iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers).
|
|
|
- Jesse Kennedy
- 5 years ago
- Views:
Transcription
1 Supplementary Materials Supplementary Methods iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers). IDH1 R132H and R132L Forward: 5 -ACGTTGGATGCAACATGACTTACTTGATCCCC-3 Reverse: 5 -ACGTTGGATGAAAAATATCCCCCGGCTTGTG-3 Extension: 5 -GTAAAACCTATCATCATAGGTC-3 IDH1 R132C, R132G, and R132S Forward: 5 -ACGTTGGATGAAAAATATCCCCCGGCTTGTG-3 Reverse: 5 -ACGTTGGATGCAACATGACTTACTTGATCCCC-3 Extension: 5 -ATCCCCATAAGCATGAC-3 IDH2 R172G Forward: 5 -ACGTTGGATGAAAAACATCCCACGCCTAGTC-3 Reverse: 5 -ACGTTGGATGTCAGTGGATCCCCTCTCCACC-3 Extension: 5 -GGTCGCCATGGGCGTGCC-3 IDH2 R172K and R172M Forward: 5 -ACGTTGGATGTCAGTGGATCCCCTCTCCACC-3 Reverse: 5 -ACGTTGGATGAAAAACATCCCACGCCTAGTC-3 Extension: 5 -GCCCATCACCATTGGCA-3 Multiplexed PCR was performed in 5 µl containing 0.1 U of Kapa 2G Fast Taq polymerase (Kapa Biosystems, Capetown, South Africa), 10 ng of genomic DNA, 2.5 pmol of each PCR primer and 2.5 mmol of dntp. An amplification
2 round to generate mutational hot spot regions was carried out using the following thermocycling conditions are as follows: 1 cycle of 95 C for 5 min; 3 cycles of 95 C for 30 sec, 64 C for 30 sec, 72 C for 60 sec; 3 cycles of 95 C for 30 sec, 62 C for 30 sec, 72 C for 60 sec; 3 cycles of 95 C for 30 sec, 60 C for 30 sec, 72 C for 60 sec; 37 cycles of 95 C for 30 sec, 58 C for 30 sec, 72 C for 60 sec, and one final cycle of 70 C for 5 min. One ml of a 1/10 dilution of final product was used as a template for a second round of amplification using the PCR conditions described above. Unincorporated dntps were deactivated using 0.3 U of shrimp alkaline phosphatase at 37º C for 40 minutes, followed by heat inactivation at 85º C for 5 minutes. Single-base primer extension was carried out using 5.4 pmol of each extension primer, 50 mmol of the appropriate dntp/ddntp combination and 0.5 units of Thermosequenase DNA polymerase (iplex enzyme). Extension reactions were cycled using a 200-short-cycle program that uses two cycling loops as follows: the sample is denatured at 94º C, annealing takes place at 52º C for 5 seconds and extension at 80º C for 5 seconds. The annealing and extension cycle is repeated four more times for a total of five cycles and then looped back to a 94º C denaturing step for 5 seconds and then enters the 5 cycle annealing and extension loop again. The five annealing and extension steps with the single denaturing step are repeated 40 times. The 40 cycles of the 5 cycle annealing and extension steps equate to a total of 200 cycles. A final extension is done at 72º C for three minutes and then the sample is cooled to 4º C. After the addition of a cation exchange resin to remove residual salt from the reactions, 10 nl of the
3 purified primer extension reaction were loaded onto a matrix pad (3- hydroxypicoloinic acid) of a SpectroCHIP (Sequenom). SpectroCHIPs were analyzed using a Bruker Biflex III matrix-assisted laser desorption/ionization time of flight (MALDI-TOF) mass spectrometer (SpectroREADER, Sequenom). All results were manually inspected, using the Mass ARRAY TyperAnalyzer v4.0 software. Automated mutation calling for TP53 sequencing Bi-directional reads and mapping tables (to link read names to sample identifiers, gene names, read direction, and amplicon) were subjected to a QC filter which excludes reads that have an average phred score of < 10 for bases Passing reads were assembled against the reference sequences for each gene, containing all coding and UTR exons including 5Kb upstream and downstream of the gene, using command line Consed 16.0 (PMID: ). Assemblies were passed on to Polyphred 6.02b (PMID: ), which generated a list of putative candidate mutations, and to Polyscan 3.0 (PMID: ) which generated a second list of putative mutations. The lists were merged together into a combined report, and the putative mutation calls were normalized to + genomic coordinates and annotated using the Genomic Mutation Consequence Calculator (PMID: ). The resulting list of annotated putative mutations was loaded into a Postgres database along with select assembly details for each mutation call (assembly position, coverage, and methods supporting mutation call). To reduce the number of false positives generated by the mutation detection software packages, only point mutations which are supported by at least one bi-directional
4 read pair and at least one sample mutation called by Polyphred were considered, and only the putative mutations which are annotated as having non-synonymous coding effects, occur within 11 bp of an exon boundary, or have a conservation score > ( were included in the final candidate list. Indels called by any method were manually reviewed and included in the candidate list if found to hit an exon. All traces for mutation calls were manually reviewed.
5 Supplementary Table Legends Table S1: Gene list of refined signature designating expression subclasses in lower-grade diffuse astrocytic gliomas from MSKCC sample set. ROC scores are shown. Table S2: Modified MSKCC sample set gene list used to analyze REMBRANDT expression data. ROC scores are shown. Table S3: Gene list of unified expression signature developed from subclass assignments in both MSKCC and REMBRANDT array data. Summed ROC scores are shown. Table S4: Ingenuity analysis results for canonical pathways using NB subclass Table S5: Ingenuity analysis results for functional annotations using NB subclass Table S6: Ingenuity analysis results for canonical pathways using EPL subclass Table S7: Ingenuity analysis results for functional annotations using EPL subclass
6 Table S8: Ingenuity analysis results for canonical pathways using PG subclass Table S9: Ingenuity analysis results for functional annotations using PG subclass Table S10: Demographic and molecular features of diffuse astrocytic gliomas (MSKCC sample set) stratified by primary/recurrent tumor status. P values are shown. Mt: mutant, Wt: wild-type, IHC: immunohistochemistry, unkn: unknown, FISH: fluorescence in-situ hybridization, NB: neuroblastic, EPL: early progenitorlike, PG: pre-gbm. Table S11: Histopathological features of diffuse astrocytic gliomas (MSKCC sample set) stratified by molecular subclass, WHO grade, and primary/recurrent tumor status. P values are shown. Pro: prominent (characterizing >50% of tumor), Pre: present, Ab: absent, NB: neuroblastic, EPL: early progenitor-like, PG: pre-gbm. Table S12: Statistically significant copy number alterations as determined by the GISTIC algorithm for REMBRANDT diffuse astrocytic gliomas of NB subclass. Table S13: Statistically significant copy number alterations as determined by the GISTIC algorithm for REMBRANDT diffuse astrocytic gliomas of EPL subclass.
7 Table S14: Statistically significant copy number alterations as determined by the GISTIC algorithm for REMBRANDT diffuse astrocytic gliomas of PG subclass. Supplementary Figure Legends FIG. S1: Kaplan-Meier curves showing overall survival for recurrent MSKCC diffuse astrocytic gliomas stratified by WHO grade and prior treatment modality (Surg: surgery only; Surg + Adj: surgery plus adjuvant radiation therapy and/or chemotherapy). Sample sizes are in parentheses. P=0.083 for WHO II tumors and P=0.363 for WHO III tumors. FIG. S2: Representative immunohistochemical stains and scoring. All photomicrographs were taken at 400X magnification. A, p53 immunopositivity required nuclear staining in >30% of tumor cells. B, PDGFRA immunopositivity required detectable staining, and was frequently seen in only a subset of tumor cells, particularly in IDH mt tumors. C, Strong p-pras40 immunostaining required robust signal in the majority of tumor cells. D, IDH R132H immunopositivity required detectable staining. FIG. S3: Consensus k-means clustering demonstrated 3 robust subclasses of lower-grade diffuse astrocytic glioma. Cluster stability for filtered sets of 200, 250, 300, and 350 genes was maximal at k=3 as measured by the Δ(Gini) (A-D).
8 FIG. S4: Positive silhouette width for initial clusters of diffuse astrocytic glioma determined core subclass members, which were then used for downstream SAM and ROC analysis. FIG. S5: Kaplan-Meier curves showing overall survival for REMBRANDT diffuse astrocytic gliomas stratified by WHO grade. Sample sizes are in parentheses. P value shown.
Nature Genetics: doi: /ng.2995
 Supplementary Figure 1 Kaplan-Meier survival curves of patients with brainstem tumors. (a) Comparison of patients with PPM1D mutation versus wild-type PPM1D. (b) Comparison of patients with PPM1D mutation
Supplementary Figure 1 Kaplan-Meier survival curves of patients with brainstem tumors. (a) Comparison of patients with PPM1D mutation versus wild-type PPM1D. (b) Comparison of patients with PPM1D mutation
Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3.
 Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3. MN1 (Accession No. NM_002430) MN1-1514F 5 -GGCTGTCATGCCCTATTGAT Exon 1 MN1-1882R 5 -CTGGTGGGGATGATGACTTC Exon
Table S1. Primers and PCR protocols for mutation screening of MN1, NF2, KREMEN1 and ZNRF3. MN1 (Accession No. NM_002430) MN1-1514F 5 -GGCTGTCATGCCCTATTGAT Exon 1 MN1-1882R 5 -CTGGTGGGGATGATGACTTC Exon
WHO Prequalification of In Vitro Diagnostics PUBLIC REPORT. Product: Alere q HIV-1/2 Detect WHO reference number: PQDx
 WHO Prequalification of In Vitro Diagnostics PUBLIC REPORT Product: Alere q HIV-1/2 Detect WHO reference number: PQDx 0226-032-00 Alere q HIV-1/2 Detect with product codes 270110050, 270110010 and 270300001,
WHO Prequalification of In Vitro Diagnostics PUBLIC REPORT Product: Alere q HIV-1/2 Detect WHO reference number: PQDx 0226-032-00 Alere q HIV-1/2 Detect with product codes 270110050, 270110010 and 270300001,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
Nature Genetics: doi: /ng Supplementary Figure 1. Details of sequencing analysis.
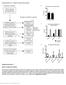 Supplementary Figure 1 Details of sequencing analysis. (a) Flow chart showing which patients fall into each category and were used for analysis. (b) Graph showing the average and median coverage for all
Supplementary Figure 1 Details of sequencing analysis. (a) Flow chart showing which patients fall into each category and were used for analysis. (b) Graph showing the average and median coverage for all
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Aliccia Bollig-Fischer, PhD Department of Oncology, Wayne State University Associate Director Genomics Core Molecular Therapeutics Program Karmanos
 Aliccia Bollig-Fischer, PhD Department of Oncology, Wayne State University Associate Director Genomics Core Molecular Therapeutics Program Karmanos Cancer Institute Development of a multiplexed assay to
Aliccia Bollig-Fischer, PhD Department of Oncology, Wayne State University Associate Director Genomics Core Molecular Therapeutics Program Karmanos Cancer Institute Development of a multiplexed assay to
Frequency(%) KRAS G12 KRAS G13 KRAS A146 KRAS Q61 KRAS K117N PIK3CA H1047 PIK3CA E545 PIK3CA E542K PIK3CA Q546. EGFR exon19 NFS-indel EGFR L858R
 Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Frequency(%) 1 a b ALK FS-indel ALK R1Q HRAS Q61R HRAS G13R IDH R17K IDH R14Q MET exon14 SS-indel KIT D8Y KIT L76P KIT exon11 NFS-indel SMAD4 R361 IDH1 R13 CTNNB1 S37 CTNNB1 S4 AKT1 E17K ERBB D769H ERBB
Corporate Medical Policy
 Corporate Medical Policy Analysis of MGMT Promoter Methylation in Malignant Gliomas File Name: Origination: Last CAP Review: Next CAP Review: Last Review: analysis_of_mgmt_promoter_methylation_in_malignant_gliomas
Corporate Medical Policy Analysis of MGMT Promoter Methylation in Malignant Gliomas File Name: Origination: Last CAP Review: Next CAP Review: Last Review: analysis_of_mgmt_promoter_methylation_in_malignant_gliomas
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Case Presentation: USCAP Jason T. Huse, MD, PhD Assistant Member Department of Pathology Memorial Sloan Kettering Cancer Center
 Case Presentation: USCAP 2016 Jason T. Huse, MD, PhD Assistant Member Department of Pathology Memorial Sloan Kettering Cancer Center Case History 53 year old female with a long standing history of migraines
Case Presentation: USCAP 2016 Jason T. Huse, MD, PhD Assistant Member Department of Pathology Memorial Sloan Kettering Cancer Center Case History 53 year old female with a long standing history of migraines
gliomas. Fetal brain expected who each low-
 Supplementary Figure S1. Grade-specificity aberrant expression of HOXA genes in gliomas. (A) Representative RT-PCR analyses of HOXA gene expression in human astrocytomas. Exemplified glioma samples include
Supplementary Figure S1. Grade-specificity aberrant expression of HOXA genes in gliomas. (A) Representative RT-PCR analyses of HOXA gene expression in human astrocytomas. Exemplified glioma samples include
Analysis with SureCall 2.1
 Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
The absence of the ERBB4 hotspot mutations in melanomas in patients from southern China
 Brief Report The absence of the ERBB4 hotspot mutations in melanomas in patients from southern China Qi-Ming Zhou 1,2,5*, Wei Li 6*, Yuan-Xiang Guan 1,3, Xing Zhang 1,2, Xin-Chun Chen 6, Ya Ding 1,2, Xi-Zhi
Brief Report The absence of the ERBB4 hotspot mutations in melanomas in patients from southern China Qi-Ming Zhou 1,2,5*, Wei Li 6*, Yuan-Xiang Guan 1,3, Xing Zhang 1,2, Xin-Chun Chen 6, Ya Ding 1,2, Xi-Zhi
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH
 EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
EPIGENETIC RE-EXPRESSION OF HIF-2α SUPPRESSES SOFT TISSUE SARCOMA GROWTH Supplementary Figure 1. Supplementary Figure 1. Characterization of KP and KPH2 autochthonous UPS tumors. a) Genotyping of KPH2
Results and Discussion of Receptor Tyrosine Kinase. Activation
 Results and Discussion of Receptor Tyrosine Kinase Activation To demonstrate the contribution which RCytoscape s molecular maps can make to biological understanding via exploratory data analysis, we here
Results and Discussion of Receptor Tyrosine Kinase Activation To demonstrate the contribution which RCytoscape s molecular maps can make to biological understanding via exploratory data analysis, we here
Polymorphism of the PAI-1gene (4G/5G) may be linked with Polycystic Ovary Syndrome and associated pregnancy disorders in South Indian Women
 www.bioinformation.net Volume 13(5) Hypothesis Polymorphism of the PAI-1gene (4G/5G) may be linked with Polycystic Ovary Syndrome and associated pregnancy disorders in South Indian Women Maniraja Jesintha
www.bioinformation.net Volume 13(5) Hypothesis Polymorphism of the PAI-1gene (4G/5G) may be linked with Polycystic Ovary Syndrome and associated pregnancy disorders in South Indian Women Maniraja Jesintha
Detection of copy number variations in PCR-enriched targeted sequencing data
 Detection of copy number variations in PCR-enriched targeted sequencing data German Demidov Parseq Lab, Saint-Petersburg University of Russian Academy of Sciences, current: Center for Genomic Regulation
Detection of copy number variations in PCR-enriched targeted sequencing data German Demidov Parseq Lab, Saint-Petersburg University of Russian Academy of Sciences, current: Center for Genomic Regulation
BWA alignment to reference transcriptome and genome. Convert transcriptome mappings back to genome space
 Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
MRC-Holland MLPA. Description version 08; 30 March 2015
 SALSA MLPA probemix P351-C1 / P352-D1 PKD1-PKD2 P351-C1 lot C1-0914: as compared to the previous version B2 lot B2-0511 one target probe has been removed and three reference probes have been replaced.
SALSA MLPA probemix P351-C1 / P352-D1 PKD1-PKD2 P351-C1 lot C1-0914: as compared to the previous version B2 lot B2-0511 one target probe has been removed and three reference probes have been replaced.
MRC-Holland MLPA. Description version 06; 23 December 2016
 SALSA MLPA probemix P417-B2 BAP1 Lot B2-1216. As compared to version B1 (lot B1-0215), two reference probes have been added and two target probes have a minor change in length. The BAP1 (BRCA1 associated
SALSA MLPA probemix P417-B2 BAP1 Lot B2-1216. As compared to version B1 (lot B1-0215), two reference probes have been added and two target probes have a minor change in length. The BAP1 (BRCA1 associated
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
DOI: 10.1038/ncb2607 Figure S1 Elf5 loss promotes EMT in mammary epithelium while Elf5 overexpression inhibits TGFβ induced EMT. (a, c) Different confocal slices through the Z stack image. (b, d) 3D rendering
SALSA MLPA probemix P315-B1 EGFR
 SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
SUPPLEMENTARY FIGURES: Supplementary Figure 1
 SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
Supplementary Figure 1
 A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
A B D Relative TAp73 mrna p73 Supplementary Figure 1 25 2 15 1 5 p63 _-tub. MDA-468 HCC1143 HCC38 SUM149 MDA-468 HCC1143 HCC38 SUM149 HCC-1937 MDA-MB-468 ΔNp63_ TAp73_ TAp73β E C Relative ΔNp63 mrna TAp73
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplemental Information. Molecular, Pathological, Radiological, and Immune. Profiling of Non-brainstem Pediatric High-Grade
 Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
Cancer Cell, Volume 33 Supplemental Information Molecular, Pathological, Radiological, and Immune Profiling of Non-brainstem Pediatric High-Grade Glioma from the HERBY Phase II Randomized Trial Alan Mackay,
6/12/2018. Disclosures. Clinical Genomics The CLIA Lab Perspective. Outline. COH HopeSeq Heme Panels
 Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Supplementary Material
 Supplementary Material 2 4 6 Single-locus gene screening methods We screened for single nucleotide polymorphisms (SNPs) at 16 candidate immune genes (Table S1) by initially sequencing three captive-bred
Supplementary Material 2 4 6 Single-locus gene screening methods We screened for single nucleotide polymorphisms (SNPs) at 16 candidate immune genes (Table S1) by initially sequencing three captive-bred
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits
 AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
Monitoring for Drug Resistance by Genotyping. Urvi M Parikh, PhD MTN Virology Core Lab
 Monitoring for Drug Resistance by Genotyping Urvi M Parikh, PhD MTN Virology Core Lab Outline What is Drug Resistance? Genotyping Algorithm Standard vs Sensitive Resistance Testing Sequencing Protocols
Monitoring for Drug Resistance by Genotyping Urvi M Parikh, PhD MTN Virology Core Lab Outline What is Drug Resistance? Genotyping Algorithm Standard vs Sensitive Resistance Testing Sequencing Protocols
Assessment performed on Friday, September 18, 2015, at Vancouver General Hospital
 Assessors report for ciqc Run 49: ATRX (June 2015) Assessors: S Yip and J Won (recorder) Assessment performed on Friday, September 18, 2015, at Vancouver General Hospital Background The combined application
Assessors report for ciqc Run 49: ATRX (June 2015) Assessors: S Yip and J Won (recorder) Assessment performed on Friday, September 18, 2015, at Vancouver General Hospital Background The combined application
Molecular Markers. Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC
 Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Complete Genomics, Inc.
 Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Evaluation of MIA FORA NGS HLA test and software. Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group
 Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
2/10/2016. Evaluation of MIA FORA NGS HLA test and software. Disclosure. NGS-HLA typing requirements for the Stanford Blood Center
 Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
Evaluation of MIA FORA NGS HLA test and software Lisa Creary, PhD Department of Pathology Stanford Blood Center Research & Development Group Disclosure Alpha and Beta Studies Sirona Genomics Reagents,
OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES
 OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES EVERY STEP OF THE WAY 1 EVERY STEP OF THE WAY MICROBIAL IDENTIFICATION METHODS DNA RNA Genotypic Sequencing of ribosomal RNA regions of bacteria
OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES EVERY STEP OF THE WAY 1 EVERY STEP OF THE WAY MICROBIAL IDENTIFICATION METHODS DNA RNA Genotypic Sequencing of ribosomal RNA regions of bacteria
Supplementary Note. Nature Genetics: doi: /ng.2928
 Supplementary Note Loss of heterozygosity analysis (LOH). We used VCFtools v0.1.11 to extract only singlenucleotide variants with minimum depth of 15X and minimum mapping quality of 20 to create a ped
Supplementary Note Loss of heterozygosity analysis (LOH). We used VCFtools v0.1.11 to extract only singlenucleotide variants with minimum depth of 15X and minimum mapping quality of 20 to create a ped
MRC-Holland MLPA. Description version 12; 13 January 2017
 SALSA MLPA probemix P219-B3 PAX6 Lot B3-0915: Compared to version B2 (lot B2-1111) two reference probes have been replaced and one additional reference probe has been added. In addition, one flanking probe
SALSA MLPA probemix P219-B3 PAX6 Lot B3-0915: Compared to version B2 (lot B2-1111) two reference probes have been replaced and one additional reference probe has been added. In addition, one flanking probe
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Figure 1
 Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Count Count Supplementary Figure 1 Coverage per amplicon for error-corrected sequencing experiments. Errorcorrected consensus sequence (ECCS) coverage was calculated for each of the 568 amplicons in the
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Bredel M, Scholtens DM, Yadav AK, et al. NFKBIA deletion in
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Bredel M, Scholtens DM, Yadav AK, et al. NFKBIA deletion in
Total genomic DNA was extracted from either 6 DBS punches (3mm), or 0.1ml of peripheral or
 Material and methods Measurement of telomere length (TL) Total genomic DNA was extracted from either 6 DBS punches (3mm), or 0.1ml of peripheral or cord blood using QIAamp DNA Mini Kit and a Qiacube (Qiagen).
Material and methods Measurement of telomere length (TL) Total genomic DNA was extracted from either 6 DBS punches (3mm), or 0.1ml of peripheral or cord blood using QIAamp DNA Mini Kit and a Qiacube (Qiagen).
A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis
 APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
Data Mining for Research and Diagnostics
 UCL DEPARTMENT OF GEOGRAPHY UCL INSTITUTE OF NEUROLOGY Data Mining for Research and Diagnostics Sebastian Brandner, UCL Institute of Neurology, London Data mining for Research and Diagnostics Our setup
UCL DEPARTMENT OF GEOGRAPHY UCL INSTITUTE OF NEUROLOGY Data Mining for Research and Diagnostics Sebastian Brandner, UCL Institute of Neurology, London Data mining for Research and Diagnostics Our setup
AZOOSPERMIA Chromosome Y
 AZOOSPERMIA Chromosome Y M i c r o d e l e t i o n Ref.: PI EDP003024-40 testspi EDP002024 1. INTRODUCTION In 1976, Tiepolo and Zuffardi reported de novo, microscopically detectable deletions of the distal
AZOOSPERMIA Chromosome Y M i c r o d e l e t i o n Ref.: PI EDP003024-40 testspi EDP002024 1. INTRODUCTION In 1976, Tiepolo and Zuffardi reported de novo, microscopically detectable deletions of the distal
Molecular biomarker profile of EGFR copy number, KRAS and BRAF mutations in colorectal carcinoma
 ORIGINAL ARTICLE Molecular biomarker profile of EGFR copy number, KRAS and BRAF mutations in colorectal carcinoma Rong Rong, Jamie Tull, Shengle Zhang Department of Pathology, SUNY Upstate University,
ORIGINAL ARTICLE Molecular biomarker profile of EGFR copy number, KRAS and BRAF mutations in colorectal carcinoma Rong Rong, Jamie Tull, Shengle Zhang Department of Pathology, SUNY Upstate University,
A novel isothermal amplification approach for rapid identification of BCR-ABL fusion genes at onset:
 A novel isothermal amplification approach for rapid identification of BCR-ABL fusion genes at onset: Josh Glason: Sales Manager, DiaSorin Australia Pty Ltd September 6, 2014 This product is not currently
A novel isothermal amplification approach for rapid identification of BCR-ABL fusion genes at onset: Josh Glason: Sales Manager, DiaSorin Australia Pty Ltd September 6, 2014 This product is not currently
Nature Biotechnology: doi: /nbt.1904
 Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
PBZ FT01_PBZ FT01_TZ FT01_NZ. interface zone (I) tumor zone (TZ) necrotic zone (NZ)
 Oncotarget, Supplementary Materials www.impactjournals.com/oncotarget/ SUPPLEMENTRY FLES ndividuals factor map (P) FT_ FT_ FT_ Dim (.%) Dim (.%) >% peripheral brain zone () around % interface zone () FT
Oncotarget, Supplementary Materials www.impactjournals.com/oncotarget/ SUPPLEMENTRY FLES ndividuals factor map (P) FT_ FT_ FT_ Dim (.%) Dim (.%) >% peripheral brain zone () around % interface zone () FT
SUPPLEMENTARY INFORMATION
 doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
Characterisation of structural variation in breast. cancer genomes using paired-end sequencing on. the Illumina Genome Analyser
 Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
HIV-1 Genemer Detection Kit Ready to Use Amplification Kit for HIV-1 Specific DNA Fragment Analysis
 Product Manual HIV-1 Genemer Detection Kit Ready to Use Amplification Kit for HIV-1 Specific DNA Fragment Analysis For research use only. Not for use in diagnostic procedures for clinical purposes Catalog
Product Manual HIV-1 Genemer Detection Kit Ready to Use Amplification Kit for HIV-1 Specific DNA Fragment Analysis For research use only. Not for use in diagnostic procedures for clinical purposes Catalog
Detection of Tamiflu resistant Swine H1N1 Influenza Human Pandemic Strain
 PrimerDesign TM Ltd Detection of Tamiflu resistant Swine H1N1 Influenza Human Pandemic Strain H275Y mutation For general laboratory and research use only Contents Introduction to Swine H1N1 Influenza Human
PrimerDesign TM Ltd Detection of Tamiflu resistant Swine H1N1 Influenza Human Pandemic Strain H275Y mutation For general laboratory and research use only Contents Introduction to Swine H1N1 Influenza Human
UNIVERSITI TEKNOLOGI MARA COPY NUMBER VARIATIONS OF ORANG ASLI (NEGRITO) FROM PENINSULAR MALAYSIA
 UNIVERSITI TEKNOLOGI MARA COPY NUMBER VARIATIONS OF ORANG ASLI (NEGRITO) FROM PENINSULAR MALAYSIA SITI SHUHADA MOKHTAR Thesis submitted in fulfillment of the requirements for the degree of Master of Science
UNIVERSITI TEKNOLOGI MARA COPY NUMBER VARIATIONS OF ORANG ASLI (NEGRITO) FROM PENINSULAR MALAYSIA SITI SHUHADA MOKHTAR Thesis submitted in fulfillment of the requirements for the degree of Master of Science
NGS in tissue and liquid biopsy
 NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery
 CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery PacBio CoLab Session October 20, 2017 For Research Use Only. Not for use in diagnostics procedures. Copyright 2017 by Pacific Biosciences
CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery PacBio CoLab Session October 20, 2017 For Research Use Only. Not for use in diagnostics procedures. Copyright 2017 by Pacific Biosciences
MRC-Holland MLPA. Description version 18; 09 September 2015
 SALSA MLPA probemix P090-A4 BRCA2 Lot A4-0715, A4-0714, A4-0314, A4-0813, A4-0712: Compared to lot A3-0710, the 88 and 96 nt control fragments have been replaced (QDX2). This product is identical to the
SALSA MLPA probemix P090-A4 BRCA2 Lot A4-0715, A4-0714, A4-0314, A4-0813, A4-0712: Compared to lot A3-0710, the 88 and 96 nt control fragments have been replaced (QDX2). This product is identical to the
Supplementary methods:
 Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
Supplementary methods: Primers sequences used in real-time PCR analyses: β-actin F: GACCTCTATGCCAACACAGT β-actin [11] R: AGTACTTGCGCTCAGGAGGA MMP13 F: TTCTGGTCTTCTGGCACACGCTTT MMP13 R: CCAAGCTCATGGGCAGCAACAATA
NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation
 NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation Michael R. Rossi, PhD, FACMG Assistant Professor Division of Cancer Biology, Department of Radiation Oncology Department
NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation Michael R. Rossi, PhD, FACMG Assistant Professor Division of Cancer Biology, Department of Radiation Oncology Department
MALDI Mass Spectrometry: A Label-Free Solution for Ultra-High-Throughput Screening
 MALDI Mass Spectrometry: A Label-Free Solution for Ultra-High-Throughput Screening MIPTEC Conference 2016 Meike Hamester, Jens Fuchser Bruker Daltonik, Bremen 1 MALDI PharmaPulse for: Biochemical Screening
MALDI Mass Spectrometry: A Label-Free Solution for Ultra-High-Throughput Screening MIPTEC Conference 2016 Meike Hamester, Jens Fuchser Bruker Daltonik, Bremen 1 MALDI PharmaPulse for: Biochemical Screening
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
SALSA MLPA probemix P241-D2 MODY mix 1 Lot D As compared to version D1 (lot D1-0911), one reference probe has been replaced.
 mix P241-D2 MODY mix 1 Lot D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent diabetes
mix P241-D2 MODY mix 1 Lot D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent diabetes
Title:Validation study of candidate single nucleotide polymorphisms associated with left ventricular hypertrophy in the Korean population
 Author's response to reviews Title:Validation study of candidate single nucleotide polymorphisms associated with left ventricular hypertrophy in the Korean population Authors: Jin-Kyu Park (cardiohy@gmail.com)
Author's response to reviews Title:Validation study of candidate single nucleotide polymorphisms associated with left ventricular hypertrophy in the Korean population Authors: Jin-Kyu Park (cardiohy@gmail.com)
A novel and universal method for microrna RT-qPCR data normalization
 A novel and universal method for microrna RT-qPCR data normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle 4 th International qpcr Symposium Weihenstephan, March 1,
A novel and universal method for microrna RT-qPCR data normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle 4 th International qpcr Symposium Weihenstephan, March 1,
Bioimaging and Functional Genomics
 Bioimaging and Functional Genomics Elisa Ficarra, EPF Lausanne Giovanni De Micheli, EPF Lausanne Sungroh Yoon, Stanford University Luca Benini, University of Bologna Enrico Macii,, Politecnico di Torino
Bioimaging and Functional Genomics Elisa Ficarra, EPF Lausanne Giovanni De Micheli, EPF Lausanne Sungroh Yoon, Stanford University Luca Benini, University of Bologna Enrico Macii,, Politecnico di Torino
Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples.
 Supplementary files Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples. Table S3. Specificity of AGO1- and AGO4-preferred 24-nt
Supplementary files Table S1. Relative abundance of AGO1/4 proteins in different organs. Table S2. Summary of smrna datasets from various samples. Table S3. Specificity of AGO1- and AGO4-preferred 24-nt
Lynch Syndrome and COLARIS Testing
 Lynch Syndrome and COLARIS Testing Webinar Objectives Review of Lynch syndrome as a multi-gene disorder COLARIS Enhancements Technical Overview COLARIS test offerings Test development and validation process
Lynch Syndrome and COLARIS Testing Webinar Objectives Review of Lynch syndrome as a multi-gene disorder COLARIS Enhancements Technical Overview COLARIS test offerings Test development and validation process
Genesis of cerebellar interneurons and the prevention of neural DNA damage require XRCC1.
 Genesis of cerebellar interneurons and the prevention of neural DNA damage require XRCC1. Youngsoo Lee, Sachin Katyal, Yang Li, Sherif F. El-Khamisy, Helen R. Russell, Keith W. Caldecott and Peter J. McKinnon.
Genesis of cerebellar interneurons and the prevention of neural DNA damage require XRCC1. Youngsoo Lee, Sachin Katyal, Yang Li, Sherif F. El-Khamisy, Helen R. Russell, Keith W. Caldecott and Peter J. McKinnon.
IDH1 R132H/ATRX Immunohistochemical validation
 IDH1 R132H/ATRX Immunohistochemical validation CIQC/DSM 2016 12 June 2016 0835-0905 Stephen Yip, M.D., Ph.D., FRCPC University of British Columbia Disclosure Statement I have nothing to disclose I will
IDH1 R132H/ATRX Immunohistochemical validation CIQC/DSM 2016 12 June 2016 0835-0905 Stephen Yip, M.D., Ph.D., FRCPC University of British Columbia Disclosure Statement I have nothing to disclose I will
Role of Paired Box9 (PAX9) (rs ) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis
 EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
Nature Medicine: doi: /nm.3967
 Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Supplementary Figure 1. Network clustering. (a) Clustering performance as a function of inflation factor. The grey curve shows the median weighted Silhouette widths for varying inflation factors (f [1.6,
Session 6: Integration of epigenetic data. Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016
 Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
a) SSR with core motif > 2 and repeats number >3. b) MNR with repeats number>5.
 1 2 APPENDIX Legends to figures 3 4 5 Figure A1: Distribution of perfect SSR along chromosome 1 of V. cholerae (El-Tor N191). a) SSR with core motif > 2 and repeats number >3. b) MNR with repeats number>5.
1 2 APPENDIX Legends to figures 3 4 5 Figure A1: Distribution of perfect SSR along chromosome 1 of V. cholerae (El-Tor N191). a) SSR with core motif > 2 and repeats number >3. b) MNR with repeats number>5.
Enterprise Interest None
 Enterprise Interest None Heterogeneous chromosomal profiles in a unique series of DIPG in children and young adults European Congress of Pathology Amsterdam, 6 th September 2017 Charlotte Dufour, Romain
Enterprise Interest None Heterogeneous chromosomal profiles in a unique series of DIPG in children and young adults European Congress of Pathology Amsterdam, 6 th September 2017 Charlotte Dufour, Romain
MRC-Holland MLPA. Description version 29;
 SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
Human Immunodeficiency Virus-1 (HIV-1) Genemer. Primer Pair for amplification of HIV-1 Specific DNA Fragment
 Product Manual Human Immunodeficiency Virus-1 (HIV-1) Genemer Primer Pair for amplification of HIV-1 Specific DNA Fragment Catalog No.: 60-2002-10 Store at 20 o C For research use only. Not for use in
Product Manual Human Immunodeficiency Virus-1 (HIV-1) Genemer Primer Pair for amplification of HIV-1 Specific DNA Fragment Catalog No.: 60-2002-10 Store at 20 o C For research use only. Not for use in
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
EXPression ANalyzer and DisplayER
 EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols.
 Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols. A-tailed DNA was ligated to T-tailed dutp adapters, circularized
Supplementary Figure 1. Schematic diagram of o2n-seq. Double-stranded DNA was sheared, end-repaired, and underwent A-tailing by standard protocols. A-tailed DNA was ligated to T-tailed dutp adapters, circularized
Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University
 Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
AVENIO ctdna Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB
 Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
Biomarker development in the era of precision medicine. Bei Li, Interdisciplinary Technical Journal Club
 Biomarker development in the era of precision medicine Bei Li, 23.08.2016 Interdisciplinary Technical Journal Club The top ten highest-grossing drugs in the United States help between 1 in 25 and 1 in
Biomarker development in the era of precision medicine Bei Li, 23.08.2016 Interdisciplinary Technical Journal Club The top ten highest-grossing drugs in the United States help between 1 in 25 and 1 in
MRC-Holland MLPA. Description version 29; 31 July 2015
 SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0114. As compared to the previous B2 version (lot 0813 and 0912), 11 target probes are replaced or added, and 10 new reference probes are included. P082
SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0114. As compared to the previous B2 version (lot 0813 and 0912), 11 target probes are replaced or added, and 10 new reference probes are included. P082
SALSA MLPA probemix P241-D2 MODY mix 1 Lot D2-0716, D As compared to version D1 (lot D1-0911), one reference probe has been replaced.
 mix P241-D2 MODY mix 1 Lot D2-0716, D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent
mix P241-D2 MODY mix 1 Lot D2-0716, D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Probe. Hind III Q,!?R'!! /0!!!!D1"?R'! vector. Homologous recombination
 Supple-Zhang Page 1 Wild-type locus Targeting construct Targeted allele Exon Exon3 Exon Probe P1 P P3 FRT FRT loxp loxp neo vector amh I Homologous recombination neo P1 P P3 FLPe recombination Q,!?R'!!
Supple-Zhang Page 1 Wild-type locus Targeting construct Targeted allele Exon Exon3 Exon Probe P1 P P3 FRT FRT loxp loxp neo vector amh I Homologous recombination neo P1 P P3 FLPe recombination Q,!?R'!!
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy
 Kavya Puchhalapalli CALS Honors Project Report Spring 2017 Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy Abstract Malignant brain tumors including medulloblastomas and primitive neuroectodermal
Kavya Puchhalapalli CALS Honors Project Report Spring 2017 Studying The Role Of DNA Mismatch Repair In Brain Cancer Malignancy Abstract Malignant brain tumors including medulloblastomas and primitive neuroectodermal
Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY
 Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY Methodology The test is based on the reported 50-gene classifier algorithm originally named PAM50 and is performed on the ncounter Dx Analysis System
Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY Methodology The test is based on the reported 50-gene classifier algorithm originally named PAM50 and is performed on the ncounter Dx Analysis System
Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY
 Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY GENE EXPRESSION PROFILING WITH PROSIGNA What is Prosigna? Prosigna Breast Cancer Prognostic Gene Signature Assay is an FDA-approved assay which provides
Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY GENE EXPRESSION PROFILING WITH PROSIGNA What is Prosigna? Prosigna Breast Cancer Prognostic Gene Signature Assay is an FDA-approved assay which provides
Journal: Nature Methods
 Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma.
 Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Supplementary Figure 1 Breeding scheme, transgenes, histological analysis and site distribution of SB-mutagenized osteosarcoma. (a) Breeding scheme. R26-LSL-SB11 homozygous mice were bred to Trp53 LSL-R270H/+
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer.
 Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Assessment performed on Tuesday, July 29, 2014, at Lions Gate Hospital, North Vancouver
 Assessors report for ciqc Run 37: BRAF V600E (April 2014) Assessors: B Gilks, R Wolber, K Ung, P Tavassoli, J Garratt and J Won (recorder) Assessment performed on Tuesday, July 29, 2014, at Lions Gate
Assessors report for ciqc Run 37: BRAF V600E (April 2014) Assessors: B Gilks, R Wolber, K Ung, P Tavassoli, J Garratt and J Won (recorder) Assessment performed on Tuesday, July 29, 2014, at Lions Gate
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Schlenk RF, Döhner K, Krauter J, et al. Mutations and treatment
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Schlenk RF, Döhner K, Krauter J, et al. Mutations and treatment
