Sterol regulatory element binding protein-1c (SREBP-1c) (1),
|
|
|
- Elinor Garrison
- 6 years ago
- Views:
Transcription
1 Unsaturated fatty acids inhibit transcription of the sterol regulatory element-binding protein-1c (SREBP-1c) gene by antagonizing liganddependent activation of the LXR Jiafu Ou*, Hua Tu, Bei Shan, Alvin Luk, Russell A. DeBose-Boyd*, Yuriy Bashmakov*, Joseph L. Goldstein*, and Michael S. Brown* *Department of Molecular Genetics, University of Texas Southwestern Medical Center, Dallas, TX ; and Tularik, Inc., Two Corporate Drive, South San Francisco, CA Contributed by Joseph L. Goldstein, March 20, 2001 Sterol regulatory element-binding protein-1c (SREBP-1c) enhances transcription of genes encoding enzymes of unsaturated fatty acid biosynthesis in liver. SREBP-1c mrna is known to increase when cells are treated with agonists of liver X receptor (LXR), a nuclear hormone receptor, and to decrease when cells are treated with unsaturated fatty acids, the end products of SREBP-1c action. Here we show that unsaturated fatty acids lower SREBP-1c mrna levels in part by antagonizing the actions of LXR. In cultured rat hepatoma cells, arachidonic acid and other fatty acids competitively inhibited activation of the endogenous SREBP-1c gene by an LXR ligand. Arachidonate also blocked the activation of a synthetic LXR-dependent promoter in transfected human embryonic kidney- 293 cells. In vitro, arachidonate and other unsaturated fatty acids competitively blocked activation of LXR, as reflected by a fluorescence polarization assay that measures ligand-dependent binding of LXR to a peptide derived from a coactivator. These data offer a potential mechanism that partially explains the long-known ability of dietary unsaturated fatty acids to decrease the synthesis and secretion of fatty acids and triglycerides in livers of humans and other animals. fatty acid synthesis arachidonic acid nuclear receptors Sterol regulatory element binding protein-1c (SREBP-1c) (1), also known as ADD-1 (2), is a membrane-bound transcription factor that functions at the interface between sterol and fatty acid metabolism. SREBP-1c enhances fatty acid synthesis by activating the genes encoding acetyl CoA carboxylase, fatty acid synthase (FAS), stearoyl CoA desaturase-1 (SCD-1), and other enzymes (3). The SREBP-1c gene is activated by insulin, which helps to explain the classic ability of insulin to enhance the conversion of glucose to fatty acids (4 6). In contrast, mrna levels for SREBP-1c are decreased when cells are exposed to unsaturated fatty acids, the end products of SREBP-1c action (7 9). Recent experiments have revealed that SREBP-1c mrna levels are also controlled by oxysterols. In rodent liver and hepatoma cells, transcription of the SREBP-1c gene is stimulated by oxysterols that bind to the and forms of the liver X receptor (LXR and ), two closely related members of the nuclear hormone receptor superfamily (10, 11). LXR activators include two naturally occurring sterols, 24(S),25-epoxycholesterol and 22(R)-hydroxycholesterol (12). LXR is also activated by T , a synthetic nonsteroidal compound (10, 13). The levels of SREBP-1c mrna declined dramatically when cultured rat hepatoma cells were treated with inhibitors of 3-hydroxy-3- methylglutaryl coenzyme reductase, which block the synthesis of endogenous LXR ligands (11). This inhibition was reversed when the cells were incubated with either T or mevalonate, the product of the reductase reaction. These data indicate that basal transcription of the SREBP-1c gene requires an endogenous sterol that activates LXR. The LXRs enhance transcription of the SREBP-1c gene by binding to a consensus recognition sequence in the enhancer region (10). SREBP-1c belongs to a three-member family that includes SREBP-1a and SREBP-2. SREBP-1a and -1c are produced from a common gene that uses alternative promoters to produce different first exons that are spliced into a common second exon (14). The encoded proteins differ at their extreme NH 2 termini, and this sequence difference affects their abilities to stimulate transcription of target genes. SREBP-1a is a strong activator of cholesterol and fatty acid biosynthesis, whereas SREBP-1c is selective for the fatty acid pathway (3, 15, 16). SREBP-2 is encoded by a separate gene, and it is relatively selective for cholesterol biosynthesis. Nonhepatic cells in tissue culture produce much more mrna for SREBP-1a than for -1c (17). In contrast, the liver, which synthesizes lipids for export as well as for membrane maintenance, produces up to 10-fold more SREBP-1c than -1a (17). All three SREBPs are synthesized as membrane-bound precursors of 1,150 aa in length (14). The NH 2 -terminal domain of 480 aa is a basic helix loop helix-leucine-zipper transcription factor. This domain is followed by a membrane attachment domain of 80 aa and a COOH-terminal regulatory domain of 590 aa. The SREBP precursors are initially bound to membranes of the endoplasmic reticulum. In sterol-depleted cells, they travel to the Golgi apparatus, where the NH 2 -terminal domains are released by two-step proteolysis, allowing them to enter the nucleus and activate transcription (14). In view of the opposing effects of LXR agonists and unsaturated fatty acids on SREBP-1c mrna levels, the current experiments were designed to test the hypothesis that unsaturated fatty acids act as direct antagonists of LXR. Consistent with this hypothesis, we show that the inhibitory effects of unsaturated fatty acids are reversed by LXR agonists in cultured rat hepatoma cells. We also present data to show that unsaturated fatty acids prevent agonists from enhancing the binding of LXR to a peptide derived from a coactivator that mediates transcriptional activation. Together, these findings provide evidence that unsaturated fatty acids reduce SREBP-1c mrna by antagonizing the activation of LXR by endogenous ligands. Abbreviations: ER, estrogen receptor; FAS, fatty acid synthase; GST, glutathione S- transferase; HEK-293 cells, human embryonic kidney-293 cells; LXR, liver X receptor; RXR, retinoid X receptor; SCD-1, stearoyl-coa desaturase-1; SREBP, sterol regulatory elementbinding protein. jgolds@mednet.swmed.edu or mbrown1@mednet.swmed.edu. The publication costs of this article were defrayed in part by page charge payment. This article must therefore be hereby marked advertisement in accordance with 18 U.S.C solely to indicate this fact. BIOCHEMISTRY cgi doi pnas PNAS May 22, 2001 vol. 98 no
2 Materials and Methods Materials. We obtained fatty acids from Sigma-Aldrich and Nu Chek Prep, Elysian, MN; defatted BSA, 9-cis retinoic acid, chenodeoxycholic acid, and 17 -estradiol from Sigma-Aldrich; N-acetyl-leucinal-leucinal-norleucinal from Calbiochem; and [ - 32 P]CTP (800 Ci mmol), redivue [ - 32 P]dCTP (3,000 Ci mmol), and [ - 32 P]UTP (800 Ci mmol) from Amersham Pharmacia. Delipidated FCS, sodium mevalonate, and sodium compactin were prepared as described (7, 18, 19). 24(S),25- Epoxycholesterol was synthesized by Julio Medina at Tularik. Preparation of Albumin-Bound Fatty Acids. A 5 or 10 mm stock solution of each fatty acid was prepared by diluting the free fatty acid in ethanol and precipitating it with the addition of NaOH (final concentration of 0.25 M) (7). Excess ethanol was evaporated under nitrogen gas, and the precipitated sodium salt was reconstituted with 0.9% (wt vol) NaCl and stirred at room temperature for 10 min with defatted BSA [final concentration at 10% (wt vol) in 0.15 M NaCl]. Each solution was adjusted to ph 7.4 with NaOH and stored in multiple aliquots at 20 C protected from light in tubes evacuated under nitrogen gas. Culture and Fractionation of FTO-2B Cells. The FTO-2B rat hepatoma cell line (obtained from R. E. Fournier, Fred Hutchinson Cancer Research Center, Seattle, WA) (20) was maintained in monolayer culture at 37 C in an 8% CO 2 incubator. On day 0, cells were set up at a density of cells per 100-mm dish in medium A (1:1 mixture of Ham s F-12 medium and DMEM containing 100 units ml penicillin and 100 g ml streptomycin sulfate) supplemented with 5% (vol vol) FCS. On day 3, cells were washed with PBS and refed with medium B (medium A supplemented with 5% delipidated FCS) containing the appropriate additions. After incubation for 5hat37 C, cells received 25 g ml N-acetyl-leucinal-leucinal-norleucinal and were further incubated for 1 h. Cells were harvested and fractionated into nuclear extract and membrane fractions as described (21). SDS PAGE and Immunoblot Analysis. Immunoblot analysis of endogenous rat SREBP-1 and -2 was carried out with 5 g ml of rabbit polyclonal antibodies against the NH 2 -terminal domains of either SREBP-1 (IgG-295) or SREBP-2 (IgG-302) (5). Bound antibodies were visualized with peroxidase-conjugated, affinitypurified donkey anti-rabbit IgG (Jackson ImmunoResearch), with the use of the SuperSignal CL-HRP substrate according to the manufacturer s instructions. SDS PAGE gels (8%) were calibrated with prestained molecular weight markers (Bio-Rad). Blots were exposed to Kodak X-Omat film at room temperature. Northern Blot Hybridization of mrna. Total RNA was isolated from two pooled dishes of FTO-2B cells with the use of an RNA Stat-60 kit (Tel-Test, Friendswood, TX). Cells were cultured as described above, except that N-acetyl-leucinal-leucinalnorleucinal was not added to the medium. Aliquots of total RNA (20 g) from each sample were denatured with RNA sample loading buffer containing 50 g ml ethidium bromide, subjected to electrophoresis in a 1% agarose formaldehyde gel, and transferred to Hybond N membranes. The cdna probes (22) were labeled with [ - 32 P]dCTP with the use of a Rediprimer II Random Labeling Kit (Amersham Pharmacia). The membranes were hybridized with the indicated 32 P-labeled probe ( cpm ml)for2hat65 C with the use of Rapid-Hyb buffer (Amersham Pharmacia), washed twice with 0.1% (wt vol) SDS 2 SSC at room temperature for 10 min, followed by 0.1% (wt vol) SDS 0.1 SSC at 65 C for 15 min, and exposed at 80 C to Kodak X-Omat film with BioMax intensifying screens for the indicated time (1 SSC 0.15 M sodium chloride M sodium citrate, ph 7). RNase Protection Assay. Aliquots of total RNA (20 g) were hybridized with [ - 32 P]CTP-labeled rat SREBP-1, SREBP-2, and -actin crna probes at 68 C for 10 min with the use of a HybSpeed RPA Kit (Ambion, Austin, TX) as described (5, 11). The mixture was digested with RNase A T1, and protected fragments corresponding to rat SREBP-1a (257 bp), -1c (160 bp), and -2 (110 bp) were separated on 8 M urea 5% polyacrylamide gels. Gels were dried and subjected to autoradiography with reflection film and BioMax intensifying screens at 80 C. Nuclear Run-On Assay. FTO-2B cells were set up at a density of cells per 150-mm dish in medium A supplemented with 5% FCS. After 60 h, the cells were washed with PBS and refed with medium B with or without 100 M sodium arachidonate and 1 M T After incubation for 6 h, the cells from four dishes were pooled and lysed in 0.5% (vol vol) Nonidet P-40 lysis buffer (23). Nuclei were stored until use at 80 C in buffer containing 75 mm Hepes (ph 7.5), 40% (vol vol) glycerol, 60 mm KCl, 5 mm MgCl 2, 0.1 mm sodium EDTA, 0.5 mm spermine, 1 mm spermidine, and 0.5 mm DTT. Nuclear run-on assays were performed as described by labeling nuclei with [ - 32 P]UTP (24). Extracted RNA ( cpm) was hybridized at 42 C for 72 h to nitrocellulose membranes containing linearized cdnas for rat SREBP-1 (3.4 kb), rat FAS (7.2 kb), and rat phosphoenolpyruvate carboxykinase (2.26 kb). Membranes were washed overnight in 0.1% SDS 0.1 SSC at 65 C. Fluorescence Polarization Assay. We used a previously described fluorescence polarization assay to measure agonist binding to LXR and other nuclear receptors (13). The assay measures the reduction in the tumbling rate that occurs when a specific rhodamine-labeled peptide (8 aa) containing the coactivator signature motif LXXLL (25) interacts with a ligand-bound nuclear receptor. The ligand-binding domains of nuclear receptors LXR, FXR, retinoid X receptor (RXR ), and estrogen receptor (ER ) were produced in Escherichia coli as glutathione S-transferase (GST) fusion proteins and purified as described (26). A rhodamine-labeled LXXLL-containing peptide (10 nm) with the amino acid sequence ILRKLLQE was incubated with 500 nm GST-LXR, GST-FXR, GST-RXR,or GST-ER in 50 l of buffer (10 mm Hepes 150 mm NaCl 2 mm MgCl 2 5 mm DTT at ph 7.9) in 384-well black polypropylene plates as described (13). BSA-conjugated fatty acids were added to the reaction mix just before one of the following agonists was added: 300 nm 24(S),25-epoxycholesterol, 20 M chenodeoxycholic acid, 100 nm 9-cis retinoic acid, or 100 nm 17- estradiol. After overnight incubation at room temperature, fluorescence polarization (mp) was measured with a CRI Symmetry Fluorescent Plate Reader (CRI, Woburn, MA). Gal4-Ligand Binding Domain Transient Transfection Assay. Human embryonic kidney-293 (HEK-293) cells were transiently transfected in 24-well plates with the use of Effectene Transfection Reagent (Qiagen, Chatsworth, CA). Each well received ng of a luciferase reporter construct containing four copies of the yeast transcription factor Gal4 response element and a TATA element derived from the thymidine kinase promoter together with 40 ng of an expression vector containing the Gal4 DNA binding domain fused to the ligand-binding domain of LXR (amino acids ), FXR (amino acids ), or RXR (amino acids 1 462) (13). An expression plasmid containing the -galactosidase gene driven by the cytomegalovirus promoter (4 ng well) was cotransfected for normalization. Results To measure the effects of fatty acids and LXR agonists on SREBP-1c mrna levels, we studied FTO-2B cells, a line of cultured rat hepatoma cells that resembles the liver in producing cgi doi pnas Ou et al.
3 Fig. 1. SREBP mrna expression in FTO-2B cells treated with different fatty acids in the absence or presence of LXR agonist T , as measured by RNase protection. On day 0, FTO-2B cells were set up in medium A containing 5% FCS as described in Materials and Methods. On day 3, cells were switched to medium B containing 100 M of the indicated fatty acid (added in a final concentration of 0.1% BSA) in the presence or absence of the indicated concentration of LXR agonist T (added in a final concentration of 0.1% DMSO). All dishes of cells received 0.1% BSA and 0.1% DMSO. After incubation for 6hat37 C, the cells from three dishes were harvested and pooled for RNA isolation. Aliquots of pooled total RNA (20 g) were hybridized for 10 min at 68 C to the indicated 32 P-labeled crna probe. RNaseprotected fragments were separated by gel electrophoresis and exposed to film at 80 C for 6 h. relatively high levels of the SREBP-1c transcript (SREBP-1c:1a ratio of 3:1 when grown in 5% delipidated FCS). The cells were incubated for 6 h in medium containing delipidated FCS, which is nearly devoid of fatty acids ( 0.5 M) and cholesterol ( 0.5 g ml). We supplemented the medium with varying concentrations of fatty acids and then measured the amounts of the three SREBP transcripts with the use of an RNase protection assay (11). In the absence of fatty acids, the amount of SREBP-1c mrna was high (Fig. 1, lane l), and it was reduced in the presence of a variety of fatty acids added at 100 M concentrations (Fig. 1, lanes 2 6). Active fatty acids included stearate (18 carbons, 0 double bonds) as well as monounsaturated and polyunsaturated fatty acids of 18, 20, or 22 carbons. The effectiveness of stearate was unexpected on the basis of prior studies with cultured kidney-derived cells from humans (HEK- 293 cells) (7) and monkeys (CV-1 cells) (27). The effect of stearate in the FTO-2B cells was reproducible in multiple experiments, and it may be attributable to the high levels of SCD-1 activity in these cells, which allows them to convert stearate to oleate. When the fatty acids were added to the FTO-2B cells together with the LXR ligand T , their ability to suppress SREBP-1c mrna was blocked (Fig. 1). This block was evident at 0.05 M T (Fig. 1, lanes 8 12) and complete at 1 M (Fig. 1, lanes 14 18). All of these effects were relatively specific for SREBP-1c. The SREBP-1a transcript was only slightly affected by fatty acids and T (Fig. 1), and the SREBP-2 transcript was not affected at all (see Fig. 3B). Fig. 2 shows that the LXR ligand reversed the fatty acid suppression in rapid fashion. In this experiment, cells were preincubated with arachidonate (20:4) for 6 h, during which time the SREBP-1c mrna declined (lane 2). The mrna rose markedly within1haftertheaddition of T , even though the arachidonate was maintained in the culture medium (lane 4). The increase persisted at 3 and 6 h (lanes 6 and 8). Fig. 3 A and B show the effects of various concentrations of arachidonate on SREBP-1c mrna levels as measured by RNase protection (A) and on the levels of mrna for two target genes, FAS and SCD-1, as measured by Northern blotting (B). In the absence of an LXR ligand, arachidonate suppressed the SREBP-1c mrna maximally at 10 M, and a slightly higher Fig. 2. T rapidly reverses the suppressive effect of arachidonate on SREBP-1c mrna. On day 0, FTO-2B cells were set up in medium A containing 5% FCS. On day 3, the cells were switched to medium B and preincubated for 6 h in the absence or presence of 100 M sodium arachidonate in a final concentration of 0.1% BSA. Two groups of cells were then harvested (lanes 1 and 2), and the remaining cells (lanes 3 8) were treated with either 1 M T in 0.1% DMSO or 0.1% DMSO alone for the indicated time. Total RNA from three dishes was pooled and isolated. Aliquots of total RNA (20 g) were hybridized for 10 min at 68 C to the indicated 32 P-labeled crna probe. RNase-protected fragments were separated by gel electrophoresis and exposed to film at 80 C for 6 h. concentration (30 M) gave maximal suppression of the FAS and SCD-1 mrnas. T at concentrations of 0.1 and 10 M overcame the inhibitory effect of arachidonate on the SREBP-1c mrna and the mrnas for the two target genes. Arachidonate had minor effects on the SREBP-1a mrna (A), and it had no effect on the SREBP-2 mrna (B). As controls, we studied the mrna for -actin (A) and cyclophilin (B), neither of which was affected by arachidonate or T Fig. 3C shows that arachidonate behaved as a competitive inhibitor of T in inducing SREBP-1c mrnas. We preincubated the cells with compactin, an inhibitor of 3-hydroxy- 3-methylglutaryl CoA reductase, and a low concentration of mevalonate, which permits synthesis of nonsterol isoprenes but not sterols. This treatment deprives cells of an endogenous sterol ligand for LXR, and therefore it lowers SREBP-1c mrna levels (11). In the absence of arachidonate, T increased SREBP-1c mrna maximally at 1 M. In the presence of 100 M arachidonate, the T dose response curve was shifted 10-fold to the right, but the maximal level of stimulation was unchanged. Previous studies in HEK-293 cells (which produce SREBP-1a predominantly) indicated that unsaturated fatty acids decrease nuclear SREBP-1 protein levels by two mechanisms: mrna suppression and partial inhibition of proteolytic cleavage (7). To determine the effects of T on these responses, we measured the amounts of the membrane-bound precursor and the nuclear forms of SREBP-1 in FTO-2B cells by immunoblotting with the best available antibody, which does not distinguish between the SREBP-1a and -1c isoforms (Fig. 4A). When added for 6 h in the absence of T , arachidonate markedly decreased the amount of nuclear SREBP-1 (Fig. 4A Upper, lane 2). The amount of the membrane-bound precursor form of SREBP-1 did not change. T increased both the precursor and nuclear forms of SREBP-1. In the presence of T , arachidonate decreased the nuclear form of SREBP-1c only partially (Fig. 4A, lane 4). As expected, in the same experiment arachidonate decreased the SREBP-1c mrna in the absence, but not the presence, of T (Fig. 4B). The partial reduction of nuclear SREBP-1 protein by arachidonate in the presence of T may represent the arachidonate-mediated inhibition of SREBP processing. Arachidonate had minor effects on BIOCHEMISTRY Ou et al. PNAS May 22, 2001 vol. 98 no
4 Fig. 4. Immunoblot analysis (A) and RNase protection assay (B) of SREBPs in FTO-2B cells incubated with arachidonate and LXR agonist T On day 0, cells were set up in medium A supplemented with 5% FCS. On day 3, cells were switched to medium B supplemented with 100 M sodium arachidonate and 1 M T as indicated. The medium in all cultures contained 0.1% BSA and 0.1% DMSO. (A) After a 5-h incubation at 37 C, cells received a direct addition of N-acetyl-leucinal-leucinal-norleucinal at 25 g ml and were harvested 1 h later. Membrane and nuclear extract fractions were prepared and subjected to SDS PAGE (68 g protein per lane) and immunoblot analysis as described in Materials and Methods. Filters were exposed to film at room temperature for 5 s. P and N denote precursor and nuclear cleaved forms of SREBPs, respectively. (B) After incubation at 37 C for 6 h, cells were harvested, and aliquots of total RNA from pooled dishes (20 g) were hybridized for 10 min at 68 C to 32 P-labeled crna probes for rat SREBP-1, SREBP-2, and -actin as described in Materials and Methods. RNase protected fragments were separated by gel electrophoresis and exposed to film for 6hat 80 C. Fig. 3. Suppression of mrnas encoding SREBP-1c and two target genes by varying concentrations of arachidonate in the absence and presence of T On day 0, FTO-2B cells were set up in medium A containing 5% FCS. (A and B) On day 3, cells were switched to medium B containing the indicated concentration of sodium arachidonate and T All dishes of cells received 0.1% BSA and 0.1% DMSO. After incubation for 6 h, the total RNA from two dishes was pooled and isolated. (A) Aliquots of total RNA (20 g) were hybridized for 10 min at 68 C to the indicated 32 P-labeled crna probe. RNase-protected fragments were separated by gel electrophoresis and exposed to film at 80 C for 8 h (SREBP-1a and -1c) or6h( -actin). (B) Aliquots of total RNA (20 g) were subjected to electrophoresis and Northern blot hybridization with the indicated 32 P-labeled probe, and filters were exposed to film at 80 C for 3 h (FAS), 4 h (SCD-1), or 8 h (SREBP-2 and cyclophilin). (C) On day 2, cells were switched to medium B containing 50 M compactin and 50 M sodium mevalonate. On day 3, cells were refed with the same medium containing the indicated concentrations of T and arachidonate. All dishes of cells received 0.1% BSA and 0.1% DMSO. After incubation for 6 h, total RNA from duplicate dishes was separately isolated, aliquots (20 g) were analyzed as described in A, and the bands for SREBP-1c were quantified with a Fuji Bio-Imaging analyzer and normalized to the -actin control. Values for each of the duplicate incubations are shown. the amount of nuclear SREBP-2 in the absence and presence of T (Fig. 4A Lower). To demonstrate directly that arachidonate and T affect SREBP-1c gene transcription, we incubated FTO-2B cells with these reagents for 6 h, isolated nuclei, and pulse-labeled them with [ - 32 P]UTP. The data showed that arachidonate reduced SREBP-1 transcription by 68%, and this reduction was reversed by T (Fig. 5). Arachidonate had a similar inhibitory effect on transcription of the FAS gene, which is a target of SREBP-1. There was no effect on the phosphoenolpyruvate carboxykinase gene, which served as a control (Fig. 5). To test directly the hypothesis that arachidonate blocks ligandmediated activation of LXR, we performed a series of in vitro assays. Previous studies showed that activation of nuclear hormone receptors increases the ability of the receptors to bind to coactivators that contain the consensus sequence leucine-x-xleucine-leucine (LXXLL), where X is any amino acid (25). This Fig. 5. Nuclear run-on assays of nascent 32 P-labeled RNA transcripts isolated from the nuclei of FTO-2B cells treated with arachidonate in the absence or presence of T Cells were set up for experiments as described in Materials and Methods. After incubation for 6hinmedium B with or without 100 M sodium arachidonate (A) in the absence or presence of 1 M T (T) as indicated, the cells from four 150-mm dishes were pooled, lysed, and subjected to nuclear run-on assay as described in Materials and Methods. After hybridization with the indicated cdna probe, the washed filters were exposed to Kodak X-Omat film with a Bio-Max (Kodak) intensifying screen at 80 C for 24 h. Relative levels of the transcripts for each gene (shown at right) were quantified with a Fuji Bio-Imaging analyzer. A blank value corresponding to the vector band was subtracted from all values cgi doi pnas Ou et al.
5 Fig. 6. Fatty acids block activation of LXR, but not other nuclear receptors, as measured in a fluorescence polarization assay. The indicated BSAconjugated fatty acid was evaluated for its ability to inhibit the binding of an LXXLL-containing peptide to LXR, RXR, FXR, or ER in the presence of the corresponding agonist: (A) 3 M 24(S),25-epoxycholesterol for LXR ; (B) 100 nm 9-cis retinoic acid for RXR ; (C) 20 M chenodeoxycholic acid for FXR; or (D) 100 nm 17- estradiol for ER. Data are presented as the increase in fluorescence polarization (mp) normalized to the BSA control. binding can be studied in vitro with the use of a fluorescence polarization assay that measures the binding of a fluorescently labeled LXXLL peptide to the nuclear receptor (13). Binding decreases the tumbling rate of the fluorophore, and thus it leads to enhanced fluorescence polarization. Fig. 6A shows an assay in which the binding of the LXXLL peptide to LXR was stimulated by the natural ligand 24(S),25-epoxycholesterol. The addition of three polyunsaturated fatty acids (18:2, 18:3, and 20:4) decreased binding of the peptide, as manifested by a decrease in fluorescence polarization (mp change). A noticeable effect was observed at fatty acid concentrations of 1 10 M. Oleate (18:1) was less effective, and stearate (18:0) had no effect. The specificity of this response was tested by showing that the fatty acids did not affect binding of the LXXLL peptide to three other nuclear hormone receptors: RXR (stimulated by 9-cis retinoic acid) (Fig. 6B), FXR (stimulated by chenodeoxycholic acid) (Fig. 6C), and the ER (stimulated by 17- estradiol) (Fig. 6D). To determine whether arachidonate acts as a competitive antagonist of the activating ligand, we measured fluorescence polarization as a function of the concentration of 24(S),25- epoxycholesterol at varying concentrations of arachidonate (Fig. 7A). The data showed that arachidonate had its greatest inhibiting effect at low concentrations of 24(S),25-epoxycholesterol, but high concentrations of the activator were able to overcome the arachidonate effect. An Eadie Hofstee plot of these data (Fig. 7B) showed that all lines converged at a common intercept on the vertical axis, indicating that arachidonate was indeed acting as a competitive antagonist of 24(S),25-epoxycholesterol. We calculated that the effect of arachidonate was half-maximal at 1.5 M. A way to test the effects of regulators on nuclear hormone receptors in vivo is through the use of nuclear receptor-gal4 fusion proteins (13). Accordingly, we transfected HEK-293 cells with a cdna encoding a chimeric protein that contains the DNA binding domain of Gal4 and the ligand-binding and transcription-activating domains of LXR. We also transfected a cdna encoding luciferase driven by a promoter containing four copies Fig. 7. Arachidonate competitively inhibits LXR activation by 24(S),25- epoxycholesterol as measured by fluorescence polarization. (A) The indicated concentration of sodium arachidonate (20:4) was evaluated for its ability to inhibit the binding of an LXXLL-containing peptide to LXR in the presence of the indicated concentration of 24(S),25-epoxycholesterol. Data are presented as the increase in fluorescence polarization (mp) normalized to the BSA control. (B) Eadie Hofstee plot of the data in A. A blank value for fluorescence polarization in the absence of 24,25-epoxycholesterol was subtracted from all measured values. of a Gal4 binding sequence. When the transfected cells were incubated with the LXR ligand 24(S),25-epoxycholesterol, luciferase activity rose by 4- to 5-fold (Fig. 8A). Addition of each of the unsaturated fatty acids lowered luciferase activity to the basal level. Stearate (18:0) had no effect, most likely because HEK-293 cells have low levels of desaturase activity. As controls, we studied Gal4 fusion proteins containing the ligand-binding domain of FXR (Fig. 8B) and RXR (Fig. 8C). When incubated with their respective ligands, both of these proteins stimulated the luciferase reporter gene. The fatty acids had no significant effect on these activations. Fig. 8. Fatty acids selectively antagonize transcription-stimulating activity of LXR in HEK-293 cells. Cells were transiently cotransfected with a Gal4-driven luciferase reporter construct; an expression vector containing the Gal4 DNA binding domain fused to the ligand-binding domain of LXR (A), FXR (B), or RXR (C) as indicated; and a control plasmid containing the -galactosidase gene driven by the cytomegalovirus promoter. Six to eight hours after transfection, the natural ligands (5 M 24(S),25-epoxycholesterol for LXR,25 M chenodeoxycholic acid for FXR, and 50 nm 9-cis retinoic acid for RXR ) and the indicated fatty acids were added to the medium in complex with BSA as described in the legend to Fig. 1. After incubation for 20 h, the cells were harvested for assay. Data are presented as relative luciferase activity normalized to -galactosidase activity. BIOCHEMISTRY Ou et al. PNAS May 22, 2001 vol. 98 no
6 Discussion The current results establish that unsaturated fatty acids can act as competitive antagonists of LXR in cultured rat hepatoma and human HEK-293 cells and in cell-free assays that reflect LXR activation. This antagonism appears to explain, at least partially, the ability of unsaturated fatty acids to lower the levels of mrna for SREBP-1c (7 9), the transcription of which has been shown to depend on an endogenous LXR ligand (11). The lowered SREBP-1c, in turn, leads to a fall in mrnas for enzymes responsible for synthesizing unsaturated fatty acids, thus completing a feedback loop. A noteworthy difference between the in vivo and in vitro assays of LXR antagonism relates to the effect of stearate (18:0), which acted as an LXR antagonist in intact FTO-2B cells (Fig. 1), but not in the in vitro assays (Fig. 6) or in the HEK-293 cells (Fig. 8). This difference may reflect a higher level of desaturase activity in the FTO-2B cells, which convert stearate to oleate. This hypothesis has yet to be tested. The in vitro assays were performed with fatty acids bound to albumin. Under these conditions, arachidonate had a halfmaximal effect at 1.5 M, which is lower than the 10 M concentration that was required when added to intact cells (Fig. 3). The levels of free fatty acids within cells are generally thought to be low, and they are largely bound to intracellular binding proteins. It is possible that fatty acids are more effective when bound to these physiological carriers. It is also possible that the true antagonists are not free fatty acids, but rather are their CoA or carnitine thioesters. We attempted to inhibit LXR activation in vitro with the use of fatty acyl CoAs, obtaining negative results in the fluorescence polarization assay. In addition to transcriptional inhibition mediated by LXR antagonism, unsaturated fatty acids also appear to lower SREBP-1 mrna by accelerating its degradation. This effect of fatty acids has been demonstrated directly by studies of mrna decay rates in the presence of -amanitin in freshly isolated rat hepatocytes (9). It is likely that such an effect also occurred in the current studies, inasmuch as arachidonate inhibited SREBP-1 transcription by about 3-fold, as determined by the nuclear run-on assay (Fig. 5) at a time when the mrna level declined by 6-fold, as determined by Northern blotting (data not shown). Arachidonate also inhibits the proteolytic processing of SREBP-1a and -1c in HEK-293 cells (7). Thus, arachidonate appears to lower nuclear SREBP-1c levels by at least three mechanisms in animal cells. When fed to experimental animals, polyunsaturated acids of the n-3 and n-6 classes have long been known to inhibit fatty acid and triglyceride synthesis and to lower plasma triglyceride levels (28). Accumulating evidence, some of which is presented here, strongly suggests that these effects are mediated by a reduction in the levels of nuclear SREBP-1c in liver. In contrast, LXR agonists raise plasma triglycerides in experimental animals, apparently through induction of hepatic SREBP-1c (13). Agents that lower nuclear SREBP-1c levels, either by antagonizing LXR action in the liver or by some other means, should be effective in lowering plasma triglyceride levels in humans. We thank Jay Horton and David Mangelsdorf for their critical review of the manuscript and Voe Hannah for helpful suggestions in the initial phases of the study. Lisa Beatty and Christine Alvares provided invaluable help with cultured cells. This research was made possible by grants from the National Institutes of Health (HL20948) and the Perot Family Foundation. R.A.D.-B. is the recipient of a postdoctoral fellowship from the Jane Coffin Childs Memorial Fund for Medical Research. 1. Yokoyama, C., Wang, X., Briggs, M. R., Admon, A., Wu, J., Hua, X., Goldstein, J. L. & Brown, M. S. (1993) Cell 75, Kim, J. B. & Spiegelman, B. M. (1996) Genes Dev. 10, Shimano, H., Horton, J. D., Shimomura, I., Hammer, R. E., Brown, M. S. & Goldstein, J. L. (1997) J. Clin. Invest. 99, Foretz, M., Guichard, C., Ferre, P. & Foufelle, F. (1999) Proc. Natl. Acad. Sci. USA 96, Shimomura, I., Bashmakov, Y., Ikemoto, S., Horton, J. D., Brown, M. S. & Goldstein, J. L. (1999) Proc. Natl. Acad. Sci. USA 96, Shimomura, I., Hammer, R. E., Ikemoto, S., Brown, M. S. & Goldstein, J. L. (1999) Nature (London) 401, Hannah, V. C., Ou, J., Luong, A., Goldstein, J. L. & Brown, M. S. (2001) J. Biol. Chem. 276, Mater, M. K., Thelen, A. P., Pan, D. A. & Jump, D. B. (1999) J. Biol. Chem. 274, Xu, J., Teran-Garcia, M., Park, J. H. Y., Nakamura, M. T. & Clarke, S. D. (2001) J. Biol. Chem. 276, Repa, J. J., Liang, G., Ou, J., Bashmakov, Y., Lobaccaro, J.-M. A., Shimomura, I., Shan, B., Brown, M. S., Goldstein, J. L. & Mangelsdorf, D. J. (2000) Genes Dev. 14, DeBose-Boyd, R. A., Ou, J., Goldstein, J. L. & Brown, M. S. (2001) Proc. Natl. Acad. Sci. USA 98, Janowski, B. A., Grogan, M. J., Jones, S. A., Wisely, G. B., Kliewer, S. A., Corey, E. J. & Mangelsdorf, D. J. (1999) Proc. Natl. Acad. Sci. USA 96, Schultz, J. R., Tu, H., Luk, A., Repa, J. J., Medina, J. C., Li, L., Schwendner, S., Wang, S., Thoolen, M., Mangelsdorf, D. J., et al. (2000) Genes Dev. 14, Brown, M. S. & Goldstein, J. L. (1997) Cell 89, Pai, J., Guryev, O., Brown, M. S. & Goldstein, J. L. (1998) J. Biol. Chem. 273, Horton, J. D., Shimomura, I., Brown, M. S., Hammer, R. E., Goldstein, J. L. & Shimano, H. (1998) J. Clin. Invest. 101, Shimomura, I., Shimano, H., Horton, J. D., Goldstein, J. L. & Brown, M. S. (1997) J. Clin. Invest. 99, Brown, M. S., Faust, J. R., Goldstein, J. L., Kaneko, I. & Endo, A. (1978) J. Biol. Chem. 253, Goldstein, J. L., Basu, S. K. & Brown, M. S. (1983) Methods Enzymol. 98, Wynshaw-Boris, A., Lugo, T. G., Short, J. M., Fournier, R. E. K. & Hanson, R. W. (1984) J. Biol. Chem. 259, DeBose-Boyd, R. A., Brown, M. S., Li, W.-P., Nohturfft, A., Goldstein, J. L. & Espenshade, P. J. (1999) Cell 99, Shimano, H., Horton, J. D., Hammer, R. E., Shimomura, I., Brown, M. S. & Goldstein, J. L. (1996) J. Clin. Invest. 98, Sambrook, J. & Russell, D. W. (2001) Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Lab. Press, Plainview, NY). 24. Zhang, J., Ou, J., Bashmakov, Y., Horton, J. D., Brown, M. S. & Goldstein, J. L. (2001) Proc. Natl. Acad. Sci. USA 98, (First Published March 20, 2001, pnas ) 25. Heery, D. M., Kalkhoven, E., Hoare, S. & Parker, M. G. (1997) Nature (London) 387, Smith, D. B. & Johnson, K. S. (1988) Gene 67, Worgall, T. S., Sturley, S. L., Seo, T., Osborne, T. F. & Deckelbaum, R. J. (1998) J. Biol. Chem. 273, Jump, D. B. & Clarke, S. D. (1999) Annu. Rev. Nutr. 19, cgi doi pnas Ou et al.
Sterol regulatory element-binding proteins (SREBPs) are a
 Expression of sterol regulatory element-binding protein 1c (SREBP-1c) mrna in rat hepatoma cells requires endogenous LXR ligands Russell A. DeBose-Boyd*, Jiafu Ou*, Joseph L. Goldstein, and Michael S.
Expression of sterol regulatory element-binding protein 1c (SREBP-1c) mrna in rat hepatoma cells requires endogenous LXR ligands Russell A. DeBose-Boyd*, Jiafu Ou*, Joseph L. Goldstein, and Michael S.
LY (4 -allylcholestan-3 -ol) was identified for its ability
 The hypocholesterolemic agent LY295427 upregulates INSIG-1, identifying the INSIG-1 protein as a mediator of cholesterol homeostasis through SREBP Bethany A. Janowski* Department of Molecular Genetics,
The hypocholesterolemic agent LY295427 upregulates INSIG-1, identifying the INSIG-1 protein as a mediator of cholesterol homeostasis through SREBP Bethany A. Janowski* Department of Molecular Genetics,
Central role for liver X receptor in insulin-mediated activation of Srebp-1c transcription and stimulation of fatty acid synthesis in liver.
 University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange Nutrition Publications and Other Works Nutrition 8-3-2004 Central role for liver X receptor in insulin-mediated activation
University of Tennessee, Knoxville Trace: Tennessee Research and Creative Exchange Nutrition Publications and Other Works Nutrition 8-3-2004 Central role for liver X receptor in insulin-mediated activation
Critical review. SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver
 SPOTLIGHT Critical review SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver Jay D. Horton, 1,2 Joseph L. Goldstein, 1 and Michael S. Brown 1 1 Department of
SPOTLIGHT Critical review SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver Jay D. Horton, 1,2 Joseph L. Goldstein, 1 and Michael S. Brown 1 1 Department of
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
ab SREBP-2 Translocation Assay Kit (Cell-Based)
 ab133114 SREBP-2 Translocation Assay Kit (Cell-Based) Instructions for Use For analysis of translocation of SREBP-2 into nuclei. This product is for research use only and is not intended for diagnostic
ab133114 SREBP-2 Translocation Assay Kit (Cell-Based) Instructions for Use For analysis of translocation of SREBP-2 into nuclei. This product is for research use only and is not intended for diagnostic
Cholesterol feeding reduces nuclear forms of sterol regulatory element binding proteins in hamster liver
 Proc. Natl. Acad. Sci. USA Vol. 94, pp. 12354 12359, November 1997 Biochemistry Cholesterol feeding reduces nuclear forms of sterol regulatory element binding proteins in hamster liver (cholesterol biosynthesis
Proc. Natl. Acad. Sci. USA Vol. 94, pp. 12354 12359, November 1997 Biochemistry Cholesterol feeding reduces nuclear forms of sterol regulatory element binding proteins in hamster liver (cholesterol biosynthesis
Distinct roles of insulin and liver X receptor in the induction and cleavage of sterol regulatory elementbinding
 Distinct roles of insulin and liver X receptor in the induction and cleavage of sterol regulatory elementbinding protein-1c Bronwyn D. Hegarty*, Alexandre Bobard*, Isabelle Hainault, Pascal Ferré, Pascale
Distinct roles of insulin and liver X receptor in the induction and cleavage of sterol regulatory elementbinding protein-1c Bronwyn D. Hegarty*, Alexandre Bobard*, Isabelle Hainault, Pascal Ferré, Pascale
Blunted Feedback Suppression of SREBP Processing by Dietary Cholesterol in Transgenic Mice Expressing Sterol-Resistant SCAP(D443N)
 Blunted Feedback Suppression of SREBP Processing by Dietary Cholesterol in Transgenic Mice Expressing Sterol-Resistant SCAP(D443N) Bobby S. Korn,* Iichiro Shimomura,* Yuriy Bashmakov,* Robert E. Hammer,
Blunted Feedback Suppression of SREBP Processing by Dietary Cholesterol in Transgenic Mice Expressing Sterol-Resistant SCAP(D443N) Bobby S. Korn,* Iichiro Shimomura,* Yuriy Bashmakov,* Robert E. Hammer,
Human LDL Receptor / LDLR ELISA Pair Set
 Human LDL Receptor / LDLR ELISA Pair Set Catalog Number : SEK10231 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized in
Human LDL Receptor / LDLR ELISA Pair Set Catalog Number : SEK10231 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized in
Cleavage of Sterol Regulatory Element-binding Proteins (SREBPs) at Site-1 Requires Interaction with SREBP Cleavage-activating Protein
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 273, No. 10, Issue of March 6, pp. 5785 5793, 1998 1998 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Cleavage of Sterol
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 273, No. 10, Issue of March 6, pp. 5785 5793, 1998 1998 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Cleavage of Sterol
Protocol for Gene Transfection & Western Blotting
 The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
Fatty acid synthase (FAS) plays a central role in de novo
 Nutritional regulation of the fatty acid synthase promoter in vivo: Sterol regulatory element binding protein functions through an upstream region containing a sterol regulatory element Maria-Jesus Latasa,
Nutritional regulation of the fatty acid synthase promoter in vivo: Sterol regulatory element binding protein functions through an upstream region containing a sterol regulatory element Maria-Jesus Latasa,
Abstract. Introduction
 Activation of Cholesterol Synthesis in Preference to Fatty Acid Synthesis in Liver and Adipose Tissue of Transgenic Mice Overproducing Sterol Regulatory Element-binding Protein-2 Jay D. Horton,* Iichiro
Activation of Cholesterol Synthesis in Preference to Fatty Acid Synthesis in Liver and Adipose Tissue of Transgenic Mice Overproducing Sterol Regulatory Element-binding Protein-2 Jay D. Horton,* Iichiro
Intramembrane glycine mediates multimerization of Insig-2, a requirement for sterol regulation in Chinese hamster ovary cells
 Intramembrane glycine mediates multimerization of Insig-2, a requirement for sterol regulation in Chinese hamster ovary cells Peter C. W. Lee * and Russell A. DeBose-Boyd 1, Howard Hughes Medical Institute
Intramembrane glycine mediates multimerization of Insig-2, a requirement for sterol regulation in Chinese hamster ovary cells Peter C. W. Lee * and Russell A. DeBose-Boyd 1, Howard Hughes Medical Institute
Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes
 Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes The major site of acetoacetate and 3-hydorxybutyrate production is in the liver. 3-hydorxybutyrate is the
Companion to Biosynthesis of Ketones & Cholesterols, Regulation of Lipid Metabolism Lecture Notes The major site of acetoacetate and 3-hydorxybutyrate production is in the liver. 3-hydorxybutyrate is the
SREBP-1 integrates the actions of thyroid hormone, insulin, camp, and medium-chain fatty acids on ACC transcription in hepatocytes
 SREBP-1 integrates the actions of thyroid hormone, insulin, camp, and medium-chain fatty acids on ACC transcription in hepatocytes Yanqiao Zhang, Liya Yin, and F. Bradley Hillgartner 1 Department of Biochemistry
SREBP-1 integrates the actions of thyroid hormone, insulin, camp, and medium-chain fatty acids on ACC transcription in hepatocytes Yanqiao Zhang, Liya Yin, and F. Bradley Hillgartner 1 Department of Biochemistry
20S Proteasome Activity Assay Kit
 20S Proteasome Activity Assay Kit For 100 Assays Cat. No. APT280 FOR RESEARCH USE ONLY NOT FOR USE IN DIAGNOSTIC PROCEDURES USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23
20S Proteasome Activity Assay Kit For 100 Assays Cat. No. APT280 FOR RESEARCH USE ONLY NOT FOR USE IN DIAGNOSTIC PROCEDURES USA & Canada Phone: +1(800) 437-7500 Fax: +1 (951) 676-9209 Europe +44 (0) 23
Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer. Application Note
 Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer Application Note Odile Sismeiro, Jean-Yves Coppée, Christophe Antoniewski, and Hélène Thomassin
Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer Application Note Odile Sismeiro, Jean-Yves Coppée, Christophe Antoniewski, and Hélène Thomassin
2.5. AMPK activity
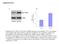 Supplement Fig. A 3 B phos-ampk 2.5 * Control AICAR AMPK AMPK activity (Absorbance at 45 nm) 2.5.5 Control AICAR Supplement Fig. Effects of AICAR on AMPK activation in macrophages. J774. macrophages were
Supplement Fig. A 3 B phos-ampk 2.5 * Control AICAR AMPK AMPK activity (Absorbance at 45 nm) 2.5.5 Control AICAR Supplement Fig. Effects of AICAR on AMPK activation in macrophages. J774. macrophages were
The Schedule and the Manual of Basic Techniques for Cell Culture
 The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
SUPPLEMENTARY INFORMATION
 Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
SREBF2 Transcription Factor Assay Kit ( Cat # KA1379 V.01 ) 1
 Background Lipid homeostasis in vertebrate cells is regulated by a family of transcription factors called sterol regulatory element binding proteins (SREBP s). SREBP s directly activate the expression
Background Lipid homeostasis in vertebrate cells is regulated by a family of transcription factors called sterol regulatory element binding proteins (SREBP s). SREBP s directly activate the expression
Influenza B Hemagglutinin / HA ELISA Pair Set
 Influenza B Hemagglutinin / HA ELISA Pair Set Catalog Number : SEK11053 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized
Influenza B Hemagglutinin / HA ELISA Pair Set Catalog Number : SEK11053 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized
Influenza A H7N9 (A/Anhui/1/2013) Hemagglutinin / HA ELISA Pair Set
 Influenza A H7N9 (A/Anhui/1/2013) Hemagglutinin / HA ELISA Pair Set Catalog Number : SEK40103 To achieve the best assay results, this manual must be read carefully before using this product and the assay
Influenza A H7N9 (A/Anhui/1/2013) Hemagglutinin / HA ELISA Pair Set Catalog Number : SEK40103 To achieve the best assay results, this manual must be read carefully before using this product and the assay
Suppression of fatty acid synthase promoter by polyunsaturated fatty acids
 Suppression of fatty acid synthase promoter by polyunsaturated fatty acids Yang Soo Moon, 1 Maria-Jesus Latasa, Michael J. Griffin, and Hei Sook Sul 2 Department of Nutritional Sciences, University of
Suppression of fatty acid synthase promoter by polyunsaturated fatty acids Yang Soo Moon, 1 Maria-Jesus Latasa, Michael J. Griffin, and Hei Sook Sul 2 Department of Nutritional Sciences, University of
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Luminescent platforms for monitoring changes in the solubility of amylin and huntingtin in living cells
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
MCB130 Midterm. GSI s Name:
 1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
1. Peroxisomes are small, membrane-enclosed organelles that function in the degradation of fatty acids and in the degradation of H 2 O 2. Peroxisomes are not part of the secretory pathway and peroxisomal
Islet viability assay and Glucose Stimulated Insulin Secretion assay RT-PCR and Western Blot
 Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
TFEB-mediated increase in peripheral lysosomes regulates. Store Operated Calcium Entry
 TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
Jay D. Horton, Iichiro Shimomura, Shinji Ikemoto **, Yuriy Bashmakov, and Robert E. Hammer
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 278, No. 38, Issue of September 19, pp. 36652 36660, 2003 2003 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Overexpression
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 278, No. 38, Issue of September 19, pp. 36652 36660, 2003 2003 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Overexpression
Total Phosphatidic Acid Assay Kit
 Product Manual Total Phosphatidic Acid Assay Kit Catalog Number MET- 5019 100 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Phosphatidic Acid (PA) is a critical precursor
Product Manual Total Phosphatidic Acid Assay Kit Catalog Number MET- 5019 100 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Phosphatidic Acid (PA) is a critical precursor
The SCAP/SREBP Pathway: A Mediator of Hepatic Steatosis
 Review Article Endocrinol Metab 2017;32:6-10 https://doi.org/10.3803/enm.2017.32.1.6 pissn 2093-596X eissn 2093-5978 The SCAP/SREBP Pathway: A Mediator of Hepatic Steatosis Young-Ah Moon Department of
Review Article Endocrinol Metab 2017;32:6-10 https://doi.org/10.3803/enm.2017.32.1.6 pissn 2093-596X eissn 2093-5978 The SCAP/SREBP Pathway: A Mediator of Hepatic Steatosis Young-Ah Moon Department of
Chapter 26 Biochemistry 5th edition. phospholipids. Sphingolipids. Cholesterol. db=books&itool=toolbar
 http://www.ncbi.nlm.nih.gov/sites/entrez? db=books&itool=toolbar 1 The surface of a soap bubble is a bilayer formed by detergent molecules 2 Chapter 26 Biochemistry 5th edition phospholipids Sphingolipids
http://www.ncbi.nlm.nih.gov/sites/entrez? db=books&itool=toolbar 1 The surface of a soap bubble is a bilayer formed by detergent molecules 2 Chapter 26 Biochemistry 5th edition phospholipids Sphingolipids
Dual regulation of mouse 5 - and 6 -desaturase gene expression by SREBP-1 and PPAR
 Dual regulation of mouse 5 - and 6 -desaturase gene expression by SREBP-1 and PPAR Takashi Matsuzaka, Hitoshi Shimano, 1, Naoya Yahagi,* Michiyo Amemiya-Kudo,* Tomohiro Yoshikawa,* Alyssa H. Hasty,* Yoshiaki
Dual regulation of mouse 5 - and 6 -desaturase gene expression by SREBP-1 and PPAR Takashi Matsuzaka, Hitoshi Shimano, 1, Naoya Yahagi,* Michiyo Amemiya-Kudo,* Tomohiro Yoshikawa,* Alyssa H. Hasty,* Yoshiaki
Amplification of the gene for SCAP, coupled with Insig-1 deficiency, confers sterol resistance in mutant Chinese hamster ovary cells
 Amplification of the gene for SCAP, coupled with Insig-1 deficiency, confers sterol resistance in mutant Chinese hamster ovary cells Peter C. W. Lee,* Pingsheng Liu, Wei-Ping Li, and Russell A. DeBose-Boyd
Amplification of the gene for SCAP, coupled with Insig-1 deficiency, confers sterol resistance in mutant Chinese hamster ovary cells Peter C. W. Lee,* Pingsheng Liu, Wei-Ping Li, and Russell A. DeBose-Boyd
Overexpression of SREBP-1a in Mouse Adipose Tissue Produces Adipocyte Hypertrophy, Increased Fatty Acid Secretion, and Fatty Liver
 JBC Papers in Press. Published on July 10, 2003 as Manuscript M306540200 1 Overexpression of SREBP-1a in Mouse Adipose Tissue Produces Adipocyte Hypertrophy, Increased Fatty Acid Secretion, and Fatty Liver
JBC Papers in Press. Published on July 10, 2003 as Manuscript M306540200 1 Overexpression of SREBP-1a in Mouse Adipose Tissue Produces Adipocyte Hypertrophy, Increased Fatty Acid Secretion, and Fatty Liver
Cholesterol determination using protein-templated fluorescent gold nanocluster probes
 Electronic Supplementary Information for Cholesterol determination using protein-templated fluorescent gold nanocluster probes Xi Chen and Gary A. Baker* Department of Chemistry, University of Missouri-Columbia,
Electronic Supplementary Information for Cholesterol determination using protein-templated fluorescent gold nanocluster probes Xi Chen and Gary A. Baker* Department of Chemistry, University of Missouri-Columbia,
Human Obestatin ELISA
 K-ASSAY Human Obestatin ELISA For the quantitative determination of obestatin in human serum and plasma Cat. No. KT-495 For Research Use Only. 1 Rev. 081309 K-ASSAY PRODUCT INFORMATION Human Obestatin
K-ASSAY Human Obestatin ELISA For the quantitative determination of obestatin in human serum and plasma Cat. No. KT-495 For Research Use Only. 1 Rev. 081309 K-ASSAY PRODUCT INFORMATION Human Obestatin
Kit for assay of thioredoxin
 FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
(A) PCR primers (arrows) designed to distinguish wild type (P1+P2), targeted (P1+P2) and excised (P1+P3)14-
 1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
Free Fatty Acid Assay Kit (Fluorometric)
 Product Manual Free Fatty Acid Assay Kit (Fluorometric) Catalog Number STA-619 100 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Triglycerides (TAG) are a type of lipid
Product Manual Free Fatty Acid Assay Kit (Fluorometric) Catalog Number STA-619 100 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Triglycerides (TAG) are a type of lipid
Insig-dependent Ubiquitination and Degradation of Mammalian 3-Hydroxy-3-methylglutaryl-CoA Reductase Stimulated by Sterols and Geranylgeraniol*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 278, No. 52, Issue of December 26, pp. 52479 52490, 2003 2003 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Insig-dependent
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 278, No. 52, Issue of December 26, pp. 52479 52490, 2003 2003 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Insig-dependent
ab SREBP-1 Transcription Factor Assay Kit
 ab133125 SREBP-1 Transcription Factor Assay Kit Instructions for Use For the detection of specific transcription factor DNA binding activity in nuclear extracts. This product is for research use only and
ab133125 SREBP-1 Transcription Factor Assay Kit Instructions for Use For the detection of specific transcription factor DNA binding activity in nuclear extracts. This product is for research use only and
RayBio Human PPAR-alpha Transcription Factor Activity Assay Kit
 RayBio Human PPAR-alpha Transcription Factor Activity Assay Kit Catalog #: TFEH-PPARa User Manual Jan 5, 2018 3607 Parkway Lane, Suite 200 Norcross, GA 30092 Tel: 1-888-494-8555 (Toll Free) or 770-729-2992,
RayBio Human PPAR-alpha Transcription Factor Activity Assay Kit Catalog #: TFEH-PPARa User Manual Jan 5, 2018 3607 Parkway Lane, Suite 200 Norcross, GA 30092 Tel: 1-888-494-8555 (Toll Free) or 770-729-2992,
Regulation of mouse sterol regulatory element-binding protein-1c gene (SREBP-1c) by oxysterol receptors, LXR and LXR
 Regulation of mouse sterol regulatory element-binding protein-1c gene (SREBP-1c) by oxysterol receptors, LXR and LXR Joyce J. Repa, 1,4 Guosheng Liang, 2,4 Jiafu Ou, 2 Yuriy Bashmakov, 2 Jean-Marc A. Lobaccaro,
Regulation of mouse sterol regulatory element-binding protein-1c gene (SREBP-1c) by oxysterol receptors, LXR and LXR Joyce J. Repa, 1,4 Guosheng Liang, 2,4 Jiafu Ou, 2 Yuriy Bashmakov, 2 Jean-Marc A. Lobaccaro,
JBC Papers in Press. Published on September 27, 2001 as Manuscript M
 JBC Papers in Press. Published on September 27, 2001 as Manuscript M108348200 THE HYPOCHOLESTEROLEMIC AGENT LY295427 REVERSES SUPPRESSION OF SREBP PROCESSING MEDIATED BY OXYSTEROLS* Bethany A. Janowski,
JBC Papers in Press. Published on September 27, 2001 as Manuscript M108348200 THE HYPOCHOLESTEROLEMIC AGENT LY295427 REVERSES SUPPRESSION OF SREBP PROCESSING MEDIATED BY OXYSTEROLS* Bethany A. Janowski,
INTERACTION DRUG BODY
 INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
INTERACTION DRUG BODY What the drug does to the body What the body does to the drug Receptors - intracellular receptors - membrane receptors - Channel receptors - G protein-coupled receptors - Tyrosine-kinase
human Total Cathepsin B Catalog Number: DY2176
 human Total Cathepsin B Catalog Number: DY2176 This DuoSet ELISA Development kit contains the basic components required for the development of sandwich ELISAs to measure natural and recombinant human Total
human Total Cathepsin B Catalog Number: DY2176 This DuoSet ELISA Development kit contains the basic components required for the development of sandwich ELISAs to measure natural and recombinant human Total
SREBP-2 Transcription Factor Assay Kit
 SREBP-2 Transcription Factor Assay Kit Item No. 10007819 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL
SREBP-2 Transcription Factor Assay Kit Item No. 10007819 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL
J. M. Hsu and S. T. Ding* Department of Animal Science, National Taiwan University, Taipei 106, Taiwan
 British Journal of Nutrition (2003), 90, 507 513 q The Authors 2003 DOI: 10.1079/BJN2003918 Effect of polyunsaturated fatty acids on the expression of transcription factor adipocyte determination and differentiation-dependent
British Journal of Nutrition (2003), 90, 507 513 q The Authors 2003 DOI: 10.1079/BJN2003918 Effect of polyunsaturated fatty acids on the expression of transcription factor adipocyte determination and differentiation-dependent
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna
 Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Regulating Hepatic Cellular Cholesterol
 Under circumstances of cholesterol deficiency, Sterol Regulatory Element Binding Proteins (SREBPs) via binding to DNA nuclear response elements set off genomic production of proteins and enzymes that induce
Under circumstances of cholesterol deficiency, Sterol Regulatory Element Binding Proteins (SREBPs) via binding to DNA nuclear response elements set off genomic production of proteins and enzymes that induce
SREBP-1 Transcription Factor Assay Kit
 SREBP-1 Transcription Factor Assay Kit Item No. 10010854 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL
SREBP-1 Transcription Factor Assay Kit Item No. 10010854 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL
Cholesterol and its transport. Alice Skoumalová
 Cholesterol and its transport Alice Skoumalová 27 carbons Cholesterol - structure Cholesterol importance A stabilizing component of cell membranes A precursor of bile salts A precursor of steroid hormones
Cholesterol and its transport Alice Skoumalová 27 carbons Cholesterol - structure Cholesterol importance A stabilizing component of cell membranes A precursor of bile salts A precursor of steroid hormones
Human Immunodeficiency Virus type 1 (HIV-1) gp120 / Glycoprotein 120 ELISA Pair Set
 Human Immunodeficiency Virus type 1 (HIV-1) gp120 / Glycoprotein 120 ELISA Pair Set Catalog Number : SEK11233 To achieve the best assay results, this manual must be read carefully before using this product
Human Immunodeficiency Virus type 1 (HIV-1) gp120 / Glycoprotein 120 ELISA Pair Set Catalog Number : SEK11233 To achieve the best assay results, this manual must be read carefully before using this product
Unit IV Problem 3 Biochemistry: Cholesterol Metabolism and Lipoproteins
 Unit IV Problem 3 Biochemistry: Cholesterol Metabolism and Lipoproteins - Cholesterol: It is a sterol which is found in all eukaryotic cells and contains an oxygen (as a hydroxyl group OH) on Carbon number
Unit IV Problem 3 Biochemistry: Cholesterol Metabolism and Lipoproteins - Cholesterol: It is a sterol which is found in all eukaryotic cells and contains an oxygen (as a hydroxyl group OH) on Carbon number
ANSC/NUTR 618 Lipids & Lipid Metabolism
 I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
TECHNICAL BULLETIN. R 2 GlcNAcβ1 4GlcNAcβ1 Asn
 GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
Product Manual. Omni-Array Sense Strand mrna Amplification Kit, 2 ng to 100 ng Version Catalog No.: Reactions
 Genetic Tools and Reagents Universal mrna amplification, sense strand amplification, antisense amplification, cdna synthesis, micro arrays, gene expression, human, mouse, rat, guinea pig, cloning Omni-Array
Genetic Tools and Reagents Universal mrna amplification, sense strand amplification, antisense amplification, cdna synthesis, micro arrays, gene expression, human, mouse, rat, guinea pig, cloning Omni-Array
HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation
 SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
Sterol-regulated ubiquitination and degradation of Insig-1 creates a convergent mechanism for feedback control of cholesterol synthesis and uptake
 Sterol-regulated ubiquitination and degradation of Insig-1 creates a convergent mechanism for feedback control of cholesterol synthesis and uptake Yi Gong, 1 Joon No Lee, 1 Peter C.W. Lee, 1 Joseph L.
Sterol-regulated ubiquitination and degradation of Insig-1 creates a convergent mechanism for feedback control of cholesterol synthesis and uptake Yi Gong, 1 Joon No Lee, 1 Peter C.W. Lee, 1 Joseph L.
Notch Signaling Pathway Notch CSL Reporter HEK293 Cell line Catalog #: 60652
 Notch Signaling Pathway Notch CSL Reporter HEK293 Cell line Catalog #: 60652 Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. Notch signaling
Notch Signaling Pathway Notch CSL Reporter HEK293 Cell line Catalog #: 60652 Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. Notch signaling
Validation & Assay Performance Summary
 Validation & Assay Performance Summary LanthaScreen IGF-1R GripTite Cells Cat. no. K1834 Modification Detected: Phosphorylation of Multiple Tyr Residues on IGF-1R LanthaScreen Cellular Assay Validation
Validation & Assay Performance Summary LanthaScreen IGF-1R GripTite Cells Cat. no. K1834 Modification Detected: Phosphorylation of Multiple Tyr Residues on IGF-1R LanthaScreen Cellular Assay Validation
A protocol for enhancement of the AAV-mediated expression of transgenes
 A protocol for enhancement of the AAV-mediated expression of transgenes Hiroaki Mizukami, Takeharu Kanazawa, Takashi Okada, and Keiya Ozawa Division of Genetic Therapeutics, Center for Molecular Medicine,
A protocol for enhancement of the AAV-mediated expression of transgenes Hiroaki Mizukami, Takeharu Kanazawa, Takashi Okada, and Keiya Ozawa Division of Genetic Therapeutics, Center for Molecular Medicine,
Western Immunoblotting Preparation of Samples:
 Western Immunoblotting Preparation of Samples: Total Protein Extraction from Culture Cells: Take off the medium Wash culture with 1 x PBS 1 ml hot Cell-lysis Solution into T75 flask Scrap out the cells
Western Immunoblotting Preparation of Samples: Total Protein Extraction from Culture Cells: Take off the medium Wash culture with 1 x PBS 1 ml hot Cell-lysis Solution into T75 flask Scrap out the cells
Influenza A H1N1 HA ELISA Pair Set
 Influenza A H1N1 HA ELISA Pair Set for H1N1 ( A/Puerto Rico/8/1934 ) HA Catalog Number : SEK11684 To achieve the best assay results, this manual must be read carefully before using this product and the
Influenza A H1N1 HA ELISA Pair Set for H1N1 ( A/Puerto Rico/8/1934 ) HA Catalog Number : SEK11684 To achieve the best assay results, this manual must be read carefully before using this product and the
SUPPLEMENTARY INFORMATION
 Glucose is a ligand for the oxysterol receptor LXR Supplementary Figure 1 Schematic representation of pathways influencing glucose fate in the liver. Glucose induces insulin secretion, suppressing hepatic
Glucose is a ligand for the oxysterol receptor LXR Supplementary Figure 1 Schematic representation of pathways influencing glucose fate in the liver. Glucose induces insulin secretion, suppressing hepatic
General Laboratory methods Plasma analysis: Gene Expression Analysis: Immunoblot analysis: Immunohistochemistry:
 General Laboratory methods Plasma analysis: Plasma insulin (Mercodia, Sweden), leptin (duoset, R&D Systems Europe, Abingdon, United Kingdom), IL-6, TNFα and adiponectin levels (Quantikine kits, R&D Systems
General Laboratory methods Plasma analysis: Plasma insulin (Mercodia, Sweden), leptin (duoset, R&D Systems Europe, Abingdon, United Kingdom), IL-6, TNFα and adiponectin levels (Quantikine kits, R&D Systems
Sensitization of the HIV-1-LTR upon Long Term Low Dose Oxidative Stress*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 271, No. 36, Issue of September 6, pp. 21798 21802, 1996 1996 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Sensitization
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 271, No. 36, Issue of September 6, pp. 21798 21802, 1996 1996 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Sensitization
Cholesterol metabolism. Function Biosynthesis Transport in the organism Hypercholesterolemia
 Cholesterol metabolism Function Biosynthesis Transport in the organism Hypercholesterolemia - component of all cell membranes - precursor of bile acids steroid hormones vitamin D Cholesterol Sources: dietary
Cholesterol metabolism Function Biosynthesis Transport in the organism Hypercholesterolemia - component of all cell membranes - precursor of bile acids steroid hormones vitamin D Cholesterol Sources: dietary
Caspase-3 Assay Cat. No. 8228, 100 tests. Introduction
 Introduction Caspase-3 Assay Cat. No. 8228, 100 tests Caspase-3 is a member of caspases that plays a key role in mediating apoptosis, or programmed cell death. Upon activation, it cleaves a variety of
Introduction Caspase-3 Assay Cat. No. 8228, 100 tests Caspase-3 is a member of caspases that plays a key role in mediating apoptosis, or programmed cell death. Upon activation, it cleaves a variety of
Supporting Information Table of content
 Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
Protein MultiColor Stable, Low Range
 Product Name: DynaMarker Protein MultiColor Stable, Low Range Code No: DM670L Lot No: ******* Size: 200 μl x 3 (DM670 x 3) (120 mini-gel lanes) Storage: 4 C Stability: 12 months at 4 C Storage Buffer:
Product Name: DynaMarker Protein MultiColor Stable, Low Range Code No: DM670L Lot No: ******* Size: 200 μl x 3 (DM670 x 3) (120 mini-gel lanes) Storage: 4 C Stability: 12 months at 4 C Storage Buffer:
Supplemental Table 1 Primer sequences (mouse) used for real-time qrt-pcr studies
 Supplemental Table 1 Primer sequences (mouse) used for real-time qrt-pcr studies Gene symbol Forward primer Reverse primer ACC1 5'-TGAGGAGGACCGCATTTATC 5'-GCATGGAATGGCAGTAAGGT ACLY 5'-GACACCATCTGTGATCTTG
Supplemental Table 1 Primer sequences (mouse) used for real-time qrt-pcr studies Gene symbol Forward primer Reverse primer ACC1 5'-TGAGGAGGACCGCATTTATC 5'-GCATGGAATGGCAGTAAGGT ACLY 5'-GACACCATCTGTGATCTTG
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
LIPID METABOLISM. Sri Widia A Jusman Department of Biochemistry & Molecular Biology FMUI
 LIPID METABOLISM Sri Widia A Jusman Department of Biochemistry & Molecular Biology FMUI Lipid metabolism is concerned mainly with fatty acids cholesterol Source of fatty acids from dietary fat de novo
LIPID METABOLISM Sri Widia A Jusman Department of Biochemistry & Molecular Biology FMUI Lipid metabolism is concerned mainly with fatty acids cholesterol Source of fatty acids from dietary fat de novo
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
Mouse Adiponectin Immunoassay Kit
 Antibody and Immunoassay Services Li Ka Shing Faculty of Medicine, The University of Hong Kong Mouse Adiponectin Immunoassay Kit Catalogue Number: 32010 For the quantitative determination of mouse adiponectin
Antibody and Immunoassay Services Li Ka Shing Faculty of Medicine, The University of Hong Kong Mouse Adiponectin Immunoassay Kit Catalogue Number: 32010 For the quantitative determination of mouse adiponectin
FATTY ACID SYNTHESIS
 FATTY ACID SYNTHESIS Malonyl- CoA inhibits Carni1ne Palmitoyl Transferase I. Malonyl- CoA is a precursor for fa=y acid synthesis. Malonyl- CoA is produced from acetyl- CoA by the enzyme Acetyl- CoA Carboxylase.
FATTY ACID SYNTHESIS Malonyl- CoA inhibits Carni1ne Palmitoyl Transferase I. Malonyl- CoA is a precursor for fa=y acid synthesis. Malonyl- CoA is produced from acetyl- CoA by the enzyme Acetyl- CoA Carboxylase.
THE GLUCOSE-FATTY ACID-KETONE BODY CYCLE Role of ketone bodies as respiratory substrates and metabolic signals
 Br. J. Anaesth. (1981), 53, 131 THE GLUCOSE-FATTY ACID-KETONE BODY CYCLE Role of ketone bodies as respiratory substrates and metabolic signals J. C. STANLEY In this paper, the glucose-fatty acid cycle
Br. J. Anaesth. (1981), 53, 131 THE GLUCOSE-FATTY ACID-KETONE BODY CYCLE Role of ketone bodies as respiratory substrates and metabolic signals J. C. STANLEY In this paper, the glucose-fatty acid cycle
Fatty acids synthesis
 Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
BIOLOGICAL MOLECULES REVIEW-UNIT 1 1. The factor being tested in an experiment is the A. data. B. variable. C. conclusion. D. observation. 2.
 BIOLOGICAL MOLECULES REVIEW-UNIT 1 1. The factor being tested in an experiment is the A. data. B. variable. C. conclusion. D. observation. 2. A possible explanation for an event that occurs in nature is
BIOLOGICAL MOLECULES REVIEW-UNIT 1 1. The factor being tested in an experiment is the A. data. B. variable. C. conclusion. D. observation. 2. A possible explanation for an event that occurs in nature is
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION
 Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
Supplementary Information POLO-LIKE KINASE 1 FACILITATES LOSS OF PTEN-INDUCED PROSTATE CANCER FORMATION X. Shawn Liu 1, 3, Bing Song 2, 3, Bennett D. Elzey 3, 4, Timothy L. Ratliff 3, 4, Stephen F. Konieczny
supplementary information
 Figure S1 Nucleotide binding status of RagA mutants. Wild type and mutant forms of MycRagA was transfected into HEK293 cells and the transfected cells were labeled with 32 Pphosphate. MycRagA was immunoprecipitated
Figure S1 Nucleotide binding status of RagA mutants. Wild type and mutant forms of MycRagA was transfected into HEK293 cells and the transfected cells were labeled with 32 Pphosphate. MycRagA was immunoprecipitated
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism SUPPLEMENTARY FIGURES, LEGENDS AND METHODS
 A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
number Done by Corrected by Doctor Faisal Al-Khatibe
 number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
Insulin-induced gene 1 (insig-1) mrna increases dramatically in
 Insig-1 brakes lipogenesis in adipocytes and inhibits differentiation of preadipocytes Jinping Li*, Kiyosumi Takaishi*, William Cook*, Sara Kay McCorkle*, and Roger H. Unger* *Touchstone Center for Diabetes
Insig-1 brakes lipogenesis in adipocytes and inhibits differentiation of preadipocytes Jinping Li*, Kiyosumi Takaishi*, William Cook*, Sara Kay McCorkle*, and Roger H. Unger* *Touchstone Center for Diabetes
Human Urokinase / PLAU / UPA ELISA Pair Set
 Human Urokinase / PLAU / UPA ELISA Pair Set Catalog Number : SEK10815 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized
Human Urokinase / PLAU / UPA ELISA Pair Set Catalog Number : SEK10815 To achieve the best assay results, this manual must be read carefully before using this product and the assay is run as summarized
Application Note. Introduction
 Simultaneously Measuring Oxidation of Exogenous and Endogenous Fatty Acids Using the XF Palmitate-BSA FAO Substrate with the Agilent Seahorse XF Cell Mito Stress Test Application Note Introduction The
Simultaneously Measuring Oxidation of Exogenous and Endogenous Fatty Acids Using the XF Palmitate-BSA FAO Substrate with the Agilent Seahorse XF Cell Mito Stress Test Application Note Introduction The
Lecithin Cholesterol Acyltransferase (LCAT) ELISA Kit
 Product Manual Lecithin Cholesterol Acyltransferase (LCAT) ELISA Kit Catalog Number STA-616 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Cholesterol is a lipid sterol
Product Manual Lecithin Cholesterol Acyltransferase (LCAT) ELISA Kit Catalog Number STA-616 96 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Cholesterol is a lipid sterol
Data Sheet. Notch Pathway Reporter Kit Catalog # 60509
 Data Sheet Notch Pathway Reporter Kit Catalog # 60509 6042 Cornerstone Court W, Ste B Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. NOTCH signaling
Data Sheet Notch Pathway Reporter Kit Catalog # 60509 6042 Cornerstone Court W, Ste B Background The Notch signaling pathway controls cell fate decisions in vertebrate and invertebrate tissues. NOTCH signaling
Human Cathepsin D ELISA Kit
 GenWay Biotech, Inc. 6777 Nancy Ridge Drive San Diego, CA 92121 Phone: 858.458.0866 Fax: 858.458.0833 Email: techline@genwaybio.com http://www.genwaybio.com Human Cathepsin D ELISA Kit Catalog No. GWB-J4JVV9
GenWay Biotech, Inc. 6777 Nancy Ridge Drive San Diego, CA 92121 Phone: 858.458.0866 Fax: 858.458.0833 Email: techline@genwaybio.com http://www.genwaybio.com Human Cathepsin D ELISA Kit Catalog No. GWB-J4JVV9
EXOTESTTM. ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids
 DATA SHEET EXOTESTTM ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids INTRODUCTION Exosomes are small endosome-derived lipid nanoparticles
DATA SHEET EXOTESTTM ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids INTRODUCTION Exosomes are small endosome-derived lipid nanoparticles
