Chemically targeting the PI3K family
|
|
|
- Albert Hodge
- 5 years ago
- Views:
Transcription
1 3rd Focused Meeting on PI3K Signalling and Disease 245 Chemically targeting the PI3K family Z.A. Knight and K.M. Shokat 1 Howard Hughes Medical Institute, Department of Cellular and Molecular Pharmacology, University of California, San Francisco, CA, U.S.A. Abstract PI3K (phosphoinositide 3-kinase) is a key regulator of cell growth, metabolism and survival. The frequent activation of the PI3K pathway in cancer has stimulated widespread interest in identifying potent and selective inhibitors of PI3K isoforms. The present paper highlights recent progress in identifying such molecules and the challenges that remain for efforts to pharmacologically target the PI3K family. Introduction PI3K (phosphoinositide 3-kinase) is activated by receptor tyrosine kinases and other cell-surface receptors to synthesize the lipid second messenger PIP 3 (phosphatidylinositol-3,4,5- trisphosphate) [1]. PIP 3, in turn, acts as a docking site at the plasma membrane that recruits and activates proteins containing phospholipid-binding domains. These downstream PI3K effectors include (i) protein kinases that promote cell growth, survival and proliferation [such as Akt, PDK1 (phosphoinositide-dependent kinase 1) and the Tec family kinases]; (ii) GAPs (GTPase-activating proteins) and GEFs (guanine nucleotide-exchange factors) that regulate GTPases mediating cell motility and membrane trafficking; and (iii) scaffolding proteins that nucleate the assembly of key signalling complexes. In this way, PI3K utilizes a single second messenger to link extracellular hormonal cues to intracellular signalling proteins that control diverse cellular processes. The PI3K family consists of 15 proteins that share sequence homology within their kinase domains but have distinct substrate specificities and modes of regulation [1]. The most well characterized members of this family are the four class I PI3Ks that link PI3K activity to receptor tyrosine kinases (p110α, p110β and p110δ) or G-protein-coupled receptors (p110γ ). These proteins synthesize exclusively PIP 3 and mediate most of the known signalling functions attributed to PI3K. In addition to these enzymes, class II and III PI3Ks have been described that also generate 3-phosphorylated inositol lipids, although the physiological roles of these proteins remain elusive. The PIKKs (PI3K-related protein kinases) are a subfamily of serine/threonine kinases that share sequence homology with PI3K but phosphorylate proteins rather than lipids. The PIKKs include key kinases that monitor genomic integrity [ATM (ataxia telangiectasia mutated), ATR (ATM- and Rad3-related), Smg-1 and DNA-PK (DNAdependent protein kinase)] as well as mtor (mammalian Key words: cancer, mammalian target of rapamycin (mtor), p110, phosphatidylinositol, phosphoinositide 3-kinase (PI3K). Abbreviations used: ATM, ataxia telangiectasia mutated; ATR, ATM- and Rad3-related; DNA-PK, DNA-dependent protein kinase; mtor, mammalian target of rapamycin; PI3K, phosphoinositide 3-kinase; PIKK, PI3K-related protein kinase; PIP 3, phosphatidylinositol-3,4,5-trisphosphate. 1 To whom correspondence should be addressed ( shokat@cmp.ucsf.edu). target of rapamycin), a kinase that integrates nutritional and growth factor signals in order to control protein synthesis and cell growth. Recent interest in PI3K signalling has been fuelled by evidence that the PI3K pathway is among the most commonly activated signalling pathways in cancer. Several observations have been critical in establishing this connection. First, the p110α isoform of PI3K is activated by mutation at high frequency in a range of primary tumours [2]. Indeed, a recent analysis reported that p110α mutations occur at a cumulative frequency of 15% across all cancer types surveyed to date [3], suggesting that p110α is likely to be the most commonly mutated kinase in the human genome. Secondly, the phosphatase PTEN (phosphatase and tensin homologue deleted on chromosome 10), which antagonizes PI3K signalling by dephosphorylating PIP 3, is a well-characterized tumour suppressor that is frequently inactivated by mutation, gene deletion or epigenetic silencing [4]. In addition, PI3K is allosterically activated by the oncogene Ras [5], and many tyrosine kinases that activate PI3K are themselves the target of mutations or amplification in cancer. Together, these observations reveal a nexus of genetic alterations in cancer that each stimulate PI3K signalling, suggesting that PI3K activation is likely to be an essential step in tumorigenesis. The widespread activation of PI3K in cancer has stimulated intense interest in developing drugs that target the PI3K pathway. Nonetheless, significant uncertainties surround efforts to therapeutically target these kinases. Unlike protein kinases, class I PI3Ks all phosphorylate the same substrate in order to generate a single lipid product, PIP 3. For this reason, the kinase activity of individual PI3K isoforms cannot be readily distinguished in cells. As class I PI3Ks are essential for a range of normal physiological processes, including glucose homoeostasis and the immune response, broad spectrum PI3K inhibition is likely to be associated with significant toxicity. Isoform-selective PI3K inhibitors may overcome these difficulties, but in many cases it has been difficult to define the isoforms that mediate PI3K s critical physiological functions [6]. Moreover, the highly conserved ATP-binding site of the class I PI3Ks presents a significant challenge for medicinal chemists seeking to identify molecules that display high selectivity between these isoforms. This review briefly
2 246 Biochemical Society Transactions (2007) Volume 35, part 2 summarizes recent progress in developing isoform-selective PI3K inhibitors as well as challenges that remain for efforts to pharmacologically target this enzyme family. The prototype PI3K inhibitors wortmannin and LY The discovery of the pan-specific PI3K inhibitors wortmannin and LY was a critical early event that enabled the rapid exploration of PI3K signalling. Wortmannin is a fungal natural product that was originally described as a potent inhibitor of the respiratory burst in neutrophils and monocytes [7]. PI3K was subsequently identified as the molecular target of wortmannin [8], and the mechanism of inhibition was shown to involve covalent attack of Lys 833 within the ATP-binding site of PI3K at an electrophilic site on the natural product [9]. Shortly after the identification of PI3K as the target of wortmannin, the synthetic PI3K inhibitor LY was reported [10]. The discovery of LY demonstrated the feasibility of inhibiting PI3K with a reversible, drug-like small molecule at a time (1994) when few kinase inhibitors were known. Subsequent crystal structures of LY and wortmannin bound to p110γ confirmed that both molecules occupy the ATP-binding site [11]. Despite the limitations of these inhibitors (wortmannin is a reactive electrophile, whereas LY has micromolar affinity), the early availability of these molecules played an essential role in shaping our current understanding of PI3K signalling. The development of isoform-specific PI3K inhibitors LY and wortmannin show little selectivity within the PI3K family. For this reason, these molecules cannot be used to probe signalling by specific PI3Ks, and their structures provide little guidance for the design of selective inhibitors. Nonetheless, growing appreciation of the therapeutic potential of PI3K inhibitors has led to significant efforts in recent years within the pharmaceutical industry to identify new PI3K inhibitors with enhanced potency, selectivity and pharmacological properties. Recently, the first of these molecules has been reported. IC87114 was the first isoform-selective PI3K inhibitor to be described [12] (Figure 1). This quinazolinone purine inhibits p110δ at mid-nanomolar concentrations and shows fold selectivity between p110δ and theother class I PI3Ks (p110α, p110β and p110γ ). The selectivity of IC87114 is remarkable given that the residues that line the ATPbinding pocket of the class I PI3Ks are highly conserved. The co-crystal structure of an IC87114 analogue (PIK-39) bound to p110γ was recently reported, and this structure suggests a mechanism for the selectivity of PIK-39 [13]. This structure revealed that PIK-39 binding induces a conformational change in the kinase whereby Met 804, which forms the upper surface of the ATP-binding pocket, shifts downward to form a novel pocket at the entrance to the kinase active site. This induced pocket buries most of the hydrophobic surface area of the PIK-39 quinazolinone ring system and therefore is likely to be critical for the high-affinity binding of this compound to p110δ. The importance of this conformational shift for PIK-39 binding was tested by introducing mutations into p110δ that replace Met 804 with β-branched amino acids (valine or isoleucine) that were predicted to restrict formation of the induced pocket [13]. Consistent with this prediction, these mutations impaired the binding of IC87114 and PIK- 39 to p110δ, but had no effect on the binding of three other chemotypes of PI3K inhibitors that do not access the induced pocket formed by Met 804. Together, these results support a model in which quinazolinone purines such as IC87114 and PIK-39 selectively inhibit p110δ by targeting a residue (Met 804 ) that shows differential conformational mobility among PI3K isoforms. Modelling suggests that additional chemotypes of PI3K inhibitors may achieve their selectivity via similar interactions with this inducible pocket, [13] but this hypothesis remains to be crystallographically verified. Finally, it is worth noting that this mechanism of inhibitor binding shares similarities with protein kinase inhibitors such as imatinib, which distinguish between closely related kinases such as Abl and Src by selectively stabilizing differentially populated conformational states of these proteins [14]. In the three years since the description of IC87114, a series of small molecules have been reported that show varying degrees of selectivity for p110α [13,15,16], p110β [17,18], p110γ [19,20], DNA-PK [21 23] and ATM [24,25] (Figure 1). All of these compounds are drug-like, reversible inhibitors that are ATP-competitive and possess nanomolar affinity against their primary target. The extended biochemical selectivity of representatives from several of these compound classes has been determined [13] and co-crystal structures of several compounds bound to p110γ have been reported [13,19]. While no compound shows absolute selectivity for its primary target, impressive 100-fold selectivity has been achieved in a several cases. A detailed discussion of each chemotype is beyond the scope of the present paper, but several themes have emerged from this analysis. First, PI3K inhibitors share many features in common with wellcharacterized inhibitors of protein kinases. Like proteinkinase inhibitors, all known PI3K inhibitors satisfy the hydrogen bond that is made between the N1 of adenine in ATP and the hinge region of the kinase [11,13,19]. In protein kinases, the size of a single residue in the interior of the ATP-binding pocket, termed the gatekeeper, has been shown to control the potency of a wide range of inhibitors by regulating their access to a deeper hydrophobic pocket [26,27]. Co-crystal structures of potent PI3K inhibitors bound to p110γ revealed that these molecules also access a deeper hydrophobic pocket in lipid kinases that is essential for their high-affinity binding [13]. However, a slight shift in the orientation of the gatekeeper residue in PI3Ks relative to protein kinases appears to make this pocket accessible throughout the family, irrespective of the size of the gatekeeper residue [28]. Interestingly, despite these structural similarities between the ATP-binding sites of protein and lipid kinases,
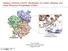
3 3rd Focused Meeting on PI3K Signalling and Disease 247 Figure 1 Chemical structures of PI3K inhibitors (A) Structure of LY294002, indicating hydrogen bonds to Val 882 and Lys 833 of p110γ.(b) Structures of arylmorpholine PI3K inhibitors. (C) Structures of PI3K inhibitors from other chemical classes. potent PI3K inhibitors from diverse chemotypes generally do not inhibit a wide range of protein kinases. This presumably reflects the fact that, despite their superficial similarities, PI3Ks and protein kinases share no sequence homology and therefore present unique surfaces within their ATP-binding sites that are recognized by high-affinity inhibitors. Analysis of the selectivity profiles of chemically diverse PI3K inhibitors has revealed trends among molecules that target this enzyme family. In particular, certain PI3K family members show similar patterns of sensitivity to existing small molecule inhibitors. We have referred to these pairs of kinases as pharmalogs to reflect the fact that they show homology in their sensitivity to chemical inhibitors [13]. For example, among the class I PI3Ks, inhibitors of p110β often potently inhibit p110δ, whereas inhibitorsof p110α often potentlyinhibit p110γ. In a similar fashion, inhibitors of class I PI3Ks
4 248 Biochemical Society Transactions (2007) Volume 35, part 2 frequently inhibit DNA-PK potently, even though the same molecules show little or no activity against the PIKKs that are most closely related to DNA-PK in amino acid sequence (such as mtor, ATM and ATR). The latter example is particularly striking because sequence alignments do not suggest obvious similarities between the class I PI3Ks and DNA-PK in the residues that line their ATP-binding pockets. This suggests that these kinases share a cryptic similarity in the three-dimensional structures of their active sites that is not obvious from sequence analysis. Alternatively, DNA-PK may be generally more druggable than other members of the PI3K family and therefore shows enhanced sensitivity to inhibitors that target any PI3K family member. Distinguishing between these possibilities will require the detailed biochemical characterization of chemically diverse inhibitors that target PI3K family members other than the class I PI3Ks. Comparison of the chemical structures of PI3K inhibitors reveals that certain privileged scaffolds are frequently found in molecules targeting this enzyme family. The most common of these is the arylmorpholine, which forms the core structure of the original PI3K inhibitor LY (Figure 1A). The crystal structure of LY bound to p110γ revealed that the oxygen atom of morpholine satisfies the hinge region hydrogen bond to Val 882 [11]. Furthermore, the morpholine ring makes extensive hydrophobic interactions with residues that form the top and bottom of the ATP-binding pocket. These extended interactions densely pack against the morpholine ring, explaining why subtle alterations in the morpholine structure result in a dramatic loss of affinity [10,29]. The versatility of this chemical class is highlighted by the diversity of arylmorpholines that have recently been described to inhibit p110α, p110β/δ, mtor, ATMor DNA-PK (Figure 1B). Questions for the development of PI3K inhibitors The past three years have witnessed an explosion of information about small molecules that target the PI3K family. The original inhibitors LY and wortmannin have now been joined by at least 15 new chemical classes of inhibitors, many of which have crystallographically defined binding modes and biochemically enumerated target selectivities. As a result of extensive medicinal chemistry efforts within the pharmaceutical industry, relatively selective inhibitors now exist for each of the class I PI3Ks as well as the PIKKs ATM and DNA-PK. Despite these advances, our understanding of the rules that determine the selectivity and potency of PI3K inhibitors remains in its infancy. In this regard, protein kinase inhibitors, whose clinical development has preceded lipid kinase inhibitors by a decade, may provide useful guidance. For example, a key discovery in the development of protein kinase inhibitors was the finding that protein kinases shuttle between multiple conformational states, and that highly selective kinase inhibitors such as imatinib act by selectively trapping these states [30]. For many protein kinases, the equilibrium between these conformations is believed to be controlled by phosphorylation of key residues in the activation loop, such that these states correspond to natural active or inactive conformations of the kinase. By contrast, PI3K is not known to undergo activation loop phosphorylation, and the conformational changes that accompany PI3K activation are largely unknown. Do current PI3K inhibitors sense the activation status of these enzymes? If not, is it possible to identify molecules that stabilize the inactive conformation of a lipid kinase, in a manner analogous to protein kinase inhibitors that bind to protein kinases in a DFG out orientation? The crystal structure of PIK-39 provides the first evidence that PI3K inhibitors can exploit the conformational flexibility of a lipid kinase to achieve highly selective binding, but the broader significance of this Met 804 down conformation, for the binding of other classes of inhibitors or for normal PI3K regulation, is unknown. The biochemical and structural exploration of these questions is likely to provide important insights into strategies for selectively targeting the PI3K family. References 1 Vanhaesebroeck, B., Leevers, S.J., Ahmadi, K., Timms, J., Katso, R., Driscoll, P.C., Woscholski, R., Parker, P.J. and Waterfield, M.D. (2001) Annu. Rev. Biochem. 70, Samuels, Y., Wang, Z., Bardelli, A., Silliman, N., Ptak, J., Szabo, S., Yan, H., Gazdar, A., Powell, S.M., Riggins, G.J. et al. (2004) Science 304, Karakas, B., Bachman, K.E. and Park, B.H. (2006) Br. J. Cancer 94, Cantley, L.C. and Neel, B.G. (1999) Proc. Natl. Acad. Sci. U.S.A. 96, Rodriguez-Viciana, P., Warne, P.H., Dhand, R., Vanhaesebroeck, B., Gout, I., Fry, M.J., Waterfield, M.D. and Downward, J. (1994) Nature 370, Vanhaesebroeck, B., Rohn, J.L. and Waterfield, M.D. (2004) Cell 118, Baggiolini, M., Dewald, B., Schnyder, J., Ruch, W., Cooper, P.H. and Payne, T.G. (1987) Exp. Cell Res. 169, Arcaro, A. and Wymann, M.P. (1993) Biochem. J. 296, Wymann, M.P., Bulgarelli-Leva, G., Zvelebil, M.J., Pirola, L., Vanhaesebroeck, B., Waterfield, M.D. and Panayotou, G. (1996) Mol. Cell. Biol. 16, Vlahos, C.J., Matter, W.F., Hui, K.Y. and Brown, R.F. (1994) J. Biol. Chem. 269, Walker, E.H., Pacold, M.E., Perisic, O., Stephens, L., Hawkins, P.T., Wymann, M.P. and Williams, R.L. (2000) Mol. Cell 6, Sadhu, C., Masinovsky, B., Dick, K., Sowell, C.G. and Staunton, D.E. (2003) J. Immunol. 170, Knight, Z.A., Gonzalez, B., Feldman, M.E., Zunder, E.R., Goldenberg, D.D., Williams, O., Loewith, R., Stokoe, D., Balla, A., Toth, B. et al. (2006) Cell 125, Schindler, T., Bornmann, W., Pellicena, P., Miller, W.T., Clarkson, B. and Kuriyan, J. (2000) Science 289, Hayakawa, M., Kaizawa, H., Kawaguchi, K.I., Ishikawa, N., Koizumi, T., Ohishi, T., Yamano, M., Okada, M., Ohta, M., Tsukamoto, S.I. et al. (2007) Bioorg. Med. Chem. 15, Hayakawa, M., Kaizawa, H., Moritomo, H., Koizumi, T., Ohishi, T., Okada, M., Ohta, M., Tsukamoto, S., Parker, P., Workman, P. and Waterfield, M. (2006) Bioorg. Med. Chem. 14, Knight, Z.A., Chiang, G.G., Alaimo, P.J., Kenski, D.M., Ho, C.B., Coan, K., Abraham, R.T. and Shokat, K.M. (2004) Bioorg. Med. Chem. 12, Jackson, S.P., Schoenwaelder, S.M., Goncalves, I., Nesbitt, W.S., Yap, C.L., Wright, C.E., Kenche, V., Anderson, K.E., Dopheide, S.M., Yuan, Y. et al. (2005) Nat. Med. 11,
5 3rd Focused Meeting on PI3K Signalling and Disease Camps, M., Ruckle, T., Ji, H., Ardissone, V., Rintelen, F., Shaw, J., Ferrandi, C., Chabert, C., Gillieron, C., Francon, B. et al. (2005) Nat. Med. 11, Pomel, V., Klicic, J., Covini, D., Church, D.D., Shaw, J.P., Roulin, K., Burgat-Charvillon, F., Valognes, D., Camps, M., Chabert, C. et al. (2006) J. Med. Chem. 49, Griffin, R.J., Fontana, G., Golding, B.T., Guiard, S., Hardcastle, I.R., Leahy, J.J., Martin, N., Richardson, C., Rigoreau, L., Stockley, M. and Smith, G.C. (2005) J. Med. Chem. 48, Kashishian, A., Douangpanya, H., Clark, D., Schlachter, S.T., Eary, C.T., Schiro, J.G., Huang, H., Burgess, L.E., Kesicki, E.A. and Halbrook, J. (2003) Mol. Cancer Ther. 2, Leahy, J.J., Golding, B.T., Griffin, R.J., Hardcastle, I.R., Richardson, C., Rigoreau, L. and Smith, G.C. (2004) Bioorg. Med. Chem. Lett. 14, Lau, A., Swinbank, K.M., Ahmed, P.S., Taylor, D.L., Jackson, S.P., Smith, G.C. and O Connor, M.J. (2005) Nat. Cell Biol. 7, Hickson, I., Zhao, Y., Richardson, C.J., Green, S.J., Martin, N.M., Orr, A.I., Reaper, P.M., Jackson, S.P., Curtin, N.J. and Smith, G.C. (2004) Cancer Res. 64, Liu, Y., Bishop, A., Witucki, L., Kraybill, B., Shimizu, E., Tsien, J., Ubersax, J., Blethrow, J., Morgan, D.O. and Shokat, K.M. (1999) Chem. Biol. 6, Schindler, T., Sicheri, F., Pico, A., Gazit, A., Levitzki, A. and Kuriyan, J. (1999) Mol. Cell 3, Alaimo, P.J., Knight, Z.A. and Shokat, K.M. (2005) Bioorg. Med. Chem. 13, Hardcastle, I.R., Cockcroft, X., Curtin, N.J., El-Murr, M.D., Leahy, J.J., Stockley, M., Golding, B.T., Rigoreau, L., Richardson, C., Smith, G.C. and Griffin, R.J. (2005) J. Med. Chem. 48, Knight, Z.A. and Shokat, K.M. (2005) Chem. Biol. 12, Received 23 October 2006
Available Online through
 ISS: 0975-766X CDE: IJPTFI Available nline through Research Article www.ijptonline.com RELATISHIP BETWEE SCRIG FUCTI AD THE IC 50 IHIBITRY F PI3K IHIBITRS Awantika Shrivastava*, Dr.K.Durga Prasad 1, Dr.
ISS: 0975-766X CDE: IJPTFI Available nline through Research Article www.ijptonline.com RELATISHIP BETWEE SCRIG FUCTI AD THE IC 50 IHIBITRY F PI3K IHIBITRS Awantika Shrivastava*, Dr.K.Durga Prasad 1, Dr.
Article. Reference. A pharmacological map of the PI3-K family defines a role for p110alpha in insulin signaling. KNIGHT, Zachary A, et al.
 Article A pharmacological map of the PI3-K family defines a role for p110alpha in insulin signaling KNIGHT, Zachary A, et al. Abstract Phosphoinositide 3-kinases (PI3-Ks) are an important emerging class
Article A pharmacological map of the PI3-K family defines a role for p110alpha in insulin signaling KNIGHT, Zachary A, et al. Abstract Phosphoinositide 3-kinases (PI3-Ks) are an important emerging class
The PI3K/AKT axis. Dr. Lucio Crinò Medical Oncology Division Azienda Ospedaliera-Perugia. Introduction
 The PI3K/AKT axis Dr. Lucio Crinò Medical Oncology Division Azienda Ospedaliera-Perugia Introduction Phosphoinositide 3-kinase (PI3K) pathway are a family of lipid kinases discovered in 1980s. They have
The PI3K/AKT axis Dr. Lucio Crinò Medical Oncology Division Azienda Ospedaliera-Perugia Introduction Phosphoinositide 3-kinase (PI3K) pathway are a family of lipid kinases discovered in 1980s. They have
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
A Pharmacological Map of the PI3-K Family Defines a Role for p110a in Insulin Signaling
 A Pharmacological Map of the PI3-K Family Defines a Role for p110a in Insulin Signaling Zachary A. Knight, 1 Beatriz Gonzalez, 4,7 Morri E. Feldman, 1 Eli R. Zunder, 1 David D. Goldenberg, 2 Olusegun Williams,
A Pharmacological Map of the PI3-K Family Defines a Role for p110a in Insulin Signaling Zachary A. Knight, 1 Beatriz Gonzalez, 4,7 Morri E. Feldman, 1 Eli R. Zunder, 1 David D. Goldenberg, 2 Olusegun Williams,
Signal Transduction Pathways. Part 2
 Signal Transduction Pathways Part 2 GPCRs G-protein coupled receptors > 700 GPCRs in humans Mediate responses to senses taste, smell, sight ~ 1000 GPCRs mediate sense of smell in mouse Half of all known
Signal Transduction Pathways Part 2 GPCRs G-protein coupled receptors > 700 GPCRs in humans Mediate responses to senses taste, smell, sight ~ 1000 GPCRs mediate sense of smell in mouse Half of all known
Enzymes Part III: regulation II. Dr. Mamoun Ahram Summer, 2017
 Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Signal Transduction: G-Protein Coupled Receptors
 Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
KEY CONCEPT QUESTIONS IN SIGNAL TRANSDUCTION
 Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
Research Topic Seminar
 Research Topic Seminar Dr. Claire Coleman The Chemistry and Biology of Wortmannin Claire Coleman @ Wipf Group 1 3/13/2004 ff white to pale yellow solid Hygroscopic/Light sensitive Small molecule natural
Research Topic Seminar Dr. Claire Coleman The Chemistry and Biology of Wortmannin Claire Coleman @ Wipf Group 1 3/13/2004 ff white to pale yellow solid Hygroscopic/Light sensitive Small molecule natural
G-Protein Signaling. Introduction to intracellular signaling. Dr. SARRAY Sameh, Ph.D
 G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor!
 Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Biol220 Cell Signalling Cyclic AMP the classical secondary messenger
 Biol220 Cell Signalling Cyclic AMP the classical secondary messenger The classical secondary messenger model of intracellular signalling A cell surface receptor binds the signal molecule (the primary
Biol220 Cell Signalling Cyclic AMP the classical secondary messenger The classical secondary messenger model of intracellular signalling A cell surface receptor binds the signal molecule (the primary
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Tala Saleh. Ahmad Attari. Mamoun Ahram
 23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
Chapter 15: Signal transduction
 Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy se
 A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
The elements of G protein-coupled receptor systems
 The elements of G protein-coupled receptor systems Prostaglandines Sphingosine 1-phosphate a receptor that contains 7 membrane-spanning domains a coupled trimeric G protein which functions as a switch
The elements of G protein-coupled receptor systems Prostaglandines Sphingosine 1-phosphate a receptor that contains 7 membrane-spanning domains a coupled trimeric G protein which functions as a switch
Propagation of the Signal
 OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Lecture: CHAPTER 13 Signal Transduction Pathways
 Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
Validation & Assay Performance Summary
 Validation & Assay Performance Summary CellSensor DBE-bla MDA-MB-468 Cell Line Cat. no. K1814 Pathway Description The phosphatidylinositol-3-kinase (PI3K) signaling cascade is essential for cell growth
Validation & Assay Performance Summary CellSensor DBE-bla MDA-MB-468 Cell Line Cat. no. K1814 Pathway Description The phosphatidylinositol-3-kinase (PI3K) signaling cascade is essential for cell growth
Principles of Genetics and Molecular Biology
 Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Vets 111/Biov 111 Cell Signalling-2. Secondary messengers the cyclic AMP intracellular signalling system
 Vets 111/Biov 111 Cell Signalling-2 Secondary messengers the cyclic AMP intracellular signalling system The classical secondary messenger model of intracellular signalling A cell surface receptor binds
Vets 111/Biov 111 Cell Signalling-2 Secondary messengers the cyclic AMP intracellular signalling system The classical secondary messenger model of intracellular signalling A cell surface receptor binds
Signaling. Dr. Sujata Persad Katz Group Centre for Pharmacy & Health research
 Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Regulation of cell function by intracellular signaling
 Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
Mechanisms of Hormone Action
 Mechanisms of Hormone Action General principles: 1. Signals act over different ranges. 2. Signals have different chemical natures. 3. The same signal can induce a different response in different cells.
Mechanisms of Hormone Action General principles: 1. Signals act over different ranges. 2. Signals have different chemical natures. 3. The same signal can induce a different response in different cells.
Chem Lecture 10 Signal Transduction
 Chem 452 - Lecture 10 Signal Transduction 111130 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
Chem 452 - Lecture 10 Signal Transduction 111130 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
Membrane associated receptor transfers the information. Second messengers relay information
 Membrane associated receptor transfers the information Most signals are polar and large Few of the signals are nonpolar Receptors are intrinsic membrane proteins Extracellular and intracellular domains
Membrane associated receptor transfers the information Most signals are polar and large Few of the signals are nonpolar Receptors are intrinsic membrane proteins Extracellular and intracellular domains
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
Introduction! Introduction! Introduction! Chem Lecture 10 Signal Transduction & Sensory Systems Part 2
 Chem 452 - Lecture 10 Signal Transduction & Sensory Systems Part 2 Questions of the Day: How does the hormone insulin trigger the uptake of glucose in the cells that it targets. Introduction! Signal transduction
Chem 452 - Lecture 10 Signal Transduction & Sensory Systems Part 2 Questions of the Day: How does the hormone insulin trigger the uptake of glucose in the cells that it targets. Introduction! Signal transduction
PHSI3009 Frontiers in Cellular Physiology 2017
 Overview of PHSI3009 L2 Cell membrane and Principles of cell communication L3 Signalling via G protein-coupled receptor L4 Calcium Signalling L5 Signalling via Growth Factors L6 Signalling via small G-protein
Overview of PHSI3009 L2 Cell membrane and Principles of cell communication L3 Signalling via G protein-coupled receptor L4 Calcium Signalling L5 Signalling via Growth Factors L6 Signalling via small G-protein
Biosignals, Chapter 8, rearranged, Part I
 Biosignals, Chapter 8, rearranged, Part I Nicotinic Acetylcholine Receptor: A Ligand-Binding Ion Channel Classes of Receptor Proteins in Eukaryotes, Heterotrimeric G Proteins Signaling View the Heterotrimeric
Biosignals, Chapter 8, rearranged, Part I Nicotinic Acetylcholine Receptor: A Ligand-Binding Ion Channel Classes of Receptor Proteins in Eukaryotes, Heterotrimeric G Proteins Signaling View the Heterotrimeric
ZSTK474 is an ATP-competitive inhibitor of class I. phosphatidylinositol 3 kinase isoforms
 ZSTK474 is an ATP-competitive inhibitor of class I Blackwell Publishing Asia phosphatidylinositol 3 kinase isoforms Dexin Kong and Takao Yamori 1 Division of Molecular Pharmacology, Cancer Chemotherapy
ZSTK474 is an ATP-competitive inhibitor of class I Blackwell Publishing Asia phosphatidylinositol 3 kinase isoforms Dexin Kong and Takao Yamori 1 Division of Molecular Pharmacology, Cancer Chemotherapy
BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney
 BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney Page 2: Immune Mechanisms & Molecular Biology of Host Defence (Prof Campbell) Page 45: Infection and Implications for Cell
BCHM3972 Human Molecular Cell Biology (Advanced) 2013 Course University of Sydney Page 2: Immune Mechanisms & Molecular Biology of Host Defence (Prof Campbell) Page 45: Infection and Implications for Cell
Biol403 MAP kinase signalling
 Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
PI3K Background. The SignalRx R & D pipeline is shown below followed by a brief description of each program:
 PI3K Background The phosphatidylinositol 3-kinase (PI3K) pathway is a key cell signaling node whose dysregulation commonly results in the transformation of normal cells into cancer cells. The role of PI3K
PI3K Background The phosphatidylinositol 3-kinase (PI3K) pathway is a key cell signaling node whose dysregulation commonly results in the transformation of normal cells into cancer cells. The role of PI3K
1. Activated receptor tyrosine kinases (RTKs) phosphorylates themselves
 Enzyme-coupled receptors Transmembrane proteins Ligand-binding domain on the outer surface Cytoplasmic domain acts as an enzyme itself or forms a complex with enzyme 1. Activated receptor tyrosine kinases
Enzyme-coupled receptors Transmembrane proteins Ligand-binding domain on the outer surface Cytoplasmic domain acts as an enzyme itself or forms a complex with enzyme 1. Activated receptor tyrosine kinases
Neurotransmitter Systems II Receptors. Reading: BCP Chapter 6
 Neurotransmitter Systems II Receptors Reading: BCP Chapter 6 Neurotransmitter Systems Normal function of the human brain requires an orderly set of chemical reactions. Some of the most important chemical
Neurotransmitter Systems II Receptors Reading: BCP Chapter 6 Neurotransmitter Systems Normal function of the human brain requires an orderly set of chemical reactions. Some of the most important chemical
Signal-Transduction Cascades - 2. The Phosphoinositide Cascade
 Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
LQB383 Testbank. Week 8 Cell Communication and Signaling Mechanisms
 LQB383 Testbank Week 8 Cell Communication and Signaling Mechanisms Terms to learn match the terms to the definitions --------------------------------------------------------------------------------------------------------------------------
LQB383 Testbank Week 8 Cell Communication and Signaling Mechanisms Terms to learn match the terms to the definitions --------------------------------------------------------------------------------------------------------------------------
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
A dual PI3 kinase/mtor inhibitor reveals emergent efficacy in glioma
 Supplemental data A dual PI3 kinase/mtor inhibitor reveals emergent efficacy in glioma Qi-Wen Fan, Zachary A. Knight, David D. Goldenberg, Wei Yu, Keith E. Mostov, David Stokoe, Kevan M. Shokat, and William
Supplemental data A dual PI3 kinase/mtor inhibitor reveals emergent efficacy in glioma Qi-Wen Fan, Zachary A. Knight, David D. Goldenberg, Wei Yu, Keith E. Mostov, David Stokoe, Kevan M. Shokat, and William
Lecture 34. Carbohydrate Metabolism 2. Glycogen. Key Concepts. Biochemistry and regulation of glycogen degradation
 Lecture 34 Carbohydrate Metabolism 2 Glycogen Key Concepts Overview of Glycogen Metabolism Biochemistry and regulation of glycogen degradation Biochemistry and regulation of glycogen synthesis What mechanisms
Lecture 34 Carbohydrate Metabolism 2 Glycogen Key Concepts Overview of Glycogen Metabolism Biochemistry and regulation of glycogen degradation Biochemistry and regulation of glycogen synthesis What mechanisms
Enzymes: The Catalysts of Life
 Chapter 6 Enzymes: The Catalysts of Life Lectures by Kathleen Fitzpatrick Simon Fraser University Activation Energy and the Metastable State Many thermodynamically feasible reactions in a cell that could
Chapter 6 Enzymes: The Catalysts of Life Lectures by Kathleen Fitzpatrick Simon Fraser University Activation Energy and the Metastable State Many thermodynamically feasible reactions in a cell that could
Statin inhibition of HMG-CoA reductase: a 3-dimensional view
 Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Chapter 10. Regulatory Strategy
 Chapter 10 Regulatory Strategy Regulation of enzymatic activity: 1. Allosteric Control. Allosteric proteins have a regulatory site(s) and multiple functional sites Activity of proteins is regulated by
Chapter 10 Regulatory Strategy Regulation of enzymatic activity: 1. Allosteric Control. Allosteric proteins have a regulatory site(s) and multiple functional sites Activity of proteins is regulated by
Protein kinases are enzymes that add a phosphate group to proteins according to the. ATP + protein OH > Protein OPO 3 + ADP
 Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Protein kinase Protein kinases are enzymes that add a phosphate group to proteins according to the following equation: 2 ATP + protein OH > Protein OPO 3 + ADP ATP represents adenosine trisphosphate, ADP
Lecture 15. Signal Transduction Pathways - Introduction
 Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Chapter 20. Cell - Cell Signaling: Hormones and Receptors. Three general types of extracellular signaling. endocrine signaling. paracrine signaling
 Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
Identification of Oral Bioavailable, Type2 Inhibitors of Discoidin Domain-containing Receptor 1/2 (DDR1/DDR2) using Back-to-Front X-Ray FBDD
 Identification of ral Bioavailable, Type2 Inhibitors of Discoidin Domain-containing Receptor 1/2 (DDR1/DDR2) using Back-to-Front X-Ray FBDD Emiliano Tamanini 26 th Symposium on Medicinal Chemistry in Eastern
Identification of ral Bioavailable, Type2 Inhibitors of Discoidin Domain-containing Receptor 1/2 (DDR1/DDR2) using Back-to-Front X-Ray FBDD Emiliano Tamanini 26 th Symposium on Medicinal Chemistry in Eastern
Life History of A Drug
 DRUG ACTION & PHARMACODYNAMIC M. Imad Damaj, Ph.D. Associate Professor Pharmacology and Toxicology Smith 652B, 828-1676, mdamaj@hsc.vcu.edu Life History of A Drug Non-Specific Mechanims Drug-Receptor Interaction
DRUG ACTION & PHARMACODYNAMIC M. Imad Damaj, Ph.D. Associate Professor Pharmacology and Toxicology Smith 652B, 828-1676, mdamaj@hsc.vcu.edu Life History of A Drug Non-Specific Mechanims Drug-Receptor Interaction
Lecture 36: Review of membrane function
 Chem*3560 Lecture 36: Review of membrane function Membrane: Lipid bilayer with embedded or associated proteins. Bilayers: 40-70% neutral phospholipid 10-20% negative phospholipid 10-30% cholesterol 10-30%
Chem*3560 Lecture 36: Review of membrane function Membrane: Lipid bilayer with embedded or associated proteins. Bilayers: 40-70% neutral phospholipid 10-20% negative phospholipid 10-30% cholesterol 10-30%
WHY TARGETTING SIGNALLING PATHWAYS?
 WHY TARGETTING SIGNALLING PATHWAYS? Cancer cells are particularly sensitive to stress therefore sensitive to inhibition of their hyper activated signaling proteins the re instatement of lost tumor suppressors.
WHY TARGETTING SIGNALLING PATHWAYS? Cancer cells are particularly sensitive to stress therefore sensitive to inhibition of their hyper activated signaling proteins the re instatement of lost tumor suppressors.
PI3K. 1
 PI3K PI3K (Phosphatidylinositol-4,5-bisphosphate 3-kinase) are a family of enzymes involved in cellular functions such as cell growth, proliferation, differentiation, motility, survival and intracellular
PI3K PI3K (Phosphatidylinositol-4,5-bisphosphate 3-kinase) are a family of enzymes involved in cellular functions such as cell growth, proliferation, differentiation, motility, survival and intracellular
Cellular Physiology (PHSI3009) Contents:
 Cellular Physiology (PHSI3009) Contents: Cell membranes and communication 2 nd messenger systems G-coupled protein signalling Calcium signalling Small G-protein signalling o RAS o MAPK o PI3K RHO GTPases
Cellular Physiology (PHSI3009) Contents: Cell membranes and communication 2 nd messenger systems G-coupled protein signalling Calcium signalling Small G-protein signalling o RAS o MAPK o PI3K RHO GTPases
Chapter 11. Cell Communication
 Chapter 11 Cell Communication Overview: The Cellular Internet Cell-to-cell communication Is absolutely essential for multicellular organisms Concept 11.1: External signals are converted into responses
Chapter 11 Cell Communication Overview: The Cellular Internet Cell-to-cell communication Is absolutely essential for multicellular organisms Concept 11.1: External signals are converted into responses
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
Signal Transduction I
 Signal Transduction I Prof. Tianhua Zhou Department of Cell Biology Zhejiang University School of Medicine Office hours by appointment tzhou@zju.edu.cn Signal transduction: Key contents for signal transduction:
Signal Transduction I Prof. Tianhua Zhou Department of Cell Biology Zhejiang University School of Medicine Office hours by appointment tzhou@zju.edu.cn Signal transduction: Key contents for signal transduction:
Signal Transduction SS Gerhild van Echten-Deckert
 Signal Transduction SS 2018 Gerhild van Echten-Deckert Tel. 73 2703 E-mail: g.echten.deckert@uni-bonn.de https://www.limes-institut-bonn.de/forschung/ Focus on 2 classes of cell-surface receptors (Growth
Signal Transduction SS 2018 Gerhild van Echten-Deckert Tel. 73 2703 E-mail: g.echten.deckert@uni-bonn.de https://www.limes-institut-bonn.de/forschung/ Focus on 2 classes of cell-surface receptors (Growth
Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression
 Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression Question No. 1 of 10 1. Which statement about cell signaling is correct? Question #1 (A) Cell signaling involves receiving
Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression Question No. 1 of 10 1. Which statement about cell signaling is correct? Question #1 (A) Cell signaling involves receiving
Previous Class. Today. Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism. Protein Kinase Catalytic Properties
 Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis
 Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis MUDr. Jiří Vachtenheim, CSc. CELL CYCLE - SUMMARY Basic terminology: Cyclins conserved proteins with homologous regions; their cellular
Cell cycle, signaling to cell cycle, and molecular basis of oncogenesis MUDr. Jiří Vachtenheim, CSc. CELL CYCLE - SUMMARY Basic terminology: Cyclins conserved proteins with homologous regions; their cellular
Enzymes: Regulation 2-3
 Enzymes: Regulation 2-3 Reversible covalent modification Association with regulatory proteins Irreversible covalent modification/proteolytic cleavage Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter
Enzymes: Regulation 2-3 Reversible covalent modification Association with regulatory proteins Irreversible covalent modification/proteolytic cleavage Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia
 Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Chapter 11: Enzyme Catalysis
 Chapter 11: Enzyme Catalysis Matching A) high B) deprotonated C) protonated D) least resistance E) motion F) rate-determining G) leaving group H) short peptides I) amino acid J) low K) coenzymes L) concerted
Chapter 11: Enzyme Catalysis Matching A) high B) deprotonated C) protonated D) least resistance E) motion F) rate-determining G) leaving group H) short peptides I) amino acid J) low K) coenzymes L) concerted
Cell Communication. Chapter 11. PowerPoint Lectures for Biology, Seventh Edition. Lectures by Chris Romero. Neil Campbell and Jane Reece
 Chapter 11 Cell Communication PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: The Cellular Internet Cell-to-cell communication Is absolutely
Chapter 11 Cell Communication PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: The Cellular Internet Cell-to-cell communication Is absolutely
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Fundamentals of Pharmacology
 Fundamentals of Pharmacology Topic Page Receptors 2 Ion channels / GABA 4 GPCR s 6 TK receptors 8 Basics of PK 11 ADR s / Clinical study design 13 Introduction to the ANS 16 Cholinergic Pharmacology 20
Fundamentals of Pharmacology Topic Page Receptors 2 Ion channels / GABA 4 GPCR s 6 TK receptors 8 Basics of PK 11 ADR s / Clinical study design 13 Introduction to the ANS 16 Cholinergic Pharmacology 20
2013 W. H. Freeman and Company. 12 Signal Transduction
 2013 W. H. Freeman and Company 12 Signal Transduction CHAPTER 12 Signal Transduction Key topics: General features of signal transduction Structure and function of G protein coupled receptors Structure
2013 W. H. Freeman and Company 12 Signal Transduction CHAPTER 12 Signal Transduction Key topics: General features of signal transduction Structure and function of G protein coupled receptors Structure
The Tissue Engineer s Toolkit
 The Tissue Engineer s Toolkit Stimuli Detection and Response Ken Webb, Ph. D. Assistant Professor Dept. of Bioengineering Clemson University Environmental Stimulus-Cellular Response Environmental Stimuli
The Tissue Engineer s Toolkit Stimuli Detection and Response Ken Webb, Ph. D. Assistant Professor Dept. of Bioengineering Clemson University Environmental Stimulus-Cellular Response Environmental Stimuli
Plasma membranes. Plasmodesmata between plant cells. Gap junctions between animal cells Cell junctions. Cell-cell recognition
 Cell Communication Cell Signaling Cell-to-cell communication is essential for multicellular organisms Communicate by chemical messengers Animal and plant cells have cell junctions that directly connect
Cell Communication Cell Signaling Cell-to-cell communication is essential for multicellular organisms Communicate by chemical messengers Animal and plant cells have cell junctions that directly connect
Carbohydrate Metabolism 2 Supplemental Reading
 Carbohydrate Metabolism 2 Supplemental Reading Key Concepts - Overview of glycogen metabolism - Biochemistry and regulation glycogen degradation - Biochemistry and regulation of glycogen synthesis - Control
Carbohydrate Metabolism 2 Supplemental Reading Key Concepts - Overview of glycogen metabolism - Biochemistry and regulation glycogen degradation - Biochemistry and regulation of glycogen synthesis - Control
A Homogeneous Phosphoinositide 3-Kinase Assay on Phospholipid FlashPlate Platforms. Busi Maswoswe, Hao Xie, Pat Kasila and Li-an Yeh
 A Homogeneous Phosphoinositide 3-Kinase Assay on Phospholipid FlashPlate Platforms Busi Maswoswe, Hao Xie, Pat Kasila and Li-an Yeh Abstract Phosphoinositide 3-kinases (PI 3-kinase) consist of a family
A Homogeneous Phosphoinositide 3-Kinase Assay on Phospholipid FlashPlate Platforms Busi Maswoswe, Hao Xie, Pat Kasila and Li-an Yeh Abstract Phosphoinositide 3-kinases (PI 3-kinase) consist of a family
Defects in glucose utilization & GLP-1
 Defects in glucose utilization & GLP-1 Song, Dae-Kyu Department of Physiology & Chronic Disease Research Center Keimyung University School of Medicine 2800 dalgubeol-daero, Dalseo-gu, Daegu, 704-701, Korea
Defects in glucose utilization & GLP-1 Song, Dae-Kyu Department of Physiology & Chronic Disease Research Center Keimyung University School of Medicine 2800 dalgubeol-daero, Dalseo-gu, Daegu, 704-701, Korea
Principles of cell signaling Lecture 4
 Principles of cell signaling Lecture 4 Johan Lennartsson Molecular Cell Biology (1BG320), 2014 Johan.Lennartsson@licr.uu.se 1 Receptor tyrosine kinase-induced signal transduction Erk MAP kinase pathway
Principles of cell signaling Lecture 4 Johan Lennartsson Molecular Cell Biology (1BG320), 2014 Johan.Lennartsson@licr.uu.se 1 Receptor tyrosine kinase-induced signal transduction Erk MAP kinase pathway
6.5 Enzymes. Enzyme Active Site and Substrate Specificity
 180 Chapter 6 Metabolism 6.5 Enzymes By the end of this section, you will be able to: Describe the role of enzymes in metabolic pathways Explain how enzymes function as molecular catalysts Discuss enzyme
180 Chapter 6 Metabolism 6.5 Enzymes By the end of this section, you will be able to: Describe the role of enzymes in metabolic pathways Explain how enzymes function as molecular catalysts Discuss enzyme
Phospho-AKT Sampler Kit
 Phospho-AKT Sampler Kit E 0 5 1 0 0 3 Kits Includes Cat. Quantity Application Reactivity Source Akt (Ab-473) Antibody E021054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit Akt (Phospho-Ser473) Antibody
Phospho-AKT Sampler Kit E 0 5 1 0 0 3 Kits Includes Cat. Quantity Application Reactivity Source Akt (Ab-473) Antibody E021054-1 50μg/50μl IHC, WB Human, Mouse, Rat Rabbit Akt (Phospho-Ser473) Antibody
PHRM20001 NOTES PART 1 Lecture 1 History of Pharmacology- Key Principles
 PHRM20001 NOTES PART 1 Lecture 1 History of Pharmacology- Key Principles Hippocrates (5 th century BCE):... benefit my patients according to my greatest ability and judgment, and I will do no harm or injustice
PHRM20001 NOTES PART 1 Lecture 1 History of Pharmacology- Key Principles Hippocrates (5 th century BCE):... benefit my patients according to my greatest ability and judgment, and I will do no harm or injustice
Lecture Series 2 Macromolecules: Their Structure and Function
 Lecture Series 2 Macromolecules: Their Structure and Function Reading Assignments Read Chapter 4 (Protein structure & Function) Biological Substances found in Living Tissues The big four in terms of macromolecules
Lecture Series 2 Macromolecules: Their Structure and Function Reading Assignments Read Chapter 4 (Protein structure & Function) Biological Substances found in Living Tissues The big four in terms of macromolecules
Chemical Mechanism of Enzymes
 Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
Lecture Series 2 Macromolecules: Their Structure and Function
 Lecture Series 2 Macromolecules: Their Structure and Function Reading Assignments Read Chapter 4 (Protein structure & Function) Biological Substances found in Living Tissues The big four in terms of macromolecules
Lecture Series 2 Macromolecules: Their Structure and Function Reading Assignments Read Chapter 4 (Protein structure & Function) Biological Substances found in Living Tissues The big four in terms of macromolecules
A. Lipids: Water-Insoluble Molecules
 Biological Substances found in Living Tissues Lecture Series 3 Macromolecules: Their Structure and Function A. Lipids: Water-Insoluble Lipids can form large biological molecules, but these aggregations
Biological Substances found in Living Tissues Lecture Series 3 Macromolecules: Their Structure and Function A. Lipids: Water-Insoluble Lipids can form large biological molecules, but these aggregations
A Self-Propelled Biological Process Plk1-Dependent Product- Activated, Feed-Forward Mechanism
 A Self-Propelled Biological Process Plk1-Dependent Product- Activated, Feed-Forward Mechanism The Harvard community has made this article openly available. Please share how this access benefits you. Your
A Self-Propelled Biological Process Plk1-Dependent Product- Activated, Feed-Forward Mechanism The Harvard community has made this article openly available. Please share how this access benefits you. Your
PCB 3023 Exam 4 - Form A First and Last Name
 PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
PCB 3023 Exam 4 - Form A First and Last Name Student ID # (U Number) A Before beginning this exam, please complete the following instructions: 1) Write your name and U number on the first page of this
CHAPTER 10: REGULATORY STRATEGIES. Traffic signals control the flow of traffic
 CHAPTER 10: REGULATORY STRATEGIES Traffic signals control the flow of traffic INTRODUCTION CHAPTER 10 The activity of enzymes must often be regulated so that they function at the proper time and place.
CHAPTER 10: REGULATORY STRATEGIES Traffic signals control the flow of traffic INTRODUCTION CHAPTER 10 The activity of enzymes must often be regulated so that they function at the proper time and place.
Hormones and Signal Transduction. Dr. Kevin Ahern
 Dr. Kevin Ahern Signaling Outline Signaling Outline Background Signaling Outline Background Membranes Signaling Outline Background Membranes Hormones & Receptors Signaling Outline Background Membranes
Dr. Kevin Ahern Signaling Outline Signaling Outline Background Signaling Outline Background Membranes Signaling Outline Background Membranes Hormones & Receptors Signaling Outline Background Membranes
Signaling Through Immune System Receptors (Ch. 7)
 Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
Objectives. Carbon Bonding. Carbon Bonding, continued. Carbon Bonding
 Biochemistry Table of Contents Objectives Distinguish between organic and inorganic compounds. Explain the importance of carbon bonding in biological molecules. Identify functional groups in biological
Biochemistry Table of Contents Objectives Distinguish between organic and inorganic compounds. Explain the importance of carbon bonding in biological molecules. Identify functional groups in biological
Cell Communication. Cell Communication. Communication between cells requires: ligand: the signaling molecule
 Cell Communication Cell Communication Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the ligand binds (may be on the plasma membrane or within
Cell Communication Cell Communication Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the ligand binds (may be on the plasma membrane or within
Which DNA sequence is most likely to form a hairpin structure? x indicates any nucleotide.
 Which DNA sequence is most likely to form a hairpin structure? x indicates any nucleotide. A. xxxgtcagtxxxxtatgcgxxx B. xxxtcgtatxxxxgtccgaxxx C. xxxcactgtxxxxgtactgxxx D. xxxgtcagtxxxxcctagaxxx E. xxxgtcatcxxxxgatgacxxx
Which DNA sequence is most likely to form a hairpin structure? x indicates any nucleotide. A. xxxgtcagtxxxxtatgcgxxx B. xxxtcgtatxxxxgtccgaxxx C. xxxcactgtxxxxgtactgxxx D. xxxgtcagtxxxxcctagaxxx E. xxxgtcatcxxxxgatgacxxx
Sarah Jaar Marah Al-Darawsheh
 22 Sarah Jaar Marah Al-Darawsheh Faisal Mohammad Receptors can be membrane proteins (for water-soluble hormones/ligands) or intracellular (found in the cytosol or nucleus and bind to DNA, for lipid-soluble
22 Sarah Jaar Marah Al-Darawsheh Faisal Mohammad Receptors can be membrane proteins (for water-soluble hormones/ligands) or intracellular (found in the cytosol or nucleus and bind to DNA, for lipid-soluble
Lecture Series 2 Macromolecules: Their Structure and Function
 Lecture Series 2 Macromolecules: Their Structure and Function Reading Assignments Read Chapter 4 (Protein structure & Function) Biological Substances found in Living Tissues The big four in terms of macromolecules
Lecture Series 2 Macromolecules: Their Structure and Function Reading Assignments Read Chapter 4 (Protein structure & Function) Biological Substances found in Living Tissues The big four in terms of macromolecules
Lecture #27 Lecturer A. N. Koval
 Lecture #27 Lecturer A. N. Koval Hormones Transduce Signals to Affect Homeostatic Mechanisms Koval A. (C), 2011 2 Lipophilic hormones Classifying hormones into hydrophilic and lipophilic molecules indicates
Lecture #27 Lecturer A. N. Koval Hormones Transduce Signals to Affect Homeostatic Mechanisms Koval A. (C), 2011 2 Lipophilic hormones Classifying hormones into hydrophilic and lipophilic molecules indicates
NIH Public Access Author Manuscript Nat Chem Biol. Author manuscript; available in PMC 2010 June 3.
 NIH Public Access Author Manuscript Published in final edited form as: Nat Chem Biol. 2008 November ; 4(11): 691 699. doi:10.1038/nchembio.117. Targeted polypharmacology: Discovery of dual inhibitors of
NIH Public Access Author Manuscript Published in final edited form as: Nat Chem Biol. 2008 November ; 4(11): 691 699. doi:10.1038/nchembio.117. Targeted polypharmacology: Discovery of dual inhibitors of
Easy50 PI3K Inhibitor Array*
 Easy50 PI3K Inhibitor Array* Catalog # IAP001 Instruction Manual For Research Use Only *Patent pending Introduction This Easy50 PI3K Inhibitor Array is a panel of serial dilutions of inhibitors that are
Easy50 PI3K Inhibitor Array* Catalog # IAP001 Instruction Manual For Research Use Only *Patent pending Introduction This Easy50 PI3K Inhibitor Array is a panel of serial dilutions of inhibitors that are
PRC2 crystal clear. Matthieu Schapira
 PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
PRC2 crystal clear Matthieu Schapira Epigenetic mechanisms control the combination of genes that are switched on and off in any given cell. In turn, this combination, called the transcriptional program,
