Transcriptional Units for Ribosomal Proteins of Escherichia coli
|
|
|
- Emory McGee
- 5 years ago
- Views:
Transcription
1 Eur. J. Biochem. 52, (1975) Transcriptional Units for Ribosomal Proteins of Escherichia coli Monica HIRSCH-KAUFFMANN, Manfred SCHWEIGER, Peter HERRLICH, Helmut PONTA, Hans-Jobst RAHMSDORF, Shou-Hsang PAI, and Heinz-Georg WITTMANN Max-Planck-lnstitut fur Molekulare Genetik, Berlin-Dahlem (Received September 7, 1974) Transcriptional units for ribosomal proteins in Escherichiu cofi were measured using the ultraviolet sensitivities of the rates of synthesis of individual ribosomal proteins. The ultraviolet sensitivities of gene transcriptions are proportional to the distances from the promoters. The longest transcriptional units for ribosomal proteins are 3.6 x 10' of DNA molecular weight corresponding to 1.8 x 10' of RNA or to of protein. The length would cover genes of ribosomal proteins (of an average M, of ). The synthesis of the ribosomal constituents seems to be highly coordinated. The coordinate control is suggested by the apparent constancy of ribosomes relative to growth rate [1,2] and the small pool of both the 55 individual ribosomal proteins [3] and the three ribosomal RNAs [4]. The mechanisms are unknown by which the synthesis of these many ribosomal components is regulated. An appealing possibility would be the arrangement of the ribosomal RNA genes in one transcriptional unit and of the ribosomal protein genes in another large transcriptional unit, a "ribosomal protein operon". The RNA genes indeed appear to be transcribed from the promoter in the order S - 5s RNA [4-71. The localization of many genes for ribosomal proteins in one region of the E. coli chromosome (between 64 and 66 min) [8] would be consistent with a ribosomal protein operon. It found further support by the polar effects of p-insertions on the expression of several ribosomal marker proteins [9]. However, experiments measuring the read-out time of mrna for ribosomal proteins after rifampicin addition, and the kinetics of synthesis of ribosomal proteins after rifampicin removal, indicated that not all genes for ribosomal proteins could be transcribed from one promoter [lo]. The experiments described here determine the size of the transcriptional units for ribosomal proteins by measuring the ultraviolet sensitivities of the rate of synthesis of the ribosomal proteins. It is shown that at least six transcriptional units for ribosomal proteins exist in E. coli, none of which carries more than ribosomal protein cistrons of average size. MATERIALS AND METHODS Radioactive and Other Chemicals ''C-labelled and 3H-labelled protein hydrolysate and ['4C]galactose were purchased from Radiochemicals, Amersham ; D-fucose was obtained from Sigma, St. Louis. Phage T7 wild type was grown and purified as described [I I]. Gro#tli of Bacteria E. coli Bs- [12] were grown at 30 "C in M9 medium supplemented with 0.4 % glucose and M MgSO,. Ultraviolet Irradiation At A600 = 0.35 the cells were irradiated in dim yellow light with various doses of ultraviolet light. 10-ml aliquots were placed in open glass petri dishes (r = 4.5 cm) under a 35-W ultraviolet lamp (Quarzlampenges. Hanau). The dose of irradiation was varied with the time of irradiation at a constant distance of 40 cm. The ultraviolet doses were roughly calibrated by comparing their effect on phage survival and on
2 470 E. coli Ribosomal Proteins : Transcriptional Units known phage marker genes [ll]. 25 s of ultraviolet reduced the surviving T7 phage of a suspension of the same A260 by lo3. The cultures were allowed to shake at 30 C for 10 min after the irradiation. Aliquots were then used for the following determinations : the rate of synthesis of galactokinase, the rate of synthesis of individual ribosomal proteins, and the rate of synthesis of various E. coli proteins. Rate of Synthesis of Various E. coli Proteins 0.2 ml aliquots of irradiated and non-irradiated cells were pulse-labelled with 14C-labelled amino acids (5 pci) for 5 min. A chase of 2 min with tryptone broth (1 % tryptone, 0.5% NaC1) followed. The cells were pelleted and treated with dodecylsulphate sample buffer [I 31. The total protein was separated by dodecylsulphate - polyacrylamide slab-gel electrophoresis [13]. The gels were dried and autoradiographed. The radioactivity in each protein band was determined by densitometry of the X-ray films. Galac tok inase Synthesis Galactokinase synthesis was determined by the rate of accumulation of enzyme activity upon induction with D-fucose. The enzyme was induced and assayed as described [14]. Rate of Synthesis of Individual Ribosomal Proteins 20-ml aliquots of irradiated and non-irradiated cells were made radioactive with 14C-labelled amino acids (1.24 pci/ml; 500 pci/pmol) for 5 min. The cells were harvested on ice and pelleted together with the same volume of 3H-labelled control cells. The control cells were not irradiated, but incubated with 3H-labelled amino acids (2.5 pci/ml; 1000 pci/pmol) for 15 min. Proteins were extracted with 67% acetic acid in the presence of 0.07 M magnesium acetate [15] from whole-cell homogenates (sonication) and resolved by the standard two-dimensional polyacrylamide gel electrophoresis [16]. In some cases carrier ribosomal protein was added before electrophoresis. The ribosomal protein spots were detected by staining with Coomassie blue and the radioactivity in each protein determined by cutting the spots out of the gel and separating 3H and 14C counts in a Packard autooxidizer. RESULTS AND DISCUSSION Principle of Promoter Mapping by Ultraviolet Irradiation Irradiation of whole cells with ultraviolet light interferes with replication and transcription. Both types of inhibition are caused by lesions in the DNA [17,18]. The ultraviolet lesions in the DNA cause termination of RNA chains prematurely under release of RNA polymerase. Ultraviolet lesions are distributed randomly; therefore, the probability to hit a stretch of DNA increases with its length : the target size. If transcription is observed, the transcriptional unit is the relevant target. The promoter-distal portion of a transcriptional unit is affected first and, with increasing doses of ultraviolet, the average length of transcribed RNA is reduced. From the portion beyond the ultraviolet lesion neither RNA nor protein is synthesized. The rate of synthesis of a protein thus is a measure of the surviving activity of the corresponding gene (provided that time is allowed to pass after the irradiation, so that preformed messenger can decay). Since, in most cases, it is difficult to define RNA species, the rate of synthesis of protein is determined. The ultraviolet sensitivity of transcription and subsequent translation depends on the distance of the end of the gene from its promoter. If a gene is located adjacent to the promoter, the ultraviolet sensitivity of its expression will be proportional to the molecular weight of the gene product. The ultraviolet sensitivity of a gene in a distal position of a transcriptional unit corresponds to the sum of the molecular weights of this gene product and of those proteins arising from promoter-proximal genes. The ultraviolet sensitivity is plotted as an inactivation curve and expressed as the inactivation tangent (as e.g. shown in Fig.3). The inactivation tangent is proportional to the target size, i.e. stretch of DNA transcribed. The inactivation tangent is plotted against the molecular weight (as in Fig. 2). This ultraviolet method has been worked out and tested in the bacteriophage T7 system [ll, 191. Ultraviolet Sensitivity of the Rate of Synthesis of Various E. coli Proteins In order to obtain a calibration curve for the ribosomal protein cistrons of E. coli, the ultraviolet sensitivity of the synthesis of various E. coli proteins was determined. Aliquots of cultures, which had been irradiated and which had been incubated subsequently for 10 min, were pulse-labelled with 14C-labelled amino acids and either used for the isolation and separation of ribosomal proteins, or directly for the separation of total protein by dodecylsulphate-polyacrylamide gel (Fig. 1). Even a superficial glance over autoradiograms of these gels shows that the proteins with high molecular weights disappear at lower doses ultraviolet irradiation than do those that had moved further in the gel. By densitometry of the autoradiograms the inactivation of synthesis by ultraviolet irradiation was
3 M. Hirsch-Kauffmann. M. Schweiger, P. Herrlich, H. Ponta, H.-J. Rahmsdorf, S.-H. Pai, and H.-G. Wittmann 471 / Fig. 1. Wltruriolet inuctirution of'thr rate qf'synthesis of E. coli proteins. E. coli Bs-, were irradiated with ultraviolet. The ultraviolet dose was varied by the time of irradiation under a source of constant ultraviolet light. After further incubation for 10 min, aliquots were taken for the following determinations: the rate of synthesis of galactokinase upon induction by D-fUCOSe (shown in Fig.2), the rate of synthesis of individual ribosomal proteins (Fig. 3 and Table I), and the rate of synthesis of various E. coli proteins. For the latter determination 0.2-ml aliquots were labelled with 5 pci "C-labelled amino acids. A chase with cold amino acids followed and the proteins were separated by dodecylsulphate- polyacrylamide slab-gel electrophoresis (10-20 "i, gradient). An autoradiogram is shown here. For standardization of the molecular weights, marker proteins were applied to the same slab gel. As an example, T7 proteins are shown (0-10-min pulse-label with '4C-labelled amino acids after infection of hcavily irradiated B,-, with T7'). All experiments of one set were performed with the same series of ultraviolet-irradiated cultures. 4 independent sets have been evaluated quantitated and the tangent plotted against the molecular weight, which was obtained by co-electrophoresis of marker proteins (Fig.2). The majority of proteins which were detected on dodecylsulphate gels plot on a straight line (Fig. 2). These proteins are apparently synthesized from genes that are located adjacent to promoters. Several proteins, however, show plot positions which are clearly above the line; for instance galactokinase synthesis, which was measured by its characteristic enzyme reaction, is inactivated at lower doses than would be expected from its molecular weight. The inactivation rate of galactokinase corre- I I I I IO-~ M, Fig. 2. Internul calihrcition standard for trunscriptionul-unit mapping with ultraviolet. As described in the text and as demonstrated in Fig. 3, inactivation curves for various proteins were obtained. Proteins were selected at random and only identified by their molecular weight. The slope of each curve, as a measure of the inactivation of the rate of synthesis by ultraviolet light, was expressed as the inactivation tangent, r. The tangent (a) of each protein was plotted against the molecular weight of the protein. The straight line served as a calibration curve as described in Results and Discussion. (0) Represents the value obtained for galactokinase synthesis. Galactokinase would have a molecular weight of if it were synthesized from a monocistronic transcriptional unit sponds to a DNA stretch of approximately x lo6 daltons or an RNA of 1.2 x lo6 daltons or a total protein of daltons. Galactokinase is coded by the third gene of the galactose operon [20]. Each of the three proteins of the gal operon has a molecular weight of [20] adding up to a total of The ultraviolet data thus say that all gal operon genes are transcribed from one promoter which has indeed been shown earlier by genetic methods [20]. With this plot we have obtained a calibration curve which was then used for the internal standardization of ultraviolet data with unknown transcriptional units, such as those for ribosomal proteins. Ultraviolet Sensitivity of the Rate of Synthesis of Individual Ribosomal Proteins The ribosomal proteins from E. coli have been isolated and their molecular weights have been
4 E. coli Ribosomal Proteins: Transcriptional Units lo I m I 1, 1 8 I c I Ultraviolet irradiation (5) Fig. 3. Ultraviolet sensitivit). oj the rute oj synthesis oj ribosomal proteins. After irradiation (see Fig. l), 20-ml aliquots of the cultures were pulse-labelled with 14C-labelled amino acids and mixed with control cells that had been labelled with 3H-labelled amino acids. The ribosomal proteins were separated and the rate of synthesis of each protein plotted as a percentage of control rate versus ultraviolet dose. The tangent a for each Ultraviolet irradiation ( s) protein was taken from these plots and evaluated in Table 1. (A) Example of proteins from the small ribosomal subunit; (B) examples of proteins from the large ribosomal subunit. The slope of inactivation of a gene, which is located at the end of a large transcriptional unit comprising all 55 ribosomal protein genes, is marked determined [21]. The average is to With this information, the ultraviolet method yields the size of the transcriptional unit or units. In order to measure rate of synthesis and not rate of assembly of ribosomal proteins, the ribosomal proteins were isolated from total-cell extracts (and not from isolated ribosomes) and separated by twodimensional gel electrophoresis. The locations of ribosomal proteins were detected by cochromatography of carrier protein and staining of the gels. The 14C counts (irradiated cells) in each spot were taken as a measure of the remaining synthesis rate as compared to the control rate (3H counts or 14C counts of nonirradiated sample). For each ribosomal protein ultraviolet-inactivation rates were plotted (Fig. 3). The theoretical curve for a unique operon containing all 55 ribosomal protein genes is marked. None of the inactivation rates of individual ribosomal proteins comes close to the curve calculated for the operon. The highest observed inactivation rate of an individual ribosomal protein was approximately 18-20% of that expected for the ribosomal operon. From the calibration curve (Fig. 2), the longest transcriptional unit has been calculated to be equivalent to 3.6 x lo6 daltons of DNA or 1.8 x lo6 daltons of RNA or about daltons of protein. At an average molecular weight of ribosomal proteins of to [21], the longest transcriptional unit would comprise about genes of average size. If all genes for ribosomal proteins were organized in such transcriptional units of maximal size, at least five transcriptional units would be needed for 55 ribosomal proteins. From the inactivation rates the relative positions of each gene with respect to its promoter were calculated (Table 1). As expected from the number of transcriptional units, at least six proteins exhibit ultraviolet sensitivities compatible with a location close to a promoter. Some of the ribosomal proteins with the lowest ultraviolet sensitivities of synthesis (such as L20) show at low doses of ultraviolet irradiation an increase of synthesis, and only at higher ultraviolet doses an inactivation (Fig. 3). These shoulders could be due to the inactivation of negative control mechanisms or simply reflect an artefact caused by the restriction of transcription to smaller stretches of DNA by the ultraviolet irradiation.
5 M. Hirsch-Kauffmann, M. Schweiger, P. Herrlich, H. Ponta, H.-J. Rahmsdorf, S.-H. Pai, and H.-G. Wittmann 473 Table 1. Distances of ribosomal protein genes from their promoters From the inactivation tangent data, such as those shown in Fig. 3, and from the calibration curve of Fig. 2, the distance of each gene from its promoter was obtained (in molecular weight of protein and of DNA that would correspond to the stretch between promoter and promoter-distal end of the gene). The data show good correlation in 4 independent experiments with the exception of those in brackets, which were more variable Protein s2 s3 s4 S6 s7 S8 s9 [SIO s11 [S13 S14 S15 S16 S17 S18 S19 s20 s21 L1 L2 L3 L4 L5 L6 L7 L8 L9 I L10 L11 L12 L13 L14 L15 L16 L17 L18 L19 L21 L22 [L23 L24 L25 L27 L29 L30 L32 Distance from promoter in DNA in protein (10-6 x M,) (10-3 x M,) The ultraviolet mapping of S18 is particularly interesting because S18 has been found by genetic methods to map at 84min [22], outside the region where other known ribosomal markers map. The ultraviolet sensitivity of S18 synthesis indicates that the gene for S18 is not close to a promoter. There would be space for four ribosomal protein genes or for other genes between the promoter and the S18 gene. The experiments reported here indicate the existence of several transcriptional units for ribosomal proteins. Our data are in agreement with those obtained by kinetic measurements [lo]. Our data would also explain the well-known flexibility of cells in the adjustment to nutrients and in the change of growth rate. At an average transcription rate of 50 nucleotides/s it would take 10min to express the whole ribosomal operon. The longest unit observed here would be expressed within 100 s. How the transcription of ribosomal protein cistrons is regulated is yet unknown. The ultraviolet data exclude, however, ribosomal RNA synthesis from being a controlling element. Because of the large size of the ribosomal RNA operon, ribosomal RNA synthesis is blocked at low doses of ultraviolet light, when little effect on the synthesis of ribosomal proteins was observed. REFERENCES 1. Maalqie, 0. (1969) Dev. Bid. Suppl. 3, Schleif, R. (1967) J. Mol. Biol. 27, Gausing, K. (1974) Mol. Gen. Genet. 129, Nikolaev, N., Silengo, L. & Schlessinger, D. (1973) Proc. Natl Acad. Sci. U.S.A. 70, Kossman, C. R., Stamato, T. D.&Pettijohn, D. E. (1971) Nut. New Biol. 234, Doolittle, W. F. & Pace, N. R. (1970) Nature (Lond.) 228, Bremer, H. & Berry, L. (1971) Nat. New Biol. 234, Osawa, S., Otaka, E., Takata, R., Dekio, S., Matsubara, M., Itoh, T. & Muto, A. (1972) in Functional Units in Protein Biosynthesis (Cox, R. A. & Hadjiolov, A. A., eds) pp , Academic Press, London. 9. Nomura, M. & Engback, F. (1972) Proc. Nut1 Acud. Sci. U.S.A. 69, Molin, S., von Meyenburg, K., Gullev, K. & Maalee, 0. (1974) Mol. Gen. Genet. 129, Herrlich, P., Rahmsdorf, H. J., Pai, S. H. & Schweiger, M. (1974) Proc. NatlAcad. Sci. U.S.A. 71, Hill, R. F. (1958) Biochim. Biophys. Acta, 30, Studier, F. W. (1973) J. Mol. Biol. 79, Williams. B. & Paigen, K. (1969) J. Bacterial. 97, Hardy, S. J. S., Kurland, C. G., Voynar, P. & Mom, P. (1969) Biochemistry, 8, Kaltschmidt, E. & Wittmann, H. G. (1970) Anal. Biochem. 36,
6 474 M. Hirsch-Kauffmann et al. : E. coli Ribosomal Proteins: Transcriptional Units 17. Michalke, H. & Bremer, H. (1969)J. Mol. Bid. 41, Wilson, D. B. & Hogness, D. S. (1974) J. Bid. Chrm. 18. Sauerbier, W., Millette, R. L. & Hackett, P. B. (1970) 244,2143. Biochim. Biophi~.s. Acta, 209, Dzionara, M., Kaltschmidt, E. & Wittmann, H. G. (1970) 19. Scherzinger. E.. Hcrrlich, P., Schweiger, H. & Schuster, H. Proc. Nut1 Acud. Sci. U.S.A. 67, (1972) Eui. J. Biochern. 25, Herzog, A,, C,abezon, T., Wilde, M. & Bollen, A. (1974) Mol. Gen. Genet. in press. M. Hirsch-Kauffmann, M. Schweiger, P. Herrlich, H. Ponta, H.-J. Rahmsdorf, S.-H. Pai, and H. G. Wittmann. Max-Planck-lnstitut fur Molekulare Genetik, D-1000 Berlin (West)-Dahlem 33, HarnackstraBe 23
Ribosomal Proteins of Escherichia coli*
 Proceedings of the National Academy of Sciences Vol. 67, No. 4, pp. 1909-1913, December 1970 Ribosomal Proteins, XIII. Molecular Weights of Isolated Ribosomal Proteins of Escherichia coli* M. Dzionara,
Proceedings of the National Academy of Sciences Vol. 67, No. 4, pp. 1909-1913, December 1970 Ribosomal Proteins, XIII. Molecular Weights of Isolated Ribosomal Proteins of Escherichia coli* M. Dzionara,
Problem-solving Test: The Mechanism of Protein Synthesis
 Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Yeast Ribosomal Proteins are Synthesized on Small Polysomes
 Eur. J. Biochem. 62, 193-197 (1976) Yeast Ribosomal Proteins are Synthesized on Small Polysomes Willem H. MAGER and Rudi J. PLANTA Biochemisch Laboratorium, Vrije Universiteit, Amsterdam (Received September
Eur. J. Biochem. 62, 193-197 (1976) Yeast Ribosomal Proteins are Synthesized on Small Polysomes Willem H. MAGER and Rudi J. PLANTA Biochemisch Laboratorium, Vrije Universiteit, Amsterdam (Received September
The Blueprint of Life: DNA to Protein. What is genetics? DNA Structure 4/27/2011. Chapter 7
 The Blueprint of Life: NA to Protein Chapter 7 What is genetics? The science of heredity; includes the study of genes, how they carry information, how they are replicated, how they are expressed NA Structure
The Blueprint of Life: NA to Protein Chapter 7 What is genetics? The science of heredity; includes the study of genes, how they carry information, how they are replicated, how they are expressed NA Structure
The Blueprint of Life: DNA to Protein
 The Blueprint of Life: NA to Protein Chapter 7 What is genetics? The science of heredity; includes the y; study of genes, how they carry information, how they are replicated, how they are expressed 1 NA
The Blueprint of Life: NA to Protein Chapter 7 What is genetics? The science of heredity; includes the y; study of genes, how they carry information, how they are replicated, how they are expressed 1 NA
hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide gel electrophoresis/genetics)
 Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
Separation of the Proteins in Human Tonsillar Cytoplasmic Ribosomes by Two-Dimensional Polyacrylamide-Gel Electrophoresis
 Eur. J. Biochem. 50, 183 189 (1974) Separation of the Proteins in Human Tonsillar Cytoplasmic Ribosomes by TwoDimensional PolyacrylamideGel Electrophoresis Ugur UCER and Engin BERMEK Abteilung Molekulare
Eur. J. Biochem. 50, 183 189 (1974) Separation of the Proteins in Human Tonsillar Cytoplasmic Ribosomes by TwoDimensional PolyacrylamideGel Electrophoresis Ugur UCER and Engin BERMEK Abteilung Molekulare
Characterization of the amino acids phosphorylated in E. coli proteins
 FEMS Microbiology Letters 17 (1983) 87-91 87 Published by Elsevier Biomedical Press Characterization of the amino acids phosphorylated in E. coli proteins 1. INTRODUCTION M. Manai and A.J. Cozzone Laboratory
FEMS Microbiology Letters 17 (1983) 87-91 87 Published by Elsevier Biomedical Press Characterization of the amino acids phosphorylated in E. coli proteins 1. INTRODUCTION M. Manai and A.J. Cozzone Laboratory
An Introduction to Carbohydrates
 An Introduction to Carbohydrates Carbohydrates are a large class of naturally occurring polyhydroxy aldehydes and ketones. Monosaccharides also known as simple sugars, are the simplest carbohydrates containing
An Introduction to Carbohydrates Carbohydrates are a large class of naturally occurring polyhydroxy aldehydes and ketones. Monosaccharides also known as simple sugars, are the simplest carbohydrates containing
DNA codes for RNA, which guides protein synthesis.
 Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
-are poly-hydroxylated aldehydes and ketones -can cyclise -can form polymeric chains
 CARBOHYDRATES -compounds of C, H and O -originally thought of as hydrates of carbon e.g. glucose C 6 H 12 O 6 thought to be C(H 2 O) carbohydrates: -are poly-hydroxylated aldehydes and ketones -can cyclise
CARBOHYDRATES -compounds of C, H and O -originally thought of as hydrates of carbon e.g. glucose C 6 H 12 O 6 thought to be C(H 2 O) carbohydrates: -are poly-hydroxylated aldehydes and ketones -can cyclise
COLI THE SYNTHESIS OF RIBOSOMES IN E. IV. THE SYNTHESIS OF RIBOSOMAL PROTEIN
 THE SYNTHESIS OF RIBOSOMES IN E. IV. THE SYNTHESIS OF RIBOSOMAL PROTEIN AND THE ASSEMBLY OF RIBOSOMES COLI R. J. BRITTEN, B. J. MCCARTHY, and R. B. ROBERTS From the Carnegie Institution of Washington,
THE SYNTHESIS OF RIBOSOMES IN E. IV. THE SYNTHESIS OF RIBOSOMAL PROTEIN AND THE ASSEMBLY OF RIBOSOMES COLI R. J. BRITTEN, B. J. MCCARTHY, and R. B. ROBERTS From the Carnegie Institution of Washington,
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
1) DNA unzips - hydrogen bonds between base pairs are broken by special enzymes.
 Biology 12 Cell Cycle To divide, a cell must complete several important tasks: it must grow, during which it performs protein synthesis (G1 phase) replicate its genetic material /DNA (S phase), and physically
Biology 12 Cell Cycle To divide, a cell must complete several important tasks: it must grow, during which it performs protein synthesis (G1 phase) replicate its genetic material /DNA (S phase), and physically
rrna Maturation as a Quality Control Step in Ribosomal Subunit Assembly in Dictyostelium discoideum*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 272, No. 44, Issue of October 31, pp. 27818 27822, 1997 1997 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. rrna Maturation
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 272, No. 44, Issue of October 31, pp. 27818 27822, 1997 1997 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. rrna Maturation
The Pools of Ribosomal Proteins and Ribosomal Ribonucleic Acids During Relaxed Control of Escherichia coli A19 (Hfr, re1 met ms)
 ~~ ~~ ~ Journal of General Microbiology (1979), 112, 149-159. Printed in Great Britain 149 The Pools of Ribosomal Proteins and Ribosomal Ribonucleic Acids During Relaxed Control of Escherichia coli A19
~~ ~~ ~ Journal of General Microbiology (1979), 112, 149-159. Printed in Great Britain 149 The Pools of Ribosomal Proteins and Ribosomal Ribonucleic Acids During Relaxed Control of Escherichia coli A19
Protein Synthesis
 Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
Protein Synthesis 10.6-10.16 Objectives - To explain the central dogma - To understand the steps of transcription and translation in order to explain how our genes create proteins necessary for survival.
RNA (Ribonucleic acid)
 RNA (Ribonucleic acid) Structure: Similar to that of DNA except: 1- it is single stranded polunucleotide chain. 2- Sugar is ribose 3- Uracil is instead of thymine There are 3 types of RNA: 1- Ribosomal
RNA (Ribonucleic acid) Structure: Similar to that of DNA except: 1- it is single stranded polunucleotide chain. 2- Sugar is ribose 3- Uracil is instead of thymine There are 3 types of RNA: 1- Ribosomal
Central Dogma. Central Dogma. Translation (mrna -> protein)
 Central Dogma Central Dogma Translation (mrna -> protein) mrna code for amino acids 1. Codons as Triplet code 2. Redundancy 3. Open reading frames 4. Start and stop codons 5. Mistakes in translation 6.
Central Dogma Central Dogma Translation (mrna -> protein) mrna code for amino acids 1. Codons as Triplet code 2. Redundancy 3. Open reading frames 4. Start and stop codons 5. Mistakes in translation 6.
 Effect of a Selenium Analogue of [L Title Transport of Candida pelliculosa (C Dedicated to Professor Masaya Okano Retirement) Author(s) Shimizu, Eiichi; Yamana, Ryutaro; T Kenji Citation Bulletin of the
Effect of a Selenium Analogue of [L Title Transport of Candida pelliculosa (C Dedicated to Professor Masaya Okano Retirement) Author(s) Shimizu, Eiichi; Yamana, Ryutaro; T Kenji Citation Bulletin of the
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins.
 Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Alternative RNA processing: Two examples of complex eukaryotic transcription units and the effect of mutations on expression of the encoded proteins. The RNA transcribed from a complex transcription unit
Student Number: THE UNIVERSITY OF MANITOBA April 10, 2006, 1:30 AM - 4:30 PM Page 1 (of 4) Biochemistry II Laboratory Section Final Examination
 Name: Student Number: April 10, 2006, 1:30 AM - 4:30 PM Page 1 (of 4) Biochemistry II Laboratory Section Final Examination Examiner: Dr. A. Scoot 1. Answer ALL questions in the space provided. 2. The back
Name: Student Number: April 10, 2006, 1:30 AM - 4:30 PM Page 1 (of 4) Biochemistry II Laboratory Section Final Examination Examiner: Dr. A. Scoot 1. Answer ALL questions in the space provided. 2. The back
Similarity in the Size and Number of Ribosomal
 JOURNAL OF BACTERIOLOGY, Aug. 1972, p. 474-480 Copyright i 1972 American Society for Microbiology Vol. 111, No. 2 Printed in U.S.A. Similarity in the Size and Number of Ribosomal Proteins from Different
JOURNAL OF BACTERIOLOGY, Aug. 1972, p. 474-480 Copyright i 1972 American Society for Microbiology Vol. 111, No. 2 Printed in U.S.A. Similarity in the Size and Number of Ribosomal Proteins from Different
BIL 256 Cell and Molecular Biology Lab Spring, Tissue-Specific Isoenzymes
 BIL 256 Cell and Molecular Biology Lab Spring, 2007 Background Information Tissue-Specific Isoenzymes A. BIOCHEMISTRY The basic pattern of glucose oxidation is outlined in Figure 3-1. Glucose is split
BIL 256 Cell and Molecular Biology Lab Spring, 2007 Background Information Tissue-Specific Isoenzymes A. BIOCHEMISTRY The basic pattern of glucose oxidation is outlined in Figure 3-1. Glucose is split
Ultrastructure of Mycoplasmatales Virus laidlawii x
 J. gen. Virol. (1972), I6, 215-22I Printed in Great Britain 2I 5 Ultrastructure of Mycoplasmatales Virus laidlawii x By JUDY BRUCE, R. N. GOURLAY, AND D. J. GARWES R. HULL* Agricultural Research Council,
J. gen. Virol. (1972), I6, 215-22I Printed in Great Britain 2I 5 Ultrastructure of Mycoplasmatales Virus laidlawii x By JUDY BRUCE, R. N. GOURLAY, AND D. J. GARWES R. HULL* Agricultural Research Council,
CHAPTER TWO MECHANISMS OF RADIATION EFFECTS
 10-2 densely ionizing radiation CHAPTER TWO MECHANISMS OF RADIATION EFFECTS 2.0 INTRODUCTION Cell survival curves describe the relationship between the fractional survival, S, of a population of radiated
10-2 densely ionizing radiation CHAPTER TWO MECHANISMS OF RADIATION EFFECTS 2.0 INTRODUCTION Cell survival curves describe the relationship between the fractional survival, S, of a population of radiated
Single Essential Amino Acids (valine/histidine/methiotiine/high-temperature inhibition)
 Proc. Nat. Acad. Sci. USA Vol. 68, No. 9, pp. 2057-2061, September 1971 Regulation of Protein Synthesis Initiation in HeLa Cells Deprived of Single ssential Amino Acids (valine/histidine/methiotiine/high-temperature
Proc. Nat. Acad. Sci. USA Vol. 68, No. 9, pp. 2057-2061, September 1971 Regulation of Protein Synthesis Initiation in HeLa Cells Deprived of Single ssential Amino Acids (valine/histidine/methiotiine/high-temperature
(44) is microconidiating, fluffy, inositolless, isoleucineless, and valineless.
 THE EFFECT OF PHOTOREACTIVATION ON MUTATION FREQUENCY IN NEUROSPORA' JEANETTE SNYDER BROWN Stanford University, Stanford, California Received for publication April 1, 1951 Kelner (1949a) first reported
THE EFFECT OF PHOTOREACTIVATION ON MUTATION FREQUENCY IN NEUROSPORA' JEANETTE SNYDER BROWN Stanford University, Stanford, California Received for publication April 1, 1951 Kelner (1949a) first reported
Electron Microscopy of Small Cells: Mycoplasma hominis
 JOURNAL of BAcTRiowOY, Dc. 1969, p. 1402-1408 Copyright 0 1969 American Society for Microbiology Vol. 100, No. 3 Printed In U.S.A. NOTES Electron Microscopy of Small Cells: Mycoplasma hominis JACK MANILOFF
JOURNAL of BAcTRiowOY, Dc. 1969, p. 1402-1408 Copyright 0 1969 American Society for Microbiology Vol. 100, No. 3 Printed In U.S.A. NOTES Electron Microscopy of Small Cells: Mycoplasma hominis JACK MANILOFF
Glycoprotein Synthesis by D-Glucosamine Hydrochloride
 JOURNAL OF VIROLOGY, Apr. 1974, p. 775-779 Copyright 0 1974 American Society for Microbiology Vol. 13, No. 4 Printed in U.S.A. Selective Inhibition of Newcastle Disease Virus-Induced Glycoprotein Synthesis
JOURNAL OF VIROLOGY, Apr. 1974, p. 775-779 Copyright 0 1974 American Society for Microbiology Vol. 13, No. 4 Printed in U.S.A. Selective Inhibition of Newcastle Disease Virus-Induced Glycoprotein Synthesis
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION TITLE: Structural Basis of Signal Sequence Surveillance and Selection by the SRP-SR Complex AUTHORS and AFFILIATIONS Ottilie von Loeffelholz 1,2, Kèvin Knoops 1,2,6, Aileen Ariosa
SUPPLEMENTARY INFORMATION TITLE: Structural Basis of Signal Sequence Surveillance and Selection by the SRP-SR Complex AUTHORS and AFFILIATIONS Ottilie von Loeffelholz 1,2, Kèvin Knoops 1,2,6, Aileen Ariosa
SelenoMet Dream TM Media Kit (MD12-506)
 SelenoMet Dream TM Media Kit (MD12-506) For the Overnight Expression of Recombinant Proteins About the Kit: MD12-506 contains: (*each packet contains enough to make up 1L of media) Description For the
SelenoMet Dream TM Media Kit (MD12-506) For the Overnight Expression of Recombinant Proteins About the Kit: MD12-506 contains: (*each packet contains enough to make up 1L of media) Description For the
Estimations of the Molecular Weight of the Influenza Virus Genome
 o r. gem Viral. &97I), H, Io3-Io9 103 Printed in Great Britain Estimations of the Molecular Weight of the Influenza Virus Genome By J. J. SKEHEL National Institute for Medical Research, Mill Hill, London
o r. gem Viral. &97I), H, Io3-Io9 103 Printed in Great Britain Estimations of the Molecular Weight of the Influenza Virus Genome By J. J. SKEHEL National Institute for Medical Research, Mill Hill, London
Control of Prokaryotic (Bacterial) Genes
 Control of Prokaryotic (Bacterial) Genes 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have enough of a product, need to stop production
Control of Prokaryotic (Bacterial) Genes 2007-2008 Bacterial metabolism Bacteria need to respond quickly to changes in their environment STOP GO if they have enough of a product, need to stop production
Point total. Page # Exam Total (out of 90) The number next to each intermediate represents the total # of C-C and C-H bonds in that molecule.
 This exam is worth 90 points. Pages 2- have questions. Page 1 is for your reference only. Honor Code Agreement - Signature: Date: (You agree to not accept or provide assistance to anyone else during this
This exam is worth 90 points. Pages 2- have questions. Page 1 is for your reference only. Honor Code Agreement - Signature: Date: (You agree to not accept or provide assistance to anyone else during this
Loss of Sensitivity to EDTA by Pseudomonas aeruginosa Grown under Conditions of Mg-Limitation
 J. gen. Microbiol. (1g6g), 54, 439-444 Printed in Great Britain 439 Loss of Sensitivity to EDTA by Pseudomonas aeruginosa Grown under Conditions of Mg-Limitation By M. R. W. BROWN AND J. MELLING Pharmaceutical
J. gen. Microbiol. (1g6g), 54, 439-444 Printed in Great Britain 439 Loss of Sensitivity to EDTA by Pseudomonas aeruginosa Grown under Conditions of Mg-Limitation By M. R. W. BROWN AND J. MELLING Pharmaceutical
Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples:
 Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
The Synthesis of Vitamin B, by some Mutant Strains of Escherichia coli
 597 MORRIS, J. G. (1959). J. gen. Mimobiol. 20, 5 974 The Synthesis of Vitamin B, by some Mutant Strains of Escherichia coli BY J. G. MORRIS Microbiology Unit, Department of Biochemistry, University of
597 MORRIS, J. G. (1959). J. gen. Mimobiol. 20, 5 974 The Synthesis of Vitamin B, by some Mutant Strains of Escherichia coli BY J. G. MORRIS Microbiology Unit, Department of Biochemistry, University of
THE QUANTITATIVE GLUCOSE AND MINERAL NUTRIENT REQUIREMENTS OF MOUSE LS (SUSPENSION) CELLS IN CHEMICALLY DEFINED MEDIUM
 J. Cell Sci. 8, 693-700 (1971) Printed in Great Britain THE QUANTITATIVE GLUCOSE AND MINERAL NUTRIENT REQUIREMENTS OF MOUSE LS (SUSPENSION) CELLS IN CHEMICALLY DEFINED MEDIUM J. R. BIRCH* AND S. J. PIRT
J. Cell Sci. 8, 693-700 (1971) Printed in Great Britain THE QUANTITATIVE GLUCOSE AND MINERAL NUTRIENT REQUIREMENTS OF MOUSE LS (SUSPENSION) CELLS IN CHEMICALLY DEFINED MEDIUM J. R. BIRCH* AND S. J. PIRT
Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding
 SUPPLEMENTARY INFORMATION: Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding Gong Zhang 1,2, Magdalena Hubalewska 1 & Zoya Ignatova 1,2 1 Department of Cellular
SUPPLEMENTARY INFORMATION: Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding Gong Zhang 1,2, Magdalena Hubalewska 1 & Zoya Ignatova 1,2 1 Department of Cellular
Virus and Prokaryotic Gene Regulation - 1
 Virus and Prokaryotic Gene Regulation - 1 We have discussed the molecular structure of DNA and its function in DNA duplication and in transcription and protein synthesis. We now turn to how cells regulate
Virus and Prokaryotic Gene Regulation - 1 We have discussed the molecular structure of DNA and its function in DNA duplication and in transcription and protein synthesis. We now turn to how cells regulate
Overview of the Expressway Cell-Free Expression Systems. Expressway Mini Cell-Free Expression System
 Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Objectives: Prof.Dr. H.D.El-Yassin
 Protein Synthesis and drugs that inhibit protein synthesis Objectives: 1. To understand the steps involved in the translation process that leads to protein synthesis 2. To understand and know about all
Protein Synthesis and drugs that inhibit protein synthesis Objectives: 1. To understand the steps involved in the translation process that leads to protein synthesis 2. To understand and know about all
130327SCH4U_biochem April 09, 2013
 Option B: B1.1 ENERGY Human Biochemistry If more energy is taken in from food than is used up, weight gain will follow. Similarly if more energy is used than we supply our body with, weight loss will occur.
Option B: B1.1 ENERGY Human Biochemistry If more energy is taken in from food than is used up, weight gain will follow. Similarly if more energy is used than we supply our body with, weight loss will occur.
previously1 were used with minor modifications in the studies described
 INHIBITION BY CERTAIN PTERIDINES OF RIBOSOMAL RNA AND DNA SYNTHESIS IN DEVELOPING ONCOPELTUS EGGS* BY S. E. HARRIS AND H. S. FORREST GENETICS FOUNDATION, THE UNIVERSITY OF TEXAS, AUSTIN Communicated by
INHIBITION BY CERTAIN PTERIDINES OF RIBOSOMAL RNA AND DNA SYNTHESIS IN DEVELOPING ONCOPELTUS EGGS* BY S. E. HARRIS AND H. S. FORREST GENETICS FOUNDATION, THE UNIVERSITY OF TEXAS, AUSTIN Communicated by
bacteria (burst size). It consists in diluting a suspension of infected2 Only a small fraction of the samples will then contain more than one infected
 THE BURST SIZE DISTRIBUTION IN THE GROWTH OF BACTERIAL VIRUSES (BACTERIOPHAGES)l M. DELBRtrCK Departments of Physics and Biology, Vanderbilt University, Nashville, Tennessee Received for publication January
THE BURST SIZE DISTRIBUTION IN THE GROWTH OF BACTERIAL VIRUSES (BACTERIOPHAGES)l M. DELBRtrCK Departments of Physics and Biology, Vanderbilt University, Nashville, Tennessee Received for publication January
X-RAY-INDUCED CELL KILLING AND MUTATIONS IN CULTURED HUMAN CELL LINES (XERODERMA PIGMENTOSUM CELLS AND HELA 53 CELLS)
 JAPAN. J. GENETICS Vol. 54, No. 6: 415-426 (1979) X-RAY-INDUCED CELL KILLING AND MUTATIONS IN CULTURED HUMAN CELL LINES (XERODERMA PIGMENTOSUM CELLS AND HELA 53 CELLS) YOSHIHIRO MURAII~, SACHIKO TATSUKAWAZ~
JAPAN. J. GENETICS Vol. 54, No. 6: 415-426 (1979) X-RAY-INDUCED CELL KILLING AND MUTATIONS IN CULTURED HUMAN CELL LINES (XERODERMA PIGMENTOSUM CELLS AND HELA 53 CELLS) YOSHIHIRO MURAII~, SACHIKO TATSUKAWAZ~
Pre-mRNA has introns The splicing complex recognizes semiconserved sequences
 Adding a 5 cap Lecture 4 mrna splicing and protein synthesis Another day in the life of a gene. Pre-mRNA has introns The splicing complex recognizes semiconserved sequences Introns are removed by a process
Adding a 5 cap Lecture 4 mrna splicing and protein synthesis Another day in the life of a gene. Pre-mRNA has introns The splicing complex recognizes semiconserved sequences Introns are removed by a process
Chemistry 107 Exam 4 Study Guide
 Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
Chemistry 107 Exam 4 Study Guide Chapter 10 10.1 Recognize that enzyme catalyze reactions by lowering activation energies. Know the definition of a catalyst. Differentiate between absolute, relative and
Early Repair Processes in Marrow Cells Irradiated and Proliferating in Vivo1
 RADIATION RESEARCH 18, 96-105 (1963) Early Repair Processes in Marrow Cells Irradiated and Proliferating in Vivo1 J. E. TILL AND E. A. McCULLOCH Department of Medical Biophysics, University of Toronto,
RADIATION RESEARCH 18, 96-105 (1963) Early Repair Processes in Marrow Cells Irradiated and Proliferating in Vivo1 J. E. TILL AND E. A. McCULLOCH Department of Medical Biophysics, University of Toronto,
Transcriptional Mapping of Rabies Virus In Vivo
 JOURNAL OF VIROLOGY, Nov. 1978, P. 518-523 0022-538X/78/0028-0518$02.00/0 Copyright 1978 American Society for Microbiology Transcriptional Mapping of Rabies Virus In Vivo Vol. 28, No. 2 Printed in U.S.A.
JOURNAL OF VIROLOGY, Nov. 1978, P. 518-523 0022-538X/78/0028-0518$02.00/0 Copyright 1978 American Society for Microbiology Transcriptional Mapping of Rabies Virus In Vivo Vol. 28, No. 2 Printed in U.S.A.
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
number Done by Corrected by Doctor Ashraf
 number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
AP Biology Reading Guide. Concept 19.1 A virus consists of a nucleic acid surrounded by a protein coat
 AP Biology Reading Guide Name Chapter 19: Viruses Overview Experimental work with viruses has provided important evidence that genes are made of nucleic acids. Viruses were also important in working out
AP Biology Reading Guide Name Chapter 19: Viruses Overview Experimental work with viruses has provided important evidence that genes are made of nucleic acids. Viruses were also important in working out
Supplementary methods 1: Parameter estimation
 Supplementary methods : Parameter estimation A number of estimates of the parameters in model () of the main text can be obtained from the literature. However these estimates are not necessarily obtained
Supplementary methods : Parameter estimation A number of estimates of the parameters in model () of the main text can be obtained from the literature. However these estimates are not necessarily obtained
10 mm KCl in a Ti-15 zonal rotor at 35,000 rpm for 16 hr at
 Proc. Nat. Acad. SCi. USA Vol. 68, No. 11, pp. 2752-2756, November 1971 Translation of Exogenous Messenger RNA for Hemoglobin on Reticulocyte and Liver Ribosomes (initiation factors/9s RNA/liver factors/reticulocyte
Proc. Nat. Acad. SCi. USA Vol. 68, No. 11, pp. 2752-2756, November 1971 Translation of Exogenous Messenger RNA for Hemoglobin on Reticulocyte and Liver Ribosomes (initiation factors/9s RNA/liver factors/reticulocyte
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis
 JOURNAL OF PROTEOMICS 74 (2011) 431 441 available at www.sciencedirect.com www.elsevier.com/locate/jprot N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis Masahiro Kamita
JOURNAL OF PROTEOMICS 74 (2011) 431 441 available at www.sciencedirect.com www.elsevier.com/locate/jprot N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis Masahiro Kamita
PROTEIN SYNTHESIS. It is known today that GENES direct the production of the proteins that determine the phonotypical characteristics of organisms.
 PROTEIN SYNTHESIS It is known today that GENES direct the production of the proteins that determine the phonotypical characteristics of organisms.» GENES = a sequence of nucleotides in DNA that performs
PROTEIN SYNTHESIS It is known today that GENES direct the production of the proteins that determine the phonotypical characteristics of organisms.» GENES = a sequence of nucleotides in DNA that performs
Student Number: THE UNIVERSITY OF MANITOBA April 11, 2011, 1:00 PM - 4:00 PM Page 1 (of 3)
 Name: Student Number: THE UNIVERSITY OF MANITOBA April 11, 2011, 1:00 PM - 4:00 PM Page 1 (of 3) Biochemistry II Laboratory Section Examiners: Drs. J. Galka 1. Answer ALL questions in the space provided.
Name: Student Number: THE UNIVERSITY OF MANITOBA April 11, 2011, 1:00 PM - 4:00 PM Page 1 (of 3) Biochemistry II Laboratory Section Examiners: Drs. J. Galka 1. Answer ALL questions in the space provided.
GLYCATION OF PROTEINS IN ESCHERICHIA COLI: EFFECT OF NUTRIENT BROTH INGREDIENTS ON GLYCATION
 Industry GLYCATION OF PROTEINS IN ESCHERICHIA COLI: EFFECT OF NUTRIENT BROTH INGREDIENTS ON GLYCATION R. Dimitrova, R. Mironova, I. Ivanov Institute of Molecular biology, Bulgarian Academy of Sciences
Industry GLYCATION OF PROTEINS IN ESCHERICHIA COLI: EFFECT OF NUTRIENT BROTH INGREDIENTS ON GLYCATION R. Dimitrova, R. Mironova, I. Ivanov Institute of Molecular biology, Bulgarian Academy of Sciences
MODULE 3: TRANSCRIPTION PART II
 MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
An Introduction to Carbohydrates
 An Introduction to Carbohydrates Carbohydrates are a large class of naturally occurring polyhydroxy aldehydes and ketones. Monosaccharides also known as simple sugars, are the simplest carbohydrates containing
An Introduction to Carbohydrates Carbohydrates are a large class of naturally occurring polyhydroxy aldehydes and ketones. Monosaccharides also known as simple sugars, are the simplest carbohydrates containing
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
Student Number: THE UNIVERSITY OF MANITOBA April 16, 2007, 9:00 AM -12:00 PM Page 1 (of 4) Biochemistry II Laboratory Section Final Examination
 Name: Student Number: THE UNIVERSITY OF MANITOBA April 16, 2007, 9:00 AM -12:00 PM Page 1 (of 4) Biochemistry II Laboratory Section Final Examination MBIO / CHEM.2370 Examiner: Dr. A. Scoot 1. Answer ALL
Name: Student Number: THE UNIVERSITY OF MANITOBA April 16, 2007, 9:00 AM -12:00 PM Page 1 (of 4) Biochemistry II Laboratory Section Final Examination MBIO / CHEM.2370 Examiner: Dr. A. Scoot 1. Answer ALL
Blocking by Histones of Accessibility to DNA in Chromatin (DNase/RNA polymerase/dna polymerase)
 Proc. Nat. Acad. Sci. USA Vol. 69, No. 8, pp. 2115-2119, August 1972 Blocking by Histones of Accessibility to in Chromatin (/RNA polymerase/ polymerase) ALFRED E. MIRSKY AND BERT SILVERMAN The Rockefeller
Proc. Nat. Acad. Sci. USA Vol. 69, No. 8, pp. 2115-2119, August 1972 Blocking by Histones of Accessibility to in Chromatin (/RNA polymerase/ polymerase) ALFRED E. MIRSKY AND BERT SILVERMAN The Rockefeller
Ch. 18 Regulation of Gene Expression
 Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Ch. 18 Regulation of Gene Expression 1 Human genome has around 23,688 genes (Scientific American 2/2006) Essential Questions: How is transcription regulated? How are genes expressed? 2 Bacteria regulate
Very stable prokaryotic messenger RNA in chromosomeless Escherichia coli minicells
 Proc. Nat. Acad. Sci. USA Vol. 7, No. 8, pp. 900-904, August 1975 Biochemistry Very stable prokaryotic messenger RNA in chromosomeless Escherichia coli minicells (outer membrane protein/lipoprotein/r plasmid)
Proc. Nat. Acad. Sci. USA Vol. 7, No. 8, pp. 900-904, August 1975 Biochemistry Very stable prokaryotic messenger RNA in chromosomeless Escherichia coli minicells (outer membrane protein/lipoprotein/r plasmid)
3. What law of heredity explains that traits, like texture and color, are inherited independently of each other?
 Section 2: Genetics Chapter 11 pg. 308-329 Part 1: Refer to the table of pea plant traits on the right. Then complete the table on the left by filling in the missing information for each cross. 6. What
Section 2: Genetics Chapter 11 pg. 308-329 Part 1: Refer to the table of pea plant traits on the right. Then complete the table on the left by filling in the missing information for each cross. 6. What
Spore Formation Induced by Glycerol, Dimethyl Sulfoxide,
 JOURNAL OF BACTERIOLOGY, Dec. 1980, p. 1076-1082 0021-9193/80/12-1076/07$2.00/0 Vol. 144, No. 3 Patterns of Protein Production in Myxococcus xanthus During Spore Formation Induced by Glycerol, Dimethyl
JOURNAL OF BACTERIOLOGY, Dec. 1980, p. 1076-1082 0021-9193/80/12-1076/07$2.00/0 Vol. 144, No. 3 Patterns of Protein Production in Myxococcus xanthus During Spore Formation Induced by Glycerol, Dimethyl
brought to you by and REFERENCES
 brought to you by www.thebacteriophages.org and www.phage.org REFERENCES 1. Butcher, S. J., T. Dokland, P. M. Ojala, D. H. Bamford, and S. D. Fuller. 1997. Intermediates in the assembly pathway of the
brought to you by www.thebacteriophages.org and www.phage.org REFERENCES 1. Butcher, S. J., T. Dokland, P. M. Ojala, D. H. Bamford, and S. D. Fuller. 1997. Intermediates in the assembly pathway of the
Student Number: To form the polar phase when adsorption chromatography was used.
 Name: Student Number: April 14, 2001, 1:30 AM - 4:30 PM Page 1 (of 4) Biochemistry II Lab Section Final Examination Examiner: Dr. A. Scoot 1. Answer ALL questions in the space provided.. 2. The last page
Name: Student Number: April 14, 2001, 1:30 AM - 4:30 PM Page 1 (of 4) Biochemistry II Lab Section Final Examination Examiner: Dr. A. Scoot 1. Answer ALL questions in the space provided.. 2. The last page
17 Cell Differentiation and Gene Expression investigation and modeling 1 2 class sessions
 17 Cell Differentiation and Gene Expression investigation and modeling 1 2 class sessions ACTIVITY OVERVIEW Students investigate gene expression as it relates to cell differentiation in four human cell
17 Cell Differentiation and Gene Expression investigation and modeling 1 2 class sessions ACTIVITY OVERVIEW Students investigate gene expression as it relates to cell differentiation in four human cell
Supplementary material: Materials and suppliers
 Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Proteins. Amino acids, structure and function. The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka
 Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Insulin mrna to Protein Kit
 Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
f(x) = x R² = RPKM (M8.MXB) f(x) = x E-014 R² = 1 RPKM (M31.
 14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
14 12 f(x) = 1.633186874x - 21.46732234 R² =.995616541 RPKM (M8.MXA) 1 8 6 4 2 2 4 6 8 1 12 14 RPKM (M8.MXB) 14 12 f(x) =.821767782x - 4.192595677497E-14 R² = 1 RPKM (M31.XA) 1 8 6 4 2 2 4 6 8 1 12 14
3) How many different amino acids are proteogenic in eukaryotic cells? A) 12 B) 20 C) 25 D) 30 E) None of the above
 Suggesting questions for Biochemistry 1 and 2 and clinical biochemistry 1) Henderson Hasselbalch Equation shows: A) The relationship between ph and the concentration of an acid and its conjugate base B)
Suggesting questions for Biochemistry 1 and 2 and clinical biochemistry 1) Henderson Hasselbalch Equation shows: A) The relationship between ph and the concentration of an acid and its conjugate base B)
ELECTROPHORETIC STUDIES OF SONIC EXTRACTS OF PROTEUS VULGARIS
 ELECTROPHORETIC STUDIES OF SONIC EXTRACTS OF PROTEUS VULGARIS I. EFFECT OF GROWTH ENVIRONMENT ON ELECTROPHORETIC PATTERNS' SIDNEY D. RODENBERG Laboratory of Microbiology, Division of Biology, University
ELECTROPHORETIC STUDIES OF SONIC EXTRACTS OF PROTEUS VULGARIS I. EFFECT OF GROWTH ENVIRONMENT ON ELECTROPHORETIC PATTERNS' SIDNEY D. RODENBERG Laboratory of Microbiology, Division of Biology, University
Viruses defined acellular organisms genomes nucleic acid replicate inside host cells host metabolic machinery ribosomes
 The Viruses Viruses Viruses may be defined as acellular organisms whose genomes consist of nucleic acid, obligately replicate inside host cells using host metabolic machinery and ribosomes to form a pool
The Viruses Viruses Viruses may be defined as acellular organisms whose genomes consist of nucleic acid, obligately replicate inside host cells using host metabolic machinery and ribosomes to form a pool
Paper Reference. Paper Reference(s) 6101/01 Edexcel GCE Biology Biology (Human) Advanced Subsidiary Unit Test 1
 Centre No. Candidate No. Paper Reference(s) 6101/01 Edexcel GCE Biology Biology (Human) Advanced Subsidiary Unit Test 1 Monday 4 June 2007 Morning Time: 1 hour Materials required for examination Ruler
Centre No. Candidate No. Paper Reference(s) 6101/01 Edexcel GCE Biology Biology (Human) Advanced Subsidiary Unit Test 1 Monday 4 June 2007 Morning Time: 1 hour Materials required for examination Ruler
SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons
 LECTROPHORES/S Revised November 1992 SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons I NTRODUCTION Electrophoresis in polyacrylamide gels in the presence of sodium dodecyl sulfate (SDS), an anionic detergent,
LECTROPHORES/S Revised November 1992 SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons I NTRODUCTION Electrophoresis in polyacrylamide gels in the presence of sodium dodecyl sulfate (SDS), an anionic detergent,
Reconstitution of Neutral Amino Acid Transport From Partially Purified Membrane Components From Ehrlich Ascites Tumor Cells
 Journal of Supramolecular Structure 7:481-487 (1977) Molecular Aspects of Membrane Transport 5 1 1-5 17 Reconstitution of Neutral Amino Acid Transport From Partially Purified Membrane Components From Ehrlich
Journal of Supramolecular Structure 7:481-487 (1977) Molecular Aspects of Membrane Transport 5 1 1-5 17 Reconstitution of Neutral Amino Acid Transport From Partially Purified Membrane Components From Ehrlich
VIRUSES. 1. Describe the structure of a virus by completing the following chart.
 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
Biochemistry 2000 Sample Question Transcription, Translation and Lipids. (1) Give brief definitions or unique descriptions of the following terms:
 (1) Give brief definitions or unique descriptions of the following terms: (a) exon (b) holoenzyme (c) anticodon (d) trans fatty acid (e) poly A tail (f) open complex (g) Fluid Mosaic Model (h) embedded
(1) Give brief definitions or unique descriptions of the following terms: (a) exon (b) holoenzyme (c) anticodon (d) trans fatty acid (e) poly A tail (f) open complex (g) Fluid Mosaic Model (h) embedded
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
AS BIOLOGY 7401/2. Paper 2
 Please write clearly, in block capitals. Centre number Candidate number Surname Forename(s) Candidate signature AS BIOLOGY Paper 2 Specimen materials (set 2) 1 hour 30 minutes Materials For this paper
Please write clearly, in block capitals. Centre number Candidate number Surname Forename(s) Candidate signature AS BIOLOGY Paper 2 Specimen materials (set 2) 1 hour 30 minutes Materials For this paper
Section 9. Junaid Malek, M.D.
 Section 9 Junaid Malek, M.D. Mutation Objective: Understand how mutations can arise, and how beneficial ones can alter populations Mutation= a randomly produced, heritable change in the nucleotide sequence
Section 9 Junaid Malek, M.D. Mutation Objective: Understand how mutations can arise, and how beneficial ones can alter populations Mutation= a randomly produced, heritable change in the nucleotide sequence
Ali Alabbadi. Bann. Bann. Dr. Belal
 31 Ali Alabbadi Bann Bann Dr. Belal Topics to be discussed in this sheet: Particles-to-PFU Single-step and multi-step growth cycles Multiplicity of infection (MOI) Physical measurements of virus particles
31 Ali Alabbadi Bann Bann Dr. Belal Topics to be discussed in this sheet: Particles-to-PFU Single-step and multi-step growth cycles Multiplicity of infection (MOI) Physical measurements of virus particles
Expression constructs
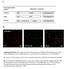 Gene expressed in bebe3 ZmBEa Expression constructs 35S ZmBEa Pnos:Hygromycin r 35S Pnos:Hygromycin r 35S ctp YFP Pnos:Hygromycin r B -1 Chl YFP- Merge Supplemental Figure S1: Constructs Used for the Expression
Gene expressed in bebe3 ZmBEa Expression constructs 35S ZmBEa Pnos:Hygromycin r 35S Pnos:Hygromycin r 35S ctp YFP Pnos:Hygromycin r B -1 Chl YFP- Merge Supplemental Figure S1: Constructs Used for the Expression
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
DNA, RNA, protein and DNases in developing rat cerebellum: Effects of early postnatal nutritional deprivation
 Biosci.,Vol. 4, Number 4, December 1982, pp. 391-400. Printed in India. DNA, RNA, protein and DNases in developing rat cerebellum: Effects of early postnatal nutritional deprivation K. V. SUBBA RAO AND
Biosci.,Vol. 4, Number 4, December 1982, pp. 391-400. Printed in India. DNA, RNA, protein and DNases in developing rat cerebellum: Effects of early postnatal nutritional deprivation K. V. SUBBA RAO AND
A protocol for enhancement of the AAV-mediated expression of transgenes
 A protocol for enhancement of the AAV-mediated expression of transgenes Hiroaki Mizukami, Takeharu Kanazawa, Takashi Okada, and Keiya Ozawa Division of Genetic Therapeutics, Center for Molecular Medicine,
A protocol for enhancement of the AAV-mediated expression of transgenes Hiroaki Mizukami, Takeharu Kanazawa, Takashi Okada, and Keiya Ozawa Division of Genetic Therapeutics, Center for Molecular Medicine,
GENE EXPRESSION. Amoeba Sisters video 3pk9YVo. Individuality & Mutations
 Amoeba Sisters video https://www.youtube.com/watch?v=giez 3pk9YVo GENE EXPRESSION Individuality & Mutations Complete video handout http://www.amoebasisters.com/uploads/ 2/1/9/0/21902384/video_recap_of_muta
Amoeba Sisters video https://www.youtube.com/watch?v=giez 3pk9YVo GENE EXPRESSION Individuality & Mutations Complete video handout http://www.amoebasisters.com/uploads/ 2/1/9/0/21902384/video_recap_of_muta
Microbiology - Problem Drill 16: Antibiotics. Question No. 1 of 10. Question. Feedback. Question
 Microbiology - Problem Drill 16: Antibiotics No. 1 of 10 1. An effective chemotherapeutic drug should have. (A) Low therapeutic index (B) More toxicity (C) Selective toxicity (D) Mutation inducing properties
Microbiology - Problem Drill 16: Antibiotics No. 1 of 10 1. An effective chemotherapeutic drug should have. (A) Low therapeutic index (B) More toxicity (C) Selective toxicity (D) Mutation inducing properties
