Home Brewed Personalized Genomics
|
|
|
- Alexander Lester
- 5 years ago
- Views:
Transcription
1 Home Brewed Personalized Genomics The Quest for Meaningful Analysis Results of a 23andMe Exome Pilot Trio of Myself, Wife, and Son February 22, 2013 Gabe Rudy, Vice President of Product Development
2 Exome Sequencing in Consumer Genomics Gabe (me) Erin JIA Ethan Exomes done as part of Pilot Program 80x coverage Raw data with no interpretation
3 Overview 1 Consumer genetics data: research or clinical grade? 2 Treating my healthy self to a Mendelian disease analysis 3 Using exome data to explain a rare autoimmune disorder
4 Consumer exomes: done using best practices Sequencing done on HiSeq bp PE - Agilent SureSelect exome capture Aligned and called with BWA/GATK - Broad s Best Practices with GATK Guide - Indel realignment - UnifiedGenotyper called samples concurrently Deliverables - BAM (minus indel realignments) - VCF (some filters applied) - PDF of Summary Report
5 Research or clinical grade? Total Reads 140M Unique Align 87% Mean Target 105x % Target at 2x 97% % Target at 10x 94% % Target at 20x 89% % Target at 30x 83%
6 Clinical grade Gabe Trio Unfiltered Provided RD>10 & GQ>20 Exonic SNPs InDels Ts/Tv Mendel Errors
7 Summary Report
8 Updated VCF and report at end of October GATK is a Research Tool. Clinics Beware.
9 Overview 1 Consumer genetics data: research or clinical grade? 2 Treating my healthy self to a Mendelian disease analysis 3 Rare Variant Analysis: The Hammer Using exome data to explain a rare autoimmune disorder My Exome: The Nail
10 Filtering and analysis strategy Follow best practices for highimpact variants Weed out falsepositives Use functional prediction Interpretation more open-ended Everything Low quality & phantom 85K Rare 17K Non-synonymous or LoF 1,698 Now what? 151K
11 Population catalog and variant classification Non-coding Coding Variant Rare (Novel) Intergenic 8,462 3,609 (1,130) Intronic 48,826 7,516 (4,418) UTR 3/5 4, (303) Non-coding 1, (183) Variant Rare (Novel) Splicing (17) Frameshift Ins/Del (118) Stop gain/loss (9) Non-synonymous 10,252 1,501 (447) In-frame Ins/Del (89) Synonymous 11, (215) Unknown (36) Loss-of-function: 197 Nonsynonymous: 1,501
12 More filtering strategies Regions of Chromosomal Duplication (SuperDups) Look at genes in OMIM (most) Use predictions of genes as recessive/haploinsufficient to weed out low-priority genes For nonsynonymous missense variants can use functional prediction (SIFT/Polyphen2) to annotate
13 Genes of interest and homozygous variants Loss of Function Rare!Dups OMIM Rec Genes Splicing (2) 0 (0) Frameshift Del (4) 1 (0) Frameshift Ins (5) 3 (1) Stop gain (0) 0 (0) Non-Synonymous Rare!Dups OMIM Rec Genes Damaging (3/3) (0) 9 (0) Damaging (2/3) (0) 7 (0) Damaging (1/3) (1) 12 (0) Tolerated/Unk (4) 10 (1) Homozygous: in OMIM: 16 Heterozygous in Rec Genes: 40
14 Homozygous variants Note Variant AD DP GQ Gene(s) Classification HGVS Coding 1 common 1: Ins 50, CDCP2 Frameshift Ins c.1224_1225insc reference 2: Ins 106, CD207 Splicing in-wife 5: Ins 6, CYFIP2 Frameshift Ins c.279_280insc bad-call 6: Del 120, AARS2 Frameshift Del c.2607delg bad-call? 10: SNV 21, GPRIN2 Nonsyn SNV c.724a>g common 12: Ins 95, ITPR2 Splicing in-wife 14: Ins 3, GPHB5 Frameshift Ins c.156_157insc bad-call 17: Del 161, WRAP53 Frameshift Del c.1565delc bad-call 19: Del 142, CNOT3 Frameshift Del c.729delt in-wife 22: Ins 6, CLTCL1 Frameshift Ins c.3601_3602insg VUS X: SNV 0, CTPS2 Nonsyn SNV c.1342a>c pathogenic X: SNV 0, OTC Nonsyn SNV c.148g>a VUS X: SNV 0, DRP2 Nonsyn SNV c.380c>t VUS/in-5-M X: SNV 0, NRK Nonsyn SNV c.2912a>g wrong-geno X: Ins 61, AMOT Frameshift Ins c.3080_3081inscc VUS/common X: Del 106, GPR50 Frameshift Del c.1504_1514delaccact GGCCA
15 Review Homozygous Variants in SNP & Variation X: X: Suite and GenomeBrowse
16 Overview 1 Consumer genetics data: research or clinical grade? 2 Treating my healthy self to a Mendelian disease analysis 3 Using exome data to explain a rare autoimmune disorder
17 Juvenile Idiopathic Arthritis (JIA) Unknown cause, onset before 16 Between 8 and 150 of every 100,000 children 50% have pauciarticular JIA 40% have polyarticular JIA Polyarticular RF negative sub-phenotype has heritability similar to Rheumatoid Arthritis (RA) - RA is expected to be 60% heritable - 51% explained through current genetic associations - 36% of heritability in HLA
18 IVA Rare deleterious variance within 1 hop of JIA
19 1KG: AAF NHLBI: AA=0/AG=7/GG=4293
20 Altered T Cell and B Cell Signaling Mostly HLA-DRB1
21 Variants in the Alerted T Cell and B Cell Signaling Pathway fall in HLA-DRB1/HLA- DQA1/HLA-DQB1 and one in NFKB1
22 HLA Typing with Omixon Target HLA 32 HLA genes, types based on IMGT/HLA nomenclature HLA-A*02 predisposes to early-onset JIA HLA-DRB1*08 and HLA-DPB1*03 predispose to poly RF- JIA HLA-DRB1*04, HLA-DQA1*03 and HLA-DQB1*03 predispose to poly RF+JIA HLA-A A*24:02:01:01 A*03:01:01:01 HLA-DRB1 DRB1*04:07:01 DRB1*14:54:01 HLA-DQA1 DQA1*01:04:01 DQA1*03:03:01 HLA-DPB1 DPB1*04:01:01 DPB1*04:01:01
23 Predicting Lifetime Risk from Population Studies
24 GWAS for JIA
25 rs has R^2 of 1 with rs which is 537bp away
26 UK 2012
27 CCHMC 2010
28 CCHMC 2012
29 Combined
30 Final thoughts No smoking gun, but variants of interest Ongoing RA research with population level WGS and family NGS To be seen how much of autoimmune disorder heritability is explained by rare variants with higher effect sizes. Most promising signal is in genotype SNPs that might be tagging for functional mutations in regulatory regions. Rare sub-classifications like JIA polyarticular RF negative may be difficult to nail down with population studies Family studies looking at shared biomarkers along with symptoms may be better suited to find cause-effect relationships Wife s nuclear family has diagnosed cases of: Pheochromocytoma: 2-8 per 1,000,000 Guillain-Barré: per 100,000
31 Acknowledgements Dr. Peter Gregersen - Director, Robert S. Boas Center for Genomics and Human Genetics, The Feinstein Institute Dr. Gerald Nepom - Director, Benaroya Research Institute, Director, Immune Tolerance Network Golden Helix - SNP & Variation Suite - GenomeBrowse Sean Scott - Ingenuity - Variant Analysis Tim Hague - Omixon - Target HLA Typing Dr. Brian Naughton, 23andMe - Trio exome sequencing Dr. Peter Gregersen Dr. Gerald Nepom
Variant Classification. Author: Mike Thiesen, Golden Helix, Inc.
 Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Identifying Mutations Responsible for Rare Disorders Using New Technologies
 Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Analysis with SureCall 2.1
 Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Golden Helix s End-to-End Solution for Clinical Labs
 Golden Helix s End-to-End Solution for Clinical Labs Steven Hystad - Field Application Scientist Nathan Fortier Senior Software Engineer 20 most promising Biotech Technology Providers Top 10 Analytics
Golden Helix s End-to-End Solution for Clinical Labs Steven Hystad - Field Application Scientist Nathan Fortier Senior Software Engineer 20 most promising Biotech Technology Providers Top 10 Analytics
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE
 CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
Rare Variant Burden Tests. Biostatistics 666
 Rare Variant Burden Tests Biostatistics 666 Last Lecture Analysis of Short Read Sequence Data Low pass sequencing approaches Modeling haplotype sharing between individuals allows accurate variant calls
Rare Variant Burden Tests Biostatistics 666 Last Lecture Analysis of Short Read Sequence Data Low pass sequencing approaches Modeling haplotype sharing between individuals allows accurate variant calls
Investigating rare diseases with Agilent NGS solutions
 Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Genetic Testing and Analysis. (858) MRN: Specimen: Saliva Received: 07/26/2016 GENETIC ANALYSIS REPORT
 GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future?
 Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future? Tony Marinakii Purine Research Laboratory Biochemical Sciences Whole exome sequencing No doubt
Whole exome sequencing as a first line test: Is there even a role for metabolic biochemists in the future? Tony Marinakii Purine Research Laboratory Biochemical Sciences Whole exome sequencing No doubt
Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection
 Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection Dr Elaine Kenny Neuropsychiatric Genetics Research Group Institute of Molecular Medicine Trinity College Dublin
Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection Dr Elaine Kenny Neuropsychiatric Genetics Research Group Institute of Molecular Medicine Trinity College Dublin
Genome Summary. Sequencing Coverage: Variation Counts: Known Phenotype Summary: 131 disease or trait variations are found in this genome
 Report Date: August 2, 215 Software Annotation Version: 8 Genome Summary Name: Greg Mendel Genome ID: genome_greg_mendel_dad Sequencing Provider: 23andMe Sequencing Type: Genotyping SNP Array Sequencing
Report Date: August 2, 215 Software Annotation Version: 8 Genome Summary Name: Greg Mendel Genome ID: genome_greg_mendel_dad Sequencing Provider: 23andMe Sequencing Type: Genotyping SNP Array Sequencing
Welcome to the Genetic Code: An Overview of Basic Genetics. October 24, :00pm 3:00pm
 Welcome to the Genetic Code: An Overview of Basic Genetics October 24, 2016 12:00pm 3:00pm Course Schedule 12:00 pm 2:00 pm Principles of Mendelian Genetics Introduction to Genetics of Complex Disease
Welcome to the Genetic Code: An Overview of Basic Genetics October 24, 2016 12:00pm 3:00pm Course Schedule 12:00 pm 2:00 pm Principles of Mendelian Genetics Introduction to Genetics of Complex Disease
PERSONALIZED GENETIC REPORT CLIENT-REPORTED DATA PURPOSE OF THE X-SCREEN TEST
 INCLUDED IN THIS REPORT: REVIEW OF YOUR GENETIC INFORMATION RELEVANT TO ENDOMETRIOSIS PERSONAL EDUCATIONAL INFORMATION RELEVANT TO YOUR GENES INFORMATION FOR OBTAINING YOUR ENTIRE X-SCREEN DATA FILE PERSONALIZED
INCLUDED IN THIS REPORT: REVIEW OF YOUR GENETIC INFORMATION RELEVANT TO ENDOMETRIOSIS PERSONAL EDUCATIONAL INFORMATION RELEVANT TO YOUR GENES INFORMATION FOR OBTAINING YOUR ENTIRE X-SCREEN DATA FILE PERSONALIZED
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
TOWARDS ACCURATE GERMLINE AND SOMATIC INDEL DISCOVERY WITH MICRO-ASSEMBLY. Giuseppe Narzisi, PhD Bioinformatics Scientist
 TOWARDS ACCURATE GERMLINE AND SOMATIC INDEL DISCOVERY WITH MICRO-ASSEMBLY Giuseppe Narzisi, PhD Bioinformatics Scientist July 29, 2014 Micro-Assembly Approach to detect INDELs 2 Outline 1 Detecting INDELs:
TOWARDS ACCURATE GERMLINE AND SOMATIC INDEL DISCOVERY WITH MICRO-ASSEMBLY Giuseppe Narzisi, PhD Bioinformatics Scientist July 29, 2014 Micro-Assembly Approach to detect INDELs 2 Outline 1 Detecting INDELs:
Spectrum of mutations in monogenic diabetes genes identified from high-throughput DNA sequencing of 6888 individuals
 Bansal et al. BMC Medicine (2017) 15:213 DOI 10.1186/s12916-017-0977-3 RESEARCH ARTICLE Spectrum of mutations in monogenic diabetes genes identified from high-throughput DNA sequencing of 6888 individuals
Bansal et al. BMC Medicine (2017) 15:213 DOI 10.1186/s12916-017-0977-3 RESEARCH ARTICLE Spectrum of mutations in monogenic diabetes genes identified from high-throughput DNA sequencing of 6888 individuals
Variant Annotation and Functional Prediction
 Variant Annotation and Functional Prediction Copyrighted 2018 Isabelle Schrauwen and Suzanne M. Leal This exercise touches on several functionalities of the program ANNOVAR to annotate and interpret candidate
Variant Annotation and Functional Prediction Copyrighted 2018 Isabelle Schrauwen and Suzanne M. Leal This exercise touches on several functionalities of the program ANNOVAR to annotate and interpret candidate
Nature Neuroscience: doi: /nn Supplementary Figure 1. Missense damaging predictions as a function of allele frequency
 Supplementary Figure 1 Missense damaging predictions as a function of allele frequency Percentage of missense variants classified as damaging by eight different classifiers and a classifier consisting
Supplementary Figure 1 Missense damaging predictions as a function of allele frequency Percentage of missense variants classified as damaging by eight different classifiers and a classifier consisting
Introduction to genetic variation. He Zhang Bioinformatics Core Facility 6/22/2016
 Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
Shashikant Kulkarni, M.S (Medicine)., Ph.D., FACMG Associate Professor of Pathology & Immunology Associate Professor of Pediatrics and Genetics
 Shashikant Kulkarni, M.S (Medicine)., Ph.D., FACMG Associate Professor of Pathology & Immunology Associate Professor of Pediatrics and Genetics Director of Cytogenomics and Molecular Pathology Evidence-based
Shashikant Kulkarni, M.S (Medicine)., Ph.D., FACMG Associate Professor of Pathology & Immunology Associate Professor of Pediatrics and Genetics Director of Cytogenomics and Molecular Pathology Evidence-based
6/12/2018. Disclosures. Clinical Genomics The CLIA Lab Perspective. Outline. COH HopeSeq Heme Panels
 Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11396 A total of 2078 samples from a large sequencing project at decode were used in this study, 219 samples from 78 trios with two grandchildren who were not also members of other trios,
doi:10.1038/nature11396 A total of 2078 samples from a large sequencing project at decode were used in this study, 219 samples from 78 trios with two grandchildren who were not also members of other trios,
SCALPEL MICRO-ASSEMBLY APPROACH TO DETECT INDELS WITHIN EXOME-CAPTURE DATA. Giuseppe Narzisi, PhD Schatz Lab
 SCALPEL MICRO-ASSEMBLY APPROACH TO DETECT INDELS WITHIN EXOME-CAPTURE DATA Giuseppe Narzisi, PhD Schatz Lab November 14, 2013 Micro-Assembly Approach to detect INDELs 2 Outline Scalpel micro-assembly pipeline
SCALPEL MICRO-ASSEMBLY APPROACH TO DETECT INDELS WITHIN EXOME-CAPTURE DATA Giuseppe Narzisi, PhD Schatz Lab November 14, 2013 Micro-Assembly Approach to detect INDELs 2 Outline Scalpel micro-assembly pipeline
PROGRESS: Beginning to Understand the Genetic Predisposition to PSC
 PROGRESS: Beginning to Understand the Genetic Predisposition to PSC Konstantinos N. Lazaridis, MD Associate Professor of Medicine Division of Gastroenterology and Hepatology Associate Director Center for
PROGRESS: Beginning to Understand the Genetic Predisposition to PSC Konstantinos N. Lazaridis, MD Associate Professor of Medicine Division of Gastroenterology and Hepatology Associate Director Center for
Concurrent Practical Session ACMG Classification
 Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
ACE ImmunoID Biomarker Discovery Solutions ACE ImmunoID Platform for Tumor Immunogenomics
 ACE ImmunoID Biomarker Discovery Solutions ACE ImmunoID Platform for Tumor Immunogenomics Precision Genomics for Immuno-Oncology Personalis, Inc. ACE ImmunoID When one biomarker doesn t tell the whole
ACE ImmunoID Biomarker Discovery Solutions ACE ImmunoID Platform for Tumor Immunogenomics Precision Genomics for Immuno-Oncology Personalis, Inc. ACE ImmunoID When one biomarker doesn t tell the whole
CITATION FILE CONTENT/FORMAT
 CITATION For any resultant publications using please cite: Matthew A. Field, Vicky Cho, T. Daniel Andrews, and Chris C. Goodnow (2015). "Reliably detecting clinically important variants requires both combined
CITATION For any resultant publications using please cite: Matthew A. Field, Vicky Cho, T. Daniel Andrews, and Chris C. Goodnow (2015). "Reliably detecting clinically important variants requires both combined
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Assessing Laboratory Performance for Next Generation Sequencing Based Detection of Germline Variants through Proficiency Testing
 Assessing Laboratory Performance for Next Generation Sequencing Based Detection of Germline Variants through Proficiency Testing Karl V. Voelkerding, MD Professor of Pathology University of Utah Medical
Assessing Laboratory Performance for Next Generation Sequencing Based Detection of Germline Variants through Proficiency Testing Karl V. Voelkerding, MD Professor of Pathology University of Utah Medical
GENOME-WIDE ASSOCIATION STUDIES
 GENOME-WIDE ASSOCIATION STUDIES SUCCESSES AND PITFALLS IBT 2012 Human Genetics & Molecular Medicine Zané Lombard IDENTIFYING DISEASE GENES??? Nature, 15 Feb 2001 Science, 16 Feb 2001 IDENTIFYING DISEASE
GENOME-WIDE ASSOCIATION STUDIES SUCCESSES AND PITFALLS IBT 2012 Human Genetics & Molecular Medicine Zané Lombard IDENTIFYING DISEASE GENES??? Nature, 15 Feb 2001 Science, 16 Feb 2001 IDENTIFYING DISEASE
Genetics and Genomics in Medicine Chapter 8 Questions
 Genetics and Genomics in Medicine Chapter 8 Questions Linkage Analysis Question Question 8.1 Affected members of the pedigree above have an autosomal dominant disorder, and cytogenetic analyses using conventional
Genetics and Genomics in Medicine Chapter 8 Questions Linkage Analysis Question Question 8.1 Affected members of the pedigree above have an autosomal dominant disorder, and cytogenetic analyses using conventional
TP53 mutational profile in CLL : A retrospective study of the FILO group.
 TP53 mutational profile in CLL : A retrospective study of the FILO group. Fanny Baran-Marszak Hopital Avicenne Bobigny France 2nd ERIC workshop on TP53 analysis in CLL, Stresa 2017 TP53 abnormalities :
TP53 mutational profile in CLL : A retrospective study of the FILO group. Fanny Baran-Marszak Hopital Avicenne Bobigny France 2nd ERIC workshop on TP53 analysis in CLL, Stresa 2017 TP53 abnormalities :
Evaluation of MIA FORA NGS HLA test and software. Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group
 Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
2/10/2016. Evaluation of MIA FORA NGS HLA test and software. Disclosure. NGS-HLA typing requirements for the Stanford Blood Center
 Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
New Enhancements: GWAS Workflows with SVS
 New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
No mutations were identified.
 Hereditary High Cholesterol Test ORDERING PHYSICIAN PRIMARY CONTACT SPECIMEN Report date: Aug 1, 2017 Dr. Jenny Jones Sample Medical Group 123 Main St. Sample, CA Kelly Peters Sample Medical Group 123
Hereditary High Cholesterol Test ORDERING PHYSICIAN PRIMARY CONTACT SPECIMEN Report date: Aug 1, 2017 Dr. Jenny Jones Sample Medical Group 123 Main St. Sample, CA Kelly Peters Sample Medical Group 123
BWA alignment to reference transcriptome and genome. Convert transcriptome mappings back to genome space
 Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS
 Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Identification of genomic alterations in cervical cancer biopsies by exome sequencing
 Chapter- 4 Identification of genomic alterations in cervical cancer biopsies by exome sequencing 105 4.1 INTRODUCTION Athough HPV has been identified as the prime etiological factor for cervical cancer,
Chapter- 4 Identification of genomic alterations in cervical cancer biopsies by exome sequencing 105 4.1 INTRODUCTION Athough HPV has been identified as the prime etiological factor for cervical cancer,
Nature Genetics: doi: /ng Supplementary Figure 1. Rates of different mutation types in CRC.
 Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Supplementary Figure 1 Rates of different mutation types in CRC. (a) Stratification by mutation type indicates that C>T mutations occur at a significantly greater rate than other types. (b) As for the
Genomic approach for drug target discovery and validation
 Genomic approach for drug target discovery and validation Hong-Hee Won, Ph.D. SAIHST, SKKU wonhh@skku.edu Well-known example of genetic findings triggering drug development Genetics Mendelian randomisation
Genomic approach for drug target discovery and validation Hong-Hee Won, Ph.D. SAIHST, SKKU wonhh@skku.edu Well-known example of genetic findings triggering drug development Genetics Mendelian randomisation
MutationTaster & RegulationSpotter
 Practical MutationTaster & RegulationSpotter Daniela Hombach & Jana Marie Schwarz Exzellenzcluster NeuroCure Charité Universitätsmedizin Berlin AG Translational Genomics 08.11.2017 Rare diseases 7000 8000rare
Practical MutationTaster & RegulationSpotter Daniela Hombach & Jana Marie Schwarz Exzellenzcluster NeuroCure Charité Universitätsmedizin Berlin AG Translational Genomics 08.11.2017 Rare diseases 7000 8000rare
Downloaded for Anonymous User (n/a) at University of Washington - Seattle - WSC from ClinicalKey.com by Elsevier on March 04, 2019.
 Accepted Manuscript The MALT1 locus and Peanut Avoidance in the Risk for Peanut Allergy Alexandra Winters, PhD, Henry T. Bahnson, MPH, Ingo Ruczinski, PhD, Meher P. Boorgula, MS, Claire Malley, MS, Ali
Accepted Manuscript The MALT1 locus and Peanut Avoidance in the Risk for Peanut Allergy Alexandra Winters, PhD, Henry T. Bahnson, MPH, Ingo Ruczinski, PhD, Meher P. Boorgula, MS, Claire Malley, MS, Ali
CPT Codes for Pharmacogenomic Tests
 CPT s for Pharmacogenomic Tests The table below lists CPT codes and lab fee information for pharmacogenomic tests as established by the Centers for Medicare and Medicaid Services. It was compiled by the
CPT s for Pharmacogenomic Tests The table below lists CPT codes and lab fee information for pharmacogenomic tests as established by the Centers for Medicare and Medicaid Services. It was compiled by the
MutationTaster & RegulationSpotter
 MutationTaster & RegulationSpotter Pathogenicity Prediction of Sequence Variants: Past, Present and Future Dr. rer. nat. Jana Marie Schwarz Klinik für Pädiatrie m. S. Neurologie Exzellenzcluster NeuroCure
MutationTaster & RegulationSpotter Pathogenicity Prediction of Sequence Variants: Past, Present and Future Dr. rer. nat. Jana Marie Schwarz Klinik für Pädiatrie m. S. Neurologie Exzellenzcluster NeuroCure
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13908 Supplementary Tables Supplementary Table 1: Families in this study (.xlsx) All families included in the study are listed. For each family, we show: the genders of the probands and
doi:10.1038/nature13908 Supplementary Tables Supplementary Table 1: Families in this study (.xlsx) All families included in the study are listed. For each family, we show: the genders of the probands and
Epigenetics. Jenny van Dongen Vrije Universiteit (VU) Amsterdam Boulder, Friday march 10, 2017
 Epigenetics Jenny van Dongen Vrije Universiteit (VU) Amsterdam j.van.dongen@vu.nl Boulder, Friday march 10, 2017 Epigenetics Epigenetics= The study of molecular mechanisms that influence the activity of
Epigenetics Jenny van Dongen Vrije Universiteit (VU) Amsterdam j.van.dongen@vu.nl Boulder, Friday march 10, 2017 Epigenetics Epigenetics= The study of molecular mechanisms that influence the activity of
Whole Exome Sequencing (WES): Questions and Answers for Providers
 Whole Exome Sequencing (WES): Questions and Answers for Providers 1. What is Whole Exome Sequencing?... 2 2. What is the difference between Whole Exome Sequencing (WES) and Whole Exome Sequencing Plus
Whole Exome Sequencing (WES): Questions and Answers for Providers 1. What is Whole Exome Sequencing?... 2 2. What is the difference between Whole Exome Sequencing (WES) and Whole Exome Sequencing Plus
HLA and new technologies. Vicky Van Sandt
 HLA and new technologies. Vicky Van Sandt Life-threatning malignant and non malignant blood disorders can be cured by hematopoetic stem cell transplantation (HSCT). GVHD is the 2nd most prevalent cause
HLA and new technologies. Vicky Van Sandt Life-threatning malignant and non malignant blood disorders can be cured by hematopoetic stem cell transplantation (HSCT). GVHD is the 2nd most prevalent cause
!"##"$%#"&!'&$'()$(%&'*& Terapia Pediatrica e Farmacologia dello Sviluppo +,-./&01,23&34,53& :&;.<&2-.=;:3&;.;2>6-6&-.&;&
 !!! "#$%&'($)*!+&,-$!.)/+$!+$!01,-$1'$!!"##"$%#"&!'&$'()$(%&'*& Terapia Pediatrica e Farmacologia dello Sviluppo 0$2-3!44566!! +,-./&01,23&34,53&63783.9-.:&;.6-6&-.&;& 582/-:3.3?;/-,.;2&@;5-2>&63:?3:;/-.:&#>A3&B&!-;C3/36&!.&))3'&!(2$&#)$7$23!+$(2$8-$#1'&!+$!177&'[!!
!!! "#$%&'($)*!+&,-$!.)/+$!+$!01,-$1'$!!"##"$%#"&!'&$'()$(%&'*& Terapia Pediatrica e Farmacologia dello Sviluppo 0$2-3!44566!! +,-./&01,23&34,53&63783.9-.:&;.6-6&-.&;& 582/-:3.3?;/-,.;2&@;5-2>&63:?3:;/-.:&#>A3&B&!-;C3/36&!.&))3'&!(2$&#)$7$23!+$(2$8-$#1'&!+$!177&'[!!
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
WHOLE EXOME SEQUENCING PIPELINE EVALUATION AND MUTATION DETECTION IN ESOPHAGEAL CANCER PATIENTS
 WHOLE EXOME SEQUENCING PIPELINE EVALUATION AND MUTATION DETECTION IN ESOPHAGEAL CANCER PATIENTS SUMMARY Tran Thi Bich Ngoc 1 ; Ho Viet Hoanh 2 ; Vu Phuong Nhung 1 ; Nguyen Hai Ha 1 Nguyen Van Ba 2 ; Nguyen
WHOLE EXOME SEQUENCING PIPELINE EVALUATION AND MUTATION DETECTION IN ESOPHAGEAL CANCER PATIENTS SUMMARY Tran Thi Bich Ngoc 1 ; Ho Viet Hoanh 2 ; Vu Phuong Nhung 1 ; Nguyen Hai Ha 1 Nguyen Van Ba 2 ; Nguyen
UNIVERSITY OF TORINO DEPARTMENT OF ONCOLOGY. Giorgio V. Scagliotti University of Torino Dipartment of Oncology
 Giorgio V. Scagliotti University of Torino Dipartment of Oncology giorgio.scagliotti@unito.it Molecular landscape of MM not fully characterized to allow personalized treatment Recurrent genetic alterations
Giorgio V. Scagliotti University of Torino Dipartment of Oncology giorgio.scagliotti@unito.it Molecular landscape of MM not fully characterized to allow personalized treatment Recurrent genetic alterations
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Systematic investigation of cancer-associated somatic point mutations in SNP databases HyunChul Jung 1,2, Thomas Bleazard 3, Jongkeun Lee 1 and Dongwan Hong 1 1. Cancer Genomics
SUPPLEMENTARY INFORMATION Systematic investigation of cancer-associated somatic point mutations in SNP databases HyunChul Jung 1,2, Thomas Bleazard 3, Jongkeun Lee 1 and Dongwan Hong 1 1. Cancer Genomics
The Role of Next Generation Sequencing in Solid Tumor Mutation Testing
 The Role of Next Generation Sequencing in Solid Tumor Mutation Testing Allie H. Grossmann MD PhD Department of Pathology, University of Utah Division of Anatomic Pathology & Oncology, ARUP Laboratories
The Role of Next Generation Sequencing in Solid Tumor Mutation Testing Allie H. Grossmann MD PhD Department of Pathology, University of Utah Division of Anatomic Pathology & Oncology, ARUP Laboratories
Variant interpretation exercise. ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18
 Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Frequency(%) KRAS G12 KRAS G13 KRAS A146 KRAS Q61 KRAS K117N PIK3CA H1047 PIK3CA E545 PIK3CA E542K PIK3CA Q546. EGFR exon19 NFS-indel EGFR L858R
 Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Functional analysis of DNA variants
 Functional analysis of DNA variants GS011143, Introduction to Bioinformatics The University of Texas GSBS program, Fall 2012 Ken Chen, Ph.D. Department of Bioinformatics and Computational Biology UT MD
Functional analysis of DNA variants GS011143, Introduction to Bioinformatics The University of Texas GSBS program, Fall 2012 Ken Chen, Ph.D. Department of Bioinformatics and Computational Biology UT MD
Supplementary Information
 1 Supplementary Information MATERIALS AND METHODS Study subjects Exome-sequenced HBOC patients Clinical characteristics of 24 exome-sequenced HBOC patients are presented in Table 1. Table 1. Clinical characteristics
1 Supplementary Information MATERIALS AND METHODS Study subjects Exome-sequenced HBOC patients Clinical characteristics of 24 exome-sequenced HBOC patients are presented in Table 1. Table 1. Clinical characteristics
VARIANT PRIORIZATION AND ANALYSIS INCORPORATING PROBLEMATIC REGIONS OF THE GENOME ANIL PATWARDHAN
 VARIANT PRIORIZATION AND ANALYSIS INCORPORATING PROBLEMATIC REGIONS OF THE GENOME ANIL PATWARDHAN Email: apatwardhan@personalis.com MICHAEL CLARK Email: michael.clark@personalis.com ALEX MORGAN Email:
VARIANT PRIORIZATION AND ANALYSIS INCORPORATING PROBLEMATIC REGIONS OF THE GENOME ANIL PATWARDHAN Email: apatwardhan@personalis.com MICHAEL CLARK Email: michael.clark@personalis.com ALEX MORGAN Email:
Exomes and Beyond Addressing the Genome with OneSeq. Madhuri Hegde, PhD, FACMG Emory University Emory Genetics Laboratory Atlanta, GA
 Exomes and Beyond Addressing the Genome with OneSeq Madhuri Hegde, PhD, FACMG Emory University Emory Genetics Laboratory Atlanta, GA Technologies to Detect Various Types of Mutations RESOLUTION Targeted
Exomes and Beyond Addressing the Genome with OneSeq Madhuri Hegde, PhD, FACMG Emory University Emory Genetics Laboratory Atlanta, GA Technologies to Detect Various Types of Mutations RESOLUTION Targeted
Implementation of the DDD/ClinGen OGT (CytoSure v3) Microarray
 Implementation of the DDD/ClinGen OGT (CytoSure v3) Microarray OGT UGM Birmingham 08/09/2016 Dom McMullan Birmingham Women's NHS Trust WM chromosomal microarray (CMA) testing Population of ~6 million (10%)
Implementation of the DDD/ClinGen OGT (CytoSure v3) Microarray OGT UGM Birmingham 08/09/2016 Dom McMullan Birmingham Women's NHS Trust WM chromosomal microarray (CMA) testing Population of ~6 million (10%)
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep
 Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
AD (Leave blank) TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients
 AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
Computational genomics and genetics of developmental disorders
 Computational genomics and genetics of developmental disorders Hongjian Qi Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and
Computational genomics and genetics of developmental disorders Hongjian Qi Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and
MEDICAL GENOMICS LABORATORY. Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG)
 Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
Supplementary Information. Data Identifies FAN1 at 15q13.3 as a Susceptibility. Gene for Schizophrenia and Autism
 Supplementary Information A Scan-Statistic Based Analysis of Exome Sequencing Data Identifies FAN1 at 15q13.3 as a Susceptibility Gene for Schizophrenia and Autism Iuliana Ionita-Laza 1,, Bin Xu 2, Vlad
Supplementary Information A Scan-Statistic Based Analysis of Exome Sequencing Data Identifies FAN1 at 15q13.3 as a Susceptibility Gene for Schizophrenia and Autism Iuliana Ionita-Laza 1,, Bin Xu 2, Vlad
The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation
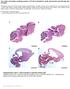 The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation Anita Becker-Heck#, Irene Zohn#, Noriko Okabe#, Andrew Pollock#, Kari Baker Lenhart,
The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation Anita Becker-Heck#, Irene Zohn#, Noriko Okabe#, Andrew Pollock#, Kari Baker Lenhart,
Molecular Diagnostic Laboratory 18 Sequencing St, Gene Town, ZY Tel: Fax:
 Molecular Diagnostic Laboratory 18 Sequencing St, Gene Town, ZY 01234 Tel: 555-920-3333 Fax: 555-920-3334 www.moldxlaboratory.com Patient Name: Jane Doe Specimen type: Blood, peripheral DOB: 04/05/1990
Molecular Diagnostic Laboratory 18 Sequencing St, Gene Town, ZY 01234 Tel: 555-920-3333 Fax: 555-920-3334 www.moldxlaboratory.com Patient Name: Jane Doe Specimen type: Blood, peripheral DOB: 04/05/1990
Immunogenetics of juvenile idiopathic arthritis: A comprehensive review
 Immunogenetics of juvenile idiopathic arthritis: A comprehensive review Aimee O. Hersh, University of Utah School of Medicine Sampath Prahalad, Emory University Journal Title: Journal of Autoimmunity Volume:
Immunogenetics of juvenile idiopathic arthritis: A comprehensive review Aimee O. Hersh, University of Utah School of Medicine Sampath Prahalad, Emory University Journal Title: Journal of Autoimmunity Volume:
Molecular Characterization of Tumors Using Next-Generation Sequencing
 Molecular Characterization of Tumors Using Next-Generation Sequencing Using BaseSpace to visualize molecular changes in cancer. Tumor-Normal Sequencing Data in BaseSpace To enable researchers new to next-generation
Molecular Characterization of Tumors Using Next-Generation Sequencing Using BaseSpace to visualize molecular changes in cancer. Tumor-Normal Sequencing Data in BaseSpace To enable researchers new to next-generation
Personalis ACE Clinical Exome The First Test to Combine an Enhanced Clinical Exome with Genome- Scale Structural Variant Detection
 Personalis ACE Clinical Exome The First Test to Combine an Enhanced Clinical Exome with Genome- Scale Structural Variant Detection Personalis, Inc. 1350 Willow Road, Suite 202, Menlo Park, California 94025
Personalis ACE Clinical Exome The First Test to Combine an Enhanced Clinical Exome with Genome- Scale Structural Variant Detection Personalis, Inc. 1350 Willow Road, Suite 202, Menlo Park, California 94025
CHAPTER IV RESULTS Microcephaly General description
 47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
MSI positive MSI negative
 Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Role of next generation sequencing in clinical care
 Role of next generation sequencing in clinical care Anne Slavotinek Division of Genetics, Department of Pediatrics, UCSF Structure 1. Next generation technologies 2. Consent and secondary findings 3. Genetics
Role of next generation sequencing in clinical care Anne Slavotinek Division of Genetics, Department of Pediatrics, UCSF Structure 1. Next generation technologies 2. Consent and secondary findings 3. Genetics
Variant interpretation using the ACMG-AMP guidelines
 Variant interpretation using the ACMG-AMP guidelines 20 TH A P R I L 2 0 1 7 S T A C E Y D A N I E L S W E S S E X R E G I O N A L G E N E T I C S L A B O R A T O R Y Talk outline Overview of the ACMG-AMP
Variant interpretation using the ACMG-AMP guidelines 20 TH A P R I L 2 0 1 7 S T A C E Y D A N I E L S W E S S E X R E G I O N A L G E N E T I C S L A B O R A T O R Y Talk outline Overview of the ACMG-AMP
Genotype-Phenotype in Egyptian Patients with Nephropathic Cystinosis. (December 2012 report)
 Genotype-Phenotype in Egyptian Patients with Nephropathic Cystinosis (December 2012 report) This is the first study of the genotype of Nephropathic Cystinosis (NC) patients in Egypt and the region of North
Genotype-Phenotype in Egyptian Patients with Nephropathic Cystinosis (December 2012 report) This is the first study of the genotype of Nephropathic Cystinosis (NC) patients in Egypt and the region of North
RNA-Seq guided gene therapy for vision loss. Michael H. Farkas
 RNA-Seq guided gene therapy for vision loss Michael H. Farkas The retina is a complex tissue Many cell types Neural retina vs. RPE Each highly dependent on the other Graw, Nature Reviews Genetics, 2003
RNA-Seq guided gene therapy for vision loss Michael H. Farkas The retina is a complex tissue Many cell types Neural retina vs. RPE Each highly dependent on the other Graw, Nature Reviews Genetics, 2003
NGS panels in clinical diagnostics: Utrecht experience. Van Gijn ME PhD Genome Diagnostics UMCUtrecht
 NGS panels in clinical diagnostics: Utrecht experience Van Gijn ME PhD Genome Diagnostics UMCUtrecht 93 Gene panels UMC Utrecht Cardiovascular disease (CAR) (5 panels) Epilepsy (EPI) (11 panels) Hereditary
NGS panels in clinical diagnostics: Utrecht experience Van Gijn ME PhD Genome Diagnostics UMCUtrecht 93 Gene panels UMC Utrecht Cardiovascular disease (CAR) (5 panels) Epilepsy (EPI) (11 panels) Hereditary
Intron p Hybrid with p.leu245val p. Gly336Cys. p.leu62phe p. Ala137Val p. Asn152Thr hybrid with p.leu245val p.
 RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
Supplementary Figure 1 Overall study design
 Supplementary Figure 1 Overall study design We obtained 19 tumour specimens from 14 men with prostate cancer who harboured pathogenic germline BRCA2-mutations. Germline DNA was available for five patients
Supplementary Figure 1 Overall study design We obtained 19 tumour specimens from 14 men with prostate cancer who harboured pathogenic germline BRCA2-mutations. Germline DNA was available for five patients
variant led to a premature stop codon p.k316* which resulted in nonsense-mediated mrna decay. Although the exact function of the C19L1 is still
 157 Neurological disorders primarily affect and impair the functioning of the brain and/or neurological system. Structural, electrical or metabolic abnormalities in the brain or neurological system can
157 Neurological disorders primarily affect and impair the functioning of the brain and/or neurological system. Structural, electrical or metabolic abnormalities in the brain or neurological system can
Leveraging the Exome Sequencing Project: Creating a WHI Resource Through Exome Imputation in SHARe. Chris Carlson on behalf of WHISP May 5, 2011
 Leveraging the Exome Sequencing Project: Creating a WHI Resource Through Exome Imputation in SHARe Chris Carlson on behalf of WHISP May 5, 2011 Exome Sequencing Project (ESP) Three cohort-based groups
Leveraging the Exome Sequencing Project: Creating a WHI Resource Through Exome Imputation in SHARe Chris Carlson on behalf of WHISP May 5, 2011 Exome Sequencing Project (ESP) Three cohort-based groups
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Complete Genomics, Inc.
 Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery
 CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery PacBio CoLab Session October 20, 2017 For Research Use Only. Not for use in diagnostics procedures. Copyright 2017 by Pacific Biosciences
CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery PacBio CoLab Session October 20, 2017 For Research Use Only. Not for use in diagnostics procedures. Copyright 2017 by Pacific Biosciences
Nature Genetics: doi: /ng Supplementary Figure 1. Country distribution of GME samples and designation of geographical subregions.
 Supplementary Figure 1 Country distribution of GME samples and designation of geographical subregions. GME samples collected across 20 countries and territories from the GME. Pie size corresponds to the
Supplementary Figure 1 Country distribution of GME samples and designation of geographical subregions. GME samples collected across 20 countries and territories from the GME. Pie size corresponds to the
Benefits and pitfalls of new genetic tests
 Benefits and pitfalls of new genetic tests Amanda Krause Division of Human Genetics, NHLS and University of the Witwatersrand Definition of Genetic Testing the analysis of human DNA, RNA, chromosomes,
Benefits and pitfalls of new genetic tests Amanda Krause Division of Human Genetics, NHLS and University of the Witwatersrand Definition of Genetic Testing the analysis of human DNA, RNA, chromosomes,
NGS in tissue and liquid biopsy
 NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
A Likelihood-Based Framework for Variant Calling and De Novo Mutation Detection in Families
 A Likelihood-Based Framework for Variant Calling and De Novo Mutation Detection in Families Bingshan Li 1 *, Wei Chen 2, Xiaowei Zhan 3, Fabio Busonero 3,4, Serena Sanna 4, Carlo Sidore 4, Francesco Cucca
A Likelihood-Based Framework for Variant Calling and De Novo Mutation Detection in Families Bingshan Li 1 *, Wei Chen 2, Xiaowei Zhan 3, Fabio Busonero 3,4, Serena Sanna 4, Carlo Sidore 4, Francesco Cucca
Germline Testing for Hereditary Cancer with Multigene Panel
 Germline Testing for Hereditary Cancer with Multigene Panel Po-Han Lin, MD Department of Medical Genetics National Taiwan University Hospital 2017-04-20 Disclosure No relevant financial relationships with
Germline Testing for Hereditary Cancer with Multigene Panel Po-Han Lin, MD Department of Medical Genetics National Taiwan University Hospital 2017-04-20 Disclosure No relevant financial relationships with
Clinical Genetics. Functional validation in a diagnostic context. Robert Hofstra. Leading the way in genetic issues
 Clinical Genetics Leading the way in genetic issues Functional validation in a diagnostic context Robert Hofstra Clinical Genetics Leading the way in genetic issues Future application of exome sequencing
Clinical Genetics Leading the way in genetic issues Functional validation in a diagnostic context Robert Hofstra Clinical Genetics Leading the way in genetic issues Future application of exome sequencing
SVIM: Structural variant identification with long reads DAVID HELLER MAX PLANCK INSTITUTE FOR MOLECULAR GENETICS, BERLIN JUNE 2O18, SMRT LEIDEN
 SVIM: Structural variant identification with long reads DAVID HELLER MAX PLANCK INSTITUTE FOR MOLECULAR GENETICS, BERLIN JUNE 2O18, SMRT LEIDEN Structural variation (SV) Variants larger than 50bps Affect
SVIM: Structural variant identification with long reads DAVID HELLER MAX PLANCK INSTITUTE FOR MOLECULAR GENETICS, BERLIN JUNE 2O18, SMRT LEIDEN Structural variation (SV) Variants larger than 50bps Affect
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13394 Genome sequencing identifies major causes of severe intellectual disability Table of Contents Participants...4 Patient recruitment... 4 Patient selection... 4 Cohort representativeness...
doi:10.1038/nature13394 Genome sequencing identifies major causes of severe intellectual disability Table of Contents Participants...4 Patient recruitment... 4 Patient selection... 4 Cohort representativeness...
Supplemental Data. De Novo Truncating Mutations in WASF1. Cause Intellectual Disability with Seizures
 The American Journal of Human Genetics, Volume 13 Supplemental Data De Novo Truncating Mutations in WASF1 Cause Intellectual Disability with Seizures Yoko Ito, Keren J. Carss, Sofia T. Duarte, Taila Hartley,
The American Journal of Human Genetics, Volume 13 Supplemental Data De Novo Truncating Mutations in WASF1 Cause Intellectual Disability with Seizures Yoko Ito, Keren J. Carss, Sofia T. Duarte, Taila Hartley,
CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY
 CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY More genetic information requires cutting-edge interpretation techniques Whole Exome Sequencing For certain patients the combination of symptoms does not allow
CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY More genetic information requires cutting-edge interpretation techniques Whole Exome Sequencing For certain patients the combination of symptoms does not allow
Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) cancer DNA
 www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
UW359 Ovary 3c 3 Serous Recurrent 68 BRCA1 816delGT BRCA1 del exon 1-2. UW417 Ovary 3c 3 Serous Primary 38 BRCA1 1675delA
 Supplementary Table 1. Cases with deleterious germline mutations, somatic HR mutations, and somatic PTEN mutations. ID Site Stage Grade Histology Tumor Age Germline mutation(s) a Somatic HR mutation(s)
Supplementary Table 1. Cases with deleterious germline mutations, somatic HR mutations, and somatic PTEN mutations. ID Site Stage Grade Histology Tumor Age Germline mutation(s) a Somatic HR mutation(s)
CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY
 CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY More genetic information requires cutting-edge interpretation techniques Whole Exome Sequencing For some patients, the combination of symptoms does not allow
CentoXome FUTURE'S KNOWLEDGE APPLIED TODAY More genetic information requires cutting-edge interpretation techniques Whole Exome Sequencing For some patients, the combination of symptoms does not allow
