Supplementary Material
|
|
|
- Erick Stafford
- 5 years ago
- Views:
Transcription
1 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total of 200 l of reaction mixture containing 0.5% salicin and enzyme (10 l diluted stored-enzyme in 50 mm HEPES and 100 mm NaCl, ph 8.0) in 100 mm phosphate-citrate buffer was incubated at ph 5.5 at 313 K for 30 minutes. Next, 3,5-dinitrosalicylic acid reagent (200 l) was added, and the solution was incubated at 368 K for five minutes. Finally, 600 l distilled water was added, and the absorbance was measured at 540 nm using a Lambda Bio 40 spectrometer (Perkin Elmer). The enzymatic activity of -glucosidase toward cellobiose was measured according to the protocol of an Amplex red glucose/glucose oxidase assay kit (Invitrogen) using glucose as the standard. The total reaction mixture was 200 l and contained 0.5% cellobiose and enzyme (10 l diluted stored-enzyme in 50 mm HEPES and 100 mm NaCl, ph 8.0) in 100 mm phosphate-citrate buffer, and this was incubated at ph 5.5 at 313 K for 30 minutes; subsequently, the solution was incubated at 368 K for five minutes to stop the reaction. Finally, the solution was diluted and manipulated according to the protocol of an Amplex red glucose/glucose oxidase assay kit and monitored at excitation and emission wavelengths of 570 nm and 585 nm, respectively, with a Fluorolog-3 spectrofluorometer (Horiba). One unit of enzyme activity corresponded to one and two mol of glucose per minute in the reaction for NkBgl toward salicin and cellobiose, respectively. Protein concentrations were determined as described by Bradford (1976) using the Bio-Rad Protein Assay Kit with BSA as the standard. The experiments were performed in triplicate using a single prepared protein sample. 1
2 Table S1. Alternative conformations in the crystal structures of NkBgl. Crystals (Gluconolactone + (1-Deoxynojirimycin + (Glycerol) (Bis-Tris) E193A (Glucose) (EPPS-1-glucoside) (Opipramol-1-glucoside) Residues No E34 1 O E66 1 O I79 3 O O O E89 1 O K92 1 O E96 4 O O O O V O O O O O O O O O O O R103 3 O O O E114 8 O O O O O O O O Q122 6 O O O O O O L189 1 O D199 6 O O O O O O E204 1 O I205 1 O M O O O O O O O O O O I211 9 O O O O O O O O O R231 8 O O O O O O O O Q238 6 O O O O O O E243 2 O O K247 3 O O O E265 1 O E271 2 O O L280 9 O O O O O O O O O E288 1 O L O O O O O O O O O O S O O O O O O O O O O K343 2 O O K344 2 O O R O O O O O O O O O O Q O O O O O O O O O O O S373 1 O K390 2 O O N412 1 O T O O O O O O O O O O S445 2 O O M O O O O O O O O O O O I475 2 O O K491 1 O E494 1 O 2
3 Table S2. The root-mean-square differences (Å) among NkBgl models. (Gluconolactone + (1-Deoxynojirimycin + (Glycerol) (Bis-Tris) (Tris) E193A (Glucose) (EPPS-1-glucoside) (Opipramol-1-glucoside) (pnpg) (Gluconolactone + (1-Deoxynojirimycin + (Glycerol) (Bis-Tris) (Tris) E193A (Glucose) (EPPS-1-glucoside) (Opipramol-1-glucoside) (pnpg)
4 Table S3. The distance (Å) between the C atom of residue-193 and the C atoms of Tyr337, Trp374, Glu402, Trp444, and Glu451 and the C 3 atom of Trp374. The distance (Å) between 193C and (Gluconolactone + (1-Deoxynojirimycin + (Glycerol) (Bis-Tris) (Tris) (EPPS-1-glucoside) (Opipramol-1-glucoside) (pnpg) E193A (Glucose) E402C E451C Y337C W374C W374C 3 W444C Average
5 Figure S1. A stereoview of the C traces of thirteen superimposed NkBgl structures. The view is from the entrance of the active site pocket. 5
6 6
7 Figure S2. A stereoview of the ligand molecules and their surrounding residues in the active sites of NkBgl structures. The wild-type NkBgl is in complex with (a) glycerol, and (b) Bis-Tris. (c) The NkBgl- mutant is in complex with salicin. (d) The NkBgl-E193A mutant is in complex with glucose. (e) The NkBgl- mutant is in complex with HEPES-1-glucoside, which is a new adduct generated from salicin mixing with HEPES. Fo - Fc difference Fourier maps for the ligand molecules contoured at 3.0 are shown in light blue. 2 Fo - Fc maps for the side chains of selective residues contoured at 2.0 are shown in gray, whereas water molecules contoured at 1.5 are shown in yellow-orange. The side chains of residues around the active site of the NkBgl are shown as line models. Ligand molecules are shown as stick models. The E402 residue labeled in orange is the catalytic nucleophile of NkBgl. The acid-base residues of wild-type or mutated NkBgl are labeled in magenta. Hydrogen-bond interactions between residues and ligand molecules are labeled in magenta dotted lines. Carbon atoms of proteins and ligand molecules are shown in cyan and green, respectively. Protein oxygen atoms are shown in red, nitrogen in blue and sulfur in gold. Next to the carboxyl group of Asp193, a water molecule, presumably involved in both the hydrolysis and transglycosylation reactions is indicated with the magenta asterisk. 7
8 Figure S3. Activity assays of the wild-type NkBgl toward salicin or cellobiose substrates. The measurements were analyzed in triplicate with error bars indicated. Error bars represent the standard deviation of three replicates from a single prepared protein sample. 8
9 Figure S4. A MALDI mass spectrometric analysis of the new glucopyranosidic product opipramol-1-glucoside. References Bradford, M. M. (1976). Anal Biochem 72, Miller, G. L. (1959). Anal Chem 31,
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Structural analysis of fungus-derived FAD glucose dehydrogenase
 Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Structure of the measles virus hemagglutinin bound to the CD46 receptor. César Santiago, María L. Celma, Thilo Stehle and José M.
 Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Guided Inquiry Skills Lab. Additional Lab 1 Making Models of Macromolecules. Problem. Introduction. Skills Focus. Materials.
 Additional Lab 1 Making Models of Macromolecules Guided Inquiry Skills Lab Problem How do monomers join together to form polymers? Introduction A small number of elements make up most of the mass of your
Additional Lab 1 Making Models of Macromolecules Guided Inquiry Skills Lab Problem How do monomers join together to form polymers? Introduction A small number of elements make up most of the mass of your
Data are contained in multiple tabs in Excel spreadsheets and in CSV files.
 Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
SUPPLEMENTARY INFORMATION FOR. (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation. by COX-2
 SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
SUPPLEMENTARY INFORMATION FOR (R)-Profens Are Substrate-Selective Inhibitors of Endocannabinoid Oxygenation by COX-2 Kelsey C. Duggan, Daniel J. Hermanson, Joel Musee, Jeffery J. Prusakiewicz, Jami L.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Information
 Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
B. 1% (w/v) Salicin Substrate Solution (Salicin) (Prepare 50 ml in Reagent A using Salicin, Sigma Prod. No. S-0625.)
 SIGMA QUALITY CONTROL TEST PROCEDURE (Q]\PDWLFÃ$VVD\ÃRIÃ */8&26,'$6( PRINCIPLE: 'Glucoside + H 2 O Glucosidase > D-Glucose + an Alcohol CONDITIONS: T = 37 C, ph = 5.0, A 540nm, Light path = 1 cm METHOD:
SIGMA QUALITY CONTROL TEST PROCEDURE (Q]\PDWLFÃ$VVD\ÃRIÃ */8&26,'$6( PRINCIPLE: 'Glucoside + H 2 O Glucosidase > D-Glucose + an Alcohol CONDITIONS: T = 37 C, ph = 5.0, A 540nm, Light path = 1 cm METHOD:
Supporting Information
 Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Previous Class. Today. Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism. Protein Kinase Catalytic Properties
 Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Excerpt from J. Mol. Biol. (2002) 320, :
 Excerpt from J. Mol. Biol. (2002) 320, 1095 1108: Crystal Structure of the Ternary Complex of the Catalytic Domain of Human Phenylalanine Hydroxylase with Tetrahydrobiopterin and 3-(2-Thienyl)-L-alanine,
Excerpt from J. Mol. Biol. (2002) 320, 1095 1108: Crystal Structure of the Ternary Complex of the Catalytic Domain of Human Phenylalanine Hydroxylase with Tetrahydrobiopterin and 3-(2-Thienyl)-L-alanine,
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
Soil Enzyme Assays K. DeAngelis October 24, 2005
 Soil Enzyme Assays K. DeAngelis October 24, 2005 Materials N-AcetylGlucosamidine-MUB Sigma #M2133 Nuclease - Phosphate-MUB Sigma #3168 MUB (for standards) Sigma #1381 Phenol Oxidase - L-DihydrOxyPhenylAlanine
Soil Enzyme Assays K. DeAngelis October 24, 2005 Materials N-AcetylGlucosamidine-MUB Sigma #M2133 Nuclease - Phosphate-MUB Sigma #3168 MUB (for standards) Sigma #1381 Phenol Oxidase - L-DihydrOxyPhenylAlanine
2.2 Properties of Water
 2.2 Properties of Water I. Water s unique properties allow life to exist on Earth. A. Life depends on hydrogen bonds in water. B. Water is a polar molecule. 1. Polar molecules have slightly charged regions
2.2 Properties of Water I. Water s unique properties allow life to exist on Earth. A. Life depends on hydrogen bonds in water. B. Water is a polar molecule. 1. Polar molecules have slightly charged regions
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants
 Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
Experiment 3: Activity Determination
 Experiment 3: Activity Determination Introduction: Specific activity is a method for measuring enzymatic activity and the enzyme purity in a mixture. In order to determine the specific activity of an enzyme,
Experiment 3: Activity Determination Introduction: Specific activity is a method for measuring enzymatic activity and the enzyme purity in a mixture. In order to determine the specific activity of an enzyme,
Interactions of Polyethylenimines with Zwitterionic and. Anionic Lipid Membranes
 Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Analysis & Interpretation. Analysis Questions answer questions on a separate sheet of paper. Name(s): Period: Date:
 Name(s): Period: Date: Dehydration Synthesis and Hydrolysis The chemical reactions that bond together macromolecules are similar and require water. When macromolecules are consumed, they must be broken
Name(s): Period: Date: Dehydration Synthesis and Hydrolysis The chemical reactions that bond together macromolecules are similar and require water. When macromolecules are consumed, they must be broken
Chymotrypsin Lecture. Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin
 Chymotrypsin Lecture Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin What s so great about enzymes? They accomplish large rate accelerations (10 10-10
Chymotrypsin Lecture Aims: to understand (1) the catalytic strategies used by enzymes and (2) the mechanism of chymotrypsin What s so great about enzymes? They accomplish large rate accelerations (10 10-10
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
Supplementary Information for Structural basis for the inhibition of Polo-like kinase 1
 Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Chemical Formulas. Chemical Formula CH 3 COCHCHOCHClCHNH Lewis Dot Structure
 Biochemistry . Chemical Formulas A chemical formula represents the chemical makeup of a compound. It shows the numbers and kinds of atoms present in a compound. It is a kind of shorthand that scientists
Biochemistry . Chemical Formulas A chemical formula represents the chemical makeup of a compound. It shows the numbers and kinds of atoms present in a compound. It is a kind of shorthand that scientists
Serrata) Alkaline Phosphatase
 Vol. 41, No. 5, April 1997 BIOCHEMISTRY and MOLECULAR BIOLOGY INTERNATIONAL Pages 951-959 An Essential Tryptophan Residue of Green Crab (Syclla Serrata) Alkaline Phosphatase Wen-Zhu Zheng 1, Qing-Xi Chen
Vol. 41, No. 5, April 1997 BIOCHEMISTRY and MOLECULAR BIOLOGY INTERNATIONAL Pages 951-959 An Essential Tryptophan Residue of Green Crab (Syclla Serrata) Alkaline Phosphatase Wen-Zhu Zheng 1, Qing-Xi Chen
Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids
 Supporting Information Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids Phansiri Boonnoy 1, Viwan Jarerattanachat 1,2, Mikko Karttunen 3*, and Jirasak Wongekkabut 1* 1 Department of
Supporting Information Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids Phansiri Boonnoy 1, Viwan Jarerattanachat 1,2, Mikko Karttunen 3*, and Jirasak Wongekkabut 1* 1 Department of
Enzymatic Assay of ß-GLUCOSIDASE (EC )
 PRINCIPLE: ß-D-Glucoside + H 2 O ß-Glucosidase > D-Glucose + an Alcohol CONDITIONS: T = 37 C, ph = 5.0, A 540nm, Light path = 1 cm METHOD: Colorimetric 1 REAGENTS: A. 100 mm Sodium Acetate Buffer, ph 5.0
PRINCIPLE: ß-D-Glucoside + H 2 O ß-Glucosidase > D-Glucose + an Alcohol CONDITIONS: T = 37 C, ph = 5.0, A 540nm, Light path = 1 cm METHOD: Colorimetric 1 REAGENTS: A. 100 mm Sodium Acetate Buffer, ph 5.0
Crystallization-grade After D After V3 cocktail. Time (s) Time (s) Time (s) Time (s) Time (s) Time (s)
 Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Elements & Macromolecules in Organisms
 Name: Period: Date: Elements & Macromolecules in Organisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight.
Name: Period: Date: Elements & Macromolecules in Organisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight.
CHAPTER 2- BIOCHEMISTRY I. WATER (VERY IMPORTANT TO LIVING ORGANISMS) A. POLAR COMPOUND- 10/4/ H O KENNEDY BIOLOGY 1AB
 CHAPTER 2- BIOCHEMISTRY KENNEDY BIOLOGY 1AB I. WATER (VERY IMPORTANT TO LIVING ORGANISMS) WATER S UNIQUE PROPERTIES MAKE IT ESSENTIAL FOR ALL LIFE FUNCTIONS IT IS POLAR, AND HAS BOTH ADHESIVE AND COHESIVE
CHAPTER 2- BIOCHEMISTRY KENNEDY BIOLOGY 1AB I. WATER (VERY IMPORTANT TO LIVING ORGANISMS) WATER S UNIQUE PROPERTIES MAKE IT ESSENTIAL FOR ALL LIFE FUNCTIONS IT IS POLAR, AND HAS BOTH ADHESIVE AND COHESIVE
Kinetics analysis of β-fructofuranosidase enzyme. 1-Effect of Time Incubation On The Rate Of An Enzymatic Reaction
 Kinetics analysis of β-fructofuranosidase enzyme 1-Effect of Time Incubation On The Rate Of An Enzymatic Reaction Enzyme kinetics It is the study of the chemical reactions that are catalyzed by enzymes.
Kinetics analysis of β-fructofuranosidase enzyme 1-Effect of Time Incubation On The Rate Of An Enzymatic Reaction Enzyme kinetics It is the study of the chemical reactions that are catalyzed by enzymes.
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors
 Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
The Structure and Function of Biomolecules
 The Structure and Function of Biomolecules The student is expected to: 9A compare the structures and functions of different types of biomolecules, including carbohydrates, lipids, proteins, and nucleic
The Structure and Function of Biomolecules The student is expected to: 9A compare the structures and functions of different types of biomolecules, including carbohydrates, lipids, proteins, and nucleic
2 3 Carbon Compounds (Macromolecules)
 2 3 Carbon Compounds (Macromolecules) Slide 1 of 37 Organic Chemistry Organic chemistry is the study of all compounds that contain bonds between carbon atoms. Slide 2 of 37 Carbon Living organisms are
2 3 Carbon Compounds (Macromolecules) Slide 1 of 37 Organic Chemistry Organic chemistry is the study of all compounds that contain bonds between carbon atoms. Slide 2 of 37 Carbon Living organisms are
UNIVERSITY OF GUELPH CHEM 4540 ENZYMOLOGY Winter 2005 Quiz #2: March 24, 2005, 11:30 12:50 Instructor: Prof R. Merrill ANSWERS
 UNIVERSITY F GUELPH CHEM 4540 ENZYMLGY Winter 2005 Quiz #2: March 24, 2005, 11:30 12:50 Instructor: Prof R. Merrill ANSWERS Instructions: Time allowed = 80 minutes. Total marks = 30. This quiz represents
UNIVERSITY F GUELPH CHEM 4540 ENZYMLGY Winter 2005 Quiz #2: March 24, 2005, 11:30 12:50 Instructor: Prof R. Merrill ANSWERS Instructions: Time allowed = 80 minutes. Total marks = 30. This quiz represents
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information.
 Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Carbon. p Has four valence electrons p Can bond with many elements p Can bond to other carbon atoms
 Organic Compounds Carbon p Has four valence electrons p Can bond with many elements p Can bond to other carbon atoms n Gives carbon the ability to form chains that are almost unlimited in length. p Organic
Organic Compounds Carbon p Has four valence electrons p Can bond with many elements p Can bond to other carbon atoms n Gives carbon the ability to form chains that are almost unlimited in length. p Organic
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Biology. Slide 1 of 37. End Show. Copyright Pearson Prentice Hall
 Biology 1 of 37 2 of 37 The Chemistry of Carbon The Chemistry of Carbon Organic chemistry is the study of all compounds that contain bonds between carbon atoms. 3 of 37 Macromolecules Macromolecules Macromolecules
Biology 1 of 37 2 of 37 The Chemistry of Carbon The Chemistry of Carbon Organic chemistry is the study of all compounds that contain bonds between carbon atoms. 3 of 37 Macromolecules Macromolecules Macromolecules
Introduction to Protein Structure Collection
 Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Β-FRUCTOFURANOSIDASE ENZYME
 KINETICS ANALYSIS OF Β-FRUCTOFURANOSIDASE ENZYME 2-The effects of enzyme concentration on the rate of an enzyme catalyzed reaction. Systematic names and numbers β-fructofuranosidase (EC 3.2.1.26) Reactions
KINETICS ANALYSIS OF Β-FRUCTOFURANOSIDASE ENZYME 2-The effects of enzyme concentration on the rate of an enzyme catalyzed reaction. Systematic names and numbers β-fructofuranosidase (EC 3.2.1.26) Reactions
Chapter 6. X-ray structure analysis of D30N tethered HIV-1 protease. dimer/saquinavir complex
 Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
SUPPLEMENTARY INFORMATION. Supplementary Figures
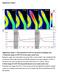 SUPPLEMENTARY INFORMATION Supplementary Figures Supplementary Figure 1: Characterization of CerTN-L15 expressed in Arabidopsis roots. a. Ratiometric images of CerTN-L15 in roots under osmotic stress Ratiometric
SUPPLEMENTARY INFORMATION Supplementary Figures Supplementary Figure 1: Characterization of CerTN-L15 expressed in Arabidopsis roots. a. Ratiometric images of CerTN-L15 in roots under osmotic stress Ratiometric
Organic Molecules. 1. The structural formulas shown represent certain organic compounds found in living cells.
 Name: ate: 1. The structural formulas shown represent certain organic compounds found in living cells. 1. (1) () (3) Which formula represents a monosaccharide? (4) (5). 1.. 3. 5. Which formula represents
Name: ate: 1. The structural formulas shown represent certain organic compounds found in living cells. 1. (1) () (3) Which formula represents a monosaccharide? (4) (5). 1.. 3. 5. Which formula represents
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69
 Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69 germline gene sequence. 2G1 and 8M2 acquired 14 and 18 mutations from the germline gene sequence, respectively. In 2G1,
Supplementary Figure 1. Heavy chain sequences of 2G1 and 8M2 aligned with V H 1-69 germline gene sequence. 2G1 and 8M2 acquired 14 and 18 mutations from the germline gene sequence, respectively. In 2G1,
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B
 3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
KDM2A. Reactions. containing. Reactions
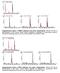 Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
The effects of ph on Type VII-NA Bovine Intestinal Mucosal Alkaline Phosphatase Activity
 The effects of ph on Type VII-NA Bovine Intestinal Mucosal Alkaline Phosphatase Activity ANDREW FLYNN, DYLAN JONES, ERIC MAN, STEPHEN SHIPMAN, AND SHERMAN TUNG Department of Microbiology and Immunology,
The effects of ph on Type VII-NA Bovine Intestinal Mucosal Alkaline Phosphatase Activity ANDREW FLYNN, DYLAN JONES, ERIC MAN, STEPHEN SHIPMAN, AND SHERMAN TUNG Department of Microbiology and Immunology,
Carbon. Has four valence electrons Can bond with many elements. Can bond to other carbon atoms. Hydrogen, Oxygen, Phosphorus, Sulfur, and Nitrogen
 Organic Compounds Carbon Has four valence electrons Can bond with many elements Hydrogen, Oxygen, Phosphorus, Sulfur, and Nitrogen Can bond to other carbon atoms Gives carbon the ability to form chains
Organic Compounds Carbon Has four valence electrons Can bond with many elements Hydrogen, Oxygen, Phosphorus, Sulfur, and Nitrogen Can bond to other carbon atoms Gives carbon the ability to form chains
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Student Manual. Background STUDENT MANUAL BACKGROUND. Enzymes
 Background Enzymes Enzymes are typically proteins (some nucleic acids have also been found to be enzymes) that act as catalysts, speeding up chemical reactions that would take far too long to occur on
Background Enzymes Enzymes are typically proteins (some nucleic acids have also been found to be enzymes) that act as catalysts, speeding up chemical reactions that would take far too long to occur on
Enzymes - Exercise 3 - Rockville
 Enzymes - Exercise 3 - Rockville Objectives -Understand the function of an enzyme. -Know what the substrate, enzyme, and the product of the reaction for this lab. -Understand how at various environments
Enzymes - Exercise 3 - Rockville Objectives -Understand the function of an enzyme. -Know what the substrate, enzyme, and the product of the reaction for this lab. -Understand how at various environments
Amprenavir complexes with HIV-1 protease and its drug-resistant mutants altering hydrophobic clusters
 Amprenavir complexes with HIV-1 protease and its drug-resistant mutants altering hydrophobic clusters Chen-Hsiang Shen 1, Yuan-Fang Wang 1, Andrey Y. Kovalevsky 1, *, Robert W. Harrison 1,2 and Irene T.
Amprenavir complexes with HIV-1 protease and its drug-resistant mutants altering hydrophobic clusters Chen-Hsiang Shen 1, Yuan-Fang Wang 1, Andrey Y. Kovalevsky 1, *, Robert W. Harrison 1,2 and Irene T.
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES
 SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
Elements & Macromolecules in Organisms
 Elements & Macromolecules in rganisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight. All compounds can be
Elements & Macromolecules in rganisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight. All compounds can be
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP
 ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
Bioanalytical chemistry. 2. Enzymes as analytical reagents
 13 Bioanalytical chemistry 2. Enzymes as analytical reagents Suggested reading: Sections 3.1 to 3.5.1.3 of Mikkelsen and Cortón, Bioanalytical Chemistry rimary Source Material Chapter 8 of Biochemistry:
13 Bioanalytical chemistry 2. Enzymes as analytical reagents Suggested reading: Sections 3.1 to 3.5.1.3 of Mikkelsen and Cortón, Bioanalytical Chemistry rimary Source Material Chapter 8 of Biochemistry:
SUPPLEMENTARY INFORMATION
 Supplementary Information Supplementary Figure 1. The serine paradox. Nature-designed AlaRSs pin-down the -NH3 + group of alanine with an acidic Asp that is critical for binding and activation of alanine.
Supplementary Information Supplementary Figure 1. The serine paradox. Nature-designed AlaRSs pin-down the -NH3 + group of alanine with an acidic Asp that is critical for binding and activation of alanine.
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Supplementary Information
 Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Molecular Docking Study of Beta-Glucosidase with Cellobiose, Cellotetraose and Cellotetriose
 www.bioinformation.net Hypothesis Volume 9(16) Molecular Docking Study of Beta-Glucosidase with Cellobiose, Cellotetraose and Cellotetriose Nurul Bahiyah Ahmad Khairudin* & Nur Shima Fadhilah Mazlan Malaysia
www.bioinformation.net Hypothesis Volume 9(16) Molecular Docking Study of Beta-Glucosidase with Cellobiose, Cellotetraose and Cellotetriose Nurul Bahiyah Ahmad Khairudin* & Nur Shima Fadhilah Mazlan Malaysia
Carbohydrates, Lipids, Proteins, and Nucleic Acids
 Carbohydrates, Lipids, Proteins, and Nucleic Acids Is it made of carbohydrates? Organic compounds composed of carbon, hydrogen, and oxygen in a 1:2:1 ratio. A carbohydrate with 6 carbon atoms would have
Carbohydrates, Lipids, Proteins, and Nucleic Acids Is it made of carbohydrates? Organic compounds composed of carbon, hydrogen, and oxygen in a 1:2:1 ratio. A carbohydrate with 6 carbon atoms would have
Project Manual Bio3055. Cholesterol Homeostasis: HMG-CoA Reductase
 Project Manual Bio3055 Cholesterol Homeostasis: HMG-CoA Reductase Bednarski 2003 Funded by HHMI Cholesterol Homeostasis: HMG-CoA Reductase Introduction: HMG-CoA Reductase is an enzyme in the cholesterol
Project Manual Bio3055 Cholesterol Homeostasis: HMG-CoA Reductase Bednarski 2003 Funded by HHMI Cholesterol Homeostasis: HMG-CoA Reductase Introduction: HMG-CoA Reductase is an enzyme in the cholesterol
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Macromolecules Cut & Paste
 Macromolecules Cut & Paste Adapted from http://mrswords.weebly.com/uploads/1/5/2/4/15244382/ch_6-3_life_molecules_cut-out_lab.pdf INTRODUCTION Many of the molecules in living cells are so large that they
Macromolecules Cut & Paste Adapted from http://mrswords.weebly.com/uploads/1/5/2/4/15244382/ch_6-3_life_molecules_cut-out_lab.pdf INTRODUCTION Many of the molecules in living cells are so large that they
C 6 H 12 O 6 + 6O 2 6CO 2 + 6H 2 O
 Sample Questions for the Individual Round Exam 1. Glucose is the most basic sugar involved in human metabolism. Its structure is provided below: a. The overall reaction of glucose metabolism is given below.
Sample Questions for the Individual Round Exam 1. Glucose is the most basic sugar involved in human metabolism. Its structure is provided below: a. The overall reaction of glucose metabolism is given below.
High resolution structural evidence suggests the Sarcoplasmic Reticulum forms microdomains with Acidic Stores (lyososomes) in the heart.
 High resolution structural evidence suggests the Sarcoplasmic Reticulum forms microdomains with Acidic Stores (lyososomes) in the heart. Daniel Aston, Rebecca A. Capel, Kerrie L. Ford, Helen C. Christian,
High resolution structural evidence suggests the Sarcoplasmic Reticulum forms microdomains with Acidic Stores (lyososomes) in the heart. Daniel Aston, Rebecca A. Capel, Kerrie L. Ford, Helen C. Christian,
Supplementary Files S1 Isolation of Monocytes S2 Haemolysis study Reagents Procedure S3 Cytotoxicity studies Trypan blue dye exclusion method
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Supplementary Files S1 Isolation of Monocytes A 3 ml volume of Histopaque 1083 solution was
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Supplementary Files S1 Isolation of Monocytes A 3 ml volume of Histopaque 1083 solution was
WHAT IS A LIPID? OBJECTIVE The objective of this worksheet is to understand the structure and function of lipids
 WHAT IS A LIPID? OBJECTIVE The objective of this worksheet is to understand the structure and function of lipids PART A: Understanding Lipids Lipids are more commonly known as fats and include triglycerides,
WHAT IS A LIPID? OBJECTIVE The objective of this worksheet is to understand the structure and function of lipids PART A: Understanding Lipids Lipids are more commonly known as fats and include triglycerides,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
B i o c h e m i s t r y N o t e s
 14 P a g e Carbon Hydrogen Nitrogen Oxygen Phosphorus Sulfur ~Major ~Found in all ~Found in most ~Found in all component of all organic organic molecules. molecules. ~Major structural atom in all organic
14 P a g e Carbon Hydrogen Nitrogen Oxygen Phosphorus Sulfur ~Major ~Found in all ~Found in most ~Found in all component of all organic organic molecules. molecules. ~Major structural atom in all organic
(5) 1. List five unusual properties of water resulting from its hydrogen bonded structure
 BCH 4053 June 1, 2001 Points HOUR TEST 1 NAME (5) 1. List five unusual properties of water resulting from its hydrogen bonded structure. Page Points 1 2 3 4 5 Total (5) 2. Draw a diagram to show how water
BCH 4053 June 1, 2001 Points HOUR TEST 1 NAME (5) 1. List five unusual properties of water resulting from its hydrogen bonded structure. Page Points 1 2 3 4 5 Total (5) 2. Draw a diagram to show how water
2. (12 pts) Given the following metabolic pathway (as it occurs in the cell):
 Answer Sheet 1 (Gold) 1. (1 pt) Write your exam ID (A) in the blank at the upper right of your answer sheet. 2. (12 pts) Given the following metabolic pathway (as it occurs in the cell): a. Would you expect
Answer Sheet 1 (Gold) 1. (1 pt) Write your exam ID (A) in the blank at the upper right of your answer sheet. 2. (12 pts) Given the following metabolic pathway (as it occurs in the cell): a. Would you expect
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Previous Class. Today. Term test I discussions. Detection of enzymatic intermediates: chymotrypsin mechanism
 Term test I discussions Previous Class Today Detection of enzymatic intermediates: chymotrypsin mechanism Mechanistic Understanding of Enzymemediated Reactions Ultimate goals: Identification of the intermediates,
Term test I discussions Previous Class Today Detection of enzymatic intermediates: chymotrypsin mechanism Mechanistic Understanding of Enzymemediated Reactions Ultimate goals: Identification of the intermediates,
Supplementary Information
 Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction
 Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Mechanisms of Enzymes
 Mechanisms of Enzymes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy How enzymes work * Chemical reactions have an energy
Mechanisms of Enzymes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy How enzymes work * Chemical reactions have an energy
Experiment 9. NATURE OF α-amylase ACTIVITY ON STARCH
 Experiment 9 NATURE OF α-amylase ACTIVITY ON STARC In Experiment 1 we described the action of α-amylase on starch as that of catalyzing the hydrolysis of α-1,4-glucosidic bonds at random in the interior
Experiment 9 NATURE OF α-amylase ACTIVITY ON STARC In Experiment 1 we described the action of α-amylase on starch as that of catalyzing the hydrolysis of α-1,4-glucosidic bonds at random in the interior
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Supporting Information. Evolution of atomically precise silver clusters to superlattices
 Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2012. Supporting Information for Part. Part. Sys. Charact., DOI: 10.1002/ppsc.((please add manuscript number)) Evolution of atomically
Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2012. Supporting Information for Part. Part. Sys. Charact., DOI: 10.1002/ppsc.((please add manuscript number)) Evolution of atomically
Gold nanocrystals at DPPC bilayer. Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad
 Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.80 Protein-Inorganic Hybrid Nanoflowers Jun Ge, Jiandu Lei, and Richard N. Zare Supporting Online Material Materials Proteins including albumin from bovine
SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.80 Protein-Inorganic Hybrid Nanoflowers Jun Ge, Jiandu Lei, and Richard N. Zare Supporting Online Material Materials Proteins including albumin from bovine
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A
 Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
SUPPLEMENTAL MATERIAL. UNC119 is required for G protein trafficking in sensory neurons
 1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
6. The catalytic mechanism of arylsulfatase A and its theoretical investigation
 6. The catalytic mechanism of arylsulfatase A and its theoretical investigation When the crystal structure of arylsulfatase A was solved, a remarkable structural analogy to another hydrolytic enzyme, the
6. The catalytic mechanism of arylsulfatase A and its theoretical investigation When the crystal structure of arylsulfatase A was solved, a remarkable structural analogy to another hydrolytic enzyme, the
