Pan-cancer network analysis identifies combinations of rare somatic mutations across pathways and protein complexes
|
|
|
- Pierce Sherman
- 6 years ago
- Views:
Transcription
1 Pan-cancer network analysis identifies combinations of rare somatic mutations across pathways and protein complexes Nature Genetics 47, (2015) doi:101038/ng3168 Max Leiserson RECOMB 2015 April 14, 2015 Raphael Lab Department of Computer Science & Center for Computational Molecular Biology
2 Identifying cancer driver genes Cancer Genome Landscapes >999% of mutations are passengers Vogelstein et al (2013) 3-8 drivers per tumor" 2
3 Identifying cancer driver genes Cancer Genome Landscapes Vogelstein et al (2013) >999% of mutations are passengers 3-8 drivers per tumor" Compare variation across tumors Tumor 1 Tumor 2 Tumor N = gene = mutation Single nucleotide variants Copy number aberrations Gene expression 2
4 Identifying cancer driver genes Cancer Genome Landscapes Vogelstein et al (2013) >999% of mutations are passengers 3-8 drivers per tumor" Compare variation across tumors = gene Tumor 1 Tumor 2 Tumor N = mutation Single nucleotide variants Copy number aberrations Gene expression Significance Score Identify cancer driver genes genes Gene Mutations weighted by: Recurrence Gene length Mutation context Expression level Replication timing N= samples long tail 2
5 Cancer driver mutations target pathways Driver mutations confer a growth advantage to the tumor driver genes are members of cancer signaling pathways 3
6 Cancer driver mutations target pathways Driver mutations confer a growth advantage to the tumor driver genes are members of cancer signaling pathways STAT PI3K MAPK TGF- RAS DNA damage control Transcriptional regulation Cell cycle/ apoptosis Selective growth advantage NOTCH Chromatin modification HH APC Vogelstein et al (Science 2013) Genome Maintenance 3
7 Cancer driver mutations target pathways Driver mutations confer a growth advantage to the tumor driver genes are members of cancer signaling pathways STAT PI3K MAPK TGF- RAS DNA damage control Transcriptional regulation Cell cycle/ apoptosis Selective growth advantage NOTCH Chromatin modification HH APC Vogelstein et al (Science 2013) Genome Maintenance Significance Score genes A B N= samples A B interacts with 3
8 Cancer driver mutations target pathways Driver mutations confer a growth advantage to the tumor driver genes are members of cancer signaling pathways STAT PI3K MAPK TGF- RAS DNA damage control Transcriptional regulation Cell cycle/ apoptosis Selective growth advantage NOTCH Chromatin modification HH APC Vogelstein et al (Science 2013) Genome Maintenance Significance Score genes A B N= samples A B interacts with TCGA Ovarian (Nature 2011) 3
9 Testing known gene sets and pathways Input data Mutation data Tumor 1 Tumor 2 Tumor N (eg most mutated genes: EGFR, KRAS, BRAF) Gene set database PI3K RAS Cell cycle/ apoptosis NOTCH STAT MAPK TGF- Selective growth advantage DNA damage control Transcriptional regulation Chromatin modification HH APC Genome Maintenance 4
10 Testing known gene sets and pathways Input data Mutation data Tumor 1 Tumor 2 Tumor N (eg most mutated genes: EGFR, KRAS, BRAF) Enrichment tests GSEA [1,2] DAVID [3,4] Gene set database PI3K RAS Cell cycle/ apoptosis NOTCH STAT MAPK TGF- Selective growth advantage DNA damage control Transcriptional regulation Chromatin modification HH APC Genome Maintenance [1] Mootha et al Nat Genet (2003) [3] Huang et al Nat Protoc (2009) [2] Subramanian et al PNAS (2005) [4] Huang et al Nucleic Acids Res (2009) 4
11 Testing known gene sets and pathways Input data Mutation data Tumor 1 Tumor 2 Tumor N (eg most mutated genes: EGFR, KRAS, BRAF) Enrichment tests GSEA [1,2] DAVID [3,4] Enriched gene sets MYC ARID1A FBXW7 APC SOX9 CTNNB1 Gene set database PI3K RAS Cell cycle/ apoptosis NOTCH STAT MAPK TGF- Selective growth advantage DNA damage control Transcriptional regulation Chromatin modification HH APC Genome Maintenance [1] Mootha et al Nat Genet (2003) [3] Huang et al Nat Protoc (2009) [2] Subramanian et al PNAS (2005) [4] Huang et al Nucleic Acids Res (2009) 4
12 Testing known gene sets and pathways Input data Mutation data Tumor 1 Tumor 2 Tumor N (eg most mutated genes: EGFR, KRAS, BRAF) Enrichment tests GSEA [1,2] DAVID [3,4] Enriched gene sets MYC ARID1A FBXW7 APC SOX9 CTNNB1 Gene set database STAT PI3K MAPK TGF- RAS DNA damage control Genome Maintenance Transcriptional regulation Cell cycle/ apoptosis Selective growth advantage NOTCH Chromatin modification HH APC Key drawbacks Novel pathways and crosstalk? Topology of interactions? Handling large and/or overlapping pathways? [1] Mootha et al Nat Genet (2003) [2] Subramanian et al PNAS (2005) [3] Huang et al Nat Protoc (2009) [4] Huang et al Nucleic Acids Res (2009) 4
13 Significantly mutated subnetworks of a protein-protein interaction network Protein-protein interaction networks Nodes: genes/protein Edges: connect genes if the proteins they encode physically interact Unweighted, undirected Goal: identify connected subnetworks with more mutations than expected by chance No Mutations 5
14 Significantly mutated subnetworks of a protein-protein interaction network Protein-protein interaction networks Nodes: genes/protein Edges: connect genes if the proteins they encode physically interact Unweighted, undirected Goal: identify connected subnetworks with more mutations than expected by chance No Mutations ~10 18 subnetworks of size k=5 5
15 Significantly mutated subnetworks of a protein-protein interaction network Protein-protein interaction networks Nodes: genes/protein Edges: connect genes if the proteins they encode physically interact Unweighted, undirected Network Nodes Edges Diameter ASP HPRD 9,205 36, HINT+HI2012 9,859 40, irefindex 12,129 91, MultiNet 14, , Goal: identify connected subnetworks with more mutations than expected by chance Low diameter Most genes have a high-scoring neighbor No Mutations ~10 18 subnetworks of size k=5 5
16 Significantly mutated subnetworks of a protein-protein interaction network Protein-protein interaction networks Nodes: genes/protein Edges: connect genes if the proteins they encode physically interact Unweighted, undirected Network Nodes Edges Diameter ASP HPRD 9,205 36, HINT+HI2012 9,859 40, irefindex 12,129 91, MultiNet 14, , Goal: identify connected subnetworks with more mutations than expected by chance Low diameter Most genes have a high-scoring neighbor Must analyze mutations and local topology simultaneously! No Mutations ~10 18 subnetworks of size k=5 5
17 Outline 1 A new algorithm, HotNet2 2 Application to TCGA Pan-Cancer data A101D A101* A101E L106T P179V E187G R252L V276T 3 Comparison of HotNet2 to other methods 6
18 Encoding mutations and graph topology with heat diffusion Heat Heat diffusion process t=0 t=1 t=n u u u Place heat on node u Heat diffuses from node u to u's neighbors Related to random walks and network propagation 7
19 Encoding mutations and graph topology with heat diffusion Heat Heat diffusion process t=0 t=1 t=n u u u Place heat on node u Heat diffuses from node u to u's neighbors Related to random walks and network propagation HotNet (Vandin et al JCB & RECOMB 2010) Mutations = heat sources Heat diffusion Extract hot subnetworks = mutated gene 7
20 HotNet applied to TCGA data TCGA Papers (~300 samples) Leukemia (NEJM 2013) Kidney (Nature 2011) Ovarian (Nature 2011) HotNet (Vandin et al JCB & RECOMB 2010) Mutations = heat sources Heat diffusion Extract hot subnetworks = mutated gene 8
21 HotNet applied to TCGA data TCGA Pan-Cancer (>3000 samples) TCGA Papers (~300 samples) Leukemia (NEJM 2013) Kidney (Nature 2011) Ovarian (Nature 2011) u HotNet finds many star subnetworks with one central, hot node HotNet (Vandin et al JCB & RECOMB 2010) Mutations = heat sources Heat diffusion Extract hot subnetworks = mutated gene 8
22 HotNet Algorithm Input Output h1 hn A = adjacency matrix h = gene scores Connected components Threshold at δ Heat kernel f(a, t) h1 0 0 hn = s11 sn1 s1n snn symmetrize r11 rn1 r1n rnn Diffusion matrix (symmetric) Time parameter t Similarity matrix (asymmetric) sij = heat on vertex i at time t given initial heat hj on vertex j at time 0 9
23 Direction of heat is important HotNet can fail HotNet s heat is symmetric Potential artifacts u sends the same heat to v even though u has much higher degree v u u Hot nodes with high degree often form large star subnetworks with many cold nodes Star graph Heat kernel f(a, t) h1 0 0 hn = s11 sn1 s1n snn symmetrize r11 rn1 r1n rnn Diffusion matrix (symmetric) Similarity matrix S (asymmetric) sij = heat on vertex i at time t given initial heat hj on vertex j at time 0 10
24 HotNet2 algorithm (HotNet diffusion oriented subnetworks) Need to consider the source of heat Encode directionality with asymmetric heat diffusion Hot genes do not necessarily implicate their neighbors Hot subnetworks have a directed path between each pair of nodes sij = heat on vertex i at equilibrium given initial heat hj on vertex j at time 0 Similarity Identify strongly connected components Similarity matrix S Asymmetric! similarity of u and v Leiserson, Vandin et al Nat Genet (2015) 11
25 HotNet2 algorithm (HotNet diffusion oriented subnetworks) Need to consider the source of heat Encode directionality with asymmetric heat diffusion Hot genes do not necessarily implicate their neighbors Hot subnetworks have a directed path between each pair of nodes sij = heat on vertex i at equilibrium given initial heat hj on vertex j at time 0 Similarity Identify strongly connected components Similarity matrix S Asymmetric! Leiserson, Vandin et al Nat Genet (2015) similarity of u and v TCGA Papers Gastric, Nature (2014) Thyroid, Cell (2014) 11
26 HotNet2 vs HotNet Input HotNet2 A = adjacency matrix h1 hn h = gene scores Strongly connected components Threshold at δ HotNet: Heat kernel f(a, t) HotNet2: Insulated heat diffusion Diffusion matrix (HotNet: symmetric HotNet2: asymmetric) h1 0 0 hn = s11 sn1 s1n snn Similarity matrix (asymmetric) symmetrize Connected components r11 rn1 r1n rnn Threshold at δ HotNet 12
27 Statistical test Evaluate graph partition with rigorously bounded False Discovery Rate (FDR) Gene scores Randomized gene scores x2 = 3 x3 = 1 x2 = 1 x3 = 0 Leiserson, Vandin et al Nat Genet (2015) Xk: number of subnetworks of size k Pr(Xk xk h, δ) 13
28 Outline 1 A new algorithm, HotNet2 2 Application to TCGA Pan-Cancer data A101D A101* A101E L106T P179V E187G R252L V276T 3 Comparison of HotNet2 to other methods 14
29 TCGA Pan-Cancer Cancer Tumor samples BLCA 99 BRCA 772 COAD/READ 224 GBM 291 HNSC 306 KIRC 417 LAML 196 LUAD 230 LUSC 178 OV 316 UCEC 248 Samples Color 3,110 tumors of 12 cancer types Weinstein et al Nature Genetics (2013) Single nucleotide variants A101D A101* A101E L106T Number of CNAs P179V E187G CCND1 MYC ERBB2 EGFR CDKN2A R252L KRAS MLL3 APC VHL Mutations V276T PTEN 11,565 mutated, expressed genes PIK3CA SNVs and CNAs in 3,110 samples among 11,565 expressed genes Hot Cold ,000 1,100 1,200 1,300 Number of SNVs Copy number aberrations Patient Chromosome arm Hot Cold TP53 15
30 HotNet2 runs on TCGA Pan-Cancer dataset Mutation & copy number data A101D A101* A101E L106T P179V E187G R252L V276T Interaction network Significantly mutated subnetworks Patient HotNet2 Chromosome arm 11,565 mutated genes in 3,110 samples 16
31 HotNet2 runs on TCGA Pan-Cancer dataset Mutation & copy number data A101D A101* A101E L106T P179V E187G R252L V276T Interaction network Significantly mutated subnetworks Patient HotNet2 Chromosome arm 11,565 mutated genes in 3,110 samples HINT+HI2012 (P < 001) irefindex 90 (P < 001) Multinet (P < 001) 40,704 interactions 9,858 proteins 91,808 interactions 12,128 proteins 109,569 interactions 14,398 proteins 16
32 HotNet2 runs on TCGA Pan-Cancer dataset Mutation & copy number data A101D A101* A101E L106T P179V E187G R252L V276T Interaction network Significantly mutated subnetworks Patient HotNet2 Chromosome arm 11,565 mutated genes in 3,110 samples HINT+HI2012 (P < 001) irefindex 90 (P < 001) Multinet (P < 001) 40,704 interactions 9,858 proteins 91,808 interactions 12,128 proteins 109,569 interactions 14,398 proteins Consensus subnetworks 16 consensus subnetworks with 4 genes (P=0004) 13 linkers between consensus subnetworks 16
33 HotNet2 Consensus HotNet2 Runs HINT+HI2012 (P < 001) irefindex 90 (P < 001) Multinet (P < 001) 1 Interaction networks Many low-confidence edges 075 TPR = Sensitivity High confidence edges High- and some lowconfidence edges FPR = 1-Specificity
34 HotNet2 Consensus HotNet2 Runs HINT+HI2012 (P < 001) irefindex 90 (P < 001) Multinet (P < 001) TPR = Sensitivity Interaction networks Multinet (110K edges)? irefindex (90K edges)?? HINT+HI2012 (40K edges) FPR = 1-Specificity
35 HotNet2 Consensus HotNet2 Runs HINT+HI2012 (P < 001) irefindex 90 (P < 001) Multinet (P < 001) Consensus 16 consensus subnetworks 13 linkers between subnetworks TPR = Sensitivity Interaction networks Multinet (110K edges)?? irefindex (90K edges) HINT+HI2012 (40K edges)? 3 Main Idea: Incorporate lowconfidence edges but give highconfidence edges more weight Consensus Linkers Consensus FPR = 1-Specificity Consensus Graph Edges connect genes identified by HotNet2 in the same subnetwork 17
36 HotNet2 Consensus Subnetworks GBM LAML Frequently and rarely mutated cancer genes OV ErbB signaling (179%) EGFR ERBB2 ERBB4 **OSMR **LRIG1 *ELF3 **AREG Core-binding factors(27%) RUNX1 CBFB **ELF4 BRCA P53 signaling(684%) TP53 CDKN2A MDM4 CCND1 PTEN NPM1 39 Others Linkers (212%) TP53INP1 WT1 **CCDC88A **RNF20 **CABLES1 *NOTCH2 PI3K signaling(326%) PIK3CA NRAS KRAS HNSC LUSC NOTCH signaling(198%) MHC Class I (312%) *HLA-A NOTCH1 **HLA-B **P4HTM B2M **CD1D *NOTCH3 **NOTCH4 **JAG1 CCNE1 **MAML111 Others CLASP and CLIP (2%) **CLIP2 **CLASP2 **CLASP1 ASCOM complex (1691%) ASXL1 MLL3 **FOXK2 MLL2 **N4BP2 KDM6A **PROSER1 **KDM1B *E2F3 BAP1 complex (71%) BAP1 **ASXL2 **ANKRD17 *FOXK1 Cohesin complex (736%) PIK3R1 MAP2K4 MAP3K1 HRAS *RASA1 ATM STK11 (19%) **BOC **CDON BRAF 8 Others SWI/SNF complex (168%) ARID1A PBRM1 SMARCA4 ARID2 *ARID1B **ADNP SMARCB1 Condensin Complex (42%) *SMC4 **NCAPG2 **NCAPD3 **NCAPH2 **NCAPD2 **SMC2 (07%) SPOP MYD88 LUAD UCEC STAG2 **PDS5A **STAG1 *SMC1A *RAD21 KEAP1, NFE2L2 (849%) KEAP1 NFE2L2 **CHD6 **WAPAL **PDS5B **PTMA **WAC BLCA COADREAD KIRC 18
37 HotNet2 Consensus Subnetworks HNSC LUSC OV *HLA-A **HLA-B B2M **P4HTM **CD1D GBM NOTCH signaling(198%) MHC Class I (312%) CLASP and CLIP (2%) **CLIP2 **CLASP2 **CLASP1 ASCOM complex (1691%) ASXL1 MLL3 **FOXK2 MLL2 **N4BP2 KDM6A **PROSER1 **KDM1B *E2F3 P53 signaling(684%) ErbB signaling (179%) EGFR ERBB2 ERBB4 **OSMR **LRIG1 BAP1 complex (71%) BAP1 **ASXL2 **ANKRD17 *FOXK1 Cohesin complex (736%) STAG2 **PDS5A **STAG1 *SMC1A *RAD21 LAML *ELF3 **AREG CBFB Linkers (212%) TP53INP1 WT1 TP53 CDKN2A MDM4 **CCDC88A CCND1 PTEN **RNF20 NPM1 39 Others **CABLES1 *NOTCH2 PIK3R1 MAP2K4 Core-binding factors(27%) (19%) **BOC **CDON KEAP1, NFE2L2 (849%) KEAP1 NFE2L2 **CHD6 RUNX1 **ELF4 PI3K signaling(326%) PIK3CA NRAS KRAS BRAF 8 Others PBRM1 SMARCA4 ARID2 *ARID1B **ADNP SMARCB1 BRCA (07%) MAP3K1 NOTCH1 HRAS SPOP MYD88 *NOTCH3 **JAG1 *RASA1 **NOTCH4 CCNE1 ATM SWI/SNF complex (168%) **MAML111 Others STK11 ARID1A Condensin Complex (42%) *SMC4 **NCAPG2 **NCAPD3 **NCAPH2 **NCAPD2 **SMC2 LUAD UCEC Frequently and rarely mutated cancer genes Well-known cancer pathways PI(3)K signaling p53 signaling NOTCH signaling ErbB signaling Linkers: HRAS, STK11, ATM **WAPAL **PDS5B **PTMA **WAC BLCA COADREAD KIRC 18
38 HotNet2 Consensus Subnetworks HNSC LUSC OV GBM NOTCH signaling(198%) MHC Class I (312%) **HLA-B **P4HTM *HLA-A B2M **CD1D CLASP and CLIP (2%) **CLASP2 **CLIP2 MLL2 **CLASP1 ASCOM complex (1691%) **N4BP2 MLL3 *E2F3 KDM6A **PROSER1 ErbB signaling (179%) Cohesin complex (736%) STAG2 **PDS5A **STAG1 *SMC1A **OSMR P53 signaling(684%) CDKN2A CCND1 NPM1 *NOTCH3 TP53 **NOTCH4 NOTCH1 MDM4 PTEN 39 Others **JAG1 CCNE1 **MAML111 Others BAP1 complex (71%) ASXL1 **FOXK2 ERBB2 BAP1 **KDM1B EGFR ERBB4 **LRIG1 *ELF3 **AREG **ASXL2 **ANKRD17 *FOXK1 *RAD21 Linkers (212%) TP53INP1 WT1 **CCDC88A **RNF20 **CABLES1 *NOTCH2 PIK3R1 MAP2K4 MAP3K1 HRAS *RASA1 ATM STK11 **BOC LAML Core-binding factors(27%) (19%) KEAP1, NFE2L2 (849%) NFE2L2 CBFB **CDON KEAP1 **CHD6 RUNX1 **ELF4 PI3K signaling(326%) NRAS BRAF SWI/SNF complex (168%) PBRM1 ARID2 PIK3CA ARID1A KRAS 8 Others SMARCA4 *ARID1B **ADNP SMARCB1 BRCA SPOP Condensin Complex (42%) *SMC4 **NCAPG2 **NCAPD3 **NCAPH2 **NCAPD2 **SMC2 (07%) MYD88 LUAD UCEC Frequently and rarely mutated cancer genes Well-known cancer pathways PI(3)K signaling p53 signaling NOTCH signaling ErbB signaling Linkers: HRAS, STK11, ATM p53 Signaling CUL9 CHD8 NPM1 TP53 PTEN MDM4 CCND1 CDKN2A EPHA3 plus ~30 others Mutations Multinet irefindex HINT+HI2012 **WAPAL **PDS5B **PTMA **WAC BLCA COADREAD KIRC 18
39 HotNet2 Consensus Subnetworks HNSC LUSC OV **HLA-B **P4HTM *HLA-A B2M **CD1D BLCA GBM NOTCH signaling(198%) MHC Class I (312%) CLASP and CLIP (2%) **CLASP2 ASCOM complex (1691%) MLL2 **N4BP2 MLL3 *E2F3 KDM6A **PROSER1 BAP1 complex (71%) ASXL1 **FOXK2 BAP1 **KDM1B **ASXL2 **ANKRD17 *FOXK1 Cohesin complex (736%) STAG2 **PDS5A **STAG1 KIRC LAML KEAP1, NFE2L2 (849%) *SMC1A NFE2L2 **CHD6 *RAD21 **WAPAL **PDS5B **PTMA **WAC CHD8 (mutated in 54 of 3110 samples) Legend **CLIP2 Legend Cancer types **CLASP1 ErbB signaling (179%) ERBB2 **OSMR P53 signaling(684%) CDKN2A CCND1 NPM1 *NOTCH3 TP53 **NOTCH4 NOTCH1 MDM4 PTEN 39 Others **JAG1 CCNE1 **MAML111 Others EGFR ERBB4 **LRIG1 *ELF3 **AREG Linkers (212%) TP53INP1 **CCDC88A **RNF20 **CABLES1 *NOTCH2 PIK3R1 MAP2K4 MAP3K1 HRAS *RASA1 ATM STK11 WT1 **BOC Core-binding factors(27%) (19%) CBFB **CDON KEAP1 RUNX1 PI3K signaling(326%) SWI/SNF complex (168%) COADREAD **ELF4 NRAS BRAF PBRM1 ARID2 PIK3CA ARID1A KRAS 8 Others SMARCA4 *ARID1B **ADNP SMARCB1 *SMC4 **NCAPG2 **NCAPD3 **NCAPH2 **NCAPD2 **SMC2 BRCA SPOP Condensin Complex (42%) (07%) MYD88 BLCA BRCA COADREAD GBM HNSC KIRC LAML LUAD LUSC OV UCEC LUAD UCEC Frequently and rarely mutated cancer genes Well-known cancer pathways PI(3)K signaling p53 signaling NOTCH signaling ErbB signaling Linkers: HRAS, STK11, ATM p53 Signaling CUL9 CHD8 NPM1 TP53 Mutation types PTEN MDM4 CCND1 CDKN2A EPHA3 plus ~30 others Inactivating SNV Amplification Mutations Multinet irefindex HINT+HI2012 SNV Deletion 18
40 HotNet2 Consensus Subnetworks HNSC LUSC OV *HLA-A **HLA-B B2M **P4HTM **CD1D GBM NOTCH signaling(198%) MHC Class I (312%) CLASP and CLIP (2%) **CLIP2 **CLASP2 **CLASP1 ASCOM complex (1691%) ASXL1 MLL3 **FOXK2 MLL2 **N4BP2 KDM6A **PROSER1 **KDM1B *E2F3 P53 signaling(684%) ErbB signaling (179%) EGFR ERBB2 ERBB4 **OSMR **LRIG1 BAP1 complex (71%) BAP1 **ASXL2 **ANKRD17 *FOXK1 Cohesin complex (736%) STAG2 **PDS5A **STAG1 *SMC1A *RAD21 LAML *ELF3 **AREG CBFB Linkers (212%) TP53INP1 WT1 TP53 CDKN2A MDM4 **CCDC88A CCND1 PTEN **RNF20 NPM1 39 Others **CABLES1 *NOTCH2 PIK3R1 MAP2K4 Core-binding factors(27%) (19%) **BOC **CDON KEAP1, NFE2L2 (849%) KEAP1 NFE2L2 **CHD6 RUNX1 **ELF4 PI3K signaling(326%) PIK3CA NRAS KRAS BRAF 8 Others PBRM1 SMARCA4 ARID2 *ARID1B **ADNP SMARCB1 BRCA (07%) MAP3K1 NOTCH1 HRAS SPOP MYD88 *NOTCH3 **JAG1 *RASA1 **NOTCH4 CCNE1 ATM SWI/SNF complex (168%) **MAML111 Others STK11 ARID1A Condensin Complex (42%) *SMC4 **NCAPG2 **NCAPD3 **NCAPH2 **NCAPD2 **SMC2 LUAD UCEC Frequently and rarely mutated cancer genes Well-known cancer pathways PI(3)K signaling p53 signaling NOTCH signaling ErbB signaling Linkers: HRAS, STK11, ATM Recently characterized complexes: SWI/SNF complex ASCOM complex BAP1 complex **WAPAL **PDS5B **PTMA **WAC BLCA COADREAD KIRC 18
41 HotNet2 Consensus Subnetworks HNSC LUSC OV *HLA-A **HLA-B B2M **P4HTM **CD1D BLCA GBM NOTCH signaling(198%) MHC Class I (312%) CLASP and CLIP (2%) **CLIP2 **CLASP2 **CLASP1 ASCOM complex (1691%) ASXL1 MLL3 **FOXK2 MLL2 **N4BP2 KDM6A **PROSER1 **KDM1B *E2F3 P53 signaling(684%) ErbB signaling (179%) EGFR ERBB2 ERBB4 **OSMR **LRIG1 BAP1 complex (71%) BAP1 **ASXL2 **ANKRD17 *FOXK1 Cohesin complex (736%) STAG2 **PDS5A **STAG1 *SMC1A *RAD21 **WAPAL **PDS5B LAML *ELF3 **AREG CBFB Linkers (212%) TP53INP1 WT1 TP53 CDKN2A MDM4 **CCDC88A CCND1 PTEN **RNF20 NPM1 39 Others **CABLES1 *NOTCH2 PIK3R1 MAP2K4 Core-binding factors(27%) (19%) **BOC **CDON KEAP1, NFE2L2 (849%) KEAP1 NFE2L2 **CHD6 **PTMA **WAC RUNX1 **ELF4 PI3K signaling(326%) COADREAD PIK3CA NRAS KRAS BRAF 8 Others PBRM1 SMARCA4 ARID2 *ARID1B **ADNP SMARCB1 BRCA (07%) MAP3K1 NOTCH1 HRAS SPOP MYD88 *NOTCH3 **JAG1 *RASA1 **NOTCH4 CCNE1 ATM SWI/SNF complex (168%) **MAML111 Others STK11 ARID1A Condensin Complex (42%) *SMC4 **NCAPG2 **NCAPD3 **NCAPH2 **NCAPD2 **SMC2 LUAD UCEC Frequently and rarely mutated cancer genes Well-known cancer pathways PI(3)K signaling p53 signaling NOTCH signaling ErbB signaling Linkers: HRAS, STK11, ATM Recently characterized complexes: SWI/SNF complex ASCOM complex BAP1 complex Potentially novel complexes: Cohesin complex Condensin complex MHC Class I proteins KIRC 18
42 HotNet2 Consensus Subnetworks HNSC LUSC OV *HLA-A **HLA-B B2M **P4HTM **CD1D BLCA GBM NOTCH signaling(198%) MHC Class I (312%) CLASP and CLIP (2%) **CLIP2 **CLASP2 **CLASP1 ASCOM complex (1691%) ASXL1 MLL3 **FOXK2 MLL2 **N4BP2 KDM6A **PROSER1 **KDM1B *E2F3 P53 signaling(684%) ErbB signaling (179%) EGFR ERBB2 ERBB4 **OSMR **LRIG1 BAP1 complex (71%) BAP1 **ASXL2 **ANKRD17 *FOXK1 Cohesin complex (736%) STAG2 **PDS5A **STAG1 *SMC1A *RAD21 **WAPAL **PDS5B LAML *ELF3 **AREG CBFB Linkers (212%) TP53INP1 WT1 TP53 CDKN2A MDM4 **CCDC88A CCND1 PTEN **RNF20 NPM1 39 Others **CABLES1 *NOTCH2 PIK3R1 MAP2K4 Core-binding factors(27%) (19%) **BOC **CDON KEAP1, NFE2L2 (849%) KEAP1 NFE2L2 **CHD6 **PTMA **WAC RUNX1 **ELF4 PI3K signaling(326%) COADREAD PIK3CA NRAS KRAS BRAF 8 Others PBRM1 SMARCA4 ARID2 *ARID1B **ADNP SMARCB1 BRCA (07%) MAP3K1 NOTCH1 HRAS SPOP MYD88 *NOTCH3 **JAG1 *RASA1 **NOTCH4 CCNE1 ATM SWI/SNF complex (168%) **MAML111 Others STK11 ARID1A Condensin Complex (42%) *SMC4 **NCAPG2 **NCAPD3 **NCAPH2 **NCAPD2 **SMC2 LUAD UCEC Frequently and rarely mutated cancer genes Well-known cancer pathways PI(3)K signaling p53 signaling NOTCH signaling ErbB signaling Linkers: HRAS, STK11, ATM Recently characterized complexes: SWI/SNF complex ASCOM complex BAP1 complex Potentially novel complexes: Cohesin complex Condensin complex MHC Class I proteins KIRC 18
43 SWI/SNF complex a SWI / SNF Complex Coverage: 168% (523 / 3110 samples) = 5 samples ARID1A (194) PBRM1 (169) SMARCA4 (67) ARID2 (56) *ARID1B (41) **ADNP (21) SMARCB1 (19) SMARCA4 LUAD: 8x10-3 ARID1A UCEC: 158x10-15 BLCA: 488x10-9 PBRM1 KIRC: 7x10-97 S10T N80Y E226Q K231R I245F I252R P427S N453S F456S R522L M523R N528I Y580C E583K M586K M586I M586T N601K L614V L618H G626V I709T Y718C M731K R760H E860K R876L P1048L P1048R R1088Q V1136L T1202K G1231R C1233W E1248G E1287D I1309T D1325N R1411C W1417G M1539I G1554R G1577C L1611P R1670P ARID1B ADNP ARID2 SMARCB1 BLCA: 001 All PPI networks MultiNet irefindex HINT HI missense Scale (AA) PBRM1 BAH_dom Bromodomain HMG_superfamily Legend Legend Cancer types BLCA BRCA COADREAD GBM HNSC KIRC LAML LUAD LUSC OV UCEC Mutation types Inactivating SNV Amplification SNV Deletion 19
44 SWI/SNF complex a SWI / SNF Complex Coverage: 168% (523 / 3110 samples) = 5 samples ARID1A (194) PBRM1 (169) SMARCA4 (67) ARID2 (56) *ARID1B (41) **ADNP (21) SMARCB1 (19) SMARCA4 LUAD: 8x10-3 ARID1A UCEC: 158x10-15 BLCA: 488x10-9 PBRM1 KIRC: 7x10-97 S10T N80Y E226Q K231R I245F I252R P427S N453S F456S R522L M523R N528I Y580C E583K M586K M586I M586T N601K L614V L618H G626V I709T Y718C M731K R760H E860K R876L P1048L P1048R R1088Q V1136L T1202K G1231R C1233W E1248G E1287D I1309T D1325N R1411C W1417G M1539I G1554R G1577C L1611P R1670P ARID1B ADNP ARID2 SMARCB1 BLCA: 001 All PPI networks MultiNet irefindex HINT HI missense Scale (AA) PBRM1 BAH_dom Bromodomain HMG_superfamily Legend Legend Cancer types BLCA BRCA COADREAD GBM HNSC KIRC LAML LUAD LUSC OV UCEC Mutation types Inactivating SNV Amplification SNV Deletion 19
45 SWI/SNF complex a SWI / SNF Complex Coverage: 168% (523 / 3110 samples) = 5 samples ARID1A (194) PBRM1 (169) SMARCA4 (67) ARID2 (56) *ARID1B (41) **ADNP (21) SMARCB1 (19) SMARCA4 LUAD: 8x10-3 ARID1A UCEC: 158x10-15 BLCA: 488x10-9 PBRM1 KIRC: 7x10-97 S10T N80Y E226Q K231R I245F I252R P427S N453S F456S R522L M523R N528I Y580C E583K M586K M586I M586T N601K L614V L618H G626V I709T Y718C M731K R760H E860K R876L P1048L P1048R R1088Q V1136L T1202K G1231R C1233W E1248G E1287D I1309T D1325N R1411C W1417G M1539I G1554R G1577C L1611P R1670P ARID1B ADNP ARID2 SMARCB1 BLCA: 001 All PPI networks MultiNet irefindex HINT HI missense Scale (AA) PBRM1 BAH_dom Bromodomain HMG_superfamily Wilson and Roberts Nature Reviews Cancer (2011) Involved in nucleosome remodeling ACTL6A/B SMARCD 1/2/3 SMARCC1 PBAF SMARCA4 SMARCB1 SMARCC2 ARID2 PBRM1 DPF 1/2/3 SMARCE1 BRD7 ACTL6A/B SMARCD 1/2/3 SMARCC1 BAF SMARCA 2/4 SMARCB1 SMARCC2 ARID1A/B DPF 1/2/3 SMARCE1 Legend Legend Cancer types BLCA BRCA COADREAD GBM HNSC KIRC LAML LUAD LUSC OV UCEC Mutation types Inactivating SNV Amplification SNV Deletion 19
46 Cohesin and condensin complexes a Cohesin Complex Coverage: 74% (229 / 3110 samples) = 5 samples STAG2 (59) SMC1A (44) **PDS5B (33) **STAG1 (31) **PDS5A (26) *RAD21 (23) **WAPAL (23) STAG2 BLCA: 5x10-3 Cohesin complex 4/5 members of complex Involved in sister chromatid cohesion and gene regulation Mutated in >4% of samples in each cancer type STAG1 SMC1A H243N S290C A428V L525R K581E Q619R S637fs V666L A704P D706G Q815* N941I R958H R958C Q994K E995Q L1001fs L1001Q exon27+2 E1027K D1059H D1059A V1075L Q1116* RAD21 PDS5B WAPAL PDS5A All PPI networks MultiNet irefindex HINT+HI frame_shift_del missense nonsense splice_site Scale (AA) STAG1 STAG Legend Legend Cancer types BLCA BRCA COADREAD GBM HNSC KIRC LAML LUAD LUSC OV UCEC Mutation types Inactivating SNV Amplification SNV Deletion 20
47 Cohesin and condensin complexes a Cohesin Complex Coverage: 74% (229 / 3110 samples) = 5 samples STAG2 (59) SMC1A (44) **PDS5B (33) **STAG1 (31) **PDS5A (26) *RAD21 (23) **WAPAL (23) STAG2 BLCA: 5x10-3 Cohesin complex 4/5 members of complex Involved in sister chromatid cohesion and gene regulation Mutated in >4% of samples in each cancer type STAG1 SMC1A H243N S290C A428V L525R K581E Q619R S637fs V666L A704P D706G Q815* N941I R958H R958C Q994K E995Q L1001fs L1001Q exon27+2 E1027K D1059H D1059A V1075L Q1116* RAD21 PDS5B WAPAL PDS5A All PPI networks MultiNet irefindex HINT+HI frame_shift_del missense nonsense splice_site Scale (AA) STAG1 STAG b Condensin Complex Coverage: 42% (131 / 3110 samples) = 5 samples **NCAPD3 (35) *SMC4 (27) **NCAPG2 (23) **NCAPH2 (19) **SMC2 (17) **NCAPD2 (14) NCAPG2 LUSC: 0015 SMC4 NCAPD3 exon2-2 I145V R176W A205V T217S P252S A369V R378Q K398T Q408* E414* R453S P529L R551S R551P S556F G569E Condensin complex 6/8 members of complex Involved in sister chromatid condensation and gene regulation Somatic mutations and expression validated using whole-genome sequencing and RNA-Seq Legend Legend Cancer types NCAPH2 SMC2 NCAPD2 All PPI networks MultiNet irefindex HINT+HI2012 BLCA BRCA COADREAD GBM HNSC KIRC LAML LUAD LUSC OV UCEC missense nonsense splice_site Mutation types Scale (AA) NCAPH2 Condensin_II_H2-like Inactivating SNV Amplification SNV Deletion 20
48 Outline 1 A new algorithm, HotNet2 2 Application to TCGA Pan-Cancer data A101D A101* A101E L106T P179V E187G R252L V276T 3 Comparison of HotNet2 to similar methods 21
49 HotNet2 outperforms other methods on real data No gold standard dataset compare methods at identifying putative cancer genes Dataset of putative cancer genes Cancer genes have: 1 20% truncating mutations; or, 2 20% mutations clustered at a locus (c) Vogelstein et al (Science, 2013) TPR = Sensitivity FPR = 1-Specificity 22
50 Summary HotNet2: Novel algorithm that analyzes topology and mutations simultaneously with asymmetric heat diffusion Identifies known and novel pathways and complexes with frequently and rarely mutated genes on TCGA Pan-Cancer data Future work: Alternate graph partitioning algorithms? Other applications: gene expression, GWAS, social networks, etc 23
51 Summary HotNet2: Novel algorithm that analyzes topology and mutations simultaneously with asymmetric heat diffusion v u Identifies known and novel pathways and complexes with frequently and rarely mutated genes on TCGA Pan-Cancer data Future work: Alternate graph partitioning algorithms? Other applications: gene expression, GWAS, social networks, etc 23
52 Acknowledgements Research Group Ben Raphael Fabio Vandin Hsin-Ta Wu Jason R Dobson Matt Reyna Jonathan Eldridge Alexandra Papoutsaki Jacob Thomas Younhun Kim Collaborators Beifang Niu Michael McLellan Li Ding Michael Lawrence Gad Getz Nuria Lopez-Bigas Abel Gonzalez-Perez David Tamborero Yuwei Chang Greg Ryslik Funding & Data NSF Travel Fellowship to RECOMB 2015
Network-based interpreta1on of perturba1ons to living systems
 Network-based interpreta1on of perturba1ons to living systems Sushmita Roy sroy@biostat.wisc.edu Computa1onal Network Biology Biosta2s2cs & Medical Informa2cs 826 Computer Sciences 838 hbps://compnetbiocourse.discovery.wisc.edu
Network-based interpreta1on of perturba1ons to living systems Sushmita Roy sroy@biostat.wisc.edu Computa1onal Network Biology Biosta2s2cs & Medical Informa2cs 826 Computer Sciences 838 hbps://compnetbiocourse.discovery.wisc.edu
Supplementary Figure 1. Copy Number Alterations TP53 Mutation Type. C-class TP53 WT. TP53 mut. Nature Genetics: doi: /ng.
 Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Supplementary Figure a Copy Number Alterations in M-class b TP53 Mutation Type Recurrent Copy Number Alterations 8 6 4 2 TP53 WT TP53 mut TP53-mutated samples (%) 7 6 5 4 3 2 Missense Truncating M-class
Characteriza*on of Soma*c Muta*ons in Cancer Genomes
 Characteriza*on of Soma*c Muta*ons in Cancer Genomes Ben Raphael Department of Computer Science Center for Computa*onal Molecular Biology Soma*c Muta*ons and Cancer Clonal Theory (Nowell 1976) Passenger
Characteriza*on of Soma*c Muta*ons in Cancer Genomes Ben Raphael Department of Computer Science Center for Computa*onal Molecular Biology Soma*c Muta*ons and Cancer Clonal Theory (Nowell 1976) Passenger
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13898 Supplementary Information Table 1 Kras mutation status of carcinogen-induced mouse lung adenomas Tumour Treatment Strain Grade Genotype Kras status (WES)* Kras status (Sanger) 32T1
doi:10.1038/nature13898 Supplementary Information Table 1 Kras mutation status of carcinogen-induced mouse lung adenomas Tumour Treatment Strain Grade Genotype Kras status (WES)* Kras status (Sanger) 32T1
Plasma-Seq conducted with blood from male individuals without cancer.
 Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Supplementary Figures Supplementary Figure 1 Plasma-Seq conducted with blood from male individuals without cancer. Copy number patterns established from plasma samples of male individuals without cancer
Targeted Agent and Profiling Utilization Registry (TAPUR ) Study. February 2018
 Targeted Agent and Profiling Utilization Registry (TAPUR ) Study February 2018 Precision Medicine Therapies designed to target the molecular alteration that aids cancer development 30 TARGET gene alterations
Targeted Agent and Profiling Utilization Registry (TAPUR ) Study February 2018 Precision Medicine Therapies designed to target the molecular alteration that aids cancer development 30 TARGET gene alterations
Sample Metrics. Allele Frequency (%) Read Depth Ploidy. Gene CDS Effect Protein Effect. LN Metastasis Tumor Purity Computational Pathology 80% 60%
 Supplemental Table 1: Estimated tumor purity, allele frequency, and independent read depth for all gene mutations classified as either potentially pathogenic or VUS in the metatastic and primary tumor
Supplemental Table 1: Estimated tumor purity, allele frequency, and independent read depth for all gene mutations classified as either potentially pathogenic or VUS in the metatastic and primary tumor
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
The Cancer Genome Atlas Pan-cancer analysis Katherine A. Hoadley
 The Cancer Genome Atlas Pan-cancer analysis Katherine A. Hoadley Department of Genetics Lineberger Comprehensive Cancer Center The University of North Carolina at Chapel Hill What is TCGA? The Cancer Genome
The Cancer Genome Atlas Pan-cancer analysis Katherine A. Hoadley Department of Genetics Lineberger Comprehensive Cancer Center The University of North Carolina at Chapel Hill What is TCGA? The Cancer Genome
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature12912 Figure S1. Distribution of mutation rates and spectra across 4,742 tumor-normal (TN) pairs from 21 tumor types, as in Figure 1 from Lawrence et al. Nature
SUPPLEMENTARY INFORMATION doi:1.138/nature12912 Figure S1. Distribution of mutation rates and spectra across 4,742 tumor-normal (TN) pairs from 21 tumor types, as in Figure 1 from Lawrence et al. Nature
Next generation histopathological diagnosis for precision medicine in solid cancers
 Next generation histopathological diagnosis for precision medicine in solid cancers from genomics to clinical application Aldo Scarpa ARC-NET Applied Research on Cancer Department of Pathology and Diagnostics
Next generation histopathological diagnosis for precision medicine in solid cancers from genomics to clinical application Aldo Scarpa ARC-NET Applied Research on Cancer Department of Pathology and Diagnostics
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples
 A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
ARTICLE. Mutational landscape and significance across12majorcancertypes. OPEN doi: /nature12634
 ARTICLE OPEN doi:1.138/nature12634 Mutational landscape and significance across12majorcancertypes Cyriac Kandoth 1 *, Michael D. McLellan 1 *, Fabio Vandin 2, Kai Ye 1,3, Beifang Niu 1, Charles Lu 1, Mingchao
ARTICLE OPEN doi:1.138/nature12634 Mutational landscape and significance across12majorcancertypes Cyriac Kandoth 1 *, Michael D. McLellan 1 *, Fabio Vandin 2, Kai Ye 1,3, Beifang Niu 1, Charles Lu 1, Mingchao
Clinical Grade Genomic Profiling: The Time Has Come
 Clinical Grade Genomic Profiling: The Time Has Come Gary Palmer, MD, JD, MBA, MPH Senior Vice President, Medical Affairs Foundation Medicine, Inc. Oct. 22, 2013 1 Why We Are Here A Shared Vision At Foundation
Clinical Grade Genomic Profiling: The Time Has Come Gary Palmer, MD, JD, MBA, MPH Senior Vice President, Medical Affairs Foundation Medicine, Inc. Oct. 22, 2013 1 Why We Are Here A Shared Vision At Foundation
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each
 Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
Supplementary Figure 1. Cytoscape bioinformatics toolset was used to create the network of protein-protein interactions between the product of each mutated gene and the panel of 125 cancer-driving genes
Ahrim Youn 1,2, Kyung In Kim 2, Raul Rabadan 3,4, Benjamin Tycko 5, Yufeng Shen 3,4,6 and Shuang Wang 1*
 Youn et al. BMC Medical Genomics (2018) 11:98 https://doi.org/10.1186/s12920-018-0425-z RESEARCH ARTICLE Open Access A pan-cancer analysis of driver gene mutations, DNA methylation and gene expressions
Youn et al. BMC Medical Genomics (2018) 11:98 https://doi.org/10.1186/s12920-018-0425-z RESEARCH ARTICLE Open Access A pan-cancer analysis of driver gene mutations, DNA methylation and gene expressions
SureSelect Cancer All-In-One Custom and Catalog NGS Assays
 SureSelect Cancer All-In-One Custom and Catalog NGS Assays Detect all cancer-relevant variants in a single SureSelect assay SNV Indel TL SNV Indel TL Single DNA input Single AIO assay Single data analysis
SureSelect Cancer All-In-One Custom and Catalog NGS Assays Detect all cancer-relevant variants in a single SureSelect assay SNV Indel TL SNV Indel TL Single DNA input Single AIO assay Single data analysis
Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser
 Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser Melissa S. Cline 1*, Brian Craft 1, Teresa Swatloski 1, Mary Goldman 1, Singer Ma 1, David Haussler 1, Jingchun Zhu 1 1 Center for Biomolecular
Exploring TCGA Pan-Cancer Data at the UCSC Cancer Genomics Browser Melissa S. Cline 1*, Brian Craft 1, Teresa Swatloski 1, Mary Goldman 1, Singer Ma 1, David Haussler 1, Jingchun Zhu 1 1 Center for Biomolecular
Cancer develops as a result of the accumulation of somatic
 Cancer-mutation network and the number and specificity of driver mutations Jaime Iranzo a,1, Iñigo Martincorena b, and Eugene V. Koonin a,1 a National Center for Biotechnology Information, National Library
Cancer-mutation network and the number and specificity of driver mutations Jaime Iranzo a,1, Iñigo Martincorena b, and Eugene V. Koonin a,1 a National Center for Biotechnology Information, National Library
Cancer gene discovery via network analysis of somatic mutation data. Insuk Lee
 Cancer gene discovery via network analysis of somatic mutation data Insuk Lee Cancer is a progressive genetic disorder. Accumulation of somatic mutations cause cancer. For example, in colorectal cancer,
Cancer gene discovery via network analysis of somatic mutation data Insuk Lee Cancer is a progressive genetic disorder. Accumulation of somatic mutations cause cancer. For example, in colorectal cancer,
Simultaneous Identification of Multiple Driver Pathways in Cancer
 Simultaneous Identification of Multiple Driver Pathways in Cancer Mark D. M. Leiserson 1, Dima Blokh 2, Roded Sharan 2., Benjamin J. Raphael 1. * 1 Department of Computer Science and Center for Computational
Simultaneous Identification of Multiple Driver Pathways in Cancer Mark D. M. Leiserson 1, Dima Blokh 2, Roded Sharan 2., Benjamin J. Raphael 1. * 1 Department of Computer Science and Center for Computational
Illumina Trusight Myeloid Panel validation A R FHAN R A FIQ
 Illumina Trusight Myeloid Panel validation A R FHAN R A FIQ G E NETIC T E CHNOLOGIST MEDICAL G E NETICS, CARDIFF To Cover Background to the project Choice of panel Validation process Genes on panel, Protocol
Illumina Trusight Myeloid Panel validation A R FHAN R A FIQ G E NETIC T E CHNOLOGIST MEDICAL G E NETICS, CARDIFF To Cover Background to the project Choice of panel Validation process Genes on panel, Protocol
Insights from Sequencing the Melanoma Exome
 Insights from Sequencing the Melanoma Exome Michael Krauthammer, MD PhD, December 2 2015 Yale University School Yof Medicine 1 2012 Exome Screens and Results Exome Sequencing of 108 sun-exposed melanomas
Insights from Sequencing the Melanoma Exome Michael Krauthammer, MD PhD, December 2 2015 Yale University School Yof Medicine 1 2012 Exome Screens and Results Exome Sequencing of 108 sun-exposed melanomas
Identification of Tissue Independent Cancer Driver Genes
 Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Nature Genetics: doi: /ng Supplementary Figure 1. Depths and coverages in whole-exome and targeted deep sequencing data.
 Supplementary Figure 1 Depths and coverages in whole-exome and targeted deep sequencing data. Depth (top) and coverage (bottom) of whole-exome sequencing for 38 independent JPN cases (mean depth = 130)
Supplementary Figure 1 Depths and coverages in whole-exome and targeted deep sequencing data. Depth (top) and coverage (bottom) of whole-exome sequencing for 38 independent JPN cases (mean depth = 130)
Clinically Useful Next Generation Sequencing and Molecular Testing in Gliomas MacLean P. Nasrallah, MD PhD
 Clinically Useful Next Generation Sequencing and Molecular Testing in Gliomas MacLean P. Nasrallah, MD PhD Neuropathology Fellow Division of Neuropathology Center for Personalized Diagnosis (CPD) Glial
Clinically Useful Next Generation Sequencing and Molecular Testing in Gliomas MacLean P. Nasrallah, MD PhD Neuropathology Fellow Division of Neuropathology Center for Personalized Diagnosis (CPD) Glial
The Center for PERSONALIZED DIAGNOSTICS
 The Center for PERSONALIZED DIAGNOSTICS Precision Diagnostics for Personalized Medicine A joint initiative between The Department of Pathology and Laboratory Medicine & The Abramson Cancer Center The (CPD)
The Center for PERSONALIZED DIAGNOSTICS Precision Diagnostics for Personalized Medicine A joint initiative between The Department of Pathology and Laboratory Medicine & The Abramson Cancer Center The (CPD)
Frequency(%) KRAS G12 KRAS G13 KRAS A146 KRAS Q61 KRAS K117N PIK3CA H1047 PIK3CA E545 PIK3CA E542K PIK3CA Q546. EGFR exon19 NFS-indel EGFR L858R
 Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Next Generation Sequencing in Haematological Malignancy: A European Perspective. Wolfgang Kern, Munich Leukemia Laboratory
 Next Generation Sequencing in Haematological Malignancy: A European Perspective Wolfgang Kern, Munich Leukemia Laboratory Diagnostic Methods Cytomorphology Cytogenetics Immunophenotype Histology FISH Molecular
Next Generation Sequencing in Haematological Malignancy: A European Perspective Wolfgang Kern, Munich Leukemia Laboratory Diagnostic Methods Cytomorphology Cytogenetics Immunophenotype Histology FISH Molecular
Patricia Aoun MD, MPH Professor and Vice-Chair for Clinical Affairs Medical Director, Clinical Laboratories Department of Pathology City of Hope
 Patricia Aoun MD, MPH Professor and Vice-Chair for Clinical Affairs Medical Director, Clinical Laboratories Department of Pathology City of Hope National Medical Center Disclosures I have no disclosures
Patricia Aoun MD, MPH Professor and Vice-Chair for Clinical Affairs Medical Director, Clinical Laboratories Department of Pathology City of Hope National Medical Center Disclosures I have no disclosures
Why Pathway/Network Analysis?
 6/8/16 More Depth on Pathway and Network Analysis Lincoln Stein Why Pathway/Network Analysis? Drama@c data size reduc@on: 1000 s of genes => dozens of pathways. Increase sta@s@cal power by reducing mul@ple
6/8/16 More Depth on Pathway and Network Analysis Lincoln Stein Why Pathway/Network Analysis? Drama@c data size reduc@on: 1000 s of genes => dozens of pathways. Increase sta@s@cal power by reducing mul@ple
Accel-Amplicon Panels
 Accel-Amplicon Panels Amplicon sequencing has emerged as a reliable, cost-effective method for ultra-deep targeted sequencing. This highly adaptable approach is especially applicable for in-depth interrogation
Accel-Amplicon Panels Amplicon sequencing has emerged as a reliable, cost-effective method for ultra-deep targeted sequencing. This highly adaptable approach is especially applicable for in-depth interrogation
Identification and clinical detection of genetic alterations of pre-neoplastic lesions Time for the PML ome? David Sidransky MD Johns Hopkins
 Identification and clinical detection of genetic alterations of pre-neoplastic lesions Time for the PML ome? David Sidransky MD Johns Hopkins February 3-5, 2016 Lansdowne Resort, Leesburg, VA Molecular
Identification and clinical detection of genetic alterations of pre-neoplastic lesions Time for the PML ome? David Sidransky MD Johns Hopkins February 3-5, 2016 Lansdowne Resort, Leesburg, VA Molecular
Hepatocellular Carcinoma (HCC): What Treatment Modality for Which Tumor?
 K. Rajender Reddy, MD, CG Hepatocellular Carcinoma (HCC): What Treatment Modality for Which Tumor? K. Rajender Reddy, MD, CG Ruimy Family President s Distinguished Professor of Medicine Director of Hepatology
K. Rajender Reddy, MD, CG Hepatocellular Carcinoma (HCC): What Treatment Modality for Which Tumor? K. Rajender Reddy, MD, CG Ruimy Family President s Distinguished Professor of Medicine Director of Hepatology
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Genomic Medicine: What every pathologist needs to know
 Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Vertical Magnetic Separation of Circulating Tumor Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients
 Vertical Magnetic Separation of Circulating Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients Chang Eun Yoo 1,2#, Jong-Myeon Park 3#, Hui-Sung Moon 1,2, Je-Gun Joung 2, Dae-Soon Son
Vertical Magnetic Separation of Circulating Cells and Somatic Genomic-Alteration Analysis in Lung Cancer Patients Chang Eun Yoo 1,2#, Jong-Myeon Park 3#, Hui-Sung Moon 1,2, Je-Gun Joung 2, Dae-Soon Son
ACTIVITY 2: EXAMINING CANCER PATIENT DATA
 OVERVIEW Refer to the Overview of Cancer Discovery Activities for Key Concepts and Learning Objectives, Curriculum Connections, and Prior Knowledge, as well as background information, references, and additional
OVERVIEW Refer to the Overview of Cancer Discovery Activities for Key Concepts and Learning Objectives, Curriculum Connections, and Prior Knowledge, as well as background information, references, and additional
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Jocelyn Chapman, MD Division of Gynecologic Oncology. Julie S. Mak, MS, MSc Genetic Counselor Hereditary Cancer Clinic
 Jocelyn Chapman, MD Division of Gynecologic Oncology Julie S. Mak, MS, MSc Genetic Counselor Hereditary Cancer Clinic Genetics underlies all cancers Somatic or tumor genetics Germline or inherited genetics
Jocelyn Chapman, MD Division of Gynecologic Oncology Julie S. Mak, MS, MSc Genetic Counselor Hereditary Cancer Clinic Genetics underlies all cancers Somatic or tumor genetics Germline or inherited genetics
Supplementary Figure 1. Screenshot of the MAGI home page and query interface.
 Supplementary Figure 1 Screenshot of the MAGI home page and query interface. (i) Users select a combination of (public or private) datasets to query. (ii) Users can enter up to 25 genes to query at once.
Supplementary Figure 1 Screenshot of the MAGI home page and query interface. (i) Users select a combination of (public or private) datasets to query. (ii) Users can enter up to 25 genes to query at once.
What is the status of the technologies of "precision medicine?
 Session 2: What is the status of the technologies of "precision medicine? Gideon Blumenthal, MD, Clinical Team Leader, Thoracic and Head/Neck Oncology, Center for Drug Evaluation and Research (CDER), U.S.
Session 2: What is the status of the technologies of "precision medicine? Gideon Blumenthal, MD, Clinical Team Leader, Thoracic and Head/Neck Oncology, Center for Drug Evaluation and Research (CDER), U.S.
Expanded View Figures
 Molecular Systems iology Tumor CNs reflect metabolic selection Nicholas Graham et al Expanded View Figures Human primary tumors CN CN characterization by unsupervised PC Human Signature Human Signature
Molecular Systems iology Tumor CNs reflect metabolic selection Nicholas Graham et al Expanded View Figures Human primary tumors CN CN characterization by unsupervised PC Human Signature Human Signature
Identification of Potential Therapeutic Targets by Molecular and Genomic Profiling of 628 Cases of Uterine Serous Carcinoma
 Identification of Potential Therapeutic Targets by Molecular and Genomic Profiling of 628 Cases of Uterine Serous Carcinoma Nathaniel L Jones 1, Joanne Xiu 2, Sandeep K. Reddy 2, Ana I. Tergas 1, William
Identification of Potential Therapeutic Targets by Molecular and Genomic Profiling of 628 Cases of Uterine Serous Carcinoma Nathaniel L Jones 1, Joanne Xiu 2, Sandeep K. Reddy 2, Ana I. Tergas 1, William
Identifying cancer type speci c oncogenes and tumor suppressors using limited size data
 Journal of Bioinformatics and Computational Biology Vol. 14, No. 6 (2016) 1650031 (16 pages) #.c The Author(s) DOI: 10.1142/S0219720016500311 Identifying cancer type speci c oncogenes and tumor suppressors
Journal of Bioinformatics and Computational Biology Vol. 14, No. 6 (2016) 1650031 (16 pages) #.c The Author(s) DOI: 10.1142/S0219720016500311 Identifying cancer type speci c oncogenes and tumor suppressors
Fluxion Biosciences and Swift Biosciences Somatic variant detection from liquid biopsy samples using targeted NGS
 APPLICATION NOTE Fluxion Biosciences and Swift Biosciences OVERVIEW This application note describes a robust method for detecting somatic mutations from liquid biopsy samples by combining circulating tumor
APPLICATION NOTE Fluxion Biosciences and Swift Biosciences OVERVIEW This application note describes a robust method for detecting somatic mutations from liquid biopsy samples by combining circulating tumor
TCGA. The Cancer Genome Atlas
 TCGA The Cancer Genome Atlas TCGA: History and Goal History: Started in 2005 by the National Cancer Institute (NCI) and the National Human Genome Research Institute (NHGRI) with $110 Million to catalogue
TCGA The Cancer Genome Atlas TCGA: History and Goal History: Started in 2005 by the National Cancer Institute (NCI) and the National Human Genome Research Institute (NHGRI) with $110 Million to catalogue
Jennifer Hauenstein Oncology Cytogenetics Emory University Hospital Atlanta, GA
 Comparison of Genomic Coverage using Affymetrix OncoScan Array and Illumina TruSight Tumor 170 NGS Panel for Detection of Copy Number Abnormalities in Clinical GBM Specimens Jennifer Hauenstein Oncology
Comparison of Genomic Coverage using Affymetrix OncoScan Array and Illumina TruSight Tumor 170 NGS Panel for Detection of Copy Number Abnormalities in Clinical GBM Specimens Jennifer Hauenstein Oncology
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester
 Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester dsg6@le.ac.uk CFDNA/CTDNA Circulating-free AS A LIQUID DNA BIOPSY (cfdna) Tumour Biopsy Liquid Biopsy
Dr David Guttery Senior PDRA Dept. of Cancer Studies and CRUK Leicester Centre University of Leicester dsg6@le.ac.uk CFDNA/CTDNA Circulating-free AS A LIQUID DNA BIOPSY (cfdna) Tumour Biopsy Liquid Biopsy
Illumina s Cancer Research Portfolio and Dedicated Workflows
 Illumina s Cancer Research Portfolio and Dedicated Workflows Michael Sohn Clinical Sales Specialist Spain&Italy 2017 2017 Illumina, Inc. All rights reserved. Illumina, 24sure, BaseSpace, BeadArray, BlueFish,
Illumina s Cancer Research Portfolio and Dedicated Workflows Michael Sohn Clinical Sales Specialist Spain&Italy 2017 2017 Illumina, Inc. All rights reserved. Illumina, 24sure, BaseSpace, BeadArray, BlueFish,
NeoTYPE Cancer Profiles
 NeoTYPE Cancer Profiles 30+ Multimethod Assays for Hematologic Diseases and Solid Tumors Molecular FISH Anatomic Pathology The next generation of diagnostic, prognostic, and therapeutic assessment What
NeoTYPE Cancer Profiles 30+ Multimethod Assays for Hematologic Diseases and Solid Tumors Molecular FISH Anatomic Pathology The next generation of diagnostic, prognostic, and therapeutic assessment What
Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making
 Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making November 20, 2014 Capturing Value in Next Generation Sequencing Symposium Douglas Johnson MD, MSCI Vanderbilt-Ingram
Next Generation Sequencing in Clinical Practice: Impact on Therapeutic Decision Making November 20, 2014 Capturing Value in Next Generation Sequencing Symposium Douglas Johnson MD, MSCI Vanderbilt-Ingram
NeoTYPE Cancer Profiles
 NeoTYPE Cancer Profiles Multimethod Analysis of 25+ Hematologic Diseases and Solid Tumors Anatomic Pathology FISH Molecular The next generation of diagnostic, prognostic, and therapeutic assessment NeoTYPE
NeoTYPE Cancer Profiles Multimethod Analysis of 25+ Hematologic Diseases and Solid Tumors Anatomic Pathology FISH Molecular The next generation of diagnostic, prognostic, and therapeutic assessment NeoTYPE
COSMIC - Catalogue of Somatic Mutations in Cancer
 COSMIC - Catalogue of Somatic Mutations in Cancer http://cancer.sanger.ac.uk/cosmic https://academic.oup.com/nar/articl e-lookup/doi/10.1093/nar/gkw1121 Data In Large-scale systematic screens Detailed
COSMIC - Catalogue of Somatic Mutations in Cancer http://cancer.sanger.ac.uk/cosmic https://academic.oup.com/nar/articl e-lookup/doi/10.1093/nar/gkw1121 Data In Large-scale systematic screens Detailed
Enabling Personalized
 Molecular Enabling Personalized Diagnostics Medicine- Targeted Sequencing: NGS-based solutions Silvia Dorn Roel Reinders- Andreas Diplas Friday, 19.06.2015 Company Overview Founded in April 2011 Development
Molecular Enabling Personalized Diagnostics Medicine- Targeted Sequencing: NGS-based solutions Silvia Dorn Roel Reinders- Andreas Diplas Friday, 19.06.2015 Company Overview Founded in April 2011 Development
Understanding Genotype- Phenotype relations in Cancer via Network Approaches
 AlgoCSB Algorithmic Methods in Computational and Systems Biology Understanding Genotype- Phenotype relations in Cancer via Network Approaches Teresa Przytycka NIH / NLM / NCBI Phenotypes Journal Wisla
AlgoCSB Algorithmic Methods in Computational and Systems Biology Understanding Genotype- Phenotype relations in Cancer via Network Approaches Teresa Przytycka NIH / NLM / NCBI Phenotypes Journal Wisla
Conditional Selection of Genomic Alterations Dictates Cancer Evolution and Oncogenic Dependencies
 Article Conditional Selection of Genomic Alterations Dictates Cancer Evolution and Oncogenic Dependencies Graphical Abstract Authors Marco Mina, Franck Raynaud, Daniele Tavernari,..., Nikolaus Schultz,
Article Conditional Selection of Genomic Alterations Dictates Cancer Evolution and Oncogenic Dependencies Graphical Abstract Authors Marco Mina, Franck Raynaud, Daniele Tavernari,..., Nikolaus Schultz,
IntelliGENSM. Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community.
 IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
IntelliGENSM Integrated Oncology is making next generation sequencing faster and more accessible to the oncology community. NGS TRANSFORMS GENOMIC TESTING Background Cancers may emerge as a result of somatically
Assessing the Economics of Genomic Medicine. Kenneth Offit, MD MPH Chief, Clinical Genetics Service Memorial Sloan-Kettering Cancer Center
 Assessing the Economics of Genomic Medicine Kenneth Offit, MD MPH Chief, Clinical Genetics Service Memorial Sloan-Kettering Cancer Center DISCLOSURES No conflicts Patents unenforced Consultancies unpaid
Assessing the Economics of Genomic Medicine Kenneth Offit, MD MPH Chief, Clinical Genetics Service Memorial Sloan-Kettering Cancer Center DISCLOSURES No conflicts Patents unenforced Consultancies unpaid
S1 Appendix: Figs A G and Table A. b Normal Generalized Fraction 0.075
 Aiello & Alter (216) PLoS One vol. 11 no. 1 e164546 S1 Appendix A-1 S1 Appendix: Figs A G and Table A a Tumor Generalized Fraction b Normal Generalized Fraction.25.5.75.25.5.75 1 53 4 59 2 58 8 57 3 48
Aiello & Alter (216) PLoS One vol. 11 no. 1 e164546 S1 Appendix A-1 S1 Appendix: Figs A G and Table A a Tumor Generalized Fraction b Normal Generalized Fraction.25.5.75.25.5.75 1 53 4 59 2 58 8 57 3 48
Multiplatform Analysis of 12 Cancer Types Reveals Molecular Classification within and across Tissues of Origin
 Resource Multiplatform Analysis of 12 Cancer Types Reveals Molecular Classification within and across Tissues of Origin Katherine A. Hoadley, 1,20 Christina Yau, 2,20 Denise M. Wolf, 3,20 Andrew D. Cherniack,
Resource Multiplatform Analysis of 12 Cancer Types Reveals Molecular Classification within and across Tissues of Origin Katherine A. Hoadley, 1,20 Christina Yau, 2,20 Denise M. Wolf, 3,20 Andrew D. Cherniack,
Looking Beyond the Standard-of- Care : The Clinical Trial Option
 1 Looking Beyond the Standard-of- Care : The Clinical Trial Option Terry Mamounas, M.D., M.P.H., F.A.C.S. Medical Director, Comprehensive Breast Program UF Health Cancer Center at Orlando Health Professor
1 Looking Beyond the Standard-of- Care : The Clinical Trial Option Terry Mamounas, M.D., M.P.H., F.A.C.S. Medical Director, Comprehensive Breast Program UF Health Cancer Center at Orlando Health Professor
Changing demographics of smoking and its effects during therapy
 Changing demographics of smoking and its effects during therapy Egbert F. Smit MD PhD. Dept. Pulmonary Diseases, Vrije Universiteit Medical Centre, Amsterdam, The Netherlands Smoking prevalence adults
Changing demographics of smoking and its effects during therapy Egbert F. Smit MD PhD. Dept. Pulmonary Diseases, Vrije Universiteit Medical Centre, Amsterdam, The Netherlands Smoking prevalence adults
NGS in tissue and liquid biopsy
 NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
Nature Genetics: doi: /ng Supplementary Figure 1. Workflow of CDR3 sequence assembly from RNA-seq data.
 Supplementary Figure 1 Workflow of CDR3 sequence assembly from RNA-seq data. Paired-end short-read RNA-seq data were mapped to human reference genome hg19, and unmapped reads in the TCR regions were extracted
Supplementary Figure 1 Workflow of CDR3 sequence assembly from RNA-seq data. Paired-end short-read RNA-seq data were mapped to human reference genome hg19, and unmapped reads in the TCR regions were extracted
Precision medicine: How to exploit the growing knowledge on the evolving genomes of cells to improve cancer prevention and therapy.
 Precision medicine: How to exploit the growing knowledge on the evolving genomes of cells to improve cancer prevention and therapy Joe Costello, PhD Department of Neurological Surgery A more accurate and
Precision medicine: How to exploit the growing knowledge on the evolving genomes of cells to improve cancer prevention and therapy Joe Costello, PhD Department of Neurological Surgery A more accurate and
Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder Labs August 30, 2012
 Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Table S1. Demographics of patients and tumor characteristics.
 Supplemental Tables for: A Clinical Actionability of Comprehensive Genomic Profiling for Management of Rare or Refractory Cancers Kim Hirshfield et al. Table S1. Demographics of patients and tumor characteristics.
Supplemental Tables for: A Clinical Actionability of Comprehensive Genomic Profiling for Management of Rare or Refractory Cancers Kim Hirshfield et al. Table S1. Demographics of patients and tumor characteristics.
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION Frampton et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing Supplementary Figure 1A Base substitution detection
SUPPLEMENTARY INFORMATION Frampton et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing Supplementary Figure 1A Base substitution detection
Voting-Based Cancer Module Identification by Combining Topological and Data-Driven Properties
 by Combining Topological and Data-Driven Properties A. K. M. Azad, Hyunju Lee* School of Information and Communications, Gwangju Institute of Science and Technology, Gwangju, South Korea Abstract Recently,
by Combining Topological and Data-Driven Properties A. K. M. Azad, Hyunju Lee* School of Information and Communications, Gwangju Institute of Science and Technology, Gwangju, South Korea Abstract Recently,
APPLICATIONS OF NEXT GENERATION SEQUENCING IN SOLID TUMORS - PATHOLOGIST PROSPECTIVE
 AMP COMPANION MEETING SYMPOSIUM AT USCAP 2015 NEXT-GENERATION OF PATHOLOGY: ROLE OF PATHOLOGIST IN NGS-BASED PERSONALIZED MEDICINE APPLICATIONS OF NEXT GENERATION SEQUENCING IN SOLID TUMORS - PATHOLOGIST
AMP COMPANION MEETING SYMPOSIUM AT USCAP 2015 NEXT-GENERATION OF PATHOLOGY: ROLE OF PATHOLOGIST IN NGS-BASED PERSONALIZED MEDICINE APPLICATIONS OF NEXT GENERATION SEQUENCING IN SOLID TUMORS - PATHOLOGIST
Cumulative Haploinsufficiency and Triplosensitivity Drive Aneuploidy Patterns and Shape the Cancer Genome
 Resource Cumulative Haploinsufficiency and Triplosensitivity Drive Aneuploidy Patterns and Shape the Cancer Genome Teresa Davoli, 1,2,5 Andrew Wei Xu, 2,4,5 Kristen E. Mengwasser, 1,2 Laura M. Sack, 1,2
Resource Cumulative Haploinsufficiency and Triplosensitivity Drive Aneuploidy Patterns and Shape the Cancer Genome Teresa Davoli, 1,2,5 Andrew Wei Xu, 2,4,5 Kristen E. Mengwasser, 1,2 Laura M. Sack, 1,2
Genomic landsccape of poorly differentiated and anaplastic thyroid carcinomas: Clues for better classification, risk stratification and therapy
 ACCME/Disclosures The USCAP requires that anyone in a position to influence or control the content of CME disclose any relevant financial relationship WITH COMMERCIAL INTERESTS which they or their spouse/partner
ACCME/Disclosures The USCAP requires that anyone in a position to influence or control the content of CME disclose any relevant financial relationship WITH COMMERCIAL INTERESTS which they or their spouse/partner
Supplemental Information. Integrated Genomic Analysis of the Ubiquitin. Pathway across Cancer Types
 Cell Reports, Volume 23 Supplemental Information Integrated Genomic Analysis of the Ubiquitin Pathway across Zhongqi Ge, Jake S. Leighton, Yumeng Wang, Xinxin Peng, Zhongyuan Chen, Hu Chen, Yutong Sun,
Cell Reports, Volume 23 Supplemental Information Integrated Genomic Analysis of the Ubiquitin Pathway across Zhongqi Ge, Jake S. Leighton, Yumeng Wang, Xinxin Peng, Zhongyuan Chen, Hu Chen, Yutong Sun,
Molecular Subtyping of Endometrial Cancer: A ProMisE ing Change
 Molecular Subtyping of Endometrial Cancer: A ProMisE ing Change Charles Matthew Quick, M.D. Associate Professor of Pathology Director of Gynecologic Pathology University of Arkansas for Medical Sciences
Molecular Subtyping of Endometrial Cancer: A ProMisE ing Change Charles Matthew Quick, M.D. Associate Professor of Pathology Director of Gynecologic Pathology University of Arkansas for Medical Sciences
Susanne Schnittger. Workflow of molecular investigations in JAK2-negative MPNs - the Munich experience
 Susanne Schnittger Workflow of molecular investigations in JAK2negative MPNs the Munich experience Cohort single centre experience to apply new markers in a daily diagnostic work flow total: 20,547 cases
Susanne Schnittger Workflow of molecular investigations in JAK2negative MPNs the Munich experience Cohort single centre experience to apply new markers in a daily diagnostic work flow total: 20,547 cases
Secuenciación masiva: papel en la toma de decisiones
 Secuenciación masiva: papel en la toma de decisiones Cancer is a Genetic Disease Development of cancer is driven by the acquisition of somatic genetic alterations: Nonsynonymous point mutations: missense.
Secuenciación masiva: papel en la toma de decisiones Cancer is a Genetic Disease Development of cancer is driven by the acquisition of somatic genetic alterations: Nonsynonymous point mutations: missense.
Tratamiento neoadyuvante: Enfermedad residual como marcador de resistencia Carlos L. Arteaga, MD Vanderbilt-Ingram Cancer Center Vanderbilt
 Tratamiento neoadyuvante: Enfermedad residual como marcador de resistencia Carlos L. Arteaga, MD Vanderbilt-Ingram Cancer Center Vanderbilt University Neoadjuvant (preoperative) therapy Surgery Systemic
Tratamiento neoadyuvante: Enfermedad residual como marcador de resistencia Carlos L. Arteaga, MD Vanderbilt-Ingram Cancer Center Vanderbilt University Neoadjuvant (preoperative) therapy Surgery Systemic
The mutations that drive cancer. Paul Edwards. Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge
 The mutations that drive cancer Paul Edwards Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge Previously on Cancer... hereditary predisposition Normal Cell Slightly
The mutations that drive cancer Paul Edwards Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge Previously on Cancer... hereditary predisposition Normal Cell Slightly
Clinical Grade Biomarkers in the Genomic Era Observations & Challenges
 Clinical Grade Biomarkers in the Genomic Era Observations & Challenges IOM Committee on Policy Issues in the Clinical Development & Use of Biomarkers for Molecularly Targeted Therapies March 31-April 1,
Clinical Grade Biomarkers in the Genomic Era Observations & Challenges IOM Committee on Policy Issues in the Clinical Development & Use of Biomarkers for Molecularly Targeted Therapies March 31-April 1,
Supplementary Figure 1 Weight and body temperature of ferrets inoculated with
 Supplementary Figure 1 Weight and body temperature of ferrets inoculated with A/Anhui/1/2013 (H7N9) influenza virus. (a) Body temperature and (b) weight change of ferrets after intranasal inoculation with
Supplementary Figure 1 Weight and body temperature of ferrets inoculated with A/Anhui/1/2013 (H7N9) influenza virus. (a) Body temperature and (b) weight change of ferrets after intranasal inoculation with
Clustered mutations of oncogenes and tumor suppressors.
 Supplementary Figure 1 Clustered mutations of oncogenes and tumor suppressors. For each oncogene (red dots) and tumor suppressor (blue dots), the number of mutations found in an intramolecular cluster
Supplementary Figure 1 Clustered mutations of oncogenes and tumor suppressors. For each oncogene (red dots) and tumor suppressor (blue dots), the number of mutations found in an intramolecular cluster
Personalized Healthcare Update
 Dr. Kai - Oliver Wesche Market Development Manager, Personalized Healthcare QIAGEN Personalized Healthcare Update Pioneering Personalized Medicine through Partnering TOMTOVOK BKM120 Zelboraf QIAGEN partners:
Dr. Kai - Oliver Wesche Market Development Manager, Personalized Healthcare QIAGEN Personalized Healthcare Update Pioneering Personalized Medicine through Partnering TOMTOVOK BKM120 Zelboraf QIAGEN partners:
Comprehensive Analyses of Circulating Cell- Free Tumor DNA
 Comprehensive Analyses of Circulating Cell- Free Tumor DNA Boston, MA June 28th, 2016 Derek Murphy, Ph.D. Scientist, Research and Development Personal Genome Diagnostics Acquisition of Somatic Alterations
Comprehensive Analyses of Circulating Cell- Free Tumor DNA Boston, MA June 28th, 2016 Derek Murphy, Ph.D. Scientist, Research and Development Personal Genome Diagnostics Acquisition of Somatic Alterations
modified dye uptake assay including formazan test EC 90 not tested plaque reduction assay
 Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
Sauerbrei A, Bohn-Wippert K, Kaspar M, Krumbholz A, Karrasch M, Zell R. 2015. Database on natural polymorphisms and resistance-related non-synonymous mutations in thymidine kinase and DNA polymerase genes
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication
 Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
Virological failure to Protease inhibitors in Monotherapy is linked to the presence of signature mutations in Gag without changes in HIV-1 replication Oscar Blanch-Lombarte Rome, 7-9 June, 2017 European
5 th July 2016 ACGS Dr Michelle Wood Laboratory Genetics, Cardiff
 5 th July 2016 ACGS Dr Michelle Wood Laboratory Genetics, Cardiff National molecular screening of patients with lung cancer for a national trial of multiple novel agents. 2000 NSCLC patients/year (late
5 th July 2016 ACGS Dr Michelle Wood Laboratory Genetics, Cardiff National molecular screening of patients with lung cancer for a national trial of multiple novel agents. 2000 NSCLC patients/year (late
Identifying driver mutations in sequenced cancer genomes: computational approaches to enable precision medicine
 Raphael et al. Genome Medicine 2014, 6:5 REVIEW Identifying driver mutations in sequenced cancer genomes: computational approaches to enable precision medicine Benjamin J Raphael 1,2*, Jason R Dobson 1,2,3,
Raphael et al. Genome Medicine 2014, 6:5 REVIEW Identifying driver mutations in sequenced cancer genomes: computational approaches to enable precision medicine Benjamin J Raphael 1,2*, Jason R Dobson 1,2,3,
Characterisation of structural variation in breast. cancer genomes using paired-end sequencing on. the Illumina Genome Analyser
 Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies
 Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
A Robust Method for Identifying Mutated Driver Pathways in Cancer
 , pp.392-397 http://dx.doi.org/10.14257/astl.2016. A Robust Method for Identifying Mutated Driver Pathways in Cancer Can-jun Hu and Shu-Lin Wang * College of computer science and electronic engineering,
, pp.392-397 http://dx.doi.org/10.14257/astl.2016. A Robust Method for Identifying Mutated Driver Pathways in Cancer Can-jun Hu and Shu-Lin Wang * College of computer science and electronic engineering,
Machine-Learning on Prediction of Inherited Genomic Susceptibility for 20 Major Cancers
 Machine-Learning on Prediction of Inherited Genomic Susceptibility for 20 Major Cancers Sung-Hou Kim University of California Berkeley, CA Global Bio Conference 2017 MFDS, Seoul, Korea June 28, 2017 Cancer
Machine-Learning on Prediction of Inherited Genomic Susceptibility for 20 Major Cancers Sung-Hou Kim University of California Berkeley, CA Global Bio Conference 2017 MFDS, Seoul, Korea June 28, 2017 Cancer
Dr Yvonne Wallis Consultant Clinical Scientist West Midlands Regional Genetics Laboratory
 Dr Yvonne Wallis Consultant Clinical Scientist West Midlands Regional Genetics Laboratory Personalised Therapy/Precision Medicine Selection of a therapeutic drug based on the presence or absence of a specific
Dr Yvonne Wallis Consultant Clinical Scientist West Midlands Regional Genetics Laboratory Personalised Therapy/Precision Medicine Selection of a therapeutic drug based on the presence or absence of a specific
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E,
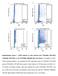 Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
Supplementary Figure 1. FACS analysis of cells infected with TY93/H5N1 GFP-627E, TY93/H5N1 GFP-627K, or the TY93/H5N1 PB2(588-759) virus library. To establish our GFP- FACS screening platform, we compared
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL SUPPLEMENTAL MATERIAL Supplemental Methods Phenotype Prediction Analyses In order to assess the phylogenetic properties of nssnvs, sequence conservation analysis was conducted using
SUPPLEMENTAL MATERIAL SUPPLEMENTAL MATERIAL Supplemental Methods Phenotype Prediction Analyses In order to assess the phylogenetic properties of nssnvs, sequence conservation analysis was conducted using
e-driver: A novel method to identify protein regions driving cancer Eduard Porta-Pardo 1, Adam Godzik 1,* 1
 Original Paper e-driver: A novel method to identify protein regions driving cancer Eduard Porta-Pardo 1, Adam Godzik 1,* 1 Bioinformatics and Systems Biology Program, Sanford-Burnham Medical Research Institute,
Original Paper e-driver: A novel method to identify protein regions driving cancer Eduard Porta-Pardo 1, Adam Godzik 1,* 1 Bioinformatics and Systems Biology Program, Sanford-Burnham Medical Research Institute,
SAPLING: A Tool for Gene Network Analysis focusing on Psychiatric Genetics
 SAPLING: A Tool for Gene Network Analysis focusing on Psychiatric Genetics sapling.cshl.edu Wim Verleyen, Ph.D. Gillis Lab Outline Motivation Disease-gene analysis Enrichment analysis Gene network analysis:
SAPLING: A Tool for Gene Network Analysis focusing on Psychiatric Genetics sapling.cshl.edu Wim Verleyen, Ph.D. Gillis Lab Outline Motivation Disease-gene analysis Enrichment analysis Gene network analysis:
