr7 # C T KCNH2 NM_ D N chr A C ZNF564 NM_ I S chr T C TCHH NM_ K E # the mother of the patient did not have coverage for this variant, as such it was validated it appeared to be maternally inherited after Sanger sequencing MR trio 3 chr G T YY1 NM_ D Y chr A G CNNM2 NM_ E G chr G A BPIL3 NM_ R H MR trio 4 chr CG CACG PRPF8 NM_ G A chr T C SSX2IP NM_ K R chr TT TTT GATAD2B NM_ N K chrx C G MAGEE1 NM_ P A chr T C PGA5 NM_ V A chr CTCCTCCT CTCCTCCTCCT AHNAK NM_ T RX chr TC ERRFI1 NM_ R E MR trio 5 chr A G HS3ST2 NM_ I V chr A C DEAF1 NM_ I S chr TGAAAAAAAAA TGAAAAAAAA ABCB4 NM_ S Q chr CTGAGTC CTGAGTCGTC EPHX2 NM_ N NX chry G A PCDH11Y NM_ V M chr ACTTTTTTTTCT ACTTTTTTTCT KRR1 NM_ K S chr G T ALMS1 NM_ A S
9 MR trio 6 chr C T CIC NM_ R W chr TTAT TTATTAT HOXC13 NM_ L LX MR trio 7 chr CAGT CAGTGAC CCDC36 NM_ L PX chr T C MOCOS NM_ F S MR trio 8 chr G A INPPL1 NM_ E K chr ACTGTGCC ACTGCC SYNGAP1 NM_ TV TA chr GCCTCCTCTC GCCTCTC TRIOBP NM_ AS ASX chr C T OR4A5 NM_ E K chr CTATTGA CTTTGA EMR3 NM_ N K chr T C ZNF540 NM_ F L MR trio 9 chr T TT KCNJ12 NM_ V F chr GGGGGC GGGGGGC CELA1 NM_ L R chr GGGGGGGA GGGGGGGGA ACTR8 NM_ chr TGTGTTTTTTTT TGTGTTTTTTT MYH10 NM_ chr GCTTTTTTTTTG GCTTTTTTTTG KCNMA1 NM_ K S chr T C RGS22 NM_ T A MR trio 10 chr G A LRBA NM_ R W chr T G IPO13 NM_ F C chr A G DNAH7 NM_ M T chr C A TMEM150B NM_ W C chr TGTGTGTGGG TGTGTGGG WBSCR27 NM_ chr AGGAGGTGGG AGGTGGG PRB1 NM_ PP PGX chr C T HIST1H1A NM_ A T Validated, de novo Validated, not de novo Not validated 9
10 Supplementary Figures Supplementary Figure 1: Coverage plots of all 30 individuals Figure legend Coverage for all exons targeted by enrichment was evaluated. The median coverage for all 30 individuals was 42-fold, with on average 90% of all targets covered at least 10-fold. The numbers in the figure legends refer to the corresponding MR trios; M: Mother; F: Father; C: Child 10
11 Supplementary Figure 2: Variant calling per trio 30,000 non genic intronic exonic 25,000 20,000 Number of variants 15,000 10,000 5,000 0 Figure legend Schematic representation of number of variants identified per individual, per trio. On average, 22,272 variants are identified per individual, of which 19,212 are located in genes (intronic and exonic together), and 12,341 are located within the exons and/or canonical splice-sites. The number on the x-axis refer to the corresponding MR trios, whereas M to Mother, F to Father and C to child. 11
12 Supplementary Figure 3: IGV browser overview for validated de novo mutations a b c 12
13 d e f 13
14 g h i 14
15 j Figure Legend IGV browser plots for the nine de novo and one X-linked inherited mutations identified in the patients with mental retardation. (a) DYNC1H1 - trio 1, (b) ZNF599 trio 1, (c) RAB39B trio 2; (d) YY1 trio 3; (e) BPIL3 trio 3; (f) PGA5 trio 4; (g) DEAF1 trio 5; (h) CIC trio 6; (i) SYNGAP1 trio 8; (j) JARID1C trio 10. For each overview, top panel show chromosome with mutation location indicated by a red box. The second panel from the top shows the relative coverage per base pair for patient, mother and father. Colored coverage columns indicate the location of the mutation with the relative height of the color representing the percentage of variant reads observed. Middle panels show the individual reads (zoom-in to ~10-15 reads of all reads) with bases deviant from the wildtype sequence highlighted in colors. Lower panel shows wildtype sequence and translated amino acids. 15
16 Supplementary Figure 4: Sanger validation of de novo mutations a Patient Father Mother b Patient Father Mother c Patient Father Mother d Patient Father Mother 16
17 e Patient Father Mother f Patient Father Mother g Patient Father Mother Figure legend Partial electropherograms showing the de novo occurrence for the nine mutations identified. Mutations are highlighted in yellow. (a) trio 1, left panel DYNC1H1 with mutation c.11465a>c leading to p.his3822pro and right panel ZNF599 with mutation c.532c>t leading to p.leu187phe. (b) trio 2, RAB39B with mutation c.557g>a leading to a premature stop p.trp186x. (c) trio 3, left panel YY1 with mutation c.1138g>t leading to p.asp380tyr and right panel BPIL3 with mutation c.887g>a leading to p.arg269his. (d) trio 4 with mutation c. 1058T>C in PGA5, leading to Val353Ala. (e) trio 5, with mutation c.683t>g in DEAF1 leading to p.ile228ser. (f) trio 6 with mutation c.1474c>t in CIC leading to p.arg492trp and (g) trio 8, with mutation c.998_999del in SYNGAP1 leading to p.val333alsfs70x. 17
18 Supplementary Figure 5: Distribution of PhyloP and Grantham scores for dbsnp, HGMD and the de novo mutations identified in this study a dbsnp HGMD b 200 Relative frequency Grantham score 'benign' 'pathogenic' PhyloP score PhyloP score c d min max 18
19 Figure legends (a) Relative distribution of PhyloP scores for non-synonymous in dbsnp (green surface) and HGMD (red surface) to represent evolutionary scores observed for benign and pathogenic variants. (b) Evolutionary conservations for the 3 functionally benign (green diamonds) and 4 functionally pathogenic de novo mutations, as well as the X-linked inherited mutation (red diamonds). This analysis was further extended to analyse these combined scores for all nonsynonymous variants in dbsnp and HGMD to visualize the probability for the de novo variants to be pathogenic or benign. Heat maps for distribution and frequency for the combined PhyloP and Grantham scores in dbsnp (c) and HGMD (d). Functionally benign and pathogenic de novo mutations, as well as the inherited JARID1C mutation, are represented by squares and triangles respectively. Benign de novo variants show a better overlap with dbsnp as indicated by the their location mostly restricted to dark green areas in c and not in d, whereas the pathogenic de novo variants show a better overlap with HGMD, as indicated by their location mostly being in dark green areas in d and not in c. Statistical probability scores for these analyses are provided in Table 2. Note: Two de novo mutations, a nonsense and frameshift mutation, identified in our patient cohort are not depicted because such mutations do not have representative Grantham scores. Their functional impact on the protein level will however be most severe and both are located in known mental retardation genes. 19
20 References to Supplementary Information 1. Marth,G.T. et al. A general approach to single-nucleotide polymorphism discovery. Nat Genet. 23, (1999). 2. Ng,S.B. et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 461, (2009). 3. Pushkarev,D., Neff,N.F., & Quake,S.R. Single-molecule sequencing of an individual human genome. Nat Biotechnol. 27, (2009). 4. Wang,J. et al. The diploid genome sequence of an Asian individual. Nature 456, (2008). 5. Venables, W.N. and Ripley, B D. Modern Applied Statistics with S. Springer, Fourth edition (2002) Lilliefors, H. On the Kolmogorov Smirnov test for normality with mean and variance unknown", Journal of the American Statistical Association 62, (1967). Web resources For variant visualization we used the IGV browser developed at: For in-house variant database: For genome browser annotations: For variant effect of identified mutations:
Van test naar diagnose naar
 Van test naar diagnose naar V therapie op maat Marjolein Kriek, LUMC Joris Veltman, RUNMC Exome diagnostics in genetically heterogeneous disease Joris Veltman, PhD Department of Human Genetics Radboud
Van test naar diagnose naar V therapie op maat Marjolein Kriek, LUMC Joris Veltman, RUNMC Exome diagnostics in genetically heterogeneous disease Joris Veltman, PhD Department of Human Genetics Radboud
Variant Detection & Interpretation in a diagnostic context. Christian Gilissen
 Variant Detection & Interpretation in a diagnostic context Christian Gilissen c.gilissen@gen.umcn.nl 28-05-2013 So far Sequencing Johan den Dunnen Marja Jakobs Ewart de Bruijn Mapping Victor Guryev Variant
Variant Detection & Interpretation in a diagnostic context Christian Gilissen c.gilissen@gen.umcn.nl 28-05-2013 So far Sequencing Johan den Dunnen Marja Jakobs Ewart de Bruijn Mapping Victor Guryev Variant
CHAPTER IV RESULTS Microcephaly General description
 47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
Dysmorphology. Sue White. Diagnostic Dysmorphology, Aase. Victorian Clinical Genetics Services
 Dysmorphology Sue White www.rch.unimelb.edu.au/nets/handbook Diagnostic Dysmorphology, Aase Dysmorphology Assessment Algorithm no Are the features familial? yes Recognised syndrome yes no AD/XL syndrome
Dysmorphology Sue White www.rch.unimelb.edu.au/nets/handbook Diagnostic Dysmorphology, Aase Dysmorphology Assessment Algorithm no Are the features familial? yes Recognised syndrome yes no AD/XL syndrome
Supplemental Data: Detailed Characteristics of Patients with MKRN3. Patient 1 was born after an uneventful pregnancy. She presented in our
 1 2 Supplemental Data: Detailed Characteristics of Patients with MKRN3 Mutations 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Patient 1 was born after an uneventful pregnancy. She presented
1 2 Supplemental Data: Detailed Characteristics of Patients with MKRN3 Mutations 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Patient 1 was born after an uneventful pregnancy. She presented
Supplementary Clinical Information
 Supplementary Clinical Information Patient 1 This female was born as the second child of a twin after a pregnancy duration of 32 weeks. The birth was otherwise uncomplicated and her twin brother was healthy.
Supplementary Clinical Information Patient 1 This female was born as the second child of a twin after a pregnancy duration of 32 weeks. The birth was otherwise uncomplicated and her twin brother was healthy.
Identification of a novel in-frame de novo mutation in SPTAN1 in intellectual disability and pontocerebellar atrophy
 Hamdan et al., Identification of a novel in-frame de novo mutation in SPTAN1 in intellectual disability and pontocerebellar atrophy Supplementary Information Gene screening and bioinformatics PCR primers
Hamdan et al., Identification of a novel in-frame de novo mutation in SPTAN1 in intellectual disability and pontocerebellar atrophy Supplementary Information Gene screening and bioinformatics PCR primers
Faravareh Khordadpoor (PhD in molecular genetics) 1- Tehran Medical Genetics Laboratory 2- Science and research branch, Islamic Azad University
 Faravareh Khordadpoor (PhD in molecular genetics) 1- Tehran Medical Genetics Laboratory 2- Science and research branch, Islamic Azad University 1395 21 مشاوره ژنتیک و نقش آن در پیش گیری از معلولیت ها 20
Faravareh Khordadpoor (PhD in molecular genetics) 1- Tehran Medical Genetics Laboratory 2- Science and research branch, Islamic Azad University 1395 21 مشاوره ژنتیک و نقش آن در پیش گیری از معلولیت ها 20
Seven cases of intellectual disability analysed by genomewide SNP analysis. Rodney J. Scott
 Seven cases of intellectual disability analysed by genomewide SNP analysis Rodney J. Scott Ability to interrogate Genomic Material ~1930 Crude analyses 2012 Highly precise analyses A (very) brief history
Seven cases of intellectual disability analysed by genomewide SNP analysis Rodney J. Scott Ability to interrogate Genomic Material ~1930 Crude analyses 2012 Highly precise analyses A (very) brief history
Approach to the Genetic Diagnosis of Neurological Disorders
 Approach to the Genetic Diagnosis of Neurological Disorders Dr Wendy Jones MBBS MRCP Great Ormond Street Hospital for Children National Hospital for Neurology and Neurosurgery What is a genetic diagnosis?
Approach to the Genetic Diagnosis of Neurological Disorders Dr Wendy Jones MBBS MRCP Great Ormond Street Hospital for Children National Hospital for Neurology and Neurosurgery What is a genetic diagnosis?
Concurrent Practical Session ACMG Classification
 Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation
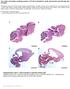 The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation Anita Becker-Heck#, Irene Zohn#, Noriko Okabe#, Andrew Pollock#, Kari Baker Lenhart,
The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation Anita Becker-Heck#, Irene Zohn#, Noriko Okabe#, Andrew Pollock#, Kari Baker Lenhart,
CHAPTER IV RESULTS. The goal of this study was to identify the underlying genetic defect in patients with MR
 CHAPTER IV RESULTS The goal of this study was to identify the underlying genetic defect in patients with MR and epilepsy. Mutation analysis from the syndromic patients were performed, from the non syndromic
CHAPTER IV RESULTS The goal of this study was to identify the underlying genetic defect in patients with MR and epilepsy. Mutation analysis from the syndromic patients were performed, from the non syndromic
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier/Additional Provider
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier/Additional Provider TEST DISEASE/CONDITION POPULATION TRIAD Submitting laboratory: Manchester RGC Approved: September 2013
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier/Additional Provider TEST DISEASE/CONDITION POPULATION TRIAD Submitting laboratory: Manchester RGC Approved: September 2013
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016
 Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
The Genetics of Parenthood
 The Genetics of Parenthood Introduction Why do people, even closely related people, look slightly different from each other? The reason for these differences in physical characteristics (called phenotype)
The Genetics of Parenthood Introduction Why do people, even closely related people, look slightly different from each other? The reason for these differences in physical characteristics (called phenotype)
MutationTaster & RegulationSpotter
 MutationTaster & RegulationSpotter Pathogenicity Prediction of Sequence Variants: Past, Present and Future Dr. rer. nat. Jana Marie Schwarz Klinik für Pädiatrie m. S. Neurologie Exzellenzcluster NeuroCure
MutationTaster & RegulationSpotter Pathogenicity Prediction of Sequence Variants: Past, Present and Future Dr. rer. nat. Jana Marie Schwarz Klinik für Pädiatrie m. S. Neurologie Exzellenzcluster NeuroCure
Clinical Genetics. Functional validation in a diagnostic context. Robert Hofstra. Leading the way in genetic issues
 Clinical Genetics Leading the way in genetic issues Functional validation in a diagnostic context Robert Hofstra Clinical Genetics Leading the way in genetic issues Future application of exome sequencing
Clinical Genetics Leading the way in genetic issues Functional validation in a diagnostic context Robert Hofstra Clinical Genetics Leading the way in genetic issues Future application of exome sequencing
Exploding Genetic Knowledge in Developmental Disabilities. Disclosures. The Genetic Principle
 Exploding Genetic Knowledge in Developmental Disabilities How to acquire the data and how to make use of it Elliott H. Sherr MD PhD Professor of Neurology & Pediatrics UCSF Disclosures InVitae: clinical
Exploding Genetic Knowledge in Developmental Disabilities How to acquire the data and how to make use of it Elliott H. Sherr MD PhD Professor of Neurology & Pediatrics UCSF Disclosures InVitae: clinical
Objectives. Genetics and Rett syndrome: As easy as apple pie! Chromosome to gene to protein
 Genetics and Rett syndrome: As easy as apple pie! Victoria Mok Siu M.D., FRCPC, FCCMG ORSA conference Ottawa April 24, 2016 Objectives Review chromosomes and genes Understand s Explore the reasons behind
Genetics and Rett syndrome: As easy as apple pie! Victoria Mok Siu M.D., FRCPC, FCCMG ORSA conference Ottawa April 24, 2016 Objectives Review chromosomes and genes Understand s Explore the reasons behind
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
UNIT IX: GENETIC DISORDERS
 UNIT IX: GENETIC DISORDERS Younas Masih Lecturer New Life College Of Nursing Karachi 3/4/2016 1 Objectives By the end of this session the Learners will be able to, 1. Know the basic terms related genetics
UNIT IX: GENETIC DISORDERS Younas Masih Lecturer New Life College Of Nursing Karachi 3/4/2016 1 Objectives By the end of this session the Learners will be able to, 1. Know the basic terms related genetics
Identifying Mutations Responsible for Rare Disorders Using New Technologies
 Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Investigating rare diseases with Agilent NGS solutions
 Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Case 1B. 46,XY,-14,+t(14;21)
 Case 1B 46,XY,-14,+t(14;21) G-banded Chromosome telomere centromere G-dark bands AT-rich few genes G-pale bands GC-rich many genes telomere ideograms ideograms Conventional (light microscopy) p = short
Case 1B 46,XY,-14,+t(14;21) G-banded Chromosome telomere centromere G-dark bands AT-rich few genes G-pale bands GC-rich many genes telomere ideograms ideograms Conventional (light microscopy) p = short
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
RACP Congress 2017 Genetics of Intellectual Disability and Autism: Past Present and Future 9 th May 2017
 RACP Congress 2017 Genetics of Intellectual Disability and Autism: Past Present and Future 9 th May 2017 Why causation? Explanation for family Prognosis Recurrence risk and reproductive options Guide medical
RACP Congress 2017 Genetics of Intellectual Disability and Autism: Past Present and Future 9 th May 2017 Why causation? Explanation for family Prognosis Recurrence risk and reproductive options Guide medical
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13908 Supplementary Tables Supplementary Table 1: Families in this study (.xlsx) All families included in the study are listed. For each family, we show: the genders of the probands and
doi:10.1038/nature13908 Supplementary Tables Supplementary Table 1: Families in this study (.xlsx) All families included in the study are listed. For each family, we show: the genders of the probands and
This fact sheet describes the condition Fragile X and includes a discussion of the symptoms, causes and available testing.
 11111 Fact Sheet 54 FRAGILE X SYNDROME This fact sheet describes the condition Fragile X and includes a discussion of the symptoms, causes and available testing. In summary Fragile X is a condition caused
11111 Fact Sheet 54 FRAGILE X SYNDROME This fact sheet describes the condition Fragile X and includes a discussion of the symptoms, causes and available testing. In summary Fragile X is a condition caused
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY.
 SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: LOSS,
SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: LOSS,
Understanding The Genetics of Diamond Blackfan Anemia
 Understanding The Genetics of Diamond Blackfan Anemia Jason Farrar, MD jefarrar@ About Me Assistant Professor of Pediatrics at University of Arkansas for Medical Sciences & Arkansas Children s Hospital
Understanding The Genetics of Diamond Blackfan Anemia Jason Farrar, MD jefarrar@ About Me Assistant Professor of Pediatrics at University of Arkansas for Medical Sciences & Arkansas Children s Hospital
Klinefelter syndrome ( 47, XXY )
 Sex Chromosome Abnormalities, Sex Chromosome Aneuploidy It has been estimated that, overall, approximately one in 400 infants have some form of sex chromosome aneuploidy. A thorough discussion of sex chromosomes
Sex Chromosome Abnormalities, Sex Chromosome Aneuploidy It has been estimated that, overall, approximately one in 400 infants have some form of sex chromosome aneuploidy. A thorough discussion of sex chromosomes
Approach to Mental Retardation and Developmental Delay. SR Ghaffari MSc MD PhD
 Approach to Mental Retardation and Developmental Delay SR Ghaffari MSc MD PhD Introduction Objectives Definition of MR and DD Classification Epidemiology (prevalence, recurrence risk, ) Etiology Importance
Approach to Mental Retardation and Developmental Delay SR Ghaffari MSc MD PhD Introduction Objectives Definition of MR and DD Classification Epidemiology (prevalence, recurrence risk, ) Etiology Importance
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier/Additional Provider
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier/Additional Provider TEST DISORDER/CONDITION POPULATION TRIAD Submitting laboratory: Exeter RGC Approved: Sept 2013 1. Disorder/condition
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier/Additional Provider TEST DISORDER/CONDITION POPULATION TRIAD Submitting laboratory: Exeter RGC Approved: Sept 2013 1. Disorder/condition
Neonatal Hypotonia Guideline Prepared by Dan Birnbaum MD August 27, 2012
 Neonatal Hypotonia Guideline Prepared by Dan Birnbaum MD August 27, 2012 Hypotonia: reduced tension or resistance to range of motion Localization can be central (brain), peripheral (spinal cord, nerve,
Neonatal Hypotonia Guideline Prepared by Dan Birnbaum MD August 27, 2012 Hypotonia: reduced tension or resistance to range of motion Localization can be central (brain), peripheral (spinal cord, nerve,
6/12/2018. Disclosures. Clinical Genomics The CLIA Lab Perspective. Outline. COH HopeSeq Heme Panels
 Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Lab Activity 36. Principles of Heredity. Portland Community College BI 233
 Lab Activity 36 Principles of Heredity Portland Community College BI 233 Terminology of Chromosomes Homologous chromosomes: A pair, of which you get one from mom, and one from dad. Example: the pair of
Lab Activity 36 Principles of Heredity Portland Community College BI 233 Terminology of Chromosomes Homologous chromosomes: A pair, of which you get one from mom, and one from dad. Example: the pair of
STUDENT WORKSHEET. The Genetics of Parenthood Data Sheet. Parents and CHILD'S GENOTYPE ALLELE FROM DAD. H h I i J j K k.
 STUDENT WORKSHEET The Genetics of Parenthood Data Sheet Parents and Child's gender Child's name Fill in data table as you determine each trait described in the Guidebook. Do not simply flip the coin for
STUDENT WORKSHEET The Genetics of Parenthood Data Sheet Parents and Child's gender Child's name Fill in data table as you determine each trait described in the Guidebook. Do not simply flip the coin for
CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE. Dr. Bahar Naghavi
 2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY.
 SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: GAIN,
SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: GAIN,
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE
 CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
variant led to a premature stop codon p.k316* which resulted in nonsense-mediated mrna decay. Although the exact function of the C19L1 is still
 157 Neurological disorders primarily affect and impair the functioning of the brain and/or neurological system. Structural, electrical or metabolic abnormalities in the brain or neurological system can
157 Neurological disorders primarily affect and impair the functioning of the brain and/or neurological system. Structural, electrical or metabolic abnormalities in the brain or neurological system can
Neonatal Hypotonia. Encephalopathy acute No encephalopathy. Neurology Chapter of IAP
 The floppy infant assumes a frog legged position. On ventral suspension, the baby can not maintain limb posture against gravity and assumes the position of a rag doll. Encephalopathy acute No encephalopathy
The floppy infant assumes a frog legged position. On ventral suspension, the baby can not maintain limb posture against gravity and assumes the position of a rag doll. Encephalopathy acute No encephalopathy
CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY
 CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY More genetic information requires cutting-edge interpretation techniques Whole Exome Sequencing For some patients, the combination of symptoms does not allow
CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY More genetic information requires cutting-edge interpretation techniques Whole Exome Sequencing For some patients, the combination of symptoms does not allow
How many disease-causing variants in a normal person? Matthew Hurles
 How many disease-causing variants in a normal person? Matthew Hurles Summary What is in a genome? What is normal? Depends on age What is a disease-causing variant? Different classes of variation Final
How many disease-causing variants in a normal person? Matthew Hurles Summary What is in a genome? What is normal? Depends on age What is a disease-causing variant? Different classes of variation Final
Pathophysiology of the Phenylketonuria
 Problem 4. Pathophysiology of the Phenylketonuria Readings for this problem are found on pages: 79-82, 84, 85-6, 945-6 and 1019 of your Pathophysiology (5 th edition) textbook. (This problem was based
Problem 4. Pathophysiology of the Phenylketonuria Readings for this problem are found on pages: 79-82, 84, 85-6, 945-6 and 1019 of your Pathophysiology (5 th edition) textbook. (This problem was based
Dysmorphology And The Paediatric Eye. Jill Clayton-Smith Manchester Centre For Genomic Medicine
 Dysmorphology And The Paediatric Eye Jill Clayton-Smith Manchester Centre For Genomic Medicine Why Make A Syndrome Diagnosis? Why did it happen? What does the future hold? How can you treat/manage it?
Dysmorphology And The Paediatric Eye Jill Clayton-Smith Manchester Centre For Genomic Medicine Why Make A Syndrome Diagnosis? Why did it happen? What does the future hold? How can you treat/manage it?
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Leber congenital amaurosis OMIM number for disease 204000 Disease alternative
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Leber congenital amaurosis OMIM number for disease 204000 Disease alternative
Human Genetic Disorders
 Human Genetic Disorders HOMOLOGOUS CHROMOSOMES Human somatic cells have 23 pairs of homologous chromosomes 23 are inherited from the mother and 23 from the father HOMOLOGOUS CHROMOSOMES Autosomes o Are
Human Genetic Disorders HOMOLOGOUS CHROMOSOMES Human somatic cells have 23 pairs of homologous chromosomes 23 are inherited from the mother and 23 from the father HOMOLOGOUS CHROMOSOMES Autosomes o Are
The Floppy Baby. Clare Betteridge
 The Floppy Baby Clare Betteridge The floppy baby Identification Evaluation Investigation Diagnosis Examples What is a floppy baby? Elbows and knees loosely extended. Head control is usually poor or absent.
The Floppy Baby Clare Betteridge The floppy baby Identification Evaluation Investigation Diagnosis Examples What is a floppy baby? Elbows and knees loosely extended. Head control is usually poor or absent.
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13394 Genome sequencing identifies major causes of severe intellectual disability Table of Contents Participants...4 Patient recruitment... 4 Patient selection... 4 Cohort representativeness...
doi:10.1038/nature13394 Genome sequencing identifies major causes of severe intellectual disability Table of Contents Participants...4 Patient recruitment... 4 Patient selection... 4 Cohort representativeness...
Nature Genetics: doi: /ng Supplementary Figure 1. PCA for ancestry in SNV data.
 Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Genetics and Genomics: Applications to Developmental Disability
 Tuesday, 12:30 2:00, B1 Objective: Genetics and Genomics: Applications to Developmental Disability Helga Toriello 616-234-2712 toriello@msu.edu Identify advances in clinical assessment and management of
Tuesday, 12:30 2:00, B1 Objective: Genetics and Genomics: Applications to Developmental Disability Helga Toriello 616-234-2712 toriello@msu.edu Identify advances in clinical assessment and management of
'Translating massively parallel sequencing into clinical diagnostics.
 'Translating massively parallel sequencing into clinical diagnostics. SA Pathology at the Women s and and The University of Adelaide Adelaide, Australia MPS is revolutionising genetics & biology - >100
'Translating massively parallel sequencing into clinical diagnostics. SA Pathology at the Women s and and The University of Adelaide Adelaide, Australia MPS is revolutionising genetics & biology - >100
Analysis with SureCall 2.1
 Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Information leaflet for patients and families. Uniparental Disomy (UPD)
 Information leaflet for patients and families Uniparental Disomy (UPD) What are genes and chromosomes? Our genes are the unique set of instructions inside every cell of our body which make each of us an
Information leaflet for patients and families Uniparental Disomy (UPD) What are genes and chromosomes? Our genes are the unique set of instructions inside every cell of our body which make each of us an
Genetic Testing and Analysis. (858) MRN: Specimen: Saliva Received: 07/26/2016 GENETIC ANALYSIS REPORT
 GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
When to suspect Prader Willi Syndrome and how to diagnose it?
 When to suspect Prader Willi Syndrome and how to diagnose it? Dr Chirita-Emandi Adela Dr Dobrescu Andreea Victor Babes University of Medicine and Pahrmacy Timisoara Emergency Hospital for Children Louis
When to suspect Prader Willi Syndrome and how to diagnose it? Dr Chirita-Emandi Adela Dr Dobrescu Andreea Victor Babes University of Medicine and Pahrmacy Timisoara Emergency Hospital for Children Louis
Improving genomic diagnoses through accurate, specific phenotype information
 Improving genomic diagnoses through accurate, specific phenotype information Lisa Ewans Clinical Geneticist, RPAH, Sydney PhD student in genomics, KCCG, Garvan; UNSW https://vlab.org Overview Phenotype
Improving genomic diagnoses through accurate, specific phenotype information Lisa Ewans Clinical Geneticist, RPAH, Sydney PhD student in genomics, KCCG, Garvan; UNSW https://vlab.org Overview Phenotype
Welcome to the Genetic Code: An Overview of Basic Genetics. October 24, :00pm 3:00pm
 Welcome to the Genetic Code: An Overview of Basic Genetics October 24, 2016 12:00pm 3:00pm Course Schedule 12:00 pm 2:00 pm Principles of Mendelian Genetics Introduction to Genetics of Complex Disease
Welcome to the Genetic Code: An Overview of Basic Genetics October 24, 2016 12:00pm 3:00pm Course Schedule 12:00 pm 2:00 pm Principles of Mendelian Genetics Introduction to Genetics of Complex Disease
Investigation of chromosomal abnormalities and microdeletions in 50 patients with multiple congenital anomalies
 Investigation of chromosomal abnormalities and microdeletions in 50 patients with multiple congenital anomalies Akbar Mohammadzadeh MD, PhD candidate in Medical Genetics Genetics Research Center University
Investigation of chromosomal abnormalities and microdeletions in 50 patients with multiple congenital anomalies Akbar Mohammadzadeh MD, PhD candidate in Medical Genetics Genetics Research Center University
CANCER GENETICS PROVIDER SURVEY
 Dear Participant, Previously you agreed to participate in an evaluation of an education program we developed for primary care providers on the topic of cancer genetics. This is an IRB-approved, CDCfunded
Dear Participant, Previously you agreed to participate in an evaluation of an education program we developed for primary care providers on the topic of cancer genetics. This is an IRB-approved, CDCfunded
PROGRESS: Beginning to Understand the Genetic Predisposition to PSC
 PROGRESS: Beginning to Understand the Genetic Predisposition to PSC Konstantinos N. Lazaridis, MD Associate Professor of Medicine Division of Gastroenterology and Hepatology Associate Director Center for
PROGRESS: Beginning to Understand the Genetic Predisposition to PSC Konstantinos N. Lazaridis, MD Associate Professor of Medicine Division of Gastroenterology and Hepatology Associate Director Center for
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
JULY 21, Genetics 101: SCN1A. Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology
 JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
Genetic mates SNPedia write-ups Final Next-gen sequencing Mike Snyder QA: new technology and the future
 Genetic mates SNPedia write-ups Final Next-gen sequencing Mike Snyder QA: new technology and the future Genetic Mates Jonathan Mortensen/Francisco Gimenez Will need class to upload data Tuesday night Analyze
Genetic mates SNPedia write-ups Final Next-gen sequencing Mike Snyder QA: new technology and the future Genetic Mates Jonathan Mortensen/Francisco Gimenez Will need class to upload data Tuesday night Analyze
Genotype to Phenotype Simulation Booklet. Combining germ cells to create a new baby human
 Genotype to Phenotype Simulation Booklet Combining germ cells to create a new baby human 1 A Genetic Simulation Making A Face: Converting Genotype Into Phenotype by Simulating Meiosis and Fertilization
Genotype to Phenotype Simulation Booklet Combining germ cells to create a new baby human 1 A Genetic Simulation Making A Face: Converting Genotype Into Phenotype by Simulating Meiosis and Fertilization
NGS panels in clinical diagnostics: Utrecht experience. Van Gijn ME PhD Genome Diagnostics UMCUtrecht
 NGS panels in clinical diagnostics: Utrecht experience Van Gijn ME PhD Genome Diagnostics UMCUtrecht 93 Gene panels UMC Utrecht Cardiovascular disease (CAR) (5 panels) Epilepsy (EPI) (11 panels) Hereditary
NGS panels in clinical diagnostics: Utrecht experience Van Gijn ME PhD Genome Diagnostics UMCUtrecht 93 Gene panels UMC Utrecht Cardiovascular disease (CAR) (5 panels) Epilepsy (EPI) (11 panels) Hereditary
No mutations were identified.
 Hereditary High Cholesterol Test ORDERING PHYSICIAN PRIMARY CONTACT SPECIMEN Report date: Aug 1, 2017 Dr. Jenny Jones Sample Medical Group 123 Main St. Sample, CA Kelly Peters Sample Medical Group 123
Hereditary High Cholesterol Test ORDERING PHYSICIAN PRIMARY CONTACT SPECIMEN Report date: Aug 1, 2017 Dr. Jenny Jones Sample Medical Group 123 Main St. Sample, CA Kelly Peters Sample Medical Group 123
Supplementary Note. Phenotype details in ten patients with deletions of 15q13
 Supplementary Note. Phenotype details in ten patients with deletions of 15q13 IMR338, IMR338M, and IMR338Cc familial 3.95 Mb deletion BP3-BP5 The proband, IMR338, was initially reported by Sharp et al.
Supplementary Note. Phenotype details in ten patients with deletions of 15q13 IMR338, IMR338M, and IMR338Cc familial 3.95 Mb deletion BP3-BP5 The proband, IMR338, was initially reported by Sharp et al.
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep
 Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
Nature Neuroscience: doi: /nn Supplementary Figure 1. Missense damaging predictions as a function of allele frequency
 Supplementary Figure 1 Missense damaging predictions as a function of allele frequency Percentage of missense variants classified as damaging by eight different classifiers and a classifier consisting
Supplementary Figure 1 Missense damaging predictions as a function of allele frequency Percentage of missense variants classified as damaging by eight different classifiers and a classifier consisting
The Genetics of VHL. Proper tissue growth - controlled traffic. How human cells and tissue grow and die?
 How human cells and tissue grow and die? The Genetics of VHL Xia Wang MD PhD Oct, 2017 Proper tissue growth - controlled traffic Normal tissue growth is regulated by many genetic factors Safe traffic is
How human cells and tissue grow and die? The Genetics of VHL Xia Wang MD PhD Oct, 2017 Proper tissue growth - controlled traffic Normal tissue growth is regulated by many genetic factors Safe traffic is
CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY
 CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY More genetic information requires cutting-edge interpretation techniques Whole Exome Sequencing For certain patients the combination of symptoms does not allow
CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY More genetic information requires cutting-edge interpretation techniques Whole Exome Sequencing For certain patients the combination of symptoms does not allow
(i) Family 1. The male proband (1.III-1) from European descent was referred at
 1 Supplementary Note Clinical descriptions of families (i) Family 1. The male proband (1.III-1) from European descent was referred at age 14 because of scoliosis. He had normal development. Physical evaluation
1 Supplementary Note Clinical descriptions of families (i) Family 1. The male proband (1.III-1) from European descent was referred at age 14 because of scoliosis. He had normal development. Physical evaluation
Challenges of CGH array testing in children with developmental delay. Dr Sally Davies 17 th September 2014
 Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
Clinical utility of the functional mrna evaluation of rare genetic variants in diagnostic practice
 Clinical utility of the functional mrna evaluation of rare genetic variants in diagnostic practice Celia Duff-Farrier: STP MSc project 2017 Funding provided by the Showering Fund mrna functional evaluation
Clinical utility of the functional mrna evaluation of rare genetic variants in diagnostic practice Celia Duff-Farrier: STP MSc project 2017 Funding provided by the Showering Fund mrna functional evaluation
PedsCases Podcast Scripts
 PedsCases Podcast Scripts This is a text version of a podcast from Pedscases.com on the Approach to Pediatric Anemia and Pallor. These podcasts are designed to give medical students an overview of key
PedsCases Podcast Scripts This is a text version of a podcast from Pedscases.com on the Approach to Pediatric Anemia and Pallor. These podcasts are designed to give medical students an overview of key
Clinical evaluation of microarray data
 Clinical evaluation of microarray data David Amor 19 th June 2011 Single base change Microarrays 3-4Mb What is a microarray? Up to 10 6 bits of Information!! Highly multiplexed FISH hybridisations. Microarray
Clinical evaluation of microarray data David Amor 19 th June 2011 Single base change Microarrays 3-4Mb What is a microarray? Up to 10 6 bits of Information!! Highly multiplexed FISH hybridisations. Microarray
MEDICAL GENOMICS LABORATORY. Non-NF1 RASopathy panel by Next-Gen Sequencing and Deletion/Duplication Analysis of SPRED1 (NNP-NG)
 Non-NF1 RASopathy panel by Next-Gen Sequencing and Deletion/Duplication Analysis of SPRED1 (NNP-NG) Ordering Information Acceptable specimen types: Blood (3-6ml EDTA; no time limitations associated with
Non-NF1 RASopathy panel by Next-Gen Sequencing and Deletion/Duplication Analysis of SPRED1 (NNP-NG) Ordering Information Acceptable specimen types: Blood (3-6ml EDTA; no time limitations associated with
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Epileptic encephalopathy, early infantile 4. OMIM number for disease 612164 Disease
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Epileptic encephalopathy, early infantile 4. OMIM number for disease 612164 Disease
Most severely affected will be the probe for exon 15. Please keep an eye on the D-fragments (especially the 96 nt fragment).
 SALSA MLPA probemix P343-C3 Autism-1 Lot C3-1016. As compared to version C2 (lot C2-0312) five reference probes have been replaced, one reference probe added and several lengths have been adjusted. Warning:
SALSA MLPA probemix P343-C3 Autism-1 Lot C3-1016. As compared to version C2 (lot C2-0312) five reference probes have been replaced, one reference probe added and several lengths have been adjusted. Warning:
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Choroideremia OMIM number for disease 303100 Disease alternative names please
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name Choroideremia OMIM number for disease 303100 Disease alternative names please
What dysmorphologists should keep in mind during evaluation of patients with blepharophimosis and mental disability?
 What dysmorphologists should keep in mind during evaluation of patients with blepharophimosis and mental disability? Aleksandra Jezela-Stanek, M.D., Ph. D. The Children s Memorial Health Institute, Warsaw
What dysmorphologists should keep in mind during evaluation of patients with blepharophimosis and mental disability? Aleksandra Jezela-Stanek, M.D., Ph. D. The Children s Memorial Health Institute, Warsaw
Diagnostic Exome Sequencing in Persons with Severe Intellectual Disability
 original article Diagnostic Exome Sequencing in Persons with Severe Intellectual Disability Joep de Ligt, M.Sc., Marjolein H. Willemsen, M.D., Bregje W.M. van Bon, M.D., Ph.D., Tjitske Kleefstra, M.D.,
original article Diagnostic Exome Sequencing in Persons with Severe Intellectual Disability Joep de Ligt, M.Sc., Marjolein H. Willemsen, M.D., Bregje W.M. van Bon, M.D., Ph.D., Tjitske Kleefstra, M.D.,
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name OMIM number for disease 147920 Disease alternative names Please provide any alternative
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier Test Disease Population Triad Disease name OMIM number for disease 147920 Disease alternative names Please provide any alternative
WDR62 is associated with the spindle pole and mutated in human microcephaly
 WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
WDR62 is associated with the spindle pole and mutated in human microcephaly Adeline K. Nicholas, Maryam Khurshid, Julie Désir, Ofélia P. Carvalho, James J. Cox, Gemma Thornton, Rizwana Kausar, Muhammad
and duodenal atresia. Two other families reported may have had the same syndrome. Case reports CASE 1 (III.8, FAMILY 1)
 JMedGenet 1991; 28: 389-394 Autosomal dominant inheritance of abnormalities of the hands and feet with short palpebral fissures, variable microcephaly with learning disability, and oesophageal/duodenal
JMedGenet 1991; 28: 389-394 Autosomal dominant inheritance of abnormalities of the hands and feet with short palpebral fissures, variable microcephaly with learning disability, and oesophageal/duodenal
Merging single gene-level CNV with sequence variant interpretation following the ACMGG/AMP sequence variant guidelines
 Merging single gene-level CNV with sequence variant interpretation following the ACMGG/AMP sequence variant guidelines Tracy Brandt, Ph.D., FACMG Disclosure I am an employee of GeneDx, Inc., a wholly-owned
Merging single gene-level CNV with sequence variant interpretation following the ACMGG/AMP sequence variant guidelines Tracy Brandt, Ph.D., FACMG Disclosure I am an employee of GeneDx, Inc., a wholly-owned
Human Congenital Diseases with Mixed Modes of Inheritance Have a Shortage of Recessive Disease. A Demographic Scenario?
 doi: 10.1111/j.1469-1809.2011.00679.x Human Congenital Diseases with Mixed Modes of Inheritance Have a Shortage of Recessive Disease. A Demographic Scenario? N. Avrion Mitchison 1, Shomi Bhattacharya 1
doi: 10.1111/j.1469-1809.2011.00679.x Human Congenital Diseases with Mixed Modes of Inheritance Have a Shortage of Recessive Disease. A Demographic Scenario? N. Avrion Mitchison 1, Shomi Bhattacharya 1
What is the relationship between genes and chromosomes? Is twinning genetic or can a person choose to have twins?
 WHAT WILL YOU KNOW? What is the relationship between genes and chromosomes? Is twinning genetic or can a person choose to have twins? How could a person have the gene for something that is never apparent?
WHAT WILL YOU KNOW? What is the relationship between genes and chromosomes? Is twinning genetic or can a person choose to have twins? How could a person have the gene for something that is never apparent?
Home Brewed Personalized Genomics
 Home Brewed Personalized Genomics The Quest for Meaningful Analysis Results of a 23andMe Exome Pilot Trio of Myself, Wife, and Son February 22, 2013 Gabe Rudy, Vice President of Product Development Exome
Home Brewed Personalized Genomics The Quest for Meaningful Analysis Results of a 23andMe Exome Pilot Trio of Myself, Wife, and Son February 22, 2013 Gabe Rudy, Vice President of Product Development Exome
Author's response to reviews
 Author's response to reviews Title: Dystonia, facial dysmorphism, intellectual disability and breast cancer associated with a chromosome 13q34 duplication and overexpression of TFDP1: Case report Authors:
Author's response to reviews Title: Dystonia, facial dysmorphism, intellectual disability and breast cancer associated with a chromosome 13q34 duplication and overexpression of TFDP1: Case report Authors:
Biology 3A Laboratory Mendelian, Human & Population Genetics Worksheet
 Biology 3A Laboratory Mendelian, Human & Population Genetics Worksheet Name: Lab Day & Time: A. UNDERSTANDING MEIOSIS & CHROMOSOME SEGREGATION 1. Meiosis activity: Diagram the process of meiosis using
Biology 3A Laboratory Mendelian, Human & Population Genetics Worksheet Name: Lab Day & Time: A. UNDERSTANDING MEIOSIS & CHROMOSOME SEGREGATION 1. Meiosis activity: Diagram the process of meiosis using
Rapid testing and prenatal diagnosis for at-risk couples presenting during pregnancy without a diagnosis
 Rapid testing and prenatal diagnosis for at-risk couples presenting during pregnancy without a diagnosis Stephen Abbs East Anglian Medical Genetics Service Cambridge University Hospitals NHS Foundation
Rapid testing and prenatal diagnosis for at-risk couples presenting during pregnancy without a diagnosis Stephen Abbs East Anglian Medical Genetics Service Cambridge University Hospitals NHS Foundation
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Complete Genomics, Inc.
 Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
