Acta Crystallographica Section D
|
|
|
- Maria Davis
- 5 years ago
- Views:
Transcription
1 Supporting information Acta Crystallographica Section D Volume 70 (2014) Supporting information for article: A conformational landscape for alginate secretion across the outer membrane of Pseudomonas aeruginosa Jingquan Tan, Sarah L. Rouse, Dianfan Li, Valerie E. Pye, Lutz Vogeley, Alette R. Brinth, John C. Whitney, P. Lynne Howell, Mark S. P. Sansom and Martin Caffrey
2 Supporting information, sup-1 Supporting Information A Conformational Landscape for Alginate Secretion Across the Outer Membrane of Pseudomonas aeruginosa Jingquan Tan 1,a, Sarah L. Rouse 2,a, Dianfan Li 1,a, Valerie E. Pye 1,3, Lutz Vogeley 1, Alette R. Brinth 1, John C. Whitney 4, P. Lynne Howell 4, Mark S. P. Sansom 2, Martin Caffrey 1* 1 Schools of Medicine and Biochemistry & Immunology, Trinity College, Dublin, Ireland; 2 Department of Biochemistry, University of Oxford, South Parks Road, Oxford, UK; 3 Current Address: Clare Hall Laboratories, Cancer Research UK, Herts., UK; 4 Program in Molecular Structure & Function, The Hospital for Sick Children, Toronto, Canada & University of Toronto, Toronto, Ontario, Canada. a These authors contributed equally to this work. *Corresponding author. martin.caffrey@tcd.ie Supplementary Figures and Tables Fig. S1-S9 Table S1
3 Supporting information, sup-2 Figure S1 Purification, spectroscopic characterization and crystallization of AlgE. (Ai) Size exclusion chromatographic analysis. V o, V e and V t mark the void, elution and total column volumes, respectively. The near Gaussian-shaped elution profile is consistent with a monodisperse protein preparation suitable for crystallization trials. (Aii) SDS-PAGE analysis. Protein purity of AlgE postsize exclusion chromatography was estimated at >90 % based on a loading series analysis with Coomassie Blue staining. The following AlgE loadings were used: 100 g, 10 g, 1 g and 0.1 g
4 Supporting information, sup-3 in lanes 1-4, respectively. Molecular weight standards are in lane 5. The hallmark heat modifiability of outer membrane -barrel proteins is illustrated in lanes 6-9 where AlgE samples were incubated at 50 o C for 0 min, 5 min, 20 min and 120 min, respectively, before being electrophoresed. (B) Spectrophotometric analysis of AlgE. (Bi) UV-visible, (Bii) fluorescence and (Biii) circular dichroism spectra of AlgE in a detergent micellar solution at 20 C. Spectra were recorded in Buffer B at 0.5, 0.1 and 1 mg AlgE/mL, respectively. An excitation wavelength of 280 nm was used for fluorescence data collection. Maximum absorbance and fluorescence emission was observed at 281 and 339 nm, respectively. The amino acid composition of AlgE is as follows: Ala, 37, 8.1%; Arg, 33, 7.2%; Asn, 20, 4.4%;, Asp, 46, 10.0%; Cys, 0, 0%; Gln, 20, 4.4%; Glu, 26, 5.6%; Gly, 51, 11.1%; His, 11, 2.4%; Ile, 16, 3.5%; Lys, 13, 2.5%; Leu, 36, 7.9%; Met, 4, 09%; Phe, 21, 4.6%; Pro, 14, 3.1%; Ser, 28, 6.1%; Thr, 34, 7.4%; Trp, 15, 3.3%; Tyr, 15, 3.3%; Val, 18, 3.9%. Analysis of the circular dichroism spectrum revealed a -strand, -helix, turn and coil content of 50, 20, 6 and 24 %, respectively, in reasonable agreement with the crystal structure. (C) Crystals of AlgE growing in the lipidic mesophase formed by 7.8 MAG at 20 o C. Trials were conducted as described (refs) using 10 mg AlgE/mL. The precipitant included (Ci) 0.1 M Bis-Tris, ph 7.0, 0.1 M sodium citrate, 21 %(v/v) PEG400; (Cii) 0.1 M sodium cacodylate, ph 6.5, 0.9 M sodium acetate; and (Ciii) 0.1 M Tris-HCl, ph 7.5, 0.1 M sodium citrate, 18 %(v/v) PEG 400. The crystal in (Ciii) gave best resolution to 1.9 Å. It continued to grow en route to the synchrotron and at time of data collection measured 150 m X 150 m. C, M and P refer to crystal, mesophase and precipitant, respectively. The image in (Ciii) was recorded with a different imager to that used in (Ci) and (Cii) which accounts for the difference in background color.
5 Supporting information, sup-4 Figure S2 Superposition of the in surfo and in meso AlgE structures. Chain B of AlgE-3RBH (red) and AlgE-2.4B (blue) were used for comparison because they represent the most complete structures. Structures are shown viewed from within the membrane (A), from outside the cell (B) and from the periplasm (C).
6 Supporting information
7 Supporting information, sup-6
8 Supporting information, sup-7 Figure S3 Membrane topology, sequence and secondary structure of in meso and in surfo AlgE models. (A) The model shown is based on the AlgE-1.9 in meso crystal structure. Squares correspond to residues in strands that, for the most part, constitute barrel staves. Blue shading identifies residues whose side chains face the lumen of the barrel. Residues that lack electron density are identified by red shading. Turns and loops are denoted T and L, respectively. -Strands are numbered sequentially toward the membrane mid-plane. The dashed horizontal lines show the approximate location of the membrane boundary as calculated in the PPM server ( (Lomize et al., 2012). The representation shown has been adjusted for clarity and legibility by orienting strands vertically and aligning them sequentially left to right. Note that the S5-L3-S6 segment between residues R150 and E166 folds into the lumen of the barrel and that sequential residues K129 and E130 both have side chains facing the lumen. Gray, round cornered boxes identify loops and turns that, when present in density, fold back into the barrel lumen. (B) Membrane topology, sequence and secondary structure comparison of in meso and in surfo AlgE models. Red coloring identifies residues that lack electron density and are assumed to be in disordered segments. Scanning across the panels it is apparent that the P-gate (T8) is present and presumed closed in models in iv-ix. The P-gate is open in ii and iii. By contrast, across the series from ii to ix the E-gate (L2) has varying stretches of the loop disordered corresponding to different degrees to which the gate is opened/closed. i-v correspond to in meso models reported in this work. vi-ix correspond to Molecules A-D in the in surfo model, 3RBH (Whitney et al., 2011).
9 Supporting information Figure S4 Packing arrangement in crystals of AlgE colored by asymmetric unit. (A) Crystal packing in AlgE-1.9, space group C2, which has one molecule per asymmetric unit. (B) Crystal packing in AlgE-2.4, space group P , which has two molecules per asymmetric unit. (C) Crystal packing in AlgE-2.8, P2 1, which has two molecules per asymmetric unit. Packing is of Type I, as observed with other structures solved with crystals grown by the in meso method (Caffrey et al., 2012).
10 Supporting information, sup-9 Figure S5 Superposition of the citrate from AlgE-1.9A on AlgE-2.8A illustrating the partial overlap in the positions of citrate and residues on T8 and L2 in the proposed alginate conduction pore. (A) Cutaway view of AlgE-2.8A is shown to reveal the channel interior. (B) Expanded view of the boxed area in (A).
11 Supporting information, sup-10 Figure S6 B-factor distribution within the AlgE structure shown in putty representation. AlgE-2.4B is shown and was chosen because of its completeness. The B-factor scale (Å 2 ), which runs from 12 (blue) to 125 (red), is shown on the right.
12 Supporting information, sup-11 Figure S7 Interactions of AlgE with lipids. (A) Average positions of lipid phosphate particles over a 1 μs coarse-grained (CG) simulation shown as a surface. Surface is coloured in terms of deformation compared to the mean lipid head group height over the course of the simulation. Some thinning of the bilayer is observed in the region of the protein. Residues Phe187 and Tyr190 (shown in magenta space-filling representation) sit above the short S5 and S6 strands. These hydrophobic residues compensate for the short β-strands such that the membrane does not significantly deform in this region. (B) Final snapshot of a CG simulation in which the bilayer has self-assembled around the AlgE protein. This snapshot is converted to atomistic representation (AT) for further simulation. The protein is coloured by secondary structure (β-sheet, yellow; α-helix, blue; turn, cyan; coil, white). Lipid phosphate groups are shown as tan spheres. Lipid tails and water molecules are not shown. A bound cardiolipin molecule is shown in both CG and AT representations. (C) Residues in contact (defined as within a 6 Å cut-off) with cardiolipin head groups during the course of the 1 μs CGMD simulation. Three binding sites in which a cardiolipin molecule is bound for over half the simulation are indicated. The inset shows the protein as spheres coloured according to the average time in contact with a cardiolipin head group (green = 0 % through white up to blue = 55 %). (D) The three binding sites identified from analysis in (C) are shown here with CG lipids coloured as transparent spheres mapped on to the 4AFK crystal structure with resolved bound lipids shown in stick representation. We propose that the outer-leaflet binding sites would be occupied by LPS.
13 Supporting information, sup-12 Figure S8 Water flux as a function of extracellular (E) gate opening. (A) The E gate opening may be measured in terms of the distance between residues Thr103 and Ser351 that form a hydrogen bonded lock when the E gate is not fully opened (upper panel). The number of water molecules crossing the pore over the simulation is shown as a cumulative count (lower panel). The gradients are shown when the lock is closed, when the lock is first broken, as the citrate moves through the constriction site and after the lock closes again. As citrate passes through the pore at time ~245 ns, coincident with L2 motion (increase in distance between Thr103 on L2 and Ser351 on L7) there is a sharp increase in the rate of water passing through. Citrate exits the pore in a hydrated state. As the L2 loop moves back towards its resting configuration (time 380 ns) the rate of water crossing reduces to close to the original level (0.40 by the final 10 ns of the trajectory, compared to 0.38 at the start). The black line corresponds to water molecules crossing the pore from the periplasmic side to the extracellular side, i.e., travelling in the direction of citrate. The red line corresponds to water molecules travelling in the reverse direction. (B) Snapshots corresponding to the lock closed (upper panel) and open (lower panel) states as indicated. The Thr103 and Ser351 side chains as well as the citrate molecule are shown in stick representation. Hydrogen bonds between these residues and citrate are shown as dashed lines.
14 Supporting information, sup-13 Figure S9 Steered molecular dynamics simulations of alginate being pushed and pulled through the AlgE pore. (A) The alginate is moved through the pore by attaching a harmonic spring to the terminal saccharides (coloured black for the push case and blue for the pulling case). The force is applied to the harmonic spring and directed along the z-axis in the direction from the extracellular side to the periplasmic side, indicated by arrows. (B) The force profiles (coloured as in A) give an indication of how much energy is required to move the alginate as it travels through the pore. The data corresponding to the push simulation is shown as In both cases the maximal applied force is similar (200 kj mol -1 nm -2 ). (C) The position along the pore of the sugar subunit proximal to the periplasmic side (coloured dark grey) is shown over time for both pushing and pulling simulations (coloured as in A and B). It is apparent from the non-linearity of these plots that the motion of alginate is not smooth, rather it moves through the pore in a step-like manner.
15 Supporting information Table S1 Comparison of topology and model completeness between AlgE structures. AlgE-1.9 (4AFK) a AlgE-2.4A (4B61) a AlgE-2.4B (4B61) a AlgE-2.8A (4AZL) a AlgE-2.8B (4AZL) a AlgE-2.3A (3BRH) AlgE-2.3B (3BRH) AlgE-2.3C (3BRH) AlgE-2.3D (3BRH) Region Missing b Missing Missing Missing Missing Missing Missing Missing Missing N-term / / / / / / / / / / / / / / / / / /7 Strand / / / / / / / / /13 - Loop / / / / / / / / / / /11 - Strand / / / / / / / / /16 - Turn / / / / / / / / /2 - Strand / / / / / / / / /12 - Loop 2 c / / / / / / / / / / / / / / / / / /17 Strand / / / / / / / / /12 - Turn 2 d / / / / / / / / /8 - Strand 5 d / / / / / / / / /11 - Loop / / / / / / / / /7 - Strand / / / / / / / / /12 - Turn / / / / / / / / /3 - Strand / / / / / / / / /9 - Loop 4 d / / / / / / / / /18 - Strand 8 d / / / / / / / / /13 - Turn / / / / / / / / /2 - Strand 9 d / / / / / / / / /12 - Loop 5 d / / / / / / / / / / / / /10 Strand / / / / / / / / /14 - Turn / / / / / / / / /10 - Strand 11 d / / / / / / / / /16 - Loop 6 d,e / / / / / / / / / / / / / / /14 Strand 12 d / / / / / / / / /16 - Turn / / / / / / / / /4 - Strand / / / / / / / / /10 - Loop / / / / / / / / /43 - Strand / / / / / / / / /12 -
16 Supporting information, sup-15 Turn / / / / / / / / /3 - Strand / / / / / / / / /12 - Loop / / / / / / / / /21 - Strand / / / / / / / / /13 - Turn 8 c / / / / / / / / / / /19 - Strand / / / / / / / / /11 - Loop / / / / / / / / / /11 - Strand / / / / / / / / /11 - C-term. 490/1-490/1-490/1-490/1-490/1-490/1-490/1-490/1-490/1 a Structure solved as part of this work. b Refers to residues that lack electron density. c Loop 2 and Turn 8, the two regions where most of the discussion is focused, are underlined. d Regions that vary in topology between the different models. Differences with AlgE-1.9 are highlighted in bold. e Loop and strand designations are based on the AlgE-1.9 model. References Caffrey, M., D. Li & A. Dukkipati (2012). Biochemistry, 51, Lomize, M. A., I. D. Pogozheva, H. Joo, H. I. Mosberg & A. L. Lomize (2012). Nucleic Acids Res., 40, D370-D376. Whitney, J. C., I. D. Hay, C. H. Li, P. D. W. Eckford, H. Robinson, M. F. Amaya, L. F. Wood, D. E. Ohman, C. E. Bear, B. H. Rehm & P. L. Howell (2011). Proc. Natl. Acad. Sci. U.S.A., 108,
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information.
 Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
Structural analysis of fungus-derived FAD glucose dehydrogenase
 Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Biomolecules: amino acids
 Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Structure of the measles virus hemagglutinin bound to the CD46 receptor. César Santiago, María L. Celma, Thilo Stehle and José M.
 Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Introduction to Protein Structure Collection
 Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
The Basics: A general review of molecular biology:
 The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
Biology. Lectures winter term st year of Pharmacy study
 Biology Lectures winter term 2008 1 st year of Pharmacy study 3 rd Lecture Chemical composition of living matter chemical basis of life. Atoms, molecules, organic compounds carbohydrates, lipids, proteins,
Biology Lectures winter term 2008 1 st year of Pharmacy study 3 rd Lecture Chemical composition of living matter chemical basis of life. Atoms, molecules, organic compounds carbohydrates, lipids, proteins,
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A
 Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Review II: The Molecules of Life
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 4 Protein Sequence
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 4 Protein Sequence 2 3 4 Are You Getting It?? A molecule of hemoglobin is compared with a molecule of lysozyme. Which characteristics do they share?
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 4 Protein Sequence 2 3 4 Are You Getting It?? A molecule of hemoglobin is compared with a molecule of lysozyme. Which characteristics do they share?
paper and beads don t fall off. Then, place the beads in the following order on the pipe cleaner:
 Beady Pipe Cleaner Proteins Background: Proteins are the molecules that carry out most of the cell s dayto-day functions. While the DNA in the nucleus is "the boss" and controls the activities of the cell,
Beady Pipe Cleaner Proteins Background: Proteins are the molecules that carry out most of the cell s dayto-day functions. While the DNA in the nucleus is "the boss" and controls the activities of the cell,
Short polymer. Dehydration removes a water molecule, forming a new bond. Longer polymer (a) Dehydration reaction in the synthesis of a polymer
 HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
BIO 311C Spring Lecture 15 Friday 26 Feb. 1
 BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Lecture 33 Membrane Proteins
 Lecture 33 Membrane Proteins Reading for today: Chapter 4, section D Required reading for next Wednesday: Chapter 14, sections A and 14.19 to the end Kuriyan, J., and Eisenberg, D. (2007) The origin of
Lecture 33 Membrane Proteins Reading for today: Chapter 4, section D Required reading for next Wednesday: Chapter 14, sections A and 14.19 to the end Kuriyan, J., and Eisenberg, D. (2007) The origin of
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Reading from the NCBI
 Reading from the NCBI http://www.ncbi.nlm.nih.gov/books/bv.fcgi?highlight=thermodyn amics&rid=stryer.section.156#167 http://www.ncbi.nlm.nih.gov/books/bv.fcgi?highlight=stability,pr otein&rid=stryer.section.365#371
Reading from the NCBI http://www.ncbi.nlm.nih.gov/books/bv.fcgi?highlight=thermodyn amics&rid=stryer.section.156#167 http://www.ncbi.nlm.nih.gov/books/bv.fcgi?highlight=stability,pr otein&rid=stryer.section.365#371
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL
 Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al.
 Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Properties of amino acids in proteins
 Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
CHAPTER 21: Amino Acids, Proteins, & Enzymes. General, Organic, & Biological Chemistry Janice Gorzynski Smith
 CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
Supplementary Information
 Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Electronic Supplementary Information. Table of Contents
 Electronic Supplementary Information Examination of native chemical ligation using peptidyl prolyl thioester Takahiro Nakamura, Akira Shigenaga, Kohei Sato, Yusuke Tsuda, Ken Sakamoto, and Akira Otaka*
Electronic Supplementary Information Examination of native chemical ligation using peptidyl prolyl thioester Takahiro Nakamura, Akira Shigenaga, Kohei Sato, Yusuke Tsuda, Ken Sakamoto, and Akira Otaka*
The Structure and Function of Macromolecules
 The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
Copyright 2008 Pearson Education, Inc., publishing as Pearson Benjamin Cummings
 Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
Methionine (Met or M)
 Fig. 5-17 Nonpolar Fig. 5-17a Nonpolar Glycine (Gly or G) Alanine (Ala or A) Valine (Val or V) Leucine (Leu or L) Isoleucine (Ile or I) Methionine (Met or M) Phenylalanine (Phe or F) Polar Trypotphan (Trp
Fig. 5-17 Nonpolar Fig. 5-17a Nonpolar Glycine (Gly or G) Alanine (Ala or A) Valine (Val or V) Leucine (Leu or L) Isoleucine (Ile or I) Methionine (Met or M) Phenylalanine (Phe or F) Polar Trypotphan (Trp
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
Levels of Protein Structure:
 Levels of Protein Structure: PRIMARY STRUCTURE (1 ) - Defined, non-random sequence of amino acids along the peptide backbone o Described in two ways: Amino acid composition Amino acid sequence M-L-D-G-C-G
Levels of Protein Structure: PRIMARY STRUCTURE (1 ) - Defined, non-random sequence of amino acids along the peptide backbone o Described in two ways: Amino acid composition Amino acid sequence M-L-D-G-C-G
PROTEINS. Building blocks, structure and function. Aim: You will have a clear picture of protein construction and their general properties
 PROTEINS Building blocks, structure and function Aim: You will have a clear picture of protein construction and their general properties Reading materials: Compendium in Biochemistry, page 13-49. Microbiology,
PROTEINS Building blocks, structure and function Aim: You will have a clear picture of protein construction and their general properties Reading materials: Compendium in Biochemistry, page 13-49. Microbiology,
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
Coarse Grained Molecular Dynamics Simulations of Transmembrane Protein-Lipid Systems
 Int. J. Mol. Sci. 2010, 11, 2393-2420; doi:10.3390/ijms11062393 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Article Coarse Grained Molecular Dynamics
Int. J. Mol. Sci. 2010, 11, 2393-2420; doi:10.3390/ijms11062393 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Article Coarse Grained Molecular Dynamics
LAB#23: Biochemical Evidence of Evolution Name: Period Date :
 LAB#23: Biochemical Evidence of Name: Period Date : Laboratory Experience #23 Bridge Worth 80 Lab Minutes If two organisms have similar portions of DNA (genes), these organisms will probably make similar
LAB#23: Biochemical Evidence of Name: Period Date : Laboratory Experience #23 Bridge Worth 80 Lab Minutes If two organisms have similar portions of DNA (genes), these organisms will probably make similar
Lipids: diverse group of hydrophobic molecules
 Lipids: diverse group of hydrophobic molecules Lipids only macromolecules that do not form polymers li3le or no affinity for water hydrophobic consist mostly of hydrocarbons nonpolar covalent bonds fats
Lipids: diverse group of hydrophobic molecules Lipids only macromolecules that do not form polymers li3le or no affinity for water hydrophobic consist mostly of hydrocarbons nonpolar covalent bonds fats
G protein coupled receptor interactions with cholesterol deep in the membrane
 G protein coupled receptor interactions with cholesterol deep in the membrane Samuel Genheden a,b, Jonathan W. Essex a,c, & Anthony G. Lee c,d * a School of Chemistry, University of Southampton, Southampton,
G protein coupled receptor interactions with cholesterol deep in the membrane Samuel Genheden a,b, Jonathan W. Essex a,c, & Anthony G. Lee c,d * a School of Chemistry, University of Southampton, Southampton,
Protein Secondary Structure
 Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
PROTEINS. Amino acids are the building blocks of proteins. Acid L-form * * Lecture 6 Macromolecules #2 O = N -C -C-O.
 Proteins: Linear polymers of amino acids workhorses of the cell tools, machines & scaffolds Lecture 6 Macromolecules #2 PRTEINS 1 Enzymes catalysts that mediate reactions, increase reaction rate Structural
Proteins: Linear polymers of amino acids workhorses of the cell tools, machines & scaffolds Lecture 6 Macromolecules #2 PRTEINS 1 Enzymes catalysts that mediate reactions, increase reaction rate Structural
Judy Wieber. Department of Computational Biology. May 27, 2008
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2008 Department of Computational Biology University it of Pittsburgh School of Medicine i May 27, 2008 Outline Introduction Proteins Carbohydrates
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2008 Department of Computational Biology University it of Pittsburgh School of Medicine i May 27, 2008 Outline Introduction Proteins Carbohydrates
Chemistry 121 Winter 17
 Chemistry 121 Winter 17 Introduction to Organic Chemistry and Biochemistry Instructor Dr. Upali Siriwardane (Ph.D. Ohio State) E-mail: upali@latech.edu Office: 311 Carson Taylor Hall ; Phone: 318-257-4941;
Chemistry 121 Winter 17 Introduction to Organic Chemistry and Biochemistry Instructor Dr. Upali Siriwardane (Ph.D. Ohio State) E-mail: upali@latech.edu Office: 311 Carson Taylor Hall ; Phone: 318-257-4941;
Practice Problems 3. a. What is the name of the bond formed between two amino acids? Are these bonds free to rotate?
 Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Moorpark College Chemistry 11 Fall Instructor: Professor Gopal. Examination # 5: Section Five May 7, Name: (print)
 Moorpark College Chemistry 11 Fall 2013 Instructor: Professor Gopal Examination # 5: Section Five May 7, 2013 Name: (print) Directions: Make sure your examination contains TEN total pages (including this
Moorpark College Chemistry 11 Fall 2013 Instructor: Professor Gopal Examination # 5: Section Five May 7, 2013 Name: (print) Directions: Make sure your examination contains TEN total pages (including this
Macromolecules of Life -3 Amino Acids & Proteins
 Macromolecules of Life -3 Amino Acids & Proteins Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Amino Acids Proteins
Macromolecules of Life -3 Amino Acids & Proteins Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Amino Acids Proteins
AP Bio. Protiens Chapter 5 1
 Concept.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 0% of the dry mass of most cells Protein functions include structural support, storage, transport,
Concept.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 0% of the dry mass of most cells Protein functions include structural support, storage, transport,
The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5
 Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Modeling and Molecular Dynamics of Membrane Proteins
 Modeling and Molecular Dynamics of Membrane Proteins Emad Tajkhorshid Department of Biochemistry, Center for Biophysics and Computational Biology, and Beckman Institute University of Illinois at Urbana-Champaign
Modeling and Molecular Dynamics of Membrane Proteins Emad Tajkhorshid Department of Biochemistry, Center for Biophysics and Computational Biology, and Beckman Institute University of Illinois at Urbana-Champaign
Protein Structure Monday, March
 Michael Morales ffice: 204/202 ary; knock hard if you visit as my office is some door. Lab: 238 & 205 ary ffice hours Either make an appointment or just drop by Phone: 829-3965 E-Mail: moralesm@buffalo.edu
Michael Morales ffice: 204/202 ary; knock hard if you visit as my office is some door. Lab: 238 & 205 ary ffice hours Either make an appointment or just drop by Phone: 829-3965 E-Mail: moralesm@buffalo.edu
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Lipid Bilayers Are Excellent For Cell Membranes
 Lipid Bilayers Are Excellent For Cell Membranes ydrophobic interaction is the driving force Self-assembly in water Tendency to close on themselves Self-sealing (a hole is unfavorable) Extensive: up to
Lipid Bilayers Are Excellent For Cell Membranes ydrophobic interaction is the driving force Self-assembly in water Tendency to close on themselves Self-sealing (a hole is unfavorable) Extensive: up to
P450 CYCLE. All P450s follow the same catalytic cycle of;
 P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
Proteins. Amino acids, structure and function. The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka
 Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Amino Acids. Amino Acids. Fundamentals. While their name implies that amino acids are compounds that contain an NH. 3 and CO NH 3
 Fundamentals While their name implies that amino acids are compounds that contain an 2 group and a 2 group, these groups are actually present as 3 and 2 respectively. They are classified as α, β, γ, etc..
Fundamentals While their name implies that amino acids are compounds that contain an 2 group and a 2 group, these groups are actually present as 3 and 2 respectively. They are classified as α, β, γ, etc..
Wheat Amino acids & Peptides for Hair Care. INCI Name EU/USA CAS # EINECS # Hydrolyzed wheat protein
 KELYAMIN Wheat Amino acids & Peptides for Hair Care Identification INCI Name EU/USA CAS # EINECS # Hydrolyzed wheat protein 70084-87-6 305-225-0 Composition % Liquid Powder Aqua Hydrolyzed wheat protein
KELYAMIN Wheat Amino acids & Peptides for Hair Care Identification INCI Name EU/USA CAS # EINECS # Hydrolyzed wheat protein 70084-87-6 305-225-0 Composition % Liquid Powder Aqua Hydrolyzed wheat protein
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Four melanocyte-stimulating hormones have the following amino acid sequences:
 Assignment 14: Melanocyte-stimulating hormone belongs to a group called the melanocortins. This group includes ACTH, alpha-msh, beta-msh and gamma-msh; these peptides are all cleavage products of a large
Assignment 14: Melanocyte-stimulating hormone belongs to a group called the melanocortins. This group includes ACTH, alpha-msh, beta-msh and gamma-msh; these peptides are all cleavage products of a large
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Cells. Variation and Function of Cells
 Cells Variation and Function of Cells Plasma Membrane= the skin of a cell, it protects and nourishes the cell while communicating with other cells at the same time. Lipid means fat and they are hydrophobic
Cells Variation and Function of Cells Plasma Membrane= the skin of a cell, it protects and nourishes the cell while communicating with other cells at the same time. Lipid means fat and they are hydrophobic
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction
 Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector
 kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
Life Sciences 1a. Practice Problems 4
 Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
1. Describe the relationship of dietary protein and the health of major body systems.
 Food Explorations Lab I: The Building Blocks STUDENT LAB INVESTIGATIONS Name: Lab Overview In this investigation, you will be constructing animal and plant proteins using beads to represent the amino acids.
Food Explorations Lab I: The Building Blocks STUDENT LAB INVESTIGATIONS Name: Lab Overview In this investigation, you will be constructing animal and plant proteins using beads to represent the amino acids.
Macromolecules Structure and Function
 Macromolecules Structure and Function Within cells, small organic molecules (monomers) are joined together to form larger molecules (polymers). Macromolecules are large molecules composed of thousands
Macromolecules Structure and Function Within cells, small organic molecules (monomers) are joined together to form larger molecules (polymers). Macromolecules are large molecules composed of thousands
Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids
 Supporting Information Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids Phansiri Boonnoy 1, Viwan Jarerattanachat 1,2, Mikko Karttunen 3*, and Jirasak Wongekkabut 1* 1 Department of
Supporting Information Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids Phansiri Boonnoy 1, Viwan Jarerattanachat 1,2, Mikko Karttunen 3*, and Jirasak Wongekkabut 1* 1 Department of
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Quantitative LC-MS/MS Analysis of Glucagon. Veniamin Lapko, Ph.D June 21, 2011
 Quantitative LC-MS/MS Analysis of Glucagon Veniamin Lapko, Ph.D June 21, 2011 Contents Comparison with small molecule LC-MS/MS LC-MS/MS sensitivity of peptides detection Stability: neat vs. matrix solutions
Quantitative LC-MS/MS Analysis of Glucagon Veniamin Lapko, Ph.D June 21, 2011 Contents Comparison with small molecule LC-MS/MS LC-MS/MS sensitivity of peptides detection Stability: neat vs. matrix solutions
1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled
 Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Catalysis & specificity: Proteins at work
 Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total. The ph at which a peptide has no net charge is its isoelectric point.
 BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
Protein-Lipid Interactions: Structural and Functional Effects Anthony Lee (Southampton)
 Saulieu ctober 2004 Protein-Lipid Interactions: Structural and Functional Effects Anthony Lee (Southampton) The membrane as a system Co-evolution of lipids and membrane proteins R P - R Phosphatidylcholine
Saulieu ctober 2004 Protein-Lipid Interactions: Structural and Functional Effects Anthony Lee (Southampton) The membrane as a system Co-evolution of lipids and membrane proteins R P - R Phosphatidylcholine
Gold nanocrystals at DPPC bilayer. Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad
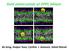 Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
SUPPORTING INFORMATION. Lysine Carbonylation is a Previously Unrecognized Contributor. to Peroxidase Activation of Cytochrome c by Chloramine-T
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Maha AbuAjamieh. Tamara Wahbeh. Mamoon Ahram
 12 Maha AbuAjamieh Tamara Wahbeh Mamoon Ahram - - Go to this sheet s last page for definitions of the words with an asterisk above them (*) - You should memorise the 3-letter abbreviations, of all the
12 Maha AbuAjamieh Tamara Wahbeh Mamoon Ahram - - Go to this sheet s last page for definitions of the words with an asterisk above them (*) - You should memorise the 3-letter abbreviations, of all the
SIMPLE BASIC METABOLISM
 SIMPLE BASIC METABOLISM When we eat food such as a tuna fish sandwich, the polysaccharides, lipids, and proteins are digested to smaller molecules that are absorbed into the cells of our body. As these
SIMPLE BASIC METABOLISM When we eat food such as a tuna fish sandwich, the polysaccharides, lipids, and proteins are digested to smaller molecules that are absorbed into the cells of our body. As these
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
(65 pts.) 27. (10 pts.) 28. (15 pts.) 29. (10 pts.) TOTAL (100 points) Moorpark College Chemistry 11 Spring Instructor: Professor Gopal
 Moorpark College Chemistry 11 Spring 2012 Instructor: Professor Gopal Examination # 5: Section Five May 1, 2012 Name: (print) GOOD LUCK! Directions: Make sure your examination contains TWELVE total pages
Moorpark College Chemistry 11 Spring 2012 Instructor: Professor Gopal Examination # 5: Section Five May 1, 2012 Name: (print) GOOD LUCK! Directions: Make sure your examination contains TWELVE total pages
Four Classes of Biological Macromolecules. Biological Macromolecules. Lipids
 Biological Macromolecules Much larger than other par4cles found in cells Made up of smaller subunits Found in all cells Great diversity of func4ons Four Classes of Biological Macromolecules Lipids Polysaccharides
Biological Macromolecules Much larger than other par4cles found in cells Made up of smaller subunits Found in all cells Great diversity of func4ons Four Classes of Biological Macromolecules Lipids Polysaccharides
Supplementary Information
 Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Cells N5 Homework book
 1 Cells N5 Homework book 2 Homework 1 3 4 5 Homework2 Cell Ultrastructure and Membrane 1. Name and give the function of the numbered organelles in the cell below: A E B D C 2. Name 3 structures you might
1 Cells N5 Homework book 2 Homework 1 3 4 5 Homework2 Cell Ultrastructure and Membrane 1. Name and give the function of the numbered organelles in the cell below: A E B D C 2. Name 3 structures you might
Head. Tail. Carboxyl group. group. group. air water. Hydrocarbon chain. lecture 5-sa Seth Copen Goldstein 2.
 Lipids Some lipid structures Organic compounds Amphipathic Polar head group (hydrophilic) Non-polar tails (hydrophobic) Lots of uses Energy storage Membranes Hormones Vitamins HO O C H 2 C CH 2 H 2 C CH
Lipids Some lipid structures Organic compounds Amphipathic Polar head group (hydrophilic) Non-polar tails (hydrophobic) Lots of uses Energy storage Membranes Hormones Vitamins HO O C H 2 C CH 2 H 2 C CH
Amino acids-incorporated nanoflowers with an
 Amino acids-incorporated nanoflowers with an intrinsic peroxidase-like activity Zhuo-Fu Wu 1,2,+, Zhi Wang 1,+, Ye Zhang 3, Ya-Li Ma 3, Cheng-Yan He 4, Heng Li 1, Lei Chen 1, Qi-Sheng Huo 3, Lei Wang 1,*
Amino acids-incorporated nanoflowers with an intrinsic peroxidase-like activity Zhuo-Fu Wu 1,2,+, Zhi Wang 1,+, Ye Zhang 3, Ya-Li Ma 3, Cheng-Yan He 4, Heng Li 1, Lei Chen 1, Qi-Sheng Huo 3, Lei Wang 1,*
Organic Chemistry 3540
 rganic Chemistry 3540 December 8, 2004 (8 Pages, 13 Parts) ame 1. (8%) Many organic compounds found in living systems are complex molecules which can be characterized, in part, by simply listing the chemical
rganic Chemistry 3540 December 8, 2004 (8 Pages, 13 Parts) ame 1. (8%) Many organic compounds found in living systems are complex molecules which can be characterized, in part, by simply listing the chemical
