Supplemental Data. Takahashi et al. Plant Cell (2014) /tpc
|
|
|
- Rosamond West
- 5 years ago
- Views:
Transcription
1 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc Fructose Glucose Sucrose Gentiobiose Gentianose Fructose STDs +GLU +INV Gentianose Supplemental Figure. TLC analysis of sugars hydrolyzed from gentianose. Gentianose (GF) were hydrolyzed by β-glucosidase obtained from Ustilago esculenta (GLU; Nakajima et al., ) and purified recombinant protein of INV. After hydrolysis, sugars were separated by silica gel TLC with ethyl acetate-acetic acid-water (3::) as the mobile phase and stained thymol-sufuric acid reagent. Fructose was stained pink color, but glucose was stained violet color.
2 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc cv. Maciry cv. SpB nmol mg - DW 5 5 G 3 G nmol mg - DW GF.... nmol mg - DW 5 5 GF Relative expression.... nmo mg - DW Relative expression INV INV Supplemental Figure. Variety comparison of changes in gentiooligosaccharide concentrations and INV expression in OWBs. The concentration of gentiobiose (G) and gentianose (GF), as well as the expression level of INV in OWBs of cv. Maciry and cv. SpB were compared. Each value represents the mean ± SD calculated from three independent experiments.
3 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc A [kda] [kda] M INV M INV CBB anti-myc antibody B GF hydrolytic activity (%) ph Temperature ( o C) Supplemental Figure 3. Characterization of recombinant INV protein. (A) SDS-PAGE and western-blot analysis of recombinant INV. Purified INV protein produced in P. pastoris was detected by Coomassie brilliant blue (CBB) staining or westernblotting using the anti-myc antibody. (B) The effect of ph and temperature on gentianose (GF) hydrolytic activity of INV. Each value represents the mean ± SD calculated from triplicate measurements.
4 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc GFP BF Merge At-GRP5 INV Control Supplemental Figure 4. Subcellular localization of INV-sGFP. Transiently expressed GFP (control, upper-row) and GFP-fusion protein (INV, middle row) in Nicotiana benthamiana leaves. GFP channel (left), bright field (BF, center) and merged image (right) are shown. Bars indicate 5 µm. The subcellular localization of the GFP fusion protein was examined 3 days after incubation under constitutive dark conditions. The At-GFP5 protein was used as a vacuolar protein marker (Mangeon et al, 9).
5 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc T B G-AP (.5 mg/ml) T B T B T B IOWB IOWB IOWB 3 Supplemental Figure 5. Uptake of G-AP into IOWBs. IOWBs were cultured on solid MS medium containing.5% G-AP at o C with a 6/8 h light/dark photoperiod. After 4 h, IOWBs (n=3) were divided in half, and metabolites in the top (T) and bottom (B) halves of IOWBs were extracted as described in METHODS. G-AP was separated by TLC and the fluorescence of G-AP was detected.
6 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc Relative expression 4 3 PDH nmol mg - DW G+GF Supplemental Figure 6. Changes in PDH expression and the sum concentrations of gentiobiose and gentianose in field grown OWBs. The expression level of PDH and the sum of gentiobiose (G) and gentianose (GF) concentrations in OWBs of cv. SpB was measured. Each value represents the mean ± SD calculated from three independent experiments.
7 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc SiR.5 OASL.5 OASL 3 OASL3.5 GCL Relative expressions GSHS. EGS. CBL 4 MeS 6 MeS SAMS 8 SAMS 4 SAMDC.4 ACO 3 SAHH SAHH 5 5 DHAR DHAR 5 5 GR 5 5 GR Supplemental Figure 7. Changes in gentiobiose-affected gene expression in field-grown OWBs before and during budbreak. Expression levels of genes that gentiobiose induced in IOWBs were compared in field grown OWBs. Each value represents the mean ± SD calculated from three independent experiments.
8 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc Control week 4 µm ethephon week 5 mm ACC week Control 3 weeks 4 µm ethephon 3 weeks 5 mm ACC 3 weeks Supplemental Figure 8. Effect of treatment of ethylene precursors on budbreak of gentian OWBs. Field-grown endodormant OWBs (G. triflora cv. SpB) harvested in September were treated with water (Control), 4 µm ethephon, or 5 mm ACC. After incubation at oc for 3 weeks, budbreak of OWBs was measured. Scale bars indicate cm.
9 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc SF ( day) % G ( day) G (nmol mg - DW) Hours Met (nmol mg - DW) Hours SF ( days) SF (3 days) % G ( days) % G (3 days) SAM (nmol mg - DW) Glu-cys (nmol mg - DW) Hours 4 36 Hours Cys (nmol mg - DW) GSH (nmol mg - DW) Hours 4 36 Hours Supplemental Figure 9. Effect of gentiobiose treatment on germination of Arabidopsis seeds. Arabidopsis seeds were placed on sugar-free (SF) or % gentiobiose (G)-containing MS agar medium and incubated at 4 o C for 4 days before transferring to a growth chamber at o C under continuous light (6 µmol m - s - ). Metabolites in the seeds were extracted and quantified as described in METHODS. Each value represents the mean ± SD calculated from three independent experiments.
10 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc Supplemental Table. Metabolite profiles in OWBs (G. triflora x G. scabra (5-543)) harvested in October, uary, and ch Concentrations (nmol g - DW) PC loadings October uary ch PC PC Gly 44. ± ± ± Ala 4.8 ± ± ± Asn ± ± ± Asp ± ± ± Lys 3.53 ± ± ± Gln ± ± ± Glu ± ± ± His ± ± ± Arg ± ± ± Ser 3.8 ± ± ± Pro ± ± ± Val ± ± ± Thr 495. ± ± ± Leu/Ile 4.4 ± ± ± Met.34 ± ± ± Phe 6.7 ± ± ± Tyr. ± ± ± Trp ± ± ± Putrescine ± ± ± GABA ± ± ± Spermidine ± ± ± Citrulline 6.7 ± ± ± Spermine 9.68 ± ± ± Arg-suc 4.3 ± ±..38 ± SAM 5.44 ± ± ± Glucose 583. ± ± ± Sucrose ± ± ± Gentiobiose ± ± ± Gentianose ± ± ± Arabinose 8. ± ± ± Galactose 6.67 ± ± ± AMP 4.75 ± ± ± ADP 79.4 ± ± ± ATP 83.4 ± ± ± GMP 8.9 ± ±.64.4 ± GDP 3. ± ± ± GTP 36.7 ± ± ± UDPG ± ± ± NAD ± ± ± NADP 5.35 ± ± ± NADPH ± ± ± Pyruvate ± ± ± Fumarate ± ± ± Succinate 5.78 ± ±.79.4 ± OG ± ± ± PEP.55 ± ± ± DHAP 3.74 ± ± ± Aconitate 6.66 ± ± ± Shikimate.9 ± ± ± PGA 69.9 ± ± ± Ru5P/R5P 366. ± ± ± G6P ± ± ± PG.3 ± ± ± RuBP 3.33 ± ± ± FBP 4.7 ± ± ± Isocitrate ± ± ± Citrate ± ± ± Malate ± ± ± Data presented are means ± SD of five independent experiments. DW, dry weight; GABA, γ-amino butyrate; SAM, S- adenosyl methionine; UDPG, UDP glucose; 3PGA, 3-phosphoglycerate; PEP, phosphoenolpyruvate; DHAP, dihydroxyacetone phosphate; Ru5P, ribulose 5-phosphate; R5P, Rib 5-phosphate; G6P, Glc 6-phosphate; 6PG, 6- phosphogluconate; RuBP, ribulose,5-bisphosphate; FBP, Fru,6-bisphosphate.
11 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc Supplemental Table. Metabolite concentrations in IOWBs (G. triflora (cv. Ihatovo)) cultured with sugar free (SF) or with % gentiobiose containing MS medium (% G). Metabolites AVR (nmol mg - DW) SD Ratio F-test T-test SF % G SF % G G/SF Alanine Arginine Asparagine Aspartate Citrulline Cysteine E-4 GABA Glu-Cys Glutamate Glutamine Glycine Histidine Homocysteine Isoleucine Leucine Lysine Methionine Ornithine Phenylalanine Proline SAM Serine Spermidine Spermine Threonine Tryptophan Tyrosine Valine ACC GSH E-5 GSSG OG PGA PG Aconitate Ascorbate E-6 DHA Citrate DHAP Fumarate G6P Malate NAD NADH NADP NADPH PEP Shikimate Succinate Data presented are calculated from four independent experiments. DW, dry weight; GABA, γ-amino butyrate; Glu-Cys, γ-glutamyl cysteine; SAM, S-adenosyl methionine; ACC, -Aminocyclopropane-- carboxylic acid; OG, -oxoglutarate; 3PGA, 3-phosphoglycerate; 6PG, 6-phosphogluconate; DHAP, dihydroxyacetone phosphate; G6P, Glc 6-phosphate; PEP, phosphoenolpyruvate.
12 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc Supplementary Table 3. List of primers used in this study Forward (5 3 ) Reverse (5 3 ) Cloning INV ATGCCATCCAAAAATTCAAAATCT ATAGGATCTAATGTCGGCGGAGTT HXK ATGGGGAAGGTAGCTGTGACGGCGG AGACTCCTCTAAACCAATGTACTG PGM ATGACATTCTGTACC TGTTATCACTGTTGGCTTCTC UGP ATGGAAACTCCTGCAA TATGTCTTGAGGACTGCTAAC SPS ATGGCGGGAAATGATTGGATAAACA GAGAACTTGTAGCTTCTCTAACGATG SUS ATGGCGCCTAAGTTAGAGAAAATA ATGCTCATCATCCACAGCAAG SiR ATGGCGACGTCGTATGGAGCAGCAGC CTTCTCAGCAAAACGGGAAGAGTGAG OASL ATGGCACAGGAAAAGATTGGTATT GGGATCAAAGGTCATGTTCTCTGC OASL ATGGCATCATCTTCTTCTTCTTC CAATCAGGGCACTGGTACCATCTT OASL3 ATGGCTGCTGCTGCTTTAGCAAATCT ATCAACTGACACAGGCTGCATATTC ECS ATGGCACTTATGTCACAAGCGGGATC GTATAGTAGCTCTTCGAAAACAGGT GSHS ATGGGTTCTGCATATTCTTCTTCTGA CCAACCGTAACAGCTGCATTTGCAC CBL ATGGCGTCTTCCTTGTTCCGTCAA AGCCGGTGCAGTTCTTAGTGCGTAAT CGS ATGGCCGTCTCCAGCTGCGCTAGGGT ATGGTCTCCAAAGCCTGCAGGATA MeS ATGGCATCACACATCGTTGGATACCC TTTGGCACTTGAGAGCTGTGTGCGG MeS ATGGCATCCCACATCGTTGGATATC TTTAGCACTGGCAAGCTGTGTGCGG SAMS ATGGACACTTTTCTATTCACCTCA AGCTTTGGGCTTGAGGATCTTGACA SAMS ATGGATACCTTCTTGTTCACTTCTG TGGCTTGTCGCTTATGAGGGGCTTG SAMDC ATGTCGGAAATGCCGGTCTCTGCGA CTCCTTTTCTTCATCTTCATCCTCCT ACO ATGGCATTCCCAATAGTAAACATGGAGAAG TCAAACGGGGGCAATGGGTCCCAAATTAATG SAHH ATGTCTCTCTCCGTCGAGAAGACCA ATACCTATAGGCCAAAGGCTTGTAT SAHH ATGTCTTTGCTCGTGGACAAAACCA ATACCTATAGTGAAGAGGCTTATAA DHAR ATGTCGACCGCCAAGTTAACATCA ACCACCCATCACCTTCGGCCGCCA DHAR ATGGCTCCGCCCGTTGAAGTAGCT TGCCTCAACCTTTGGCTTCCATCC GR ATGTCGGGTTTGGGAAGGAAGATG AAGGTTTGTTTTCGGTTTAACCGC GR ATGGCGACGTCTCTTGCTACGCCG GACCCCAGCAGTGGCTTTGATCTC AtGRP5 ATGGCTTCCAAGTCACTCTTTCTTGT ATGATGTCCACCACCGAAACCACCT Real-time PCR INV GTGGGTCTCGGGTTGAGATA AATCCAGCCCCATAAAATCC HXK TGTGATATTGGGAACGGGAACTA ATCTTGCCACTTGGGAATCG PGM TTGGGTGGTGATGGTCGAT ACCATTGCCTGCGGAAATT UGP TGAGTTGGGACCCGAGTTTAAG CACGATACTGGGAATGGATTTGA SPS CCTTTGGAGAATGTCGACCTTT CTCATCGATATTATCCCGGTTACC SUS CTGGAGTTTATGGCTTCTGGAAAT GAACATCTCAAGGTAACGCCTTGT SiR CCACTTTGTCCTTTGGCAATC CCCTCTTAAGCACATCCGGTAT OASTL AAAACCTGGCCCGCATAAA CAAAACACCAGGAATGAATCCA OASTL TTTCCTGGCCATCAGATTTGT AAGGTAGCCAACCCAATGTCA OASTL3 TCTTCCCTAGTCACTCCATTTGC ATTCGAACTGTGGGTCCAGAA ECS AACCCAACGGTTATCTCAGCAT TCCGATCTCCATCTGTGTCAAG GSHS GCTTCCCTTCCTTCGAATCTG CTGCCTTCTCGGTCTCCATTT CBL TTGGAAGCGTAAAGTCCCTCAT GGTATGCTCGCGTGTGACA CGS GGGCCTAAAAAGTCATCCTGAA CACCACCAAAACCAGTCATCTG MeS GCAATCTTATGGTTCTCGTTGTGT GCTTGGGTCGACTCACATCA MeS GGGTCAACCCGGATTGTG CAGAGCCGGCTTGACTTCTC SAMS TTCGGGATACCTGCAGAGAGA TCAGCATCCAGGCCAACA SAMS TGCCTCTGAGCCATGTCCTT TCTTGCGGACTTCGGTCAA SAMDC AATTATACACGTGGGAGCTTCATCT GAAGCTACGATGCGGAAACG ACO GAAGCTGTTGAGGGTGAAGTCA TCAGGGATTTCAGAGATGTTGGA SAHH TCGACATGCTCGGGTTAGAGA GGTTTGTGGCTTGATCGTGAT SAHH GAATTCAAAACGTCTGGGACAAT ACGATCTGGAACTCAGCATTATCA DHAR GGGAAGCCGGCGAGTAA CGCCGACTTCGCCATTAAT DHAR AGATCCCAATGATGGCTCAGA GATGCTCATCCAATGCTTTCAA GR GCCTGTTGCCTTGATGGAA TCGGCTGCTCACCAAAAAC GR CCTTCGAAGCCCAAAAAGATT CCAGCAAACTCAACAGCGATAT UBQ GGAAGCAGTTGGAAGATGGC GGCCAAAGTTCGTCCGTCCTC Vector construction for protein expression CCGCTCGAGAAAAGAATGCCATCCAAAAATT INV CAAAATCTC TGCTCTAGAAGATAGGATCTAATGTCGGCGGA GTT
13 Supplemental Data. Takahashi et al. Plant Cell (4).5/tpc Supplemental references Nakajima, M., Yamashita, T., Takahashi, M., Nakano, Y., Takeda, T. (). Identification, cloning, and characterization of β-glucosidase from Ustilago esculenta. Appl. Microbiol. Biotechnol. 93: Mangeon, A., Magioli, C., Menezes-Salgueiro, A.D., Cardeal, V., de Oliveira, C., Galvao, V.C., gis, R., Engler, G., and Sachetto-tins, G. (9). AtGRP5, a vacuole-located glycine-rich protein involved in cell elongation. Planta 3:
Amino Acids. Amino Acids. Fundamentals. While their name implies that amino acids are compounds that contain an NH. 3 and CO NH 3
 Fundamentals While their name implies that amino acids are compounds that contain an 2 group and a 2 group, these groups are actually present as 3 and 2 respectively. They are classified as α, β, γ, etc..
Fundamentals While their name implies that amino acids are compounds that contain an 2 group and a 2 group, these groups are actually present as 3 and 2 respectively. They are classified as α, β, γ, etc..
1. Describe the relationship of dietary protein and the health of major body systems.
 Food Explorations Lab I: The Building Blocks STUDENT LAB INVESTIGATIONS Name: Lab Overview In this investigation, you will be constructing animal and plant proteins using beads to represent the amino acids.
Food Explorations Lab I: The Building Blocks STUDENT LAB INVESTIGATIONS Name: Lab Overview In this investigation, you will be constructing animal and plant proteins using beads to represent the amino acids.
LAB#23: Biochemical Evidence of Evolution Name: Period Date :
 LAB#23: Biochemical Evidence of Name: Period Date : Laboratory Experience #23 Bridge Worth 80 Lab Minutes If two organisms have similar portions of DNA (genes), these organisms will probably make similar
LAB#23: Biochemical Evidence of Name: Period Date : Laboratory Experience #23 Bridge Worth 80 Lab Minutes If two organisms have similar portions of DNA (genes), these organisms will probably make similar
Amino acids-incorporated nanoflowers with an
 Amino acids-incorporated nanoflowers with an intrinsic peroxidase-like activity Zhuo-Fu Wu 1,2,+, Zhi Wang 1,+, Ye Zhang 3, Ya-Li Ma 3, Cheng-Yan He 4, Heng Li 1, Lei Chen 1, Qi-Sheng Huo 3, Lei Wang 1,*
Amino acids-incorporated nanoflowers with an intrinsic peroxidase-like activity Zhuo-Fu Wu 1,2,+, Zhi Wang 1,+, Ye Zhang 3, Ya-Li Ma 3, Cheng-Yan He 4, Heng Li 1, Lei Chen 1, Qi-Sheng Huo 3, Lei Wang 1,*
Journal of Cell Science Supplementary information. Arl8b +/- Arl8b -/- Inset B. electron density. genotype
 J. Cell Sci. : doi:.4/jcs.59: Supplementary information E9. A Arl8b /- Arl8b -/- Arl8b Arl8b non-specific band Gapdh Tbp E7.5 HE Inset B D Control al am hf C E Arl8b -/- al am hf E8.5 F low middle high
J. Cell Sci. : doi:.4/jcs.59: Supplementary information E9. A Arl8b /- Arl8b -/- Arl8b Arl8b non-specific band Gapdh Tbp E7.5 HE Inset B D Control al am hf C E Arl8b -/- al am hf E8.5 F low middle high
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
9/6/2011. Amino Acids. C α. Nonpolar, aliphatic R groups
 Amino Acids Side chains (R groups) vary in: size shape charge hydrogen-bonding capacity hydrophobic character chemical reactivity C α Nonpolar, aliphatic R groups Glycine (Gly, G) Alanine (Ala, A) Valine
Amino Acids Side chains (R groups) vary in: size shape charge hydrogen-bonding capacity hydrophobic character chemical reactivity C α Nonpolar, aliphatic R groups Glycine (Gly, G) Alanine (Ala, A) Valine
Midterm 2 Results. Standard Deviation:
 Midterm 2 Results High: Low: Mean: Standard Deviation: 97.5% 16% 58% 16.3 Lecture 17 Amino Acid Metabolism Urea Cycle N and S assimilation Last cofactors: THF and SAM Dietary (Exogenous) Proteins Hydrolyzed
Midterm 2 Results High: Low: Mean: Standard Deviation: 97.5% 16% 58% 16.3 Lecture 17 Amino Acid Metabolism Urea Cycle N and S assimilation Last cofactors: THF and SAM Dietary (Exogenous) Proteins Hydrolyzed
Page 8/6: The cell. Where to start: Proteins (control a cell) (start/end products)
 Page 8/6: The cell Where to start: Proteins (control a cell) (start/end products) Page 11/10: Structural hierarchy Proteins Phenotype of organism 3 Dimensional structure Function by interaction THE PROTEIN
Page 8/6: The cell Where to start: Proteins (control a cell) (start/end products) Page 11/10: Structural hierarchy Proteins Phenotype of organism 3 Dimensional structure Function by interaction THE PROTEIN
Short polymer. Dehydration removes a water molecule, forming a new bond. Longer polymer (a) Dehydration reaction in the synthesis of a polymer
 HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
Quantitative analysis of hydrophilic metabolite using ion-paring chromatography with a high-speed triple quadrupole mass spectrometer
 Quantitative analysis of hydrophilic metabolite using ion-paring chromatography with a high-speed triple ASMS 2012 ThP11-251 Zanariah Hashim 1 ; Yudai Dempo1; Tairo Ogura 2 ; Ichiro Hirano 2 ; Junko Iida
Quantitative analysis of hydrophilic metabolite using ion-paring chromatography with a high-speed triple ASMS 2012 ThP11-251 Zanariah Hashim 1 ; Yudai Dempo1; Tairo Ogura 2 ; Ichiro Hirano 2 ; Junko Iida
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A
 Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
GL Science Inertsearch for LC Inertsil Applications - Acids. Data No. Column Data Title Solutes Eluent Detection Data No.
 GL Science Inertsearch for LC Inertsil Applications: Acids For complete Product Description, Chromatograms Price & Delivery in Australia & New Zealand contact info@winlab.com.au or call 61 (0)7 3205 1209
GL Science Inertsearch for LC Inertsil Applications: Acids For complete Product Description, Chromatograms Price & Delivery in Australia & New Zealand contact info@winlab.com.au or call 61 (0)7 3205 1209
AMINO ACID METABOLISM. Sri Widia A Jusman Dept. of Biochemistry & Molecular Biology FMUI
 AMINO ACID METABOLISM Sri Widia A Jusman Dept. of Biochemistry & Molecular Biology FMUI Amino acids derived from dietary protein absorbed from intestine through blood taken up by tissues used for biosynthesis
AMINO ACID METABOLISM Sri Widia A Jusman Dept. of Biochemistry & Molecular Biology FMUI Amino acids derived from dietary protein absorbed from intestine through blood taken up by tissues used for biosynthesis
Identification of free amino acids in several crude extracts of two legumes
 1 2 Identification of free amino acids in several crude extracts of two legumes using Thin Layer Chromatography 3 Authors 4 5 6 7 8 9 Taghread Hudaib Key words 10 11 12 13 14 15 16 17 18 19 20 Amino acids;
1 2 Identification of free amino acids in several crude extracts of two legumes using Thin Layer Chromatography 3 Authors 4 5 6 7 8 9 Taghread Hudaib Key words 10 11 12 13 14 15 16 17 18 19 20 Amino acids;
Integrative Metabolism: Significance
 Integrative Metabolism: Significance Energy Containing Nutrients Carbohydrates Fats Proteins Catabolism Energy Depleted End Products H 2 O NH 3 ADP + Pi NAD + NADP + FAD + Pi NADH+H + NADPH+H + FADH2 Cell
Integrative Metabolism: Significance Energy Containing Nutrients Carbohydrates Fats Proteins Catabolism Energy Depleted End Products H 2 O NH 3 ADP + Pi NAD + NADP + FAD + Pi NADH+H + NADPH+H + FADH2 Cell
AP Bio. Protiens Chapter 5 1
 Concept.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 0% of the dry mass of most cells Protein functions include structural support, storage, transport,
Concept.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 0% of the dry mass of most cells Protein functions include structural support, storage, transport,
Midterm 2. Low: 14 Mean: 61.3 High: 98. Standard Deviation: 17.7
 Midterm 2 Low: 14 Mean: 61.3 High: 98 Standard Deviation: 17.7 Lecture 17 Amino Acid Metabolism Review of Urea Cycle N and S assimilation Last cofactors: THF and SAM Synthesis of few amino acids Dietary
Midterm 2 Low: 14 Mean: 61.3 High: 98 Standard Deviation: 17.7 Lecture 17 Amino Acid Metabolism Review of Urea Cycle N and S assimilation Last cofactors: THF and SAM Synthesis of few amino acids Dietary
Properties of amino acids in proteins
 Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Metabolism of amino acids. Vladimíra Kvasnicová
 Metabolism of amino acids Vladimíra Kvasnicová Classification of proteinogenic AAs -metabolic point of view 1) biosynthesis in a human body nonessential (are synthesized) essential (must be present in
Metabolism of amino acids Vladimíra Kvasnicová Classification of proteinogenic AAs -metabolic point of view 1) biosynthesis in a human body nonessential (are synthesized) essential (must be present in
Four Classes of Biological Macromolecules. Biological Macromolecules. Lipids
 Biological Macromolecules Much larger than other par4cles found in cells Made up of smaller subunits Found in all cells Great diversity of func4ons Four Classes of Biological Macromolecules Lipids Polysaccharides
Biological Macromolecules Much larger than other par4cles found in cells Made up of smaller subunits Found in all cells Great diversity of func4ons Four Classes of Biological Macromolecules Lipids Polysaccharides
Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version]
![Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version] Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version]](/thumbs/84/90818314.jpg) Earth/matriX: SCIENCE TODAY Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version] By Charles William Johnson Earth/matriX Editions P.O.
Earth/matriX: SCIENCE TODAY Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version] By Charles William Johnson Earth/matriX Editions P.O.
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Moorpark College Chemistry 11 Fall Instructor: Professor Gopal. Examination # 5: Section Five May 7, Name: (print)
 Moorpark College Chemistry 11 Fall 2013 Instructor: Professor Gopal Examination # 5: Section Five May 7, 2013 Name: (print) Directions: Make sure your examination contains TEN total pages (including this
Moorpark College Chemistry 11 Fall 2013 Instructor: Professor Gopal Examination # 5: Section Five May 7, 2013 Name: (print) Directions: Make sure your examination contains TEN total pages (including this
Lipids: diverse group of hydrophobic molecules
 Lipids: diverse group of hydrophobic molecules Lipids only macromolecules that do not form polymers li3le or no affinity for water hydrophobic consist mostly of hydrocarbons nonpolar covalent bonds fats
Lipids: diverse group of hydrophobic molecules Lipids only macromolecules that do not form polymers li3le or no affinity for water hydrophobic consist mostly of hydrocarbons nonpolar covalent bonds fats
The Structure and Function of Macromolecules
 The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
For questions 1-4, match the carbohydrate with its size/functional group name:
 Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the free response
Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the free response
Cells. Variation and Function of Cells
 Cells Variation and Function of Cells Plasma Membrane= the skin of a cell, it protects and nourishes the cell while communicating with other cells at the same time. Lipid means fat and they are hydrophobic
Cells Variation and Function of Cells Plasma Membrane= the skin of a cell, it protects and nourishes the cell while communicating with other cells at the same time. Lipid means fat and they are hydrophobic
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Macromolecules Structure and Function
 Macromolecules Structure and Function Within cells, small organic molecules (monomers) are joined together to form larger molecules (polymers). Macromolecules are large molecules composed of thousands
Macromolecules Structure and Function Within cells, small organic molecules (monomers) are joined together to form larger molecules (polymers). Macromolecules are large molecules composed of thousands
(30 pts.) 16. (24 pts.) 17. (20 pts.) 18. (16 pts.) 19. (5 pts.) 20. (5 pts.) TOTAL (100 points)
 Moorpark College Chemistry 11 Spring 2009 Instructor: Professor Torres Examination # 5: Section Five April 30, 2009 ame: (print) ame: (sign) Directions: Make sure your examination contains TWELVE total
Moorpark College Chemistry 11 Spring 2009 Instructor: Professor Torres Examination # 5: Section Five April 30, 2009 ame: (print) ame: (sign) Directions: Make sure your examination contains TWELVE total
The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5
 Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Copyright 2008 Pearson Education, Inc., publishing as Pearson Benjamin Cummings
 Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
Biomolecules: amino acids
 Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Methionine (Met or M)
 Fig. 5-17 Nonpolar Fig. 5-17a Nonpolar Glycine (Gly or G) Alanine (Ala or A) Valine (Val or V) Leucine (Leu or L) Isoleucine (Ile or I) Methionine (Met or M) Phenylalanine (Phe or F) Polar Trypotphan (Trp
Fig. 5-17 Nonpolar Fig. 5-17a Nonpolar Glycine (Gly or G) Alanine (Ala or A) Valine (Val or V) Leucine (Leu or L) Isoleucine (Ile or I) Methionine (Met or M) Phenylalanine (Phe or F) Polar Trypotphan (Trp
Amino acid metabolism
 Amino acid metabolism The important reaction commonly employed in the breakdown of an amino acid is always the removal of its -amino group. The product ammonia is excreted after conversion to urea or other
Amino acid metabolism The important reaction commonly employed in the breakdown of an amino acid is always the removal of its -amino group. The product ammonia is excreted after conversion to urea or other
If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out.
 Sign In Forgot Password Register username username password password Sign In If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out. ChemWiki
Sign In Forgot Password Register username username password password Sign In If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out. ChemWiki
CH395G FINAL (3 rd ) EXAM Kitto/Hackert - Fall 2003
 CH395G FINAL (3 rd ) EXAM Kitto/Hackert - Fall 2003 1. A cell in an active, catabolic state has a. a high (ATP/ADP) and a high (NADH/NAD + ) ratio b. a high (ATP/ADP) and a low (NADH/NAD + ) ratio c. a
CH395G FINAL (3 rd ) EXAM Kitto/Hackert - Fall 2003 1. A cell in an active, catabolic state has a. a high (ATP/ADP) and a high (NADH/NAD + ) ratio b. a high (ATP/ADP) and a low (NADH/NAD + ) ratio c. a
Chemistry 121 Winter 17
 Chemistry 121 Winter 17 Introduction to Organic Chemistry and Biochemistry Instructor Dr. Upali Siriwardane (Ph.D. Ohio State) E-mail: upali@latech.edu Office: 311 Carson Taylor Hall ; Phone: 318-257-4941;
Chemistry 121 Winter 17 Introduction to Organic Chemistry and Biochemistry Instructor Dr. Upali Siriwardane (Ph.D. Ohio State) E-mail: upali@latech.edu Office: 311 Carson Taylor Hall ; Phone: 318-257-4941;
18 Amino Acid Oxidation and Production of Urea W. H. Freeman and Company
 18 Amino Acid Oxidation and Production of Urea 2013 W. H. Freeman and Company 1 Last Class of Biomolecules For Energy 1. Production of acetyl-coa. Glucose. To pyruvate via glycolysis. To acetyl-coa by
18 Amino Acid Oxidation and Production of Urea 2013 W. H. Freeman and Company 1 Last Class of Biomolecules For Energy 1. Production of acetyl-coa. Glucose. To pyruvate via glycolysis. To acetyl-coa by
(65 pts.) 27. (10 pts.) 28. (15 pts.) 29. (10 pts.) TOTAL (100 points) Moorpark College Chemistry 11 Spring Instructor: Professor Gopal
 Moorpark College Chemistry 11 Spring 2012 Instructor: Professor Gopal Examination # 5: Section Five May 1, 2012 Name: (print) GOOD LUCK! Directions: Make sure your examination contains TWELVE total pages
Moorpark College Chemistry 11 Spring 2012 Instructor: Professor Gopal Examination # 5: Section Five May 1, 2012 Name: (print) GOOD LUCK! Directions: Make sure your examination contains TWELVE total pages
Amino acid metabolism II
 Amino acid metabolism II Fates of amino acid carbon skeleton degradation to common intermediates pyruvate, intermediates of citric acid cycle, acetyl-coa Glucogenic AA precursors of glucose - degradation
Amino acid metabolism II Fates of amino acid carbon skeleton degradation to common intermediates pyruvate, intermediates of citric acid cycle, acetyl-coa Glucogenic AA precursors of glucose - degradation
Supporting information for. Time-dependent Responses of Earthworms to Soil. Contaminated with Low Levels of Lead as Detected using 1 H
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2017 Supporting information for Time-dependent Responses of Earthworms to Soil Contaminated with
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2017 Supporting information for Time-dependent Responses of Earthworms to Soil Contaminated with
1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids
 Amino acids 1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids 5-To understand amino acids synthesis Amino
Amino acids 1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids 5-To understand amino acids synthesis Amino
Amino Acid Analyzer AAA400
 Amino Acid Analyzer AAA400 Determination of amino acid of hydrolyzates (food and feed) Column: LG ANB OSTION 3.6x340 12μm Eluents: sodium-citrate buffers, 0.2 M NaOH Aspartic Acid, Threonine, Serine, Glutamic
Amino Acid Analyzer AAA400 Determination of amino acid of hydrolyzates (food and feed) Column: LG ANB OSTION 3.6x340 12μm Eluents: sodium-citrate buffers, 0.2 M NaOH Aspartic Acid, Threonine, Serine, Glutamic
Cells N5 Homework book
 1 Cells N5 Homework book 2 Homework 1 3 4 5 Homework2 Cell Ultrastructure and Membrane 1. Name and give the function of the numbered organelles in the cell below: A E B D C 2. Name 3 structures you might
1 Cells N5 Homework book 2 Homework 1 3 4 5 Homework2 Cell Ultrastructure and Membrane 1. Name and give the function of the numbered organelles in the cell below: A E B D C 2. Name 3 structures you might
number Done by Corrected by Doctor Dr.Diala
 number 32 Done by Mousa Salah Corrected by Bahaa Najjar Doctor Dr.Diala 1 P a g e In the last lecture we talked about the common processes between all amino acids which are: transamination, deamination,
number 32 Done by Mousa Salah Corrected by Bahaa Najjar Doctor Dr.Diala 1 P a g e In the last lecture we talked about the common processes between all amino acids which are: transamination, deamination,
1. (38 pts.) 2. (25 pts.) 3. (15 pts.) 4. (12 pts.) 5. (10 pts.) Bonus (12 pts.) TOTAL (100 points)
 Moorpark College Chemistry 11 Spring 2010 Instructor: Professor Torres Examination #5: Section Five May 4, 2010 ame: (print) ame: (sign) Directions: Make sure your examination contains TWELVE total pages
Moorpark College Chemistry 11 Spring 2010 Instructor: Professor Torres Examination #5: Section Five May 4, 2010 ame: (print) ame: (sign) Directions: Make sure your examination contains TWELVE total pages
SIMPLE BASIC METABOLISM
 SIMPLE BASIC METABOLISM When we eat food such as a tuna fish sandwich, the polysaccharides, lipids, and proteins are digested to smaller molecules that are absorbed into the cells of our body. As these
SIMPLE BASIC METABOLISM When we eat food such as a tuna fish sandwich, the polysaccharides, lipids, and proteins are digested to smaller molecules that are absorbed into the cells of our body. As these
Chemical Nature of the Amino Acids. Table of a-amino Acids Found in Proteins
 Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
Catabolism of Carbon skeletons of Amino acids. Amino acid metabolism
 Catabolism of Carbon skeletons of Amino acids Amino acid metabolism Carbon skeleton Carbon Skeleton a carbon skeleton is the internal structure of organic molecules. Carbon Arrangements The arrangement
Catabolism of Carbon skeletons of Amino acids Amino acid metabolism Carbon skeleton Carbon Skeleton a carbon skeleton is the internal structure of organic molecules. Carbon Arrangements The arrangement
Chapter 5: Structure and Function of Macromolecules AP Biology 2011
 Chapter 5: Structure and Function of Macromolecules AP Biology 2011 1 Macromolecules Fig. 5.1 Carbohydrates Lipids Proteins Nucleic Acids Polymer - large molecule consisting of many similar building blocks
Chapter 5: Structure and Function of Macromolecules AP Biology 2011 1 Macromolecules Fig. 5.1 Carbohydrates Lipids Proteins Nucleic Acids Polymer - large molecule consisting of many similar building blocks
Reactions and amino acids structure & properties
 Lecture 2: Reactions and amino acids structure & properties Dr. Sameh Sarray Hlaoui Common Functional Groups Common Biochemical Reactions AH + B A + BH Oxidation-Reduction A-H + B-OH + energy ª A-B + H
Lecture 2: Reactions and amino acids structure & properties Dr. Sameh Sarray Hlaoui Common Functional Groups Common Biochemical Reactions AH + B A + BH Oxidation-Reduction A-H + B-OH + energy ª A-B + H
Midterm 1 Last, First
 Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
Biochemistry 423 Final Examination NAME:
 Biochemistry 423 Final Examination NAME: 1 Circle the single BEST answer (3 points each) 1. At equilibrium the free energy of a reaction G A. depends only on the temperature B. is positive C. is 0 D. is
Biochemistry 423 Final Examination NAME: 1 Circle the single BEST answer (3 points each) 1. At equilibrium the free energy of a reaction G A. depends only on the temperature B. is positive C. is 0 D. is
Amino Acid Metabolism
 Amino Acid Metabolism Fate of Dietary Protein Dietary protein Stomach: l, pepsin Denatured and partially hydrolyzed protein (large polypeptides) small intestine: proteases Amino acids and dipeptides intestinal
Amino Acid Metabolism Fate of Dietary Protein Dietary protein Stomach: l, pepsin Denatured and partially hydrolyzed protein (large polypeptides) small intestine: proteases Amino acids and dipeptides intestinal
Fate of Dietary Protein
 Fate of Dietary Protein Dietary protein Stomach: l, pepsin Denatured and partially hydrolyzed protein (large polypeptides) small intestine: proteases Amino acids and dipeptides intestinal lining: proteases
Fate of Dietary Protein Dietary protein Stomach: l, pepsin Denatured and partially hydrolyzed protein (large polypeptides) small intestine: proteases Amino acids and dipeptides intestinal lining: proteases
E.coli Core Model: Metabolic Core
 1 E.coli Core Model: Metabolic Core 2 LEARNING OBJECTIVES Each student should be able to: Describe the glycolysis pathway in the core model. Describe the TCA cycle in the core model. Explain gluconeogenesis.
1 E.coli Core Model: Metabolic Core 2 LEARNING OBJECTIVES Each student should be able to: Describe the glycolysis pathway in the core model. Describe the TCA cycle in the core model. Explain gluconeogenesis.
Biology. Lectures winter term st year of Pharmacy study
 Biology Lectures winter term 2008 1 st year of Pharmacy study 3 rd Lecture Chemical composition of living matter chemical basis of life. Atoms, molecules, organic compounds carbohydrates, lipids, proteins,
Biology Lectures winter term 2008 1 st year of Pharmacy study 3 rd Lecture Chemical composition of living matter chemical basis of life. Atoms, molecules, organic compounds carbohydrates, lipids, proteins,
Molecular Biology. general transfer: occurs normally in cells. special transfer: occurs only in the laboratory in specific conditions.
 Chapter 9: Proteins Molecular Biology replication general transfer: occurs normally in cells transcription special transfer: occurs only in the laboratory in specific conditions translation unknown transfer:
Chapter 9: Proteins Molecular Biology replication general transfer: occurs normally in cells transcription special transfer: occurs only in the laboratory in specific conditions translation unknown transfer:
For questions 1-4, match the carbohydrate with its size/functional group name:
 Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 ANSWERS For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the
Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 ANSWERS For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the
Krebs cycle Energy Petr Tůma Eva Samcová
 Krebs cycle Energy - 215 Petr Tůma Eva Samcová Overview of Citric Acid Cycle Key Concepts The citric acid cycle (Krebs cycle) is a multistep catalytic process that converts acetyl groups derived from carbohydrates,
Krebs cycle Energy - 215 Petr Tůma Eva Samcová Overview of Citric Acid Cycle Key Concepts The citric acid cycle (Krebs cycle) is a multistep catalytic process that converts acetyl groups derived from carbohydrates,
Study of Amino Acids in DDGS
 Study of Amino Acids in DDGS Y. Zhang, J. V. Simpson and B. A. Wrenn National Corn-to-Ethanol Research Center Edwardsville, IL 62025 Hans Stein University of Illinois Urbana Champaign Gerald C. Shurson
Study of Amino Acids in DDGS Y. Zhang, J. V. Simpson and B. A. Wrenn National Corn-to-Ethanol Research Center Edwardsville, IL 62025 Hans Stein University of Illinois Urbana Champaign Gerald C. Shurson
AMINO ACIDS Description of Amino acids and their functions in plants
 AMINO ACIDS Description of Amino acids and their functions in plants Technical Report 1. Introduction... 3 Proteic amino acids... 5 Amino acids in plants... 6 2. Amino acids functions.... 6 Aliphatic...
AMINO ACIDS Description of Amino acids and their functions in plants Technical Report 1. Introduction... 3 Proteic amino acids... 5 Amino acids in plants... 6 2. Amino acids functions.... 6 Aliphatic...
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Classification of amino acids: -
 Page 1 of 8 P roteinogenic amino acids, also known as standard, normal or primary amino acids are 20 amino acids that are incorporated in proteins and that are coded in the standard genetic code (subunit
Page 1 of 8 P roteinogenic amino acids, also known as standard, normal or primary amino acids are 20 amino acids that are incorporated in proteins and that are coded in the standard genetic code (subunit
CHM333 LECTURE 6: 1/25/12 SPRING 2012 Professor Christine Hrycyna AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID:
 AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID: - The R group side chains on amino acids are VERY important. o Determine the properties of the amino acid itself o Determine
AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID: - The R group side chains on amino acids are VERY important. o Determine the properties of the amino acid itself o Determine
LC-MS Analysis of Amino Acids on a Novel Mixed-Mode HPLC Column
 Liquid Chromatography Mass Spectrometry SSI-LCMS-022 LC-MS Analysis of Amino Acids on a ovel Mixed-Mode PLC Column LCMS-8040 Background There are four established methods for analyzing amino acids: prelabeled,
Liquid Chromatography Mass Spectrometry SSI-LCMS-022 LC-MS Analysis of Amino Acids on a ovel Mixed-Mode PLC Column LCMS-8040 Background There are four established methods for analyzing amino acids: prelabeled,
Nitrogen Metabolism. Overview
 Nitrogen Metabolism Pratt and Cornely Chapter 18 Overview Nitrogen assimilation Amino acid biosynthesis Nonessential aa Essential aa Nucleotide biosynthesis Amino Acid Catabolism Urea Cycle Juicy Steak
Nitrogen Metabolism Pratt and Cornely Chapter 18 Overview Nitrogen assimilation Amino acid biosynthesis Nonessential aa Essential aa Nucleotide biosynthesis Amino Acid Catabolism Urea Cycle Juicy Steak
استاذ الكيمياءالحيوية
 قسم الكيمياء الحيوية د.دولت على سالمه استاذ الكيمياءالحيوية ٢٠١٥-٢٠١٤ الرمز الكودي : ٥١٢ المحاضرة األولى ١ Content : Definition of proteins Definition of amino acids Definition of peptide bond General
قسم الكيمياء الحيوية د.دولت على سالمه استاذ الكيمياءالحيوية ٢٠١٥-٢٠١٤ الرمز الكودي : ٥١٢ المحاضرة األولى ١ Content : Definition of proteins Definition of amino acids Definition of peptide bond General
Introduction to Protein Structure Collection
 Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Biomolecules Amino Acids & Protein Chemistry
 Biochemistry Department Date: 17/9/ 2017 Biomolecules Amino Acids & Protein Chemistry Prof.Dr./ FAYDA Elazazy Professor of Biochemistry and Molecular Biology Intended Learning Outcomes ILOs By the end
Biochemistry Department Date: 17/9/ 2017 Biomolecules Amino Acids & Protein Chemistry Prof.Dr./ FAYDA Elazazy Professor of Biochemistry and Molecular Biology Intended Learning Outcomes ILOs By the end
Introduction to Peptide Sequencing
 Introduction to Peptide equencing Quadrupole Ion Traps tructural Biophysics Course December 3, 2014 12/8/14 Introduction to Peptide equencing - athan Yates 1 Why are ion traps used to sequence peptides?
Introduction to Peptide equencing Quadrupole Ion Traps tructural Biophysics Course December 3, 2014 12/8/14 Introduction to Peptide equencing - athan Yates 1 Why are ion traps used to sequence peptides?
Welcome to Class 14! Class 14: Outline and Objectives. Overview of amino acid catabolism! Introductory Biochemistry!
 Welcome to Class 14 Introductory Biochemistry Class 14: Outline and Objectives Amino Acid Catabolism Fates of amino groups transamination urea cycle Fates of carbon skeletons important cofactors metabolic
Welcome to Class 14 Introductory Biochemistry Class 14: Outline and Objectives Amino Acid Catabolism Fates of amino groups transamination urea cycle Fates of carbon skeletons important cofactors metabolic
7.014 Problem Set 2 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions will be posted on the web.
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Name: Section : 7.014 Problem Set 2 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 Name: Section : 7.014 Problem Set 2 Answers to this problem set are to be turned in. Problem sets will not be accepted late. Solutions
Protein Investigator. Protein Investigator - 3
 Protein Investigator Objectives To learn more about the interactions that govern protein structure. To test hypotheses regarding protein structure and function. To design proteins with specific shapes.
Protein Investigator Objectives To learn more about the interactions that govern protein structure. To test hypotheses regarding protein structure and function. To design proteins with specific shapes.
Chapter 4: Information and Knowledge in the Protein Insulin
 Chapter 4: Information and Knowledge in the Protein Insulin This chapter will calculate the information and molecular knowledge in a real protein. The techniques discussed in this chapter to calculate
Chapter 4: Information and Knowledge in the Protein Insulin This chapter will calculate the information and molecular knowledge in a real protein. The techniques discussed in this chapter to calculate
Use of RapidFire/MS for Metabolomics. - from untargeted to targeted and Pathway driven analysis. RapidFire/MS. Moritz Wagner Agilent Technologies
 Use of RapidFire/MS for Metabolomics RapidFire/MS - from untargeted to targeted and Pathway driven analysis Moritz Wagner Agilent Technologies 1 Agilent RapidFire TM High-throughput Mass Spectrometry System
Use of RapidFire/MS for Metabolomics RapidFire/MS - from untargeted to targeted and Pathway driven analysis Moritz Wagner Agilent Technologies 1 Agilent RapidFire TM High-throughput Mass Spectrometry System
AMINO ACID METABOLISM
 AMINO ACID METABOLISM PHL-285 Biochemistry-2 Mahmoud N. Nagi, Ph.D. Professor of Biochemistry Overview of amino acid metabolism. Classification of amino acids. Biosynthesis of nonessential amino acids.
AMINO ACID METABOLISM PHL-285 Biochemistry-2 Mahmoud N. Nagi, Ph.D. Professor of Biochemistry Overview of amino acid metabolism. Classification of amino acids. Biosynthesis of nonessential amino acids.
Biochemistry 2 Recita0on Amino Acid Metabolism
 Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Regulation. 1. Short term control 8-1
 Regulation Several aspects of regulation have been alluded to or described in detail as we have progressed through the various sections of the course. These include: (a) compartmentation: This was not
Regulation Several aspects of regulation have been alluded to or described in detail as we have progressed through the various sections of the course. These include: (a) compartmentation: This was not
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Proteins are a major component of dissolved organic nitrogen (DON) leached from terrestrially aged Eucalyptus camaldulensis leaves
 Environ. Chem. 216, 13, 877 887 doi:1.171/en165_ac CSIRO 216 Supplementary material Proteins are a major component of dissolved organic nitrogen (DON) leached from terrestrially aged Eucalyptus camaldulensis
Environ. Chem. 216, 13, 877 887 doi:1.171/en165_ac CSIRO 216 Supplementary material Proteins are a major component of dissolved organic nitrogen (DON) leached from terrestrially aged Eucalyptus camaldulensis
Lecture 4. Grouping Amino Acid 7/1/10. Proteins. Amino Acids. Where Are Proteins Located. Nonpolar Amino Acids
 Proteins Lecture 4 Proteins - Composition of Proteins (Amino Acids) Chapter 21 ection 1-6! Proteins are compounds of high molar mass consisting almost entirely of amino acid chain(s)! Molar masses range
Proteins Lecture 4 Proteins - Composition of Proteins (Amino Acids) Chapter 21 ection 1-6! Proteins are compounds of high molar mass consisting almost entirely of amino acid chain(s)! Molar masses range
Chapter 13 Carbohydrate Metabolism
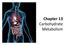 Chapter 13 Carbohydrate Metabolism Metabolism of Foods Food is broken down into carbohydrates, lipids, and proteins and sent through catabolic pathways to produce energy. Glycolysis glucose 2 P i 2 ADP
Chapter 13 Carbohydrate Metabolism Metabolism of Foods Food is broken down into carbohydrates, lipids, and proteins and sent through catabolic pathways to produce energy. Glycolysis glucose 2 P i 2 ADP
A Design Automation Framework for Computational Bioenergetics in Biological Networks
 This journal is The Royal Society of Chemistry 3 A Design Automation Framework for Computational Bioenergetics in Biological Networks Claudio Angione, a Jole Costanza, b Giovanni Carapezza, a Pietro Lió
This journal is The Royal Society of Chemistry 3 A Design Automation Framework for Computational Bioenergetics in Biological Networks Claudio Angione, a Jole Costanza, b Giovanni Carapezza, a Pietro Lió
Moorpark College Chemistry 11 Fall Instructor: Professor Gopal. Examination #5: Section Five December 7, Name: (print) Section:
 Moorpark College Chemistry 11 Fall 2011 Instructor: Professor Gopal Examination #5: Section Five December 7, 2011 Name: (print) Section: alkene < alkyne < amine < alcohol < ketone < aldehyde < amide
Moorpark College Chemistry 11 Fall 2011 Instructor: Professor Gopal Examination #5: Section Five December 7, 2011 Name: (print) Section: alkene < alkyne < amine < alcohol < ketone < aldehyde < amide
Amino Acid Oxidation and the Urea Cycle
 Amino Acid Oxidation and the Urea Cycle Amino Acids: Final class of biomolecules whose oxidation contributes significantly to the generation of energy Undergo oxidation in three metabolic circumstances
Amino Acid Oxidation and the Urea Cycle Amino Acids: Final class of biomolecules whose oxidation contributes significantly to the generation of energy Undergo oxidation in three metabolic circumstances
Amino Acid Metabolism
 Amino Acid Metabolism Last Week Most of the Animal Kingdom = Lazy - Most higher organisms in the animal kindom don t bother to make all of the amino acids. - Instead, we eat things that make the essential
Amino Acid Metabolism Last Week Most of the Animal Kingdom = Lazy - Most higher organisms in the animal kindom don t bother to make all of the amino acids. - Instead, we eat things that make the essential
PROTEINS. Amino acids are the building blocks of proteins. Acid L-form * * Lecture 6 Macromolecules #2 O = N -C -C-O.
 Proteins: Linear polymers of amino acids workhorses of the cell tools, machines & scaffolds Lecture 6 Macromolecules #2 PRTEINS 1 Enzymes catalysts that mediate reactions, increase reaction rate Structural
Proteins: Linear polymers of amino acids workhorses of the cell tools, machines & scaffolds Lecture 6 Macromolecules #2 PRTEINS 1 Enzymes catalysts that mediate reactions, increase reaction rate Structural
Proximate composition, amino acid and fatty acid composition of fish maws. Department of Biology, Lingnan Normal University, Zhanjiang, , China
 SUPPLEMENTARY MATERIAL Proximate composition, amino acid and fatty acid composition of fish maws Jing Wen a, Ling Zeng b *, Youhou Xu c, Yulin Sun a, Ziming Chen b and Sigang Fan d a Department of Biology,
SUPPLEMENTARY MATERIAL Proximate composition, amino acid and fatty acid composition of fish maws Jing Wen a, Ling Zeng b *, Youhou Xu c, Yulin Sun a, Ziming Chen b and Sigang Fan d a Department of Biology,
The human DEK oncogene: metabolic reprogramming of engineered epidermis
 The human DEK oncogene: metabolic reprogramming of engineered epidermis Markey Cancer Center, CESB, University of Kentucky 218 Metabolomics Symposium NMR Based Metabolomics Core CESB Co-director, UKY Marion
The human DEK oncogene: metabolic reprogramming of engineered epidermis Markey Cancer Center, CESB, University of Kentucky 218 Metabolomics Symposium NMR Based Metabolomics Core CESB Co-director, UKY Marion
Supporting Information
 Supporting Information Chaleckis et al. 10.1073/pnas.1603023113 SI Materials and Methods Human Subject Characteristics. Thirty healthy male and female volunteers participated in this study (Table S2).
Supporting Information Chaleckis et al. 10.1073/pnas.1603023113 SI Materials and Methods Human Subject Characteristics. Thirty healthy male and female volunteers participated in this study (Table S2).
Authors. Abstract. Introduction. Food
 Improved and Simplified Liquid Chromatography/Atmospheric Pressure Chemical Ionization Mass Spectrometry Method for the Analysis of Underivatized Free Ao Acids in Various Foods Application Food Authors
Improved and Simplified Liquid Chromatography/Atmospheric Pressure Chemical Ionization Mass Spectrometry Method for the Analysis of Underivatized Free Ao Acids in Various Foods Application Food Authors
Nitrogen Metabolism. Overview
 Nitrogen Metabolism Pratt and Cornely Chapter 18 Overview Nitrogen assimilation Amino acid biosynthesis Nonessential aa Essential aa Nucleotide biosynthesis Amino Acid Catabolism Urea Cycle Juicy Steak
Nitrogen Metabolism Pratt and Cornely Chapter 18 Overview Nitrogen assimilation Amino acid biosynthesis Nonessential aa Essential aa Nucleotide biosynthesis Amino Acid Catabolism Urea Cycle Juicy Steak
Chapter 3: Amino Acids and Peptides
 Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
9/16/15. Properties of Water. Benefits of Water. More properties of water
 Properties of Water Solid/Liquid Density Water is densest at 4⁰C Ice floats Allows life under the ice Hydrogen bond Ice Hydrogen bonds are stable Liquid water Hydrogen bonds break and re-form Benefits
Properties of Water Solid/Liquid Density Water is densest at 4⁰C Ice floats Allows life under the ice Hydrogen bond Ice Hydrogen bonds are stable Liquid water Hydrogen bonds break and re-form Benefits
Amino acid Catabolism
 Enzymatic digestion of dietary proteins in gastrointestinal-tract. Amino acid Catabolism Amino acids: 1. There are 20 different amino acid, they are monomeric constituents of proteins 2. They act as precursors
Enzymatic digestion of dietary proteins in gastrointestinal-tract. Amino acid Catabolism Amino acids: 1. There are 20 different amino acid, they are monomeric constituents of proteins 2. They act as precursors
Metabolism of amino acids I. Josef Fontana
 Metabolism of amino acids I Josef Fontana EC Overview of the lecture Introduction to protein and amino acids metabolism Metabolic pathways of amino acids Transamination Conversion glutamate - glutamine
Metabolism of amino acids I Josef Fontana EC Overview of the lecture Introduction to protein and amino acids metabolism Metabolic pathways of amino acids Transamination Conversion glutamate - glutamine
